prompt
stringlengths 19
879k
| completion
stringlengths 3
53.8k
| api
stringlengths 8
59
|
|---|---|---|
import wave
import numpy as np
import matplotlib.pyplot as plt
def show_wave(s):
with wave.open(s, "rb") as f:
params = f.getparams()
nchannels, sampwidth, framerate, nframes = params[:4]
str_data = f.readframes(nframes)
wave_data =
|
np.frombuffer(str_data, dtype=np.short)
|
numpy.frombuffer
|
from __future__ import absolute_import
from __future__ import division
from __future__ import print_function
from __future__ import unicode_literals
import unittest
import numpy as np
import tensorflow as tf
from onnx_tf.backend import run_node
from onnx_tf.backend import supports_device
from onnx import helper
from onnx.onnx_pb2 import TensorProto
class TestNode(unittest.TestCase):
""" Tests for nodes
"""
def _get_rnd(self, shape, low=-1.0, high=1.0):
return np.random.uniform(low, high, np.prod(shape)) \
.reshape(shape) \
.astype(np.float32)
def _elu(self, x):
# f(x) = alpha * (exp(x) - 1.) for x < 0,
# f(x) = x for x >= 0
if x < 0.:
return np.expm1(x)
return x
def _leaky_relu(self, x, alpha):
# f(x) = alpha * x for x < 0,
# f(x) = x for x >= 0
if x < 0.:
return alpha * x
return x
def test_abs(self):
node_def = helper.make_node("Abs", ["X"], ["Y"])
x = self._get_rnd([1000])
output = run_node(node_def, [x])
np.testing.assert_almost_equal(output["Y"], np.abs(x))
def test_add(self):
node_def = helper.make_node("Add", ["X", "Y"], ["Z"], broadcast=1, axis=1)
x = self._get_rnd([5, 10, 5, 5])
y = self._get_rnd([10])
output = run_node(node_def, [x, y])
np.testing.assert_almost_equal(output["Z"], np.add(x, y.reshape([1, 10, 1, 1])))
# node_def = helper.make_node("Add", ["A", "B"], ["C"], broadcast=1)
# a = self._get_rnd([10, 10])
# b = self._get_rnd([10, 10])
# output = run_node(node_def, [a, b])
# np.testing.assert_almost_equal(output["C"], np.add(a, b))
# node_def = helper.make_node("Add", ["A", "B"], ["C"], broadcast=1)
# a = self._get_rnd([10, 10])
# b = self._get_rnd([10,])
# output = run_node(node_def, [a, b])
# np.testing.assert_almost_equal(output["C"], np.add(a, b))
def test_arg_max(self):
# TODO: need to fix this test
return
for axis in [0, 1]:
node_def = helper.make_node("ArgMax", ["data"], ["reduced"],
axis=axis,
keepdims=0)
data = self._get_rnd([10, 10])
output = run_node(node_def, [data])
np.testing.assert_almost_equal(output["reduced"],
np.argmax(data, axis=axis))
def test_arg_min(self):
# TODO: need to fix this test
return
for axis in [0, 1]:
node_def = helper.make_node("ArgMin", ["data"], ["reduced"],
axis=axis, keepdims=0)
data = self._get_rnd([10, 10])
output = run_node(node_def, [data])
np.testing.assert_almost_equal(output["reduced"],
|
np.argmin(data, axis=axis)
|
numpy.argmin
|
#!/usr/bin/env python3
# -*- coding: utf-8 -*-
"""Plotting functions."""
import logging
import logging.config
import math
import os
import warnings
from pathlib import Path
import cartopy.crs as ccrs
import iris
import matplotlib as mpl
import matplotlib.colors as colors
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
from matplotlib.colors import from_levels_and_colors
from tqdm import tqdm
from ..data import dummy_lat_lon_cube
from ..logging_config import LOGGING
from ..utils import (
CoordinateSystemError,
in_360_longitude_system,
multiline,
select_valid_subset,
translate_longitude_system,
)
logger = logging.getLogger(__name__)
class PlottingError(Exception):
"""Base class for exception in the plotting module."""
class MaskedDataError(PlottingError):
"""Raised when trying to plot fully-masked data."""
class FigureSaver:
"""Save figures using pre-defined options and directories.
If `debug`, `debug_options` will be used. Otherwise `options` are used.
Figure(s) are saved automatically:
with FigureSaver("filename"):
plt.figure()
...
with FigureSaver(("filename1", "filename2")):
plt.figure()
...
plt.figure()
...
Manual saving using the defined options is also possible.
with FigureSaver() as saver:
fig = plt.figure()
...
saver.save_figure(fig, "plot_name")
"""
debug = False
directory = "."
# These options serve as default value that may be overridden during
# initialisation.
options = {
"bbox_inches": "tight",
"transparent": True,
"filetype": "pdf",
"dpi": 600,
}
debug_options = {
"bbox_inches": "tight",
"transparent": False,
"filetype": "png",
"dpi": 350,
}
def __init__(self, filenames=None, *, directories=None, debug=None, **kwargs):
"""Initialise figure saver.
The initialised FigureSaver instance can be used repeatedly (as a context
manager) to save figures by calling it with at least filenames and optionally
directories, debug state, and saving options.
Args:
filenames ((iterable of) str or pathlib.Path, or None): If None, the
FigureSaver instance must be called with a list of filenames and used
as a context manager for automatic saving. Otherwise, the number of
strings or Paths given must match the number of figures opened within
the context.
directory ((iterable of) str or pathlib.Path, or None): The directory to
save figures in. If None, use `FigureSaver.directory`. New directories
will be created if they do not exist.
debug (bool or None): Select the pre-set settings with which figures will
be saved. If None, use `FigureSaver.debug`.
**kwargs: Optional kwargs which are passed to plt.savefig().
"""
# Backwards compatibility.
if "filename" in kwargs:
warnings.warn(
"The `filename` argument is deprecated in favour of the `filenames` "
"argument, which takes precedence.",
FutureWarning,
)
# Only use the deprecated argument if the updated version is not used.
if filenames is None:
filenames = kwargs.pop("filename")
if "directory" in kwargs:
warnings.warn(
"The `directory` argument is deprecated in favour of the `directories` "
"argument, which takes precedence.",
FutureWarning,
)
# Only use the deprecated argument if the updated version is not used.
if directories is None:
directories = kwargs.pop("directory")
# Set instance defaults.
if debug is not None:
self.debug = debug
directories = directories if directories is not None else self.directory
self.directories = (
(directories,) if isinstance(directories, (str, Path)) else directories
)
if filenames is not None:
self.filenames = (
(filenames,) if isinstance(filenames, (str, Path)) else filenames
)
if len(self.directories) != 1 and len(self.directories) != len(
self.filenames
):
raise ValueError(
multiline(
f"""If multiple directories are given, their number has to match
the number of file names, but got {len(self.directories)}
directories and {len(self.filenames)} file names."""
)
)
# Make sure to resolve the home directory.
self.directories = tuple(map(os.path.expanduser, self.directories))
self.options = self.debug_options.copy() if self.debug else self.options.copy()
self.options.update(kwargs)
def __call__(self, filenames=None, sub_directory=None, **kwargs):
"""Return a copy containing the given filenames for figure saving.
An optional sub-directory can also be specified for figures saved by the
returned object.
This is meant to be used as a context manager:
>>> figure_saver = FigureSaver(**options) # doctest: +SKIP
>>> with figure_saver("filename"): # doctest: +SKIP
... plt.figure() # doctest: +SKIP
Directories, options, etc... which the FigureSaver instance was initialised
with will be used to save the figures.
Args:
filenames ((iterable of) str): Filenames used to save created figures.
sub_directory (str): If given, figures will be saved in a sub-directory
`sub_directory` of the pre-specified directory/directories.
**kwargs: Optional kwargs which are passed to plt.savefig().
"""
new_inst = type(self)(
filenames,
directories=[os.path.join(orig, sub_directory) for orig in self.directories]
if sub_directory is not None
else self.directories,
debug=self.debug,
)
new_inst.options = {**self.options, **kwargs}
return new_inst
@property
def suffix(self):
return (
self.options["filetype"]
if "." in self.options["filetype"]
else "." + self.options["filetype"]
)
def __enter__(self):
self.old_fignums = plt.get_fignums()
return self
def __exit__(self, type, value, traceback):
if type is not None:
return False # Re-raise exception.
new_figure_numbers = plt.get_fignums()
if new_figure_numbers == self.old_fignums:
raise RuntimeError("No new figures detected.")
fignums_save = [
num for num in new_figure_numbers if num not in self.old_fignums
]
if len(fignums_save) != len(self.filenames):
raise RuntimeError(
f"Expected {len(self.filenames)} figures, but got {len(fignums_save)}."
)
saved_figures = [
num
if not plt.figure(num).get_label()
else (num, plt.figure(num).get_label())
for num in fignums_save
]
logger.debug(f"Saving figures {saved_figures}.")
if len(self.directories) == 1:
# Adapt to the number of figures.
directories = [self.directories[0]] * len(self.filenames)
else:
directories = self.directories
for fignum, directory, filename in zip(
fignums_save, directories, self.filenames
):
fig = plt.figure(fignum)
self.save_figure(fig, filename, directory)
def save_figure(self, fig, filename, directory=None, sub_directory=None, **kwargs):
"""Save a single figure.
Args:
fig (matplotlib.figure.Figure): Figure to save.
filename (str): Filename where the figure will be saved.
directory (str): Directory to save the figure in.
sub_directory (str): If given, figures will be saved in a sub-directory
`sub_directory` of the pre-specified directory/directories.
**kwargs: Optional kwargs which are passed to plt.savefig().
Raises:
ValueError: If multiple default directories were specified and no explicit
directory is supplied here. Since only one figure is being saved here,
it is unclear which directory to choose. In this case, the context
manager interface is to be used.
"""
if directory is None:
if len(self.directories) > 1:
raise ValueError("More than 1 default directory specified.")
# Use default.
directory = self.directories[0]
if sub_directory is not None:
directory = os.path.join(directory, sub_directory)
os.makedirs(directory, exist_ok=True)
if "." in filename:
filename = "".join(filename.split(".")[:-1])
filepath = (
os.path.expanduser(
os.path.abspath(os.path.expanduser(os.path.join(directory, filename)))
)
+ self.suffix
)
logger.debug("Saving figure to '{}'.".format(filepath))
fig.savefig(
filepath,
**{
**{
option: value
for option, value in self.options.items()
if option != "filetype"
},
**kwargs,
},
)
def get_cubes_vmin_vmax(cubes, vmin_vmax_percentiles=(0.0, 100.0)):
"""Get vmin and vmax from a list of cubes given two percentiles.
Args:
cubes (iris.cube.CubeList): List of cubes.
vmin_vmax_percentiles (tuple or None): The two percentiles, used to set the minimum
and maximum values on the colorbar. If `None`, use the minimum and maximum
of the data (equivalent to percentiles of (0, 100)).
Returns:
tuple: tuple of floats (vmin, vmax) if `vmin_vmax_percentiles` is not (0, 100) in which case
(None, None) will be returned.
"""
if vmin_vmax_percentiles is None or np.all(
np.isclose(np.array(vmin_vmax_percentiles), np.array([0, 100]))
):
return None, None
limits = []
for cube in cubes:
if isinstance(cube.data, np.ma.core.MaskedArray):
if isinstance(cube.data.mask, np.ndarray):
valid_data = cube.data.data[~cube.data.mask]
elif cube.data.mask:
raise ValueError("All data is masked.")
else:
valid_data = cube.data.data
else:
valid_data = cube.data
limits.append(np.percentile(valid_data, vmin_vmax_percentiles))
output = []
if np.isclose(vmin_vmax_percentiles[0], 0):
output.append(None)
else:
output.append(min(limit[0] for limit in limits))
if np.isclose(vmin_vmax_percentiles[1], 100):
output.append(None)
else:
output.append(max(limit[1] for limit in limits))
return output
def map_model_output(ba_predicted, ba_data, model_name, coast_linewidth):
"""Plotting of burned area data & predictions.
Args:
ba_predicted: predicted burned area
ba_data: observed
model_name (str): Name of the run.
Returns:
tuple: The created figures.
"""
figs = []
# Plotting params.
figsize = (5, 3.33)
mpl.rcParams["figure.figsize"] = figsize
vmin = min((np.min(ba_predicted), np.min(ba_data)))
vmax = max((np.max(ba_predicted), np.max(ba_data)))
boundaries = [1e-7, 1e-6, 1e-5, 1e-4, 1e-3, 1e-2, 1e-1]
# Plotting predicted.
fig = cube_plotting(
ba_predicted,
cmap="brewer_RdYlBu_11_r",
title=None,
log=True,
vmin=vmin,
vmax=vmax,
min_edge=vmin,
extend="min",
boundaries=boundaries,
)
figs.append(fig)
# Plotting observed.
fig = cube_plotting(
ba_data,
cmap="brewer_RdYlBu_11_r",
title=None,
log=True,
vmin=vmin,
vmax=vmax,
min_edge=vmin,
extend="min",
boundaries=boundaries,
colorbar_kwargs={"label": "Burnt Area Fraction"},
coastline_kwargs={"linewidth": coast_linewidth},
)
figs.append(fig)
# Plotting differences.
# https://blogs.sas.com/content/iml/2014/07/14/log-transformation-of-pos-neg.html
# (Do not!) use log-modulus transformation
perc_diffs = (ba_data - ba_predicted) / ba_data
diff_boundaries = [-1e3, -1e0, -1e-1, -1e-2, 0, 1e-1, 5e-1, 1e0]
fig = cube_plotting(
perc_diffs,
cmap="brewer_RdYlBu_11_r",
title=None,
log=True,
boundaries=diff_boundaries,
extend="min" if np.max(perc_diffs) <= max(diff_boundaries) else "both",
colorbar_kwargs={"label": "(Observed - Predicted) / Observed"},
coastline_kwargs={"linewidth": coast_linewidth},
)
figs.append(fig)
return figs
def _get_log_bin_edges(vmin, vmax, n_bins):
if isinstance(n_bins, (int, np.integer)):
return np.geomspace(vmin, vmax, n_bins + 1)
else:
edges = []
# "Auto" bins.
vmax_log = np.log10(vmax)
vmin_log = np.log10(vmin)
# Convert to float here explicitly as np.float32 does not have an
# `is_integer()` method, for example.
if not float(vmax_log).is_integer():
edges.append(vmax)
vmax_log = math.floor(vmax_log)
# To ensure it is an integer (round() does not turn a numpy float into an int).
vmax_log = int(round(vmax_log))
if not float(vmin_log).is_integer():
edges.append(vmin)
vmin_log = math.ceil(vmin_log)
# To ensure it is an integer (round() does not turn a numpy float into an int).
vmin_log = int(round(vmin_log))
edges.extend(np.logspace(vmin_log, vmax_log, vmax_log - vmin_log + 1))
return sorted(edges)
def get_bin_edges(
data=None,
vmin=None,
vmax=None,
n_bins=11,
log=True,
min_edge_type="manual",
min_edge=None,
simple_lin_bins=False,
):
"""Get bin edges.
Args:
data (numpy.ndarray or None): Data array to determine limits.
vmin (float or None): Minimum bin edge.
vmax (float or None): Maximum bin edge.
n_bins (int or str): If "auto" (only applies when `log=True`), determine bin
edges using the data (see `min_edge`).
log (bool): If `log`, bin edges are computed in log space (base 10).
min_edge_type (str): If "manual", the supplied `min_edge` will be used for the
minimum edge(s), and `vmin` and `vmax` for the upper limits. If "auto",
determine `min_edge` from the data. If "symmetric", determine `min_edge`
from the data, but use the same value for the positive and negative edges.
If either `vmin` or `vmax` is very close to (or equal to) 0, `min_edge` is
also required as the starting point of the bin edges (see examples).
min_edge (float, iterable of float, or None): If None, the minimum (absolute)
data value will be used. Otherwise the supplied float will be used to set
the minimum exponent of the log bins. If two floats are given, they will
be used for the positive and negative ranges, respectively.
simple_lin_bins (bool): If True, simply create `n_bins` divisions from vmin
(minimum data) to vmax (maximum data). If False (default), explicitly
structure the bins around 0 if needed. If there is only positive or only
negative data, the two cases are equivalent.
Returns:
list: The bin edges.
Examples:
>>> get_bin_edges(vmin=0, vmax=100, n_bins=2, log=False)
[0.0, 50.0, 100.0]
>>> get_bin_edges(vmin=1, vmax=100, n_bins=2, log=True, min_edge=1)
[1.0, 10.0, 100.0]
>>> get_bin_edges(vmin=-100, vmax=100, n_bins="auto", log=True, min_edge=1,
... min_edge_type="manual")
[-100.0, -10.0, -1.0, 0.0, 1.0, 10.0, 100.0]
>>> get_bin_edges(vmin=-150, vmax=150, n_bins="auto", log=True, min_edge=1,
... min_edge_type="manual")
[-150.0, -100.0, -10.0, -1.0, 0.0, 1.0, 10.0, 100.0, 150.0]
>>> get_bin_edges(vmin=-1000, vmax=100, n_bins=7, log=True, min_edge=1,
... min_edge_type="manual")
[-1000.0, -100.0, -10.0, -1.0, 0.0, 1.0, 10.0, 100.0]
>>> get_bin_edges(vmin=-100, vmax=1000, n_bins="auto", log=True, min_edge=1,
... min_edge_type="manual")
[-100.0, -10.0, -1.0, 0.0, 1.0, 10.0, 100.0, 1000.0]
>>> get_bin_edges(vmin=0, vmax=100, n_bins=3, log=True, min_edge=1)
[0.0, 1.0, 10.0, 100.0]
>>> get_bin_edges(np.array([0, 1000]), n_bins=4, min_edge=1.0, log=True)
[0.0, 1.0, 10.0, 100.0, 1000.0]
>>> get_bin_edges(np.array([0, 1, 1000]), n_bins=4, min_edge_type="auto", log=True)
[0.0, 1.0, 10.0, 100.0, 1000.0]
>>> get_bin_edges(np.array([0, 10, 100]), n_bins=2, min_edge_type="auto", log=True)
[0.0, 10.0, 100.0]
>>> get_bin_edges(vmin=-20, vmax=80, n_bins=5, log=False,
... simple_lin_bins=True)
[-20.0, 0.0, 20.0, 40.0, 60.0, 80.0]
>>> get_bin_edges(vmin=-20, vmax=80, n_bins=5, log=False,
... simple_lin_bins=False)
[-20.0, 0.0, 20.0, 40.0, 60.0, 80.0]
>>> np.all(
... np.isclose(
... get_bin_edges(
... vmin=-20,
... vmax=77,
... n_bins=9,
... log=False,
... simple_lin_bins=True,
... ),
... np.linspace(-20, 77, 10),
... )
... )
True
>>> get_bin_edges(vmin=-20, vmax=77, n_bins=9, log=False,
... simple_lin_bins=False)
[-20.0, -10.0, 0.0, 11.0, 22.0, 33.0, 44.0, 55.0, 66.0, 77.0]
"""
if not log:
assert isinstance(
n_bins, (int, np.integer)
), f"Bin number must be an integer if `log=False`. Got {repr(n_bins)} instead."
if any(vlim is None for vlim in (vmin, vmax)) and data is None:
raise ValueError("Need data when vmin & vmax are not supplied (are None).")
if all(vlim is not None for vlim in (vmin, vmax)) and vmin > vmax:
raise ValueError(f"vmin ({vmin}) must not be larger than vmax ({vmax}).")
if log and all(vlim is not None for vlim in (vmin, vmax)) and data is None:
assert (
min_edge is not None
), "When not supplying data, `min_edge` needs to be given."
assert (
min_edge_type == "manual"
), "When not supplying data, `min_edge_type` needs to be 'manual'."
vmin = vmin if vmin is not None else np.min(data)
vmax = vmax if vmax is not None else np.max(data)
n_close_lim = sum(np.isclose(vlim, 0) for vlim in (vmin, vmax))
assert n_close_lim <= 1, "At most one limit should be close to 0."
if not log:
if simple_lin_bins or not (vmin < 0 and vmax > 0):
# Handle simple cases where the bin edges do not cross 0.
return list(np.linspace(vmin, vmax, n_bins + 1))
else:
vrange = vmax - vmin
pos_edges = math.ceil((n_bins + 1) * np.abs(vmin) / vrange)
neg_edges = math.ceil((n_bins + 1) * vmax / vrange)
# The desired number of edges is bins + 2, since we are removing one edge
# (the duplicated 0) and thus we are left with bins + 1, the number of
# edges required to form 'bins' number of bins as desired.
if pos_edges + neg_edges != n_bins + 2:
assert pos_edges + neg_edges == n_bins + 1, (
"Expecting at most 1 missing edge. Got "
f"{pos_edges + neg_edges}, expected {n_bins + 1} or {n_bins + 2}."
)
# Determine which side to add an edge to.
ideal_pos_ratio = vmax / vrange
if pos_edges / (pos_edges + neg_edges) >= ideal_pos_ratio:
# We have too many positive edges already, so increment the number
# of negative edges.
neg_edges += 1
else:
pos_edges += 1
return list(np.linspace(vmin, 0, pos_edges)) + list(
np.linspace(0, vmax, neg_edges)[1:]
)
# Only the log case remains here.
if min_edge_type not in ("symmetric", "auto", "manual"):
raise ValueError(f"Unexpected `min_edge_type` {min_edge_type}.")
if min_edge_type == "manual":
assert min_edge is not None, "Need valid `min_edge` for 'manual' edge type."
if min_edge is not None:
if isinstance(min_edge, (float, int, np.float, np.integer)):
min_edge = [min_edge, min_edge]
min_edge = np.abs(np.asarray(min_edge, dtype=np.float64))
# Handle the positive and negative data ranges separately.
# Positive and negative data are handled the same way (after taking the absolute
# value) and the resulting bins are then stitched together if needed.
if data is not None:
zero_mask = ~np.isclose(data, 0)
split_data = (
data[(data > 0) & zero_mask],
np.abs(data[(data < 0) & zero_mask]),
)
else:
split_data = (None, None)
if vmin >= 0:
split_data = split_data[:1]
elif vmax <= 0:
split_data = split_data[1:]
if min_edge is not None:
if min_edge_type != "manual":
raise ValueError(
"Value for `min_edge` supplied even though `min_edge_type` was set to "
f"'{min_edge_type}'."
)
if min_edge_type in ("symmetric", "auto"):
# This guarantees that data is not None, meaning that we can use the zero
# mask.
if min_edge_type == "symmetric":
# Use the same minimum edge for both data ranges if needed.
min_edge_type = "manual"
min_edge = [np.min(np.abs(data[zero_mask]))] * 2
else:
# If edge type is auto, data is needed to infer the bounds.
if not all(map(np.any, split_data)):
raise ValueError(
f"Insufficient data for `min_edge_type` '{min_edge_type}'."
)
# Use the data to infer the minimum edge separately for each data range.
min_edge = list(map(np.min, split_data))
# List that holds the output bin edges.
bin_edges = []
if vmin >= 0:
multipliers = (1,)
limits = [[vmin, vmax]]
if np.isclose(limits[0][0], 0):
bin_edges.append(limits[0][0])
if isinstance(n_bins, (int, np.integer)):
n_bins -= 1 # Compensate for the additional edge above.
limits[0][0] = min_edge[0]
elif vmax <= 0:
multipliers = (-1,)
limits = [[np.abs(vmax), np.abs(vmin)]]
if np.isclose(limits[0][0], 0):
bin_edges.append(limits[0][0])
if isinstance(n_bins, (int, np.integer)):
n_bins -= 1 # Compensate for the additional edge above.
limits[0][0] = min_edge[1]
else:
multipliers = (1, -1)
limits = [(min_edge[0], vmax), (min_edge[1], np.abs(vmin))]
if vmin >= 0 or vmax <= 0:
# Only a single bin number is required.
bins = [n_bins]
else:
if isinstance(n_bins, (int, np.integer)):
contributions = np.array(
[np.ptp(np.log10(s_limits)) for s_limits in limits]
)
total = np.sum(contributions)
# To arrive at `n_bins` bins (ie. `n_bins + 1` edges), we need to consider
# the number of bins for the -ve and +ve part, and the 0-edge in the
# middle. Concatenating the two ranges (which do not include 0 as opposed
# to the linear case above) adds 1 bin, while adding the 0-edge adds
# another bin. Thus, `n_bins - 2` bins need to be provided by the two
# parts.
bins = [math.ceil(n_bin) for n_bin in (n_bins - 2) * contributions / total]
# If we have 1 bin too few (remember that we want `n_bins - 2` bins at
# this stage.
if sum(bins) < n_bins - 2:
ideal_pos_ratio = contributions[0] / total
if bins[0] / np.sum(bins) >= ideal_pos_ratio:
# We have too many positive bins already, so increment the number
# of negative bins.
bins[0] += 1
else:
bins[1] += 1
else:
# Ie. use "auto" bins for both ranges.
bins = [n_bins, n_bins]
if n_close_lim == 1:
assert (
len(split_data) == 1
), "There should be only 1 split dataset if one of the limits is (close to) 0."
# Now that the ranges have been segregated, handle them each individually and
# combine the resulting bin edges.
# Get the relevant limits for the data in both the positive and negative case.
# Do this by checking the sign and magnitude of the predefined limits, and
# using max/min of the selected data if necessary.
for multiplier, s_limits, s_bins in zip(multipliers, limits, bins):
bin_edges.extend(multiplier * np.asarray(_get_log_bin_edges(*s_limits, s_bins)))
if len(split_data) == 2:
bin_edges.append(0)
return sorted(float(edge) for edge in bin_edges)
def cube_plotting(
cube,
log=False,
dummy_lat_lims=(-90, 90),
dummy_lon_lims=(-180, 180),
vmin=None,
vmax=None,
vmin_vmax_percentiles=(0, 100),
nbins=10,
log_auto_bins=True,
boundaries=None,
min_edge=None,
extend=None,
projection=None,
cmap="viridis",
cmap_midpoint=None,
cmap_symmetric=False,
fig=None,
ax=None,
title=None,
average_first_coord=True,
select_valid=False,
transform_vmin_vmax=False,
animation_output=False,
mesh=None,
title_text=None,
return_cbar=False,
colorbar_kwargs=None,
coastline_kwargs=None,
gridline_kwargs=None,
**kwargs,
):
"""Plotting of cubes.
Eg. for temperature, use
cmap='Reds'
colorbar_kwargs{"label": r"T ($\degree$C)"}
Args:
cube: Cube to plot.
log: True to log.
dummy_lat_lims: Tuple passed to dummy_lat_lon_cube function in case the input
argument is not a cube.
dummy_lon_lims: Tuple passed to dummy_lat_lon_cube function in case the input
argument is not a cube.
vmin: Minimum value for colorbar.
vmax: Maximum value for colorbar.
vmin_vmax_percentiles (tuple or None): The two percentiles used to set the
minimum and maximum values on the colorbar. If `None`, use the minimum and
maximum of the data (equivalent to percentiles of (0, 100)). `vmin` and
`vmax` parameters take precedence.
nbins (int): Number of bins. Does not apply if `log` and `log_auto_bins`.
log_auto_bins (bool): Make log bins stick to integers.
boundaries (iterable or None): If None, bin boundaries will be computed
automatically. If given, this supersedes all other options relating to
boundary creation, like `log' or `vmin'.
min_edge (float or None): Minimum log bin exponent. See `get_bin_edges`.
extend (None or {"neither", "min", "max"}): The colormap extension. If `None`,
`extend` is determined based on `boundaries` (which may be automatically
determined from other arguments if not given) and `cube`.
projection: A projection as defined in `cartopy.crs`. If None (default),
`cartopy.crs.Robinson()` will be used, where the central longitude will be
defined as the average of the cube longitudes (see `select_valid`).
cmap (matplotlib Colormap or str): Colormap to use, e.g. 'Reds',
'Reds_r', etc. 'viridis' is used by default.
cmap_midpoint (float or None): The value corresponding to the middle of the
colormap range will be the value in `boundaries` closest to
`cmap_midpoint`.
cmap_symmetric (bool): If True, the coverage of the colormap relative to the
`cmap_midpoint` will depend on `boundaries`. Only applies if
`cmap_midpoint` is not `None`.
fig (matplotlib Figure): Figure to plot onto. If `None`, a new Figure will be
created. If `fig` is `None` but `ax` is not None, the Figure `ax` belongs
to will be used.
ax (matplotlib Axes): Axis to plot onto. If None, a new axis with `projection`
will be created using the current Figure (see `fig`).
title (str or None): Title text. If None, will be created automatically from
`cube`. If `False`, no title will be plotted.
average_first_coord (bool): Take the mean across the first coordinate if there
are 3 dimensions.
select_valid (bool): If True, select the central contiguous unmasked subset of
data.
transform_vmin_vmax (bool): If True and `log` is True, apply the log function
used to transform the data to `vmin` and `vmax` as well.
animation_output (bool): If `True`, additional variables required to create an
animation are returned (Figure, Axes, QuadMesh, Text).
mesh (matplotlib.collections.QuadMesh): If given, update the mesh instead of
creating a new one.
title_text (matplotlib.text.Text): Title text instance. When `title` is not
`False`, `title_text` will be updated using either `title` or the
automatically generated title (if `title` is `None`).
return_cbar (bool): If True, return the colorbar.
colorbar_kwargs (dict, bool, or None): If `None`, create a new colorbar using
internal default options. These options may be altered by supplying a
corresponding dict. Colorbar creation is disabled (e.g. for animation) by
giving `False`.
The following values are not given to `colorbar()`:
cbar_tick_size: Colorbar tick param text size.
cbar_label_size: Colorbar label size.
These will be given to `colorbar()`:
label: `cube.units` by default.
orientation: `vertical` by default.
fraction: 0.15 by default.
pad: 0.07 by default.
shrink: 0.9 if `orientation='horizontal'` and 0.7 otherwise by default.
aspect: 30 by default.
anchor: (0.5, 1.0) by default.
panchor: (0.5, 0.0) by default.
format: '%.1e' if `log` and `None` otherwise by default.
ax: Parent axis which will be resized to make room for the colorbar.
cax: Axes into which the colorbar will be drawn.
coastline_kwargs (dict, bool, or None): If `None`, draw coastlines using
internal defaults. If False, do not draw coastlines. Otherwise override
defaults for coastline plotting using `ax.coastlines()`.
gridline_kwargs (dict, bool, or None): If `None`, draw gridlines using
internal defaults. If False, do not draw gridlines. Otherwise override
defaults for gridline plotting using `ax.gridlines()`.
**kwargs: Additional keyword arguments are given to `pcolormesh()`.
Returns:
matplotlib Figure: Figure used for plotting. See `fig`.
If `return_cbar`:
matplotlib Figure: Figure used for plotting. See `fig`.
matplotlib colorbar:
If `animation_output`:
matplotlib Figure: Figure used for plotting. See `fig`.
matplotlib Axes:
matplotlib QuadMesh:
matplotlib Text: Title Text.
Raises:
MaskedDataError: If all input data is masked.
ValueError: If `cmap` is a str and `extend` is not in {'neither', 'min',
'max', 'both'}.
"""
if not isinstance(cube, iris.cube.Cube):
cube = dummy_lat_lon_cube(
cube, lat_lims=dummy_lat_lims, lon_lims=dummy_lon_lims
)
if hasattr(cube.data, "mask"):
if np.all(cube.data.mask):
raise MaskedDataError("All data is masked.")
if not hasattr(cube.data, "mask"):
cube.data = np.ma.MaskedArray(cube.data, mask=False)
if colorbar_kwargs is None:
colorbar_kwargs = {}
elif colorbar_kwargs is False:
colorbar_kwargs = {"_disabled": True}
if coastline_kwargs is None:
coastline_kwargs = {}
elif coastline_kwargs is False:
coastline_kwargs = {"_disabled": True}
if gridline_kwargs is None:
gridline_kwargs = dict(zorder=0, alpha=0.4, linestyle="--", linewidth=0.3)
if select_valid and not np.all(~cube.data.mask):
cube, tr_longitudes = select_valid_subset(
cube, longitudes=cube.coord("longitude").points
)
central_longitude = np.mean(tr_longitudes)
else:
longitudes = cube.coord("longitude").points
try:
if in_360_longitude_system(longitudes):
# Translate longitudes to centre the map on Africa when all longitudes are
# present and a choice has to be made.
logger.debug("Translating longitudes from [0, 360] to [-180, 180].")
longitudes = translate_longitude_system(
longitudes, return_indices=False
)
except CoordinateSystemError:
# Assume that unusual longitudes are by design, e.g. for contiguity.
pass
finally:
central_longitude = np.mean(longitudes)
if title is None:
# Construct a default title.
title_list = [cube.name()]
try:
time_coord = cube.coord("time")
min_time = (
time_coord.cell(0).bound[0]
if time_coord.cell(0).bound is not None
else time_coord.cell(0).point
)
max_time = (
time_coord.cell(-1).bound[1]
if time_coord.cell(-1).bound is not None
else time_coord.cell(-1).point
)
if min_time == max_time:
title_list.append(f"{min_time}")
else:
title_list.append(f"{min_time} - {max_time}")
except iris.exceptions.CoordinateNotFoundError:
# No time coordinate was found, so we cannot use it in the title.
pass
title = "\n".join(title_list)
if projection is None:
logger.debug(f"Central longitude ({title}): {central_longitude:0.2f}")
projection = ccrs.Robinson(central_longitude=central_longitude)
if average_first_coord and len(cube.shape) == 3:
cube = cube.collapsed(cube.coords()[0], iris.analysis.MEAN)
if fig is None and ax is None:
fig = plt.figure()
elif fig is None:
fig = ax.get_figure()
if ax is None:
ax = plt.axes(projection=projection)
if mesh is not None:
mesh.set_array(cube.data.ravel())
else:
for coord_name in ["latitude", "longitude"]:
if not cube.coord(coord_name).has_bounds():
cube.coord(coord_name).guess_bounds()
gridlons = cube.coord("longitude").contiguous_bounds()
gridlats = cube.coord("latitude").contiguous_bounds()
if vmin is None and vmax is None:
data_vmin, data_vmax = get_cubes_vmin_vmax([cube], vmin_vmax_percentiles)
if vmin is None:
vmin = data_vmin
if vmax is None:
vmax = data_vmax
if "norm" not in kwargs:
if boundaries is None:
boundaries = get_bin_edges(
cube.data,
vmin,
vmax,
"auto" if log and log_auto_bins else nbins,
log,
"symmetric" if min_edge is None else "manual",
min_edge=min_edge,
)
if extend is None:
# Determine automatically from boundaries and the input cube.
ext_min = boundaries[0] > np.min(cube.data)
ext_max = boundaries[-1] < np.max(cube.data)
if ext_min and not ext_max:
extend = "min"
elif ext_max and not ext_min:
extend = "max"
elif ext_min and ext_max:
extend = "both"
else:
extend = "neither"
if isinstance(cmap, str):
# Allow manual flipping of colormap.
cmap_slice = slice(None)
try:
orig_cmap = plt.get_cmap(cmap)
except ValueError:
logger.debug(f"Exception while trying to access cmap '{cmap}'.")
if isinstance(cmap, str) and "_r" in cmap:
# Try to reverse the colormap manually, in case a reversed
# colormap was requested using the '_r' suffix, but this is
# not available.
cmap = cmap[:-2]
orig_cmap = plt.get_cmap(cmap)
# Flip limits to achieve reversal effect.
cmap_slice = slice(None, None, -1)
logger.debug(f"Manually reversing cmap '{cmap}'.")
else:
raise
logger.debug(f"Boundaries:{boundaries}.")
if extend == "neither":
n_colors = len(boundaries) - 1
elif extend == "min":
n_colors = len(boundaries)
elif extend == "max":
n_colors = len(boundaries)
elif extend == "both":
n_colors = len(boundaries) + 1
else:
raise ValueError(f"Unknown value for `extend` {repr(extend)}.")
if n_colors > orig_cmap.N:
logger.warning(
f"Expected at most {orig_cmap.N} colors, but got {n_colors}."
)
if n_colors <= 20:
orig_cmap = plt.get_cmap("tab20")
else:
orig_cmap = plt.get_cmap("viridis")
logger.warning(f"Reverting colormap to {orig_cmap.name}.")
cmap_sample_lims = [0, 1]
if cmap_symmetric and cmap_midpoint is not None:
# Adjust the colormap sample limits such that the deviation from
# 0.5 is proportional to the magnitude of the maximum deviation
# from the midpoint.
diffs = np.array(
(boundaries[0] - cmap_midpoint, boundaries[-1] - cmap_midpoint)
)
max_diff = max(np.abs(diffs))
scaled = diffs / max_diff
cmap_sample_lims = 0.5 + scaled * 0.5
logger.debug(f"cmap_midpoint: {cmap_midpoint}. n_colors: {n_colors}.")
if cmap_midpoint is None:
colors = orig_cmap(
np.linspace(*cmap_sample_lims[cmap_slice], n_colors)
)
logger.debug(f"No explicit midpoint, {len(colors)} colors.")
else:
# Find closest boundary.
closest_bound_index = np.argmin(
np.abs(np.asarray(boundaries[cmap_slice]) - cmap_midpoint)
)
lower_range = 0.5 - cmap_sample_lims[0]
n_lower = closest_bound_index + (
1 if extend in ("min", "both") else 0
)
upper_range = cmap_sample_lims[1] - 0.5
n_upper = (
len(boundaries)
- 1
- closest_bound_index
+ (1 if extend in ("max", "both") else 0)
)
colors = np.vstack(
(
orig_cmap(
cmap_sample_lims[0]
+
|
np.arange(n_lower)
|
numpy.arange
|
import math
import numpy as np
from random import shuffle
from itertools import product
from collections import defaultdict
import sys
# Read available tiles
tiles = {}
current_id = None
current_tile = []
try:
while True:
line = input()
if not line:
tiles[current_id] = np.array(current_tile)
current_id = None
current_tile = []
elif line[:4] == 'Tile':
current_id = int(line[5:-1])
else:
current_tile.append(list(line))
except EOFError:
pass
dir_up = 0
dir_right = 1
dir_down = 2
dir_left = 3
dirs = (dir_up, dir_right, dir_down, dir_left)
dir_vecs = {dir_up: (-1, 0), dir_right: (0, 1), dir_down: (1, 0), dir_left: (0, -1)}
rot_0 = 0
rot_90 = 1
rot_180 = 2
rot_270 = 3
rots = (rot_0, rot_90, rot_180, rot_270)
flip_0 = False
flip_lr = 0
flip_ud = 1
flips = (flip_0, flip_lr, flip_ud)
print('Finding tile neighbors…')
neighbors = defaultdict(lambda: defaultdict(set))
keys = list(tiles.keys())
for rot1, flip1, rot2, flip2 in product(rots, flips, repeat=2):
for i in range(len(keys)):
key1 = keys[i]
tile1 = np.rot90(tiles[key1], k=rot1)
if flip1 != False: tile1 = np.flip(tile1, axis=flip1)
for j in range(i + 1, len(keys)):
key2 = keys[j]
tile2 = np.rot90(tiles[key2], k=rot2)
if flip2 != False: tile2 = np.flip(tile2, axis=flip2)
if np.array_equal(tile1[0, :], tile2[-1, :]):
neighbors[key1][(rot1, flip1)].add((dir_up, key2, rot2, flip2))
neighbors[key2][(rot2, flip2)].add((dir_down, key1, rot1, flip1))
elif
|
np.array_equal(tile1[:, -1], tile2[:, 0])
|
numpy.array_equal
|
# Copyright 2018 The TensorFlow Authors. All Rights Reserved.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# ==============================================================================
"""Tests for compiled Model subclassing."""
import tensorflow.compat.v2 as tf
import os
import numpy as np
import keras
from keras import keras_parameterized
from keras import testing_utils
from keras.tests import model_subclassing_test_util as model_util
try:
import h5py # pylint:disable=g-import-not-at-top
except ImportError:
h5py = None
@keras_parameterized.run_all_keras_modes
class ModelSubclassCompiledTest(keras_parameterized.TestCase):
def test_single_io_workflow_with_np_arrays(self):
num_classes = 2
num_samples = 100
input_dim = 50
model = testing_utils.SmallSubclassMLP(
num_hidden=32, num_classes=num_classes, use_dp=True, use_bn=True)
model.compile(
loss='mse',
optimizer='rmsprop',
metrics=['acc', keras.metrics.CategoricalAccuracy()],
run_eagerly=testing_utils.should_run_eagerly())
x = np.ones((num_samples, input_dim))
y = np.zeros((num_samples, num_classes))
model.fit(x, y, epochs=2, batch_size=32, verbose=0)
_ = model.evaluate(x, y, verbose=0)
def test_multi_io_workflow_with_np_arrays(self):
num_classes = (2, 3)
num_samples = 1000
input_dim = 50
model = model_util.get_multi_io_subclass_model(
num_classes=num_classes, use_dp=True, use_bn=True)
model.compile(
loss='mse',
optimizer='rmsprop',
metrics=['acc'],
run_eagerly=testing_utils.should_run_eagerly())
x1 = np.ones((num_samples, input_dim))
x2 = np.ones((num_samples, input_dim))
y1 = np.zeros((num_samples, num_classes[0]))
y2 = np.zeros((num_samples, num_classes[1]))
model.fit([x1, x2], [y1, y2], epochs=2, batch_size=32, verbose=0)
_ = model.evaluate([x1, x2], [y1, y2], verbose=0)
def test_single_io_workflow_with_datasets(self):
num_classes = 2
num_samples = 10
input_dim = 50
with self.cached_session():
model = testing_utils.SmallSubclassMLP(
num_hidden=32, num_classes=num_classes, use_dp=True, use_bn=True)
model.compile(
loss='mse',
optimizer='rmsprop',
run_eagerly=testing_utils.should_run_eagerly())
x = np.ones((num_samples, input_dim), dtype=np.float32)
y = np.zeros((num_samples, num_classes), dtype=np.float32)
dataset = tf.data.Dataset.from_tensor_slices((x, y))
dataset = dataset.repeat(100)
dataset = dataset.batch(10)
model.fit(dataset, epochs=2, steps_per_epoch=10, verbose=0)
_ = model.evaluate(dataset, steps=10, verbose=0)
def test_attributes(self):
# layers, weights, trainable_weights, non_trainable_weights, inputs, outputs
num_classes = (2, 3)
num_samples = 100
input_dim = 50
model = model_util.get_multi_io_subclass_model(
num_classes=num_classes, use_bn=True)
x1 = np.ones((num_samples, input_dim))
x2 = np.ones((num_samples, input_dim))
y1 = np.zeros((num_samples, num_classes[0]))
y2 = np.zeros((num_samples, num_classes[1]))
self.assertEqual(model.name, 'test_model')
self.assertEqual(model.built, False)
self.assertEqual(len(model.weights), 0)
model.compile(
loss='mse',
optimizer='rmsprop',
run_eagerly=testing_utils.should_run_eagerly())
model.train_on_batch([x1, x2], [y1, y2])
self.assertEqual(model.built, True)
self.assertEqual(len(model.layers), 4)
self.assertEqual(len(model.weights), 10)
self.assertEqual(len(model.trainable_weights), 8)
self.assertEqual(len(model.non_trainable_weights), 2)
def test_updates(self):
# test that updates get run during training
num_samples = 100
input_dim = 50
class BNNet(keras.Model):
def __init__(self):
super(BNNet, self).__init__()
self.bn = keras.layers.BatchNormalization(beta_initializer='ones',
gamma_initializer='ones')
def call(self, inputs):
return self.bn(inputs)
x = np.ones((num_samples, input_dim))
y = np.ones((num_samples, input_dim))
model = BNNet()
model.compile(
loss='mse',
optimizer='rmsprop',
run_eagerly=testing_utils.should_run_eagerly())
y_ref = model.predict(x)
model.train_on_batch(x, y)
y_new = model.predict(x)
self.assertGreater(np.sum(np.abs(y_ref - y_new)), 0.1)
def test_training_and_inference_behavior(self):
# test that dropout is applied in training and not inference
num_samples = 100
input_dim = 50
class DPNet(keras.Model):
def __init__(self):
super(DPNet, self).__init__()
self.dp = keras.layers.Dropout(0.5)
self.dense = keras.layers.Dense(1,
use_bias=False,
kernel_initializer='ones')
def call(self, inputs):
x = self.dp(inputs)
return self.dense(x)
model = DPNet()
x = np.ones((num_samples, input_dim))
y = model.predict(x)
self.assertEqual(np.sum(y), np.sum(x))
model.compile(
loss='mse',
optimizer='rmsprop',
run_eagerly=testing_utils.should_run_eagerly())
loss = model.train_on_batch(x, y)
self.assertGreater(loss, 0.1)
def test_training_methods(self):
# test fit, train_on_batch
# on different input types: list, dict
num_classes = (2, 3)
num_samples = 100
input_dim = 50
x1 = np.ones((num_samples, input_dim))
x2 = np.ones((num_samples, input_dim))
y1 = np.zeros((num_samples, num_classes[0]))
y2 = np.zeros((num_samples, num_classes[1]))
model = model_util.get_multi_io_subclass_model(
num_classes=num_classes, use_bn=True)
model.compile(
loss='mse',
optimizer='rmsprop',
run_eagerly=testing_utils.should_run_eagerly())
model.fit([x1, x2], [y1, y2], epochs=2, batch_size=32, verbose=0)
model.fit({'input_1': x1, 'input_2': x2},
{'output_1': y1, 'output_2': y2},
epochs=2, batch_size=32)
model.fit([x1, x2], [y1, y2], epochs=2, batch_size=32, verbose=0,
validation_data=([x1, x2], [y1, y2]))
model = model_util.get_multi_io_subclass_model(
num_classes=num_classes, use_bn=True)
model.compile(
loss='mse',
optimizer='rmsprop',
run_eagerly=testing_utils.should_run_eagerly())
model.train_on_batch([x1, x2], [y1, y2])
model.train_on_batch({'input_1': x1, 'input_2': x2},
{'output_1': y1, 'output_2': y2})
def test_inference_methods(self):
# test predict, evaluate, test_on_batch, predict_on_batch
# on different input types: list, dict
num_classes = (2, 3)
num_samples = 100
input_dim = 50
x1 = np.ones((num_samples, input_dim))
x2 = np.ones((num_samples, input_dim))
y1 =
|
np.zeros((num_samples, num_classes[0]))
|
numpy.zeros
|
# The application object sets up and provides access to the facenet neural network interfaces
import numpy as np
import tensorflow as tf
from facenet.src.align import detect_face
from models import Face, db
from facenet.src import facenet
from scipy import misc
class Application:
def detect_faces(self, img):
# convert img into an array
im = np.array(img.data)
bounding_boxes, points = self.detect_faces_nn(im)
detected_faces = self.get_detected_faces(bounding_boxes, points)
detected_faces_json = []
# the number of boxes indicates the number of faces detected
for i in range(len(detected_faces)):
#self.store_face(detected_faces[i])
detected_faces_json.append(detected_faces[i].json())
return detected_faces_json
# returns the L2 distribution representing the differences between faces
# if the dist is < .99, then it's the same person
def compare_faces(self, img1, img2):
bounding_boxes, points = self.detect_faces_nn(img1)
aligned_img1 = self.align_img(img1, bounding_boxes, self.image_size, self.margin)
bounding_boxes, points = self.detect_faces_nn(img2)
aligned_img2 = self.align_img(img2, bounding_boxes, self.image_size, self.margin)
aligned_img_list = [None] * 2
aligned_img_list[0] = aligned_img1
aligned_img_list[1] = aligned_img2
images = np.stack(aligned_img_list)
emb = self.get_embeddings(images)
result = False
for i in range(len(images)):
print('%1d ' % i, end='')
dist = 0.0
for j in range(len(images)):
dist = np.sqrt(np.sum(np.square(np.subtract(emb[i, :], emb[j, :]))))
print(' %1.4f ' % dist, end='')
if dist > 0.0 and dist <= self.compare_threshold:
result = True
break
return result
# detects faces using a neural network
def detect_faces_nn(self, img):
bounding_boxes, points = detect_face.detect_face(img, self.minsize, self.pnet, self.rnet,
self.onet, self.threshold, self.factor)
return bounding_boxes, points
# converts the detected faces from the NN to a data model
def get_detected_faces(self, bounding_boxes, points):
detected_faces = []
# the number of boxes indicates the number of faces detected
for i in range(len(bounding_boxes)):
bbox = bounding_boxes[i][:4]
score = bounding_boxes[i][4:]
detected_face = Face()
detected_face.face_rectangle = bbox.tolist()
landmarks = []
for p in range(len(points)):
landmarks.append(float(points[p][i]))
detected_face.face_landmarks = landmarks
detected_face.confidence = score[0]
detected_faces.append(detected_face)
return detected_faces
# aligns and prewhitens the detected faces from the image
def align_img(self, img, bounding_boxes, image_size, margin):
img_size =
|
np.asarray(img.shape)
|
numpy.asarray
|
# Copyright (c) 2019, NVIDIA CORPORATION. All rights reserved.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
from __future__ import absolute_import
from __future__ import division
from __future__ import print_function
from absl import flags
import os
os.environ['TF_CPP_MIN_LOG_LEVEL'] = '3'
import absl.logging as _logging # pylint: disable=unused-import
import sys
import getopt
import json
from datetime import datetime
import tensorflow as tf
from tensorflow.python.client import timeline
import numpy as np
import modeling
def getInput(input_file, batch_size, i):
data = np.load(input_file)
arr_input_ids=data["input_ids:0"]
arr_input_mask=data["input_mask:0"]
arr_segment_ids=data["segment_ids:0"]
input_ids=np.transpose(arr_input_ids[i*batch_size:(i+1)*batch_size,:])
input_mask=
|
np.transpose(arr_input_mask[i*batch_size:(i+1)*batch_size,:])
|
numpy.transpose
|
# MIT License - Copyright <NAME> and contributors
# See the LICENSE.md file included in this source code package
"""A mixture distribution that has no analytical expression for MI."""
from ennemi import estimate_mi
from scipy.integrate import dblquad
from scipy.stats import norm, multivariate_normal as mvnorm
import numpy as np
import unittest
class TestMixtureDistribution(unittest.TestCase):
def setUp(self) -> None:
# A mixture of two normal distributions
self.cov1 = np.asarray([[1, 0.6], [0.6, 1]])
self.cov2 = np.asarray([[1, -0.4], [-0.4, 1]])
self.mean1 = np.asarray([-1, -1])
self.mean2 = np.asarray([2, 0.5])
def test_mi(self) -> None:
# Estimate the actual MI by numerical integration
expected = self.integrate_mi()
# Create random samples from the distribution
rng = np.random.default_rng(0)
small_k1 = []
small_k3 = []
full_k2 = []
full_k40 = []
for _ in range(5):
full_sample = np.concatenate((
rng.multivariate_normal(self.mean1, self.cov1, 2000),
rng.multivariate_normal(self.mean2, self.cov2, 2000),
))
small_sample = rng.choice(full_sample, 200, replace=False)
# Estimate the MI with two k values and sample sizes
small_k1.append(estimate_mi(small_sample[:,0], small_sample[:,1], k=1))
small_k3.append(estimate_mi(small_sample[:,0], small_sample[:,1], k=3))
full_k2.append(estimate_mi(full_sample[:,0], full_sample[:,1], k=2))
full_k40.append(estimate_mi(full_sample[:,0], full_sample[:,1], k=40))
small_k1a = np.asarray(small_k1)
small_k3a = np.asarray(small_k3)
full_k2a = np.asarray(full_k2)
full_k40a = np.asarray(full_k40)
# With low sample size, increasing k should increase accuracy
self.assertLess(np.mean(np.abs(small_k3a - expected)), np.mean(np.abs(small_k1a - expected)))
self.assertAlmostEqual(
|
np.median(small_k3a)
|
numpy.median
|
# -*- coding: utf-8 -*-
# Author: <NAME> <<EMAIL>>
# License: BSD 3 clause
"""
Functions to craft features.
"""
import warnings
import numpy as np
from scipy import ndimage as ndi
from skimage.morphology import binary_opening
from skimage.morphology.selem import disk
import bigfish.stack as stack
from .input_preparation import prepare_extracted_data
# ### Main functions ###
def compute_features(cell_mask, nuc_mask, ndim, rna_coord, smfish=None,
voxel_size_yx=None, foci_coord=None,
centrosome_coord=None,
compute_distance=False, compute_intranuclear=False,
compute_protrusion=False, compute_dispersion=False,
compute_topography=False, compute_foci=False,
compute_area=False, compute_centrosome=False,
return_names=False):
"""Compute requested features.
Parameters
----------
cell_mask : np.ndarray, np.uint, np.int or bool
Surface of the cell with shape (y, x).
nuc_mask: np.ndarray, np.uint, np.int or bool
Surface of the nucleus with shape (y, x).
ndim : int
Number of spatial dimensions to consider (2 or 3).
rna_coord : np.ndarray, np.int64
Coordinates of the detected spots with shape (nb_spots, 4) or
(nb_spots, 3). One coordinate per dimension (zyx or yx dimensions)
plus the index of the cluster assigned to the spot. If no cluster was
assigned, value is -1. If cluster id is not provided foci related
features are not computed.
smfish : np.ndarray, np.uint
Image of RNAs, with shape (y, x).
voxel_size_yx : int, float or None
Size of a voxel on the yx plan, in nanometer.
foci_coord : np.ndarray, np.int64
Array with shape (nb_foci, 5) or (nb_foci, 4). One coordinate per
dimension for the foci centroid (zyx or yx coordinates), the number of
spots detected in the foci and its index.
centrosome_coord : np.ndarray, np.int64
Coordinates of the detected centrosome with shape (nb_elements, 3) or
(nb_elements, 2). One coordinate per dimension (zyx or yx dimensions).
These coordinates are mandatory to compute centrosome related features.
compute_distance : bool
Compute distance related features.
compute_intranuclear : bool
Compute nucleus related features.
compute_protrusion : bool
Compute protrusion related features.
compute_dispersion : bool
Compute dispersion indices.
compute_topography : bool
Compute topographic features.
compute_foci : bool
Compute foci related features.
compute_area : bool
Compute area related features.
compute_centrosome : bool
Compute centrosome related features.
return_names : bool
Return features names.
Returns
-------
features : np.ndarray, np.float32
Array of features.
"""
# check parameters
stack.check_parameter(voxel_size_yx=(int, float, type(None)),
compute_distance=bool,
compute_intranuclear=bool,
compute_protrusion=bool,
compute_dispersion=bool,
compute_topography=bool,
compute_foci=bool,
compute_area=bool,
compute_centrosome=bool,
return_names=bool)
if smfish is not None:
stack.check_array(smfish, ndim=[2, 3], dtype=[np.uint8, np.uint16])
if smfish.ndim == 3:
smfish = stack.maximum_projection(smfish)
if foci_coord is not None:
stack.check_array(foci_coord, ndim=2, dtype=np.int64)
# prepare input data
(cell_mask,
distance_cell, distance_cell_normalized,
centroid_cell, distance_centroid_cell,
nuc_mask, cell_mask_out_nuc,
distance_nuc, distance_nuc_normalized,
centroid_nuc, distance_centroid_nuc,
rna_coord_out_nuc,
centroid_rna, distance_centroid_rna,
centroid_rna_out_nuc, distance_centroid_rna_out_nuc,
distance_centrosome) = prepare_extracted_data(
cell_mask, nuc_mask, ndim, rna_coord, centrosome_coord)
# initialization
features = ()
names_features_distance = False
names_features_intranuclear = False
names_features_protrusion = False
names_features_dispersion = False
names_features_topography = False
names_features_foci = False
names_features_area = False
names_features_centrosome = False
# distance related features
if compute_distance:
features += features_distance(
rna_coord, distance_cell, distance_nuc, cell_mask, ndim, False)
names_features_distance = True
# nucleus related features
if compute_intranuclear:
features += features_in_out_nucleus(
rna_coord, rna_coord_out_nuc, False)
names_features_intranuclear = True
# protrusion related features
if compute_protrusion:
features += features_protrusion(
rna_coord, cell_mask, nuc_mask, ndim, voxel_size_yx, False)
names_features_protrusion = True
# dispersion indices
if compute_dispersion and smfish is not None:
features += features_dispersion(
smfish, rna_coord, centroid_rna, cell_mask, centroid_cell,
centroid_nuc, ndim, False)
names_features_dispersion = True
elif compute_dispersion and smfish is None:
raise ValueError("Dispersion features can't be computed because "
"'smfish' is not provided.")
# topographic features
if compute_topography and voxel_size_yx is not None:
features += features_topography(
rna_coord, cell_mask, nuc_mask, cell_mask_out_nuc, ndim,
voxel_size_yx, False)
names_features_topography = True
elif compute_topography and voxel_size_yx is None:
raise ValueError("Topographic features can't be computed because "
"'voxel_size_yx' is not provided.")
# foci related features
if compute_foci and foci_coord is not None:
features += features_foci(
rna_coord, foci_coord, ndim, False)
names_features_foci = True
elif compute_foci and foci_coord is None:
raise ValueError("Foci related features can't be computed because "
"'foci_coord' is not provided.")
# area related features
if compute_area:
features += features_area(
cell_mask, nuc_mask, cell_mask_out_nuc, False)
names_features_area = True
# centrosome related features
if (compute_centrosome and centrosome_coord is not None
and voxel_size_yx is not None and smfish is not None):
features += features_centrosome(
smfish, rna_coord, distance_centrosome, cell_mask, ndim,
voxel_size_yx, False)
names_features_centrosome = True
elif compute_centrosome and centrosome_coord is None:
raise ValueError("Centrosome related features can't be computed "
"because 'centrosome_coord' is not provided.")
elif compute_centrosome and voxel_size_yx is None:
raise ValueError("Centrosome related features can't be computed "
"because 'voxel_size_yx' is not provided.")
elif compute_centrosome and smfish is None:
raise ValueError("Centrosome related features can't be computed "
"because 'smfish' is not provided.")
# format features
features = np.array(features, dtype=np.float32)
features = np.round(features, decimals=2)
if return_names:
features_names = get_features_name(
names_features_distance=names_features_distance,
names_features_intranuclear=names_features_intranuclear,
names_features_protrusion=names_features_protrusion,
names_features_dispersion=names_features_dispersion,
names_features_topography=names_features_topography,
names_features_foci=names_features_foci,
names_features_area=names_features_area,
names_features_centrosome=names_features_centrosome)
return features, features_names
return features
def get_features_name(names_features_distance=False,
names_features_intranuclear=False,
names_features_protrusion=False,
names_features_dispersion=False,
names_features_topography=False,
names_features_foci=False,
names_features_area=False,
names_features_centrosome=False):
"""Return the current list of features names.
Parameters
----------
names_features_distance : bool
Return names of features related to distances from nucleus or cell
membrane.
names_features_intranuclear : bool
Return names of features related to nucleus.
names_features_protrusion : bool
Return names of features related to protrusions.
names_features_dispersion : bool
Return names of features used to quantify mRNAs dispersion within the
cell.
names_features_topography : bool
Return names of topographic features of the cell.
names_features_foci : bool
Return names of features related to foci.
names_features_area : bool
Return names of features related to area of the cell.
names_features_centrosome : bool
Return names of features related to centrosome.
Returns
-------
features_name : List[str]
A list of features name.
"""
# check parameters
stack.check_parameter(names_features_distance=bool,
names_features_intranuclear=bool,
names_features_protrusion=bool,
names_features_dispersion=bool,
names_features_topography=bool,
names_features_foci=bool,
names_features_area=bool,
names_features_centrosome=bool)
# initialization
features_name = []
# get feature names
if names_features_distance:
features_name += ["index_mean_distance_cell",
"index_median_distance_cell",
"index_mean_distance_nuc",
"index_median_distance_nuc"]
if names_features_intranuclear:
features_name += ["proportion_rna_in_nuc",
"nb_rna_out_nuc",
"nb_rna_in_nuc"]
if names_features_protrusion:
features_name += ["index_rna_protrusion",
"proportion_rna_protrusion",
"protrusion_area"]
if names_features_dispersion:
features_name += ["index_polarization",
"index_dispersion",
"index_peripheral_dispersion"]
if names_features_topography:
features_name += ["index_rna_nuc_edge",
"proportion_rna_nuc_edge"]
a = 500
for b in range(1000, 3001, 500):
features_name += ["index_rna_nuc_radius_{}_{}".format(a, b),
"proportion_rna_nuc_radius_{}_{}".format(a, b)]
a = b
a = 0
for b in range(500, 3001, 500):
features_name += ["index_rna_cell_radius_{}_{}".format(a, b),
"proportion_rna_cell_radius_{}_{}".format(a, b)]
a = b
if names_features_foci:
features_name += ["proportion_rna_in_foci"]
if names_features_area:
features_name += ["proportion_nuc_area",
"cell_area",
"nuc_area",
"cell_area_out_nuc"]
if names_features_centrosome:
features_name += ["index_mean_distance_centrosome",
"index_median_distance_centrosome",
"index_rna_centrosome",
"proportion_rna_centrosome",
"index_centrosome_dispersion"]
return features_name
# ### Features functions ###
def features_distance(rna_coord, distance_cell, distance_nuc, cell_mask, ndim,
check_input=True):
"""Compute distance related features.
Parameters
----------
rna_coord : np.ndarray, np.int64
Coordinates of the detected RNAs with zyx or yx coordinates in the
first 3 or 2 columns.
distance_cell : np.ndarray, np.float32
Distance map from the cell with shape (y, x).
distance_nuc : np.ndarray, np.float32
Distance map from the nucleus with shape (y, x).
cell_mask : np.ndarray, bool
Surface of the cell with shape (y, x).
ndim : int
Number of spatial dimensions to consider.
check_input : bool
Check input validity.
Returns
-------
index_mean_dist_cell : float
Normalized mean distance of RNAs to the cell membrane.
index_median_dist_cell : float
Normalized median distance of RNAs to the cell membrane.
index_mean_dist_nuc : float
Normalized mean distance of RNAs to the nucleus.
index_median_dist_nuc : float
Normalized median distance of RNAs to the nucleus.
"""
# check parameters
stack.check_parameter(check_input=bool)
if check_input:
stack.check_parameter(ndim=int)
if ndim not in [2, 3]:
raise ValueError("'ndim' should be 2 or 3, not {0}.".format(ndim))
stack.check_array(rna_coord, ndim=2, dtype=np.int64)
stack.check_array(distance_cell, ndim=2,
dtype=[np.float16, np.float32, np.float64])
stack.check_array(distance_nuc, ndim=2,
dtype=[np.float16, np.float32, np.float64])
stack.check_array(cell_mask, ndim=2, dtype=bool)
# case where no mRNAs are detected
if len(rna_coord) == 0:
features = (1., 1., 1., 1.)
return features
# compute mean and median distance to cell membrane
rna_distance_cell = distance_cell[rna_coord[:, ndim - 2],
rna_coord[:, ndim - 1]]
expected_distance = np.mean(distance_cell[cell_mask])
index_mean_dist_cell = np.mean(rna_distance_cell) / expected_distance
expected_distance = np.median(distance_cell[cell_mask])
index_median_dist_cell = np.median(rna_distance_cell) / expected_distance
features = (index_mean_dist_cell, index_median_dist_cell)
# compute mean and median distance to nucleus
rna_distance_nuc = distance_nuc[rna_coord[:, ndim - 2],
rna_coord[:, ndim - 1]]
expected_distance = np.mean(distance_nuc[cell_mask])
index_mean_dist_nuc =
|
np.mean(rna_distance_nuc)
|
numpy.mean
|
"""Deviation preserving reduction"""
import numpy as np
from gameanalysis import paygame
from gameanalysis import restrict
from gameanalysis import rsgame
from gameanalysis import utils
from gameanalysis.reduction import _common
from gameanalysis.reduction import hierarchical
def _devs(game, num_profs):
"""Return an array of the player counts after deviation"""
return np.tile(
np.repeat(
game.num_role_players - np.eye(game.num_roles, dtype=int),
game.num_role_strats,
0,
),
(num_profs, 1),
)
def reduce_game(full_game, red_players): # pylint: disable=too-many-locals
"""Reduce a game using deviation preserving reduction
Parameters
----------
full_game : Game
The game to reduce.
red_players : ndarray-like
The reduced number of players for each role. This will be coerced
into the proper shape if necessary.
"""
red_game = rsgame.empty_names(
full_game.role_names, red_players, full_game.strat_names
)
utils.check(
np.all((red_game.num_role_players > 1) | (full_game.num_role_players == 1)),
"all reduced players must be greater than zero",
)
utils.check(
np.all(full_game.num_role_players >= red_game.num_role_players),
"all full counts must not be less than reduced counts",
)
if full_game.is_empty():
return red_game
elif full_game.num_profiles < red_game.num_all_dpr_profiles:
full_profiles = full_game.profiles()
full_payoffs = full_game.payoffs()
else:
full_profiles = expand_profiles(full_game, red_game.all_profiles())
full_payoffs = full_game.get_payoffs(full_profiles)
valid = ~np.all(np.isnan(full_payoffs) | (full_profiles == 0), 1)
full_profiles = full_profiles[valid]
full_payoffs = full_payoffs[valid]
# Reduce
red_profiles, red_inds, full_inds, strat_inds = _reduce_profiles(
red_game, full_profiles, True
)
if red_profiles.size == 0: # Empty reduction
return red_game
# Build mapping from payoffs to reduced profiles, and use bincount
# to count the number of payoffs mapped to a specific location, and
# sum the number of payoffs mapped to a specific location
cum_inds = red_inds * full_game.num_strats + strat_inds
payoff_vals = full_payoffs[full_inds, strat_inds]
red_payoffs = np.bincount(cum_inds, payoff_vals, red_profiles.size).reshape(
red_profiles.shape
)
red_payoff_counts = np.bincount(cum_inds, minlength=red_profiles.size).reshape(
red_profiles.shape
)
mask = red_payoff_counts > 1
red_payoffs[mask] /= red_payoff_counts[mask]
unknown = (red_profiles > 0) & (red_payoff_counts == 0)
red_payoffs[unknown] = np.nan
valid = ~np.all((red_profiles == 0) | np.isnan(red_payoffs), 1)
return paygame.game_replace(red_game, red_profiles[valid], red_payoffs[valid])
def expand_profiles(full_game, profiles): # pylint: disable=too-many-locals
"""Expand profiles using dpr
Parameters
----------
full_game : Game
Game that expanded profiles will be valid for.
profiles : ndarray-like
The profiles to expand
return_contributions : bool, optional
If specified, returns a boolean array matching the shape is
returned indicating the payoffs that are needed for the initial
profiles.
"""
profiles = np.asarray(profiles, int)
utils.check(
profiles.shape[-1] == full_game.num_strats, "profiles not a valid shape"
)
if not profiles.size:
return np.empty((0, full_game.num_strats), int)
profiles = profiles.reshape((-1, full_game.num_strats))
all_red_players = np.add.reduceat(profiles, full_game.role_starts, 1)
red_players = all_red_players[0]
utils.check(np.all(all_red_players == red_players), "profiles must be valid")
num_profs = profiles.shape[0]
dev_profs = profiles[:, None] - np.eye(full_game.num_strats, dtype=int)
dev_profs = np.reshape(dev_profs, (-1, full_game.num_strats))
dev_full_players = _devs(full_game, num_profs)
mask = ~np.any(dev_profs < 0, 1)
devs = (
np.eye(full_game.num_strats, dtype=bool)[None]
.repeat(num_profs, 0)
.reshape((-1, full_game.num_strats))[mask]
)
dev_full_profs = (
_common.expand_profiles(full_game, dev_full_players[mask], dev_profs[mask])
+ devs
)
ids = utils.axis_to_elem(dev_full_profs)
return dev_full_profs[np.unique(ids, return_index=True)[1]]
def reduce_profiles(red_game, profiles):
"""Reduce profiles using dpr
Parameters
----------
red_game : Game
Game that reduced profiles will be profiles for.
profiles : ndarray-like
The profiles to reduce.
"""
return _reduce_profiles(red_game, profiles, False)
def _reduce_profiles(
red_game, profiles, return_contributions
): # pylint: disable=too-many-locals
"""Reduce profiles using dpr
Parameters
----------
red_game : Game
Game that reduced profiles will be profiles for.
profiles : ndarray-like
The profiles to reduce.
return_contributions : bool, optional
If true return ancillary information about where the payoffs come
from.
"""
profiles = np.asarray(profiles, int)
utils.check(profiles.shape[-1] == red_game.num_strats, "profiles not a valid shape")
if not profiles.size:
return np.empty((0, red_game.num_strats), int)
profiles = profiles.reshape((-1, red_game.num_strats))
all_full_players = np.add.reduceat(profiles, red_game.role_starts, 1)
full_players = all_full_players[0]
utils.check(np.all(all_full_players == full_players), "profiles must be valid")
num_profs = profiles.shape[0]
dev_profs = profiles.repeat(np.sum(profiles > 0, 1), 0)
_, strat_inds = profiles.nonzero()
dev_profs[np.arange(dev_profs.shape[0]), strat_inds] -= 1
dev_red_players = _devs(red_game, num_profs)
mask = (profiles > 0).ravel()
red_profs, reduced = _common.reduce_profiles(
red_game, dev_red_players[mask], dev_profs
)
rstrat_inds = strat_inds[reduced]
red_profs[np.arange(red_profs.shape[0]), rstrat_inds] += 1
red_profs, red_inds = np.unique(utils.axis_to_elem(red_profs), return_inverse=True)
red_profs = utils.axis_from_elem(red_profs)
if not return_contributions:
return red_profs
full_inds = np.arange(num_profs).repeat(red_game.num_strats)[mask][reduced]
return red_profs, red_inds, full_inds, rstrat_inds
def expand_deviation_profiles(full_game, rest, red_players, role_index=None):
"""Expand all deviation profiles from a restriction
Parameters
----------
full_game : Game
The game the deviations profiles will be valid for.
rest : [bool]
The restriction to get deviations from.
red_players : [int]
The number of players in each role in the reduced game.
role_index : int, optional
If specified , only expand deviations for the role selected.
"""
rest =
|
np.asarray(rest, bool)
|
numpy.asarray
|
import cv2 as cv
import numpy as np
import time
import sys, os
def load_yolo():
return cv.dnn.readNetFromDarknet("../yolo/yolov4.cfg", "../yolo/yolov4.weights")
def load_tiny_yolo():
return cv.dnn.readNetFromDarknet("../yolo/yolov4-tiny.cfg", "../yolo/yolov4-tiny.weights")
def yolo_detect_pic(ynn, img, min_area_k=0.001, min_score=0.1):
rows = img.shape[0]
cols = img.shape[1]
r =
|
np.array([cols, rows, cols, rows])
|
numpy.array
|
#!/usr/bin/env python
#
# Author: <NAME> <<EMAIL>>
#
import numpy
import ctypes
import pyscf.lib
BLKSIZE = 96 # needs to be the same to lib/gto/grid_ao_drv.c
ANG_OF = 1
NPRIM_OF = 2
NCTR_OF = 3
KAPPA_OF = 4
PTR_EXP = 5
PTR_COEFF = 6
BAS_SLOTS = 8
libcgto = pyscf.lib.load_library('libcgto')
def eval_gto(eval_name, atm, bas, env, coords,
comp=1, shls_slice=None, non0tab=None, out=None):
'''Evaluate AO function value on the given grids,
Args:
eval_name : str
========================== ========= =======================
Function type Expression
========================== ========= =======================
"GTOval_sph" spherical |AO>
"GTOval_ip_sph" spherical nabla |AO>
"GTOval_ig_sph" spherical (#C(0 1) g) |AO>
"GTOval_ipig_sph" spherical (#C(0 1) nabla g) |AO>
"GTOval_cart" cart |AO>
"GTOval_ip_cart" cart nabla |AO>
"GTOval_ig_cart" cart (#C(0 1) g)|AO>
========================== ========= =======================
atm : int32 ndarray
libcint integral function argument
bas : int32 ndarray
libcint integral function argument
env : float64 ndarray
libcint integral function argument
coords : 2D array, shape (N,3)
The coordinates of the grids.
Kwargs:
shls_slice : 2-element list
(shl_start, shl_end).
If given, only part of AOs (shl_start <= shell_id < shl_end) are
evaluated. By default, all shells defined in mol will be evaluated.
non0tab : 2D bool array
mask array to indicate whether the AO values are zero. The mask
array can be obtained by calling :func:`make_mask`
out : ndarray
If provided, results are written into this array.
Returns:
2D array of shape (N,nao) Or 3D array of shape (\*,N,nao) for AO values
Examples:
>>> mol = gto.M(atom='O 0 0 0; H 0 0 1; H 0 1 0', basis='ccpvdz')
>>> coords = numpy.random.random((100,3)) # 100 random points
>>> ao_value = eval_gto("GTOval_sph", mol._atm, mol._bas, mol._env, coords)
>>> print(ao_value.shape)
(100, 24)
>>> ao_value = eval_gto("GTOval_ig_sph", mol._atm, mol._bas, mol._env, coords, comp=3)
>>> print(ao_value.shape)
(3, 100, 24)
'''
atm = numpy.asarray(atm, dtype=numpy.int32, order='C')
bas = numpy.asarray(bas, dtype=numpy.int32, order='C')
env = numpy.asarray(env, dtype=numpy.double, order='C')
coords = numpy.asarray(coords, dtype=numpy.double, order='C')
natm = atm.shape[0]
nbas = bas.shape[0]
ngrids = coords.shape[0]
if shls_slice is None:
shls_slice = (0, nbas)
bastart, basend = shls_slice
bascount = basend - bastart
if '_cart' in eval_name:
dtype = numpy.double
l = bas[bastart:basend,ANG_OF]
nao = ((l+1)*(l+2)//2 * bas[bastart:basend,NCTR_OF]).sum()
elif '_sph' in eval_name:
dtype = numpy.double
l = bas[bastart:basend,ANG_OF]
nao = ((l*2+1) * bas[bastart:basend,NCTR_OF]).sum()
else:
raise NotImplementedError(eval_name)
if out is None:
ao = numpy.empty((comp,ngrids,nao))
else:
ao = numpy.ndarray((comp,ngrids,nao), buffer=out)
if non0tab is None:
non0tab = numpy.ones(((ngrids+BLKSIZE-1)//BLKSIZE,nbas),
dtype=numpy.int8)
drv = getattr(libcgto, eval_name)
drv(ctypes.c_int(nao), ctypes.c_int(ngrids),
ctypes.c_int(BLKSIZE), ctypes.c_int(bastart), ctypes.c_int(bascount),
ao.ctypes.data_as(ctypes.c_void_p),
coords.ctypes.data_as(ctypes.c_void_p),
non0tab.ctypes.data_as(ctypes.c_void_p),
atm.ctypes.data_as(ctypes.c_void_p), ctypes.c_int(natm),
bas.ctypes.data_as(ctypes.c_void_p), ctypes.c_int(nbas),
env.ctypes.data_as(ctypes.c_void_p))
if comp == 1:
return ao.reshape(ngrids,nao)
else:
return ao
if __name__ == '__main__':
from pyscf import gto
mol = gto.M(atom='O 0 0 0; H 0 0 1; H 0 1 0', basis='ccpvdz')
coords =
|
numpy.random.random((100,3))
|
numpy.random.random
|
from sklearn.cluster import KMeans
from sklearn.decomposition import PCA
import numpy as np
import pandas as pd
import scipy.sparse.linalg
import scipy.linalg
import matplotlib.pyplot as plt
from sigclust.constrained_kmeans import ConstrainedKMeans
import sigclust.helper_functions as helper
import sigclust.avg_2means as avg_2means
import sigclust.soft_thresholding as soft_thresholding
import sigclust.sdp_clustering as sdp_clustering
class SigClust(object):
def __init__(self, num_simulations=1000, covariance_method='soft_thresholding'):
self.num_simulations = num_simulations
self.simulated_cluster_indices = None
self.sample_cluster_index = None
self.p_value = None
self.z_score = None
self.covariance_method = covariance_method
def fit(self, data, labels):
"""Fit the SigClust object.
Currently only implementing the version of
SigClust that uses the sample covariance matrix.
data: a matrix where rows are observations and cols are features.
labels: a list or array of cluster labels. Must have two unique members.
"""
n, d = data.shape
eigenvalues = get_eigenvalues(data, self.covariance_method)
self.sample_cluster_index = helper.compute_cluster_index(data, labels)
def simulate_cluster_index():
# Recall that using invariance, it suffices to simulate
# mean 0 MVNs with diagonal covariance matrix of sample
# eigenvalues
simulated_matrix = np.random.standard_normal(size=(n, d)) *
|
np.sqrt(eigenvalues)
|
numpy.sqrt
|
import os
import h5py
import pandas as pd
import copy
from RAMAC import extract_feature
import torchvision
import torchvision.transforms as transforms
import torch.utils.data as data
import numpy as np
from PIL import Image, ImageChops
import torch
from diffusion import Diffusion
from resnet import resnet101
from cirtorch.networks.imageretrievalnet import init_network, extract_vectors
def get_imlist(path):
imlist = [os.path.join(path,f) for f in os.listdir(path) if f.endswith('.jpg')]
imlist.sort()
return imlist
def pad(im, imsize):
if im.size[0]>im.size[1]:
im = im.resize((imsize, int(imsize*im.size[1]/im.size[0])), Image.ANTIALIAS)
elif im.size[1]>im.size[0]:
im = im.resize((int(imsize*im.size[0]/im.size[1]), imsize), Image.ANTIALIAS)
else:
im = im.resize((imsize, imsize), Image.ANTIALIAS)
new_im = Image.new(im.mode,(imsize, imsize), 'white')
new_im.paste(im, (int((imsize-im.size[0])/2),
int((imsize-im.size[1])/2)))
return new_im
def read_image(path):
#if memory is insufficient, you can try to save in your computer firstly.
img_list = []
file = os.listdir(path)
file.sort()
for file_name in file:
#print(class_id, file_name)
img = Image.open(path+file_name)
img = pad(img,256)
pix_array = np.array(img)
if len(pix_array.shape) == 2:
pix_array.resize((pix_array.shape[0], pix_array.shape[1], 1))
pix_array = np.repeat(pix_array, 3, 2)
if pix_array.shape[2]==4:
pix_array=pix_array[:,:,:3]
img_list.append(pix_array)
return img_list
class DataLoader(data.Dataset):
"""Metric Learning Dataset.
"""
def __init__(self, path, transform=None, target_transform=None, nnIndex = None):
self.path = path
self.img_data = read_image(self.path)
self.transform = transform
self.target_transform = target_transform
self.nnIndex = nnIndex
def __getitem__(self, index):
img = self.img_data[index]
img = self.transform(img)
return img, index
def __len__(self):
return len(self.img_data)
def obtainf(net, testloader):
net.cuda()
net.eval()
ptr =0
test_size = testloader.dataset.__len__()
test_features = np.zeros((test_size,128))
with torch.no_grad():
for batch_idx, (inputs, indexes) in enumerate(testloader):
batchSize = inputs.size(0)
real_size = min(batchSize, 32)
batch_feat = net(inputs.cuda())
test_features[ptr:ptr+real_size,:] = np.asarray(batch_feat.cpu())
ptr += real_size
query_feats = np.array(test_features).astype('float32')
return test_features
def main():
arg_parser = argparse.ArgumentParser()
arg_parser.add_argument('train_images_path')
arg_parser.add_argument('test_images_path')
arg_parser.add_argument('predictions_path')
args = arg_parser.parse_args()
imsize=480
train_path=args.train_images_path
test_path=args.test_images_path
outfile=args.predictions_path
##read data##
data_list=os.listdir(ipath)
train_images = get_imlist(train_path)
test_images = get_imlist(test_path)
##RAMAC##
RAMAC = extract_feature(train_images, 'resnet101', imsize)
RAMAC_test = extract_feature(test_images, 'resnet101', imsize)
##UEL##
normalize = transforms.Normalize(mean=[0.485, 0.456, 0.406],
std=[0.229, 0.224, 0.225])
transform_train = transforms.Compose([
transforms.ToPILImage(),
transforms.RandomCrop(size=224),
transforms.RandomHorizontalFlip(),
transforms.ToTensor(),
normalize,
])
transform_test = transforms.Compose([
transforms.ToPILImage(),
transforms.CenterCrop(224),
transforms.ToTensor(),
normalize,
])
net = resnet101(pretrained=True,low_dim=128)
model_path = './model/UEL.t'#After training UEL
net.load_state_dict(torch.load())
imset = DataLoader(path = train_path, transform=transform_test)
train_loader = torch.utils.data.DataLoader(imset, batch_size=32, shuffle=False, num_workers=0)
UEL = obtainf(net, train_loader)
imset = DataLoader(path = test_path, transform=transform_test)
test_loader = torch.utils.data.DataLoader(imset, batch_size=32, shuffle=False, num_workers=0)
UEL_test = obtainf(net, test_loader)
##GEM##
image_size=1024
multiscale='[1, 2**(1/2), 1/2**(1/2)]'
state = torch.load('./model/retrievalSfM120k-vgg16-gem-b4dcdc6.pth')
net_params = {}
net_params['architecture'] = state['meta']['architecture']
net_params['pooling'] = state['meta']['pooling']
net_params['local_whitening'] = state['meta'].get('local_whitening', False)
net_params['regional'] = state['meta'].get('regional', False)
net_params['whitening'] = state['meta'].get('whitening', False)
net_params['mean'] = state['meta']['mean']
net_params['std'] = state['meta']['std']
net_params['pretrained'] = False
# load network
net = init_network(net_params)
net.load_state_dict(state['state_dict'])
# if whitening is precomputed
if 'Lw' in state['meta']:
net.meta['Lw'] = state['meta']['Lw']
ms = list(eval(multiscale))
msp = net.pool.p.item()
net.cuda()
net.eval()
# set up the transform
normalize = transforms.Normalize(
mean=net.meta['mean'],
std=net.meta['std']
)
transform = transforms.Compose([
transforms.ToTensor(),
normalize
])
GEM = extract_vectors(net,train_images , 480, transform, ms=ms, msp=msp).numpy().T
GEM_test = extract_vectors(net,test_images , 480, transform, ms=ms, msp=msp).numpy().T
##Retrieval##
feats=np.concatenate((RAMAC,UEL,GEM),axis=1).astype('float32')
query_feat=
|
np.concatenate((RAMAC_test, UEL_test,GEM_test),axis=1)
|
numpy.concatenate
|
#!/bin/python
# -*- coding: utf-8 -*-
import os
import time
import tqdm
import numpy as np
from grgrlib.multiprocessing import serializer
from .mpile import get_par
from .stats import summary, pmdm_report
class PMDM(object):
"""A wrapper to have a progress par for the posterior mode maximization.
"""
name = 'PMDM'
def __init__(self, model, maxfev, tol, method, linear, update_freq, verbose):
import scipy.optimize as so
print('[pmdm:]'.ljust(15, ' ') + "WARNING: I have not used this function for quite a while, it is unmaintained and probably malfunctioning! `cmaes` is likely to do a better job.")
self.model = model
self.maxfev = maxfev
self.tol = tol
self.linear = linear
self.update_freq = update_freq
if update_freq is None:
self.update_freq = int(maxfev*.1)
self.verbose = verbose
self.n = 0
self.res_max = np.inf
if not verbose:
self.pbar = tqdm.tqdm(total=maxfev, dynamic_ncols=True)
self.report = self.pbar.write
else:
self.report = print
if linear:
self.desc_str = 'linear_'
else:
self.desc_str = ''
print()
self.opt_dict = {}
if method is None:
self.method = 'Nelder-Mead'
elif isinstance(method, int):
methodl = ["Nelder-Mead", "Powell", "BFGS", "CG",
"L-BFGS-G", "SLSQP", "trust-constr", "COBYLA", "TNC"]
# Nelder-Mead: fast and reliable, but doesn't max out the likelihood completely (not that fast if far away from max)
# Powell: provides the highes likelihood but is slow and sometimes ends up in strange corners of the parameter space (sorting effects)
# BFGS: hit and go but *can* outperform Nelder-Mead without sorting effects
# CG: *can* perform well but can also get lost in a bad region with low LL
# L-BFGS-G: leaves values untouched
# SLSQP: fast but not very precise (or just wrong)
# trust-constr: very fast but terminates too early
# COBYLA: very fast but hangs up for no good reason and is effectively unusable
# TNC: gets stuck around the initial values
self.method = methodl[method]
print('[pmdm:]'.ljust(20, ' ') +
' Available methods are %s.' % ', '.join(methodl))
if self.method == 'trust-constr':
self.opt_dict = {'maxiter': np.inf}
if self.method == 'Nelder-Mead':
self.opt_dict = {
'maxiter': np.inf,
'maxfev': np.inf
}
if not verbose:
np.warnings.filterwarnings('ignore')
print('[pmdm:]'.ljust(20, ' ') +
" Maximizing posterior mode density using '%s' (meanwhile warnings are disabled)." % self.method)
else:
print('[pmdm:]'.ljust(20, ' ') +
' Maximizing posterior mode density using %s.' % self.method)
print()
def __call__(self, pars):
self.res = -self.model.lprob(pars, self.linear, self.verbose)
self.x = pars
# better ensure we're not just running with the wolfs when maxfev is hit
if self.res < self.res_max:
self.res_max = self.res
self.x_max = self.x
self.n += 1
if not self.verbose:
# ensure displayed number is correct
self.pbar.n = self.n
self.pbar.update(0)
self.pbar.set_description(
'll: '+str(-self.res.round(5)).rjust(12, ' ')+' ['+str(-self.res_max.round(5))+']')
# prints information snapshots
if self.update_freq and not self.n % self.update_freq:
pmdm_report(self.model, self.x_max,
self.res_max, self.n, self.report)
if self.n >= self.maxfev:
raise StopIteration
return self.res
def go(self):
try:
f_val = -np.inf
self.x = get_par('best', self, linear=linear,
verbose=verbose, full=False)
res = so.minimize(self, self.x, method=self.method,
tol=self.tol, options=self.opt_dict)
if not self.verbose:
self.pbar.close()
print('')
if self.res_max < res['fun']:
print('[pmdm ('+self.desc_str+'):]'.ljust(20, ' ')+str(res['message']) +
' Maximization returned value lower than actual (known) optimum ('+str(-self.res_max)+' > '+str(-self.res)+').')
else:
print('[pmdm ('+self.desc_str+'):]'.ljust(20, ' ')+str(res['message']
)+' Log-likelihood is '+str(np.round(-res['fun'], 5))+'.')
print('')
except StopIteration:
if not self.verbose:
self.pbar.close()
print('')
print('[pmdm ('+self.desc_str+'):]'.ljust(20, ' ') +
' Maximum number of function calls exceeded, exiting. Log-likelihood is '+str(np.round(-self.res_max, 5))+'...')
print('')
except KeyboardInterrupt:
if not self.verbose:
self.pbar.close()
print('')
print('[pmdm ('+self.desc_str+'):]'.ljust(20, ' ') +
' Iteration interrupted manually. Log-likelihood is '+str(np.round(-self.res_max, 5))+'...')
print('')
return self.x_max, self.res_max
def pmdm(self, linear=None, maxfev=None, linear_pre_pmdm=False, method=None, tol=1e-2, update_freq=None, verbose=False):
print('[pmdm:]'.ljust(15, ' ') + "WARNING: I have not used this function for quite a while, it is unmaintained and probably malfunctioning! `cmaes` is likely to do a better job.")
if maxfev is None:
maxfev = 1000
if linear is None:
linear = self.linear_filter
if linear_pre_pmdm:
print('[pmdm:]'.ljust(30, ' ') +
' starting pre-maximization of linear function.')
self.fdict['mode_x'] = PMDM(self, maxfev, tol, method,
True, update_freq, verbose=verbose).go()
print('[pmdm:]'.ljust(30, ' ') +
' pre-maximization of linear function done, starting actual maximization.')
description = self.description
self.pmdm_par, fmax = PMDM(self, maxfev, tol, method, linear,
update_freq, verbose=verbose).go()
self.fdict['pmdm_x'] = self.pmdm_par
self.fdict['pmdm_f'] = fmax
if 'mode_f' in self.fdict.keys() and fmax < self.fdict['mode_f']:
print('[pmdm:]'.ljust(15, ' ') + " New mode of %s is below old mode of %s. Rejecting..." %
(fmax, self.fdict['mode_f']))
else:
self.fdict['mode_x'] = self.pmdm_par
self.fdict['mode_f'] = fmax
np.warnings.filterwarnings('default')
print()
print('[estimation:]'.ljust(30, ' ')+' posterior mode values:')
with os.popen('stty size', 'r') as rows_cols:
cols = rows_cols.read().split()[1]
lnum = (len(self.prior)*8)//(int(cols)-8) + 1
prior_chunks = np.array_split(
np.array(self.fdict['prior_names']), lnum)
vals_chunks = np.array_split([round(m_val, 3)
for m_val in self.pmdm_par], lnum)
for pchunk, vchunk in zip(prior_chunks, vals_chunks):
row_format = "{:>8}" * (len(pchunk) + 1)
print(row_format.format("", *pchunk))
print(row_format.format("", *vchunk))
print()
print()
return self.pmdm_par
def cmaes(self, p0=None, sigma=None, pop_size=None, restart_factor=2, seeds=3, seed=None, linear=None, lprob_seed=None, update_freq=1000, verbose=True, debug=False, **args):
"""Find mode using CMA-ES from grgrlib.
Parameters
----------
pop_size : int
Size of each population. (Default: number of dimensions)
seeds : in, optional
Number of different seeds tried. (Default: 3)
"""
from grgrlib.optimize import cmaes as fmin
np.random.seed(seed or self.fdict['seed'])
if isinstance(seeds, int):
seeds = np.random.randint(2**32-2, size=seeds)
bnd =
|
np.array(self.fdict['prior_bounds'])
|
numpy.array
|
import logging
import re
import warnings
import numpy as np
import pandas as pd
idl2np_dtype = {1: np.byte, 2: np.int16, 3: np.int32, 4: np.float32,
5: np.float64}
idl2struct = {4: 'f', 5:'d'}
archtype2struct={'sparc': None, 'bigend': '>', 'litend': '<',
'alpha': None, 'ppc': None, 'x86': None, 'x86_64': None}
class ReadBin52(object):
"""
Class to read a bin 52 and organize the output
Attributes
----------
bin52data : ndarray
The raw, n-dimensional KRC data
casesize : int
The number of bytes in each KRC case
date : bytes
The date the read file was created
ddd : ndarray
Raw temperature data
ddd_pd : dataframe
A pandas dataframe of surface temperature data
filename : str
The name of the KRC file that was input
header : list
The KRC header unpack to a list of ints
headerlength : int
The length, in bytes, of the KRC header
ncases : int
The number of different cases the model was run for
ndim : int
TBD
ndx : int
The number of indices for a single KRC run
nhours : int
The number of hour bins per 24 Mars hours
nlats : int
The number of valid, non-zero latitude bins
nseasons : int
The number of seasons the model was run for
nvariables : int
The number of variables contained within the KRC
lookup table
structdtype : str
Describing endianess and data type
temperature_data : dataframe
A multi-indexed dataframe of surface temperatures
transferedlayers : int
The number of KRC transfered layers
version : list
The krc verison used to create the file in the form
[major, minor, release]
words_per_krc : int
The number of bytes per krc entry
"""
def __init__(self, filename, headerlen=512):
"""
Parameters
----------
filename : str
The file to read
headerlength : int
The length, in bytes, of the text header.
Default: 512
"""
# Get or setup the logging object
self.logger = logging.getLogger(__name__)
self.filename = filename
self.readbin5(headerlen)
print(self.ncases)
assert(self.ncases == self.bin52data.shape[0])
def definekrc(self, what='KRC', endianness='<'):
"""
Defines a custom binary data structure for the KRC files.
"""
if what == 'KRC':
numfd = 96 # Size of floats of T-dependent materials
numid = 40 # size of " " integers
numld = 20 # size of " " logicals
maxn1 = 30 # dimension of layers
maxn2 = 384 * 4 # dimensions of times of day
maxn3 = 16 # dimensions of iterations days
maxn4 = self.nlats * 2 - 1 # dimensions of latitudes
maxn5 = 161 # dimensions of seasons
maxn6 = 6 # dimensions of saved years
maxnh = self.nhours # dimensions of saved times of day
maxbot = 6 # dimensions of time divisions
numtit = 20 # number of 4-byte words in TITLE
numday = 5 # number of 4-byte words in DAY
e = endianness
self.logger.debug(self.structdtype)
#Define the structure of the KRC file
if self.structdtype == '<f':
self._krcstructure= np.dtype([('fd','{}{}f'.format(e, numfd)),
('id','{}{}i'.format(e, numid)),
('ld','{}{}i'.format(e, numld)),
('title','{}{}a'.format(e, 4 * numtit)),
('daytime','{}{}a'.format(e, 4 * numday)),
('alat','{}{}f4'.format(e, maxn4)),
('elev','{}{}f4'.format(e,maxn4))])
elif self.structdtype == '<d':
self._krcstructure = np.dtype([('fd','{}{}d'.format(e, numfd)),
('alat','{}{}d'.format(e, maxn4)),
('elev','{}{}d'.format(e,maxn4) ),
('id','{}{}i'.format(e, numid)),
('ld','{}{}i'.format(e, numld)),
('title','{}{}a'.format(e, 4 * numtit) ),
('daytime','{}{}a'.format(e, 4 * numday))])
def readbin5(self, headerlen):
"""
Reads the type 52 file containing KRC output.
Tested with KRC version 2.2.2. Note that the output format
can change
Parameters
----------
filename (str) Full PATH to the file
"""
def _parse_header():
header = re.findall(b'\d+', fullheader.split(b'<<')[0])
header = list(map(int, header))
self.ndim = header[0]
self.nhours = header[1]
self.nvariables = header[2]
self.nlats = header[3]
self.nseasons = header[4] - 1
self.headerlength = header[8]
self.ncases = header[0 + self.ndim]
self.header = header
print(self.header)
# Compute how large each case is
self.casesize = self.nhours
if self.ndim > 1:
for k in range(1, self.ndim - 1):
self.casesize *= header[k + 1]
def _parse_front():
# Read the front matter
front = np.fromfile(bin5, dtype=self.structdtype, count=4).astype(np.int)
self.words_per_krc = front[0]
self.ndx = front[2]
def _read_data():
bin5.seek(self.headerlength)
self.bin52data = np.fromfile(bin5,
dtype=self.structdtype)
self.logger.debug(len(self.bin52data))
indices = arraysize[1: -1]
self.bin52data = self.bin52data.reshape(indices[: : -1])
def _read_metadata():
if self.structdtype == '<f':
j = self.headerlength + 16 # Skip header plus 1 16bit entry
elif self.structdtype == '<d':
j = self.headerlength + 32 # Skip header plus 1 32bit entry
bin5.seek(j)
self.definekrc('KRC')
structarr = np.fromfile(bin5, dtype=self._krcstructure, count=1)
if self.structdtype == '<f':
self.structarr = {'fd': structarr[0][0],
'id': structarr[0][1],
'ld': structarr[0][2],
'title': structarr[0][3],
'date': structarr[0][4],
'alat': structarr[0][5],
'elevation': structarr[0][6]
}
elif self.structdtype == '<d':
self.structarr = {'fd': structarr[0][0],
'alat': structarr[0][1],
'elevation': structarr[0][2],
'id': structarr[0][3],
'ld': structarr[0][4],
'title': structarr[0][5],
'date':structarr[0][6]}
def _get_version():
ver = re.findall(b'\d+', head)
ver = list(map(int, ver))
self.version = ver[: 3]
with open(self.filename, 'rb') as bin5:
#To handle endianness and architectures
archbytes = 8
c_end = 5
fullheader = bin5.read(headerlen)
_parse_header()
print(self.header)
arraysize = self.header[0: self.ndim + 2]
arraydtypecode = arraysize[arraysize[0] + 1]
try:
arraydtype = idl2np_dtype[arraydtypecode]
self.logger.debug("Dtype: ", arraydtype)
except KeyError:
self.logger.error("Unable to determine input datatype.")
assert(len(self.header) == self.ndim + 4)
if self.headerlength > 512:
warnings.Warn('Expected header to be 512 bytes, is {} bytes'.format(self.headerlength))
return
#Get the endianness of the input file and the data type (32 or 64 bit)
archstart = self.headerlength - (archbytes + c_end)
archend = self.headerlength - c_end
encodingarch = fullheader[archstart: archend].rstrip()
encodingarch = encodingarch.decode()
self._endianness = archtype2struct[encodingarch]
self.structdtype = self._endianness + idl2struct[arraydtypecode]
#Get the date and head debug
idx2 = fullheader.find(b'>>')
idx1 = idx2 - 21
self.date = fullheader[idx1: idx1 + 20]
head = fullheader[idx2 + 2: self.headerlength - (archbytes + c_end) - 3 - idx2]
head = head.rstrip()
# Parse the header
_get_version()
_parse_front()
_read_data()
_read_metadata()
def readcase(self, case=0):
"""
Read a single dimension (case) from a bin52 file.
Parameters
-----------
case : int
The case to be extracted
"""
def latitems2dataframe():
"""
Converts Latitude items to a dataframe
"""
columns = ['# Days to Compute Soln.',
'RMS Temp. Change on Last Day',
'Predicted Final Atmospheric Temp.',
'Predicted frost amount, [kg/m^2]',
'Frost albedo (at the last time step)',
'Mean upward heat flow into soil surface on last day, [W/m^2]']
# Grab the correct slice from the data cube and reshape
latitems = layeritems[: ,: ,: ,: ,0: 3].reshape(self.ncases,
self.nseasons,
self.nlats, len(columns))
# Multi-index generation
idxcount = self.nseasons * self.nlats * self.ncases
idxpercase = self.nseasons * self.nlats
caseidx = np.empty(idxcount)
for c in range(self.ncases):
start = c * idxpercase
caseidx[start:start+idxpercase] = np.repeat(c, idxpercase)
nseasvect = np.arange(self.nseasons)
seasonidx = np.repeat(np.arange(self.nseasons), self.nlats)
latidx = np.tile(self.latitudes.values.ravel(), self.nseasons)
# Pack the dataframe
self.latitude = pd.DataFrame(latitems.reshape(self.nseasons * self.nlats, -1),
index=[caseidx, seasonidx, latidx],
columns=columns)
self.latitude.index.names = ['Case','Season' ,'Latitude']
def layer2dataframe():
"""
Converts layeritems into
"""
columns = ['Tmin', 'Tmax']
ddd = layeritems[: ,: ,: ,: ,3: 3 + self.transferedlayers].reshape(self.ncases,
self.nseasons,
self.nlats,
len(columns),
self.transferedlayers)
idxcount = self.nseasons * self.nlats * self.transferedlayers * self.ncases
caseidx = np.empty(idxcount)
idxpercase = self.nseasons * self.nlats * self.transferedlayers
for c in range(self.ncases):
start = c * idxpercase
caseidx[start:start + idxpercase] = np.repeat(c, idxpercase)
seasonidx = np.repeat(np.arange(self.nseasons), idxcount / self.nseasons / self.ncases)
nlatidx = np.repeat(self.latitudes.values.ravel(), idxcount / self.transferedlayers / self.ncases)
tranlayeridx = np.tile(np.repeat(np.arange(self.transferedlayers), self.nlats), self.nseasons)
self.layers = pd.DataFrame(ddd.reshape(idxcount, -1),
columns=columns,
index=[caseidx, seasonidx, nlatidx, tranlayeridx])
self.layers.index.names = ['Case', 'Season', 'Latitude', 'Layer']
def latelv2dataframes():
"""
Convert the latitude and elevation arrays to dataframes
"""
#All latitudes
#Hugh made some change to the krcc format, but I cannot find documentation...
if self.structdtype == '<f':
alllats = krcc[:,prelatwords:].reshape(2, nlat_include_null, self.ncases)
elif self.structdtype == '<d':
alllats = krcc[:,96:170].reshape(2, nlat_include_null, self.ncases)
#Latitudes and elevations for each case
latelv = alllats[: ,0: nlat]
if latelv.shape[-1] == 1:
latelv = latelv[:,:,0]
self.latitudes = pd.DataFrame(latelv[0], columns=['Latitude'])
self.elevations = pd.DataFrame(latelv[1], columns=['Elevation'])
def season2dataframe():
columns = ['Current Julian date (offset from J2000.0)',
'Seasonal longitude of Sun (in degrees)',
'Current surface pressure at 0 elevation (in Pascals)',
'Mean visible opacity of dust, solar wavelengths',
'Global average columnar mass of frost [kg /m^2]']
# Build a dataframe of the seasonal information
seasitems = header[:, 4 + self.words_per_krc: k ].reshape(self.ncases,
len(columns),
self.nseasons)
caseidx = np.repeat(np.arange(self.ncases), self.nseasons)
seasonidx = np.repeat(np.arange(self.nseasons), self.ncases)
flt_seasitems = seasitems.reshape(len(columns),
self.ncases * self.nseasons)
self.seasons = pd.DataFrame(flt_seasitems.T,
index=[caseidx,seasonidx],
columns=columns)
self.seasons.index.names = ['Case', 'Season']
def hourly2dataframe():
"""
Converts the hourly 'ttt' vector to a
labelled Pandas dataframe.
"""
columns = ['Final Hourly Surface Temp.',
'Final Hourly Planetary Temp.',
'Final Hourly Atmospheric Temp.',
'Hourly net downward solar flux [W/m^2]',
'Hourly net downward thermal flux [W/m^2]']
ttt = self.bin52data[: ,self.ndx: ,: ,0: len(columns),: ].reshape(self.ncases,
self.nseasons,
self.nlats,
len(columns),
self.nhours)
reshapettt = np.swapaxes(ttt.reshape(self.ncases * self.nseasons * self.nlats,
len(columns),
self.nhours),1,2)
shp = reshapettt.shape
reshapettt = reshapettt.reshape((shp[0] * shp[1], shp[2])).T
#Indices
caseidx = np.repeat(np.arange(self.ncases), self.nseasons * self.nlats * self.nhours)
seasonidx = np.tile(np.repeat(np.arange(self.nseasons), self.nlats * self.nhours), self.ncases)
latidx = np.tile(np.repeat(self.latitudes.values.ravel(), self.nhours), self.nseasons)
houridx = np.tile(np.tile(np.tile(
|
np.arange(self.nhours)
|
numpy.arange
|
import numpy
import matplotlib.pyplot as plt
plt.ion()
try:
import icecube
within_icecube = True
except:
within_icecube = False
if within_icecube: from icecube import corsika
else: import corsika
import sys
if len(sys.argv)>1:
filename = sys.argv[1]
else:
filename = corsika.example_data_dir + '/DAT000011-proton-EHISTORY-MUPROD'
f = corsika.CorsikaShowerFile(filename)
f.find_event(1)
shower = f.current_shower
ptype = numpy.dtype([('code',int), ('pdg',int), ('px',float), ('py',float), ('pz',float), ('x',float), ('y',float), ('z_or_t', float), ('h',int), ('energy',float)])
def particle2dtype(particle, row):
row['code'] = particle.corsika_code
row['pdg'] = particle.pdg_code
row['px'] = particle.px
row['py'] = particle.px
row['pz'] = particle.px
row['x'] = particle.x
row['y'] = particle.y
row['z_or_t'] = particle.t_or_z
row['h'] = particle.hadronic_generation
row['energy'] = particle.kinetic_energy
particle_it = shower.particles
l = len([p for p in particle_it if p.has_parent])
daughters = numpy.zeros(l, dtype=ptype)
parents = numpy.zeros(l, dtype=ptype)
grand_parents = numpy.zeros(l, dtype=ptype)
particle_it.rewind()
i = 0
for p in particle_it:
if not p.has_parent: continue
particle2dtype(p, daughters[i])
particle2dtype(p.parent, parents[i])
particle2dtype(p.grand_parent, grand_parents[i])
i += 1
x_bins = sorted(set(daughters['code']))
x_names = [corsika.ParticleList().name_from_corsika(c) for c in x_bins]
y_bins = sorted(set(parents['code']))
y_names = [corsika.ParticleList().name_from_corsika(c) for c in y_bins]
yy_bins = sorted(set(grand_parents['code']))
yy_names = [corsika.ParticleList().name_from_corsika(c) for c in yy_bins]
contents = numpy.zeros((len(x_bins), len(y_bins)), dtype=int)
for i,x in enumerate(x_bins):
for j,y in enumerate(y_bins):
contents[i,j] =
|
numpy.sum((daughters['code']==x)*(parents['code']==y))
|
numpy.sum
|
"""
@file gazebo/model.py
Defines a model of the gazebo simulator aircraft
and provides functionality to linearize about
steady level and steady turning flight conditions.
"""
import numpy as np
from casadi import *
import sys
# Vehicle parameters
M = 1.5 # kg
IX = 0.19756 # kg m^2
IY = 0.14589 # kg m^2
IZ = 0.14770 # kg m^2
G = 9.8066 # m/s^2
RHO = 1.2041 # kg/m^3
D_MAX = 0.53 # rad, control surface deflection limits
T_MAX = 10 # N, max thrust
# Position vectors from CM to center of pressure of each
# element, written in body frame coordinates in meters
P_LW =
|
np.array([-0.05, -0.3, -0.05])
|
numpy.array
|
import Globals
import tkinter as tk
from tkinter import filedialog, INSERT, DISABLED, messagebox, NORMAL, simpledialog,\
PhotoImage, BOTH, Canvas, N, S, W, E, ALL, Frame, SUNKEN, Radiobutton, GROOVE, ACTIVE, \
FLAT, END, Scrollbar, HORIZONTAL, VERTICAL, ttk, TOP, RIGHT, LEFT, ttk
import os
from os.path import normpath, basename
from PIL import Image, ImageTk
import cv2
from cv2 import imread, IMREAD_ANYCOLOR, IMREAD_ANYDEPTH, imwrite
import pydicom
from matplotlib.figure import Figure
from matplotlib.backends.backend_tkagg import FigureCanvasTkAgg
import matplotlib as mpl
from matplotlib import cm
import matplotlib.pyplot as plt
from matplotlib.backends.backend_tkagg import FigureCanvasTkAgg, NavigationToolbar2Tk
import numpy as np
#Bresenham's line algorithm
def clearAll():
Globals.profiles_film_orientation.set('-')
Globals.profiles_film_orientation_menu.config(state=ACTIVE, bg = '#ffffff', width=15, relief=FLAT)
#Globals.profiles_depth.config(state=NORMAL, fg='black')
#Globals.profiles_depth.delete('1.0', END)
#Globals.profiles_depth.insert(INSERT, " ")
Globals.profiles_iscoenter_coords = []
Globals.profiles_film_isocenter = None
Globals.profiles_film_reference_point = None
Globals.profiles_mark_isocenter_up_down_line = []
Globals.profiles_mark_isocenter_right_left_line = []
Globals.profiles_mark_isocenter_oval = []
Globals.profiles_mark_reference_point_oval = []
Globals.profiles_mark_ROI_rectangle = []
Globals.profiles_ROI_coords = []
#if(Globals.profiles_isocenter_check and Globals.profiles_ROI_check):
# Globals.profiles_done_button.config(state=DISABLED)
Globals.profiles_isocenter_check = False
Globals.profiles_ROI_check = False
Globals.profiles_reference_point_check = False
Globals.profiles_ROI_reference_point_check = False
#if(Globals.profiles_film_window_open):
# Globals.profiles_film_window.destroy()
# Globals.profiles_film_window_open = False
Globals.profiles_upload_button_film.config(state=ACTIVE)
Globals.profiles_upload_button_doseplan.config(state=DISABLED)
Globals.profiles_upload_button_rtplan.config(state=DISABLED)
Globals.profiles_distance_isocenter_ROI = []
Globals.profiles_film_dataset = None
Globals.profiles_film_dataset_red_channel = None
Globals.profiles_film_dataset_ROI = None
Globals.profiles_film_dataset_ROI_red_channel = None
Globals.profiles_film_match_isocenter_dataset = np.zeros((7,7))
Globals.profiles_dataset_doseplan = None
Globals.profiles_dataset_rtplan = None
Globals.profiles_isocenter_mm = None
Globals.profiles_test_if_added_rtplan = False
Globals.profiles_test_if_added_doseplan = False
Globals.tab4_canvas.unbind("<Up>")
Globals.tab4_canvas.unbind("<Down>")
return
def getCoordsInRandomLine(x1,y1,x2,y2):
points = []
issteep = abs(y2-y1) - abs(x2-x1)
if issteep > 0:
x1, y1 = y1, x1
x2, y2 = y2, x2
rev = False
if x1 > x2:
x1, x2 = x2, x1
y1, y2 = y2, y1
rev = True
deltax = x2 - x1
deltay = abs(y2-y1)
error = int(deltax / 2)
y = y1
ystep = None
if y1 < y2:
ystep = 1
else:
ystep = -1
for x in range(x1, x2 + 1):
if issteep:
points.append((y, x))
else:
points.append((x, y))
error -= deltay
if error < 0:
y += ystep
error += deltax
# Reverse the list if the coordinates were reversed
if rev:
points.reverse()
return points
def drawProfiles(even):
if Globals.profiles_choice_of_profile_line_type.get() == 'h' or Globals.profiles_choice_of_profile_line_type.get() == 'v':
Globals.profiles_lines = []
if Globals.profiles_dataset_doseplan == None:
return
Globals.profiles_adjust_button_right.config(state=ACTIVE)
Globals.profiles_adjust_button_left.config(state=ACTIVE)
Globals.profiles_adjust_button_down.config(state=ACTIVE)
Globals.profiles_adjust_button_up.config(state=ACTIVE)
Globals.profiles_adjust_button_return.config(state=ACTIVE)
def draw(line_orient, dataset_film, dataset_doseplan):
Globals.profile_plot_canvas.delete('all')
fig= Figure(figsize=(5,3))
a = fig.add_subplot(111)
plot_canvas = FigureCanvasTkAgg(fig, master=Globals.profile_plot_canvas)
plot_canvas.get_tk_widget().grid(row=0,column=0,columnspan=4, sticky=N+E+W+S, padx=(5,0), pady=(0,0))
#annotation = a.annotate("HEI", xy=(0,0), xytext=(0,20))
#annotation.set_visible(False)
#txt = tk.Text(Globals.profile_plot_canvas, width=50, height=6)
#txt.insert(INSERT, " ")
#txt.grid(row=1, column = 1, sticky=N+E+W+S, pady=(5,0), padx=(5,0))
#txt.config(bg='#ffffff', font=('calibri', '10'), state=DISABLED, relief=FLAT, bd= 0)
cols = (' ', 'Point match', 'Distance', 'Dose', 'Rel. to max', 'Rel. to target')
listBox = ttk.Treeview(Globals.profile_plot_canvas, columns=cols, show='headings')
for col in cols:
listBox.heading(col, text=col, anchor=W)
listBox.column(col ,width=84, stretch=False, anchor=W)
listBox.grid(row=1, column=0, columnspan=4)
lst = [['Film: ', ' ', ' ', ' ', ' ', ' '],\
['Doseplan: ', ' ', ' ', ' ', ' ', ' ']]
for i, (name, m, dis, d, rdROI, rdTarget) in enumerate(lst):
listBox.insert("", "end", values=(name, m, dis, d, rdROI, rdTarget))
#a.text(0,0, "", fontsize=7, bbox=dict(facecolor='gray', alpha=0.1))
#txt.set_visible(False)
v_line = a.axvline(x=0, ymin=0, ymax=50, c='gray')
#v_line.set_visible(False)
if line_orient == 'h':
if(Globals.profiles_dataset_doseplan.PixelSpacing==[1, 1]):
dy = Globals.profiles_doseplan_dataset_ROI.shape[1]/2
elif(Globals.profiles_dataset_doseplan.PixelSpacing==[2, 2]):
dy = Globals.profiles_doseplan_dataset_ROI.shape[1]*2/2
else:
dy = Globals.profiles_doseplan_dataset_ROI.shape[1]*3/2
dx = dataset_film.shape[1]*0.2/2
x = np.linspace(-dx,dx, dataset_film.shape[1])
y = np.linspace(-dy,dy, Globals.profiles_doseplan_dataset_ROI.shape[1])
plot_film = dataset_film[Globals.profiles_coordinate_in_dataset,:]/100
plot_doseplan = dataset_doseplan[Globals.profiles_coordinate_in_dataset, :]
film = a.plot(x,plot_film, color='r', label='Film')
dose = a.plot(y,plot_doseplan, color='b', label='Doseplan')
elif line_orient == 'v':
if(Globals.profiles_dataset_doseplan.PixelSpacing==[1, 1]):
dy = Globals.profiles_doseplan_dataset_ROI.shape[0]/2
elif(Globals.profiles_dataset_doseplan.PixelSpacing==[2, 2]):
dy = Globals.profiles_doseplan_dataset_ROI.shape[0]*2/2
else:
dy = Globals.profiles_doseplan_dataset_ROI.shape[0]*3/2
dx = dataset_film.shape[0]*0.2/2
x = np.linspace(-dx,dx, dataset_film.shape[0])
y = np.linspace(-dy,dy, Globals.profiles_doseplan_dataset_ROI.shape[0])
plot_film = dataset_film[:,Globals.profiles_coordinate_in_dataset]/100
plot_doseplan = dataset_doseplan[:, Globals.profiles_coordinate_in_dataset] #Globals.profiles_doseplan_dataset_ROI
film=a.plot(x,plot_film, color='r', label='Film')
dose=a.plot(y,plot_doseplan, color='b', label='Doseplan')
elif line_orient == 'd':
start_f_x, start_f_y = Globals.profiles_line_coords_film[0]
end_f_x, end_f_y = Globals.end_point
dx=np.sqrt(((end_f_x-start_f_x)*0.2)**2 + ((end_f_y-start_f_y)*0.2)**2)/2
if(Globals.profiles_dataset_doseplan.PixelSpacing==[1, 1]):
start_d_x, start_d_y = Globals.profiles_line_coords_doseplan[0]
end_d_x, end_d_y = Globals.end_point
end_d_x=end_d_x/5; end_d_y=end_d_y/5
dy=np.sqrt(((end_d_x-start_d_x))**2 + ((end_d_y-start_d_y))**2)/2
elif(Globals.profiles_dataset_doseplan.PixelSpacing==[2, 2]):
start_d_x, start_d_y = Globals.profiles_line_coords_doseplan[0]
end_d_x, end_d_y = Globals.end_point
end_d_x=end_d_x/10; end_d_y=end_d_y/10
dy=np.sqrt(((end_d_x-start_d_x)*2)**2 + ((end_d_y-start_d_y)*2)**2)/2
else:
start_d_x, start_d_y = Globals.profiles_line_coords_doseplan[0]
end_d_x, end_d_y = Globals.end_point
end_d_x=end_d_x/15; end_d_y=end_d_y/15
dy=np.sqrt(((end_d_x-start_d_x)*3)**2 + ((end_d_y-start_d_y)*3)**2)/2
print(dx, dy)
x = np.linspace(-dx,dx,len(dataset_film))
y = np.linspace(-dy,dy,len(dataset_doseplan))
plot_film=dataset_film/100
plot_doseplan=dataset_doseplan
film = a.plot(x,plot_film, color='r', label='Film')
dose= a.plot(y,plot_doseplan, 'b', label='Doseplan')
else:
messagebox.showerror("Error", "Fatal error. Something has gone wrong, try again \n(Code: draw")
return
a.legend()
a.set_title("Profiles", fontsize=12)
a.set_ylabel("Dose (Gy)", fontsize=12)
a.set_xlabel("Distance (mm)", fontsize=12)
def mouseMove(event):
if event.inaxes == a:
dist = event.xdata
idx_film = np.searchsorted(x, dist)
idx_doseplan = np.searchsorted(y, dist)
if idx_film == 0:
idx_film = 0
elif idx_film == len(x):
idx_film = len(x)-1
else:
if abs(x[idx_film-1]-dist) < abs(x[idx_film]-dist):
idx_film = idx_film-1
else:
idx_film = idx_film
if idx_doseplan == 0:
idx_doseplan = 0
elif idx_doseplan == len(y):
idx_doseplan = len(y)-1
else:
if abs(y[idx_doseplan-1]-dist) < abs(y[idx_doseplan]-dist):
idx_doseplan = idx_doseplan-1
else:
idx_doseplan = idx_doseplan
idx_film = int(np.round(idx_film))
if idx_film < 0:
idx_film = 0
if idx_film >= len(plot_film):
idx_film = len(plot_film) - 1
#if Globals.profiles_dataset_doseplan.PixelSpacing == [1, 1]:
# idx_doseplan = int(np.round(idx_doseplan/1))
#elif Globals.profiles_dataset_doseplan.PixelSpacing == [2, 2]:
# idx_doseplan = int(np.round(idx_doseplan/2))
##else:
# idx_doseplan = np.round(idx_doseplan/3)
idx_doseplan = int(np.round(idx_doseplan))
if idx_doseplan < 0:
idx_doseplan = 0
if idx_doseplan >= len(plot_doseplan):
idx_doseplan = len(plot_doseplan) - 1
match_text = "\tGraph match: \t"
match = str(np.round(min(plot_film[idx_film], plot_doseplan[idx_doseplan])/max(plot_film[idx_film], plot_doseplan[idx_doseplan])*100, 2)) + "\n"
distance_text = "Distance:\t "
dose_text = "Dose: \t"
rel_target_dose_text = "Relative to target dose: \t "
rel_mx_dose_ROI_text = "Relative to max dose in ROI: \n"
distance = str(np.round(dist,2)) + "\n"
film = "FILM: \t"
dose_film = str(np.round(plot_film[idx_film],2)) + "\t"
rel_target_dose_film = str(np.round(100*plot_film[idx_film]/Globals.max_dose_doseplan,2)) + "\t\t\t"
rel_mx_dose_ROI_film = str(np.round(100*plot_film[idx_film]/np.max(plot_film),2)) + "\n"
doseplan = "DOSEPLAN: \t"
dose_doseplan = str(np.round(plot_doseplan[idx_doseplan],2)) + "\t"
rel_target_dose_doseplan = str(np.round(100*plot_doseplan[idx_doseplan]/Globals.max_dose_doseplan,2)) + "\t\t\t"
rel_mx_dose_ROI_doseplan = str(np.round(100*plot_doseplan[idx_doseplan]/np.max(plot_doseplan),2))
notation = match_text + distance_text + dose_text, rel_target_dose_text + rel_mx_dose_ROI_text +\
film + dose_film + rel_target_dose_film + rel_mx_dose_ROI_film+\
doseplan + dose_doseplan + rel_target_dose_doseplan + rel_mx_dose_ROI_doseplan
children = listBox.get_children()
for item in children:
listBox.delete(item)
lst = [['Film: ', match, distance, dose_film, rel_mx_dose_ROI_film, rel_target_dose_film],\
['Doseplan: ', match, distance, dose_doseplan, rel_mx_dose_ROI_doseplan, rel_target_dose_doseplan]]
for i, (name, m, dis, d, rdROI, rdTarget) in enumerate(lst):
listBox.insert("", "end", values=(name, m, dis, d, rdROI, rdTarget))
y_min = max(plot_film[idx_film], plot_doseplan[idx_doseplan])-0.3*max(np.max(plot_film), np.max(plot_doseplan))
if y_min < 0:
y_min = 0
y_max = max(plot_film[idx_film], plot_doseplan[idx_doseplan])+0.3*max(np.max(plot_film), np.max(plot_doseplan))
if y_max > max(np.max(plot_film), np.max(plot_doseplan)):
y_max = max(np.max(plot_film), np.max(plot_doseplan))
v_line.set_xdata(dist)
#v_line.set_ylim(y_min,y_max)
#v_line.set_ymax = y
#v_line.set_ymax = y_max # =
#v_line = a.axvline(x=dist, ymin=0, ymax=40, c='gray')
v_line.set_visible(True)
fig.canvas.draw_idle()
def freezeData(event):
fig.canvas.mpl_disconnect(cid)
v_line.set_visible(False)
fig.canvas.draw_idle()
def startData(event):
fig.canvas.mpl_disconnect(cid2)
fig.canvas.mpl_disconnect(cid3)
draw(line_orient, dataset_film, dataset_doseplan)
cid3 = fig.canvas.mpl_connect('button_press_event', startData)
cid2 = fig.canvas.mpl_connect('button_press_event', freezeData)
else:
return
cid3 = None
cid = fig.canvas.mpl_connect('motion_notify_event', mouseMove)
fig.tight_layout()
if even:
draw('d', Globals.profiles_dataset_film_variable_draw, Globals.profiles_dataset_doesplan_variable_draw)
return
if(Globals.profiles_choice_of_profile_line_type.get() == 'h' and Globals.profiles_dataset_doseplan.PixelSpacing == [1, 1]):
dataset_film = np.zeros(\
(Globals.profiles_doseplan_dataset_ROI.shape[0], Globals.profiles_film_dataset_ROI_red_channel_dose.shape[1]))
for i in range(dataset_film.shape[0]-1):
dataset_film[i,:] = Globals.profiles_film_dataset_ROI_red_channel_dose[int((i*5)+2),:]
try:
dataset_film[dataset_film.shape[0]-1,:] = Globals.profiles_film_dataset_ROI_red_channel_dose[int((dataset_film.shape[0]-1)*5+2), :]
except:
dataset_film[dataset_film.shape[0]-1,:] = \
Globals.profiles_film_dataset_ROI_red_channel_dose[Globals.profiles_film_dataset_ROI_red_channel_dose.shape[0]-1,:]
line_doseplan = Globals.doseplan_write_image.create_line(0,Globals.doseplan_write_image_var_x,\
Globals.doseplan_write_image_width,Globals.doseplan_write_image_var_x, fill='red')
line_film_dosemap = Globals.film_dose_write_image.create_line(0,Globals.doseplan_write_image_var_x,\
Globals.doseplan_write_image_width,Globals.doseplan_write_image_var_x, fill='red')
line_film = Globals.film_write_image.create_line(0,Globals.doseplan_write_image_var_x,\
Globals.doseplan_write_image_width,Globals.doseplan_write_image_var_x, fill='red')
Globals.profiles_lines.append(line_doseplan)
Globals.profiles_lines.append(line_film_dosemap)
Globals.profiles_lines.append(line_film)
def up_button_pressed(event):
temp_x = Globals.doseplan_write_image_var_x - 5
if(temp_x < 0):
#Outside the frame
return
#inside the frame
Globals.doseplan_write_image_var_x = temp_x
Globals.profiles_coordinate_in_dataset = int(temp_x/5)
Globals.doseplan_write_image.coords(line_doseplan,0,Globals.doseplan_write_image_var_x,\
Globals.doseplan_write_image_width,Globals.doseplan_write_image_var_x)
Globals.film_dose_write_image.coords(line_film_dosemap, 0,Globals.doseplan_write_image_var_x,\
Globals.doseplan_write_image_width,Globals.doseplan_write_image_var_x)
Globals.film_write_image.coords(line_film, 0,Globals.doseplan_write_image_var_x,\
Globals.doseplan_write_image_width,Globals.doseplan_write_image_var_x)
draw('h', dataset_film, Globals.profiles_doseplan_dataset_ROI)
def down_button_pressed(event):
temp_x = Globals.doseplan_write_image_var_x + 5
if(temp_x >= Globals.doseplan_write_image_height):
#Outside the frame
return
#Inside the frame
Globals.profiles_coordinate_in_dataset = int(temp_x/5)
Globals.doseplan_write_image_var_x = temp_x
Globals.doseplan_write_image.coords(line_doseplan,0,Globals.doseplan_write_image_var_x,\
Globals.doseplan_write_image_width,Globals.doseplan_write_image_var_x)
Globals.film_dose_write_image.coords(line_film_dosemap,0,Globals.doseplan_write_image_var_x,\
Globals.doseplan_write_image_width,Globals.doseplan_write_image_var_x)
Globals.film_write_image.coords(line_film, 0,Globals.doseplan_write_image_var_x,\
Globals.doseplan_write_image_width,Globals.doseplan_write_image_var_x)
draw('h', dataset_film, Globals.profiles_doseplan_dataset_ROI)
Globals.form.bind("<Up>", up_button_pressed)
Globals.form.bind("<Down>", down_button_pressed)
if Globals.profiles_first_time_in_drawProfiles:
Globals.profiles_first_time_in_drawProfiles = False
draw('h', dataset_film, Globals.profiles_doseplan_dataset_ROI)
elif(Globals.profiles_choice_of_profile_line_type.get()=='h' and Globals.profiles_dataset_doseplan.PixelSpacing==[2, 2]):
dataset_film = np.zeros(\
(Globals.profiles_doseplan_dataset_ROI.shape[0], Globals.profiles_film_dataset_ROI_red_channel_dose.shape[1]))
for i in range(dataset_film.shape[0]-1):
dataset_film[i,:] = Globals.profiles_film_dataset_ROI_red_channel_dose[int((i*10)+5),:]
try:
dataset_film[dataset_film.shape[0]-1,:] = Globals.profiles_film_dataset_ROI_red_channel_dose[int((dataset_film.shape[0]-1)*10+5), :]
except:
dataset_film[dataset_film.shape[0]-1,:] = \
Globals.profiles_film_dataset_ROI_red_channel_dose[Globals.profiles_film_dataset_ROI_red_channel_dose.shape[0]-1,:]
line_doseplan = Globals.doseplan_write_image.create_line(0,Globals.doseplan_write_image_var_x,\
Globals.doseplan_write_image_width,Globals.doseplan_write_image_var_x, fill='red')
line_film_dosemap = Globals.film_dose_write_image.create_line(0,Globals.doseplan_write_image_var_x,\
Globals.doseplan_write_image_width,Globals.doseplan_write_image_var_x, fill='red')
line_film = Globals.film_write_image.create_line(0,Globals.doseplan_write_image_var_x,\
Globals.doseplan_write_image_width,Globals.doseplan_write_image_var_x, fill='red')
Globals.profiles_lines.append(line_doseplan)
Globals.profiles_lines.append(line_film_dosemap)
Globals.profiles_lines.append(line_film)
def up_button_pressed(event):
temp_x = Globals.doseplan_write_image_var_x - 10
if(temp_x < 0):
#Outside the frame
return
#inside the frame
Globals.doseplan_write_image_var_x = temp_x
Globals.profiles_coordinate_in_dataset = int(temp_x/10)
Globals.doseplan_write_image.coords(line_doseplan,0,Globals.doseplan_write_image_var_x,\
Globals.doseplan_write_image_width,Globals.doseplan_write_image_var_x)
Globals.film_dose_write_image.coords(line_film_dosemap, 0,Globals.doseplan_write_image_var_x,\
Globals.doseplan_write_image_width,Globals.doseplan_write_image_var_x)
Globals.film_write_image.coords(line_film, 0,Globals.doseplan_write_image_var_x,\
Globals.doseplan_write_image_width,Globals.doseplan_write_image_var_x)
draw('h', dataset_film, Globals.profiles_doseplan_dataset_ROI)
def down_button_pressed(event):
temp_x = Globals.doseplan_write_image_var_x + 10
if(temp_x >= Globals.doseplan_write_image_height):
#Outside the frame
return
#Inside the frame
Globals.profiles_coordinate_in_dataset = int(temp_x/10)
Globals.doseplan_write_image_var_x = temp_x
Globals.doseplan_write_image.coords(line_doseplan,0,Globals.doseplan_write_image_var_x,\
Globals.doseplan_write_image_width,Globals.doseplan_write_image_var_x)
Globals.film_dose_write_image.coords(line_film_dosemap,0,Globals.doseplan_write_image_var_x,\
Globals.doseplan_write_image_width,Globals.doseplan_write_image_var_x)
Globals.film_write_image.coords(line_film, 0,Globals.doseplan_write_image_var_x,\
Globals.doseplan_write_image_width,Globals.doseplan_write_image_var_x)
draw('h', dataset_film, Globals.profiles_doseplan_dataset_ROI)
Globals.form.bind("<Up>", up_button_pressed)
Globals.form.bind("<Down>", down_button_pressed)
if Globals.profiles_first_time_in_drawProfiles:
Globals.profiles_first_time_in_drawProfiles = False
draw('h', dataset_film, Globals.profiles_doseplan_dataset_ROI)
elif(Globals.profiles_choice_of_profile_line_type.get() == 'h' and Globals.profiles_dataset_doseplan.PixelSpacing==[3, 3]):
dataset_film = np.zeros(\
(Globals.profiles_doseplan_dataset_ROI.shape[0], Globals.profiles_film_dataset_ROI_red_channel_dose.shape[1]))
for i in range(dataset_film.shape[0]-1):
dataset_film[i,:] = Globals.profiles_film_dataset_ROI_red_channel_dose[int((i*15)+7),:]
try:
dataset_film[dataset_film.shape[0]-1,:] = Globals.profiles_film_dataset_ROI_red_channel_dose[int((dataset_film.shape[0]-1)*15+7), :]
except:
dataset_film[dataset_film.shape[0]-1,:] = \
Globals.profiles_film_dataset_ROI_red_channel_dose[Globals.profiles_film_dataset_ROI_red_channel_dose.shape[0]-1,:]
line_doseplan = Globals.doseplan_write_image.create_line(0,Globals.doseplan_write_image_var_x,\
Globals.doseplan_write_image_width,Globals.doseplan_write_image_var_x, fill='red')
line_film_dosemap = Globals.film_dose_write_image.create_line(0,Globals.doseplan_write_image_var_x,\
Globals.doseplan_write_image_width,Globals.doseplan_write_image_var_x, fill='red')
line_film = Globals.film_write_image.create_line(0,Globals.doseplan_write_image_var_x,\
Globals.doseplan_write_image_width,Globals.doseplan_write_image_var_x, fill='red')
Globals.profiles_lines.append(line_doseplan)
Globals.profiles_lines.append(line_film_dosemap)
Globals.profiles_lines.append(line_film)
def up_button_pressed(event):
temp_x = Globals.doseplan_write_image_var_x - 15
if(temp_x < 0):
#Outside the frame
return
#inside the frame
Globals.doseplan_write_image_var_x = temp_x
Globals.profiles_coordinate_in_dataset = int(temp_x/15)
Globals.doseplan_write_image.coords(line_doseplan,0,Globals.doseplan_write_image_var_x,\
Globals.doseplan_write_image_width,Globals.doseplan_write_image_var_x)
Globals.film_dose_write_image.coords(line_film_dosemap, 0,Globals.doseplan_write_image_var_x,\
Globals.doseplan_write_image_width,Globals.doseplan_write_image_var_x)
Globals.film_write_image.coords(line_film, 0,Globals.doseplan_write_image_var_x,\
Globals.doseplan_write_image_width,Globals.doseplan_write_image_var_x)
draw('h', dataset_film, Globals.profiles_doseplan_dataset_ROI)
def down_button_pressed(event):
temp_x = Globals.doseplan_write_image_var_x + 15
if(temp_x >= Globals.doseplan_write_image_height):
#Outside the frame
return
#Inside the frame
Globals.profiles_coordinate_in_dataset = int(temp_x/15)
Globals.doseplan_write_image_var_x = temp_x
Globals.doseplan_write_image.coords(line_doseplan,0,Globals.doseplan_write_image_var_x,\
Globals.doseplan_write_image_width,Globals.doseplan_write_image_var_x)
Globals.film_dose_write_image.coords(line_film_dosemap,0,Globals.doseplan_write_image_var_x,\
Globals.doseplan_write_image_width,Globals.doseplan_write_image_var_x)
Globals.film_write_image.coords(line_film, 0,Globals.doseplan_write_image_var_x,\
Globals.doseplan_write_image_width,Globals.doseplan_write_image_var_x)
draw('h', dataset_film,Globals.profiles_doseplan_dataset_ROI)
Globals.form.bind("<Up>", up_button_pressed)
Globals.form.bind("<Down>", down_button_pressed)
if Globals.profiles_first_time_in_drawProfiles:
Globals.profiles_first_time_in_drawProfiles = False
draw('h', dataset_film, Globals.profiles_doseplan_dataset_ROI)
elif(Globals.profiles_choice_of_profile_line_type.get() == 'v' and Globals.profiles_dataset_doseplan.PixelSpacing == [1, 1]):
dataset_film = np.zeros(\
(Globals.profiles_film_dataset_ROI_red_channel_dose.shape[0], Globals.profiles_doseplan_dataset_ROI.shape[1]))
for i in range(dataset_film.shape[1]-1):
dataset_film[:,i] = Globals.profiles_film_dataset_ROI_red_channel_dose[:,int((i*5)+2)]
try:
dataset_film[:,dataset_film.shape[1]-1] = Globals.profiles_film_dataset_ROI_red_channel_dose[:,int((dataset_film.shape[1]-1)*5+2)]
except:
dataset_film[:,dataset_film.shape[1]-1] = \
Globals.profiles_film_dataset_ROI_red_channel_dose[:,Globals.profiles_film_dataset_ROI_red_channel_dose.shape[1]-1]
line_doseplan = Globals.doseplan_write_image.create_line(Globals.doseplan_write_image_var_y, 0,\
Globals.doseplan_write_image_var_y, Globals.doseplan_write_image_height, fill='red')
line_film_dosemap = Globals.film_dose_write_image.create_line(Globals.doseplan_write_image_var_y,0,\
Globals.doseplan_write_image_var_y, Globals.doseplan_write_image_height, fill='red')
line_film = Globals.film_write_image.create_line(Globals.doseplan_write_image_var_y,0,\
Globals.doseplan_write_image_var_y,Globals.doseplan_write_image_height, fill='red')
Globals.profiles_lines.append(line_doseplan)
Globals.profiles_lines.append(line_film_dosemap)
Globals.profiles_lines.append(line_film)
def left_button_pressed(event):
temp_y = Globals.doseplan_write_image_var_y - 5
if(temp_y < 0):
#Outside the frame
return
#inside the frame
Globals.doseplan_write_image_var_y = temp_y
Globals.profiles_coordinate_in_dataset = int(temp_y/5)
Globals.doseplan_write_image.coords(line_doseplan,Globals.doseplan_write_image_var_y, 0,\
Globals.doseplan_write_image_var_y, Globals.doseplan_write_image_height)
Globals.film_dose_write_image.coords(line_film_dosemap, Globals.doseplan_write_image_var_y, 0,\
Globals.doseplan_write_image_var_y, Globals.doseplan_write_image_height)
Globals.film_write_image.coords(line_film, Globals.doseplan_write_image_var_y, 0,\
Globals.doseplan_write_image_var_y, Globals.doseplan_write_image_height)
draw('v', dataset_film, Globals.profiles_doseplan_dataset_ROI)
def right_button_pressed(event):
temp_y = Globals.doseplan_write_image_var_y + 5
if(temp_y >= Globals.doseplan_write_image_width):
#Outside the frame
return
#Inside the frame
Globals.profiles_coordinate_in_dataset = int(temp_y/5)
Globals.doseplan_write_image_var_y = temp_y
Globals.doseplan_write_image.coords(line_doseplan,Globals.doseplan_write_image_var_y, 0,\
Globals.doseplan_write_image_var_y, Globals.doseplan_write_image_height)
Globals.film_dose_write_image.coords(line_film_dosemap,Globals.doseplan_write_image_var_y, 0,\
Globals.doseplan_write_image_var_y, Globals.doseplan_write_image_height)
Globals.film_write_image.coords(line_film, Globals.doseplan_write_image_var_y, 0,\
Globals.doseplan_write_image_var_y, Globals.doseplan_write_image_height)
draw('v', dataset_film, Globals.profiles_doseplan_dataset_ROI)
Globals.form.bind("<Left>", left_button_pressed)
Globals.form.bind("<Right>", right_button_pressed)
if Globals.profiles_first_time_in_drawProfiles:
Globals.profiles_first_time_in_drawProfiles = False
draw('v', dataset_film, Globals.profiles_doseplan_dataset_ROI)
elif(Globals.profiles_choice_of_profile_line_type.get() == 'v' and Globals.profiles_dataset_doseplan.PixelSpacing == [2, 2]):
dataset_film = np.zeros(\
(Globals.profiles_film_dataset_ROI_red_channel_dose.shape[0], Globals.profiles_doseplan_dataset_ROI.shape[1]))
for i in range(dataset_film.shape[1]-1):
dataset_film[:,i] = Globals.profiles_film_dataset_ROI_red_channel_dose[:,int((i*10)+5)]
try:
dataset_film[:,dataset_film.shape[1]-1] = Globals.profiles_film_dataset_ROI_red_channel_dose[:,int((dataset_film.shape[1]-1)*10+5)]
except:
dataset_film[:,dataset_film.shape[1]-1] = \
Globals.profiles_film_dataset_ROI_red_channel_dose[:,Globals.profiles_film_dataset_ROI_red_channel_dose.shape[1]-1]
line_doseplan = Globals.doseplan_write_image.create_line(Globals.doseplan_write_image_var_y, 0,\
Globals.doseplan_write_image_var_y, Globals.doseplan_write_image_height, fill='red')
line_film_dosemap = Globals.film_dose_write_image.create_line(Globals.doseplan_write_image_var_y,0,\
Globals.doseplan_write_image_var_y, Globals.doseplan_write_image_height, fill='red')
line_film = Globals.film_write_image.create_line(Globals.doseplan_write_image_var_y,0,\
Globals.doseplan_write_image_var_y,Globals.doseplan_write_image_height, fill='red')
Globals.profiles_lines.append(line_doseplan)
Globals.profiles_lines.append(line_film_dosemap)
Globals.profiles_lines.append(line_film)
def left_button_pressed(event):
temp_y = Globals.doseplan_write_image_var_y - 10
if(temp_y < 0):
#Outside the frame
return
#inside the frame
Globals.doseplan_write_image_var_y = temp_y
Globals.profiles_coordinate_in_dataset = int(temp_y/10)
Globals.doseplan_write_image.coords(line_doseplan,Globals.doseplan_write_image_var_y, 0,\
Globals.doseplan_write_image_var_y, Globals.doseplan_write_image_height)
Globals.film_dose_write_image.coords(line_film_dosemap, Globals.doseplan_write_image_var_y, 0,\
Globals.doseplan_write_image_var_y, Globals.doseplan_write_image_height)
Globals.film_write_image.coords(line_film, Globals.doseplan_write_image_var_y, 0,\
Globals.doseplan_write_image_var_y, Globals.doseplan_write_image_height)
draw('v', dataset_film, Globals.profiles_doseplan_dataset_ROI)
def right_button_pressed(event):
temp_y = Globals.doseplan_write_image_var_y + 10
if(temp_y >= Globals.doseplan_write_image_width):
#Outside the frame
return
#Inside the frame
Globals.profiles_coordinate_in_dataset = int(temp_y/10)
Globals.doseplan_write_image_var_y = temp_y
Globals.doseplan_write_image.coords(line_doseplan,Globals.doseplan_write_image_var_y, 0,\
Globals.doseplan_write_image_var_y, Globals.doseplan_write_image_height)
Globals.film_dose_write_image.coords(line_film_dosemap,Globals.doseplan_write_image_var_y, 0,\
Globals.doseplan_write_image_var_y, Globals.doseplan_write_image_height)
Globals.film_write_image.coords(line_film, Globals.doseplan_write_image_var_y, 0,\
Globals.doseplan_write_image_var_y, Globals.doseplan_write_image_height)
draw('v', dataset_film, Globals.profiles_doseplan_dataset_ROI)
Globals.form.bind("<Left>", left_button_pressed)
Globals.form.bind("<Right>", right_button_pressed)
if Globals.profiles_first_time_in_drawProfiles:
Globals.profiles_first_time_in_drawProfiles = False
draw('v', dataset_film, Globals.profiles_doseplan_dataset_ROI)
elif(Globals.profiles_choice_of_profile_line_type.get() == 'v' and Globals.profiles_dataset_doseplan.PixelSpacing == [3, 3]):
dataset_film = np.zeros(\
(Globals.profiles_film_dataset_ROI_red_channel_dose.shape[0], Globals.profiles_doseplan_dataset_ROI.shape[1]))
for i in range(dataset_film.shape[1]-1):
dataset_film[:,i] = Globals.profiles_film_dataset_ROI_red_channel_dose[:,int((i*15)+7)]
try:
dataset_film[:,dataset_film.shape[1]-1] = Globals.profiles_film_dataset_ROI_red_channel_dose[:,int((dataset_film.shape[1]-1)*15+7)]
except:
dataset_film[:,dataset_film.shape[1]-1] = \
Globals.profiles_film_dataset_ROI_red_channel_dose[:,Globals.profiles_film_dataset_ROI_red_channel_dose.shape[1]-1]
line_doseplan = Globals.doseplan_write_image.create_line(Globals.doseplan_write_image_var_y, 0,\
Globals.doseplan_write_image_var_y, Globals.doseplan_write_image_height, fill='red')
line_film_dosemap = Globals.film_dose_write_image.create_line(Globals.doseplan_write_image_var_y,0,\
Globals.doseplan_write_image_var_y, Globals.doseplan_write_image_height, fill='red')
line_film = Globals.film_write_image.create_line(Globals.doseplan_write_image_var_y,0,\
Globals.doseplan_write_image_var_y,Globals.doseplan_write_image_height, fill='red')
Globals.profiles_lines.append(line_doseplan)
Globals.profiles_lines.append(line_film_dosemap)
Globals.profiles_lines.append(line_film)
def left_button_pressed(event):
temp_y = Globals.doseplan_write_image_var_y - 15
if(temp_y < 0):
#Outside the frame
return
#inside the frame
Globals.doseplan_write_image_var_y = temp_y
Globals.profiles_coordinate_in_dataset = int(temp_y/15)
Globals.doseplan_write_image.coords(line_doseplan,Globals.doseplan_write_image_var_y, 0,\
Globals.doseplan_write_image_var_y, Globals.doseplan_write_image_height)
Globals.film_dose_write_image.coords(line_film_dosemap, Globals.doseplan_write_image_var_y, 0,\
Globals.doseplan_write_image_var_y, Globals.doseplan_write_image_height)
Globals.film_write_image.coords(line_film, Globals.doseplan_write_image_var_y, 0,\
Globals.doseplan_write_image_var_y, Globals.doseplan_write_image_height)
draw('v', dataset_film, Globals.profiles_doseplan_dataset_ROI)
def right_button_pressed(event):
temp_y = Globals.doseplan_write_image_var_y + 15
if(temp_y >= Globals.doseplan_write_image_width):
#Outside the frame
return
#Inside the frame
Globals.profiles_coordinate_in_dataset = int(temp_y/15)
Globals.doseplan_write_image_var_y = temp_y
Globals.doseplan_write_image.coords(line_doseplan,Globals.doseplan_write_image_var_y, 0,\
Globals.doseplan_write_image_var_y, Globals.doseplan_write_image_height)
Globals.film_dose_write_image.coords(line_film_dosemap,Globals.doseplan_write_image_var_y, 0,\
Globals.doseplan_write_image_var_y, Globals.doseplan_write_image_height)
Globals.film_write_image.coords(line_film, Globals.doseplan_write_image_var_y, 0,\
Globals.doseplan_write_image_var_y, Globals.doseplan_write_image_height)
draw('v', dataset_film, Globals.profiles_doseplan_dataset_ROI)
Globals.form.bind("<Left>", left_button_pressed)
Globals.form.bind("<Right>", right_button_pressed)
if Globals.profiles_first_time_in_drawProfiles:
Globals.profiles_first_time_in_drawProfiles = False
draw('v', dataset_film, Globals.profiles_doseplan_dataset_ROI)
elif(Globals.profiles_choice_of_profile_line_type.get() == 'd' and Globals.profiles_dataset_doseplan.PixelSpacing == [1, 1]):
start_point = [0,0]
def mousePushed(event):
start_point = [event.y, event.x]
if not len(Globals.profiles_lines)==0:
Globals.doseplan_write_image.delete(Globals.profiles_lines[0])
Globals.film_dose_write_image.delete(Globals.profiles_lines[1])
Globals.film_write_image.delete(Globals.profiles_lines[2])
Globals.profiles_lines = []
line_doseplan = Globals.doseplan_write_image.create_line(start_point[1], start_point[0],start_point[1],start_point[0], fill='red')
line_film_dosemap = Globals.film_dose_write_image.create_line(start_point[1], start_point[0],start_point[1],start_point[0], fill='red')
line_film = Globals.film_write_image.create_line(start_point[1], start_point[0],start_point[1],start_point[0], fill='red')
Globals.profiles_lines.append(line_doseplan)
Globals.profiles_lines.append(line_film_dosemap)
Globals.profiles_lines.append(line_film)
def mouseMoving(event):
Globals.doseplan_write_image.coords(line_doseplan, start_point[1], start_point[0], event.x, event.y)
Globals.film_dose_write_image.coords(line_film_dosemap, start_point[1], start_point[0], event.x, event.y)
Globals.film_write_image.coords(line_film, start_point[1], start_point[0], event.x, event.y)
Globals.film_dose_write_image.bind("<B1-Motion>", mouseMoving)
def mouseReleased(event):
Globals.end_point = [event.y, event.x]
Globals.doseplan_write_image.coords(line_doseplan, start_point[1], start_point[0], event.x, event.y)
Globals.film_dose_write_image.coords(line_film_dosemap, start_point[1], start_point[0], event.x, event.y)
Globals.film_write_image.coords(line_film, start_point[1], start_point[0], event.x, event.y)
Globals.profiles_line_coords_film = getCoordsInRandomLine(start_point[1], start_point[0], Globals.end_point[1], Globals.end_point[0])
Globals.profiles_line_coords_doseplan = getCoordsInRandomLine(int(start_point[1]/5), int(start_point[0]/5), \
int(Globals.end_point[1]/5), int(Globals.end_point[0]/5))
Globals.profiles_dataset_film_variable_draw = np.zeros(len(Globals.profiles_line_coords_film))
Globals.profiles_dataset_doesplan_variable_draw=np.zeros(len(Globals.profiles_line_coords_doseplan))
for i in range(len(Globals.profiles_dataset_film_variable_draw)):
coord = Globals.profiles_line_coords_film[i]
try:
Globals.profiles_dataset_film_variable_draw[i] = Globals.profiles_film_dataset_ROI_red_channel_dose[coord[0]-1, coord[1]-1]
except:
return
for i in range(len(Globals.profiles_dataset_doesplan_variable_draw)):
coord = Globals.profiles_line_coords_doseplan[i]
try:
Globals.profiles_dataset_doesplan_variable_draw[i] = Globals.profiles_doseplan_dataset_ROI[coord[0]-1, coord[1]-1]
except:
return
draw('d', Globals.profiles_dataset_film_variable_draw, Globals.profiles_dataset_doesplan_variable_draw)
Globals.film_dose_write_image.bind("<ButtonRelease-1>", mouseReleased)
Globals.film_dose_write_image.bind("<Button-1>", mousePushed)
elif(Globals.profiles_choice_of_profile_line_type.get() == 'd' and Globals.profiles_dataset_doseplan.PixelSpacing == [2, 2]):
start_point = [0,0]
def mousePushed(event):
start_point = [event.y, event.x]
if not len(Globals.profiles_lines)==0:
Globals.doseplan_write_image.delete(Globals.profiles_lines[0])
Globals.film_dose_write_image.delete(Globals.profiles_lines[1])
Globals.film_write_image.delete(Globals.profiles_lines[2])
Globals.profiles_lines = []
line_doseplan = Globals.doseplan_write_image.create_line(start_point[1], start_point[0],start_point[1],start_point[0], fill='red')
line_film_dosemap = Globals.film_dose_write_image.create_line(start_point[1], start_point[0],start_point[1],start_point[0], fill='red')
line_film = Globals.film_write_image.create_line(start_point[1], start_point[0],start_point[1],start_point[0], fill='red')
Globals.profiles_lines.append(line_doseplan)
Globals.profiles_lines.append(line_film_dosemap)
Globals.profiles_lines.append(line_film)
def mouseMoving(event):
Globals.doseplan_write_image.coords(line_doseplan, start_point[1], start_point[0], event.x, event.y)
Globals.film_dose_write_image.coords(line_film_dosemap, start_point[1], start_point[0], event.x, event.y)
Globals.film_write_image.coords(line_film, start_point[1], start_point[0], event.x, event.y)
Globals.film_dose_write_image.bind("<B1-Motion>", mouseMoving)
def mouseReleased(event):
Globals.end_point = [event.y, event.x]
Globals.doseplan_write_image.coords(line_doseplan, start_point[1], start_point[0], event.x, event.y)
Globals.film_dose_write_image.coords(line_film_dosemap, start_point[1], start_point[0], event.x, event.y)
Globals.film_write_image.coords(line_film, start_point[1], start_point[0], event.x, event.y)
Globals.profiles_line_coords_film = getCoordsInRandomLine(start_point[1], start_point[0], Globals.end_point[1], Globals.end_point[0])
Globals.profiles_line_coords_doseplan = getCoordsInRandomLine(int(start_point[1]/10), int(start_point[0]/10), \
int(Globals.end_point[1]/10), int(Globals.end_point[0]/10))
Globals.profiles_dataset_film_variable_draw = np.zeros(len(Globals.profiles_line_coords_film))
Globals.profiles_dataset_doesplan_variable_draw=np.zeros(len(Globals.profiles_line_coords_doseplan))
for i in range(len(Globals.profiles_dataset_film_variable_draw)):
coord = Globals.profiles_line_coords_film[i]
try:
Globals.profiles_dataset_film_variable_draw[i] = \
Globals.profiles_film_dataset_ROI_red_channel_dose[coord[0]-1, coord[1]-1]
except:
return
for i in range(len(Globals.profiles_dataset_doesplan_variable_draw)):
try:
Globals.profiles_dataset_doesplan_variable_draw[i] = \
Globals.profiles_doseplan_dataset_ROI[int(Globals.profiles_line_coords_doseplan[i][1])-1, int(Globals.profiles_line_coords_doseplan[i][0])-1]
except:
return
draw('d', Globals.profiles_dataset_film_variable_draw, Globals.profiles_dataset_doesplan_variable_draw)
Globals.film_dose_write_image.bind("<ButtonRelease-1>", mouseReleased)
Globals.film_dose_write_image.bind("<Button-1>", mousePushed)
elif(Globals.profiles_choice_of_profile_line_type.get() == 'd' and Globals.profiles_dataset_doseplan.PixelSpacing == [3, 3]):
start_point = [0,0]
def mousePushed(event):
start_point = [event.y, event.x]
if not len(Globals.profiles_lines)==0:
Globals.doseplan_write_image.delete(Globals.profiles_lines[0])
Globals.film_dose_write_image.delete(Globals.profiles_lines[1])
Globals.film_write_image.delete(Globals.profiles_lines[2])
Globals.profiles_lines = []
line_doseplan = Globals.doseplan_write_image.create_line(start_point[1], start_point[0],start_point[1],start_point[0], fill='red')
line_film_dosemap = Globals.film_dose_write_image.create_line(start_point[1], start_point[0],start_point[1],start_point[0], fill='red')
line_film = Globals.film_write_image.create_line(start_point[1], start_point[0],start_point[1],start_point[0], fill='red')
Globals.profiles_lines.append(line_doseplan)
Globals.profiles_lines.append(line_film_dosemap)
Globals.profiles_lines.append(line_film)
def mouseMoving(event):
Globals.doseplan_write_image.coords(line_doseplan, start_point[1], start_point[0], event.x, event.y)
Globals.film_dose_write_image.coords(line_film_dosemap, start_point[1], start_point[0], event.x, event.y)
Globals.film_write_image.coords(line_film, start_point[1], start_point[0], event.x, event.y)
Globals.film_dose_write_image.bind("<B1-Motion>", mouseMoving)
def mouseReleased(event):
Globals.end_point = [event.y, event.x]
Globals.doseplan_write_image.coords(line_doseplan, start_point[1], start_point[0], event.x, event.y)
Globals.film_dose_write_image.coords(line_film_dosemap, start_point[1], start_point[0], event.x, event.y)
Globals.film_write_image.coords(line_film, start_point[1], start_point[0], event.x, event.y)
Globals.profiles_line_coords_film = getCoordsInRandomLine(start_point[1], start_point[0], Globals.end_point[1], Globals.end_point[0])
Globals.profiles_line_coords_doseplan = getCoordsInRandomLine(int(start_point[1]/15), int(start_point[0]/15), \
int(Globals.end_point[1]/15), int(Globals.end_point[0]/15))
Globals.profiles_dataset_film_variable_draw = np.zeros(len(Globals.profiles_line_coords_film))
Globals.profiles_dataset_doesplan_variable_draw=np.zeros(len(Globals.profiles_line_coords_doseplan))
for i in range(len(Globals.profiles_dataset_film_variable_draw)):
coord = Globals.profiles_line_coords_film[i]
try:
Globals.profiles_dataset_film_variable_draw[i] = Globals.profiles_film_dataset_ROI_red_channel_dose[coord[0]-1, coord[1]-1]
except:
return
for i in range(len(Globals.profiles_dataset_doesplan_variable_draw)):
try:
Globals.profiles_dataset_doesplan_variable_draw[i] = Globals.profiles_doseplan_dataset_ROI[int(Globals.profiles_line_coords_doseplan[i][0])-1, \
int(Globals.profiles_line_coords_doseplan[i][1])-1]
except:
return
draw('d', Globals.profiles_dataset_film_variable_draw, Globals.profiles_dataset_doesplan_variable_draw)
Globals.film_dose_write_image.bind("<ButtonRelease-1>", mouseReleased)
Globals.film_dose_write_image.bind("<Button-1>", mousePushed)
else:
messagebox.showerror("Error", "Fatal error. Something went wrong, try again \n(Code: drawProfiles)")
return
def trace_profileLineType(var, indx, mode):
test_drawProfiles()
def test_drawProfiles():
if Globals.profiles_dataset_doseplan == None:
return
else:
Globals.doseplan_write_image.delete(Globals.profiles_lines[0])
Globals.film_dose_write_image.delete(Globals.profiles_lines[1])
Globals.film_write_image.delete(Globals.profiles_lines[2])
Globals.form.unbind("<Up>")
Globals.form.unbind("<Down>")
Globals.form.unbind("<Left>")
Globals.form.unbind("<Rigth>")
Globals.profiles_first_time_in_drawProfiles = True
drawProfiles(False)
def adjustROILeft(line_orient):
if not line_orient == 'd':
Globals.doseplan_write_image.delete(Globals.profiles_lines[0])
Globals.film_dose_write_image.delete(Globals.profiles_lines[1])
Globals.film_write_image.delete(Globals.profiles_lines[2])
if(Globals.profiles_film_variable_ROI_coords[2]-1 < 0):
messagebox.showwarning("Warning", "Reached end of film \n(Code: adjustROILeft)")
return
Globals.profiles_film_variable_ROI_coords = \
[Globals.profiles_film_variable_ROI_coords[0], Globals.profiles_film_variable_ROI_coords[1],\
Globals.profiles_film_variable_ROI_coords[2]-1, Globals.profiles_film_variable_ROI_coords[3]-1]
Globals.profiles_film_dataset_ROI_red_channel_dose = \
Globals.profiles_film_dataset_red_channel_dose\
[Globals.profiles_film_variable_ROI_coords[0]:Globals.profiles_film_variable_ROI_coords[1],\
Globals.profiles_film_variable_ROI_coords[2]:Globals.profiles_film_variable_ROI_coords[3]]
Globals.profiles_first_time_in_drawProfiles = True
if line_orient == 'd':
for i in range(len(Globals.profiles_dataset_film_variable_draw)):
coord = Globals.profiles_line_coords_film[i]
try:
Globals.profiles_dataset_film_variable_draw[i] = Globals.profiles_film_dataset_ROI_red_channel_dose[coord[0]-1, coord[1]-1]
except:
return
for i in range(len(Globals.profiles_dataset_doesplan_variable_draw)):
try:
Globals.profiles_dataset_doesplan_variable_draw[i] = Globals.profiles_doseplan_dataset_ROI[int(Globals.profiles_line_coords_doseplan[i][0])-1, int(Globals.profiles_line_coords_doseplan[i][1])-1]
except:
return
drawProfiles(True)
else:
drawProfiles(False)
def adjustROIRight(line_orient):
if not line_orient == 'd':
Globals.doseplan_write_image.delete(Globals.profiles_lines[0])
Globals.film_dose_write_image.delete(Globals.profiles_lines[1])
Globals.film_write_image.delete(Globals.profiles_lines[2])
if(Globals.profiles_film_variable_ROI_coords[3]+1 > Globals.profiles_film_dataset_red_channel_dose.shape[1]):
messagebox.showwarning("Warning", "Reached end of film \n(Code: adjustROIRight)")
return
Globals.profiles_film_variable_ROI_coords = \
[Globals.profiles_film_variable_ROI_coords[0], Globals.profiles_film_variable_ROI_coords[1],\
Globals.profiles_film_variable_ROI_coords[2]+1, Globals.profiles_film_variable_ROI_coords[3]+1]
Globals.profiles_film_dataset_ROI_red_channel_dose = \
Globals.profiles_film_dataset_red_channel_dose\
[Globals.profiles_film_variable_ROI_coords[0]:Globals.profiles_film_variable_ROI_coords[1],\
Globals.profiles_film_variable_ROI_coords[2]:Globals.profiles_film_variable_ROI_coords[3]]
Globals.profiles_first_time_in_drawProfiles = True
if line_orient == 'd':
for i in range(len(Globals.profiles_dataset_film_variable_draw)):
coord = Globals.profiles_line_coords_film[i]
try:
Globals.profiles_dataset_film_variable_draw[i] = Globals.profiles_film_dataset_ROI_red_channel_dose[coord[0]-1, coord[1]-1]
except:
return
for i in range(len(Globals.profiles_dataset_doesplan_variable_draw)):
try:
Globals.profiles_dataset_doesplan_variable_draw[i] = Globals.profiles_doseplan_dataset_ROI[int(Globals.profiles_line_coords_doseplan[i][0])-1, int(Globals.profiles_line_coords_doseplan[i][1])-1]
except:
return
drawProfiles(True)
else:
drawProfiles(False)
def adjustROIUp(line_orient):
if not line_orient == 'd':
Globals.doseplan_write_image.delete(Globals.profiles_lines[0])
Globals.film_dose_write_image.delete(Globals.profiles_lines[1])
Globals.film_write_image.delete(Globals.profiles_lines[2])
if(Globals.profiles_film_variable_ROI_coords[0]-1 < 0):
messagebox.showwarning("Warning", "Reached end of film \n(Code: adjustROIUp)")
return
Globals.profiles_film_variable_ROI_coords = \
[Globals.profiles_film_variable_ROI_coords[0]-1, Globals.profiles_film_variable_ROI_coords[1]-1,\
Globals.profiles_film_variable_ROI_coords[2], Globals.profiles_film_variable_ROI_coords[3]]
Globals.profiles_film_dataset_ROI_red_channel_dose = \
Globals.profiles_film_dataset_red_channel_dose\
[Globals.profiles_film_variable_ROI_coords[0]:Globals.profiles_film_variable_ROI_coords[1],\
Globals.profiles_film_variable_ROI_coords[2]:Globals.profiles_film_variable_ROI_coords[3]]
Globals.profiles_first_time_in_drawProfiles = True
if line_orient == 'd':
for i in range(len(Globals.profiles_dataset_film_variable_draw)):
coord = Globals.profiles_line_coords_film[i]
try:
Globals.profiles_dataset_film_variable_draw[i] = Globals.profiles_film_dataset_ROI_red_channel_dose[coord[0]-1, coord[1]-1]
except:
return
for i in range(len(Globals.profiles_dataset_doesplan_variable_draw)):
try:
Globals.profiles_dataset_doesplan_variable_draw[i] = Globals.profiles_doseplan_dataset_ROI[int(Globals.profiles_line_coords_doseplan[i][0])-1, int(Globals.profiles_line_coords_doseplan[i][1])-1]
except:
return
drawProfiles(True)
else:
drawProfiles(False)
def adjustROIDown(line_orient):
if not line_orient == 'd':
Globals.doseplan_write_image.delete(Globals.profiles_lines[0])
Globals.film_dose_write_image.delete(Globals.profiles_lines[1])
Globals.film_write_image.delete(Globals.profiles_lines[2])
if(Globals.profiles_film_variable_ROI_coords[1]+1 > Globals.profiles_film_dataset_red_channel_dose.shape[0]):
messagebox.showwarning("Warning", "Reached end of film \n(Code: adjustROIDown)")
return
Globals.profiles_film_variable_ROI_coords = \
[Globals.profiles_film_variable_ROI_coords[0]+1, Globals.profiles_film_variable_ROI_coords[1]+1,\
Globals.profiles_film_variable_ROI_coords[2], Globals.profiles_film_variable_ROI_coords[3]]
Globals.profiles_film_dataset_ROI_red_channel_dose = \
Globals.profiles_film_dataset_red_channel_dose\
[Globals.profiles_film_variable_ROI_coords[0]:Globals.profiles_film_variable_ROI_coords[1],\
Globals.profiles_film_variable_ROI_coords[2]:Globals.profiles_film_variable_ROI_coords[3]]
Globals.profiles_first_time_in_drawProfiles = True
if line_orient == 'd':
for i in range(len(Globals.profiles_dataset_film_variable_draw)):
coord = Globals.profiles_line_coords_film[i]
try:
Globals.profiles_dataset_film_variable_draw[i] = Globals.profiles_film_dataset_ROI_red_channel_dose[coord[0]-1, coord[1]-1]
except:
return
for i in range(len(Globals.profiles_dataset_doesplan_variable_draw)):
try:
Globals.profiles_dataset_doesplan_variable_draw[i] = Globals.profiles_doseplan_dataset_ROI[int(Globals.profiles_line_coords_doseplan[i][0])-1, int(Globals.profiles_line_coords_doseplan[i][1])-1]
except:
return
drawProfiles(True)
else:
drawProfiles(False)
def returnToOriginalROICoordinates(line_orient):
if not line_orient == 'd':
Globals.doseplan_write_image.delete(Globals.profiles_lines[0])
Globals.film_dose_write_image.delete(Globals.profiles_lines[1])
Globals.film_write_image.delete(Globals.profiles_lines[2])
Globals.profiles_film_variable_ROI_coords = \
[Globals.profiles_ROI_coords[0][1], Globals.profiles_ROI_coords[2][1],\
Globals.profiles_ROI_coords[0][0], Globals.profiles_ROI_coords[1][0]]
Globals.profiles_film_dataset_ROI_red_channel_dose = \
Globals.profiles_film_dataset_red_channel_dose\
[Globals.profiles_film_variable_ROI_coords[0]:Globals.profiles_film_variable_ROI_coords[1],\
Globals.profiles_film_variable_ROI_coords[2]:Globals.profiles_film_variable_ROI_coords[3]]
Globals.profiles_first_time_in_drawProfiles = True
if line_orient == 'd':
for i in range(len(Globals.profiles_dataset_film_variable_draw)):
coord = Globals.profiles_line_coords_film[i]
try:
Globals.profiles_dataset_film_variable_draw[i] = Globals.profiles_film_dataset_ROI_red_channel_dose[coord[0]-1, coord[1]-1]
except:
return
for i in range(len(Globals.profiles_dataset_doesplan_variable_draw)):
Globals.profiles_dataset_doesplan_variable_draw[i] = Globals.profiles_doseplan_dataset_ROI[int(Globals.profiles_line_coords_doseplan[i][0])-1, int(Globals.profiles_line_coords_doseplan[i][1])-1]
drawProfiles(True)
else:
drawProfiles(False)
def pixel_to_dose(P,a,b,c):
ret = c + b/(P-a)
return ret
def processDoseplan_usingReferencePoint(only_one):
################ RT Plan ######################
#Find each coordinate in mm to isocenter relative to first element in doseplan
iso_1 = abs(Globals.profiles_dataset_doseplan.ImagePositionPatient[0] - Globals.profiles_dataset_rtplan.BeamSequence[0].ControlPointSequence[0].IsocenterPosition[0])
iso_2 = abs(Globals.profiles_dataset_doseplan.ImagePositionPatient[1] - Globals.profiles_dataset_rtplan.BeamSequence[0].ControlPointSequence[0].IsocenterPosition[1])
iso_3 = abs(Globals.profiles_dataset_doseplan.ImagePositionPatient[2] - Globals.profiles_dataset_rtplan.BeamSequence[0].ControlPointSequence[0].IsocenterPosition[2])
#Given as [x,y,z] in patient coordinates
Globals.profiles_isocenter_mm = [iso_1, iso_2, iso_3]
#Reads input displacement from phantom on reference point in film
#lateral = Globals.profiles_input_lateral_displacement.get("1.0",'end-1c')
#vertical = Globals.profiles_input_vertical_displacement.get("1.0", 'end-1c')
#longit = Globals.profiles_input_longitudinal_displacement.get("1.0", 'end-1c')
#if(lateral==" "):lateral=0
#if(vertical==" "):vertical=0
#if(longit==" "):longit=0
try:
Globals.profiles_vertical = int(Globals.profiles_vertical)
except:
messagebox.showerror("Error", "Could not read the vertical displacements\n (Code: displacements to integer)")
return
try:
Globals.profiles_lateral = int(Globals.profiles_lateral)
except:
messagebox.showerror("Error", "Could not read the lateral displacements\n (Code: displacements to integer)")
return
try:
Globals.profiles_longitudinal = int(Globals.profiles_longitudinal)
except:
messagebox.showerror("Error", "Could not read the longitudinal displacements\n (Code: displacements to integer)")
return
lateral = Globals.profiles_lateral
longit = Globals.profiles_longitudinal
vertical = Globals.profiles_vertical
isocenter_px = np.zeros(3)
distance_in_doseplan_ROI_reference_point_px = []
if(Globals.profiles_dataset_doseplan.PixelSpacing==[1, 1]):
#make isocenter coordinates into pixel values
isocenter_px[0] = np.round(iso_1)
isocenter_px[1] = np.round(iso_2)
isocenter_px[2] = np.round(iso_3)
#find the pixel distance from reference point to ROI corners
distance_in_doseplan_ROI_reference_point_px.append([np.round(Globals.profiles_distance_reference_point_ROI[0][0]),\
np.round(Globals.profiles_distance_reference_point_ROI[0][1])])
distance_in_doseplan_ROI_reference_point_px.append([np.round(Globals.profiles_distance_reference_point_ROI[1][0]),\
np.round(Globals.profiles_distance_reference_point_ROI[1][1])])
distance_in_doseplan_ROI_reference_point_px.append([np.round(Globals.profiles_distance_reference_point_ROI[2][0]),\
np.round(Globals.profiles_distance_reference_point_ROI[2][1])])
distance_in_doseplan_ROI_reference_point_px.append([np.round(Globals.profiles_distance_reference_point_ROI[3][0]),\
np.round(Globals.profiles_distance_reference_point_ROI[3][1])])
#Input to px
lateral_px = np.round(lateral)
vertical_px = np.round(vertical)
longit_px = np.round(longit)
#displacment to px
doseplan_lateral_displacement_px = np.round(Globals.profiles_doseplan_lateral_displacement)
doseplan_vertical_displacement_px = np.round(Globals.profiles_doseplan_vertical_displacement)
doseplan_longitudinal_displacement_px = np.round(Globals.profiles_doseplan_longitudianl_displacement)
elif(Globals.profiles_dataset_doseplan.PixelSpacing==[2, 2]):
#make isocenter coordinates into pixel values
isocenter_px[0] = np.round(iso_1/2)
isocenter_px[1] = np.round(iso_2/2)
isocenter_px[2] = np.round(iso_3/2)
#find the pixel distance from reference point to ROI corners
distance_in_doseplan_ROI_reference_point_px.append([np.round((Globals.profiles_distance_reference_point_ROI[0][0])/2),\
np.round((Globals.profiles_distance_reference_point_ROI[0][1])/2)])
distance_in_doseplan_ROI_reference_point_px.append([np.round((Globals.profiles_distance_reference_point_ROI[1][0])/2),\
np.round((Globals.profiles_distance_reference_point_ROI[1][1])/2)])
distance_in_doseplan_ROI_reference_point_px.append([np.round((Globals.profiles_distance_reference_point_ROI[2][0])/2),\
np.round((Globals.profiles_distance_reference_point_ROI[2][1])/2)])
distance_in_doseplan_ROI_reference_point_px.append([np.round((Globals.profiles_distance_reference_point_ROI[3][0])/2),\
np.round((Globals.profiles_distance_reference_point_ROI[3][1])/2)])
#Input to px
lateral_px = np.round(lateral/2)
vertical_px = np.round(vertical/2)
longit_px = np.round(longit/2)
#displacment to pc
doseplan_lateral_displacement_px = np.round((Globals.profiles_doseplan_lateral_displacement)/2)
doseplan_vertical_displacement_px = np.round((Globals.profiles_doseplan_vertical_displacement)/2)
doseplan_longitudinal_displacement_px = np.round((Globals.profiles_doseplan_longitudianl_displacement)/2)
else:
#make isocenter coordinates into pixel values
isocenter_px[0] = np.round(iso_1/3)
isocenter_px[1] = np.round(iso_2/3)
isocenter_px[2] = np.round(iso_3/3)
#find the pixel distance from reference point to ROI corners
distance_in_doseplan_ROI_reference_point_px.append([np.round((Globals.profiles_distance_reference_point_ROI[0][0])/3),\
np.round((Globals.profiles_distance_reference_point_ROI[0][1])/3)])
distance_in_doseplan_ROI_reference_point_px.append([np.round((Globals.profiles_distance_reference_point_ROI[1][0])/3),\
|
np.round((Globals.profiles_distance_reference_point_ROI[1][1])/3)
|
numpy.round
|
import numpy as np
from skimage import restoration, data, color, img_as_float, measure
from skimage.measure import compare_psnr
from skimage.restoration._denoise import _wavelet_threshold
import pywt
from skimage._shared import testing
from skimage._shared.testing import (assert_equal, assert_almost_equal,
assert_warns, assert_)
from skimage._shared._warnings import expected_warnings
np.random.seed(1234)
astro = img_as_float(data.astronaut()[:128, :128])
astro_gray = color.rgb2gray(astro)
checkerboard_gray = img_as_float(data.checkerboard())
checkerboard = color.gray2rgb(checkerboard_gray)
def test_denoise_tv_chambolle_2d():
# astronaut image
img = astro_gray.copy()
# add noise to astronaut
img += 0.5 * img.std() * np.random.rand(*img.shape)
# clip noise so that it does not exceed allowed range for float images.
img = np.clip(img, 0, 1)
# denoise
denoised_astro = restoration.denoise_tv_chambolle(img, weight=0.1)
# which dtype?
assert_(denoised_astro.dtype in [np.float, np.float32, np.float64])
from scipy import ndimage as ndi
grad = ndi.morphological_gradient(img, size=((3, 3)))
grad_denoised = ndi.morphological_gradient(denoised_astro, size=((3, 3)))
# test if the total variation has decreased
assert_(grad_denoised.dtype == np.float)
assert_(np.sqrt((grad_denoised**2).sum()) < np.sqrt((grad**2).sum()))
def test_denoise_tv_chambolle_multichannel():
denoised0 = restoration.denoise_tv_chambolle(astro[..., 0], weight=0.1)
denoised = restoration.denoise_tv_chambolle(astro, weight=0.1,
multichannel=True)
assert_equal(denoised[..., 0], denoised0)
# tile astronaut subset to generate 3D+channels data
astro3 = np.tile(astro[:64, :64, np.newaxis, :], [1, 1, 2, 1])
# modify along tiled dimension to give non-zero gradient on 3rd axis
astro3[:, :, 0, :] = 2*astro3[:, :, 0, :]
denoised0 = restoration.denoise_tv_chambolle(astro3[..., 0], weight=0.1)
denoised = restoration.denoise_tv_chambolle(astro3, weight=0.1,
multichannel=True)
assert_equal(denoised[..., 0], denoised0)
def test_denoise_tv_chambolle_float_result_range():
# astronaut image
img = astro_gray
int_astro = np.multiply(img, 255).astype(np.uint8)
assert_(np.max(int_astro) > 1)
denoised_int_astro = restoration.denoise_tv_chambolle(int_astro,
weight=0.1)
# test if the value range of output float data is within [0.0:1.0]
assert_(denoised_int_astro.dtype == np.float)
assert_(np.max(denoised_int_astro) <= 1.0)
assert_(np.min(denoised_int_astro) >= 0.0)
def test_denoise_tv_chambolle_3d():
"""Apply the TV denoising algorithm on a 3D image representing a sphere."""
x, y, z = np.ogrid[0:40, 0:40, 0:40]
mask = (x - 22)**2 + (y - 20)**2 + (z - 17)**2 < 8**2
mask = 100 * mask.astype(np.float)
mask += 60
mask += 20 * np.random.rand(*mask.shape)
mask[mask < 0] = 0
mask[mask > 255] = 255
res = restoration.denoise_tv_chambolle(mask.astype(np.uint8), weight=0.1)
assert_(res.dtype == np.float)
assert_(res.std() * 255 < mask.std())
def test_denoise_tv_chambolle_1d():
"""Apply the TV denoising algorithm on a 1D sinusoid."""
x = 125 + 100*np.sin(np.linspace(0, 8*np.pi, 1000))
x += 20 * np.random.rand(x.size)
x = np.clip(x, 0, 255)
res = restoration.denoise_tv_chambolle(x.astype(np.uint8), weight=0.1)
assert_(res.dtype == np.float)
assert_(res.std() * 255 < x.std())
def test_denoise_tv_chambolle_4d():
""" TV denoising for a 4D input."""
im = 255 * np.random.rand(8, 8, 8, 8)
res = restoration.denoise_tv_chambolle(im.astype(np.uint8), weight=0.1)
assert_(res.dtype == np.float)
assert_(res.std() * 255 < im.std())
def test_denoise_tv_chambolle_weighting():
# make sure a specified weight gives consistent results regardless of
# the number of input image dimensions
rstate = np.random.RandomState(1234)
img2d = astro_gray.copy()
img2d += 0.15 * rstate.standard_normal(img2d.shape)
img2d = np.clip(img2d, 0, 1)
# generate 4D image by tiling
img4d = np.tile(img2d[..., None, None], (1, 1, 2, 2))
w = 0.2
denoised_2d = restoration.denoise_tv_chambolle(img2d, weight=w)
denoised_4d = restoration.denoise_tv_chambolle(img4d, weight=w)
assert_(measure.compare_ssim(denoised_2d,
denoised_4d[:, :, 0, 0]) > 0.99)
def test_denoise_tv_bregman_2d():
img = checkerboard_gray.copy()
# add some random noise
img += 0.5 * img.std() * np.random.rand(*img.shape)
img = np.clip(img, 0, 1)
out1 = restoration.denoise_tv_bregman(img, weight=10)
out2 = restoration.denoise_tv_bregman(img, weight=5)
# make sure noise is reduced in the checkerboard cells
assert_(img[30:45, 5:15].std() > out1[30:45, 5:15].std())
assert_(out1[30:45, 5:15].std() > out2[30:45, 5:15].std())
def test_denoise_tv_bregman_float_result_range():
# astronaut image
img = astro_gray.copy()
int_astro = np.multiply(img, 255).astype(np.uint8)
assert_(np.max(int_astro) > 1)
denoised_int_astro = restoration.denoise_tv_bregman(int_astro, weight=60.0)
# test if the value range of output float data is within [0.0:1.0]
assert_(denoised_int_astro.dtype == np.float)
assert_(np.max(denoised_int_astro) <= 1.0)
assert_(np.min(denoised_int_astro) >= 0.0)
def test_denoise_tv_bregman_3d():
img = checkerboard.copy()
# add some random noise
img += 0.5 * img.std() * np.random.rand(*img.shape)
img = np.clip(img, 0, 1)
out1 = restoration.denoise_tv_bregman(img, weight=10)
out2 = restoration.denoise_tv_bregman(img, weight=5)
# make sure noise is reduced in the checkerboard cells
assert_(img[30:45, 5:15].std() > out1[30:45, 5:15].std())
assert_(out1[30:45, 5:15].std() > out2[30:45, 5:15].std())
def test_denoise_bilateral_2d():
img = checkerboard_gray.copy()[:50, :50]
# add some random noise
img += 0.5 * img.std() * np.random.rand(*img.shape)
img = np.clip(img, 0, 1)
out1 = restoration.denoise_bilateral(img, sigma_color=0.1,
sigma_spatial=10, multichannel=False)
out2 = restoration.denoise_bilateral(img, sigma_color=0.2,
sigma_spatial=20, multichannel=False)
# make sure noise is reduced in the checkerboard cells
assert_(img[30:45, 5:15].std() > out1[30:45, 5:15].std())
assert_(out1[30:45, 5:15].std() > out2[30:45, 5:15].std())
def test_denoise_bilateral_zeros():
img = np.zeros((10, 10))
assert_equal(img, restoration.denoise_bilateral(img, multichannel=False))
def test_denoise_bilateral_constant():
img = np.ones((10, 10)) * 5
assert_equal(img, restoration.denoise_bilateral(img, multichannel=False))
def test_denoise_bilateral_color():
img = checkerboard.copy()[:50, :50]
# add some random noise
img += 0.5 * img.std() * np.random.rand(*img.shape)
img = np.clip(img, 0, 1)
out1 = restoration.denoise_bilateral(img, sigma_color=0.1,
sigma_spatial=10, multichannel=True)
out2 = restoration.denoise_bilateral(img, sigma_color=0.2,
sigma_spatial=20, multichannel=True)
# make sure noise is reduced in the checkerboard cells
assert_(img[30:45, 5:15].std() > out1[30:45, 5:15].std())
assert_(out1[30:45, 5:15].std() > out2[30:45, 5:15].std())
def test_denoise_bilateral_3d_grayscale():
img = np.ones((50, 50, 3))
with testing.raises(ValueError):
restoration.denoise_bilateral(img, multichannel=False)
def test_denoise_bilateral_3d_multichannel():
img = np.ones((50, 50, 50))
with expected_warnings(["grayscale"]):
result = restoration.denoise_bilateral(img, multichannel=True)
assert_equal(result, img)
def test_denoise_bilateral_multidimensional():
img = np.ones((10, 10, 10, 10))
with testing.raises(ValueError):
restoration.denoise_bilateral(img, multichannel=False)
with testing.raises(ValueError):
restoration.denoise_bilateral(img, multichannel=True)
def test_denoise_bilateral_nan():
img = np.full((50, 50), np.NaN)
# This is in fact an optional warning for our test suite.
# Python 3.5 will not trigger a warning.
with expected_warnings(['invalid|\A\Z']):
out = restoration.denoise_bilateral(img, multichannel=False)
assert_equal(img, out)
def test_denoise_nl_means_2d():
img = np.zeros((40, 40))
img[10:-10, 10:-10] = 1.
sigma = 0.3
img += sigma * np.random.randn(*img.shape)
for s in [sigma, 0]:
denoised = restoration.denoise_nl_means(img, 7, 5, 0.2, fast_mode=True,
multichannel=True, sigma=s)
# make sure noise is reduced
assert_(img.std() > denoised.std())
denoised = restoration.denoise_nl_means(img, 7, 5, 0.2,
fast_mode=False,
multichannel=True, sigma=s)
# make sure noise is reduced
assert_(img.std() > denoised.std())
def test_denoise_nl_means_2d_multichannel():
# reduce image size because nl means is slow
img = np.copy(astro[:50, :50])
img = np.concatenate((img, ) * 2, ) # 6 channels
# add some random noise
sigma = 0.1
imgn = img + sigma * np.random.standard_normal(img.shape)
imgn = np.clip(imgn, 0, 1)
for fast_mode in [True, False]:
for s in [sigma, 0]:
for n_channels in [2, 3, 6]:
psnr_noisy = compare_psnr(img[..., :n_channels],
imgn[..., :n_channels])
denoised = restoration.denoise_nl_means(imgn[..., :n_channels],
3, 5, h=0.75 * sigma,
fast_mode=fast_mode,
multichannel=True,
sigma=s)
psnr_denoised = compare_psnr(denoised[..., :n_channels],
img[..., :n_channels])
# make sure noise is reduced
assert_(psnr_denoised > psnr_noisy)
def test_denoise_nl_means_3d():
img = np.zeros((12, 12, 8))
img[5:-5, 5:-5, 2:-2] = 1.
sigma = 0.3
imgn = img + sigma * np.random.randn(*img.shape)
psnr_noisy = compare_psnr(img, imgn)
for s in [sigma, 0]:
denoised = restoration.denoise_nl_means(imgn, 3, 4, h=0.75 * sigma,
fast_mode=True,
multichannel=False, sigma=s)
# make sure noise is reduced
assert_(compare_psnr(img, denoised) > psnr_noisy)
denoised = restoration.denoise_nl_means(imgn, 3, 4, h=0.75 * sigma,
fast_mode=False,
multichannel=False, sigma=s)
# make sure noise is reduced
assert_(compare_psnr(img, denoised) > psnr_noisy)
def test_denoise_nl_means_multichannel():
# for true 3D data, 3D denoising is better than denoising as 2D+channels
img = np.zeros((13, 10, 8))
img[6, 4:6, 2:-2] = 1.
sigma = 0.3
imgn = img + sigma * np.random.randn(*img.shape)
denoised_wrong_multichannel = restoration.denoise_nl_means(
imgn, 3, 4, 0.6 * sigma, fast_mode=True, multichannel=True)
denoised_ok_multichannel = restoration.denoise_nl_means(
imgn, 3, 4, 0.6 * sigma, fast_mode=True, multichannel=False)
psnr_wrong = compare_psnr(img, denoised_wrong_multichannel)
psnr_ok = compare_psnr(img, denoised_ok_multichannel)
assert_(psnr_ok > psnr_wrong)
def test_denoise_nl_means_wrong_dimension():
img = np.zeros((5, 5, 5, 5))
with testing.raises(NotImplementedError):
restoration.denoise_nl_means(img, multichannel=True)
def test_no_denoising_for_small_h():
img = np.zeros((40, 40))
img[10:-10, 10:-10] = 1.
img += 0.3*np.random.randn(*img.shape)
# very small h should result in no averaging with other patches
denoised = restoration.denoise_nl_means(img, 7, 5, 0.01, fast_mode=True,
multichannel=True)
assert_(np.allclose(denoised, img))
denoised = restoration.denoise_nl_means(img, 7, 5, 0.01, fast_mode=False,
multichannel=True)
assert_(np.allclose(denoised, img))
def test_wavelet_denoising():
rstate = np.random.RandomState(1234)
# version with one odd-sized dimension
astro_gray_odd = astro_gray[:, :-1]
astro_odd = astro[:, :-1]
for img, multichannel, convert2ycbcr in [(astro_gray, False, False),
(astro_gray_odd, False, False),
(astro_odd, True, False),
(astro_odd, True, True)]:
sigma = 0.1
noisy = img + sigma * rstate.randn(*(img.shape))
noisy = np.clip(noisy, 0, 1)
# Verify that SNR is improved when true sigma is used
denoised = restoration.denoise_wavelet(noisy, sigma=sigma,
multichannel=multichannel,
convert2ycbcr=convert2ycbcr)
psnr_noisy = compare_psnr(img, noisy)
psnr_denoised = compare_psnr(img, denoised)
assert_(psnr_denoised > psnr_noisy)
# Verify that SNR is improved with internally estimated sigma
denoised = restoration.denoise_wavelet(noisy,
multichannel=multichannel,
convert2ycbcr=convert2ycbcr)
psnr_noisy = compare_psnr(img, noisy)
psnr_denoised = compare_psnr(img, denoised)
assert_(psnr_denoised > psnr_noisy)
# SNR is improved less with 1 wavelet level than with the default.
denoised_1 = restoration.denoise_wavelet(noisy,
multichannel=multichannel,
wavelet_levels=1,
convert2ycbcr=convert2ycbcr)
psnr_denoised_1 = compare_psnr(img, denoised_1)
assert_(psnr_denoised > psnr_denoised_1)
assert_(psnr_denoised_1 > psnr_noisy)
# Test changing noise_std (higher threshold, so less energy in signal)
res1 = restoration.denoise_wavelet(noisy, sigma=2*sigma,
multichannel=multichannel)
res2 = restoration.denoise_wavelet(noisy, sigma=sigma,
multichannel=multichannel)
assert_(
|
np.sum(res1**2)
|
numpy.sum
|
import numpy as np
from numpy import ndarray
def alpha_quartz_stiffness():
"""
Returns the stiffness matrix for α-silicon dioxide, as defined in Voigt notation.
:returns: The 6x6 stiffness matrix for α-silicon dioxide as a numpy array.
:rtype: ndarray
"""
C11, C33, C44, C66, C12, C13, C14 = 86.99, 106.39, 58.12, 40.12, 6.75, 12.17, 17.99
C = np.zeros((6, 6))
C[0, :4] = [C11, C12, C13, C14]
C[1, 1:4] = [C11, C13, -C14]
C[2, 2] = C33
C[3, 3] = C44
C[4, 4:6] = [C44, C14]
C[5, 5] = C66
C = C + np.tril(C.T, -1)
C = C * (10 ** 9) # Convert to GPa
return np.array(C)
def transform_stiffness(U, C):
"""Transform a stiffness matrix defined in the grain coordinate system into a stiffness matrix defined in the
sample coordinate system by applying a rotational matrix defined from the grain orientation matrix U.
:param U: Orientation/rotation tensor as a 3x3 numpy array.
:type U: ndarray
:param C: Stiffness matrix as a 6x6 numpy array.
:type C: ndarray
:returns: A stiffness matrix valid in the sample coordinate system as a 6x6 numpy array.
:rtype: ndarray
"""
M = _get_rotation_matrix(U)
C_rot = (M @ C @ np.transpose(M))
return C_rot
def _get_rotation_matrix(U):
"""
Private function for creating the rotational matrices used in :func:`transform_stiffness`. For further information
see <NAME>, *Acoustic fields and waves in solids*.
:param U: Orientation/rotation tensor as a 3x3 numpy array.
:type U: ndarray
:return: Rotational matrix as a 6x6 numpy array.
:rtype: ndarray
"""
M = np.array([[U[0, 0] ** 2, U[0, 1] ** 2, U[0, 2] ** 2, 2 * U[0, 1] * U[0, 2], 2 * U[0, 2] * U[0, 0],
2 * U[0, 0] * U[0, 1]],
[U[1, 0] ** 2, U[1, 1] ** 2, U[1, 2] ** 2, 2 * U[1, 1] * U[1, 2], 2 * U[1, 2] * U[1, 0],
2 * U[1, 0] * U[1, 1]],
[U[2, 0] ** 2, U[2, 1] ** 2, U[2, 2] ** 2, 2 * U[2, 1] * U[2, 2], 2 * U[2, 2] * U[2, 0],
2 * U[2, 0] * U[2, 1]],
[U[1, 0] * U[2, 0], U[1, 1] * U[2, 1], U[1, 2] * U[2, 2], U[1, 1] * U[2, 2] + U[1, 2] * U[2, 1],
U[1, 0] * U[2, 2] + U[1, 2] * U[2, 0], U[1, 1] * U[2, 0] + U[1, 0] * U[2, 1]],
[U[2, 0] * U[0, 0], U[2, 1] * U[0, 1], U[2, 2] * U[0, 2], U[0, 1] * U[2, 2] + U[0, 2] * U[2, 1],
U[0, 2] * U[2, 0] + U[0, 0] * U[2, 2], U[0, 0] * U[2, 1] + U[0, 1] * U[2, 0]],
[U[0, 0] * U[1, 0], U[0, 1] * U[1, 1], U[0, 2] * U[1, 2], U[0, 1] * U[1, 2] + U[0, 2] * U[1, 1],
U[0, 2] * U[1, 0] + U[0, 0] * U[1, 2], U[0, 0] * U[1, 1] + U[0, 1] * U[1, 0]]])
return M
def calculate_stress_by_matrix_rotation(wlsq_strain, U):
"""
Calculate the intra-granular stresses by applying the sample system stiffnes matrix for a given grain and z-slice
to the corresponding strains.
:param wlsq_strain: Strains as a list of numpy arrays, where each list contains a strain component. The order of
the strains are ["XX", "YY", "ZZ", "YZ", "XZ", "XY"].
:type wlsq_strain: list[ndarray]
:param U: The orientation matrix for a given grain and z-slice.
:type U: ndarray
:return: The intra-granular stresses in the same format as the provided intragranular strains.
:rtype: list[ndarray]
"""
# Get the stiffness matrix as measured in the grain coordinate system
C = alpha_quartz_stiffness()
# Rotate the stiffness matrix by the grain orientation matrix
C = transform_stiffness(U, C)
# Stack the strain vectors into a matrix, where each row contains the strain components for a certain element in
# the mesh which the stress will be plotted on. Make an empty matrix for the stress vectors.
strain_mat = np.column_stack(wlsq_strain)
stress_mat = np.zeros_like(strain_mat)
# Exract a row from the strain matrix, multiply the shear strain components by 2 to obtain the engineeering shear
# strain which is compatible with the Voigt notation.
for i in range(np.size(strain_mat, 0)):
strain_vector = strain_mat[i, :]
strain_vector[3:6] *= 2
# Apply the stiffness matrix to get the stress vectors and stack the stress vectors in a matrix.
stress_mat[i, :] = C @ strain_vector
# Split the stress matrix to give it the same format as wlsq_strains.
wlsq_stress = np.hsplit(stress_mat, 6)
for i, arr in enumerate(wlsq_stress):
wlsq_stress[i] = arr.reshape((-1))
return wlsq_stress
def calculate_stress_by_vector_rotation(wlsq_strain, U):
"""
Calculate the intra-granular stresses by converting the strains into a 3x3 tensor format and using the grain
orientation matrix to calculate the strains in the grain coordiante system. The stiffness matrix is then applied
in the grain coordinate system and the resulting stresses are transformed back to the sample coordiante system.
:param wlsq_strain: Strains as a list of numpy arrays, where each list contains a strain component. The order of
the strains are ["XX", "YY", "ZZ", "YZ", "XZ", "XY"].
:type wlsq_strain: list[ndarray]
:param U: The orientation matrix for a given grain and z-slice.
:type U: ndarray
:return: The intra-granular stresses in the same format as the provided intragranular strains.
:rtype: list[ndarray]
"""
# Here the stresses will be calculated using a different method where the strain vectors are rotated into the grain
# coordinate system where the stiffness matrix is applied and then the corresponding strain vector is rotated back
# by solving a system of equations.
# Get the stiffness matrix as measured in the grain coordinate system
C = alpha_quartz_stiffness()
# Stack the strain vectors into a matrix, where each row contains the strain components for a certain element in
# the mesh which the stress will be plotted on. Make an empty matrix for the stress vectors.
strain_mat = np.column_stack(wlsq_strain)
stress_mat = np.zeros_like(strain_mat)
for i in range(np.size(strain_mat, 0)):
strain_vector = strain_mat[i, :]
# Transform the strain_vector to the grain coordinate system.
strain_tensor = vec_to_tens(strain_vector)
grain_strain_tensor = U.T @ strain_tensor @ U
# Convert the grain strain vector to Voigt notation.
grain_strain_vector = tens_to_vec(grain_strain_tensor)
grain_strain_vector[3:6] *= 2
# Calculate the stress in the grain coordinate system and apply the U matrix to transform the strain back to
# the sample coordinate system.
grain_stress_vector = C @ grain_strain_vector
# Convert the stress vector to the sample coordinate system.
grain_stess_tensor = vec_to_tens(grain_stress_vector)
sample_stress_tensor = U @ grain_stess_tensor @ U.T
sample_stress_vector = tens_to_vec(sample_stress_tensor)
stress_mat[i, :] = sample_stress_vector
# Split the stress matrix to give it the same format as wlsq_strains.
wlsq_stress = np.hsplit(stress_mat, 6)
for i, arr in enumerate(wlsq_stress):
wlsq_stress[i] = arr.reshape((-1))
return wlsq_stress
def calc_principal_stresses(wlsq_stress):
"""
Calculate the principal stresses by solving the eigenvalue problem for each reconstructed strain tensor.
The result is a list of numpy arrays where each numpy array contains one of the principal stress components.
:param wlsq_stress: Stresses as a list of numpy arrays, where each list contains a stress component. The order of
the stresses should be ["XX", "YY", "ZZ", "YZ", "XZ", "XY"].
:type wlsq_stress: list[ndarray]
:return: The principal stresses as a list of numpy arrays. The order of the principal stresses is
["σ_1", "σ_2", "σ_3"], where σ_1 corresponds to the largest tensile stress and σ_3 correspond to the largest
compressive stress.
:rtype: list[ndarray]
"""
stress = np.column_stack(wlsq_stress)
nrows = np.size(stress, 0)
principal_stresses = np.zeros((nrows, 3))
for i in range(nrows):
sigma = vec_to_tens(stress[i, :])
eigenvals, eigenvects =
|
np.linalg.eig(sigma)
|
numpy.linalg.eig
|
# coding: utf-8
# # Domain Adaptation between digits
#
# #### *<NAME>, <NAME>*
#
# In this practical session we will apply on digit classification the OT based domain adaptation method proposed in
#
# <NAME>, <NAME>, <NAME>, <NAME>, "[Optimal transport for domain adaptation](http://remi.flamary.com/biblio/courty2016optimal.pdf)", Pattern Analysis and Machine Intelligence, IEEE Transactions on , 2016.
#
# 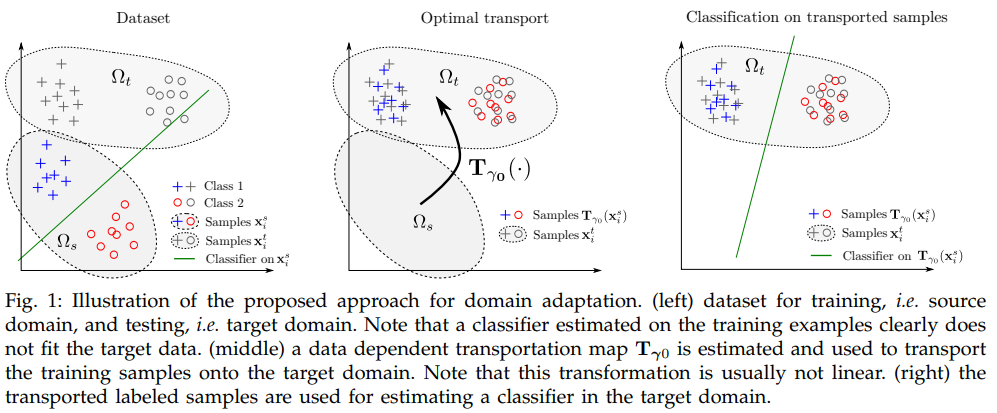
#
# To this end we will try and adapt between the MNIST and USPS datasets. Since those datasets do not have the same resolution (28x28 and 16x16 for MNSIT and USPS) we perform a zeros padding of the USPS digits
#
#
# #### Import modules
#
# First we import the relevant modules. Note that you will need ```sklearn``` to learn the Support Vector Machine cleassifier and to projet the data with TSNE.
#
# In[3]:
import numpy as np # always need it
import pylab as pl # do the plots
from sklearn.svm import SVC
from sklearn.manifold import TSNE
import ot
# ### Loading data and normalization
#
# We load the data in memory and perform a normalization of the images so that they all sum to 1.
#
# Note that every line in the ```xs``` and ```xt``` is a 28x28 image.
# In[4]:
data=np.load('data/mnist_usps.npz')
xs,ys=data['xs'],data['ys']
xt,yt=data['xt'],data['yt']
# normalization
xs=xs/xs.sum(1,keepdims=True) # every l
xt=xt/xt.sum(1,keepdims=True)
ns=xs.shape[0]
nt=xt.shape[0]
# ### Vizualizing Source (MNIST) and Target (USPS) datasets
#
#
#
#
# In[5]:
# function for plotting images
def plot_image(x):
pl.imshow(x.reshape((28,28)),cmap='gray')
pl.xticks(())
pl.yticks(())
nb=10
# Fisrt we plot MNIST
pl.figure(1,(nb,nb))
for i in range(nb*nb):
pl.subplot(nb,nb,1+i)
c=i%nb
plot_image(xs[np.where(ys==c)[0][i//nb],:])
pl.gcf().suptitle("MNIST", fontsize=20);
pl.gcf().subplots_adjust(top=0.95)
# Then we plot USPS
pl.figure(2,(nb,nb))
for i in range(nb*nb):
pl.subplot(nb,nb,1+i)
c=i%nb
plot_image(xt[
|
np.where(yt==c)
|
numpy.where
|
# -*- coding: utf-8 -*-
"""
docstring goes here.
:copyright: Copyright 2014 by the Elephant team, see AUTHORS.txt.
:license: Modified BSD, see LICENSE.txt for details.
"""
from __future__ import division, print_function
import unittest
from itertools import chain
from neo.test.generate_datasets import fake_neo
import numpy as np
from numpy.testing.utils import assert_array_equal
import quantities as pq
try:
import pandas as pd
from pandas.util.testing import assert_frame_equal, assert_index_equal
except ImportError:
HAVE_PANDAS = False
else:
import elephant.pandas_bridge as ep
HAVE_PANDAS = True
@unittest.skipUnless(HAVE_PANDAS, 'requires pandas')
class MultiindexFromDictTestCase(unittest.TestCase):
def test__multiindex_from_dict(self):
inds = {'test1': 6.5,
'test2': 5,
'test3': 'test'}
targ = pd.MultiIndex(levels=[[6.5], [5], ['test']],
labels=[[0], [0], [0]],
names=['test1', 'test2', 'test3'])
res0 = ep._multiindex_from_dict(inds)
self.assertEqual(targ.levels, res0.levels)
self.assertEqual(targ.names, res0.names)
self.assertEqual(targ.labels, res0.labels)
def _convert_levels(levels):
"""Convert a list of levels to the format pandas returns for a MultiIndex.
Parameters
----------
levels : list
The list of levels to convert.
Returns
-------
list
The the level in `list` converted to values like what pandas will give.
"""
levels = list(levels)
for i, level in enumerate(levels):
if hasattr(level, 'lower'):
try:
level = unicode(level)
except NameError:
pass
elif hasattr(level, 'date'):
levels[i] = pd.DatetimeIndex(data=[level])
continue
elif level is None:
levels[i] = pd.Index([])
continue
levels[i] = pd.Index([level])
return levels
@unittest.skipUnless(HAVE_PANDAS, 'requires pandas')
class ConvertValueSafeTestCase(unittest.TestCase):
def test__convert_value_safe__float(self):
targ = 5.5
value = targ
res = ep._convert_value_safe(value)
self.assertIs(res, targ)
def test__convert_value_safe__str(self):
targ = 'test'
value = targ
res = ep._convert_value_safe(value)
self.assertIs(res, targ)
def test__convert_value_safe__bytes(self):
targ = 'test'
value = b'test'
res = ep._convert_value_safe(value)
self.assertEqual(res, targ)
def test__convert_value_safe__numpy_int_scalar(self):
targ = 5
value = np.array(5)
res = ep._convert_value_safe(value)
self.assertEqual(res, targ)
self.assertFalse(hasattr(res, 'dtype'))
def test__convert_value_safe__numpy_float_scalar(self):
targ = 5.
value = np.array(5.)
res = ep._convert_value_safe(value)
self.assertEqual(res, targ)
self.assertFalse(hasattr(res, 'dtype'))
def test__convert_value_safe__numpy_unicode_scalar(self):
targ = u'test'
value = np.array('test', dtype='U')
res = ep._convert_value_safe(value)
self.assertEqual(res, targ)
self.assertFalse(hasattr(res, 'dtype'))
def test__convert_value_safe__numpy_str_scalar(self):
targ = u'test'
value = np.array('test', dtype='S')
res = ep._convert_value_safe(value)
self.assertEqual(res, targ)
self.assertFalse(hasattr(res, 'dtype'))
def test__convert_value_safe__quantity_scalar(self):
targ = (10., 'ms')
value = 10. * pq.ms
res = ep._convert_value_safe(value)
self.assertEqual(res, targ)
self.assertFalse(hasattr(res[0], 'dtype'))
self.assertFalse(hasattr(res[0], 'units'))
@unittest.skipUnless(HAVE_PANDAS, 'requires pandas')
class SpiketrainToDataframeTestCase(unittest.TestCase):
def test__spiketrain_to_dataframe__parents_empty(self):
obj = fake_neo('SpikeTrain', seed=0)
res0 = ep.spiketrain_to_dataframe(obj)
res1 = ep.spiketrain_to_dataframe(obj, child_first=True)
res2 = ep.spiketrain_to_dataframe(obj, child_first=False)
res3 = ep.spiketrain_to_dataframe(obj, parents=True)
res4 = ep.spiketrain_to_dataframe(obj, parents=True,
child_first=True)
res5 = ep.spiketrain_to_dataframe(obj, parents=True,
child_first=False)
res6 = ep.spiketrain_to_dataframe(obj, parents=False)
res7 = ep.spiketrain_to_dataframe(obj, parents=False, child_first=True)
res8 = ep.spiketrain_to_dataframe(obj, parents=False,
child_first=False)
targvalues = pq.Quantity(obj.magnitude, units=obj.units)
targvalues = targvalues.rescale('s').magnitude[np.newaxis].T
targindex = np.arange(len(targvalues))
attrs = ep._extract_neo_attrs_safe(obj, parents=True, child_first=True)
keys, values = zip(*sorted(attrs.items()))
values = _convert_levels(values)
self.assertEqual(1, len(res0.columns))
self.assertEqual(1, len(res1.columns))
self.assertEqual(1, len(res2.columns))
self.assertEqual(1, len(res3.columns))
self.assertEqual(1, len(res4.columns))
self.assertEqual(1, len(res5.columns))
self.assertEqual(1, len(res6.columns))
self.assertEqual(1, len(res7.columns))
self.assertEqual(1, len(res8.columns))
self.assertEqual(len(obj), len(res0.index))
self.assertEqual(len(obj), len(res1.index))
self.assertEqual(len(obj), len(res2.index))
self.assertEqual(len(obj), len(res3.index))
self.assertEqual(len(obj), len(res4.index))
self.assertEqual(len(obj), len(res5.index))
self.assertEqual(len(obj), len(res6.index))
self.assertEqual(len(obj), len(res7.index))
self.assertEqual(len(obj), len(res8.index))
assert_array_equal(targvalues, res0.values)
assert_array_equal(targvalues, res1.values)
assert_array_equal(targvalues, res2.values)
assert_array_equal(targvalues, res3.values)
assert_array_equal(targvalues, res4.values)
assert_array_equal(targvalues, res5.values)
assert_array_equal(targvalues, res6.values)
assert_array_equal(targvalues, res7.values)
assert_array_equal(targvalues, res8.values)
assert_array_equal(targindex, res0.index)
assert_array_equal(targindex, res1.index)
assert_array_equal(targindex, res2.index)
assert_array_equal(targindex, res3.index)
assert_array_equal(targindex, res4.index)
assert_array_equal(targindex, res5.index)
assert_array_equal(targindex, res6.index)
assert_array_equal(targindex, res7.index)
assert_array_equal(targindex, res8.index)
self.assertEqual(['spike_number'], res0.index.names)
self.assertEqual(['spike_number'], res1.index.names)
self.assertEqual(['spike_number'], res2.index.names)
self.assertEqual(['spike_number'], res3.index.names)
self.assertEqual(['spike_number'], res4.index.names)
self.assertEqual(['spike_number'], res5.index.names)
self.assertEqual(['spike_number'], res6.index.names)
self.assertEqual(['spike_number'], res7.index.names)
self.assertEqual(['spike_number'], res8.index.names)
self.assertEqual(keys, res0.columns.names)
self.assertEqual(keys, res1.columns.names)
self.assertEqual(keys, res2.columns.names)
self.assertEqual(keys, res3.columns.names)
self.assertEqual(keys, res4.columns.names)
self.assertEqual(keys, res5.columns.names)
self.assertEqual(keys, res6.columns.names)
self.assertEqual(keys, res7.columns.names)
self.assertEqual(keys, res8.columns.names)
for value, level in zip(values, res0.columns.levels):
assert_index_equal(value, level)
for value, level in zip(values, res1.columns.levels):
assert_index_equal(value, level)
for value, level in zip(values, res2.columns.levels):
assert_index_equal(value, level)
for value, level in zip(values, res3.columns.levels):
assert_index_equal(value, level)
for value, level in zip(values, res4.columns.levels):
assert_index_equal(value, level)
for value, level in zip(values, res5.columns.levels):
assert_index_equal(value, level)
for value, level in zip(values, res6.columns.levels):
assert_index_equal(value, level)
for value, level in zip(values, res7.columns.levels):
assert_index_equal(value, level)
for value, level in zip(values, res8.columns.levels):
assert_index_equal(value, level)
def test__spiketrain_to_dataframe__noparents(self):
blk = fake_neo('Block', seed=0)
obj = blk.list_children_by_class('SpikeTrain')[0]
res0 = ep.spiketrain_to_dataframe(obj, parents=False)
res1 = ep.spiketrain_to_dataframe(obj, parents=False,
child_first=True)
res2 = ep.spiketrain_to_dataframe(obj, parents=False,
child_first=False)
targvalues = pq.Quantity(obj.magnitude, units=obj.units)
targvalues = targvalues.rescale('s').magnitude[np.newaxis].T
targindex = np.arange(len(targvalues))
attrs = ep._extract_neo_attrs_safe(obj, parents=False,
child_first=True)
keys, values = zip(*sorted(attrs.items()))
values = _convert_levels(values)
self.assertEqual(1, len(res0.columns))
self.assertEqual(1, len(res1.columns))
self.assertEqual(1, len(res2.columns))
self.assertEqual(len(obj), len(res0.index))
self.assertEqual(len(obj), len(res1.index))
self.assertEqual(len(obj), len(res2.index))
assert_array_equal(targvalues, res0.values)
assert_array_equal(targvalues, res1.values)
assert_array_equal(targvalues, res2.values)
assert_array_equal(targindex, res0.index)
assert_array_equal(targindex, res1.index)
assert_array_equal(targindex, res2.index)
self.assertEqual(['spike_number'], res0.index.names)
self.assertEqual(['spike_number'], res1.index.names)
self.assertEqual(['spike_number'], res2.index.names)
self.assertEqual(keys, res0.columns.names)
self.assertEqual(keys, res1.columns.names)
self.assertEqual(keys, res2.columns.names)
for value, level in zip(values, res0.columns.levels):
assert_index_equal(value, level)
for value, level in zip(values, res1.columns.levels):
assert_index_equal(value, level)
for value, level in zip(values, res2.columns.levels):
assert_index_equal(value, level)
def test__spiketrain_to_dataframe__parents_childfirst(self):
blk = fake_neo('Block', seed=0)
obj = blk.list_children_by_class('SpikeTrain')[0]
res0 = ep.spiketrain_to_dataframe(obj)
res1 = ep.spiketrain_to_dataframe(obj, child_first=True)
res2 = ep.spiketrain_to_dataframe(obj, parents=True)
res3 = ep.spiketrain_to_dataframe(obj, parents=True, child_first=True)
targvalues = pq.Quantity(obj.magnitude, units=obj.units)
targvalues = targvalues.rescale('s').magnitude[np.newaxis].T
targindex = np.arange(len(targvalues))
attrs = ep._extract_neo_attrs_safe(obj, parents=True, child_first=True)
keys, values = zip(*sorted(attrs.items()))
values = _convert_levels(values)
self.assertEqual(1, len(res0.columns))
self.assertEqual(1, len(res1.columns))
self.assertEqual(1, len(res2.columns))
self.assertEqual(1, len(res3.columns))
self.assertEqual(len(obj), len(res0.index))
self.assertEqual(len(obj), len(res1.index))
self.assertEqual(len(obj), len(res2.index))
self.assertEqual(len(obj), len(res3.index))
assert_array_equal(targvalues, res0.values)
assert_array_equal(targvalues, res1.values)
assert_array_equal(targvalues, res2.values)
assert_array_equal(targvalues, res3.values)
assert_array_equal(targindex, res0.index)
assert_array_equal(targindex, res1.index)
assert_array_equal(targindex, res2.index)
assert_array_equal(targindex, res3.index)
self.assertEqual(['spike_number'], res0.index.names)
self.assertEqual(['spike_number'], res1.index.names)
self.assertEqual(['spike_number'], res2.index.names)
self.assertEqual(['spike_number'], res3.index.names)
self.assertEqual(keys, res0.columns.names)
self.assertEqual(keys, res1.columns.names)
self.assertEqual(keys, res2.columns.names)
self.assertEqual(keys, res3.columns.names)
for value, level in zip(values, res0.columns.levels):
assert_index_equal(value, level)
for value, level in zip(values, res1.columns.levels):
assert_index_equal(value, level)
for value, level in zip(values, res2.columns.levels):
assert_index_equal(value, level)
for value, level in zip(values, res3.columns.levels):
assert_index_equal(value, level)
def test__spiketrain_to_dataframe__parents_parentfirst(self):
blk = fake_neo('Block', seed=0)
obj = blk.list_children_by_class('SpikeTrain')[0]
res0 = ep.spiketrain_to_dataframe(obj, child_first=False)
res1 = ep.spiketrain_to_dataframe(obj, parents=True, child_first=False)
targvalues = pq.Quantity(obj.magnitude, units=obj.units)
targvalues = targvalues.rescale('s').magnitude[np.newaxis].T
targindex = np.arange(len(targvalues))
attrs = ep._extract_neo_attrs_safe(obj, parents=True,
child_first=False)
keys, values = zip(*sorted(attrs.items()))
values = _convert_levels(values)
self.assertEqual(1, len(res0.columns))
self.assertEqual(1, len(res1.columns))
self.assertEqual(len(obj), len(res0.index))
self.assertEqual(len(obj), len(res1.index))
assert_array_equal(targvalues, res0.values)
assert_array_equal(targvalues, res1.values)
assert_array_equal(targindex, res0.index)
assert_array_equal(targindex, res1.index)
self.assertEqual(['spike_number'], res0.index.names)
self.assertEqual(['spike_number'], res1.index.names)
self.assertEqual(keys, res0.columns.names)
self.assertEqual(keys, res1.columns.names)
for value, level in zip(values, res0.columns.levels):
assert_index_equal(value, level)
for value, level in zip(values, res1.columns.levels):
assert_index_equal(value, level)
@unittest.skipUnless(HAVE_PANDAS, 'requires pandas')
class EventToDataframeTestCase(unittest.TestCase):
def test__event_to_dataframe__parents_empty(self):
obj = fake_neo('Event', seed=42)
res0 = ep.event_to_dataframe(obj)
res1 = ep.event_to_dataframe(obj, child_first=True)
res2 = ep.event_to_dataframe(obj, child_first=False)
res3 = ep.event_to_dataframe(obj, parents=True)
res4 = ep.event_to_dataframe(obj, parents=True, child_first=True)
res5 = ep.event_to_dataframe(obj, parents=True, child_first=False)
res6 = ep.event_to_dataframe(obj, parents=False)
res7 = ep.event_to_dataframe(obj, parents=False, child_first=True)
res8 = ep.event_to_dataframe(obj, parents=False, child_first=False)
targvalues = obj.labels[:len(obj.times)][np.newaxis].T.astype('U')
targindex = obj.times[:len(obj.labels)].rescale('s').magnitude
attrs = ep._extract_neo_attrs_safe(obj, parents=True, child_first=True)
keys, values = zip(*sorted(attrs.items()))
values = _convert_levels(values)
self.assertEqual(1, len(res0.columns))
self.assertEqual(1, len(res1.columns))
self.assertEqual(1, len(res2.columns))
self.assertEqual(1, len(res3.columns))
self.assertEqual(1, len(res4.columns))
self.assertEqual(1, len(res5.columns))
self.assertEqual(1, len(res6.columns))
self.assertEqual(1, len(res7.columns))
self.assertEqual(1, len(res8.columns))
self.assertEqual(min(len(obj.times), len(obj.labels)),
len(res0.index))
self.assertEqual(min(len(obj.times), len(obj.labels)),
len(res1.index))
self.assertEqual(min(len(obj.times), len(obj.labels)),
len(res2.index))
self.assertEqual(min(len(obj.times), len(obj.labels)),
len(res3.index))
self.assertEqual(min(len(obj.times), len(obj.labels)),
len(res4.index))
self.assertEqual(min(len(obj.times), len(obj.labels)),
len(res5.index))
self.assertEqual(min(len(obj.times), len(obj.labels)),
len(res6.index))
self.assertEqual(min(len(obj.times), len(obj.labels)),
len(res7.index))
self.assertEqual(min(len(obj.times), len(obj.labels)),
len(res8.index))
assert_array_equal(targvalues, res0.values)
assert_array_equal(targvalues, res1.values)
assert_array_equal(targvalues, res2.values)
assert_array_equal(targvalues, res3.values)
assert_array_equal(targvalues, res4.values)
assert_array_equal(targvalues, res5.values)
assert_array_equal(targvalues, res6.values)
assert_array_equal(targvalues, res7.values)
assert_array_equal(targvalues, res8.values)
assert_array_equal(targindex, res0.index)
assert_array_equal(targindex, res1.index)
assert_array_equal(targindex, res2.index)
assert_array_equal(targindex, res3.index)
assert_array_equal(targindex, res4.index)
assert_array_equal(targindex, res5.index)
assert_array_equal(targindex, res6.index)
assert_array_equal(targindex, res7.index)
assert_array_equal(targindex, res8.index)
self.assertEqual(['times'], res0.index.names)
self.assertEqual(['times'], res1.index.names)
self.assertEqual(['times'], res2.index.names)
self.assertEqual(['times'], res3.index.names)
self.assertEqual(['times'], res4.index.names)
self.assertEqual(['times'], res5.index.names)
self.assertEqual(['times'], res6.index.names)
self.assertEqual(['times'], res7.index.names)
self.assertEqual(['times'], res8.index.names)
self.assertEqual(keys, res0.columns.names)
self.assertEqual(keys, res1.columns.names)
self.assertEqual(keys, res2.columns.names)
self.assertEqual(keys, res3.columns.names)
self.assertEqual(keys, res4.columns.names)
self.assertEqual(keys, res5.columns.names)
self.assertEqual(keys, res6.columns.names)
self.assertEqual(keys, res7.columns.names)
self.assertEqual(keys, res8.columns.names)
for value, level in zip(values, res0.columns.levels):
assert_index_equal(value, level)
for value, level in zip(values, res1.columns.levels):
assert_index_equal(value, level)
for value, level in zip(values, res2.columns.levels):
assert_index_equal(value, level)
for value, level in zip(values, res3.columns.levels):
assert_index_equal(value, level)
for value, level in zip(values, res4.columns.levels):
assert_index_equal(value, level)
for value, level in zip(values, res5.columns.levels):
assert_index_equal(value, level)
for value, level in zip(values, res6.columns.levels):
assert_index_equal(value, level)
for value, level in zip(values, res7.columns.levels):
assert_index_equal(value, level)
for value, level in zip(values, res8.columns.levels):
assert_index_equal(value, level)
def test__event_to_dataframe__noparents(self):
blk = fake_neo('Block', seed=42)
obj = blk.list_children_by_class('Event')[0]
res0 = ep.event_to_dataframe(obj, parents=False)
res1 = ep.event_to_dataframe(obj, parents=False, child_first=False)
res2 = ep.event_to_dataframe(obj, parents=False, child_first=True)
targvalues = obj.labels[:len(obj.times)][np.newaxis].T.astype('U')
targindex = obj.times[:len(obj.labels)].rescale('s').magnitude
attrs = ep._extract_neo_attrs_safe(obj, parents=False,
child_first=True)
keys, values = zip(*sorted(attrs.items()))
values = _convert_levels(values)
self.assertEqual(1, len(res0.columns))
self.assertEqual(1, len(res1.columns))
self.assertEqual(1, len(res2.columns))
self.assertEqual(min(len(obj.times), len(obj.labels)),
len(res0.index))
self.assertEqual(min(len(obj.times), len(obj.labels)),
len(res1.index))
self.assertEqual(min(len(obj.times), len(obj.labels)),
len(res2.index))
assert_array_equal(targvalues, res0.values)
assert_array_equal(targvalues, res1.values)
assert_array_equal(targvalues, res2.values)
assert_array_equal(targindex, res0.index)
assert_array_equal(targindex, res1.index)
assert_array_equal(targindex, res2.index)
self.assertEqual(['times'], res0.index.names)
self.assertEqual(['times'], res1.index.names)
self.assertEqual(['times'], res2.index.names)
self.assertEqual(keys, res0.columns.names)
self.assertEqual(keys, res1.columns.names)
self.assertEqual(keys, res2.columns.names)
for value, level in zip(values, res0.columns.levels):
assert_index_equal(value, level)
for value, level in zip(values, res1.columns.levels):
assert_index_equal(value, level)
for value, level in zip(values, res2.columns.levels):
assert_index_equal(value, level)
def test__event_to_dataframe__parents_childfirst(self):
blk = fake_neo('Block', seed=42)
obj = blk.list_children_by_class('Event')[0]
res0 = ep.event_to_dataframe(obj)
res1 = ep.event_to_dataframe(obj, child_first=True)
res2 = ep.event_to_dataframe(obj, parents=True)
res3 = ep.event_to_dataframe(obj, parents=True, child_first=True)
targvalues = obj.labels[:len(obj.times)][np.newaxis].T.astype('U')
targindex = obj.times[:len(obj.labels)].rescale('s').magnitude
attrs = ep._extract_neo_attrs_safe(obj, parents=True, child_first=True)
keys, values = zip(*sorted(attrs.items()))
values = _convert_levels(values)
self.assertEqual(1, len(res0.columns))
self.assertEqual(1, len(res1.columns))
self.assertEqual(1, len(res2.columns))
self.assertEqual(1, len(res3.columns))
self.assertEqual(min(len(obj.times), len(obj.labels)),
len(res0.index))
self.assertEqual(min(len(obj.times), len(obj.labels)),
len(res1.index))
self.assertEqual(min(len(obj.times), len(obj.labels)),
len(res2.index))
self.assertEqual(min(len(obj.times), len(obj.labels)),
len(res3.index))
assert_array_equal(targvalues, res0.values)
assert_array_equal(targvalues, res1.values)
assert_array_equal(targvalues, res2.values)
assert_array_equal(targvalues, res3.values)
assert_array_equal(targindex, res0.index)
assert_array_equal(targindex, res1.index)
assert_array_equal(targindex, res2.index)
assert_array_equal(targindex, res3.index)
self.assertEqual(['times'], res0.index.names)
self.assertEqual(['times'], res1.index.names)
self.assertEqual(['times'], res2.index.names)
self.assertEqual(['times'], res3.index.names)
self.assertEqual(keys, res0.columns.names)
self.assertEqual(keys, res1.columns.names)
self.assertEqual(keys, res2.columns.names)
self.assertEqual(keys, res3.columns.names)
for value, level in zip(values, res0.columns.levels):
assert_index_equal(value, level)
for value, level in zip(values, res1.columns.levels):
assert_index_equal(value, level)
for value, level in zip(values, res2.columns.levels):
assert_index_equal(value, level)
for value, level in zip(values, res3.columns.levels):
assert_index_equal(value, level)
def test__event_to_dataframe__parents_parentfirst(self):
blk = fake_neo('Block', seed=42)
obj = blk.list_children_by_class('Event')[0]
res0 = ep.event_to_dataframe(obj, child_first=False)
res1 = ep.event_to_dataframe(obj, parents=True, child_first=False)
targvalues = obj.labels[:len(obj.times)][np.newaxis].T.astype('U')
targindex = obj.times[:len(obj.labels)].rescale('s').magnitude
attrs = ep._extract_neo_attrs_safe(obj, parents=True,
child_first=False)
keys, values = zip(*sorted(attrs.items()))
values = _convert_levels(values)
self.assertEqual(1, len(res0.columns))
self.assertEqual(1, len(res1.columns))
self.assertEqual(min(len(obj.times), len(obj.labels)),
len(res0.index))
self.assertEqual(min(len(obj.times), len(obj.labels)),
len(res1.index))
assert_array_equal(targvalues, res0.values)
assert_array_equal(targvalues, res1.values)
assert_array_equal(targindex, res0.index)
assert_array_equal(targindex, res1.index)
self.assertEqual(['times'], res0.index.names)
self.assertEqual(['times'], res1.index.names)
self.assertEqual(keys, res0.columns.names)
self.assertEqual(keys, res1.columns.names)
for value, level in zip(values, res0.columns.levels):
assert_index_equal(value, level)
for value, level in zip(values, res1.columns.levels):
assert_index_equal(value, level)
@unittest.skipUnless(HAVE_PANDAS, 'requires pandas')
class EpochToDataframeTestCase(unittest.TestCase):
def test__epoch_to_dataframe__parents_empty(self):
obj = fake_neo('Epoch', seed=42)
res0 = ep.epoch_to_dataframe(obj)
res1 = ep.epoch_to_dataframe(obj, child_first=True)
res2 = ep.epoch_to_dataframe(obj, child_first=False)
res3 = ep.epoch_to_dataframe(obj, parents=True)
res4 = ep.epoch_to_dataframe(obj, parents=True, child_first=True)
res5 = ep.epoch_to_dataframe(obj, parents=True, child_first=False)
res6 = ep.epoch_to_dataframe(obj, parents=False)
res7 = ep.epoch_to_dataframe(obj, parents=False, child_first=True)
res8 = ep.epoch_to_dataframe(obj, parents=False, child_first=False)
minlen = min([len(obj.times), len(obj.durations), len(obj.labels)])
targvalues = obj.labels[:minlen][np.newaxis].T.astype('U')
targindex = np.vstack([obj.durations[:minlen].rescale('s').magnitude,
obj.times[:minlen].rescale('s').magnitude])
targvalues = targvalues[targindex.argsort()[0], :]
targindex.sort()
attrs = ep._extract_neo_attrs_safe(obj, parents=True,
child_first=True)
keys, values = zip(*sorted(attrs.items()))
values = _convert_levels(values)
self.assertEqual(1, len(res0.columns))
self.assertEqual(1, len(res1.columns))
self.assertEqual(1, len(res2.columns))
self.assertEqual(1, len(res3.columns))
self.assertEqual(1, len(res4.columns))
self.assertEqual(1, len(res5.columns))
self.assertEqual(1, len(res6.columns))
self.assertEqual(1, len(res7.columns))
self.assertEqual(1, len(res8.columns))
self.assertEqual(min(len(obj.times), len(obj.durations),
len(obj.labels)),
len(res0.index))
self.assertEqual(min(len(obj.times), len(obj.durations),
len(obj.labels)),
len(res1.index))
self.assertEqual(min(len(obj.times), len(obj.durations),
len(obj.labels)),
len(res2.index))
self.assertEqual(min(len(obj.times), len(obj.durations),
len(obj.labels)),
len(res3.index))
self.assertEqual(min(len(obj.times), len(obj.durations),
len(obj.labels)),
len(res4.index))
self.assertEqual(min(len(obj.times), len(obj.durations),
len(obj.labels)),
len(res5.index))
self.assertEqual(min(len(obj.times), len(obj.durations),
len(obj.labels)),
len(res6.index))
self.assertEqual(min(len(obj.times), len(obj.durations),
len(obj.labels)),
len(res7.index))
self.assertEqual(min(len(obj.times), len(obj.durations),
len(obj.labels)),
len(res8.index))
assert_array_equal(targvalues, res0.values)
assert_array_equal(targvalues, res1.values)
assert_array_equal(targvalues, res2.values)
assert_array_equal(targvalues, res3.values)
assert_array_equal(targvalues, res4.values)
assert_array_equal(targvalues, res5.values)
assert_array_equal(targvalues, res6.values)
assert_array_equal(targvalues, res7.values)
assert_array_equal(targvalues, res8.values)
self.assertEqual(keys, res0.columns.names)
self.assertEqual(keys, res1.columns.names)
self.assertEqual(keys, res2.columns.names)
self.assertEqual(keys, res3.columns.names)
self.assertEqual(keys, res4.columns.names)
self.assertEqual(keys, res5.columns.names)
self.assertEqual(keys, res6.columns.names)
self.assertEqual(keys, res7.columns.names)
self.assertEqual(keys, res8.columns.names)
self.assertEqual([u'durations', u'times'], res0.index.names)
self.assertEqual([u'durations', u'times'], res1.index.names)
self.assertEqual([u'durations', u'times'], res2.index.names)
self.assertEqual([u'durations', u'times'], res3.index.names)
self.assertEqual([u'durations', u'times'], res4.index.names)
self.assertEqual([u'durations', u'times'], res5.index.names)
self.assertEqual([u'durations', u'times'], res6.index.names)
self.assertEqual([u'durations', u'times'], res7.index.names)
self.assertEqual([u'durations', u'times'], res8.index.names)
self.assertEqual(2, len(res0.index.levels))
self.assertEqual(2, len(res1.index.levels))
self.assertEqual(2, len(res2.index.levels))
self.assertEqual(2, len(res3.index.levels))
self.assertEqual(2, len(res4.index.levels))
self.assertEqual(2, len(res5.index.levels))
self.assertEqual(2, len(res6.index.levels))
self.assertEqual(2, len(res7.index.levels))
self.assertEqual(2, len(res8.index.levels))
assert_array_equal(targindex, res0.index.levels)
assert_array_equal(targindex, res1.index.levels)
assert_array_equal(targindex, res2.index.levels)
assert_array_equal(targindex, res3.index.levels)
assert_array_equal(targindex, res4.index.levels)
assert_array_equal(targindex, res5.index.levels)
assert_array_equal(targindex, res6.index.levels)
assert_array_equal(targindex, res7.index.levels)
assert_array_equal(targindex, res8.index.levels)
for value, level in zip(values, res0.columns.levels):
assert_index_equal(value, level)
for value, level in zip(values, res1.columns.levels):
assert_index_equal(value, level)
for value, level in zip(values, res2.columns.levels):
assert_index_equal(value, level)
for value, level in zip(values, res3.columns.levels):
assert_index_equal(value, level)
for value, level in zip(values, res4.columns.levels):
assert_index_equal(value, level)
for value, level in zip(values, res5.columns.levels):
assert_index_equal(value, level)
for value, level in zip(values, res6.columns.levels):
assert_index_equal(value, level)
for value, level in zip(values, res7.columns.levels):
assert_index_equal(value, level)
for value, level in zip(values, res8.columns.levels):
assert_index_equal(value, level)
def test__epoch_to_dataframe__noparents(self):
blk = fake_neo('Block', seed=42)
obj = blk.list_children_by_class('Epoch')[0]
res0 = ep.epoch_to_dataframe(obj, parents=False)
res1 = ep.epoch_to_dataframe(obj, parents=False, child_first=True)
res2 = ep.epoch_to_dataframe(obj, parents=False, child_first=False)
minlen = min([len(obj.times), len(obj.durations), len(obj.labels)])
targvalues = obj.labels[:minlen][np.newaxis].T.astype('U')
targindex = np.vstack([obj.durations[:minlen].rescale('s').magnitude,
obj.times[:minlen].rescale('s').magnitude])
targvalues = targvalues[targindex.argsort()[0], :]
targindex.sort()
attrs = ep._extract_neo_attrs_safe(obj, parents=False,
child_first=True)
keys, values = zip(*sorted(attrs.items()))
values = _convert_levels(values)
self.assertEqual(1, len(res0.columns))
self.assertEqual(1, len(res1.columns))
self.assertEqual(1, len(res2.columns))
self.assertEqual(min(len(obj.times), len(obj.durations),
len(obj.labels)),
len(res0.index))
self.assertEqual(min(len(obj.times), len(obj.durations),
len(obj.labels)),
len(res1.index))
self.assertEqual(min(len(obj.times), len(obj.durations),
len(obj.labels)),
len(res2.index))
assert_array_equal(targvalues, res0.values)
assert_array_equal(targvalues, res1.values)
assert_array_equal(targvalues, res2.values)
self.assertEqual(keys, res0.columns.names)
self.assertEqual(keys, res1.columns.names)
self.assertEqual(keys, res2.columns.names)
self.assertEqual([u'durations', u'times'], res0.index.names)
self.assertEqual([u'durations', u'times'], res1.index.names)
self.assertEqual([u'durations', u'times'], res2.index.names)
self.assertEqual(2, len(res0.index.levels))
self.assertEqual(2, len(res1.index.levels))
self.assertEqual(2, len(res2.index.levels))
assert_array_equal(targindex, res0.index.levels)
assert_array_equal(targindex, res1.index.levels)
assert_array_equal(targindex, res2.index.levels)
for value, level in zip(values, res0.columns.levels):
assert_index_equal(value, level)
for value, level in zip(values, res1.columns.levels):
assert_index_equal(value, level)
for value, level in zip(values, res2.columns.levels):
assert_index_equal(value, level)
def test__epoch_to_dataframe__parents_childfirst(self):
blk = fake_neo('Block', seed=42)
obj = blk.list_children_by_class('Epoch')[0]
res0 = ep.epoch_to_dataframe(obj)
res1 = ep.epoch_to_dataframe(obj, child_first=True)
res2 = ep.epoch_to_dataframe(obj, parents=True)
res3 = ep.epoch_to_dataframe(obj, parents=True, child_first=True)
minlen = min([len(obj.times), len(obj.durations), len(obj.labels)])
targvalues = obj.labels[:minlen][np.newaxis].T.astype('U')
targindex = np.vstack([obj.durations[:minlen].rescale('s').magnitude,
obj.times[:minlen].rescale('s').magnitude])
targvalues = targvalues[targindex.argsort()[0], :]
targindex.sort()
attrs = ep._extract_neo_attrs_safe(obj, parents=True, child_first=True)
keys, values = zip(*sorted(attrs.items()))
values = _convert_levels(values)
self.assertEqual(1, len(res0.columns))
self.assertEqual(1, len(res1.columns))
self.assertEqual(1, len(res2.columns))
self.assertEqual(1, len(res3.columns))
self.assertEqual(min(len(obj.times), len(obj.durations),
len(obj.labels)),
len(res0.index))
self.assertEqual(min(len(obj.times), len(obj.durations),
len(obj.labels)),
len(res1.index))
self.assertEqual(min(len(obj.times), len(obj.durations),
len(obj.labels)),
len(res2.index))
self.assertEqual(min(len(obj.times), len(obj.durations),
len(obj.labels)),
len(res3.index))
assert_array_equal(targvalues, res0.values)
assert_array_equal(targvalues, res1.values)
assert_array_equal(targvalues, res2.values)
assert_array_equal(targvalues, res3.values)
self.assertEqual(keys, res0.columns.names)
self.assertEqual(keys, res1.columns.names)
self.assertEqual(keys, res2.columns.names)
self.assertEqual(keys, res3.columns.names)
self.assertEqual([u'durations', u'times'], res0.index.names)
self.assertEqual([u'durations', u'times'], res1.index.names)
self.assertEqual([u'durations', u'times'], res2.index.names)
self.assertEqual([u'durations', u'times'], res3.index.names)
self.assertEqual(2, len(res0.index.levels))
self.assertEqual(2, len(res1.index.levels))
self.assertEqual(2, len(res2.index.levels))
self.assertEqual(2, len(res3.index.levels))
assert_array_equal(targindex, res0.index.levels)
assert_array_equal(targindex, res1.index.levels)
assert_array_equal(targindex, res2.index.levels)
assert_array_equal(targindex, res3.index.levels)
for value, level in zip(values, res0.columns.levels):
assert_index_equal(value, level)
for value, level in zip(values, res1.columns.levels):
assert_index_equal(value, level)
for value, level in zip(values, res2.columns.levels):
assert_index_equal(value, level)
for value, level in zip(values, res3.columns.levels):
assert_index_equal(value, level)
def test__epoch_to_dataframe__parents_parentfirst(self):
blk = fake_neo('Block', seed=42)
obj = blk.list_children_by_class('Epoch')[0]
res0 = ep.epoch_to_dataframe(obj, child_first=False)
res1 = ep.epoch_to_dataframe(obj, parents=True, child_first=False)
minlen = min([len(obj.times), len(obj.durations), len(obj.labels)])
targvalues = obj.labels[:minlen][np.newaxis].T.astype('U')
targindex = np.vstack([obj.durations[:minlen].rescale('s').magnitude,
obj.times[:minlen].rescale('s').magnitude])
targvalues = targvalues[targindex.argsort()[0], :]
targindex.sort()
attrs = ep._extract_neo_attrs_safe(obj, parents=True,
child_first=False)
keys, values = zip(*sorted(attrs.items()))
values = _convert_levels(values)
self.assertEqual(1, len(res0.columns))
self.assertEqual(1, len(res1.columns))
self.assertEqual(min(len(obj.times), len(obj.durations),
len(obj.labels)),
len(res0.index))
self.assertEqual(min(len(obj.times), len(obj.durations),
len(obj.labels)),
len(res1.index))
assert_array_equal(targvalues, res0.values)
assert_array_equal(targvalues, res1.values)
self.assertEqual(keys, res0.columns.names)
self.assertEqual(keys, res1.columns.names)
self.assertEqual([u'durations', u'times'], res0.index.names)
self.assertEqual([u'durations', u'times'], res1.index.names)
self.assertEqual(2, len(res0.index.levels))
self.assertEqual(2, len(res1.index.levels))
assert_array_equal(targindex, res0.index.levels)
assert_array_equal(targindex, res1.index.levels)
for value, level in zip(values, res0.columns.levels):
assert_index_equal(value, level)
for value, level in zip(values, res1.columns.levels):
assert_index_equal(value, level)
@unittest.skipUnless(HAVE_PANDAS, 'requires pandas')
class MultiSpiketrainsToDataframeTestCase(unittest.TestCase):
def setUp(self):
if hasattr(self, 'assertItemsEqual'):
self.assertCountEqual = self.assertItemsEqual
def test__multi_spiketrains_to_dataframe__single(self):
obj = fake_neo('SpikeTrain', seed=0, n=5)
res0 = ep.multi_spiketrains_to_dataframe(obj)
res1 = ep.multi_spiketrains_to_dataframe(obj, parents=False)
res2 = ep.multi_spiketrains_to_dataframe(obj, parents=True)
res3 = ep.multi_spiketrains_to_dataframe(obj, child_first=True)
res4 = ep.multi_spiketrains_to_dataframe(obj, parents=False,
child_first=True)
res5 = ep.multi_spiketrains_to_dataframe(obj, parents=True,
child_first=True)
res6 = ep.multi_spiketrains_to_dataframe(obj, child_first=False)
res7 = ep.multi_spiketrains_to_dataframe(obj, parents=False,
child_first=False)
res8 = ep.multi_spiketrains_to_dataframe(obj, parents=True,
child_first=False)
targ = ep.spiketrain_to_dataframe(obj)
keys = ep._extract_neo_attrs_safe(obj, parents=True,
child_first=True).keys()
keys = list(keys)
targwidth = 1
targlen = len(obj)
self.assertEqual(targwidth, len(targ.columns))
self.assertEqual(targwidth, len(res0.columns))
self.assertEqual(targwidth, len(res1.columns))
self.assertEqual(targwidth, len(res2.columns))
self.assertEqual(targwidth, len(res3.columns))
self.assertEqual(targwidth, len(res4.columns))
self.assertEqual(targwidth, len(res5.columns))
self.assertEqual(targwidth, len(res6.columns))
self.assertEqual(targwidth, len(res7.columns))
self.assertEqual(targwidth, len(res8.columns))
self.assertEqual(targlen, len(targ.index))
self.assertEqual(targlen, len(res0.index))
self.assertEqual(targlen, len(res1.index))
self.assertEqual(targlen, len(res2.index))
self.assertEqual(targlen, len(res3.index))
self.assertEqual(targlen, len(res4.index))
self.assertEqual(targlen, len(res5.index))
self.assertEqual(targlen, len(res6.index))
self.assertEqual(targlen, len(res7.index))
self.assertEqual(targlen, len(res8.index))
self.assertCountEqual(keys, targ.columns.names)
self.assertCountEqual(keys, res0.columns.names)
self.assertCountEqual(keys, res1.columns.names)
self.assertCountEqual(keys, res2.columns.names)
self.assertCountEqual(keys, res3.columns.names)
self.assertCountEqual(keys, res4.columns.names)
self.assertCountEqual(keys, res5.columns.names)
self.assertCountEqual(keys, res6.columns.names)
self.assertCountEqual(keys, res7.columns.names)
self.assertCountEqual(keys, res8.columns.names)
assert_array_equal(targ.values, res0.values)
assert_array_equal(targ.values, res1.values)
assert_array_equal(targ.values, res2.values)
assert_array_equal(targ.values, res3.values)
assert_array_equal(targ.values, res4.values)
assert_array_equal(targ.values, res5.values)
assert_array_equal(targ.values, res6.values)
assert_array_equal(targ.values, res7.values)
assert_array_equal(targ.values, res8.values)
assert_frame_equal(targ, res0)
assert_frame_equal(targ, res0)
assert_frame_equal(targ, res1)
assert_frame_equal(targ, res2)
assert_frame_equal(targ, res3)
assert_frame_equal(targ, res4)
assert_frame_equal(targ, res5)
assert_frame_equal(targ, res6)
assert_frame_equal(targ, res7)
assert_frame_equal(targ, res8)
def test__multi_spiketrains_to_dataframe__unit_default(self):
obj = fake_neo('Unit', seed=0, n=5)
res0 = ep.multi_spiketrains_to_dataframe(obj)
objs = obj.spiketrains
targ = [ep.spiketrain_to_dataframe(iobj) for iobj in objs]
targ = ep._sort_inds(pd.concat(targ, axis=1), axis=1)
keys = ep._extract_neo_attrs_safe(objs[0], parents=True,
child_first=True).keys()
keys = list(keys)
targwidth = len(objs)
targlen = max(len(iobj) for iobj in objs)
self.assertGreater(len(objs), 0)
self.assertEqual(targwidth, len(targ.columns))
self.assertEqual(targwidth, len(res0.columns))
self.assertEqual(targlen, len(targ.index))
self.assertEqual(targlen, len(res0.index))
self.assertCountEqual(keys, targ.columns.names)
self.assertCountEqual(keys, res0.columns.names)
assert_array_equal(targ.values, res0.values)
assert_frame_equal(targ, res0)
def test__multi_spiketrains_to_dataframe__segment_default(self):
obj = fake_neo('Segment', seed=0, n=5)
res0 = ep.multi_spiketrains_to_dataframe(obj)
objs = obj.spiketrains
targ = [ep.spiketrain_to_dataframe(iobj) for iobj in objs]
targ = ep._sort_inds(pd.concat(targ, axis=1), axis=1)
keys = ep._extract_neo_attrs_safe(objs[0], parents=True,
child_first=True).keys()
keys = list(keys)
targwidth = len(objs)
targlen = max(len(iobj) for iobj in objs)
self.assertGreater(len(objs), 0)
self.assertEqual(targwidth, len(targ.columns))
self.assertEqual(targwidth, len(res0.columns))
self.assertEqual(targlen, len(targ.index))
self.assertEqual(targlen, len(res0.index))
self.assertCountEqual(keys, targ.columns.names)
self.assertCountEqual(keys, res0.columns.names)
assert_array_equal(targ.values, res0.values)
assert_frame_equal(targ, res0)
def test__multi_spiketrains_to_dataframe__block_noparents(self):
obj = fake_neo('Block', seed=0, n=3)
res0 = ep.multi_spiketrains_to_dataframe(obj, parents=False)
res1 = ep.multi_spiketrains_to_dataframe(obj, parents=False,
child_first=True)
res2 = ep.multi_spiketrains_to_dataframe(obj, parents=False,
child_first=False)
objs = obj.list_children_by_class('SpikeTrain')
targ = [ep.spiketrain_to_dataframe(iobj,
parents=False, child_first=True)
for iobj in objs]
targ = ep._sort_inds(pd.concat(targ, axis=1), axis=1)
keys = ep._extract_neo_attrs_safe(objs[0], parents=False,
child_first=True).keys()
keys = list(keys)
targwidth = len(objs)
targlen = max(len(iobj) for iobj in objs)
self.assertGreater(len(objs), 0)
self.assertEqual(targwidth, len(targ.columns))
self.assertEqual(targwidth, len(res0.columns))
self.assertEqual(targwidth, len(res1.columns))
self.assertEqual(targwidth, len(res2.columns))
self.assertEqual(targlen, len(targ.index))
self.assertEqual(targlen, len(res0.index))
self.assertEqual(targlen, len(res1.index))
self.assertEqual(targlen, len(res2.index))
self.assertCountEqual(keys, targ.columns.names)
self.assertCountEqual(keys, res0.columns.names)
self.assertCountEqual(keys, res1.columns.names)
self.assertCountEqual(keys, res2.columns.names)
assert_array_equal(targ.values, res0.values)
assert_array_equal(targ.values, res1.values)
assert_array_equal(targ.values, res2.values)
assert_frame_equal(targ, res0)
assert_frame_equal(targ, res1)
assert_frame_equal(targ, res2)
def test__multi_spiketrains_to_dataframe__block_parents_childfirst(self):
obj = fake_neo('Block', seed=0, n=3)
res0 = ep.multi_spiketrains_to_dataframe(obj)
res1 = ep.multi_spiketrains_to_dataframe(obj, parents=True)
res2 = ep.multi_spiketrains_to_dataframe(obj, child_first=True)
res3 = ep.multi_spiketrains_to_dataframe(obj, parents=True,
child_first=True)
objs = obj.list_children_by_class('SpikeTrain')
targ = [ep.spiketrain_to_dataframe(iobj,
parents=True, child_first=True)
for iobj in objs]
targ = ep._sort_inds(pd.concat(targ, axis=1), axis=1)
keys = ep._extract_neo_attrs_safe(objs[0], parents=True,
child_first=True).keys()
keys = list(keys)
targwidth = len(objs)
targlen = max(len(iobj) for iobj in objs)
self.assertGreater(len(objs), 0)
self.assertEqual(targwidth, len(targ.columns))
self.assertEqual(targwidth, len(res0.columns))
self.assertEqual(targwidth, len(res1.columns))
self.assertEqual(targwidth, len(res2.columns))
self.assertEqual(targwidth, len(res3.columns))
self.assertEqual(targlen, len(targ.index))
self.assertEqual(targlen, len(res0.index))
self.assertEqual(targlen, len(res1.index))
self.assertEqual(targlen, len(res2.index))
self.assertEqual(targlen, len(res3.index))
self.assertCountEqual(keys, targ.columns.names)
self.assertCountEqual(keys, res0.columns.names)
self.assertCountEqual(keys, res1.columns.names)
self.assertCountEqual(keys, res2.columns.names)
self.assertCountEqual(keys, res3.columns.names)
|
assert_array_equal(targ.values, res0.values)
|
numpy.testing.utils.assert_array_equal
|
###################################################################################
## Main sampler
## Depending on the number of MCMC states defined in the first run.
if __name__ == "__main__":
import nonstat_model_noXs.model_sim as utils
import nonstat_model_noXs.generic_samplers as sampler
import nonstat_model_noXs.priors as priors
import os
import numpy as np
import matplotlib.pyplot as plt
from matplotlib.backends.backend_pdf import PdfPages
from pickle import load
from pickle import dump
from scipy.linalg import lapack
# Check whether the 'mpi4py' is installed
test_mpi = os.system("python -c 'from mpi4py import *' &> /dev/null")
if test_mpi != 0:
import sys
sys.exit("mpi4py import is failing, aborting...")
# get rank and size
from mpi4py import MPI
comm = MPI.COMM_WORLD
rank = comm.Get_rank()
size = comm.Get_size()
thinning = 10; echo_interval = 20; n_updates = 50001
# Filename for storing the intermediate results
input_file='./nonstat_progress_'+str(rank)+'.pkl'
# Load data input
if rank==0:
with open(input_file, 'rb') as f:
Y = load(f)
cen = load(f)
cen_above = load(f)
initial_values = load(f)
sigma_m = load(f)
prop_sigma = load(f)
iter_current = load(f)
phi_trace = load(f)
tau_sqd_trace = load(f)
theta_c_trace = load(f)
beta_loc0_trace = load(f)
beta_loc1_trace = load(f)
beta_scale_trace = load(f)
beta_shape_trace = load(f)
Z_1t_trace = load(f)
R_1t_trace = load(f)
Y_onetime = load(f)
X_onetime = load(f)
X_s_onetime = load(f)
R_onetime = load(f)
Z_onetime = load(f)
f.close()
else:
with open(input_file, 'rb') as f:
Y = load(f)
cen = load(f)
cen_above = load(f)
initial_values = load(f)
sigma_m = load(f)
iter_current = load(f)
Z_1t_trace = load(f)
R_1t_trace = load(f)
Y_onetime = load(f)
X_onetime = load(f)
X_s_onetime = load(f)
R_onetime = load(f)
Z_onetime = load(f)
f.close()
# Bookkeeping
n_s = Y.shape[0]
n_t = Y.shape[1]
if n_t != size:
import sys
sys.exit("Make sure the number of cpus (N) = number of time replicates (n_t), i.e.\n srun -N python nonstat_sampler.py")
wh_to_plot_Xs = n_s*np.array([0.25,0.5,0.75])
wh_to_plot_Xs = wh_to_plot_Xs.astype(int)
# Filename for storing the intermediate results
filename='./nonstat_progress_'+str(rank)+'.pkl'
# Generate multiple independent random streams
random_generator = np.random.RandomState()
# Constants to control adaptation of the Metropolis sampler
c_0 = 10
c_1 = 0.8
offset = 3 # the iteration offset
r_opt_1d = .41
r_opt_2d = .35
eps = 1e-6 # a small number
# Hyper parameters for the prior of the mixing distribution parameters and
hyper_params_phi = np.array([0.5,0.7])
hyper_params_tau_sqd = np.array([0.1,0.1])
hyper_params_theta_c = np.array([0, 20])
hyper_params_theta_gev = 25
# hyper_params_range = np.array([0.5,1.5]) # in case where roughness is not updated
# Load latest values
initial_values = comm.bcast(initial_values,root=0) # Latest values are mostly in initial_values
phi = initial_values['phi']
gamma = initial_values['gamma']
tau_sqd = initial_values['tau_sqd']
prob_below = initial_values['prob_below']
prob_above = initial_values['prob_above']
Dist = initial_values['Dist']
theta_c = initial_values['theta_c']
Design_mat = initial_values['Design_mat']
beta_loc0 = initial_values['beta_loc0']
beta_loc1 = initial_values['beta_loc1']
Time = initial_values['Time']
beta_scale = initial_values['beta_scale']
beta_shape = initial_values['beta_shape']
n_covariates = len(beta_loc0)
if rank == 0:
X = np.empty((n_s,n_t))
X_s = np.empty((n_s,n_t))
Z = np.empty((n_s,n_t))
R = np.empty((n_t,))
# Eigendecomposition of the correlation matrix
tmp_vec = np.ones(n_s)
Cor = utils.corr_fn(Dist, theta_c)
# eig_Cor = np.linalg.eigh(Cor) #For symmetric matrices
# V = eig_Cor[1]
# d = eig_Cor[0]
cholesky_inv = lapack.dposv(Cor,tmp_vec)
# For current values of phi and gamma, obtain grids of survival probs and densities
grid = utils.density_interp_grid(phi, gamma, grid_size=800)
xp = grid[0]; den_p = grid[1]; surv_p = grid[2]
thresh_X = utils.qRW_me_interp(prob_below, xp, surv_p, tau_sqd, phi, gamma)
thresh_X_above = utils.qRW_me_interp(prob_above, xp, surv_p, tau_sqd, phi, gamma)
# Marginal GEV parameters: per location x time
loc0 = Design_mat @beta_loc0
loc1 = Design_mat @beta_loc1
Loc = np.tile(loc0, n_t) + np.tile(loc1, n_t)*np.repeat(Time,n_s)
Loc = Loc.reshape((n_s,n_t),order='F')
scale = Design_mat @beta_scale
Scale = np.tile(scale, n_t)
Scale = Scale.reshape((n_s,n_t),order='F')
Design_mat1 = np.c_[np.repeat(1,n_s), np.log(Design_mat[:,1])]
shape = Design_mat1 @beta_shape
Shape = np.tile(shape, n_t)
Shape = Shape.reshape((n_s,n_t),order='F')
# Initial trace objects
Z_1t_accept = np.zeros(n_s)
R_accept = 0
if rank == 0:
print("Number of time replicates = %d"%size)
theta_c_trace_within_thinning = np.empty((2,thinning)); theta_c_trace_within_thinning[:] = np.nan
beta_loc0_trace_within_thinning = np.empty((n_covariates,thinning)); beta_loc0_trace_within_thinning[:] = np.nan
beta_loc1_trace_within_thinning = np.empty((n_covariates,thinning)); beta_loc1_trace_within_thinning[:] = np.nan
beta_scale_trace_within_thinning =
|
np.empty((n_covariates,thinning))
|
numpy.empty
|
# Copyright 2017 The dm_control Authors.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# ============================================================================
"""Planar Stacker domain."""
from __future__ import absolute_import
from __future__ import division
from __future__ import print_function
import collections
from dm_control import mujoco
from dm_control.rl import control
from dm_control.suite import base
from dm_control.suite import common
from dm_control.utils import containers
from dm_control.utils import rewards
from dm_control.utils import xml_tools
from lxml import etree
import numpy as np
_CLOSE = .01 # (Meters) Distance below which a thing is considered close.
_CONTROL_TIMESTEP = .01 # (Seconds)
_TIME_LIMIT = 10 # (Seconds)
_ARM_JOINTS = ['arm_root', 'arm_shoulder', 'arm_elbow', 'arm_wrist',
'finger', 'fingertip', 'thumb', 'thumbtip']
SUITE = containers.TaggedTasks()
def make_model(n_boxes):
"""Returns a tuple containing the model XML string and a dict of assets."""
xml_string = common.read_model('stacker.xml')
parser = etree.XMLParser(remove_blank_text=True)
mjcf = etree.XML(xml_string, parser)
# Remove unused boxes
for b in range(n_boxes, 4):
box = xml_tools.find_element(mjcf, 'body', 'box' + str(b))
box.getparent().remove(box)
return etree.tostring(mjcf, pretty_print=True), common.ASSETS
@SUITE.add('hard')
def stack_2(fully_observable=True, time_limit=_TIME_LIMIT, random=None,
environment_kwargs=None):
"""Returns stacker task with 2 boxes."""
n_boxes = 2
physics = Physics.from_xml_string(*make_model(n_boxes=n_boxes))
task = Stack(n_boxes=n_boxes,
fully_observable=fully_observable,
random=random)
environment_kwargs = environment_kwargs or {}
return control.Environment(
physics, task, control_timestep=_CONTROL_TIMESTEP, time_limit=time_limit,
**environment_kwargs)
@SUITE.add('hard')
def stack_4(fully_observable=True, time_limit=_TIME_LIMIT, random=None,
environment_kwargs=None):
"""Returns stacker task with 4 boxes."""
n_boxes = 4
physics = Physics.from_xml_string(*make_model(n_boxes=n_boxes))
task = Stack(n_boxes=n_boxes,
fully_observable=fully_observable,
random=random)
environment_kwargs = environment_kwargs or {}
return control.Environment(
physics, task, control_timestep=_CONTROL_TIMESTEP, time_limit=time_limit,
**environment_kwargs)
class Physics(mujoco.Physics):
"""Physics with additional features for the Planar Manipulator domain."""
def bounded_joint_pos(self, joint_names):
"""Returns joint positions as (sin, cos) values."""
joint_pos = self.named.data.qpos[joint_names]
return np.vstack([np.sin(joint_pos), np.cos(joint_pos)]).T
def joint_vel(self, joint_names):
"""Returns joint velocities."""
return self.named.data.qvel[joint_names]
def body_2d_pose(self, body_names, orientation=True):
"""Returns positions and/or orientations of bodies."""
if not isinstance(body_names, str):
body_names = np.array(body_names).reshape(-1, 1) # Broadcast indices.
pos = self.named.data.xpos[body_names, ['x', 'z']]
if orientation:
ori = self.named.data.xquat[body_names, ['qw', 'qy']]
return np.hstack([pos, ori])
else:
return pos
def touch(self):
return np.log1p(self.data.sensordata)
def site_distance(self, site1, site2):
site1_to_site2 = np.diff(self.named.data.site_xpos[[site2, site1]], axis=0)
return
|
np.linalg.norm(site1_to_site2)
|
numpy.linalg.norm
|
import os.path
import random
import numpy as np
import cv2
import torch
import torch.utils.data as data
import data.util as util
import sys
sys.path.append('../codes/scripts')
sys.path.append('../codes/data')
import augmentations
class LRHRDataset(data.Dataset):
'''
Read LR and HR image pairs.
If only HR image is provided, generate LR image on-the-fly.
The pair is ensured by 'sorted' function, so please check the name convention.
'''
def __init__(self, opt):
super(LRHRDataset, self).__init__()
self.opt = opt
self.paths_LR = None
self.paths_HR = None
self.LR_env = None # environment for lmdb
self.HR_env = None
self.HR_crop = None #v
self.HR_rrot = None #v
self.LR_scale = None #v
self.LR_blur = None #v
self.HR_noise = None #v
self.LR_noise = None #v
self.LR_noise2 = None #v
self.LR_cutout = None #v
self.LR_erasing = None #v
# read image list from subset list txt
if opt['subset_file'] is not None and opt['phase'] == 'train':
with open(opt['subset_file']) as f:
self.paths_HR = sorted([os.path.join(opt['dataroot_HR'], line.rstrip('\n')) \
for line in f])
if opt['dataroot_LR'] is not None:
raise NotImplementedError('Now subset only supports generating LR on-the-fly.')
else: # read image list from lmdb or image files
self.HR_env, self.paths_HR = util.get_image_paths(opt['data_type'], opt['dataroot_HR'])
self.LR_env, self.paths_LR = util.get_image_paths(opt['data_type'], opt['dataroot_LR'])
assert self.paths_HR, 'Error: HR path is empty.'
if self.paths_LR and self.paths_HR:
assert len(self.paths_LR) == len(self.paths_HR), \
'HR and LR datasets have different number of images - {}, {}.'.format(\
len(self.paths_LR), len(self.paths_HR))
#v parse on the fly options
if opt['hr_crop']: #v variable to activate automatic crop of HR image to correct size and generate LR
self.HR_crop = True
print("Automatic crop of HR images enabled")
if opt['hr_rrot']: #v variable to activate automatic rotate HR image and generate LR
self.HR_rrot = True
print("HR random rotation enabled")
if opt['hr_noise']: #v variable to activate adding noise to HR image
self.HR_noise = True
self.hr_noise_types = opt['hr_noise_types']
print("HR_noise enabled")
print(self.hr_noise_types)
if opt['lr_downscale']: #v variable to activate automatic downscale of HR images to LR pair, controlled by the scale of the model
self.LR_scale = True
self.scale_algos = opt['lr_downscale_types']
print("LR_scale enabled")
print(self.scale_algos)
if opt['lr_blur']: #v variable to activate automatic blur of LR images
self.LR_blur = True
self.blur_algos = opt['lr_blur_types']
print("LR_blur enabled")
print(self.blur_algos)
if opt['lr_noise']: #v variable to activate adding noise to LR image
self.LR_noise = True
self.noise_types = opt['lr_noise_types']
print("LR_noise enabled")
print(self.noise_types)
if opt['lr_noise2']: #vvariable to activate adding a secondary noise to LR image
self.LR_noise2 = True
self.noise_types2 = opt['lr_noise_types2']
print("LR_noise enabled")
print(self.noise_types)
if opt['lr_cutout']: #v variable to activate random cutout
self.LR_cutout = True
print("LR cutout enabled")
if opt['lr_erasing']: #v variable to activate random erasing
self.LR_erasing = True
print("LR random erasing enabled")
#v parse on the fly options
self.random_scale_list = [1]
def __getitem__(self, index):
HR_path, LR_path = None, None
scale = self.opt['scale']
HR_size = self.opt['HR_size']
#v
if self.opt['rand_flip_LR_HR'] and self.LR_scale and self.opt['phase'] == 'train':
LRHRchance = random.uniform(0, 1)
if self.opt['flip_chance']:
flip_chance = self.opt['flip_chance']
else:
flip_chance = 0.05
#print("Random Flip Enabled")
else:
LRHRchance = 0.
flip_chance = 0.
#print("No Random Flip")
# get HR image
if LRHRchance < (1- flip_chance):
HR_path = self.paths_HR[index]
#print("HR kept")
else:
HR_path = self.paths_LR[index]
#print("HR flipped")
#v
img_HR = util.read_img(self.HR_env, HR_path)
# modcrop in the validation / test phase
if self.opt['phase'] != 'train':
img_HR = util.modcrop(img_HR, scale)
# change color space if necessary
if self.opt['color']:
img_HR = util.channel_convert(img_HR.shape[2], self.opt['color'], [img_HR])[0]
#v
if self.HR_crop and (self.HR_rrot != True):
crop_size = (HR_size, HR_size)
img_HR, _ = augmentations.random_resize_img(img_HR, crop_size)
elif self.HR_rrot and (self.HR_crop != True):
img_HR, _ = augmentations.random_rotate(img_HR)
elif self.HR_crop and self.HR_rrot:
if np.random.rand() > 0.5:
crop_size = (HR_size, HR_size)
img_HR, _ = augmentations.random_resize_img(img_HR, crop_size)
else:
img_HR, _ = augmentations.random_rotate(img_HR)
#v
#v
if self.HR_noise:
img_HR, hr_noise_algo = augmentations.noise_img(img_HR, self.hr_noise_types)
#v
# get LR image
if self.paths_LR:
if self.HR_crop or self.HR_rrot: #v
img_LR = img_HR
else:
if LRHRchance < (1- flip_chance):
LR_path = self.paths_LR[index]
#print("LR kept")
else:
LR_path = self.paths_HR[index]
#print("LR flipped")
img_LR = util.read_img(self.LR_env, LR_path)
#"""
#v scale
if self.LR_scale:
img_LR, scale_interpol_algo = augmentations.scale_img(img_LR, scale)
#"""
#"""
#v blur
if self.LR_blur:
img_LR, blur_algo, blur_kernel_size = augmentations.blur_img(img_LR, self.blur_algos)
#"""
#"""
#v noise
if self.LR_noise:
img_LR, noise_algo = augmentations.noise_img(img_LR, self.noise_types)
if self.LR_noise2:
img_LR, noise_algo2 = augmentations.noise_img(img_LR, self.noise_types2)
#"""
#"""
#v LR cutout / LR random erasing
if self.LR_cutout and (self.LR_erasing != True):
img_LR = augmentations.cutout(img_LR, img_LR.shape[0] // 2)
elif self.LR_erasing and (self.LR_cutout != True): #only do cutout or erasing, not both
img_LR = augmentations.random_erasing(img_LR)
elif self.LR_cutout and self.LR_erasing:
if
|
np.random.rand()
|
numpy.random.rand
|
#%%
import numpy as np
import math
import scipy
from scipy.optimize import curve_fit
from scipy.interpolate import interp1d
from scipy.interpolate import CloughTocher2DInterpolator
from scipy.integrate import quad
import sys
sys.path.append('../')
import SQ_calcs
# Constants
pi = math.pi
heV = 4.14e-15 # eV*s
c = 2.99792e8 # m/s
kbeV = 8.6173e-5 # eV/K
keV = 8.6173e-5 # eV/K
h = 6.626e-34
kb = 1.38065e-23
q = 1.60218e-19
#%%
# This module contains functions for Photoluminescence data analysis and modeling
def aipl(data, dark, grating):
"""
This function takes PL data in cts/second units and
converts to AIPL based on a laser power and grating calibration
file. Functionality is built in to handle both single and map files
INPUTS:
data - data matrix containing input wavelength and PL cts/sec data
if m x 2 matrix, treats as single spectra file
if m x n matrix, treats as map along m
if n x m matrix, treats as map along n
dark - can be 0
grating - specifies which grating used, a string either '500nm' or '1200nm'
or '1200nm-InGaAs'
OUTPUTS:
aipl_data - data converted to absolute units , [=] photons/m^2-s-eV
"""
#Get grating calibration file, then calculate conversion factor
def BBPhotonFluxPerNM(lam,T):
a = 2*pi/(h**3*c**2)*((h*c/(lam*1e-9))**2/(np.exp((h*c/(lam*1e-9))/(kb*T))-1))*(h*c/(lam*1e-9)**2)*1e-9
return a
if grating == '500nm':
BB1050 = np.loadtxt('../../data/PLdata/grating_calibration_files/150 500'
'blaze BB files/BB 1050 10 um hole 10x SiCCD 532 LP'
'F No Duoscan Autoscanning_2.txt')
BB_raw_photon_data = BB1050[:,1]/np.insert(BB1050[1:,0]-BB1050[:-1,0],
0,BB1050[1,0]-BB1050[0,0])
AbsFluxesPerNM = np.zeros(BB1050.shape[0])
Ts = 1050;
for ii in range(BB1050.shape[0]):
AbsFluxesPerNM[ii] = BBPhotonFluxPerNM(BB1050[ii,0],Ts+273.15)
AbsPhotonRate = pi*(10/2*1e-6)**2*AbsFluxesPerNM #photons/sec-nm
Conversion_factor = AbsPhotonRate/BB_raw_photon_data
Ave_conv_factors = np.zeros([BB1050.shape[0],2])
Ave_conv_factors[:,0] = BB1050[:,0]
Ave_conv_factors[:,1] = Conversion_factor
f2 = interp1d(Ave_conv_factors[:,0], Ave_conv_factors[:,1], kind='cubic')
elif grating == '1200nm':
BB850 = np.loadtxt('../../data/PLdata/grating_calibration_files/BB 850C 10 um hole D0 10x 150 grating CCD 532 nm NoDS.txt')
BB950 = np.loadtxt('../../data/PLdata/grating_calibration_files/BB 950C 10 um hole D0 10x 150 grating CCD 532 nm NoDS.txt')
BB1050 = np.loadtxt('../../data/PLdata/grating_calibration_files/BB 1050C 10 um hole D0 10x 150 grating CCD 532 nm NoDS.txt')
BB_raw_photon_data_1 = BB850[:,1]/np.insert(BB1050[1:,0]-BB1050[:-1,0],
0,BB1050[1,0]-BB1050[0,0])
BB_raw_photon_data_2 = BB950[:,1]/np.insert(BB1050[1:,0]-BB1050[:-1,0],
0,BB1050[1,0]-BB1050[0,0])
BB_raw_photon_data_3 = BB1050[:,1]/np.insert(BB1050[1:,0]-BB1050[:-1,0],
0,BB1050[1,0]-BB1050[0,0])
BB_raw_photon_data = np.array([BB_raw_photon_data_1,BB_raw_photon_data_2,BB_raw_photon_data_3])
AbsFluxesPerNM = np.zeros(BB_raw_photon_data.shape)
for lam in range(len(BB_raw_photon_data_1)):
tt = 0
for T in (850,950,1050):
AbsFluxesPerNM[tt,lam] = BBPhotonFluxPerNM(BB850[lam,0],T+273.15)
tt += 1
AbsPhotonRate = pi*(10/2*1e-6)**2*AbsFluxesPerNM #photons/sec-nm
Conversion_factor = AbsPhotonRate/BB_raw_photon_data
Ave_conv_factors = np.zeros([BB850.shape[0],2])
Ave_conv_factors[:,0] = BB850[:,0]
Ave_conv_factors[:,1] = np.mean(Conversion_factor,0)
f2 = interp1d(Ave_conv_factors[:,0], Ave_conv_factors[:,1], kind='cubic')
elif grating == '1200nm-InGaAs':
BB850 = np.loadtxt('../../data/PLdata/grating_calibration_files/Response_Synapse CCD2_784_150_Objective_x10_UV_0_Detector_Second_InjRej_Edge 785nm PL.txt')
BB_raw_photon_data = BB850[:,1]/np.insert(BB850[1:,0]-BB850[:-1,0],
0,BB850[1,0]-BB850[0,0])
AbsFluxesPerNM = np.zeros(BB850.shape[0])
Ts = 850;
for ii in range(BB850.shape[0]):
AbsFluxesPerNM[ii] = BBPhotonFluxPerNM(BB850[ii,0],Ts+273.15)
AbsPhotonRate = pi*(10/2*1e-6)**2*AbsFluxesPerNM #photons/sec-nm
Conversion_factor = AbsPhotonRate/BB_raw_photon_data
Ave_conv_factors = np.zeros([BB850.shape[0],2])
Ave_conv_factors[:,0] = BB850[:,0]
Ave_conv_factors[:,1] = Conversion_factor
f2 = interp1d(Ave_conv_factors[:,0], Ave_conv_factors[:,1], kind='cubic')
if data.shape[1] == 2: #single spectrum
aipl_data = data
lam = data[:,0]
Ipl_raw = data[:,1] #cts/sec
if dark == []:
Ipl_raw2 = Ipl_raw
else:
Ipl_raw = Ipl_raw - dark[:,1]
Ipl_raw2 = Ipl_raw/np.insert(lam[1:]-lam[:-1],0,lam[1]-lam[0]) #cts/sec-nm
Ipl_nm = Ipl_raw2*f2(lam) #photons/sec-nm
bandwidth_conv = np.insert(lam[1:]-lam[:-1],0,lam[1]-lam[0])/(heV*c/(lam*1e-9)**2*np.insert(lam[1:]-lam[:-1],0,lam[1]-lam[0])*1e-9)
Ipl = Ipl_nm*bandwidth_conv/(pi*(6.01e-6)**2*2*0.921) #photons/sec-eV-m^2 (divide by factor of 2 since only considering FWHM beam area) (divide by 0.921 for window)
aipl_data[:,1] = Ipl
else:
aipl_data = data
k = 0
while np.isnan(data[0,k]):
k = k + 1
lam = data[0,k:]
for ii in range(1,data.shape[0]):
Ipl_raw = data[ii,k:]
if dark == []:
Ipl_raw2 = Ipl_raw
else:
Ipl_raw = Ipl_raw - dark[:,1]
Ipl_raw2 = Ipl_raw/np.insert(lam[1:]-lam[:-1],0,lam[1]-lam[0]) #cts/sec-nm
Ipl_nm = Ipl_raw2*f2(lam) #photons/sec-nm
bandwidth_conv = np.insert(lam[1:]-lam[:-1],0,lam[1]-lam[0])/(heV*c/(lam*1e-9)**2*np.insert(lam[1:]-lam[:-1],0,lam[1]-lam[0])*1e-9)
Ipl = Ipl_nm*bandwidth_conv/(pi*(6.01e-6)**2*2*0.921) #photons/sec-eV-m^2 (divide by factor of 2 since only considering FWHM beam area) (divide by 0.921 for window)
aipl_data[ii,k:] = Ipl
return aipl_data
def plqy_ext(aipl_data, laser_power, laser, temperature):
'''
This is the simple PLQY method for determining quasi-Fermi level splitting
from PLQY, using SQ limit as reference. Presently the assumed temperature
is 350K for SQ calculation and 300K for chi calculation (this avoids
overestimation of QLFS or chi)
INPUTS:
aipl_data - PL spectrum matrix in absolute units (output from
PLtools.aipl function)
laser_power - laser powermeter reason in SI units (needed for PLQY calc)
laser - string
OUTPUTs:
All of the useful PL parameters
mean_Ipl - mean PL emission E [eV] (also called 1st moment)
peak_pos - PL peak position [eV]
FWHM - Full Width Half Max of PL peak [eV]
PLQY - Photoluminescence Quantuum Yield [fraction]
dmu_PLQY - Quasi-Fermi Level splitting from PLQY method
chi_PLQY - QFLS/SQ-max from PLQY method
dmu_PLQY_Eg - QFLS, PLQY method, using PL integrated above peak_pos only
chi_PLQY_Eg - QFLS / SQ-Max, from PLQY-Eg method
'''
DiodeReadings_1sun = laser_power
if laser == '532nm':
DiodeResponse532= 0.2741
Ep532 = 2.3305 #E per photon @532
Area785ImageJ = pi*(6.01e-6)**2 #m^2
elif laser == '785nm':
DiodeResponse532= 0.4165906265 # for 785
Ep532 = 1.59236 #E per photon @785
Area785ImageJ = 1.77e-10 #m^2
#Load data from Mathmatica calcs to determine SQ limits @ 300 K and 350 K for various
#Egs
Egs = np.loadtxt('../../data/PLdata/vocmax_data/Egs.txt',delimiter=',')
VocSQs300 = np.loadtxt('../../data/PLdata/vocmax_data/VocMaxs.txt',delimiter=',') # 300 K
Jphs = np.loadtxt('../../data/PLdata/vocmax_data/Jphs.txt',delimiter=',') #300 K
VocSQs350 = np.loadtxt('../../data/PLdata/vocmax_data/' + temperature + '/VocMaxs2.txt',delimiter=',') # 350 K
VocSQs350 = np.loadtxt('../../data/PLdata/vocmax_data/VocMaxs2.txt',delimiter=',') # 350 K
VocSQs300_fn = interp1d(Egs, VocSQs300, kind='cubic')
VocSQs350_fn = interp1d(Egs, VocSQs350, kind='cubic')
Jphs_fn = interp1d(Egs, Jphs, kind='cubic')
DiodeReading = DiodeReadings_1sun
P532 = DiodeReading/(DiodeResponse532*Area785ImageJ*10) #W/m^2
Jp532 = DiodeReading*0.925/(DiodeResponse532*Area785ImageJ*1.60218e-19*Ep532*2)
T = float(temperature[:-1])
if aipl_data.shape[1] == 2: #single spectrum
lam = aipl_data[:,0]
E = heV*c/(lam*1e-9)
Ipl = aipl_data[:,1]
maxI = np.max(Ipl)
maxI_idx = np.argmax(Ipl)
peak_pos = E[maxI_idx]
HHMax_idx = np.argmin(np.absolute(maxI/2-Ipl[:maxI_idx]))
LHMax_idx = np.argmin(np.absolute(maxI/2-Ipl[maxI_idx:]))
LHMax_idx = LHMax_idx+maxI_idx-1
FWHM = E[HHMax_idx]-E[LHMax_idx]
try:
VocSQ300 = VocSQs300_fn(E[maxI_idx])
VocSQ350 = VocSQs350_fn(E[maxI_idx])
JphSQ = Jphs_fn(E[maxI_idx])
except ValueError:
VocSQ300 = SQ_calcs.VocSQ(E[maxI_idx],300)
VocSQ350 = SQ_calcs.VocSQ(E[maxI_idx],315)
JphSQ = SQ_calcs.JphSQ(E[maxI_idx],300)
NSuns = Jp532*q/JphSQ;
VocMax300 = VocSQ300 + kb*300/q*np.log(Jp532*q/JphSQ)
VocMax350 = VocSQ350 + kb*T/q*np.log(Jp532*q/JphSQ)
TotalPL = np.mean(-E[1:-1]+E[0:-2])/2*(Ipl[0]+Ipl[-1]+2*np.sum(Ipl[1:-2]))
TotalPL = np.max([TotalPL, -TotalPL])
TotalPL_Eg = np.mean(-E[1:maxI_idx]+E[0:maxI_idx-1])/2*(Ipl[0]+Ipl[maxI_idx]+2*np.sum(Ipl[1:maxI_idx-1]))
TotalPL_Eg = np.max([TotalPL_Eg, -TotalPL_Eg])
PLQY = TotalPL/Jp532
dmu_PLQY = VocMax350-kbeV*T*np.log(1/PLQY)
chi_PLQY = dmu_PLQY/VocMax300
chi_PLQY_Eg = (VocMax350-kbeV*T*np.log(1/(TotalPL_Eg/Jp532)))/VocMax300
PLQY_Eg = TotalPL_Eg/Jp532
dmu_PLQY_Eg = VocMax350-kbeV*T*np.log(1/(TotalPL_Eg/Jp532))
mean_Ipl = np.sum(Ipl*E)/np.sum(Ipl)
else: #maps
k = 0
while np.isnan(aipl_data[0,k]):
k = k + 1
lam = aipl_data[0,k:]
E = heV*c/(lam*1e-9)
mean_Ipl = np.zeros(aipl_data.shape[0]-1)
peak_pos = np.zeros(aipl_data.shape[0]-1)
FWHM = np.zeros(aipl_data.shape[0]-1)
PLQY = np.zeros(aipl_data.shape[0]-1)
dmu_PLQY = np.zeros(aipl_data.shape[0]-1)
chi_PLQY = np.zeros(aipl_data.shape[0]-1)
dmu_PLQY_Eg = np.zeros(aipl_data.shape[0]-1)
chi_PLQY_Eg = np.zeros(aipl_data.shape[0]-1)
for ii in range(1,aipl_data.shape[0]):
Ipl = aipl_data[ii,k:]
maxI = np.max(Ipl)
maxI_idx = np.argmax(Ipl)
peak_pos[ii-1] = E[maxI_idx]
HHMax_idx = np.argmin(np.absolute(maxI/2-Ipl[:maxI_idx]))
LHMax_idx = np.argmin(np.absolute(maxI/2-Ipl[maxI_idx:]))
LHMax_idx = LHMax_idx+maxI_idx-1
FWHM[ii-1] = E[HHMax_idx]-E[LHMax_idx]
try:
VocSQ300 = VocSQs300_fn(E[maxI_idx])
VocSQ350 = VocSQs350_fn(E[maxI_idx])
JphSQ = Jphs_fn(E[maxI_idx])
except ValueError:
VocSQ300 = SQ_calcs.VocSQ(E[maxI_idx],300)
VocSQ350 = SQ_calcs.VocSQ(E[maxI_idx],315)
JphSQ = SQ_calcs.JphSQ(E[maxI_idx],300)
NSuns = Jp532*q/JphSQ;
VocMax300 = VocSQ300 + kb*300/q*
|
np.log(Jp532*q/JphSQ)
|
numpy.log
|
import os
import glob
import numpy as np
import math
import _pickle as cPickle
pred_data = "eval_results/TEST_"
pred_list = [1] # eval_id list, if you want to test the mean score of TEST_1 and TEST_2, change to [1,2]
data_dir = "My_NOCS"
synset_names = ['BG',
'bottle',
'bowl',
'camera',
'can',
'laptop',
'mug'
]
def compute_3d_iou_new(RT_1, RT_2, noc_cube_1, noc_cube_2, handle_visibility, class_name_1, class_name_2):
'''Computes IoU overlaps between two 3d bboxes.
bbox_3d_1, bbox_3d_1: [3, 8]
'''
# flatten masks
def asymmetric_3d_iou(RT_1, RT_2, noc_cube_1, noc_cube_2):
bbox_3d_1 = transform_coordinates_3d(noc_cube_1, RT_1)
bbox_3d_2 = transform_coordinates_3d(noc_cube_2, RT_2)
bbox_1_max =
|
np.amax(bbox_3d_1, axis=0)
|
numpy.amax
|
"""Methods for geodetic calculations."""
import os
import numpy
import srtm
import geopy
from geopy.distance import GeodesicDistance
from gewittergefahr.gg_utils import longitude_conversion as lng_conversion
from gewittergefahr.gg_utils import file_system_utils
from gewittergefahr.gg_utils import error_checking
RADIANS_TO_DEGREES = 180. / numpy.pi
DEGREES_TO_RADIANS = numpy.pi / 180
MIN_LATITUDE_DEG = -90.
MAX_LATITUDE_DEG = 90.
MIN_LONGITUDE_NEGATIVE_IN_WEST_DEG = -180.
MAX_LONGITUDE_NEGATIVE_IN_WEST_DEG = 180.
MIN_LONGITUDE_POSITIVE_IN_WEST_DEG = 0.
MAX_LONGITUDE_POSITIVE_IN_WEST_DEG = 360.
POSITIVE_LONGITUDE_ARG = 'positive'
NEGATIVE_LONGITUDE_ARG = 'negative'
EITHER_SIGN_LONGITUDE_ARG = 'either'
VALID_LONGITUDE_SIGN_ARGS = [
POSITIVE_LONGITUDE_ARG, NEGATIVE_LONGITUDE_ARG, EITHER_SIGN_LONGITUDE_ARG]
class ElevationFileHandler:
"""File-handler for elevation data.
This class mimics the class `FileHandler` in main.py of the `srtm` package.
"""
working_dir_name = ''
def __init__(self, working_dir_name=None):
"""Creates new instance.
:param working_dir_name: Path to working directory. Elevation files
will be read from here and, if necessary, downloaded to here. If
`working_dir_name is None`, will try to create subdirectory
".cache/srtm" in the home directory.
:raises: ValueError: if `working_dir_name is None` and this method
cannot create ".cache/srtm" in the home directory.
"""
if working_dir_name is None:
if 'HOME' in os.environ:
top_working_dir_name = os.environ['HOME']
elif 'HOMEPATH' in os.environ:
top_working_dir_name = os.environ['HOMEPATH']
else:
raise ValueError('Cannot find home directory.')
working_dir_name = '{0:s}/.cache/srtm'.format(top_working_dir_name)
file_system_utils.mkdir_recursive_if_necessary(
directory_name=working_dir_name)
self.working_dir_name = working_dir_name
def get_srtm_dir(self):
"""Returns path to working directory.
:return: working_dir_name: See doc for `__init__`.
"""
return self.working_dir_name
def exists(self, file_name):
"""Returns flag, indicating whether or not a file exists.
:param file_name: Pathless file name.
:return: does_file_exist: Boolean flag.
"""
full_file_name = '{0:s}/{1:s}'.format(self.get_srtm_dir(), file_name)
return os.path.isfile(full_file_name)
def write(self, file_name, contents):
"""Writes elevation file to working directory.
:param file_name: Pathless file name.
:param contents: Stuff to be written.
"""
full_file_name = '{0:s}/{1:s}'.format(self.get_srtm_dir(), file_name)
with open(full_file_name, 'wb') as f:
f.write(contents)
def read(self, file_name):
"""Reads elevation file from working directory.
:param file_name: Pathless file name.
:return: contents: Stuff contained in file.
"""
full_file_name = '{0:s}/{1:s}'.format(self.get_srtm_dir(), file_name)
with open(full_file_name, 'rb') as f:
return f.read()
def _get_elevation(
latitude_deg, longitude_deg, srtm_data_object=None,
working_dir_name=None):
"""Gets elevation at a single point.
WARNING: Input longitudes in western hemisphere must be negative.
If `srtm_data_object is None`, it will be created on the fly.
:param latitude_deg: Latitude (deg N).
:param longitude_deg: Longitude (deg E).
:param srtm_data_object: Instance of `srtm.data.GeoElevationData`.
:param working_dir_name: See doc for `__init__` in class
`ElevationFileHandler`.
:return: elevation_m_asl: Elevation (metres above sea level).
:return: srtm_data_object: Instance of `srtm.data.GeoElevationData`.
"""
if srtm_data_object is None:
srtm_data_object = srtm.get_data(
file_handler=ElevationFileHandler(working_dir_name))
elevation_m_asl = srtm_data_object.get_elevation(
latitude=latitude_deg, longitude=longitude_deg)
# TODO(thunderhoser): I am concerned about this hack.
if elevation_m_asl is None:
elevation_m_asl = 0.
return elevation_m_asl, srtm_data_object
def find_invalid_latitudes(latitudes_deg):
"""Returns array indices of invalid latitudes.
:param latitudes_deg: 1-D numpy array of latitudes (deg N).
:return: invalid_indices: 1-D numpy array with array indices of invalid
latitudes.
"""
error_checking.assert_is_real_numpy_array(latitudes_deg)
error_checking.assert_is_numpy_array(latitudes_deg, num_dimensions=1)
valid_flags = numpy.logical_and(
latitudes_deg >= MIN_LATITUDE_DEG, latitudes_deg <= MAX_LATITUDE_DEG)
return numpy.where(numpy.invert(valid_flags))[0]
def find_invalid_longitudes(
longitudes_deg, sign_in_western_hemisphere=POSITIVE_LONGITUDE_ARG):
"""Returns array indices of invalid longitudes.
:param longitudes_deg: 1-D numpy array of longitudes (deg E).
:param sign_in_western_hemisphere: Required sign in western hemisphere. May
be "positive", "negative", or "either".
:return: invalid_indices: 1-D numpy array with array indices of invalid
longitudes.
:raises: ValueError: if `sign_in_western_hemisphere` is not one of the 3
aforelisted options.
"""
error_checking.assert_is_real_numpy_array(longitudes_deg)
error_checking.assert_is_numpy_array(longitudes_deg, num_dimensions=1)
error_checking.assert_is_string(sign_in_western_hemisphere)
if sign_in_western_hemisphere == POSITIVE_LONGITUDE_ARG:
valid_flags = numpy.logical_and(
longitudes_deg >= MIN_LONGITUDE_POSITIVE_IN_WEST_DEG,
longitudes_deg <= MAX_LONGITUDE_POSITIVE_IN_WEST_DEG)
elif sign_in_western_hemisphere == NEGATIVE_LONGITUDE_ARG:
valid_flags = numpy.logical_and(
longitudes_deg >= MIN_LONGITUDE_NEGATIVE_IN_WEST_DEG,
longitudes_deg <= MAX_LONGITUDE_NEGATIVE_IN_WEST_DEG)
elif sign_in_western_hemisphere == EITHER_SIGN_LONGITUDE_ARG:
valid_flags = numpy.logical_and(
longitudes_deg >= MIN_LONGITUDE_NEGATIVE_IN_WEST_DEG,
longitudes_deg <= MAX_LONGITUDE_POSITIVE_IN_WEST_DEG)
else:
error_string = (
'\n\n{0:s}Valid options for `sign_in_western_hemisphere` are listed'
' above and do not include "{1:s}".'
).format(str(VALID_LONGITUDE_SIGN_ARGS), sign_in_western_hemisphere)
raise ValueError(error_string)
return numpy.where(numpy.invert(valid_flags))[0]
def get_latlng_centroid(latitudes_deg, longitudes_deg, allow_nan=True):
"""Finds centroid of lat-long points.
P = number of points
:param latitudes_deg: length-P numpy array of latitudes (deg N).
:param longitudes_deg: length-P numpy array of longitudes (deg E).
:param allow_nan: Boolean flag. If True, input arrays may contain NaN's
(however, NaN's must occur at the exact same positions in the two
arrays).
:return: centroid_lat_deg: Centroid latitude (deg N).
:return: centroid_lng_deg: Centroid longitude (deg E).
:raises: ValueError: if allow_nan = True but NaN's do not occur at the same
positions in the two arrays.
"""
error_checking.assert_is_boolean(allow_nan)
error_checking.assert_is_valid_lat_numpy_array(latitudes_deg, allow_nan)
error_checking.assert_is_numpy_array(latitudes_deg, num_dimensions=1)
num_points = len(latitudes_deg)
longitudes_deg = lng_conversion.convert_lng_positive_in_west(
longitudes_deg, allow_nan)
error_checking.assert_is_numpy_array(
longitudes_deg, exact_dimensions=numpy.array([num_points]))
nan_latitude_indices = numpy.where(numpy.isnan(latitudes_deg))[0]
nan_longitude_indices = numpy.where(numpy.isnan(longitudes_deg))[0]
if not numpy.array_equal(nan_latitude_indices, nan_longitude_indices):
error_string = (
'\nNaN''s occur at the following positions in `latitudes_deg`:\n' +
str(nan_latitude_indices) +
'\nand the following positions in `longitudes_deg`:\n' +
str(nan_longitude_indices) +
'\nNaN''s should occur at the same positions in the two arrays.')
raise ValueError(error_string)
return numpy.nanmean(latitudes_deg), numpy.nanmean(longitudes_deg)
def get_elevations(latitudes_deg, longitudes_deg, working_dir_name=None):
"""Returns elevation of each point.
N = number of points
:param latitudes_deg: length-N numpy array of latitudes (deg N).
:param longitudes_deg: length-N numpy array of longitudes (deg E).
:param working_dir_name: See doc for `__init__` in class
`ElevationFileHandler`.
:return: elevations_m_asl: length-N numpy array of elevations (metres above
sea level).
"""
error_checking.assert_is_valid_lat_numpy_array(latitudes_deg)
error_checking.assert_is_numpy_array(latitudes_deg, num_dimensions=1)
num_points = len(latitudes_deg)
longitudes_deg = lng_conversion.convert_lng_negative_in_west(
longitudes_deg, allow_nan=False)
error_checking.assert_is_numpy_array(
longitudes_deg, exact_dimensions=numpy.array([num_points]))
srtm_data_object = None
elevations_m_asl = numpy.full(num_points, numpy.nan)
for i in range(num_points):
elevations_m_asl[i], srtm_data_object = _get_elevation(
latitude_deg=latitudes_deg[i], longitude_deg=longitudes_deg[i],
srtm_data_object=srtm_data_object,
working_dir_name=working_dir_name)
return elevations_m_asl
def start_points_and_displacements_to_endpoints(
start_latitudes_deg, start_longitudes_deg, scalar_displacements_metres,
geodetic_bearings_deg):
"""Computes endpoint from each start point and displacement.
:param start_latitudes_deg: numpy array with latitudes (deg N) of start
points.
:param start_longitudes_deg: equivalent-size numpy array with longitudes
(deg E) of start points.
:param scalar_displacements_metres: equivalent-size numpy array of scalar
displacements.
:param geodetic_bearings_deg: equivalent-size numpy array of geodetic
bearings (from start point to end point, measured clockwise from due
north).
:return: end_latitudes_deg: equivalent-size numpy array with latitudes
(deg N) of endpoints.
:return: end_longitudes_deg: equivalent-size numpy array with longitudes
(deg E) of endpoints.
"""
error_checking.assert_is_valid_lat_numpy_array(
start_latitudes_deg, allow_nan=False)
start_longitudes_deg = lng_conversion.convert_lng_positive_in_west(
start_longitudes_deg, allow_nan=False)
error_checking.assert_is_numpy_array(
start_longitudes_deg,
exact_dimensions=numpy.array(start_latitudes_deg.shape))
error_checking.assert_is_geq_numpy_array(scalar_displacements_metres, 0.)
error_checking.assert_is_numpy_array(
scalar_displacements_metres,
exact_dimensions=numpy.array(start_latitudes_deg.shape))
error_checking.assert_is_geq_numpy_array(geodetic_bearings_deg, 0.)
error_checking.assert_is_leq_numpy_array(geodetic_bearings_deg, 360.)
error_checking.assert_is_numpy_array(
geodetic_bearings_deg,
exact_dimensions=numpy.array(start_latitudes_deg.shape))
end_latitudes_deg =
|
numpy.full(start_latitudes_deg.shape, numpy.nan)
|
numpy.full
|
import torch
from torch.utils.data import Dataset, DataLoader
import torchvision
from torchvision import transforms
import torch.nn as nn
import os
import glob
import numpy as np
import time
import cv2
from einops import rearrange, reduce, repeat
from PIL import Image
#from utils.augmentations import SSDAugmentation, BaseTransform, NlosTransform
MEANS = (103.94, 116.78, 123.68)
STD = (57.38, 57.12, 58.40)
class DetrDataset(Dataset):
def __init__(self, args, is_correlation=False, mode='train', byol=False):
'''
dataset 처리
rf와 이미지의 경우에는 init 할 때부터 읽어와서 메모리에 올리지만 gt는 데이터를 활용할 때마다 load함.
mode - train : 학습을 위함. rf, gt, img 다 있는 경우
test : test를 위함. rf, gt, img 다 있는 경우
valid: valid를 위함(demo). rf, img만 있는 경우
'''
self.is_correlation = is_correlation
self.load_img = False #args.vis
self.mode = mode
#self.is_gaussian = args.gaussian
self.std = 0.1
self.mean = 0
self.is_normalize = False
self.cutoff = args.cutoff
self.augmentation = None
self.augmentation_prob = 1
#self.intensity = Intensity(scale=0.05)
self.print_once = True
self.flatten = True #args.flatten
self.byol = byol
#self.transform = NlosTransform(MEANS)
#print("for byol ? = ", self.byol)
data_path = '/home/tako/save_data_ver2'
#data_path_list = os.listdir(data_path)
data_path_list = glob.glob(data_path + '/*')
#print("data list", data_path_list)
data_path_list = sorted(data_path_list)
#print(data_path_list)
rf_data = [] # rf data list
target_list = [] # ground truth target
mask_list = [] # ground truth mask
img_list = []
human_index = []
print("start - data read")
#test_dir = [8, 9] # past version - 1
test_dir = [2, 5, 10, 14, 16, 19] #, 19] # cur version - 2
#test_dir = [2, 5, 10] # los
#test_dir = [14, 16, 19] #nlos
#test_dir = [10, 19] # demo - with mask , los , nlos
#test_dir = [2]
#test_dir = [14] # 흰 옷
remove_dir = [3, 4]
#valid_dir = [25, 26, 27]
#valid_dir = [19]
# valid_dir = [28, 29] # nlos wall
valid_dir = [x for x in range(21, 40)]
#valid_dir = [x for x in range(15, 40)]
#valid_dir += [x for x in range(1, 13)]
valid_dir = [x for x in range(1, 40)] # Model test
dir_count = 0
rf_index = 0
if mode == 'train':
outlier_list = range(49500, 50000)
else:
outlier_list = range(18000, 19000)
rf_index = -1
target_index = -1
mask_index = -1
img_index = -1
for file in data_path_list:
if dir_count in remove_dir:
dir_count += 1
continue
if mode == 'train' and (dir_count in test_dir or dir_count in valid_dir):
dir_count += 1
continue
elif mode == 'test' and dir_count not in test_dir:
dir_count += 1
continue
elif mode == 'valid' and dir_count not in valid_dir:
dir_count += 1
continue
if os.path.isdir(file) is True:
# 각 폴더 안의 npy 데이터
rf_file_list = glob.glob(file + '/raw/*.npy')
rf_file_list = sorted(rf_file_list)
print('dir_count:', dir_count,'dir(raw):', file, '\t# of data :', len(rf_file_list))
#print(rf_file_list)
for rf in rf_file_list:
rf_index += 1
if rf_index in outlier_list:
continue
temp_raw_rf = np.load(rf)[:, :, self.cutoff:]
#print("raw shape", temp_raw_rf.shape)
#----- normalization ------
if self.is_normalize is True:
for i in range(temp_raw_rf.shape[0]):
for j in range(temp_raw_rf.shape[1]):
stdev = np.std(temp_raw_rf[i, j])
temp_raw_rf[i, j] = temp_raw_rf[i, j]/stdev
#print("now shape",temp_raw_rf.shape)
#temp_raw_rf = torch.tensor(temp_raw_rf).float()
#print("now shape",temp_raw_rf.shape)
#m = torch.nn.Upsample(scale_factor=3, mode='bilinear')
#---------- 2차원으로 만들기 -----------
if self.flatten:
#print("now shape",temp_raw_rf.shape)
#temp_raw_rf = rearrange(temp_raw_rf, 'tx rx len -> (tx rx) len')
temp_raw_rf = rearrange(temp_raw_rf, 'tx rx len -> tx (rx len)')
#temp_raw_rf = rearrange(temp_raw_rf, 'x (len1 len2) -> x len1 len2', len1=int(math.sqrt(temp_raw_rf.shape[1])))
#print("now shape",temp_raw_rf.shape)
temp_raw_rf = rearrange(temp_raw_rf, 'tx (len1 len2) -> tx len1 len2', len1=72)
#temp_raw_rf = repeat(temp_raw_rf, 'c h w -> c (h rep_1) (w rep_2)', rep_1=3, rep_2=3)
#temp_raw_rf = m(temp_raw_rf)
#temp_raw_rf = temp_raw_rf.unsqueeze(0)
#print("now shape",temp_raw_rf.shape)
rf_data.append(temp_raw_rf)
#print("rf shape", temp_raw_rf.shape)
if self.print_once:
#print(temp_raw_rf[0][0])
#print(re_temp_raw_rf[0][0])
print("rf shape", temp_raw_rf.shape)
self.print_once = False
#break
'''
ground truth data 읽어오기.
target : [num_obj * 5] ( box 좌표, class)
mask : [num_obj, h, w] 1 = mask, 0 = else
'''
target_file_list = glob.glob(file + '/target/*')
target_file_list = sorted(target_file_list)
print('dir(target):', file, '\t# of data :', len(target_file_list))
'''
mask_file_list = glob.glob(file + '/mask/*')
mask_file_list = sorted(mask_file_list)
print('dir(mask):', file, '\t# of data :', len(mask_file_list))
'''
img_file_list = glob.glob(file + '/img/*')
img_file_list = sorted(img_file_list)
print('dir(img):', file, '\t# of data :', len(img_file_list))
#----- gt 파일 이름명만 리스트에 넣어놓기 -----
for target in target_file_list:
if target_index == 0:
print("target_shape ",
|
np.load(target)
|
numpy.load
|
'''
<NAME>
15863
Home Work 7
Exercise 3
'''
import matplotlib.pyplot as plt
import numpy as np
from matplotlib.collections import LineCollection
a = 2227057010910366687
M = 2 ** 64 - 59
c = 0
N = 1000000
x1 = np.zeros(N + 1)
y1 =
|
np.zeros(N + 1)
|
numpy.zeros
|
# audio-offset-finder
#
# Copyright (c) 2014 British Broadcasting Corporation
# Copyright (c) 2018 <NAME>
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
from subprocess import Popen, PIPE
from scipy.io import wavfile
# from scikits.talkbox.features.mfcc import mfcc
import matplotlib.pyplot as plt
import librosa
import os, tempfile, warnings
import numpy as np
def mfcc(audio, nwin=256, nfft=512, fs=16000, nceps=13):
#return librosa.feature.mfcc(y=audio, sr=44100, hop_length=nwin, n_mfcc=nceps)
return [np.transpose(librosa.feature.mfcc(y=audio, sr=fs, n_fft=nfft, win_length=nwin,n_mfcc=nceps))]
def add_feature(mfcc1, rmsa1):
tmfcc1 = np.zeros((mfcc1.shape[0],mfcc1.shape[1]+rmsa1.shape[0]))
n = mfcc1.shape[0]
m = mfcc1.shape[1]
w = rmsa1.shape[0]
tmfcc1[0:n,0:m] = mfcc1[0:n,0:m]
tmfcc1[0:n,m:m+w] = np.transpose(rmsa1[0:w,0:n])
return tmfcc1
def get_audio(file1, fs=8000, trim=60*15):
sr = fs
tmp1 = convert_and_trim(file1, fs, trim)
# Removing warnings because of 18 bits block size
# outputted by ffmpeg
# https://trac.ffmpeg.org/ticket/1843
warnings.simplefilter("ignore", wavfile.WavFileWarning)
a1 = wavfile.read(tmp1, mmap=True)[1] / (2.0 ** 15)
# We truncate zeroes off the beginning of each signals
# (only seems to happen in ffmpeg, not in sox)
a1 = ensure_non_zero(a1)
print("%s samples: %s" % (file1,a1.shape[0]))
mfcc1 = mfcc(a1, nwin=256, nfft=512, fs=fs, nceps=26)[0]
mfcc1 = std_mfcc(mfcc1)
rmsa1 = librosa.feature.rms(a1)
cent1 = librosa.feature.spectral_centroid(y=a1, sr=fs)
rolloff1 = librosa.feature.spectral_rolloff(y=a1, sr=fs, roll_percent=0.1)
chroma_cq1 = librosa.feature.chroma_cqt(y=a1, sr=fs, n_chroma=10)
onset_env1 = librosa.onset.onset_strength(y=a1, sr=sr)
pulse1 = librosa.beat.plp(onset_envelope=onset_env1, sr=sr)
mfcc1 = add_feature(mfcc1, rmsa1)
mfcc1 = add_feature(mfcc1, rolloff1/fs)
mfcc1 = add_feature(mfcc1, cent1/fs)
mfcc1 = add_feature(mfcc1, chroma_cq1)
mfcc1 = add_feature(mfcc1, onset_env1.reshape(1,onset_env1.shape[0]))
mfcc1 = add_feature(mfcc1, pulse1.reshape(1,onset_env1.shape[0]))
return tmp1, mfcc1, a1, rmsa1
def find_offset(audio1, audio2, fs=8000, correl_nframes=1000, plotit=False):
tmp1, mfcc1, a1, rmsa1 = audio1
tmp2, mfcc2, a2, rmsa2 = audio2
c = cross_correlation(mfcc1, mfcc2, nframes=correl_nframes)
max_k_index =
|
np.argmax(c)
|
numpy.argmax
|
from fidimag.micro import Zeeman
from fidimag.common import CuboidMesh
from fidimag.micro import Sim
import numpy as np
def varying_field(pos):
return (1.2 * pos[0], 2.3 * pos[1], 0)
def test_H0_is_indexable_or_callable():
"""
Test that an exception is raised if H0 is not indexable, and that an
exception is not raised if H0 is indexable.
"""
# Test for some different accepted types.
inputSuccess = ([0., 0., 1.],
|
np.array([0., 0., 1.])
|
numpy.array
|
import nengo
import numpy as np
import pytest
from nengo_loihi import BlockShape, add_params
def test_spike_units(Simulator, seed):
with nengo.Network(seed=seed) as model:
a = nengo.Ensemble(100, 1)
p = nengo.Probe(a.neurons)
with Simulator(model) as sim:
sim.run(0.1)
values = np.unique(sim.data[p])
assert values[0] == 0
assert values[1] == int(1.0 / sim.dt)
assert len(values) == 2
@pytest.mark.parametrize("dim", [1, 3])
def test_voltage_decode(allclose, Simulator, seed, plt, dim):
with nengo.Network(seed=seed) as model:
stim = nengo.Node(lambda t: [np.sin(2 * np.pi * t) / np.sqrt(dim)] * dim)
p_stim = nengo.Probe(stim, synapse=0.01)
a = nengo.Ensemble(100 * 3, dim, intercepts=nengo.dists.Uniform(-0.95, 0.95))
nengo.Connection(stim, a)
p_a = nengo.Probe(a, synapse=0.01)
with Simulator(model, precompute=True) as sim:
sim.run(1.0)
plt.plot(sim.trange(), sim.data[p_a])
plt.plot(sim.trange(), sim.data[p_stim])
assert allclose(sim.data[p_stim], sim.data[p_a], atol=0.3)
def test_repeated_probes(Simulator):
with nengo.Network() as net:
ens = nengo.Ensemble(1024, 1)
nengo.Probe(ens.neurons)
for _ in range(5):
with Simulator(net) as sim:
sim.run(0.1)
@pytest.mark.filterwarnings("ignore:Model is precomputable.")
@pytest.mark.parametrize("precompute", [True, False])
@pytest.mark.parametrize("probe_target", ["input", "voltage"])
def test_neuron_probes(precompute, probe_target, Simulator, seed, plt, allclose):
simtime = 0.3
with nengo.Network(seed=seed) as model:
stim = nengo.Node(lambda t: [np.sin(t * 2 * np.pi / simtime)])
a = nengo.Ensemble(
1,
1,
neuron_type=nengo.LIF(min_voltage=-1),
encoders=nengo.dists.Choice([[1]]),
max_rates=nengo.dists.Choice([100]),
intercepts=nengo.dists.Choice([0.0]),
)
nengo.Connection(stim, a, synapse=None)
p_stim = nengo.Probe(stim, synapse=0.005)
p_neurons = nengo.Probe(a.neurons, probe_target)
probe_synapse = nengo.Alpha(0.01)
p_stim_f = nengo.Probe(
stim, synapse=probe_synapse.combine(nengo.Lowpass(0.005))
)
p_neurons_f = nengo.Probe(a.neurons, probe_target, synapse=probe_synapse)
with Simulator(model, precompute=precompute) as sim:
sim.run(simtime)
scale = float(sim.data[p_neurons].max())
t = sim.trange()
x = sim.data[p_stim]
xf = sim.data[p_stim_f]
y = sim.data[p_neurons] / scale
yf = sim.data[p_neurons_f] / scale
plt.plot(t, x, label="stim")
plt.plot(t, xf, label="stim filt")
plt.plot(t, y, label="loihi")
plt.plot(t, yf, label="loihi filt")
plt.legend()
if probe_target == "input":
# shape of current input should roughly match stimulus
assert allclose(y, x, atol=0.4, rtol=0) # noisy, so rough match
assert allclose(yf, xf, atol=0.05, rtol=0) # tight match
elif probe_target == "voltage":
# check for voltage fluctuations (spiking) when stimulus is positive,
# and negative voltage when stimulus is most negative
spos = (t > 0.1 * simtime) & (t < 0.4 * simtime)
assert allclose(yf[spos], 0.5, atol=0.1, rtol=0.1)
assert y[spos].std() > 0.25
sneg = (t > 0.7 * simtime) & (t < 0.9 * simtime)
assert np.all(y[sneg] < 0)
def test_neuron_probe_with_synapse(Simulator, seed, allclose):
synapse = nengo.Lowpass(0.01)
with nengo.Network(seed=seed) as net:
ens = nengo.Ensemble(10, 1)
p_nosynapse = nengo.Probe(ens.neurons, synapse=None)
p_synapse = nengo.Probe(ens.neurons, synapse=synapse)
with Simulator(net) as sim:
sim.run(0.1)
assert allclose(sim.data[p_synapse], synapse.filt(sim.data[p_nosynapse]))
@pytest.mark.parametrize("precompute", [True, False])
def test_probe_filter_twice(precompute, plt, seed, Simulator):
with nengo.Network(seed=seed) as net:
stim = nengo.Node([1])
ens = nengo.Ensemble(100, 1)
probe = nengo.Probe(ens, synapse=0.01)
nengo.Connection(stim, ens)
with Simulator(net, precompute=precompute) as sim0:
sim0.run(0.04)
with Simulator(net, precompute=precompute) as sim1:
sim1.run(0.02)
sim1.run(0.02)
plt.plot(sim0.trange(), sim0.data[probe])
plt.plot(sim1.trange(), sim1.data[probe])
assert np.all(sim0.data[probe] == sim1.data[probe])
def test_probe_split_blocks(Simulator, seed, plt):
n_neurons = 80
gain = np.ones(n_neurons)
bias = np.linspace(0, 20, n_neurons)
simtime = 0.2
with nengo.Network(seed=seed) as net:
ens = nengo.Ensemble(n_neurons, 1, gain=gain, bias=bias)
probe = nengo.Probe(ens.neurons)
probe1_slice = slice(3, 33)
probe1 = nengo.Probe(ens.neurons[probe1_slice])
probe2_slice = slice(7, 52, 3)
probe2 = nengo.Probe(ens.neurons[probe2_slice])
probe3_slice = [2, 5, 17, 21, 36, 49, 52, 69, 73] # randomly chosen inds
probe3 = nengo.Probe(ens.neurons[probe3_slice])
# run without splitting ensemble
with Simulator(net) as sim1:
assert len(sim1.model.blocks) == 1
sim1.run(simtime)
# run with splitting ensemble
with net:
add_params(net)
net.config[ens].block_shape = BlockShape((5, 4), (10, 8))
with Simulator(net) as sim2:
assert len(sim2.model.blocks) == 4
sim2.run(simtime)
for k, sim in enumerate((sim1, sim2)):
plt.subplot(2, 1, k + 1)
plt.plot(bias, sim.data[probe].mean(axis=0))
plt.plot(bias[probe1_slice], sim.data[probe1].mean(axis=0))
plt.plot(bias[probe2_slice], sim.data[probe2].mean(axis=0), ".")
plt.plot(bias[probe3_slice], sim.data[probe3].mean(axis=0), "x")
# ensure rates increase and not everything is zero
for sim in (sim1, sim2):
diffs = np.diff(sim.data[probe].mean(axis=0))
assert (diffs >= 0).all() and (diffs > 1).sum() > 10
# ensure slices match unsliced probe
for sim in (sim1, sim2):
assert np.array_equal(sim.data[probe1], sim.data[probe][:, probe1_slice])
assert np.array_equal(sim.data[probe2], sim.data[probe][:, probe2_slice])
assert np.array_equal(sim.data[probe3], sim.data[probe][:, probe3_slice])
# ensure split and unsplit simulators match
for p in (probe, probe1, probe2, probe3):
assert np.array_equal(sim1.data[p], sim2.data[p])
def piecewise_net(n_pres, pres_time, seed):
values = np.linspace(-1, 1, n_pres)
with nengo.Network(seed=seed) as net:
add_params(net)
inp = nengo.Node(nengo.processes.PresentInput(values, pres_time), size_out=1)
ens = nengo.Ensemble(100, 1)
nengo.Connection(inp, ens)
net.probe = nengo.Probe(ens, synapse=nengo.Alpha(0.01))
node = nengo.Node(size_in=1)
nengo.Connection(ens, node, synapse=nengo.Alpha(0.01))
net.node_probe = nengo.Probe(node)
return net, values
@pytest.mark.parametrize("precompute", [False, True])
def test_clear_probes(Simulator, seed, plt, allclose, precompute):
n_pres = 5
pres_time = 0.1
net, values = piecewise_net(n_pres, pres_time, seed)
outputs = {"probe": [], "node": []}
with Simulator(net, precompute=precompute) as sim:
for _ in range(n_pres):
sim.clear_probes()
sim.run(pres_time)
outputs["probe"].append(np.copy(
|
np.squeeze(sim.data[net.probe], axis=-1)
|
numpy.squeeze
|
# -*- coding: utf-8 -*-
import skimage.io
import skimage.feature
import skimage.color
import skimage.transform
import skimage.util
import skimage.segmentation
import numpy
import matplotlib.pyplot as plt
import matplotlib.patches as mpatches
import SimpleITK as sitk
import cv2
def _generate_segments(im_orig, scale, sigma, min_size):
im_mask = skimage.segmentation.felzenszwalb(
skimage.util.img_as_float(im_orig), scale=scale, sigma=sigma,
min_size=min_size)
im_orig = numpy.append(
im_orig, numpy.zeros(im_orig.shape[:2])[:, :, numpy.newaxis], axis=2)
im_orig[:, :, 3] = im_mask
return im_orig
def _sim_colour(r1, r2):
return sum([min(a, b) for a, b in zip(r1["hist_c"], r2["hist_c"])])
def _sim_texture(r1, r2):
return sum([min(a, b) for a, b in zip(r1["hist_t"], r2["hist_t"])])
def _sim_size(r1, r2, imsize):
return 1.0 - (r1["size"] + r2["size"]) / imsize
def _sim_fill(r1, r2, imsize):
bbsize = (
(max(r1["max_x"], r2["max_x"]) - min(r1["min_x"], r2["min_x"]))
* (max(r1["max_y"], r2["max_y"]) - min(r1["min_y"], r2["min_y"]))
)
return 1.0 - (bbsize - r1["size"] - r2["size"]) / imsize
def _calc_sim(r1, r2, imsize):
return (_sim_colour(r1, r2) + _sim_texture(r1, r2)
+ _sim_size(r1, r2, imsize) + _sim_fill(r1, r2, imsize))
def _calc_colour_hist(img):
BINS = 25
hist = numpy.array([])
for colour_channel in (0, 1, 2):
c = img[:, colour_channel]
hist = numpy.concatenate(
[hist] + [numpy.histogram(c, BINS, (0.0, 255.0))[0]])
hist = hist / len(img)
return hist
def _calc_texture_gradient(img):
ret = numpy.zeros((img.shape[0], img.shape[1], img.shape[2]))
for colour_channel in (0, 1, 2):
ret[:, :, colour_channel] = skimage.feature.local_binary_pattern(
img[:, :, colour_channel], 8, 1.0)
return ret
def _calc_texture_hist(img):
BINS = 10
hist =
|
numpy.array([])
|
numpy.array
|
import torch
import numpy as np
from sklearn.cluster import SpectralClustering
from cogdl.utils import spmm
from .. import BaseModel, register_model
@register_model("agc")
class AGC(BaseModel):
r"""The AGC model from the `"Attributed Graph Clustering via Adaptive Graph Convolution"
<https://arxiv.org/abs/1906.01210>`_ paper
Args:
num_clusters (int) : Number of clusters.
max_iter (int) : Max iteration to increase k
"""
@staticmethod
def add_args(parser):
# fmt: off
parser.add_argument("--num-clusters", type=int, default=7)
parser.add_argument("--max-iter", type=int, default=10)
# fmt: on
@classmethod
def build_model_from_args(cls, args):
return cls(args.num_clusters, args.max_iter, args.cpu)
def __init__(self, num_clusters, max_iter, cpu):
super(AGC, self).__init__()
self.num_clusters = num_clusters
self.max_iter = max_iter
self.device = "cuda" if torch.cuda.is_available() and not cpu else "cpu"
def forward(self, data):
data = data.to(self.device)
self.num_nodes = data.x.shape[0]
graph = data
graph.add_remaining_self_loops()
graph.sym_norm()
graph.edge_weight = data.edge_weight * 0.5
pre_intra = 1e27
pre_feat = None
for t in range(1, self.max_iter + 1):
x = data.x
for i in range(t):
x = spmm(graph, x)
k = torch.mm(x, x.t())
w = (torch.abs(k) + torch.abs(k.t())) / 2
clustering = SpectralClustering(
n_clusters=self.num_clusters, assign_labels="discretize", random_state=0
).fit(w.detach().cpu())
clusters = clustering.labels_
intra = self.compute_intra(x.cpu().numpy(), clusters)
print("iter #%d, intra = %.4lf" % (t, intra))
if intra > pre_intra:
features_matrix = pre_feat
return features_matrix
pre_intra = intra
pre_feat = w
features_matrix = w
return features_matrix.cpu()
def compute_intra(self, x, clusters):
num_nodes = x.shape[0]
intra =
|
np.zeros(self.num_clusters)
|
numpy.zeros
|
import cv2
import pytest
import numpy as np
import mira.core as mc
import mira.detectors as md
import mira.datasets as mds
import mira.detectors.experimental.pixelwise as mpx
@pytest.mark.parametrize(
"detector_class",
[md.RetinaNet, md.EfficientDet, mpx.AggregatedSegmentation, md.FasterRCNN, md.DETR],
)
def test_detector_edge_cases(detector_class):
dataset = mds.load_shapes(width=256, height=256, n_scenes=1)
base = dataset[0]
dataset = dataset.assign(
scenes=[
base, # A regular case
base.assign(
annotations=[], image=
|
np.ones_like(base.image)
|
numpy.ones_like
|
import numpy as np
import cv2
from matplotlib import pyplot as plt
import os
from tools.CreateTFRecords.generic_tf_tools.resize import resize
from tools.ProjectionTools.Gated2RGB.lib.image_transformer import ImageTransformer
# You need to import the resize class from the first Calib read and projection tools.
import json
# Projections are created in the gated frame
def pad_gated_to_psm(img_in):
img_out = np.lib.pad(img_in, ((0, 0), (216, 296), (0, 0)), mode='constant', constant_values=0)
return img_out
def process_points(DEBUG=False):
with open(os.path.join(os.path.dirname(os.path.realpath(__file__)), 'coresponding_points.txt'), 'r') as f:
points = json.load(f)
X, Y = points['pos1'], points['pos2']
if DEBUG:
for x, y in zip(X, Y):
print(x, y)
print(len(X), len(Y))
return X,Y
class WarpingClass():
def __init__(self):
self.r = resize('RGB2Gatedv2')
self.X, self.Y = process_points()
dst_pts = np.asarray([[x, y] for x, y in self.X]).astype(np.float32).reshape(-1, 1, 2)
src_pts =
|
np.asarray([[x, y - 768] for x, y in self.Y])
|
numpy.asarray
|
import numpy as np
def get_data(model_type, TRAIN, words, EMB, enforce_gen, n_side_pixl):
import numpy as np
EMBEDDINGS, OBJ_ctr_sd_enf_gen = {}, []
# 0. Get dictionary of ALL our embedding words
EMB_dict = build_emb_dict(words, EMB)
# 1. Get the RELEVANT training instances (filtering for 'predicates' and 'complete_only' variables)
OBJ_ctr_sd, rel_ids, TRAIN_relevant = get_TRAIN_relevant(TRAIN, words)
# 2. get dictionaries WORDLISTS (INDICES for the embedding layer!)
EMBEDDINGS['obj_list'] = list(set(TRAIN_relevant['obj']))
EMBEDDINGS['subj_list'] = list(set(TRAIN_relevant['subj']))
EMBEDDINGS['pred_list'] = list(set(TRAIN_relevant['rel']))
allwords = np.concatenate((EMBEDDINGS['subj_list'], EMBEDDINGS['pred_list'], EMBEDDINGS['obj_list']), axis=0)
EMBEDDINGS['allwords_list'] = list(
set(allwords)) # IMPORTANT: The order of this list is what prevails later on as index for embeddings
# 3. Get INITIALIZATION embeddings
EMBEDDINGS['subj_EMB'] = wordlist2emb_matrix(EMBEDDINGS['subj_list'], EMB_dict)
EMBEDDINGS['pred_EMB'] = wordlist2emb_matrix(EMBEDDINGS['pred_list'], EMB_dict)
EMBEDDINGS['obj_EMB'] = wordlist2emb_matrix(EMBEDDINGS['obj_list'], EMB_dict)
EMBEDDINGS['allwords_EMB'] = wordlist2emb_matrix(EMBEDDINGS['allwords_list'],EMB_dict)
# 3.1. Get RANDOM embeddings (of the size of allwords_EMB)
EMBEDDINGS['allwords_EMB_rnd'] = get_random_EMB(EMBEDDINGS['allwords_EMB'])
EMBEDDINGS['subj_EMB_rnd'] = get_random_EMB(EMBEDDINGS['subj_EMB'])
EMBEDDINGS['pred_EMB_rnd'] = get_random_EMB(EMBEDDINGS['pred_EMB'])
EMBEDDINGS['obj_EMB_rnd'] = get_random_EMB(EMBEDDINGS['obj_EMB'])
# 3.2. get ONE-HOT embeddings:
EMBEDDINGS['subj_EMB_onehot'] = np.identity(len(EMBEDDINGS['subj_list']))
EMBEDDINGS['pred_EMB_onehot'] = np.identity(len(EMBEDDINGS['pred_list']))
EMBEDDINGS['obj_EMB_onehot'] = np.identity(len(EMBEDDINGS['obj_list']))
EMBEDDINGS['allwords_EMB_onehot'] = np.identity(len(EMBEDDINGS['allwords_list']))
# 4. Get X data (i.e., get the SEQUENCES of INDICES for the embedding layer)
X, X_extra, y, y_pixl, X_extra_enf_gen, X_enf_gen, y_enf_gen, y_enf_gen_pixl, \
idx_IN_X_and_y, idx_enf_gen = relevant_instances2X_and_y(model_type, TRAIN_relevant, EMBEDDINGS, enforce_gen,
n_side_pixl)
# 5. Get the OBJ_ctr_sd_enf_gen that we need for some performance measures!
if enforce_gen['eval'] is not None:
OBJ_ctr_sd_enf_gen = OBJ_ctr_sd[idx_enf_gen]
# 6. Finally, if we have REDUCED the X and y data by ENFORCING generalization (excluding instances) we have to reduce OBJ_ctr_sd and TRAIN_relevant accordingly
if enforce_gen['eval'] is not None:
for key in TRAIN_relevant:
TRAIN_relevant[key] = np.array(TRAIN_relevant[key])
TRAIN_relevant[key] = TRAIN_relevant[key][idx_IN_X_and_y]
OBJ_ctr_sd = OBJ_ctr_sd[idx_IN_X_and_y]
rel_ids = np.array(rel_ids)
rel_ids = rel_ids[idx_IN_X_and_y]
return X, X_extra, y, y_pixl, X_extra_enf_gen, X_enf_gen, y_enf_gen, y_enf_gen_pixl, rel_ids, OBJ_ctr_sd, OBJ_ctr_sd_enf_gen, EMBEDDINGS, TRAIN_relevant
def relevant_instances2X_and_y(model_type, TRAIN_relevant, EMBEDDINGS, enforce_gen, n_side_pixl):
# OUTPUT: the X and y data, gotten by converting each word into its corresponding index
print('Getting X and y data')
X_vars = ['subj_ctr_x', 'subj_ctr_y', 'subj_sd_x', 'subj_sd_y']
y_vars = ['obj_sd_x', 'obj_sd_y', 'obj_ctr_x', 'obj_ctr_y']
subj_list, pred_list, obj_list, allwords_list = EMBEDDINGS['subj_list'], EMBEDDINGS['pred_list'], EMBEDDINGS[
'obj_list'], EMBEDDINGS['allwords_list']
# get X:
X, X_enf_gen = {}, {}
X['subj'], X['pred'], X['obj'] = [], [], []
X_enf_gen['subj'], X_enf_gen['pred'], X_enf_gen['obj'] = [], [], []
for i in range(len(TRAIN_relevant['subj'])):
triplet = (TRAIN_relevant['subj'][i], TRAIN_relevant['rel'][i], TRAIN_relevant['obj'][i])
# append to the GENERALIZED set
if (enforce_gen['eval'] == 'triplets') and (triplet in enforce_gen['triplets']):
X_enf_gen['subj'].append(subj_list.index(TRAIN_relevant['subj'][i]))
X_enf_gen['pred'].append(pred_list.index(TRAIN_relevant['rel'][i]))
X_enf_gen['obj'].append(obj_list.index(TRAIN_relevant['obj'][i]))
elif (enforce_gen['eval'] == 'words') and any(word in enforce_gen['words'] for word in triplet):
X_enf_gen['subj'].append(subj_list.index(TRAIN_relevant['subj'][i]))
X_enf_gen['pred'].append(pred_list.index(TRAIN_relevant['rel'][i]))
X_enf_gen['obj'].append(obj_list.index(TRAIN_relevant['obj'][i]))
else: # if either the triplet/word is not generalized or we aren't enforcing generalization
X['subj'].append(subj_list.index(TRAIN_relevant['subj'][i]))
X['pred'].append(pred_list.index(TRAIN_relevant['rel'][i]))
X['obj'].append(obj_list.index(TRAIN_relevant['obj'][i]))
# Reshape
X['subj'] = np.array(X['subj']).reshape((-1, 1))
X['pred'] = np.array(X['pred']).reshape((-1, 1))
X['obj'] = np.array(X['obj']).reshape((-1, 1))
# FORMAT: if we have gotten some zero shot instances
if X_enf_gen['subj'] != []:
X_enf_gen['subj'] = np.array(X_enf_gen['subj']).reshape(
(-1, 1)) # get them in the right FORMAT for the merged (SEP) model!
X_enf_gen['pred'] = np.array(X_enf_gen['pred']).reshape((-1, 1))
X_enf_gen['obj'] = np.array(X_enf_gen['obj']).reshape((-1, 1))
else:
X_enf_gen['subj'], X_enf_gen['pred'], X_enf_gen['obj'] = None, None, None
# Get Y (if model_type = PIX we output the regular y besides y_pixl!)
y, y_pixl, y_enf_gen, idx_IN_X_and_y, idx_enf_gen, y_enf_gen_pixl = [], [], [], [], [], []
for i in range(len(TRAIN_relevant['subj'])):
y_new_row = []
for k in range(len(y_vars)):
y_new_row.extend([float(TRAIN_relevant[y_vars[k]][i])]) # IMPORTANT: We assume that the variables are NUMERIC
if model_type == 'PIX':
obj_sd_x, obj_sd_y = float(TRAIN_relevant['obj_sd_x'][i]), float(TRAIN_relevant['obj_sd_y'][i])
obj_ctr_x, obj_ctr_y = float(TRAIN_relevant['obj_ctr_x'][i]), float(TRAIN_relevant['obj_ctr_y'][i])
y_pixl_new_row = coord2pixel_indiv(obj_sd_x, obj_sd_y, obj_ctr_x, obj_ctr_y, n_side_pixl)
# get stuff for the generalzed setting:
triplet = (TRAIN_relevant['subj'][i], TRAIN_relevant['rel'][i], TRAIN_relevant['obj'][i])
if (enforce_gen['eval'] == 'triplets') and (triplet in enforce_gen['triplets']):
y_enf_gen.append(y_new_row)
if model_type == 'PIX':
y_enf_gen_pixl.append(y_pixl_new_row)
idx_enf_gen.append(i)
elif (enforce_gen['eval'] == 'words') and any(word in enforce_gen['words'] for word in triplet):
y_enf_gen.append(y_new_row)
if model_type == 'PIX':
y_enf_gen_pixl.append(y_pixl_new_row)
idx_enf_gen.append(i)
else: # NON GENERALIZED
y.append(y_new_row)
if model_type == 'PIX':
y_pixl.append(y_pixl_new_row)
idx_IN_X_and_y.append(i)
y = np.array(y)
y_enf_gen = np.array(y_enf_gen) if y_enf_gen != [] else None
if model_type == 'PIX':
y_pixl = np.array(y_pixl)
y_enf_gen_pixl = np.array(y_enf_gen_pixl) if y_enf_gen_pixl != [] else None
else:
y_pixl = [[[]]] # necessary because we get the index 0 of y_pixl (if model_type != 'PIX') to save memory in learn_and_evaluate()
print('We have gotten ' + str(len(idx_IN_X_and_y)) + ' instances (for both, train & test)')
# Get X_extra
X_extra, X_extra_enf_gen = [], []
if X_vars != []:
for i in range(len(TRAIN_relevant['subj'])):
X_extra_new_row = []
for k in range(len(X_vars)): # we already ASSUME that we have at least one y-variable
X_extra_new_row.extend(
[float(TRAIN_relevant[X_vars[k]][i])]) # IMPORTANT: We assume that the variables are NUMERIC
# get stuff for the generalized:
triplet = (TRAIN_relevant['subj'][i], TRAIN_relevant['rel'][i], TRAIN_relevant['obj'][i])
if (enforce_gen['eval'] == 'triplets') and (triplet in enforce_gen['triplets']):
X_extra_enf_gen.append(X_extra_new_row)
elif (enforce_gen['eval'] == 'words') and any(word in enforce_gen['words'] for word in triplet):
X_extra_enf_gen.append(X_extra_new_row)
else:
X_extra.append(X_extra_new_row)
X_extra = np.array(X_extra) if X_extra != [] else None # IMPORTANT: we only make it a numpy array if we have something, because we use == [] as condition in models_learn
X_extra_enf_gen = np.array(X_extra_enf_gen) if X_extra_enf_gen != [] else None
return X, X_extra, y, y_pixl, X_extra_enf_gen, X_enf_gen, y_enf_gen, y_enf_gen_pixl, idx_IN_X_and_y, idx_enf_gen
def get_TRAIN_relevant(TRAIN, words):
# IMPORTANT: we preserve the ORDER of TRAIN (so that we can recover information afterwards)
TRAIN_relevant, rel_ids, OBJ_ctr_sd = {}, [], []
print('Getting *relevant* instances, from a total of: ' + str(len(TRAIN['subj'])))
var_names = [key for key in TRAIN]
# INITIALIZE TRAIN_relavant
for varname in var_names:
TRAIN_relevant[varname] = []
for i in range(len( TRAIN['subj'] )): # Samples loop
we_have_it = True if ((TRAIN['subj'][i] in words) and (TRAIN['rel'][i] in words) and (TRAIN['obj'][i] in words)) else False # if we have the complete triplet
if we_have_it == True:
for varname in var_names:
TRAIN_relevant[varname].append(TRAIN[varname][i])
rel_ids.append(TRAIN['rel_id'][i])
OBJ_ctr_sd.append([TRAIN['img_idx'][i], TRAIN['rel_id'][i], TRAIN['subj'][i], TRAIN['rel'][i],
TRAIN['obj'][i], TRAIN['subj_sd_x'][i], TRAIN['subj_sd_y'][i],
TRAIN['subj_ctr_x'][i], TRAIN['subj_ctr_y'][i], TRAIN['obj_sd_x'][i],
TRAIN['obj_sd_y'][i], TRAIN['obj_ctr_x'][i], TRAIN['obj_ctr_y'][i]])
OBJ_ctr_sd = np.array(OBJ_ctr_sd)
print('We have gotten ' + str(len(TRAIN_relevant['subj'])) + ' RELEVANT instances')
return OBJ_ctr_sd, rel_ids, TRAIN_relevant
def get_random_EMB(actual_EMB):
# Returns embedding matrix of the original shape with random normal vectors (dimension-wise)
mu, sigma, vec_size = np.mean(actual_EMB), np.mean(np.std(actual_EMB, axis=0)), len(actual_EMB[0, :])
rand_EMB = []
for i in range(actual_EMB.shape[0]): # build a dictionary of random vectors
rand_EMB.append(np.random.normal(mu, sigma, vec_size))
rand_EMB = np.array(rand_EMB)
return rand_EMB
def coord2pixel_indiv(obj_sd_x, obj_sd_y, obj_ctr_x, obj_ctr_y, n_side_pixl):
'''
This function works with an individual example (extending it to many examples, where e.g., obj_sd_x is a vector, is easy)
:param obj_sd_x (and the rest): real number (not vectors!)
:param n_side_pixl: number of pixels as output (hyperparameter)
:return y_pixl: matrix of pixels, i.e., a 2D tensor (n_side_pixl, n_side_pixl)
'''
# continuous bounding box corners (prevent problems of predictions outside [0,1])
A_left_x, A_right_x = max((obj_ctr_x - obj_sd_x), 0), min((obj_ctr_x + obj_sd_x), 1)
A_low_y, A_top_y = min((obj_ctr_y + obj_sd_y), 1), max((obj_ctr_y - obj_sd_y), 0)
# translate continuous bounding box corners into indices in a n_side_pixl x n_side_pixl matrix
i_left, i_right = np.rint( (n_side_pixl - 1)*A_left_x).astype(np.int), np.rint((n_side_pixl - 1)*A_right_x).astype(np.int)
j_low, j_top = np.rint((n_side_pixl - 1)*A_low_y).astype(np.int), np.rint((n_side_pixl - 1)*A_top_y).astype(np.int)
pixl_matr = np.zeros( (n_side_pixl, n_side_pixl) )
# add ones inside of the bounding box
i_range = range( i_left, i_right )
i_range = [i_left] if ((i_left == i_right) or (i_range == [])) else i_range # AVOID THE CASE where width is 0 AND i_range=[] (as upper bound < lower bound)
j_range = range( j_top, j_low )
j_range = [j_low] if ((j_low == j_top) or (j_range == [])) else j_range # AVOID THE CASE where height is 0 AND i_range=[] (as upper bound < lower bound)
pixl_matr[ np.array(i_range)[:, None], np.array(j_range)] = 1 # (IMPORTANT: indices must be np.arrays) put a 1 everywhere inside of the bounding box
pixl_matr = pixl_matr.reshape((-1))
return pixl_matr
def pixl_idx2coord_all_examples(y_pixl):
'''
Transforms the whole set of predicted matrices y_pixl into their continuous CENTER coordinates (Obj_ctr)
:param y_pixl: array of MATRICES with predicted heatmaps (pixels). Each matrix = 1 example
:return: PRED_obj_ctr_x, PRED_obj_ctr_y: arrays of length = number of examples
'''
PRED_obj_ctr_x, PRED_obj_ctr_y = [], []
n_side_pixl = y_pixl.shape[1] #get automatically the number of pixels from the pixel matrix side
for i in range( y_pixl.shape[0] ): # loop on number of examples
idx_maximums = get_maximums_idx(y_pixl[i]) # get indices of maximum (allow for multiple of them)
ctr_x, ctr_y = pixl_idx2coord_indiv(idx_maximums, n_side_pixl) # transform pixel indices into continuous coordinates
PRED_obj_ctr_x.append(ctr_x)
PRED_obj_ctr_y.append(ctr_y)
PRED_obj_ctr_x, PRED_obj_ctr_y =
|
np.array(PRED_obj_ctr_x)
|
numpy.array
|
import math
import numpy as np
import torch
from torch import optim
from torch import nn
import torch.utils.data
from torch.nn import (
BatchNorm1d,
Dropout,
LeakyReLU,
Linear,
Module,
ReLU,
Sequential,
Sigmoid,
)
from torch.autograd import Variable
import warnings
from .data_sampler import DataSampler
from ctgan.synthesizers import CTGANSynthesizer
from snsynth.preprocessors.data_transformer import BaseTransformer
from .privacy_utils import weights_init, pate, moments_acc
class Discriminator(Module):
def __init__(self, input_dim, discriminator_dim, loss, pac=10):
super(Discriminator, self).__init__()
torch.cuda.manual_seed(0)
torch.manual_seed(0)
dim = input_dim * pac
# print ('now dim is {}'.format(dim))
self.pac = pac
self.pacdim = dim
seq = []
for item in list(discriminator_dim):
seq += [Linear(dim, item), LeakyReLU(0.2), Dropout(0.5)]
dim = item
seq += [Linear(dim, 1)]
if loss == "cross_entropy":
seq += [Sigmoid()]
self.seq = Sequential(*seq)
def dragan_penalty(self, real_data, device="cpu", pac=10, lambda_=10):
# real_data = torch.from_numpy(real_data).to(device)
alpha = (
torch.rand(real_data.shape[0], 1, device=device)
.squeeze()
.expand(real_data.shape[0])
)
delta = torch.normal(
mean=0.0, std=float(pac), size=real_data.shape, device=device
) # 0.5 * real_data.std() * torch.rand(real_data.shape)
x_hat = Variable(
(alpha * real_data.T + (1 - alpha) * (real_data + delta).T).T,
requires_grad=True,
)
pred_hat = self(x_hat.float())
gradients = torch.autograd.grad(
outputs=pred_hat,
inputs=x_hat,
grad_outputs=torch.ones(pred_hat.size(), device=device),
create_graph=True,
retain_graph=True,
only_inputs=True,
)[0]
dragan_penalty = lambda_ * ((gradients.norm(2, dim=1) - 1) ** 2).mean()
return dragan_penalty
def forward(self, input):
assert input.size()[0] % self.pac == 0
return self.seq(input.view(-1, self.pacdim))
class Residual(Module):
def __init__(self, i, o):
super(Residual, self).__init__()
self.fc = Linear(i, o)
self.bn = BatchNorm1d(o)
self.relu = ReLU()
def forward(self, input):
out = self.fc(input)
out = self.bn(out)
out = self.relu(out)
return torch.cat([out, input], dim=1)
class Generator(Module):
def __init__(self, embedding_dim, generator_dim, data_dim):
super(Generator, self).__init__()
dim = embedding_dim
seq = []
for item in list(generator_dim):
seq += [Residual(dim, item)]
dim += item
seq.append(Linear(dim, data_dim))
self.seq = Sequential(*seq)
def forward(self, input):
data = self.seq(input)
return data
class PATECTGAN(CTGANSynthesizer):
def __init__(
self,
embedding_dim=128,
generator_dim=(256, 256),
discriminator_dim=(256, 256),
generator_lr=2e-4,
generator_decay=1e-6,
discriminator_lr=2e-4,
discriminator_decay=1e-6,
batch_size=500,
discriminator_steps=1,
log_frequency=False,
verbose=False,
epochs=300,
pac=1,
cuda=True,
epsilon=1,
binary=False,
regularization=None,
loss="cross_entropy",
teacher_iters=5,
student_iters=5,
sample_per_teacher=1000,
delta=None,
noise_multiplier=1e-3,
preprocessor_eps=1,
moments_order=100,
category_epsilon_pct=0.1,
):
assert batch_size % 2 == 0
self._embedding_dim = embedding_dim
self._generator_dim = generator_dim
self._discriminator_dim = discriminator_dim
self._generator_lr = generator_lr
self._generator_decay = generator_decay
self._discriminator_lr = discriminator_lr
self._discriminator_decay = discriminator_decay
self._batch_size = batch_size
self._discriminator_steps = discriminator_steps
self._log_frequency = log_frequency
self._verbose = verbose
self._epochs = epochs
self.pac = pac
self.preprocessor_eps = preprocessor_eps
self.epsilon = epsilon - preprocessor_eps
self._category_epsilon_pct = category_epsilon_pct
self.verbose = verbose
self.loss = loss
# PATE params
self.regularization = regularization if self.loss != "wasserstein" else "dragan"
self.teacher_iters = teacher_iters
self.student_iters = student_iters
self.pd_cols = None
self.pd_index = None
self.binary = binary
self.sample_per_teacher = sample_per_teacher
self.noise_multiplier = noise_multiplier
self.moments_order = moments_order
self.delta = delta
if not cuda or not torch.cuda.is_available():
device = "cpu"
elif isinstance(cuda, str):
device = cuda
else:
device = "cuda"
self._device = torch.device(device)
if self._log_frequency:
warnings.warn(
"log_frequency is selected. This may result in oversampling frequent "
"categories, which could cause privacy leaks."
)
def train(
self,
data,
categorical_columns=None,
ordinal_columns=None,
update_epsilon=None,
transformer=BaseTransformer,
continuous_columns_lower_upper=None,
):
if update_epsilon:
self.epsilon = update_epsilon - self.preprocessor_eps
for col in categorical_columns:
if str(data[col].dtype).startswith("float"):
raise ValueError(
"It looks like you are passing in a vector of continuous values"
f"to a categorical column at [{col}]."
"Please discretize and pass in categorical columns with"
"unsigned integer or string category names."
)
sample_per_teacher = (
self.sample_per_teacher if self.sample_per_teacher < len(data) else 1000
)
self.num_teachers = int(len(data) / sample_per_teacher) + 1
self._transformer = transformer(self.preprocessor_eps)
self._transformer.fit(
data,
discrete_columns=categorical_columns,
continuous_columns_lower_upper=continuous_columns_lower_upper,
)
train_data = self._transformer.transform(data)
data_partitions = np.array_split(train_data, self.num_teachers)
data_dim = self._transformer.output_dimensions
sampler_eps = 0.0
if categorical_columns and self._category_epsilon_pct:
sampler_eps = self.epsilon * self._category_epsilon_pct
per_col_sampler_eps = sampler_eps / len(categorical_columns)
self.epsilon = self.epsilon - sampler_eps
else:
per_col_sampler_eps = None
self.cond_generator = DataSampler(
train_data,
self._transformer.output_info_list,
self._log_frequency,
per_column_epsilon=per_col_sampler_eps,
)
spent = self.cond_generator.total_spent
if spent > sampler_eps and not np.isclose(spent, sampler_eps):
raise AssertionError(
f"The data sampler used {spent} epsilon and was budgeted for {sampler_eps}"
)
# create conditional generator for each teacher model
# Note: Previously, there existed a ConditionalGenerator object in CTGAN
# - that functionality has been subsumed by DataSampler, but switch is
# essentially 1 for 1
# don't need to count eps for each teacher, because these are disjoint partitions
cached_probs = self.cond_generator.discrete_column_category_prob
cond_generator = [
DataSampler(
d,
self._transformer.output_info_list,
self._log_frequency,
per_column_epsilon=None,
discrete_column_category_prob=cached_probs,
)
for d in data_partitions
]
self._generator = Generator(
self._embedding_dim + self.cond_generator.dim_cond_vec(),
self._generator_dim,
data_dim,
).to(self._device)
discriminator = Discriminator(
data_dim + self.cond_generator.dim_cond_vec(),
self._discriminator_dim,
self.loss,
self.pac,
).to(self._device)
student_disc = discriminator
student_disc.apply(weights_init)
teacher_disc = [discriminator for i in range(self.num_teachers)]
for i in range(self.num_teachers):
teacher_disc[i].apply(weights_init)
optimizerG = optim.Adam(
self._generator.parameters(),
lr=self._generator_lr,
betas=(0.5, 0.9),
weight_decay=self._generator_decay,
)
optimizer_s = optim.Adam(student_disc.parameters(), lr=2e-4, betas=(0.5, 0.9))
optimizer_t = [
optim.Adam(
teacher_disc[i].parameters(),
lr=self._discriminator_lr,
betas=(0.5, 0.9),
weight_decay=self._discriminator_decay,
)
for i in range(self.num_teachers)
]
noise_multiplier = self.noise_multiplier
alphas = torch.tensor(
[0.0 for i in range(self.moments_order)], device=self._device
)
l_list = 1 + torch.tensor(range(self.moments_order), device=self._device)
eps = torch.zeros(1)
mean = torch.zeros(self._batch_size, self._embedding_dim, device=self._device)
std = mean + 1
real_label = 1
fake_label = 0
criterion = nn.BCELoss() if (self.loss == "cross_entropy") else self.w_loss
if self.verbose:
print(
"using loss {} and regularization {}".format(
self.loss, self.regularization
)
)
iteration = 0
if self.delta is None:
self.delta = 1 / (train_data.shape[0] * np.sqrt(train_data.shape[0]))
while eps.item() < self.epsilon:
iteration += 1
eps = min((alphas - math.log(self.delta)) / l_list)
if eps.item() > self.epsilon:
if iteration == 1:
raise ValueError(
"Inputted epsilon parameter is too small to"
+ " create a private dataset. Try increasing epsilon and rerunning."
)
break
# train teacher discriminators
for t_2 in range(self.teacher_iters):
for i in range(self.num_teachers):
partition_data = data_partitions[i]
data_sampler = DataSampler(
partition_data,
self._transformer.output_info_list,
self._log_frequency,
per_column_epsilon=None,
discrete_column_category_prob=cached_probs,
)
fakez = torch.normal(mean, std=std).to(self._device)
condvec = cond_generator[i].sample_condvec(self._batch_size)
if condvec is None:
c1, m1, col, opt = None, None, None, None
real = data_sampler.sample_data(self._batch_size, col, opt)
else:
c1, m1, col, opt = condvec
c1 = torch.from_numpy(c1).to(self._device)
m1 = torch.from_numpy(m1).to(self._device)
fakez = torch.cat([fakez, c1], dim=1)
perm = np.arange(self._batch_size)
np.random.shuffle(perm)
real = data_sampler.sample_data(
self._batch_size, col[perm], opt[perm]
)
c2 = c1[perm]
fake = self._generator(fakez)
fakeact = self._apply_activate(fake)
real = torch.from_numpy(real.astype("float32")).to(self._device)
if c1 is not None:
fake_cat = torch.cat([fakeact, c1], dim=1)
real_cat = torch.cat([real, c2], dim=1)
else:
real_cat = real
fake_cat = fake
optimizer_t[i].zero_grad()
y_all = torch.cat(
[teacher_disc[i](fake_cat), teacher_disc[i](real_cat)]
)
label_fake = torch.full(
(int(self._batch_size / self.pac), 1),
fake_label,
dtype=torch.float,
device=self._device,
)
label_true = torch.full(
(int(self._batch_size / self.pac), 1),
real_label,
dtype=torch.float,
device=self._device,
)
labels = torch.cat([label_fake, label_true])
error_d = criterion(y_all.squeeze(), labels.squeeze())
error_d.backward()
if self.regularization == "dragan":
pen = teacher_disc[i].dragan_penalty(
real_cat, device=self._device
)
pen.backward(retain_graph=True)
optimizer_t[i].step()
###
# train student discriminator
for t_3 in range(self.student_iters):
data_sampler = DataSampler(
train_data,
self._transformer.output_info_list,
self._log_frequency,
per_column_epsilon=None,
discrete_column_category_prob=cached_probs,
)
fakez = torch.normal(mean=mean, std=std)
condvec = self.cond_generator.sample_condvec(self._batch_size)
if condvec is None:
c1, m1, col, opt = None, None, None, None
real = data_sampler.sample_data(self._batch_size, col, opt)
else:
c1, m1, col, opt = condvec
c1 = torch.from_numpy(c1).to(self._device)
m1 = torch.from_numpy(m1).to(self._device)
fakez = torch.cat([fakez, c1], dim=1)
perm =
|
np.arange(self._batch_size)
|
numpy.arange
|
#!/usr/bin/python3 -u
import os
import json
import re
import subprocess
import nibabel
from dipy.io import read_bvals_bvecs
from dipy.core.gradients import gradient_table
import math
import numpy as np
#import matplotlib
#import imageio
from scipy.ndimage import zoom
from json import encoder
encoder.FLOAT_REPR = lambda o: format(o, '.2f')
with open('config.json') as config_json:
config = json.load(config_json)
#Returns the unit vector of the vector.
def unit_vector(vector):
return vector /
|
np.linalg.norm(vector)
|
numpy.linalg.norm
|
import cv2
import numpy as np
import matplotlib.pyplot as plt
from skimage.filters import gabor
import mahotas as mt
import pandas as pd
from glob import glob
from skimage.feature import local_binary_pattern
def fun1(img_mask,Label):
count = 0
gaborenergy1 = []
gaborentropy1 = []
w1=[]
h1=[]
area1 = []
perimeter1 = []
rectArea1= []
aspectratio1 = []
rectangularity1 = []
circularity1 = []
equi_diameter1 = []
red_mean1 = []
green_mean1 = []
blue_mean1 = []
red_var1 = []
blue_var1 = []
green_var1 = []
contrast1 = []
correlation1 = []
inversedifferencemoments1 = []
entropy1 = []
Label1 = []
LBP = []
extent1= []
solidity1=[]
hull_area1=[]
equi_diameter1 = []
radius = 3
no_points = 8 * radius
img_names = glob(img_mask)
iasd=0
for fn in img_names:
#print('processing %s...' % fn,i)
print(iasd,end="\t")
iasd=iasd+1
img = cv2.imread(fn)
#cv2.imshow("original",img)
####### Converting image to grayscale #########
gs = cv2.cvtColor(img, cv2.COLOR_RGB2GRAY)
# GABOR filter....................................................................
gaborFilt_real, gaborFilt_imag = gabor(gs, frequency=0.6)
gaborFilt = (gaborFilt_real ** 2 + gaborFilt_imag ** 2) // 2
#fig, ax = plt.subplots(1, 3)
#ax[0].imshow(gaborFilt_real, cmap='gray')
#ax[1].imshow(gaborFilt_imag, cmap='gray')
#ax[2].imshow(gaborFilt, cmap='gray')
#plt.show()
# energy and entropy of GABOR filter response......................................
gabor_hist, _ = np.histogram(gaborFilt, 8)
gabor_hist = np.array(gabor_hist, dtype=float)
gabor_prob = np.divide(gabor_hist, np.sum(gabor_hist))
gabor_energy = np.sum(gabor_prob ** 2)
gabor_entropy = -np.sum(np.multiply(gabor_prob, np.log2(gabor_prob)))
#print("gabor_energy:" + str(gabor_energy))
#print("gabor_entropy:" + str(gabor_entropy))
count = count+1
#print(count)
#########################local_binary_pattern#########################
lbp = local_binary_pattern(gs, no_points, radius, method='uniform')
###### Smoothing image using Guassian filter
blur = cv2.GaussianBlur(gs, (25,25),0)
#print(gs.shape)
####Adaptive image thresholding using Otsu's thresholding method
ret_otsu,im_bw_otsu = cv2.threshold(blur,0,255,cv2.THRESH_BINARY_INV+cv2.THRESH_OTSU)
#cv2.imshow("Thresholding",im_bw_otsu)
####Boundary extraction using sobel filters
sobelx64f = cv2.Sobel(im_bw_otsu,cv2.CV_64F,1,0,ksize=5)
abs_sobel64f = np.absolute(sobelx64f)
sobel_8u = np.uint8(abs_sobel64f)
#cv2.imshow("Boundary Extraction",abs_sobel64f)
ret_sobel,im_bw_sobel = cv2.threshold(sobel_8u,1,255,cv2.THRESH_BINARY)
#cv2.imshow("boundary",im_bw_sobel)
kernel_edge = np.ones((15,15),np.uint8)
closing_edge = cv2.morphologyEx(im_bw_sobel, cv2.MORPH_CLOSE, kernel_edge)
#cv2.imshow("Closing Edge",closing_edge)
#cv2.imshow("Boundary ",im_bw_otsu)
##### Boundary extraction using contours
ret, thresh = cv2.threshold(gs, 127, 255, 0)
contours, hierarchy = cv2.findContours(im_bw_otsu, cv2.RETR_TREE, cv2.CHAIN_APPROX_SIMPLE)
len(contours)
cnt=contours[0]
len(cnt)
plottedContour = cv2.drawContours(gs,contours,-1,(0,255,0),10)
#cv2.imshow("Plotted Contour",plottedContour)
##### Shape based features
M = cv2.moments(cnt)
#print("MOments: ",M)
area = cv2.contourArea(cnt)
#print("Area",area)
perimeter = cv2.arcLength(cnt,True)
#print("Perimeter",perimeter)
rect = cv2.minAreaRect(cnt)
box = cv2.boxPoints(rect)
box = np.int0(box)
contours_im = cv2.drawContours(im_bw_otsu,[box],0,(255,255,255),2)
#cv2.imshow("best fit rect",contours_im)
#ellipse = cv2.fitEllipse(cnt)
#im = cv2.ellipse(im_bw_otsu,ellipse,(255,255,255),2)
#cv2.imshow("")
x,y,w,h = cv2.boundingRect(cnt)
aspect_ratio = float(w)/h
#print("Aspect Ratio: ",aspect_ratio)
######### Extent#############
rect_area = w * h
extent = float(area) / rect_area
######### solidity #############
hull = cv2.convexHull(cnt)
hull_area = cv2.contourArea(hull)
if hull_area != 0:
solidity = float(area) / hull_area
else:
solidity = 0
####Shape based features calculated - Aspect ratio, rectangularity, circularity
if area !=0:
rectangularity =w*h/area
circularity = ((perimeter) ** 2) / area
else:
rectangularity=0
circularity = 0
#print("rectangularity: ",rectangularity)
#print("circularity: ",circularity)
equi_diameter =
|
np.sqrt(4*area/np.pi)
|
numpy.sqrt
|
# Copyright (c) 2017, 2020 ADLINK Technology Inc.
#
# This program and the accompanying materials are made available under the
# terms of the Eclipse Public License 2.0 which is available at
# http://www.eclipse.org/legal/epl-2.0, or the Apache License, Version 2.0
# which is available at https://www.apache.org/licenses/LICENSE-2.0.
#
# SPDX-License-Identifier: EPL-2.0 OR Apache-2.0
#
# Contributors:
# ADLINK zenoh team, <<EMAIL>>
# Examples
# HOST1 $ zn_sub -l tcp/<IP HOST1>:7447
# HOST2 $ zn_pub -e tcp/<IP HOST1>:7447
import sys
from datetime import datetime
import argparse
import zenoh
from zenoh.net import config, SubInfo, Reliability, SubMode
import time
import numpy as np
# --- Command line argument parsing --- --- --- --- --- ---
parser = argparse.ArgumentParser(
prog='zn_sub',
description='zenoh-net sub example')
parser.add_argument('--mode', '-m', dest='mode',
choices=['peer', 'client'],
type=str,
help='The zenoh session mode.')
parser.add_argument('--peer', '-e', dest='peer',
metavar='LOCATOR',
action='append',
type=str,
help='Peer locators used to initiate the zenoh session.')
parser.add_argument('--listener', '-l', dest='listener',
metavar='LOCATOR',
action='append',
type=str,
help='Locators to listen on.')
parser.add_argument('--selector', '-s', dest='selector',
default='/demo/example/**',
type=str,
help='The selection of resources to subscribe.')
parser.add_argument('--config', '-c', dest='config',
metavar='FILE',
type=str,
help='A configuration file.')
args = parser.parse_args()
conf = zenoh.config_from_file(args.config) if args.config is not None else {}
if args.mode is not None:
conf["mode"] = args.mode
if args.peer is not None:
conf["peer"] = ",".join(args.peer)
if args.listener is not None:
conf["listener"] = ",".join(args.listener)
selector = args.selector
# zenoh-net code --- --- --- --- --- --- --- --- --- --- ---
# def listener(sample):
def listener(consumed_data):
############
############ For IMAGE ONLY
t0_decoding = time.time()
deserialized_bytes =
|
np.frombuffer(consumed_data.payload, dtype=np.int8)
|
numpy.frombuffer
|
#!/usr/bin/env python
from __future__ import division
import numpy as np
import cv2
from optparse import OptionParser
import copy
from scipy import optimize
import data_fit
##############################################################################################
# Circle
def estimate_circle_from_data_points(x_m, y_m):
model = data_fit.models.CircleModel()
data = np.zeros(len(x_m))
model.fit(data, [np.array(x_m), np.array(y_m)])
model.parameters['radius'] = np.abs(model.parameters['radius'])
model.parameters['center_x'] = np.abs(model.parameters['center_x'])
model.parameters['center_y'] = np.abs(model.parameters['center_y'])
print ('Circle Estimates')
print ('Center (x,y): ', model.parameters['center_x'], model.parameters['center_y'])
print ('Radius: ', model.parameters['radius'])
return model.parameters['center_x'], model.parameters['center_y'], model.parameters['radius']
# Iterative Optimization Method
#print 'Fitting Linear Model with: scipy.optimize.leastsq'
def f(parameter_values, parameter_names):
self.set_parameters(parameter_names, parameter_values)
ans = self.get_errors(data, inputs)
if len(ans.shape) == 2 and ans.shape[0] == 1:
ans = ans.reshape(ans.shape[1])
return ans
parameter_values = []
parameter_names = []
for name, value in self.parameters.items():
if name in ignore_parameter_names:
continue
else:
parameter_values.append(value)
parameter_names.append(name)
optimize.leastsq(f, parameter_values, parameter_names)
class ClickCircle(object):
def __init__(self, filename):
self.image = cv2.imread(filename)
self.display_name = "Display"
cv2.namedWindow(self.display_name)
cv2.setMouseCallback(self.display_name, self.on_mouse_click)
self.circle_points_x = []
self.circle_points_y = []
self.circle_fit = None
def on_mouse_click(self, event, x, y, flags, param):
if event == cv2.EVENT_LBUTTONUP:
self.circle_points_x.append(x)
self.circle_points_y.append(y)
if len(self.circle_points_x) >= 3:
x,y,R = estimate_circle_from_data_points(self.circle_points_x, self.circle_points_y)
self.circle_fit = [x,y,R]
def draw(self):
canvas = copy.copy(self.image)
for i in range(len(self.circle_points_x)):
cv2.circle(canvas, (self.circle_points_x[i], self.circle_points_y[i]), 2, [0,0,255], 2)
if self.circle_fit is not None:
cv2.circle(canvas, (int(self.circle_fit[0]), int(self.circle_fit[1])), int(self.circle_fit[2]), [0,255,0], 2)
cv2.imshow("Display", canvas)
#cv2.waitKey(1)
def run(self):
while (cv2.waitKey(30) != 27):
self.draw()
cv.destroyAllWindows();
##############################################################################################
# Ellipse
def estimate_ellipse_from_data_points(x_m, y_m):
points = []
for i in range(len(x_m)):
points.append((x_m[i], y_m[i]))
ellipse = cv2.fitEllipse(
|
np.array(points)
|
numpy.array
|
from typing import Text, Dict, List, Optional
import numpy as np
import pytest
from rasa.core.featurizers.single_state_featurizer import SingleStateFeaturizer
from rasa.core.featurizers.single_state_featurizer import (
IntentTokenizerSingleStateFeaturizer,
)
from rasa.core.featurizers.tracker_featurizers import (
TrackerFeaturizer as TrackerFeaturizer,
)
from rasa.core.featurizers.tracker_featurizers import MaxHistoryTrackerFeaturizer
from rasa.core.featurizers.tracker_featurizers import IntentMaxHistoryTrackerFeaturizer
from rasa.core.featurizers.tracker_featurizers import FullDialogueTrackerFeaturizer
from rasa.shared.core.domain import Domain
from tests.core.utilities import user_uttered
from rasa.shared.nlu.training_data.features import Features
from rasa.shared.nlu.constants import INTENT, ACTION_NAME
from rasa.shared.core.constants import (
ACTION_LISTEN_NAME,
ACTION_UNLIKELY_INTENT_NAME,
USER,
PREVIOUS_ACTION,
)
from rasa.shared.core.events import ActionExecuted
from rasa.shared.core.trackers import DialogueStateTracker
from rasa.utils.tensorflow.constants import LABEL_PAD_ID
from rasa.core.exceptions import InvalidTrackerFeaturizerUsageError
def test_fail_to_load_non_existent_featurizer():
assert TrackerFeaturizer.load("non_existent_class") is None
def test_persist_and_load_tracker_featurizer(tmp_path: Text, moodbot_domain: Domain):
state_featurizer = SingleStateFeaturizer()
state_featurizer.prepare_for_training(moodbot_domain)
tracker_featurizer = MaxHistoryTrackerFeaturizer(state_featurizer)
tracker_featurizer.persist(tmp_path)
loaded_tracker_featurizer = TrackerFeaturizer.load(tmp_path)
assert loaded_tracker_featurizer is not None
assert loaded_tracker_featurizer.state_featurizer is not None
def test_convert_action_labels_to_ids(domain: Domain):
trackers_as_actions = [
["utter_greet", "utter_channel"],
["utter_greet", "utter_default", "utter_goodbye"],
]
tracker_featurizer = TrackerFeaturizer()
actual_output = tracker_featurizer._convert_labels_to_ids(
trackers_as_actions, domain
)
expected_output = np.array(
[
np.array(
[
domain.action_names_or_texts.index("utter_greet"),
domain.action_names_or_texts.index("utter_channel"),
],
),
np.array(
[
domain.action_names_or_texts.index("utter_greet"),
domain.action_names_or_texts.index("utter_default"),
domain.action_names_or_texts.index("utter_goodbye"),
],
),
],
)
assert expected_output.size == actual_output.size
for expected_array, actual_array in zip(expected_output, actual_output):
assert np.all(expected_array == actual_array)
def test_convert_intent_labels_to_ids(domain: Domain):
trackers_as_intents = [
["next_intent", "nlu_fallback", "out_of_scope", "restart"],
["greet", "hello", "affirm"],
]
tracker_featurizer = IntentMaxHistoryTrackerFeaturizer()
actual_labels = tracker_featurizer._convert_labels_to_ids(
trackers_as_intents, domain
)
expected_labels = np.array(
[
[
domain.intents.index("next_intent"),
domain.intents.index("nlu_fallback"),
domain.intents.index("out_of_scope"),
domain.intents.index("restart"),
],
[
domain.intents.index("greet"),
domain.intents.index("hello"),
domain.intents.index("affirm"),
LABEL_PAD_ID,
],
],
)
assert expected_labels.size == actual_labels.size
assert expected_labels.shape == actual_labels.shape
assert np.all(expected_labels == actual_labels)
def test_featurize_trackers_raises_on_missing_state_featurizer(domain: Domain):
tracker_featurizer = TrackerFeaturizer()
with pytest.raises(InvalidTrackerFeaturizerUsageError):
tracker_featurizer.featurize_trackers([], domain, precomputations=None)
def compare_featurized_states(
states1: List[Dict[Text, List[Features]]], states2: List[Dict[Text, List[Features]]]
) -> bool:
"""Compares two lists of featurized states and returns True if they
are identical and False otherwise.
"""
if len(states1) != len(states2):
return False
for state1, state2 in zip(states1, states2):
if state1.keys() != state2.keys():
return False
for key in state1.keys():
for feature1, feature2 in zip(state1[key], state2[key]):
if np.any((feature1.features != feature2.features).toarray()):
return False
if feature1.origin != feature2.origin:
return False
if feature1.attribute != feature2.attribute:
return False
if feature1.type != feature2.type:
return False
return True
def test_featurize_trackers_with_full_dialogue_tracker_featurizer(
moodbot_tracker: DialogueStateTracker,
moodbot_domain: Domain,
moodbot_features: Dict[Text, Dict[Text, Features]],
):
state_featurizer = SingleStateFeaturizer()
tracker_featurizer = FullDialogueTrackerFeaturizer(state_featurizer)
actual_features, actual_labels, entity_tags = tracker_featurizer.featurize_trackers(
[moodbot_tracker], moodbot_domain, precomputations=None,
)
expected_features = [
[
{},
{
ACTION_NAME: [moodbot_features["actions"][ACTION_LISTEN_NAME]],
INTENT: [moodbot_features["intents"]["greet"]],
},
{ACTION_NAME: [moodbot_features["actions"]["utter_greet"]]},
{
ACTION_NAME: [moodbot_features["actions"][ACTION_LISTEN_NAME]],
INTENT: [moodbot_features["intents"]["mood_unhappy"]],
},
{ACTION_NAME: [moodbot_features["actions"]["utter_cheer_up"]]},
{ACTION_NAME: [moodbot_features["actions"]["utter_did_that_help"]]},
{
ACTION_NAME: [moodbot_features["actions"][ACTION_LISTEN_NAME]],
INTENT: [moodbot_features["intents"]["deny"]],
},
]
]
assert actual_features is not None
assert len(actual_features) == len(expected_features)
for actual, expected in zip(actual_features, expected_features):
assert compare_featurized_states(actual, expected)
expected_labels = np.array([[0, 16, 0, 13, 14, 0, 15]])
assert actual_labels is not None
assert len(actual_labels) == 1
for actual, expected in zip(actual_labels, expected_labels):
assert np.all(actual == expected)
# moodbot doesn't contain e2e entities
assert not any([any(turn_tags) for turn_tags in entity_tags])
def test_trackers_ignore_action_unlikely_intent_with_full_dialogue_tracker_featurizer(
moodbot_domain: Domain, moodbot_features: Dict[Text, Dict[Text, Features]],
):
tracker = DialogueStateTracker.from_events(
"default",
[
ActionExecuted(ACTION_LISTEN_NAME),
user_uttered("greet"),
ActionExecuted(ACTION_UNLIKELY_INTENT_NAME),
ActionExecuted("utter_greet"),
ActionExecuted(ACTION_LISTEN_NAME),
user_uttered("mood_unhappy"),
ActionExecuted(ACTION_UNLIKELY_INTENT_NAME),
ActionExecuted("utter_cheer_up"),
ActionExecuted("utter_did_that_help"),
ActionExecuted(ACTION_LISTEN_NAME),
user_uttered("deny"),
ActionExecuted(ACTION_UNLIKELY_INTENT_NAME),
ActionExecuted("utter_goodbye"),
],
domain=moodbot_domain,
)
state_featurizer = SingleStateFeaturizer()
tracker_featurizer = FullDialogueTrackerFeaturizer(state_featurizer)
actual_features, actual_labels, entity_tags = tracker_featurizer.featurize_trackers(
[tracker],
moodbot_domain,
precomputations=None,
ignore_action_unlikely_intent=True,
)
expected_features = [
[
{},
{
ACTION_NAME: [moodbot_features["actions"][ACTION_LISTEN_NAME]],
INTENT: [moodbot_features["intents"]["greet"]],
},
{ACTION_NAME: [moodbot_features["actions"]["utter_greet"]]},
{
ACTION_NAME: [moodbot_features["actions"][ACTION_LISTEN_NAME]],
INTENT: [moodbot_features["intents"]["mood_unhappy"]],
},
{ACTION_NAME: [moodbot_features["actions"]["utter_cheer_up"]]},
{ACTION_NAME: [moodbot_features["actions"]["utter_did_that_help"]]},
{
ACTION_NAME: [moodbot_features["actions"][ACTION_LISTEN_NAME]],
INTENT: [moodbot_features["intents"]["deny"]],
},
]
]
assert actual_features is not None
assert len(actual_features) == len(expected_features)
for actual, expected in zip(actual_features, expected_features):
assert compare_featurized_states(actual, expected)
expected_labels = np.array([[0, 16, 0, 13, 14, 0, 15]])
assert actual_labels is not None
assert len(actual_labels) == 1
for actual, expected in zip(actual_labels, expected_labels):
assert np.all(actual == expected)
# moodbot doesn't contain e2e entities
assert not any([any(turn_tags) for turn_tags in entity_tags])
def test_trackers_keep_action_unlikely_intent_with_full_dialogue_tracker_featurizer(
moodbot_domain: Domain, moodbot_features: Dict[Text, Dict[Text, Features]],
):
tracker = DialogueStateTracker.from_events(
"default",
[
ActionExecuted(ACTION_LISTEN_NAME),
user_uttered("greet"),
ActionExecuted(ACTION_UNLIKELY_INTENT_NAME),
ActionExecuted("utter_greet"),
ActionExecuted(ACTION_LISTEN_NAME),
user_uttered("mood_unhappy"),
ActionExecuted(ACTION_UNLIKELY_INTENT_NAME),
ActionExecuted("utter_cheer_up"),
ActionExecuted("utter_did_that_help"),
ActionExecuted(ACTION_LISTEN_NAME),
user_uttered("deny"),
ActionExecuted(ACTION_UNLIKELY_INTENT_NAME),
ActionExecuted("utter_goodbye"),
],
domain=moodbot_domain,
)
state_featurizer = SingleStateFeaturizer()
tracker_featurizer = FullDialogueTrackerFeaturizer(state_featurizer)
actual_features, actual_labels, entity_tags = tracker_featurizer.featurize_trackers(
[tracker], moodbot_domain, precomputations=None,
)
expected_features = [
[
{},
{
ACTION_NAME: [moodbot_features["actions"][ACTION_LISTEN_NAME]],
INTENT: [moodbot_features["intents"]["greet"]],
},
{ACTION_NAME: [moodbot_features["actions"][ACTION_UNLIKELY_INTENT_NAME]]},
{ACTION_NAME: [moodbot_features["actions"]["utter_greet"]]},
{
ACTION_NAME: [moodbot_features["actions"][ACTION_LISTEN_NAME]],
INTENT: [moodbot_features["intents"]["mood_unhappy"]],
},
{ACTION_NAME: [moodbot_features["actions"][ACTION_UNLIKELY_INTENT_NAME]]},
{ACTION_NAME: [moodbot_features["actions"]["utter_cheer_up"]]},
{ACTION_NAME: [moodbot_features["actions"]["utter_did_that_help"]]},
{
ACTION_NAME: [moodbot_features["actions"][ACTION_LISTEN_NAME]],
INTENT: [moodbot_features["intents"]["deny"]],
},
{ACTION_NAME: [moodbot_features["actions"][ACTION_UNLIKELY_INTENT_NAME]]},
]
]
assert actual_features is not None
assert len(actual_features) == len(expected_features)
for actual, expected in zip(actual_features, expected_features):
assert compare_featurized_states(actual, expected)
expected_labels = np.array([[0, 9, 16, 0, 9, 13, 14, 0, 9, 15]])
assert actual_labels is not None
assert len(actual_labels) == 1
for actual, expected in zip(actual_labels, expected_labels):
assert np.all(actual == expected)
# moodbot doesn't contain e2e entities
assert not any([any(turn_tags) for turn_tags in entity_tags])
def test_create_state_features_full_dialogue_tracker_featurizer(
moodbot_tracker: DialogueStateTracker,
moodbot_domain: Domain,
moodbot_features: Dict[Text, Dict[Text, Features]],
):
state_featurizer = SingleStateFeaturizer()
tracker_featurizer = FullDialogueTrackerFeaturizer(state_featurizer)
state_featurizer.prepare_for_training(moodbot_domain)
actual_features = tracker_featurizer.create_state_features(
[moodbot_tracker], moodbot_domain, precomputations=None,
)
expected_features = [
[
{},
{
ACTION_NAME: [moodbot_features["actions"][ACTION_LISTEN_NAME]],
INTENT: [moodbot_features["intents"]["greet"]],
},
{ACTION_NAME: [moodbot_features["actions"]["utter_greet"]]},
{
ACTION_NAME: [moodbot_features["actions"][ACTION_LISTEN_NAME]],
INTENT: [moodbot_features["intents"]["mood_unhappy"]],
},
{ACTION_NAME: [moodbot_features["actions"]["utter_cheer_up"]]},
{ACTION_NAME: [moodbot_features["actions"]["utter_did_that_help"]]},
{
ACTION_NAME: [moodbot_features["actions"][ACTION_LISTEN_NAME]],
INTENT: [moodbot_features["intents"]["deny"]],
},
{ACTION_NAME: [moodbot_features["actions"]["utter_goodbye"]]},
]
]
assert actual_features is not None
assert len(actual_features) == len(expected_features)
for actual, expected in zip(actual_features, expected_features):
assert compare_featurized_states(actual, expected)
def test_state_features_ignore_action_unlikely_intent_full_dialogue_tracker_featurizer(
moodbot_domain: Domain, moodbot_features: Dict[Text, Dict[Text, Features]],
):
tracker = DialogueStateTracker.from_events(
"default",
[
ActionExecuted(ACTION_LISTEN_NAME),
user_uttered("greet"),
ActionExecuted(ACTION_UNLIKELY_INTENT_NAME),
ActionExecuted("utter_greet"),
ActionExecuted(ACTION_LISTEN_NAME),
user_uttered("mood_great"),
ActionExecuted(ACTION_UNLIKELY_INTENT_NAME),
ActionExecuted("utter_happy"),
ActionExecuted(ACTION_LISTEN_NAME),
user_uttered("goodbye"),
],
domain=moodbot_domain,
)
state_featurizer = SingleStateFeaturizer()
tracker_featurizer = FullDialogueTrackerFeaturizer(state_featurizer)
state_featurizer.prepare_for_training(moodbot_domain)
actual_features = tracker_featurizer.create_state_features(
[tracker],
moodbot_domain,
precomputations=None,
ignore_action_unlikely_intent=True,
)
expected_features = [
[
{},
{
ACTION_NAME: [moodbot_features["actions"][ACTION_LISTEN_NAME]],
INTENT: [moodbot_features["intents"]["greet"]],
},
{ACTION_NAME: [moodbot_features["actions"]["utter_greet"]]},
{
ACTION_NAME: [moodbot_features["actions"][ACTION_LISTEN_NAME]],
INTENT: [moodbot_features["intents"]["mood_great"]],
},
{ACTION_NAME: [moodbot_features["actions"]["utter_happy"]]},
{
ACTION_NAME: [moodbot_features["actions"][ACTION_LISTEN_NAME]],
INTENT: [moodbot_features["intents"]["goodbye"]],
},
]
]
assert actual_features is not None
assert len(actual_features) == len(expected_features)
for actual, expected in zip(actual_features, expected_features):
assert compare_featurized_states(actual, expected)
def test_state_features_keep_action_unlikely_intent_full_dialogue_tracker_featurizer(
moodbot_domain: Domain, moodbot_features: Dict[Text, Dict[Text, Features]],
):
tracker = DialogueStateTracker.from_events(
"default",
[
ActionExecuted(ACTION_LISTEN_NAME),
user_uttered("greet"),
ActionExecuted(ACTION_UNLIKELY_INTENT_NAME),
ActionExecuted("utter_greet"),
ActionExecuted(ACTION_LISTEN_NAME),
user_uttered("mood_great"),
ActionExecuted(ACTION_UNLIKELY_INTENT_NAME),
ActionExecuted("utter_happy"),
ActionExecuted(ACTION_LISTEN_NAME),
user_uttered("goodbye"),
],
domain=moodbot_domain,
)
state_featurizer = SingleStateFeaturizer()
tracker_featurizer = FullDialogueTrackerFeaturizer(state_featurizer)
state_featurizer.prepare_for_training(moodbot_domain)
actual_features = tracker_featurizer.create_state_features(
[tracker], moodbot_domain, precomputations=None,
)
expected_features = [
[
{},
{
ACTION_NAME: [moodbot_features["actions"][ACTION_LISTEN_NAME]],
INTENT: [moodbot_features["intents"]["greet"]],
},
{ACTION_NAME: [moodbot_features["actions"][ACTION_UNLIKELY_INTENT_NAME]]},
{ACTION_NAME: [moodbot_features["actions"]["utter_greet"]]},
{
ACTION_NAME: [moodbot_features["actions"][ACTION_LISTEN_NAME]],
INTENT: [moodbot_features["intents"]["mood_great"]],
},
{ACTION_NAME: [moodbot_features["actions"][ACTION_UNLIKELY_INTENT_NAME]]},
{ACTION_NAME: [moodbot_features["actions"]["utter_happy"]]},
{
ACTION_NAME: [moodbot_features["actions"][ACTION_LISTEN_NAME]],
INTENT: [moodbot_features["intents"]["goodbye"]],
},
]
]
assert actual_features is not None
assert len(actual_features) == len(expected_features)
for actual, expected in zip(actual_features, expected_features):
assert compare_featurized_states(actual, expected)
def test_prediction_states_with_full_dialogue_tracker_featurizer(
moodbot_tracker: DialogueStateTracker, moodbot_domain: Domain
):
state_featurizer = SingleStateFeaturizer()
tracker_featurizer = FullDialogueTrackerFeaturizer(state_featurizer)
actual_states = tracker_featurizer.prediction_states(
[moodbot_tracker], moodbot_domain,
)
expected_states = [
[
{},
{
PREVIOUS_ACTION: {ACTION_NAME: ACTION_LISTEN_NAME},
USER: {INTENT: "greet"},
},
{USER: {INTENT: "greet"}, PREVIOUS_ACTION: {ACTION_NAME: "utter_greet"},},
{
PREVIOUS_ACTION: {ACTION_NAME: ACTION_LISTEN_NAME},
USER: {INTENT: "mood_unhappy"},
},
{
USER: {INTENT: "mood_unhappy"},
PREVIOUS_ACTION: {ACTION_NAME: "utter_cheer_up"},
},
{
USER: {INTENT: "mood_unhappy"},
PREVIOUS_ACTION: {ACTION_NAME: "utter_did_that_help"},
},
{
PREVIOUS_ACTION: {ACTION_NAME: ACTION_LISTEN_NAME},
USER: {INTENT: "deny"},
},
{USER: {INTENT: "deny"}, PREVIOUS_ACTION: {ACTION_NAME: "utter_goodbye"},},
]
]
assert actual_states is not None
assert len(actual_states) == len(expected_states)
for actual, expected in zip(actual_states, expected_states):
assert actual == expected
def test_prediction_states_hide_rule_states_with_full_dialogue_tracker_featurizer(
moodbot_domain: Domain,
):
state_featurizer = SingleStateFeaturizer()
tracker_featurizer = FullDialogueTrackerFeaturizer(state_featurizer)
rule_tracker = DialogueStateTracker.from_events(
"default",
[
ActionExecuted(ACTION_LISTEN_NAME),
user_uttered("greet"),
ActionExecuted("utter_greet", hide_rule_turn=True),
ActionExecuted(ACTION_LISTEN_NAME, hide_rule_turn=True),
],
domain=moodbot_domain,
)
actual_states = tracker_featurizer.prediction_states(
[rule_tracker], moodbot_domain, ignore_rule_only_turns=True,
)
expected_states = [
[
{},
{
PREVIOUS_ACTION: {ACTION_NAME: ACTION_LISTEN_NAME},
USER: {INTENT: "greet"},
},
],
]
assert actual_states is not None
assert len(actual_states) == len(expected_states)
for actual, expected in zip(actual_states, expected_states):
assert actual == expected
embedded_rule_tracker = DialogueStateTracker.from_events(
"default",
[
ActionExecuted(ACTION_LISTEN_NAME),
user_uttered("greet"),
ActionExecuted("utter_greet", hide_rule_turn=True),
ActionExecuted(ACTION_LISTEN_NAME, hide_rule_turn=True),
user_uttered("mood_great"),
ActionExecuted("utter_happy"),
ActionExecuted(ACTION_LISTEN_NAME),
],
domain=moodbot_domain,
)
actual_states = tracker_featurizer.prediction_states(
[embedded_rule_tracker], moodbot_domain, ignore_rule_only_turns=True,
)
expected_states = [
[
{},
{
PREVIOUS_ACTION: {ACTION_NAME: ACTION_LISTEN_NAME},
USER: {INTENT: "mood_great"},
},
{
USER: {INTENT: "mood_great"},
PREVIOUS_ACTION: {ACTION_NAME: "utter_happy"},
},
{
USER: {INTENT: "mood_great"},
PREVIOUS_ACTION: {ACTION_NAME: ACTION_LISTEN_NAME},
},
]
]
assert actual_states is not None
assert len(actual_states) == len(expected_states)
for actual, expected in zip(actual_states, expected_states):
assert actual == expected
def test_prediction_states_ignore_action_intent_unlikely_full_dialogue_featurizer(
moodbot_domain: Domain,
):
state_featurizer = SingleStateFeaturizer()
tracker_featurizer = FullDialogueTrackerFeaturizer(state_featurizer)
tracker = DialogueStateTracker.from_events(
"default",
[
ActionExecuted(ACTION_LISTEN_NAME),
user_uttered("greet"),
ActionExecuted(ACTION_UNLIKELY_INTENT_NAME),
ActionExecuted("utter_greet"),
ActionExecuted(ACTION_LISTEN_NAME),
user_uttered("mood_great"),
ActionExecuted(ACTION_UNLIKELY_INTENT_NAME),
ActionExecuted("utter_happy"),
ActionExecuted(ACTION_LISTEN_NAME),
user_uttered("goodbye"),
],
domain=moodbot_domain,
)
actual_states = tracker_featurizer.prediction_states(
[tracker], moodbot_domain, ignore_action_unlikely_intent=True
)
expected_states = [
[
{},
{
PREVIOUS_ACTION: {ACTION_NAME: ACTION_LISTEN_NAME},
USER: {INTENT: "greet"},
},
{USER: {INTENT: "greet"}, PREVIOUS_ACTION: {ACTION_NAME: "utter_greet"},},
{
PREVIOUS_ACTION: {ACTION_NAME: ACTION_LISTEN_NAME},
USER: {INTENT: "mood_great"},
},
{
USER: {INTENT: "mood_great"},
PREVIOUS_ACTION: {ACTION_NAME: "utter_happy"},
},
{
PREVIOUS_ACTION: {ACTION_NAME: ACTION_LISTEN_NAME},
USER: {INTENT: "goodbye"},
},
]
]
assert actual_states is not None
assert len(actual_states) == len(expected_states)
for actual, expected in zip(actual_states, expected_states):
assert actual == expected
def test_prediction_states_keeps_action_intent_unlikely_full_dialogue_featurizer(
moodbot_domain: Domain,
):
state_featurizer = SingleStateFeaturizer()
tracker_featurizer = FullDialogueTrackerFeaturizer(state_featurizer)
tracker = DialogueStateTracker.from_events(
"default",
[
ActionExecuted(ACTION_LISTEN_NAME),
user_uttered("greet"),
ActionExecuted(ACTION_UNLIKELY_INTENT_NAME),
ActionExecuted("utter_greet"),
ActionExecuted(ACTION_LISTEN_NAME),
user_uttered("mood_great"),
ActionExecuted(ACTION_UNLIKELY_INTENT_NAME),
ActionExecuted("utter_happy"),
ActionExecuted(ACTION_LISTEN_NAME),
user_uttered("goodbye"),
],
domain=moodbot_domain,
)
actual_states = tracker_featurizer.prediction_states([tracker], moodbot_domain,)
expected_states = [
[
{},
{
PREVIOUS_ACTION: {ACTION_NAME: ACTION_LISTEN_NAME},
USER: {INTENT: "greet"},
},
{
USER: {INTENT: "greet"},
PREVIOUS_ACTION: {ACTION_NAME: ACTION_UNLIKELY_INTENT_NAME},
},
{USER: {INTENT: "greet"}, PREVIOUS_ACTION: {ACTION_NAME: "utter_greet"},},
{
PREVIOUS_ACTION: {ACTION_NAME: ACTION_LISTEN_NAME},
USER: {INTENT: "mood_great"},
},
{
USER: {INTENT: "mood_great"},
PREVIOUS_ACTION: {ACTION_NAME: ACTION_UNLIKELY_INTENT_NAME},
},
{
USER: {INTENT: "mood_great"},
PREVIOUS_ACTION: {ACTION_NAME: "utter_happy"},
},
{
PREVIOUS_ACTION: {ACTION_NAME: ACTION_LISTEN_NAME},
USER: {INTENT: "goodbye"},
},
]
]
assert actual_states is not None
assert len(actual_states) == len(expected_states)
for actual, expected in zip(actual_states, expected_states):
assert actual == expected
@pytest.mark.parametrize("max_history", [None, 2])
def test_featurize_trackers_with_max_history_tracker_featurizer(
moodbot_tracker: DialogueStateTracker,
moodbot_domain: Domain,
moodbot_features: Dict[Text, Dict[Text, Features]],
max_history: Optional[int],
):
state_featurizer = SingleStateFeaturizer()
tracker_featurizer = MaxHistoryTrackerFeaturizer(
state_featurizer, max_history=max_history
)
actual_features, actual_labels, entity_tags = tracker_featurizer.featurize_trackers(
[moodbot_tracker], moodbot_domain, precomputations=None,
)
expected_features = [
[{},],
[
{},
{
ACTION_NAME: [moodbot_features["actions"][ACTION_LISTEN_NAME]],
INTENT: [moodbot_features["intents"]["greet"]],
},
],
[
{},
{
ACTION_NAME: [moodbot_features["actions"][ACTION_LISTEN_NAME]],
INTENT: [moodbot_features["intents"]["greet"]],
},
{ACTION_NAME: [moodbot_features["actions"]["utter_greet"]]},
],
[
{},
{
ACTION_NAME: [moodbot_features["actions"][ACTION_LISTEN_NAME]],
INTENT: [moodbot_features["intents"]["greet"]],
},
{ACTION_NAME: [moodbot_features["actions"]["utter_greet"]]},
{
ACTION_NAME: [moodbot_features["actions"][ACTION_LISTEN_NAME]],
INTENT: [moodbot_features["intents"]["mood_unhappy"]],
},
],
[
{},
{
ACTION_NAME: [moodbot_features["actions"][ACTION_LISTEN_NAME]],
INTENT: [moodbot_features["intents"]["greet"]],
},
{ACTION_NAME: [moodbot_features["actions"]["utter_greet"]]},
{
ACTION_NAME: [moodbot_features["actions"][ACTION_LISTEN_NAME]],
INTENT: [moodbot_features["intents"]["mood_unhappy"]],
},
{ACTION_NAME: [moodbot_features["actions"]["utter_cheer_up"]]},
],
[
{},
{
ACTION_NAME: [moodbot_features["actions"][ACTION_LISTEN_NAME]],
INTENT: [moodbot_features["intents"]["greet"]],
},
{ACTION_NAME: [moodbot_features["actions"]["utter_greet"]]},
{
ACTION_NAME: [moodbot_features["actions"][ACTION_LISTEN_NAME]],
INTENT: [moodbot_features["intents"]["mood_unhappy"]],
},
{ACTION_NAME: [moodbot_features["actions"]["utter_cheer_up"]]},
{ACTION_NAME: [moodbot_features["actions"]["utter_did_that_help"]]},
],
[
{},
{
ACTION_NAME: [moodbot_features["actions"][ACTION_LISTEN_NAME]],
INTENT: [moodbot_features["intents"]["greet"]],
},
{ACTION_NAME: [moodbot_features["actions"]["utter_greet"]]},
{
ACTION_NAME: [moodbot_features["actions"][ACTION_LISTEN_NAME]],
INTENT: [moodbot_features["intents"]["mood_unhappy"]],
},
{ACTION_NAME: [moodbot_features["actions"]["utter_cheer_up"]]},
{ACTION_NAME: [moodbot_features["actions"]["utter_did_that_help"]]},
{
ACTION_NAME: [moodbot_features["actions"][ACTION_LISTEN_NAME]],
INTENT: [moodbot_features["intents"]["deny"]],
},
],
]
if max_history is not None:
expected_features = [x[-max_history:] for x in expected_features]
assert actual_features is not None
assert len(actual_features) == len(expected_features)
for actual, expected in zip(actual_features, expected_features):
assert compare_featurized_states(actual, expected)
expected_labels = np.array([[0, 16, 0, 13, 14, 0, 15]]).T
assert actual_labels is not None
assert actual_labels.shape == expected_labels.shape
assert np.all(actual_labels == expected_labels)
# moodbot doesn't contain e2e entities
assert not any([any(turn_tags) for turn_tags in entity_tags])
@pytest.mark.parametrize("max_history", [None, 2])
def test_featurize_trackers_ignore_action_unlikely_intent_max_history_featurizer(
moodbot_domain: Domain,
moodbot_features: Dict[Text, Dict[Text, Features]],
max_history: Optional[int],
):
tracker = DialogueStateTracker.from_events(
"default",
[
ActionExecuted(ACTION_LISTEN_NAME),
user_uttered("greet"),
ActionExecuted(ACTION_UNLIKELY_INTENT_NAME),
ActionExecuted("utter_greet"),
ActionExecuted(ACTION_LISTEN_NAME),
user_uttered("mood_unhappy"),
],
domain=moodbot_domain,
)
state_featurizer = SingleStateFeaturizer()
tracker_featurizer = MaxHistoryTrackerFeaturizer(
state_featurizer, max_history=max_history,
)
actual_features, actual_labels, entity_tags = tracker_featurizer.featurize_trackers(
[tracker],
moodbot_domain,
precomputations=None,
ignore_action_unlikely_intent=True,
)
expected_features = [
[{},],
[
{},
{
ACTION_NAME: [moodbot_features["actions"][ACTION_LISTEN_NAME]],
INTENT: [moodbot_features["intents"]["greet"]],
},
],
[
{},
{
ACTION_NAME: [moodbot_features["actions"][ACTION_LISTEN_NAME]],
INTENT: [moodbot_features["intents"]["greet"]],
},
{ACTION_NAME: [moodbot_features["actions"]["utter_greet"]]},
],
]
if max_history is not None:
expected_features = [x[-max_history:] for x in expected_features]
assert actual_features is not None
assert len(actual_features) == len(expected_features)
for actual, expected in zip(actual_features, expected_features):
assert compare_featurized_states(actual, expected)
expected_labels = np.array([[0, 16, 0]]).T
assert actual_labels.shape == expected_labels.shape
for actual, expected in zip(actual_labels, expected_labels):
assert np.all(actual == expected)
# moodbot doesn't contain e2e entities
assert not any([any(turn_tags) for turn_tags in entity_tags])
@pytest.mark.parametrize("max_history", [None, 2])
def test_featurize_trackers_keep_action_unlikely_intent_max_history_featurizer(
moodbot_domain: Domain,
moodbot_features: Dict[Text, Dict[Text, Features]],
max_history: Optional[int],
):
tracker = DialogueStateTracker.from_events(
"default",
[
ActionExecuted(ACTION_LISTEN_NAME),
user_uttered("greet"),
ActionExecuted(ACTION_UNLIKELY_INTENT_NAME),
ActionExecuted("utter_greet"),
ActionExecuted(ACTION_LISTEN_NAME),
user_uttered("mood_unhappy"),
],
domain=moodbot_domain,
)
state_featurizer = SingleStateFeaturizer()
tracker_featurizer = MaxHistoryTrackerFeaturizer(
state_featurizer, max_history=max_history,
)
actual_features, actual_labels, entity_tags = tracker_featurizer.featurize_trackers(
[tracker], moodbot_domain, precomputations=None,
)
expected_features = [
[{},],
[
{},
{
ACTION_NAME: [moodbot_features["actions"][ACTION_LISTEN_NAME]],
INTENT: [moodbot_features["intents"]["greet"]],
},
],
[
{},
{
ACTION_NAME: [moodbot_features["actions"][ACTION_LISTEN_NAME]],
INTENT: [moodbot_features["intents"]["greet"]],
},
{ACTION_NAME: [moodbot_features["actions"][ACTION_UNLIKELY_INTENT_NAME]]},
],
[
{},
{
ACTION_NAME: [moodbot_features["actions"][ACTION_LISTEN_NAME]],
INTENT: [moodbot_features["intents"]["greet"]],
},
{ACTION_NAME: [moodbot_features["actions"][ACTION_UNLIKELY_INTENT_NAME]]},
{ACTION_NAME: [moodbot_features["actions"]["utter_greet"]]},
],
]
if max_history is not None:
expected_features = [x[-max_history:] for x in expected_features]
assert actual_features is not None
assert len(actual_features) == len(expected_features)
for actual, expected in zip(actual_features, expected_features):
assert compare_featurized_states(actual, expected)
expected_labels = np.array([[0, 9, 16, 0]]).T
assert actual_labels is not None
assert actual_labels.shape == expected_labels.shape
for actual, expected in zip(actual_labels, expected_labels):
assert np.all(actual == expected)
# moodbot doesn't contain e2e entities
assert not any([any(turn_tags) for turn_tags in entity_tags])
@pytest.mark.parametrize(
"remove_duplicates,max_history",
[[True, None], [True, 2], [False, None], [False, 2],],
)
def test_deduplicate_featurize_trackers_with_max_history_tracker_featurizer(
moodbot_tracker: DialogueStateTracker,
moodbot_domain: Domain,
moodbot_features: Dict[Text, Dict[Text, Features]],
remove_duplicates: bool,
max_history: Optional[int],
):
state_featurizer = SingleStateFeaturizer()
tracker_featurizer = MaxHistoryTrackerFeaturizer(
state_featurizer, max_history=max_history, remove_duplicates=remove_duplicates
)
# Add Duplicate moodbot_tracker states should get removed.
actual_features, actual_labels, entity_tags = tracker_featurizer.featurize_trackers(
[moodbot_tracker, moodbot_tracker], moodbot_domain, precomputations=None,
)
expected_features = [
[{},],
[
{},
{
ACTION_NAME: [moodbot_features["actions"][ACTION_LISTEN_NAME]],
INTENT: [moodbot_features["intents"]["greet"]],
},
],
[
{},
{
ACTION_NAME: [moodbot_features["actions"][ACTION_LISTEN_NAME]],
INTENT: [moodbot_features["intents"]["greet"]],
},
{ACTION_NAME: [moodbot_features["actions"]["utter_greet"]]},
],
[
{},
{
ACTION_NAME: [moodbot_features["actions"][ACTION_LISTEN_NAME]],
INTENT: [moodbot_features["intents"]["greet"]],
},
{ACTION_NAME: [moodbot_features["actions"]["utter_greet"]]},
{
ACTION_NAME: [moodbot_features["actions"][ACTION_LISTEN_NAME]],
INTENT: [moodbot_features["intents"]["mood_unhappy"]],
},
],
[
{},
{
ACTION_NAME: [moodbot_features["actions"][ACTION_LISTEN_NAME]],
INTENT: [moodbot_features["intents"]["greet"]],
},
{ACTION_NAME: [moodbot_features["actions"]["utter_greet"]]},
{
ACTION_NAME: [moodbot_features["actions"][ACTION_LISTEN_NAME]],
INTENT: [moodbot_features["intents"]["mood_unhappy"]],
},
{ACTION_NAME: [moodbot_features["actions"]["utter_cheer_up"]]},
],
[
{},
{
ACTION_NAME: [moodbot_features["actions"][ACTION_LISTEN_NAME]],
INTENT: [moodbot_features["intents"]["greet"]],
},
{ACTION_NAME: [moodbot_features["actions"]["utter_greet"]]},
{
ACTION_NAME: [moodbot_features["actions"][ACTION_LISTEN_NAME]],
INTENT: [moodbot_features["intents"]["mood_unhappy"]],
},
{ACTION_NAME: [moodbot_features["actions"]["utter_cheer_up"]]},
{ACTION_NAME: [moodbot_features["actions"]["utter_did_that_help"]]},
],
[
{},
{
ACTION_NAME: [moodbot_features["actions"][ACTION_LISTEN_NAME]],
INTENT: [moodbot_features["intents"]["greet"]],
},
{ACTION_NAME: [moodbot_features["actions"]["utter_greet"]]},
{
ACTION_NAME: [moodbot_features["actions"][ACTION_LISTEN_NAME]],
INTENT: [moodbot_features["intents"]["mood_unhappy"]],
},
{ACTION_NAME: [moodbot_features["actions"]["utter_cheer_up"]]},
{ACTION_NAME: [moodbot_features["actions"]["utter_did_that_help"]]},
{
ACTION_NAME: [moodbot_features["actions"][ACTION_LISTEN_NAME]],
INTENT: [moodbot_features["intents"]["deny"]],
},
],
]
if max_history is not None:
expected_features = [x[-max_history:] for x in expected_features]
if not remove_duplicates:
expected_features = expected_features * 2
assert actual_features is not None
assert len(actual_features) == len(expected_features)
for actual, expected in zip(actual_features, expected_features):
assert compare_featurized_states(actual, expected)
expected_labels = np.array([[0, 16, 0, 13, 14, 0, 15]]).T
if not remove_duplicates:
expected_labels = np.vstack([expected_labels] * 2)
assert actual_labels is not None
assert actual_labels.shape == expected_labels.shape
assert
|
np.all(actual_labels == expected_labels)
|
numpy.all
|
# Licensed to the Apache Software Foundation (ASF) under one
# or more contributor license agreements. See the NOTICE file
# distributed with this work for additional information
# regarding copyright ownership. The ASF licenses this file
# to you under the Apache License, Version 2.0 (the
# "License"); you may not use this file except in compliance
# with the License. You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing,
# software distributed under the License is distributed on an
# "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY
# KIND, either express or implied. See the License for the
# specific language governing permissions and limitations
# under the License.
import tvm
from tvm import te
import numpy as np
from tvm import relay
from tvm.contrib import graph_executor
import tvm.topi.testing
# "unquantize" a quantized tensor
def recover(data, scale, zp):
return scale * (np.asarray(data) - zp)
def generate_golden_output(x_recovered, y_recovered, scale, zp):
mul = x_recovered * y_recovered
output = np.around(mul / scale + zp)
q_min = np.iinfo(np.uint8).min
q_max = np.iinfo(np.uint8).max
return np.clip(output, q_min, q_max)
def test_tflite_same_io_qnn_params():
data_dtype = "uint8"
lhs_scale = rhs_scale = output_scale = 0.00784314
lhs_zero_point = rhs_zero_point = output_zero_point = 127
x = relay.var("x", shape=(1, 4), dtype=data_dtype)
y = relay.var("y", shape=(1, 4), dtype=data_dtype)
z = relay.qnn.op.mul(
lhs=x,
rhs=y,
lhs_scale=relay.const(lhs_scale, "float32"),
lhs_zero_point=relay.const(lhs_zero_point, "int32"),
rhs_scale=relay.const(rhs_scale, "float32"),
rhs_zero_point=relay.const(rhs_zero_point, "int32"),
output_scale=relay.const(output_scale, "float32"),
output_zero_point=relay.const(output_zero_point, "int32"),
)
func = relay.Function([x, y], z)
mod = tvm.IRModule.from_expr(func)
mod = relay.transform.InferType()(mod)
mod = relay.qnn.transform.CanonicalizeOps()(mod)
func = mod["main"]
x_datas = [
np.array((1, 153, 2, 178)).reshape((1, 4)),
np.array((25, 1, 178, 216)).reshape((1, 4)),
np.array((25, 153, 1, 165)).reshape((1, 4)),
]
y_datas = [
np.array((204, 178, 1, 8)).reshape((1, 4)),
np.array((204, 178, 191, 1)).reshape((1, 4)),
np.array((204, 178, 1, 191)).reshape((1, 4)),
]
for i in range(0, 3):
x_data = x_datas[i]
y_data = y_datas[i]
x_rec = recover(x_data, lhs_scale, lhs_zero_point)
y_rec = recover(y_data, rhs_scale, rhs_zero_point)
golden = generate_golden_output(x_rec, y_rec, output_scale, output_zero_point)
intrp = relay.create_executor("graph", device=tvm.cpu(0), target="llvm")
op_res = intrp.evaluate(func)(x_data, y_data)
np.testing.assert_equal(op_res.numpy(), np.uint8(golden))
def test_tflite_different_io_qnn_params():
data_dtype = "uint8"
lhs_scale = 0.0156863
lhs_zero_point = 127
rhs_scale = 0.0117647
rhs_zero_point = 85
output_scale = 0.0235294
output_zero_point = 128
x = relay.var("x", shape=(1, 4), dtype=data_dtype)
y = relay.var("y", shape=(1, 4), dtype=data_dtype)
z = relay.qnn.op.mul(
lhs=x,
rhs=y,
lhs_scale=relay.const(lhs_scale, "float32"),
lhs_zero_point=relay.const(lhs_zero_point, "int32"),
rhs_scale=relay.const(rhs_scale, "float32"),
rhs_zero_point=relay.const(rhs_zero_point, "int32"),
output_scale=relay.const(output_scale, "float32"),
output_zero_point=relay.const(output_zero_point, "int32"),
)
func = relay.Function([x, y], z)
mod = tvm.IRModule.from_expr(func)
mod = relay.transform.InferType()(mod)
mod = relay.qnn.transform.CanonicalizeOps()(mod)
func = mod["main"]
x_datas = [
np.array((76, 140, 153, 172)).reshape((1, 4)),
|
np.array((133, 140, 146, 153))
|
numpy.array
|
import pickle
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.patches as mpatches
CB_color_cycle = ['#377eb8', '#ff7f00', '#4daf4a',
'#f781bf', '#a65628', '#984ea3',
'#999999', '#e41a1c', '#dede00']
def __min_birth_max_death(persistence, band=0.0):
# Look for minimum birth date and maximum death date for plot optimisation
max_death = 0
min_birth = persistence[0][1][0]
for interval in reversed(persistence):
if float(interval[1][1]) != float("inf"):
if float(interval[1][1]) > max_death:
max_death = float(interval[1][1])
if float(interval[1][0]) > max_death:
max_death = float(interval[1][0])
if float(interval[1][0]) < min_birth:
min_birth = float(interval[1][0])
if band > 0.0:
max_death += band
return (min_birth, max_death)
def _array_handler(a):
if isinstance(a[0][1], np.float64) or isinstance(a[0][1], float):
return [[0, x] for x in a]
else:
return a
def plot_persistence_barcode(
persistence=[],
alpha=0.85,
max_intervals=1024,
max_barcodes=1024,
inf_delta=0.1,
legend=True,
colormap=None,
axes=None,
fontsize=14,
):
persistence = _array_handler(persistence)
if max_intervals > 0 and max_intervals < len(persistence):
# Sort by life time, then takes only the max_intervals elements
persistence = sorted(
persistence,
key=lambda life_time: life_time[1][1] - life_time[1][0],
reverse=True,
)[:max_intervals]
if colormap is None:
# colormap = plt.cm.Set1.colors
colormap = CB_color_cycle
if axes is None:
fig, axes = plt.subplots(1, 1)
persistence = sorted(persistence, key=lambda birth: birth[1][0])
(min_birth, max_death) = __min_birth_max_death(persistence)
ind = 0
delta = (max_death - min_birth) * inf_delta
# Replace infinity values with max_death + delta for bar code to be more
# readable
infinity = max_death + delta
axis_start = min_birth - delta
# Draw horizontal bars in loop
for interval in reversed(persistence):
if float(interval[1][1]) != float("inf"):
# Finite death case
axes.barh(
ind,
(interval[1][1] - interval[1][0]),
height=0.8,
left=interval[1][0],
alpha=alpha,
color=colormap[interval[0]],
linewidth=0.5,
)
else:
# Infinite death case for diagram to be nicer
axes.barh(
ind,
(infinity - interval[1][0]),
height=0.8,
left=interval[1][0],
alpha=alpha,
color=colormap[interval[0]],
linewidth=0.5,
)
ind = ind + 1
if legend:
dimensions = list(set(item[0] for item in persistence))
axes.legend(
handles=[
mpatches.Patch(color=colormap[dim], label="H"+str(dim))
for dim in dimensions
],
loc="upper right",
)
axes.set_title("Persistence barcode", fontsize=fontsize)
# Ends plot on infinity value and starts a little bit before min_birth
axes.axis([axis_start, infinity, 0, ind])
return axes
def plot_persistence_diagram(
persistence=[],
alpha=0.6,
band=0.0,
max_intervals=1024,
max_plots=1024,
inf_delta=0.1,
legend=True,
colormap=None,
axes=None,
fontsize=14,
greyblock=False
):
persistence = _array_handler(persistence)
if max_plots != 1000:
print("Deprecated parameter. It has been replaced by max_intervals")
max_intervals = max_plots
if max_intervals > 0 and max_intervals < len(persistence):
# Sort by life time, then takes only the max_intervals elements
persistence = sorted(
persistence,
key=lambda life_time: life_time[1][1] - life_time[1][0],
reverse=True,
)[:max_intervals]
if colormap is None:
# colormap = plt.cm.Set1.colors
colormap = CB_color_cycle
if axes is None:
fig, axes = plt.subplots(1, 1)
(min_birth, max_death) = __min_birth_max_death(persistence, band)
delta = (max_death - min_birth) * inf_delta
# Replace infinity values with max_death + delta for diagram to be more
# readable
infinity = max_death + delta
axis_end = max_death + delta / 2
axis_start = min_birth - delta
# bootstrap band
if band > 0.0:
x = np.linspace(axis_start, infinity, 1000)
axes.fill_between(x, x, x + band, alpha=alpha, facecolor="red")
# lower diag patch
if greyblock:
axes.add_patch(mpatches.Polygon([[axis_start, axis_start],
[axis_end, axis_start],
[axis_end, axis_end]],
fill=True,
color='lightgrey'))
# Draw points in loop
pts_at_infty = False # Records presence of pts at infty
for interval in reversed(persistence):
if float(interval[1][1]) != float("inf"):
# Finite death case
axes.scatter(
interval[1][0],
interval[1][1],
alpha=alpha,
color=colormap[interval[0]],
)
else:
pts_at_infty = True
# Infinite death case for diagram to be nicer
axes.scatter(interval[1][0],
infinity,
alpha=alpha,
color=colormap[interval[0]])
if pts_at_infty:
# infinity line and text
axes.plot([axis_start, axis_end],
[axis_start, axis_end],
linewidth=1.0,
color="k")
axes.plot([axis_start, axis_end],
[infinity, infinity],
linewidth=1.0,
color="k",
alpha=alpha)
# Infinity label
yt = axes.get_yticks()
yt = yt[
|
np.where(yt < axis_end)
|
numpy.where
|
# coding: utf-8
# Copyright (c) Pymatgen Development Team.
# Distributed under the terms of the MIT License.
"""
This module contains the main object used to identify the coordination environments in a given structure.
If you use this module, please cite the following:
<NAME>, <NAME>, <NAME>, <NAME>, <NAME>,
<NAME>, <NAME>, <NAME>, <NAME>, <NAME>, and <NAME>,
"Statistical analysis of coordination environments in oxides",
Chem. Mater., 2017, 29 (19), pp 8346–8360,
DOI: 10.1021/acs.chemmater.7b02766
"""
__author__ = "<NAME>"
__copyright__ = "Copyright 2012, The Materials Project"
__credits__ = "<NAME>"
__version__ = "2.0"
__maintainer__ = "<NAME>"
__email__ = "<EMAIL>"
__date__ = "Feb 20, 2016"
import itertools
import logging
import time
from collections import OrderedDict
from random import shuffle
import numpy as np
from numpy.linalg import norm, svd
from pymatgen.analysis.bond_valence import BVAnalyzer
from pymatgen.analysis.chemenv.coordination_environments.chemenv_strategies import (
MultiWeightsChemenvStrategy,
)
from pymatgen.analysis.chemenv.coordination_environments.coordination_geometries import (
EXPLICIT_PERMUTATIONS,
SEPARATION_PLANE,
AllCoordinationGeometries,
)
from pymatgen.analysis.chemenv.coordination_environments.structure_environments import (
ChemicalEnvironments,
LightStructureEnvironments,
StructureEnvironments,
)
from pymatgen.analysis.chemenv.coordination_environments.voronoi import (
DetailedVoronoiContainer,
)
from pymatgen.analysis.chemenv.utils.coordination_geometry_utils import (
Plane,
collinear,
separation_in_list,
sort_separation,
sort_separation_tuple,
)
from pymatgen.analysis.chemenv.utils.defs_utils import chemenv_citations
from pymatgen.core.lattice import Lattice
from pymatgen.core.periodic_table import Species
from pymatgen.core.structure import Structure
from pymatgen.symmetry.analyzer import SpacegroupAnalyzer
debug = False
DIST_TOLERANCES = [0.02, 0.05, 0.1, 0.2, 0.3]
class AbstractGeometry:
"""
Class used to describe a geometry (perfect or distorted)
"""
def __init__(
self,
central_site=None,
bare_coords=None,
centering_type="standard",
include_central_site_in_centroid=False,
optimization=None,
):
"""
Constructor for the abstract geometry
:param central_site: Coordinates of the central site
:param bare_coords: Coordinates of the neighbors of the central site
:param centering_type: How to center the abstract geometry
:param include_central_site_in_centroid: When the centering is on the centroid, the central site is included
if this parameter is set to True.
:raise: ValueError if the parameters are not consistent
"""
bcoords = np.array(bare_coords)
self.bare_centre = np.array(central_site)
self.bare_points_without_centre = bcoords
self.bare_points_with_centre = np.array(central_site)
self.bare_points_with_centre = np.concatenate(([self.bare_points_with_centre], bcoords))
self.centroid_without_centre = np.mean(self.bare_points_without_centre, axis=0)
self.centroid_with_centre = np.mean(self.bare_points_with_centre, axis=0)
self._points_wcs_csc = self.bare_points_with_centre - self.bare_centre
self._points_wocs_csc = self.bare_points_without_centre - self.bare_centre
self._points_wcs_ctwcc = self.bare_points_with_centre - self.centroid_with_centre
self._points_wocs_ctwcc = self.bare_points_without_centre - self.centroid_with_centre
self._points_wcs_ctwocc = self.bare_points_with_centre - self.centroid_without_centre
self._points_wocs_ctwocc = self.bare_points_without_centre - self.centroid_without_centre
self.centering_type = centering_type
self.include_central_site_in_centroid = include_central_site_in_centroid
self.bare_central_site = np.array(central_site)
if centering_type == "standard":
if len(bare_coords) < 5:
if include_central_site_in_centroid:
raise ValueError(
"The center is the central site, no calculation of the centroid, "
"variable include_central_site_in_centroid should be set to False"
)
if central_site is None:
raise ValueError("Centering_type is central_site, the central site should be given")
self.centre = np.array(central_site)
else:
total = np.sum(bcoords, axis=0)
if include_central_site_in_centroid:
if central_site is None:
raise ValueError("The centroid includes the central site but no central site is given")
total += self.bare_centre
self.centre = total / (np.float(len(bare_coords)) + 1.0)
else:
self.centre = total / np.float(len(bare_coords))
elif centering_type == "central_site":
if include_central_site_in_centroid:
raise ValueError(
"The center is the central site, no calculation of the centroid, "
"variable include_central_site_in_centroid should be set to False"
)
if central_site is None:
raise ValueError("Centering_type is central_site, the central site should be given")
self.centre = np.array(central_site)
elif centering_type == "centroid":
total = np.sum(bcoords, axis=0)
if include_central_site_in_centroid:
if central_site is None:
raise ValueError("The centroid includes the central site but no central site is given")
total += self.bare_centre
self.centre = total / (np.float(len(bare_coords)) + 1.0)
else:
self.centre = total / np.float(len(bare_coords))
self._bare_coords = self.bare_points_without_centre
self._coords = self._bare_coords - self.centre
self.central_site = self.bare_central_site - self.centre
self.coords = self._coords
self.bare_coords = self._bare_coords
def __str__(self):
"""
String representation of the AbstractGeometry
:return: String representation of the AbstractGeometry
"""
outs = ["\nAbstract Geometry with {n} points :".format(n=len(self.coords))]
for pp in self.coords:
outs.append(" {pp}".format(pp=pp))
if self.centering_type == "standard":
if self.include_central_site_in_centroid:
outs.append(
"Points are referenced to the central site for coordination numbers < 5"
" and to the centroid (calculated with the central site) for coordination"
" numbers >= 5 : {c}\n".format(c=self.centre)
)
else:
outs.append(
"Points are referenced to the central site for coordination numbers < 5"
" and to the centroid (calculated without the central site) for coordination"
" numbers >= 5 : {c}\n".format(c=self.centre)
)
elif self.centering_type == "central_site":
outs.append("Points are referenced to the central site : {c}\n".format(c=self.centre))
elif self.centering_type == "centroid":
if self.include_central_site_in_centroid:
outs.append(
"Points are referenced to the centroid "
"(calculated with the central site) :\n {c}\n".format(c=self.centre)
)
else:
outs.append(
"Points are referenced to the centroid"
" (calculated without the central site) :\n {c}\n".format(c=self.centre)
)
return "\n".join(outs)
@classmethod
def from_cg(cls, cg, centering_type="standard", include_central_site_in_centroid=False):
"""
:param cg:
:param centering_type:
:param include_central_site_in_centroid:
:return:
"""
central_site = cg.get_central_site()
bare_coords = [np.array(pt, np.float) for pt in cg.points]
return cls(
central_site=central_site,
bare_coords=bare_coords,
centering_type=centering_type,
include_central_site_in_centroid=include_central_site_in_centroid,
)
def points_wcs_csc(self, permutation=None):
"""
:param permutation:
:return:
"""
if permutation is None:
return self._points_wcs_csc
return np.concatenate((self._points_wcs_csc[0:1], self._points_wocs_csc.take(permutation, axis=0)))
def points_wocs_csc(self, permutation=None):
"""
:param permutation:
:return:
"""
if permutation is None:
return self._points_wocs_csc
return self._points_wocs_csc.take(permutation, axis=0)
def points_wcs_ctwcc(self, permutation=None):
"""
:param permutation:
:return:
"""
if permutation is None:
return self._points_wcs_ctwcc
return np.concatenate(
(
self._points_wcs_ctwcc[0:1],
self._points_wocs_ctwcc.take(permutation, axis=0),
)
)
def points_wocs_ctwcc(self, permutation=None):
"""
:param permutation:
:return:
"""
if permutation is None:
return self._points_wocs_ctwcc
return self._points_wocs_ctwcc.take(permutation, axis=0)
def points_wcs_ctwocc(self, permutation=None):
"""
:param permutation:
:return:
"""
if permutation is None:
return self._points_wcs_ctwocc
return np.concatenate(
(
self._points_wcs_ctwocc[0:1],
self._points_wocs_ctwocc.take(permutation, axis=0),
)
)
def points_wocs_ctwocc(self, permutation=None):
"""
:param permutation:
:return:
"""
if permutation is None:
return self._points_wocs_ctwocc
return self._points_wocs_ctwocc.take(permutation, axis=0)
@property
def cn(self):
"""
:return: Coordination number
"""
return len(self.coords)
@property
def coordination_number(self):
"""
:return: Coordination number
"""
return len(self.coords)
def symmetry_measure(points_distorted, points_perfect):
"""
Computes the continuous symmetry measure of the (distorted) set of points "points_distorted" with respect to the
(perfect) set of points "points_perfect".
:param points_distorted: List of points describing a given (distorted) polyhedron for which the symmetry measure
has to be computed with respect to the model polyhedron described by the list of points
"points_perfect".
:param points_perfect: List of "perfect" points describing a given model polyhedron.
:return: The continuous symmetry measure of the distorted polyhedron with respect to the perfect polyhedron
"""
# When there is only one point, the symmetry measure is 0.0 by definition
if len(points_distorted) == 1:
return {
"symmetry_measure": 0.0,
"scaling_factor": None,
"rotation_matrix": None,
}
# Find the rotation matrix that aligns the distorted points to the perfect points in a least-square sense.
rot = find_rotation(points_distorted=points_distorted, points_perfect=points_perfect)
# Find the scaling factor between the distorted points and the perfect points in a least-square sense.
scaling_factor, rotated_coords, points_perfect = find_scaling_factor(
points_distorted=points_distorted, points_perfect=points_perfect, rot=rot
)
# Compute the continuous symmetry measure [see Eq. 1 in Pinsky et al., Inorganic Chemistry 37, 5575 (1998)]
rotated_coords = scaling_factor * rotated_coords
diff = points_perfect - rotated_coords
num = np.tensordot(diff, diff)
denom = np.tensordot(points_perfect, points_perfect)
return {
"symmetry_measure": num / denom * 100.0,
"scaling_factor": scaling_factor,
"rotation_matrix": rot,
}
def find_rotation(points_distorted, points_perfect):
"""
This finds the rotation matrix that aligns the (distorted) set of points "points_distorted" with respect to the
(perfect) set of points "points_perfect" in a least-square sense.
:param points_distorted: List of points describing a given (distorted) polyhedron for which the rotation that
aligns these points in a least-square sense to the set of perfect points "points_perfect"
:param points_perfect: List of "perfect" points describing a given model polyhedron.
:return: The rotation matrix
"""
H = np.matmul(points_distorted.T, points_perfect)
[U, S, Vt] = svd(H)
rot = np.matmul(Vt.T, U.T)
return rot
def find_scaling_factor(points_distorted, points_perfect, rot):
"""
This finds the scaling factor between the (distorted) set of points "points_distorted" and the
(perfect) set of points "points_perfect" in a least-square sense.
:param points_distorted: List of points describing a given (distorted) polyhedron for which the scaling factor has
to be obtained.
:param points_perfect: List of "perfect" points describing a given model polyhedron.
:param rot: The rotation matrix
:return: The scaling factor between the two structures and the rotated set of (distorted) points.
"""
rotated_coords = np.matmul(rot, points_distorted.T).T
num = np.tensordot(rotated_coords, points_perfect)
denom = np.tensordot(rotated_coords, rotated_coords)
return num / denom, rotated_coords, points_perfect
class LocalGeometryFinder:
"""
Main class used to find the local environments in a structure
"""
DEFAULT_BVA_DISTANCE_SCALE_FACTOR = 1.0
BVA_DISTANCE_SCALE_FACTORS = {
"experimental": 1.0,
"GGA_relaxed": 1.015,
"LDA_relaxed": 0.995,
}
DEFAULT_SPG_ANALYZER_OPTIONS = {"symprec": 1e-3, "angle_tolerance": 5}
STRUCTURE_REFINEMENT_NONE = "none"
STRUCTURE_REFINEMENT_REFINED = "refined"
STRUCTURE_REFINEMENT_SYMMETRIZED = "symmetrized"
DEFAULT_STRATEGY = MultiWeightsChemenvStrategy.stats_article_weights_parameters()
PRESETS = {
"DEFAULT": {
"maximum_distance_factor": 2.0,
"minimum_angle_factor": 0.05,
"voronoi_normalized_distance_tolerance": 0.05,
"voronoi_normalized_angle_tolerance": 0.03,
"optimization": 2,
}
}
def __init__(
self,
permutations_safe_override=False,
plane_ordering_override=True,
debug_level=None,
plane_safe_permutations=False,
only_symbols=None,
):
"""
Constructor for the LocalGeometryFinder, initializes the list of coordination geometries
:param permutations_safe_override: If set to True, all permutations are tested (very time-consuming for large
coordination numbers!)
:param plane_ordering_override: If set to False, the ordering of the points in the plane is disabled
"""
self.allcg = AllCoordinationGeometries(
permutations_safe_override=permutations_safe_override,
only_symbols=only_symbols,
)
self.permutations_safe_override = permutations_safe_override
self.plane_ordering_override = plane_ordering_override
self.plane_safe_permutations = plane_safe_permutations
self.setup_parameters(
centering_type="centroid",
include_central_site_in_centroid=True,
bva_distance_scale_factor=None,
structure_refinement=self.STRUCTURE_REFINEMENT_NONE,
)
print(chemenv_citations())
def setup_parameters(
self,
centering_type="standard",
include_central_site_in_centroid=False,
bva_distance_scale_factor=None,
structure_refinement=STRUCTURE_REFINEMENT_REFINED,
spg_analyzer_options=None,
):
"""
Setup of the parameters for the coordination geometry finder. A reference point for the geometries has to be
chosen. This can be the centroid of the structure (including or excluding the atom for which the coordination
geometry is looked for) or the atom itself. In the 'standard' centering_type, the reference point is the central
atom for coordination numbers 1, 2, 3 and 4 and the centroid for coordination numbers > 4.
:param centering_type: Type of the reference point (centering) 'standard', 'centroid' or 'central_site'
:param include_central_site_in_centroid: In case centering_type is 'centroid', the central site is included if
this value is set to True.
:param bva_distance_scale_factor: Scaling factor for the bond valence analyzer (this might be different whether
the structure is an experimental one, an LDA or a GGA relaxed one, or any other relaxation scheme (where
under- or over-estimation of bond lengths is known).
:param structure_refinement: Refinement of the structure. Can be "none", "refined" or "symmetrized".
:param spg_analyzer_options: Options for the SpaceGroupAnalyzer (dictionary specifying "symprec"
and "angle_tolerance". See pymatgen's SpaceGroupAnalyzer for more information.
"""
self.centering_type = centering_type
self.include_central_site_in_centroid = include_central_site_in_centroid
if bva_distance_scale_factor is not None:
self.bva_distance_scale_factor = bva_distance_scale_factor
else:
self.bva_distance_scale_factor = self.DEFAULT_BVA_DISTANCE_SCALE_FACTOR
self.structure_refinement = structure_refinement
if spg_analyzer_options is None:
self.spg_analyzer_options = self.DEFAULT_SPG_ANALYZER_OPTIONS
else:
self.spg_analyzer_options = spg_analyzer_options
def setup_parameter(self, parameter, value):
"""
Setup of one specific parameter to the given value. The other parameters are unchanged. See setup_parameters
method for the list of possible parameters
:param parameter: Parameter to setup/update
:param value: Value of the parameter
"""
self.__dict__[parameter] = value
def setup_structure(self, structure):
"""
Sets up the structure for which the coordination geometries have to be identified. The structure is analyzed
with the space group analyzer and a refined structure is used
:param structure: A pymatgen Structure
"""
self.initial_structure = structure.copy()
if self.structure_refinement == self.STRUCTURE_REFINEMENT_NONE:
self.structure = structure.copy()
self.spg_analyzer = None
self.symmetrized_structure = None
else:
self.spg_analyzer = SpacegroupAnalyzer(
self.initial_structure,
symprec=self.spg_analyzer_options["symprec"],
angle_tolerance=self.spg_analyzer_options["angle_tolerance"],
)
if self.structure_refinement == self.STRUCTURE_REFINEMENT_REFINED:
self.structure = self.spg_analyzer.get_refined_structure()
self.symmetrized_structure = None
elif self.structure_refinement == self.STRUCTURE_REFINEMENT_SYMMETRIZED:
self.structure = self.spg_analyzer.get_refined_structure()
self.spg_analyzer_refined = SpacegroupAnalyzer(
self.structure,
symprec=self.spg_analyzer_options["symprec"],
angle_tolerance=self.spg_analyzer_options["angle_tolerance"],
)
self.symmetrized_structure = self.spg_analyzer_refined.get_symmetrized_structure()
def get_structure(self):
"""
Returns the pymatgen Structure that has been setup for the identification of geometries (the initial one
might have been refined/symmetrized using the SpaceGroupAnalyzer).
:return: The pymatgen Structure that has been setup for the identification of geometries (the initial one
might have been refined/symmetrized using the SpaceGroupAnalyzer).
"""
return self.structure
def set_structure(self, lattice, species, coords, coords_are_cartesian):
"""
Sets up the pymatgen structure for which the coordination geometries have to be identified starting from the
lattice, the species and the coordinates
:param lattice: The lattice of the structure
:param species: The species on the sites
:param coords: The coordinates of the sites
:param coords_are_cartesian: If set to True, the coordinates are given in cartesian coordinates
"""
self.setup_structure(Structure(lattice, species, coords, coords_are_cartesian))
def compute_coordination_environments(
self,
structure,
indices=None,
only_cations=True,
strategy=DEFAULT_STRATEGY,
valences="bond-valence-analysis",
initial_structure_environments=None,
):
"""
:param structure:
:param indices:
:param only_cations:
:param strategy:
:param valences:
:param initial_structure_environments:
:return:
"""
self.setup_structure(structure=structure)
if valences == "bond-valence-analysis":
bva = BVAnalyzer()
try:
vals = bva.get_valences(structure=structure)
except ValueError:
vals = "undefined"
else:
if valences == "undefined":
vals = valences
else:
if len(valences) != len(structure):
raise ValueError("Valences do not match the number of sites in the structure")
vals = valences
# TODO: add something to compute only the neighbors sets needed for the strategy.
se = self.compute_structure_environments(
only_cations=only_cations,
only_indices=indices,
valences=vals,
initial_structure_environments=initial_structure_environments,
)
lse = LightStructureEnvironments.from_structure_environments(strategy=strategy, structure_environments=se)
return lse.coordination_environments
def compute_structure_environments(
self,
excluded_atoms=None,
only_atoms=None,
only_cations=True,
only_indices=None,
maximum_distance_factor=PRESETS["DEFAULT"]["maximum_distance_factor"],
minimum_angle_factor=PRESETS["DEFAULT"]["minimum_angle_factor"],
max_cn=None,
min_cn=None,
only_symbols=None,
valences="undefined",
additional_conditions=None,
info=None,
timelimit=None,
initial_structure_environments=None,
get_from_hints=False,
voronoi_normalized_distance_tolerance=PRESETS["DEFAULT"]["voronoi_normalized_distance_tolerance"],
voronoi_normalized_angle_tolerance=PRESETS["DEFAULT"]["voronoi_normalized_angle_tolerance"],
recompute=None,
optimization=PRESETS["DEFAULT"]["optimization"],
):
"""
Computes and returns the StructureEnvironments object containing all the information about the coordination
environments in the structure
:param excluded_atoms: Atoms for which the coordination geometries does not have to be identified
:param only_atoms: If not set to None, atoms for which the coordination geometries have to be identified
:param only_cations: If set to True, will only compute environments for cations
:param only_indices: If not set to None, will only compute environments the atoms of the given indices
:param maximum_distance_factor: If not set to None, neighbors beyond
maximum_distance_factor*closest_neighbor_distance are not considered
:param minimum_angle_factor: If not set to None, neighbors for which the angle is lower than
minimum_angle_factor*largest_angle_neighbor are not considered
:param max_cn: maximum coordination number to be considered
:param min_cn: minimum coordination number to be considered
:param only_symbols: if not set to None, consider only coordination environments with the given symbols
:param valences: valences of the atoms
:param additional_conditions: additional conditions to be considered in the bonds (example : only bonds
between cation and anion
:param info: additional info about the calculation
:param timelimit: time limit (in secs) after which the calculation of the StructureEnvironments object stops
:param initial_structure_environments: initial StructureEnvironments object (most probably incomplete)
:param get_from_hints: whether to add neighbors sets from "hints" (e.g. capped environment => test the
neighbors without the cap)
:param voronoi_normalized_distance_tolerance: tolerance for the normalized distance used to distinguish
neighbors sets
:param voronoi_normalized_angle_tolerance: tolerance for the normalized angle used to distinguish
neighbors sets
:param recompute: whether to recompute the sites already computed (when initial_structure_environments
is not None)
:param optimization: optimization algorithm
:return: The StructureEnvironments object containing all the information about the coordination
environments in the structure
"""
time_init = time.process_time()
if info is None:
info = {}
info.update(
{
"local_geometry_finder": {
"parameters": {
"centering_type": self.centering_type,
"include_central_site_in_centroid": self.include_central_site_in_centroid,
"structure_refinement": self.structure_refinement,
"spg_analyzer_options": self.spg_analyzer_options,
}
}
}
)
if only_symbols is not None:
self.allcg = AllCoordinationGeometries(
permutations_safe_override=self.permutations_safe_override,
only_symbols=only_symbols,
)
if valences == "undefined":
firstsite = self.structure[0]
try:
sp = firstsite.specie
if isinstance(sp, Species):
self.valences = [int(site.specie.oxi_state) for site in self.structure]
else:
self.valences = valences
except AttributeError:
self.valences = valences
else:
self.valences = valences
# Get a list of indices of unequivalent sites from the initial structure
self.equivalent_sites = [[site] for site in self.structure]
self.struct_sites_to_irreducible_site_list_map = list(range(len(self.structure)))
self.sites_map = list(range(len(self.structure)))
indices = list(range(len(self.structure)))
# Get list of unequivalent sites with valence >= 0
if only_cations and self.valences != "undefined":
sites_indices = [isite for isite in indices if self.valences[isite] >= 0]
else:
sites_indices = list(indices)
# Include atoms that are in the list of "only_atoms" if it is provided
if only_atoms is not None:
sites_indices = [
isite
for isite in sites_indices
if any([at in [sp.symbol for sp in self.structure[isite].species] for at in only_atoms])
]
# Exclude atoms that are in the list of excluded atoms
if excluded_atoms:
sites_indices = [
isite
for isite in sites_indices
if not any([at in [sp.symbol for sp in self.structure[isite].species] for at in excluded_atoms])
]
if only_indices is not None:
sites_indices = [isite for isite in indices if isite in only_indices]
# Get the VoronoiContainer for the sites defined by their indices (sites_indices)
logging.debug("Getting DetailedVoronoiContainer")
if voronoi_normalized_distance_tolerance is None:
normalized_distance_tolerance = DetailedVoronoiContainer.default_normalized_distance_tolerance
else:
normalized_distance_tolerance = voronoi_normalized_distance_tolerance
if voronoi_normalized_angle_tolerance is None:
normalized_angle_tolerance = DetailedVoronoiContainer.default_normalized_angle_tolerance
else:
normalized_angle_tolerance = voronoi_normalized_angle_tolerance
self.detailed_voronoi = DetailedVoronoiContainer(
self.structure,
isites=sites_indices,
valences=self.valences,
maximum_distance_factor=maximum_distance_factor,
minimum_angle_factor=minimum_angle_factor,
additional_conditions=additional_conditions,
normalized_distance_tolerance=normalized_distance_tolerance,
normalized_angle_tolerance=normalized_angle_tolerance,
)
logging.debug("DetailedVoronoiContainer has been set up")
# Initialize the StructureEnvironments object (either from initial_structure_environments or from scratch)
if initial_structure_environments is not None:
se = initial_structure_environments
if se.structure != self.structure:
raise ValueError("Structure is not the same in initial_structure_environments")
if se.voronoi != self.detailed_voronoi:
if self.detailed_voronoi.is_close_to(se.voronoi):
self.detailed_voronoi = se.voronoi
else:
raise ValueError("Detailed Voronoi is not the same in initial_structure_environments")
se.info = info
else:
se = StructureEnvironments(
voronoi=self.detailed_voronoi,
valences=self.valences,
sites_map=self.sites_map,
equivalent_sites=self.equivalent_sites,
ce_list=[None] * len(self.structure),
structure=self.structure,
info=info,
)
# Set up the coordination numbers that have to be computed based on min_cn, max_cn and possibly the settings
# for an update (argument "recompute") of an existing StructureEnvironments
if min_cn is None:
min_cn = 1
if max_cn is None:
max_cn = 20
all_cns = range(min_cn, max_cn + 1)
do_recompute = False
if recompute is not None:
if "cns" in recompute:
cns_to_recompute = recompute["cns"]
all_cns = list(set(all_cns).intersection(cns_to_recompute))
do_recompute = True
# Variables used for checking timelimit
max_time_one_site = 0.0
breakit = False
if optimization > 0:
self.detailed_voronoi.local_planes = [None] * len(self.structure)
self.detailed_voronoi.separations = [None] * len(self.structure)
# Loop on all the sites
for isite in range(len(self.structure)):
if isite not in sites_indices:
logging.debug(
" ... in site #{:d}/{:d} ({}) : "
"skipped".format(isite, len(self.structure), self.structure[isite].species_string)
)
continue
if breakit:
logging.debug(
" ... in site #{:d}/{:d} ({}) : "
"skipped (timelimit)".format(isite, len(self.structure), self.structure[isite].species_string)
)
continue
logging.debug(
" ... in site #{:d}/{:d} ({})".format(isite, len(self.structure), self.structure[isite].species_string)
)
t1 = time.process_time()
if optimization > 0:
self.detailed_voronoi.local_planes[isite] = OrderedDict()
self.detailed_voronoi.separations[isite] = {}
se.init_neighbors_sets(
isite=isite,
additional_conditions=additional_conditions,
valences=valences,
)
to_add_from_hints = []
nb_sets_info = {}
for cn, nb_sets in se.neighbors_sets[isite].items():
if cn not in all_cns:
continue
for inb_set, nb_set in enumerate(nb_sets):
logging.debug(" ... getting environments for nb_set ({:d}, {:d})".format(cn, inb_set))
tnbset1 = time.process_time()
ce = self.update_nb_set_environments(
se=se,
isite=isite,
cn=cn,
inb_set=inb_set,
nb_set=nb_set,
recompute=do_recompute,
optimization=optimization,
)
tnbset2 = time.process_time()
if cn not in nb_sets_info:
nb_sets_info[cn] = {}
nb_sets_info[cn][inb_set] = {"time": tnbset2 - tnbset1}
if get_from_hints:
for cg_symbol, cg_dict in ce:
cg = self.allcg[cg_symbol]
# Get possibly missing neighbors sets
if cg.neighbors_sets_hints is None:
continue
logging.debug(' ... getting hints from cg with mp_symbol "{}" ...'.format(cg_symbol))
hints_info = {
"csm": cg_dict["symmetry_measure"],
"nb_set": nb_set,
"permutation": cg_dict["permutation"],
}
for nb_sets_hints in cg.neighbors_sets_hints:
suggested_nb_set_voronoi_indices = nb_sets_hints.hints(hints_info)
for inew, new_nb_set_voronoi_indices in enumerate(suggested_nb_set_voronoi_indices):
logging.debug(" hint # {:d}".format(inew))
new_nb_set = se.NeighborsSet(
structure=se.structure,
isite=isite,
detailed_voronoi=se.voronoi,
site_voronoi_indices=new_nb_set_voronoi_indices,
sources={
"origin": "nb_set_hints",
"hints_type": nb_sets_hints.hints_type,
"suggestion_index": inew,
"cn_map_source": [cn, inb_set],
"cg_source_symbol": cg_symbol,
},
)
cn_new_nb_set = len(new_nb_set)
if max_cn is not None and cn_new_nb_set > max_cn:
continue
if min_cn is not None and cn_new_nb_set < min_cn:
continue
if new_nb_set in [ta["new_nb_set"] for ta in to_add_from_hints]:
has_nb_set = True
elif cn_new_nb_set not in se.neighbors_sets[isite]:
has_nb_set = False
else:
has_nb_set = new_nb_set in se.neighbors_sets[isite][cn_new_nb_set]
if not has_nb_set:
to_add_from_hints.append(
{
"isite": isite,
"new_nb_set": new_nb_set,
"cn_new_nb_set": cn_new_nb_set,
}
)
logging.debug(" => to be computed")
else:
logging.debug(" => already present")
logging.debug(" ... getting environments for nb_sets added from hints")
for missing_nb_set_to_add in to_add_from_hints:
se.add_neighbors_set(isite=isite, nb_set=missing_nb_set_to_add["new_nb_set"])
for missing_nb_set_to_add in to_add_from_hints:
isite_new_nb_set = missing_nb_set_to_add["isite"]
cn_new_nb_set = missing_nb_set_to_add["cn_new_nb_set"]
new_nb_set = missing_nb_set_to_add["new_nb_set"]
inew_nb_set = se.neighbors_sets[isite_new_nb_set][cn_new_nb_set].index(new_nb_set)
logging.debug(
" ... getting environments for nb_set ({:d}, {:d}) - "
"from hints".format(cn_new_nb_set, inew_nb_set)
)
tnbset1 = time.process_time()
self.update_nb_set_environments(
se=se,
isite=isite_new_nb_set,
cn=cn_new_nb_set,
inb_set=inew_nb_set,
nb_set=new_nb_set,
optimization=optimization,
)
tnbset2 = time.process_time()
if cn not in nb_sets_info:
nb_sets_info[cn] = {}
nb_sets_info[cn][inew_nb_set] = {"time": tnbset2 - tnbset1}
t2 = time.process_time()
se.update_site_info(isite=isite, info_dict={"time": t2 - t1, "nb_sets_info": nb_sets_info})
if timelimit is not None:
time_elapsed = t2 - time_init
time_left = timelimit - time_elapsed
if time_left < 2.0 * max_time_one_site:
breakit = True
max_time_one_site = max(max_time_one_site, t2 - t1)
logging.debug(" ... computed in {:.2f} seconds".format(t2 - t1))
time_end = time.process_time()
logging.debug(" ... compute_structure_environments ended in {:.2f} seconds".format(time_end - time_init))
return se
def update_nb_set_environments(self, se, isite, cn, inb_set, nb_set, recompute=False, optimization=None):
"""
:param se:
:param isite:
:param cn:
:param inb_set:
:param nb_set:
:param recompute:
:param optimization:
:return:
"""
ce = se.get_coordination_environments(isite=isite, cn=cn, nb_set=nb_set)
if ce is not None and not recompute:
return ce
ce = ChemicalEnvironments()
if optimization == 2:
neighb_coords = nb_set.neighb_coordsOpt
else:
neighb_coords = nb_set.neighb_coords
self.setup_local_geometry(isite, coords=neighb_coords, optimization=optimization)
if optimization > 0:
logging.debug("Getting StructureEnvironments with optimized algorithm")
nb_set.local_planes = OrderedDict()
nb_set.separations = {}
cncgsm = self.get_coordination_symmetry_measures_optim(nb_set=nb_set, optimization=optimization)
else:
logging.debug("Getting StructureEnvironments with standard algorithm")
cncgsm = self.get_coordination_symmetry_measures()
for cg in cncgsm:
other_csms = {
"csm_wocs_ctwocc": cncgsm[cg]["csm_wocs_ctwocc"],
"csm_wocs_ctwcc": cncgsm[cg]["csm_wocs_ctwcc"],
"csm_wocs_csc": cncgsm[cg]["csm_wocs_csc"],
"csm_wcs_ctwocc": cncgsm[cg]["csm_wcs_ctwocc"],
"csm_wcs_ctwcc": cncgsm[cg]["csm_wcs_ctwcc"],
"csm_wcs_csc": cncgsm[cg]["csm_wcs_csc"],
"rotation_matrix_wocs_ctwocc": cncgsm[cg]["rotation_matrix_wocs_ctwocc"],
"rotation_matrix_wocs_ctwcc": cncgsm[cg]["rotation_matrix_wocs_ctwcc"],
"rotation_matrix_wocs_csc": cncgsm[cg]["rotation_matrix_wocs_csc"],
"rotation_matrix_wcs_ctwocc": cncgsm[cg]["rotation_matrix_wcs_ctwocc"],
"rotation_matrix_wcs_ctwcc": cncgsm[cg]["rotation_matrix_wcs_ctwcc"],
"rotation_matrix_wcs_csc": cncgsm[cg]["rotation_matrix_wcs_csc"],
"scaling_factor_wocs_ctwocc": cncgsm[cg]["scaling_factor_wocs_ctwocc"],
"scaling_factor_wocs_ctwcc": cncgsm[cg]["scaling_factor_wocs_ctwcc"],
"scaling_factor_wocs_csc": cncgsm[cg]["scaling_factor_wocs_csc"],
"scaling_factor_wcs_ctwocc": cncgsm[cg]["scaling_factor_wcs_ctwocc"],
"scaling_factor_wcs_ctwcc": cncgsm[cg]["scaling_factor_wcs_ctwcc"],
"scaling_factor_wcs_csc": cncgsm[cg]["scaling_factor_wcs_csc"],
"translation_vector_wocs_ctwocc": cncgsm[cg]["translation_vector_wocs_ctwocc"],
"translation_vector_wocs_ctwcc": cncgsm[cg]["translation_vector_wocs_ctwcc"],
"translation_vector_wocs_csc": cncgsm[cg]["translation_vector_wocs_csc"],
"translation_vector_wcs_ctwocc": cncgsm[cg]["translation_vector_wcs_ctwocc"],
"translation_vector_wcs_ctwcc": cncgsm[cg]["translation_vector_wcs_ctwcc"],
"translation_vector_wcs_csc": cncgsm[cg]["translation_vector_wcs_csc"],
}
ce.add_coord_geom(
cg,
cncgsm[cg]["csm"],
algo=cncgsm[cg]["algo"],
permutation=cncgsm[cg]["indices"],
local2perfect_map=cncgsm[cg]["local2perfect_map"],
perfect2local_map=cncgsm[cg]["perfect2local_map"],
detailed_voronoi_index={"cn": cn, "index": inb_set},
other_symmetry_measures=other_csms,
rotation_matrix=cncgsm[cg]["rotation_matrix"],
scaling_factor=cncgsm[cg]["scaling_factor"],
)
se.update_coordination_environments(isite=isite, cn=cn, nb_set=nb_set, ce=ce)
return ce
def setup_local_geometry(self, isite, coords, optimization=None):
"""
Sets up the AbstractGeometry for the local geometry of site with index isite.
:param isite: Index of the site for which the local geometry has to be set up
:param coords: The coordinates of the (local) neighbors
"""
self.local_geometry = AbstractGeometry(
central_site=self.structure.cart_coords[isite],
bare_coords=coords,
centering_type=self.centering_type,
include_central_site_in_centroid=self.include_central_site_in_centroid,
optimization=optimization,
)
def setup_test_perfect_environment(
self,
symbol,
randomness=False,
max_random_dist=0.1,
symbol_type="mp_symbol",
indices="RANDOM",
random_translation="NONE",
random_rotation="NONE",
random_scale="NONE",
points=None,
):
"""
:param symbol:
:param randomness:
:param max_random_dist:
:param symbol_type:
:param indices:
:param random_translation:
:param random_rotation:
:param random_scale:
:param points:
:return:
"""
if symbol_type == "IUPAC":
cg = self.allcg.get_geometry_from_IUPAC_symbol(symbol)
elif symbol_type in ("MP", "mp_symbol"):
cg = self.allcg.get_geometry_from_mp_symbol(symbol)
elif symbol_type == "CoordinationGeometry":
cg = symbol
else:
raise ValueError("Wrong mp_symbol to setup coordination geometry")
neighb_coords = []
if points is not None:
mypoints = points
else:
mypoints = cg.points
if randomness:
rv = np.random.random_sample(3)
while norm(rv) > 1.0:
rv = np.random.random_sample(3)
coords = [np.zeros(3, np.float) + max_random_dist * rv]
for pp in mypoints:
rv = np.random.random_sample(3)
while norm(rv) > 1.0:
rv = np.random.random_sample(3)
neighb_coords.append(np.array(pp) + max_random_dist * rv)
else:
coords = [np.zeros(3, np.float)]
for pp in mypoints:
neighb_coords.append(np.array(pp))
if indices == "RANDOM":
shuffle(neighb_coords)
elif indices == "ORDERED":
pass
else:
neighb_coords = [neighb_coords[ii] for ii in indices]
# Scaling the test environment
if random_scale == "RANDOM":
scale = 0.1 * np.random.random_sample() + 0.95
elif random_scale == "NONE":
scale = 1.0
else:
scale = random_scale
coords = [scale * cc for cc in coords]
neighb_coords = [scale * cc for cc in neighb_coords]
# Rotating the test environment
if random_rotation == "RANDOM":
uu = np.random.random_sample(3) + 0.1
uu = uu / norm(uu)
theta = np.pi * np.random.random_sample()
cc = np.cos(theta)
ss = np.sin(theta)
ux = uu[0]
uy = uu[1]
uz = uu[2]
RR = [
[
ux * ux + (1.0 - ux * ux) * cc,
ux * uy * (1.0 - cc) - uz * ss,
ux * uz * (1.0 - cc) + uy * ss,
],
[
ux * uy * (1.0 - cc) + uz * ss,
uy * uy + (1.0 - uy * uy) * cc,
uy * uz * (1.0 - cc) - ux * ss,
],
[
ux * uz * (1.0 - cc) - uy * ss,
uy * uz * (1.0 - cc) + ux * ss,
uz * uz + (1.0 - uz * uz) * cc,
],
]
elif random_rotation == "NONE":
RR = [[1.0, 0.0, 0.0], [0.0, 1.0, 0.0], [0.0, 0.0, 1.0]]
else:
RR = random_rotation
newcoords = []
for cc in coords:
newcc = np.dot(RR, cc).T
newcoords.append(newcc.ravel())
coords = newcoords
newcoords = []
for cc in neighb_coords:
newcc = np.dot(RR, cc.T)
newcoords.append(newcc.ravel())
neighb_coords = newcoords
# Translating the test environment
if random_translation == "RANDOM":
translation = 10.0 * (2.0 * np.random.random_sample(3) - 1.0)
elif random_translation == "NONE":
translation = np.zeros(3, np.float)
else:
translation = random_translation
coords = [cc + translation for cc in coords]
neighb_coords = [cc + translation for cc in neighb_coords]
coords.extend(neighb_coords)
myspecies = ["O"] * (len(coords))
myspecies[0] = "Cu"
amin = np.min([cc[0] for cc in coords])
amax = np.max([cc[0] for cc in coords])
bmin = np.min([cc[1] for cc in coords])
bmax = np.max([cc[1] for cc in coords])
cmin = np.min([cc[2] for cc in coords])
cmax = np.max([cc[2] for cc in coords])
factor = 5.0
aa = factor * max([amax - amin, bmax - bmin, cmax - cmin])
lattice = Lattice.cubic(a=aa)
structure = Structure(
lattice=lattice,
species=myspecies,
coords=coords,
to_unit_cell=False,
coords_are_cartesian=True,
)
self.setup_structure(structure=structure)
self.setup_local_geometry(isite=0, coords=neighb_coords)
self.perfect_geometry = AbstractGeometry.from_cg(cg=cg)
def setup_random_structure(self, coordination):
"""
Sets up a purely random structure with a given coordination.
:param coordination: coordination number for the random structure
"""
aa = 0.4
bb = -0.2
coords = list()
for ii in range(coordination + 1):
coords.append(
aa
* np.random.random_sample(
3,
)
+ bb
)
self.set_structure(
lattice=np.array([[10, 0, 0], [0, 10, 0], [0, 0, 10]], np.float),
species=["Si"] * (coordination + 1),
coords=coords,
coords_are_cartesian=False,
)
self.setup_random_indices_local_geometry(coordination)
def setup_random_indices_local_geometry(self, coordination):
"""
Sets up random indices for the local geometry, for testing purposes
:param coordination: coordination of the local geometry
"""
self.icentral_site = 0
self.indices = list(range(1, coordination + 1))
np.random.shuffle(self.indices)
def setup_ordered_indices_local_geometry(self, coordination):
"""
Sets up ordered indices for the local geometry, for testing purposes
:param coordination: coordination of the local geometry
"""
self.icentral_site = 0
self.indices = list(range(1, coordination + 1))
def setup_explicit_indices_local_geometry(self, explicit_indices):
"""
Sets up explicit indices for the local geometry, for testing purposes
:param explicit_indices: explicit indices for the neighbors (set of numbers
from 0 to CN-1 in a given order)
"""
self.icentral_site = 0
self.indices = [ii + 1 for ii in explicit_indices]
def get_coordination_symmetry_measures(self, only_minimum=True, all_csms=True, optimization=None):
"""
Returns the continuous symmetry measures of the current local geometry in a dictionary.
:return: the continuous symmetry measures of the current local geometry in a dictionary.
"""
test_geometries = self.allcg.get_implemented_geometries(len(self.local_geometry.coords))
if len(self.local_geometry.coords) == 1:
if len(test_geometries) == 0:
return {}
result_dict = {
"S:1": {
"csm": 0.0,
"indices": [0],
"algo": "EXPLICIT",
"local2perfect_map": {0: 0},
"perfect2local_map": {0: 0},
"scaling_factor": None,
"rotation_matrix": None,
"translation_vector": None,
}
}
if all_csms:
for csmtype in [
"wocs_ctwocc",
"wocs_ctwcc",
"wocs_csc",
"wcs_ctwocc",
"wcs_ctwcc",
"wcs_csc",
]:
result_dict["S:1"]["csm_{}".format(csmtype)] = 0.0
result_dict["S:1"]["scaling_factor_{}".format(csmtype)] = None
result_dict["S:1"]["rotation_matrix_{}".format(csmtype)] = None
result_dict["S:1"]["translation_vector_{}".format(csmtype)] = None
return result_dict
result_dict = {}
for geometry in test_geometries:
self.perfect_geometry = AbstractGeometry.from_cg(
cg=geometry,
centering_type=self.centering_type,
include_central_site_in_centroid=self.include_central_site_in_centroid,
)
points_perfect = self.perfect_geometry.points_wcs_ctwcc()
cgsm = self.coordination_geometry_symmetry_measures(
geometry, points_perfect=points_perfect, optimization=optimization
)
result, permutations, algos, local2perfect_maps, perfect2local_maps = cgsm
if only_minimum:
if len(result) > 0:
imin = np.argmin([rr["symmetry_measure"] for rr in result])
if geometry.algorithms is not None:
algo = algos[imin]
else:
algo = algos
result_dict[geometry.mp_symbol] = {
"csm": result[imin]["symmetry_measure"],
"indices": permutations[imin],
"algo": algo,
"local2perfect_map": local2perfect_maps[imin],
"perfect2local_map": perfect2local_maps[imin],
"scaling_factor": 1.0 / result[imin]["scaling_factor"],
"rotation_matrix": np.linalg.inv(result[imin]["rotation_matrix"]),
"translation_vector": result[imin]["translation_vector"],
}
if all_csms:
self._update_results_all_csms(result_dict, permutations, imin, geometry)
else:
result_dict[geometry.mp_symbol] = {
"csm": result,
"indices": permutations,
"algo": algos,
"local2perfect_map": local2perfect_maps,
"perfect2local_map": perfect2local_maps,
}
return result_dict
def _update_results_all_csms(self, result_dict, permutations, imin, geometry):
permutation = permutations[imin]
# Without central site, centered on the centroid (centroid does not include the central site)
# result_dict[geometry.mp_symbol]['csm_wocs_ctwocc'] = \
# result[imin]
pdist = self.local_geometry.points_wocs_ctwocc(permutation=permutation)
pperf = self.perfect_geometry.points_wocs_ctwocc()
sm_info = symmetry_measure(points_distorted=pdist, points_perfect=pperf)
result_dict[geometry.mp_symbol]["csm_wocs_ctwocc"] = sm_info["symmetry_measure"]
result_dict[geometry.mp_symbol]["rotation_matrix_wocs_ctwocc"] = np.linalg.inv(sm_info["rotation_matrix"])
result_dict[geometry.mp_symbol]["scaling_factor_wocs_ctwocc"] = 1.0 / sm_info["scaling_factor"]
result_dict[geometry.mp_symbol]["translation_vector_wocs_ctwocc"] = self.local_geometry.centroid_without_centre
# Without central site, centered on the centroid (centroid includes the central site)
pdist = self.local_geometry.points_wocs_ctwcc(permutation=permutation)
pperf = self.perfect_geometry.points_wocs_ctwcc()
sm_info = symmetry_measure(points_distorted=pdist, points_perfect=pperf)
result_dict[geometry.mp_symbol]["csm_wocs_ctwcc"] = sm_info["symmetry_measure"]
result_dict[geometry.mp_symbol]["rotation_matrix_wocs_ctwcc"] = np.linalg.inv(sm_info["rotation_matrix"])
result_dict[geometry.mp_symbol]["scaling_factor_wocs_ctwcc"] = 1.0 / sm_info["scaling_factor"]
result_dict[geometry.mp_symbol]["translation_vector_wocs_ctwcc"] = self.local_geometry.centroid_with_centre
# Without central site, centered on the central site
pdist = self.local_geometry.points_wocs_csc(permutation=permutation)
pperf = self.perfect_geometry.points_wocs_csc()
sm_info = symmetry_measure(points_distorted=pdist, points_perfect=pperf)
result_dict[geometry.mp_symbol]["csm_wocs_csc"] = sm_info["symmetry_measure"]
result_dict[geometry.mp_symbol]["rotation_matrix_wocs_csc"] = np.linalg.inv(sm_info["rotation_matrix"])
result_dict[geometry.mp_symbol]["scaling_factor_wocs_csc"] = 1.0 / sm_info["scaling_factor"]
result_dict[geometry.mp_symbol]["translation_vector_wocs_csc"] = self.local_geometry.bare_centre
# With central site, centered on the centroid (centroid does not include the central site)
pdist = self.local_geometry.points_wcs_ctwocc(permutation=permutation)
pperf = self.perfect_geometry.points_wcs_ctwocc()
sm_info = symmetry_measure(points_distorted=pdist, points_perfect=pperf)
result_dict[geometry.mp_symbol]["csm_wcs_ctwocc"] = sm_info["symmetry_measure"]
result_dict[geometry.mp_symbol]["rotation_matrix_wcs_ctwocc"] = np.linalg.inv(sm_info["rotation_matrix"])
result_dict[geometry.mp_symbol]["scaling_factor_wcs_ctwocc"] = 1.0 / sm_info["scaling_factor"]
result_dict[geometry.mp_symbol]["translation_vector_wcs_ctwocc"] = self.local_geometry.centroid_without_centre
# With central site, centered on the centroid (centroid includes the central site)
pdist = self.local_geometry.points_wcs_ctwcc(permutation=permutation)
pperf = self.perfect_geometry.points_wcs_ctwcc()
sm_info = symmetry_measure(points_distorted=pdist, points_perfect=pperf)
result_dict[geometry.mp_symbol]["csm_wcs_ctwcc"] = sm_info["symmetry_measure"]
result_dict[geometry.mp_symbol]["rotation_matrix_wcs_ctwcc"] = np.linalg.inv(sm_info["rotation_matrix"])
result_dict[geometry.mp_symbol]["scaling_factor_wcs_ctwcc"] = 1.0 / sm_info["scaling_factor"]
result_dict[geometry.mp_symbol]["translation_vector_wcs_ctwcc"] = self.local_geometry.centroid_with_centre
# With central site, centered on the central site
pdist = self.local_geometry.points_wcs_csc(permutation=permutation)
pperf = self.perfect_geometry.points_wcs_csc()
sm_info = symmetry_measure(points_distorted=pdist, points_perfect=pperf)
result_dict[geometry.mp_symbol]["csm_wcs_csc"] = sm_info["symmetry_measure"]
result_dict[geometry.mp_symbol]["rotation_matrix_wcs_csc"] = np.linalg.inv(sm_info["rotation_matrix"])
result_dict[geometry.mp_symbol]["scaling_factor_wcs_csc"] = 1.0 / sm_info["scaling_factor"]
result_dict[geometry.mp_symbol]["translation_vector_wcs_csc"] = self.local_geometry.bare_centre
def get_coordination_symmetry_measures_optim(
self, only_minimum=True, all_csms=True, nb_set=None, optimization=None
):
"""
Returns the continuous symmetry measures of the current local geometry in a dictionary.
:return: the continuous symmetry measures of the current local geometry in a dictionary.
"""
cn = len(self.local_geometry.coords)
test_geometries = self.allcg.get_implemented_geometries(cn)
if all([cg.algorithms[0].algorithm_type == EXPLICIT_PERMUTATIONS for cg in test_geometries]):
return self.get_coordination_symmetry_measures(
only_minimum=only_minimum, all_csms=all_csms, optimization=optimization
)
if not all(
[all([algo.algorithm_type == SEPARATION_PLANE for algo in cg.algorithms]) for cg in test_geometries]
):
raise ValueError("All algorithms should be EXPLICIT_PERMUTATIONS or SEPARATION_PLANE")
result_dict = {}
for geometry in test_geometries:
logging.log(
level=5,
msg="Getting Continuous Symmetry Measure with Separation Plane "
'algorithm for geometry "{}"'.format(geometry.ce_symbol),
)
self.perfect_geometry = AbstractGeometry.from_cg(
cg=geometry,
centering_type=self.centering_type,
include_central_site_in_centroid=self.include_central_site_in_centroid,
)
points_perfect = self.perfect_geometry.points_wcs_ctwcc()
cgsm = self.coordination_geometry_symmetry_measures_sepplane_optim(
geometry,
points_perfect=points_perfect,
nb_set=nb_set,
optimization=optimization,
)
result, permutations, algos, local2perfect_maps, perfect2local_maps = cgsm
if only_minimum:
if len(result) > 0:
imin = np.argmin([rr["symmetry_measure"] for rr in result])
if geometry.algorithms is not None:
algo = algos[imin]
else:
algo = algos
result_dict[geometry.mp_symbol] = {
"csm": result[imin]["symmetry_measure"],
"indices": permutations[imin],
"algo": algo,
"local2perfect_map": local2perfect_maps[imin],
"perfect2local_map": perfect2local_maps[imin],
"scaling_factor": 1.0 / result[imin]["scaling_factor"],
"rotation_matrix": np.linalg.inv(result[imin]["rotation_matrix"]),
"translation_vector": result[imin]["translation_vector"],
}
if all_csms:
self._update_results_all_csms(result_dict, permutations, imin, geometry)
return result_dict
def coordination_geometry_symmetry_measures(
self,
coordination_geometry,
tested_permutations=False,
points_perfect=None,
optimization=None,
):
"""
Returns the symmetry measures of a given coordination_geometry for a set of permutations depending on
the permutation setup. Depending on the parameters of the LocalGeometryFinder and on the coordination
geometry, different methods are called.
:param coordination_geometry: Coordination geometry for which the symmetry measures are looked for
:return: the symmetry measures of a given coordination_geometry for a set of permutations
:raise: NotImplementedError if the permutation_setup does not exists
"""
if tested_permutations:
tested_permutations = set()
if self.permutations_safe_override:
raise ValueError("No permutations safe override anymore")
csms = []
permutations = []
algos = []
local2perfect_maps = []
perfect2local_maps = []
for algo in coordination_geometry.algorithms:
if algo.algorithm_type == EXPLICIT_PERMUTATIONS:
return self.coordination_geometry_symmetry_measures_standard(
coordination_geometry,
algo,
points_perfect=points_perfect,
optimization=optimization,
)
if algo.algorithm_type == SEPARATION_PLANE:
cgsm = self.coordination_geometry_symmetry_measures_separation_plane(
coordination_geometry,
algo,
tested_permutations=tested_permutations,
points_perfect=points_perfect,
)
csm, perm, algo, local2perfect_map, perfect2local_map = cgsm
csms.extend(csm)
permutations.extend(perm)
algos.extend(algo)
local2perfect_maps.extend(local2perfect_map)
perfect2local_maps.extend(perfect2local_map)
return csms, permutations, algos, local2perfect_maps, perfect2local_maps
def coordination_geometry_symmetry_measures_sepplane_optim(
self, coordination_geometry, points_perfect=None, nb_set=None, optimization=None
):
"""
Returns the symmetry measures of a given coordination_geometry for a set of permutations depending on
the permutation setup. Depending on the parameters of the LocalGeometryFinder and on the coordination
geometry, different methods are called.
:param coordination_geometry: Coordination geometry for which the symmetry measures are looked for
:return: the symmetry measures of a given coordination_geometry for a set of permutations
:raise: NotImplementedError if the permutation_setup does not exists
"""
csms = []
permutations = []
algos = []
local2perfect_maps = []
perfect2local_maps = []
for algo in coordination_geometry.algorithms:
if algo.algorithm_type == SEPARATION_PLANE:
cgsm = self.coordination_geometry_symmetry_measures_separation_plane_optim(
coordination_geometry,
algo,
points_perfect=points_perfect,
nb_set=nb_set,
optimization=optimization,
)
csm, perm, algo, local2perfect_map, perfect2local_map = cgsm
csms.extend(csm)
permutations.extend(perm)
algos.extend(algo)
local2perfect_maps.extend(local2perfect_map)
perfect2local_maps.extend(perfect2local_map)
return csms, permutations, algos, local2perfect_maps, perfect2local_maps
def coordination_geometry_symmetry_measures_standard(
self, coordination_geometry, algo, points_perfect=None, optimization=None
):
"""
Returns the symmetry measures for a set of permutations (whose setup depends on the coordination geometry)
for the coordination geometry "coordination_geometry". Standard implementation looking for the symmetry
measures of each permutation
:param coordination_geometry: The coordination geometry to be investigated
:return: The symmetry measures for the given coordination geometry for each permutation investigated
"""
# permutations_symmetry_measures = np.zeros(len(algo.permutations),
# np.float)
if optimization == 2:
permutations_symmetry_measures = [None] * len(algo.permutations)
permutations = list()
algos = list()
local2perfect_maps = list()
perfect2local_maps = list()
for iperm, perm in enumerate(algo.permutations):
local2perfect_map = {}
perfect2local_map = {}
permutations.append(perm)
for iperfect, ii in enumerate(perm):
perfect2local_map[iperfect] = ii
local2perfect_map[ii] = iperfect
local2perfect_maps.append(local2perfect_map)
perfect2local_maps.append(perfect2local_map)
points_distorted = self.local_geometry.points_wcs_ctwcc(permutation=perm)
sm_info = symmetry_measure(points_distorted=points_distorted, points_perfect=points_perfect)
sm_info["translation_vector"] = self.local_geometry.centroid_with_centre
permutations_symmetry_measures[iperm] = sm_info
algos.append(str(algo))
return (
permutations_symmetry_measures,
permutations,
algos,
local2perfect_maps,
perfect2local_maps,
)
permutations_symmetry_measures = [None] * len(algo.permutations)
permutations = list()
algos = list()
local2perfect_maps = list()
perfect2local_maps = list()
for iperm, perm in enumerate(algo.permutations):
local2perfect_map = {}
perfect2local_map = {}
permutations.append(perm)
for iperfect, ii in enumerate(perm):
perfect2local_map[iperfect] = ii
local2perfect_map[ii] = iperfect
local2perfect_maps.append(local2perfect_map)
perfect2local_maps.append(perfect2local_map)
points_distorted = self.local_geometry.points_wcs_ctwcc(permutation=perm)
sm_info = symmetry_measure(points_distorted=points_distorted, points_perfect=points_perfect)
sm_info["translation_vector"] = self.local_geometry.centroid_with_centre
permutations_symmetry_measures[iperm] = sm_info
algos.append(str(algo))
return (
permutations_symmetry_measures,
permutations,
algos,
local2perfect_maps,
perfect2local_maps,
)
def coordination_geometry_symmetry_measures_separation_plane(
self,
coordination_geometry,
separation_plane_algo,
testing=False,
tested_permutations=False,
points_perfect=None,
):
"""
Returns the symmetry measures of the given coordination geometry "coordination_geometry" using separation
facets to reduce the complexity of the system. Caller to the refined 2POINTS, 3POINTS and other ...
:param coordination_geometry: The coordination geometry to be investigated
:return: The symmetry measures for the given coordination geometry for each plane and permutation investigated
"""
permutations = list()
permutations_symmetry_measures = list()
plane_separations = list()
algos = list()
perfect2local_maps = list()
local2perfect_maps = list()
if testing:
separation_permutations = list()
nplanes = 0
for npoints in range(
separation_plane_algo.minimum_number_of_points,
min(separation_plane_algo.maximum_number_of_points, 4) + 1,
):
for points_combination in itertools.combinations(self.local_geometry.coords, npoints):
if npoints == 2:
if collinear(
points_combination[0],
points_combination[1],
self.local_geometry.central_site,
tolerance=0.25,
):
continue
plane = Plane.from_3points(
points_combination[0],
points_combination[1],
self.local_geometry.central_site,
)
elif npoints == 3:
if collinear(
points_combination[0],
points_combination[1],
points_combination[2],
tolerance=0.25,
):
continue
plane = Plane.from_3points(
points_combination[0],
points_combination[1],
points_combination[2],
)
elif npoints > 3:
plane = Plane.from_npoints(points_combination, best_fit="least_square_distance")
else:
raise ValueError("Wrong number of points to initialize separation plane")
cgsm = self._cg_csm_separation_plane(
coordination_geometry=coordination_geometry,
sepplane=separation_plane_algo,
local_plane=plane,
plane_separations=plane_separations,
dist_tolerances=DIST_TOLERANCES,
testing=testing,
tested_permutations=tested_permutations,
points_perfect=points_perfect,
)
csm, perm, algo = cgsm[0], cgsm[1], cgsm[2]
if csm is not None:
permutations_symmetry_measures.extend(csm)
permutations.extend(perm)
for thisperm in perm:
p2l = {}
l2p = {}
for i_p, pp in enumerate(thisperm):
p2l[i_p] = pp
l2p[pp] = i_p
perfect2local_maps.append(p2l)
local2perfect_maps.append(l2p)
algos.extend(algo)
if testing:
separation_permutations.extend(cgsm[3])
nplanes += 1
if nplanes > 0:
break
if nplanes == 0:
return self.coordination_geometry_symmetry_measures_fallback_random(
coordination_geometry, points_perfect=points_perfect
)
if testing:
return permutations_symmetry_measures, permutations, separation_permutations
return (
permutations_symmetry_measures,
permutations,
algos,
local2perfect_maps,
perfect2local_maps,
)
def coordination_geometry_symmetry_measures_separation_plane_optim(
self,
coordination_geometry,
separation_plane_algo,
points_perfect=None,
nb_set=None,
optimization=None,
):
"""
Returns the symmetry measures of the given coordination geometry "coordination_geometry" using separation
facets to reduce the complexity of the system. Caller to the refined 2POINTS, 3POINTS and other ...
Args:
coordination_geometry: The coordination geometry to be investigated.
separation_plane_algo: Separation Plane algorithm used.
points_perfect: Points corresponding to the perfect geometry.
nb_set: Neighbor set for this set of points. (used to store already computed separation planes)
optimization: Optimization level (1 or 2).
Returns:
tuple: Continuous symmetry measures for the given coordination geometry for each plane and permutation
investigated, corresponding permutations, corresponding algorithms,
corresponding mappings from local to perfect environment and corresponding mappings
from perfect to local environment.
"""
if optimization == 2:
logging.log(level=5, msg="... using optimization = 2")
cgcsmoptim = self._cg_csm_separation_plane_optim2
elif optimization == 1:
logging.log(level=5, msg="... using optimization = 2")
cgcsmoptim = self._cg_csm_separation_plane_optim1
else:
raise ValueError("Optimization should be 1 or 2")
cn = len(self.local_geometry.coords)
permutations = list()
permutations_symmetry_measures = list()
algos = list()
perfect2local_maps = list()
local2perfect_maps = list()
if separation_plane_algo.separation in nb_set.separations:
for sep_indices, (local_plane, npsep) in nb_set.separations[separation_plane_algo.separation].items():
cgsm = cgcsmoptim(
coordination_geometry=coordination_geometry,
sepplane=separation_plane_algo,
local_plane=local_plane,
points_perfect=points_perfect,
separation_indices=npsep,
)
csm, perm, algo, _ = cgsm[0], cgsm[1], cgsm[2], cgsm[3]
permutations_symmetry_measures.extend(csm)
permutations.extend(perm)
for thisperm in perm:
p2l = {}
l2p = {}
for i_p, pp in enumerate(thisperm):
p2l[i_p] = pp
l2p[pp] = i_p
perfect2local_maps.append(p2l)
local2perfect_maps.append(l2p)
algos.extend(algo)
# Get the local planes and separations up to 3 points
for npoints in range(self.allcg.minpoints[cn], min(self.allcg.maxpoints[cn], 3) + 1):
for ipoints_combination in itertools.combinations(range(self.local_geometry.cn), npoints):
if ipoints_combination in nb_set.local_planes:
continue
# Set up new plane
nb_set.local_planes[ipoints_combination] = None
points_combination = [self.local_geometry.coords[ip] for ip in ipoints_combination]
if npoints == 2:
if collinear(
points_combination[0],
points_combination[1],
self.local_geometry.central_site,
tolerance=0.25,
):
continue
plane = Plane.from_3points(
points_combination[0],
points_combination[1],
self.local_geometry.central_site,
)
elif npoints == 3:
if collinear(
points_combination[0],
points_combination[1],
points_combination[2],
tolerance=0.25,
):
continue
plane = Plane.from_3points(
points_combination[0],
points_combination[1],
points_combination[2],
)
elif npoints > 3:
plane = Plane.from_npoints(points_combination, best_fit="least_square_distance")
else:
raise ValueError("Wrong number of points to initialize separation plane")
# Takes a lot of time and happens rarely ...
# if any([plane.is_same_plane_as(plane2) for comb2, plane2 in nb_set.local_planes.items()
# if plane2 is not None]):
# continue
nb_set.local_planes[ipoints_combination] = plane
# Get the separations for this plane
# TODO: check sensitivity to delta/delta_factor parameter
dig = plane.distances_indices_groups(points=self.local_geometry._coords, delta_factor=0.1, sign=True)
grouped_indices = dig[2]
new_seps = []
for ng in range(1, len(grouped_indices) + 1):
inplane = list(itertools.chain(*grouped_indices[:ng]))
if len(inplane) > self.allcg.maxpoints_inplane[cn]:
break
inplane = [ii[0] for ii in inplane]
outplane = list(itertools.chain(*grouped_indices[ng:]))
s1 = [ii_sign[0] for ii_sign in outplane if ii_sign[1] < 0]
s2 = [ii_sign[0] for ii_sign in outplane if ii_sign[1] > 0]
separation = sort_separation_tuple([s1, inplane, s2])
sep = tuple([len(gg) for gg in separation])
if sep not in self.allcg.separations_cg[cn]:
continue
if sep not in nb_set.separations:
nb_set.separations[sep] = {}
mysep = [
|
np.array(ss, dtype=np.int8)
|
numpy.array
|
"""
Script plots trends of various variables over the WACC period. Subplot compares
all six experiments with ERA-Interim.
Notes
-----
Author : <NAME>
Date : 20 February 2019
"""
### Import modules
import datetime
import numpy as np
import matplotlib.pyplot as plt
import cmocean
from mpl_toolkits.basemap import Basemap, addcyclic, shiftgrid
import read_MonthlyData as MOM
import read_Reanalysis as MOR
import calc_Utilities as UT
### Define time
now = datetime.datetime.now()
currentmn = str(now.month)
currentdy = str(now.day)
currentyr = str(now.year)
currenttime = currentmn + '_' + currentdy + '_' + currentyr
titletime = currentmn + '/' + currentdy + '/' + currentyr
print('\n' '----Plotting WACC Variable Trends - %s----' % titletime)
#### Alott time series
year1 = 1979
year2 = 2016
years = np.arange(year1,year2+1,1)
### Add parameters
ensembles = 10
su = [0,1,2,3,5,6,7]
period = 'AMJ'
varnames = ['T2M','SLP','Z500','Z50','U200','U10']
varnames = ['THICK']
runnames = [r'ERA-I',r'CSST',r'CSIC',r'AMIP',r'AMQ',r'AMS',r'AMQS']
runnamesm = [r'CSST',r'CSIC',r'AMIP',r'AMQ',r'AMS',r'AMQS']
### Define directories
directoryfigure = '/home/zlabe/Desktop/Trends/Trends_%s/' % period
for v in range(len(varnames)):
### Call function to read in ERA-Interim
lat,lon,time,lev,era = MOR.readDataR(varnames[v],'surface',False,True)
### Call functions to read in WACCM data
models = np.empty((len(runnamesm),ensembles,era.shape[0],era.shape[1],
era.shape[2],era.shape[3]))
for i in range(len(runnamesm)):
lat,lon,time,lev,models[i] = MOM.readDataM(varnames[v],runnamesm[i],
'surface',False,True)
### Retrieve time period of interest
if period == 'DJF':
modq = np.empty((len(runnamesm),ensembles,era.shape[0]-1,era.shape[2],
era.shape[3]))
for i in range(len(runnamesm)):
for j in range(ensembles):
modq[i,j,:,:,:] = UT.calcDecJanFeb(models[i,j,:,:,:],
lat,lon,'surface',1)
eraq = UT.calcDecJanFeb(era,lat,lon,'surface',1)
elif period == 'JF':
modq = np.nanmean(models[:,:,:,0:2,:,:],axis=3)
eraq = np.nanmean(era[:,0:2,:,:],axis=1)
elif period == 'ON':
modq = np.nanmean(models[:,:,:,9:11,:,:],axis=3)
eraq = np.nanmean(era[:,9:11,:,:],axis=1)
elif period == 'OND':
modq = np.nanmean(models[:,:,:,-3:,:,:],axis=3)
eraq = np.nanmean(era[:,-3:,:,:],axis=1)
elif period == 'S':
modq = models[:,:,:,-4,:,:].squeeze()
eraq = era[:,-4,:,:].squeeze()
elif period == 'O':
modq = models[:,:,:,-3,:,:].squeeze()
eraq = era[:,-3,:,:].squeeze()
elif period == 'N':
modq = models[:,:,:,-2,:,:].squeeze()
eraq = era[:,-2,:,:].squeeze()
elif period == 'D':
modq = models[:,:,:,-1:,:,:].squeeze()
eraq = era[:,-1:,:,:].squeeze()
elif period == 'ND':
modq = np.nanmean(models[:,:,:,-2:,:,:],axis=3)
eraq = np.nanmean(era[:,-2:,:,:],axis=1)
elif period == 'FM':
modq = np.nanmean(models[:,:,:,1:3,:,:],axis=3)
eraq = np.nanmean(era[:,1:3,:,:],axis=1)
elif period == 'JJA':
modq = np.nanmean(models[:,:,:,5:8,:,:],axis=3)
eraq = np.nanmean(era[:,5:8,:,:],axis=1)
elif period == 'AMJ':
modq = np.nanmean(models[:,:,:,3:6,:,:],axis=3)
eraq = np.nanmean(era[:,3:6,:,:],axis=1)
elif period == 'Annual':
modq = np.nanmean(models[:,:,:,:,:,:],axis=3)
eraq =
|
np.nanmean(era[:,:,:,:],axis=1)
|
numpy.nanmean
|
import numpy as np
import time
import copy
import math
import logging
logging.basicConfig(
level=logging.INFO,
format="%(asctime)s [%(levelname)s] %(message)s",
handlers=[
logging.FileHandler("debug.log"),
logging.StreamHandler()
]
)
from pynumdiff.utils import utility as utility
try:
import cvxpy
except:
logging.info('Import Error.\nCould not import cvxpy.\nTo use convex total variation regularized derivatives, install cvxpy (http://www.cvxpy.org/install/index.html)\n\
Recommended solver: MOSEK, free academic license available: https://www.mosek.com/products/academic-licenses/\nDespite this error, you can still use the iterative TVR method.\n')
from pynumdiff.total_variation_regularization import __chartrand_tvregdiff__ as __chartrand_tvregdiff__
import pynumdiff.smooth_finite_difference
from pynumdiff.utils import utility as utility
__gaussian_kernel__ = utility.__gaussian_kernel__
# Iterative total variation regularization
def iterative_velocity(x, dt, params, options={'cg_maxiter': 1000, 'scale': 'small'}):
'''
Use an iterative solver to find the total variation regularized 1st derivative.
See __chartrand_tvregdiff__.py for details, author info, and license
Methods described in: <NAME>, "Numerical differentiation of noisy, nonsmooth data,"
ISRN Applied Mathematics, Vol. 2011, Article ID 164564, 2011.
Original code (MATLAB and python): https://sites.google.com/site/dnartrahckcir/home/tvdiff-code
Inputs
------
x : (np.array of floats, 1xN) time series to differentiate
dt : (float) time step
Parameters
----------
params : (list) [iterations, (int) : Number of iterations to run the solver.
More iterations results in blockier derivatives,
which approach the convex result
gamma], (float): Regularization parameter. Larger values result
in more regularization / smoothing.
options : (dict) {'cg_maxiter': None, (int) : Max number of iterations to use in
scipy.sparse.linalg.cg
Default, None, results in maxiter = len(x)
This works well in our test examples.
'scale': 'small'} (str) : This method has two different numerical options.
From __chartrand_tvregdiff__.py:
'large' or 'small' (case insensitive). Default is
'small'. 'small' has somewhat better boundary
behavior, but becomes unwieldly for data larger than
1000 entries or so. 'large' has simpler numerics but
is more efficient for large-scale problems. 'large' is
more readily modified for higher-order derivatives,
since the implicit differentiation matrix is square.
Outputs
-------
x_hat : estimated (smoothed) x
dxdt_hat : estimated derivative of x
'''
iterations, gamma = params
dxdt_hat = __chartrand_tvregdiff__.TVRegDiff(x, iterations, gamma, dx=dt,
maxit=options['cg_maxiter'], scale=options['scale'],
ep=1e-6, u0=None, plotflag=False, diagflag=1)
x_hat = utility.integrate_dxdt_hat(dxdt_hat, dt)
x0 = utility.estimate_initial_condition(x, x_hat)
x_hat = x_hat + x0
return x_hat, dxdt_hat
# Generalized total variation regularized derivatives
def __total_variation_regularized_derivative__(x, dt, N, gamma, solver='MOSEK'):
'''
Use convex optimization (cvxpy) to solve for the Nth total variation regularized derivative.
Default solver is MOSEK: https://www.mosek.com/
Inputs
------
x : (np.array of floats, 1xN) time series to differentiate
dt : (float) time step
Parameters
----------
N : (int) 1, 2, or 3, the Nth derivative to regularize
gamma : (float) regularization parameter
solver : (string) Solver to use. Solver options include: 'MOSEK' and 'CVXOPT',
in testing, 'MOSEK' was the most robust.
Outputs
-------
x_hat : estimated (smoothed) x
dxdt_hat : estimated derivative of x
'''
# Normalize
mean = np.mean(x)
std = np.std(x)
x = (x-mean)/std
# Define the variables for the highest order derivative and the integration constants
var = cvxpy.Variable( len(x)+N )
# Recursively integrate the highest order derivative to get back to the position
derivatives = [var[N:]]
for i in range(N):
d = cvxpy.cumsum(derivatives[-1]) + var[i]
derivatives.append(d)
# Compare the recursively integration position to the noisy position
sum_squared_error = cvxpy.sum_squares(derivatives[-1] - x)
# Total variation regularization on the highest order derivative
r = cvxpy.sum( gamma*cvxpy.tv(derivatives[0]) )
#r = gamma*cvxpy.sum_squares( derivatives[0] )
# Set up and solve the optimization problem
obj = cvxpy.Minimize(sum_squared_error + r)
prob = cvxpy.Problem(obj)
prob.solve(solver=solver)
# Recursively calculate the value of each derivative
final_derivative = var.value[N:]
derivative_values = [final_derivative]
for i in range(N):
d = np.cumsum(derivative_values[-1]) + var.value[i]
derivative_values.append(d)
for i in range(len(derivative_values)):
derivative_values[i] = derivative_values[i]/(dt**(N-i))
# Extract the velocity and smoothed position
dxdt_hat = derivative_values[-2]
x_hat = derivative_values[-1]
dxdt_hat = (dxdt_hat[0:-1] + dxdt_hat[1:])/2
ddxdt_hat_f = dxdt_hat[-1] - dxdt_hat[-2]
dxdt_hat_f = dxdt_hat[-1] + ddxdt_hat_f
dxdt_hat = np.hstack((dxdt_hat, dxdt_hat_f))
# fix first point
d = dxdt_hat[2] - dxdt_hat[1]
dxdt_hat[0] = dxdt_hat[1] - d
return x_hat*std+mean, dxdt_hat*std
def velocity(x, dt, params, options={'solver': 'MOSEK'}):
'''
Use convex optimization (cvxpy) to solve for the velocity total variation regularized derivative.
Default solver is MOSEK: https://www.mosek.com/
Inputs
------
x : (np.array of floats, 1xN) time series to differentiate
dt : (float) time step
Parameters
----------
params : (list) [gamma], where gamma (float) is the regularization parameter
or if 'iterate' in options: [gamma, num_iterations]
options : (dict) {'solver': SOLVER} SOLVER options include: 'MOSEK' and 'CVXOPT',
in testing, 'MOSEK' was the most robust.
Outputs
-------
x_hat : estimated (smoothed) x
dxdt_hat : estimated derivative of x
'''
if type(params) is list:
gamma = params[0]
else:
gamma = params
return __total_variation_regularized_derivative__(x, dt, 1, gamma, solver=options['solver'])
def acceleration(x, dt, params, options={'solver': 'MOSEK'}):
'''
Use convex optimization (cvxpy) to solve for the acceleration total variation regularized derivative.
Default solver is MOSEK: https://www.mosek.com/
Inputs
------
x : (np.array of floats, 1xN) time series to differentiate
dt : (float) time step
Parameters
----------
params : (list) [gamma], where gamma (float) is the regularization parameter
or if 'iterate' in options: [gamma, num_iterations]
options : (dict) {'solver': SOLVER} SOLVER options include: 'MOSEK' and 'CVXOPT',
in testing, 'MOSEK' was the most robust.
Outputs
-------
x_hat : estimated (smoothed) x
dxdt_hat : estimated derivative of x
'''
if type(params) is list:
gamma = params[0]
else:
gamma = params
return __total_variation_regularized_derivative__(x, dt, 2, gamma, solver=options['solver'])
def jerk(x, dt, params, options={'solver': 'MOSEK'}):
'''
Use convex optimization (cvxpy) to solve for the jerk total variation regularized derivative.
Default solver is MOSEK: https://www.mosek.com/
Inputs
------
x : (np.array of floats, 1xN) time series to differentiate
dt : (float) time step
Parameters
----------
params : (list) [gamma], where gamma (float) is the regularization parameter
or if 'iterate' in options: [gamma, num_iterations]
options : (dict) {'solver': SOLVER} SOLVER options include: 'MOSEK' and 'CVXOPT',
in testing, 'MOSEK' was the most robust.
Outputs
-------
x_hat : estimated (smoothed) x
dxdt_hat : estimated derivative of x
'''
if type(params) is list:
gamma = params[0]
else:
gamma = params
return __total_variation_regularized_derivative__(x, dt, 3, gamma, solver=options['solver'])
def smooth_acceleration(x, dt, params, options={'solver': 'MOSEK'}):
gamma, window_size = params
x_hat, dxdt_hat = acceleration(x, dt, [gamma], options=options)
kernel = __gaussian_kernel__(window_size)
dxdt_hat = pynumdiff.smooth_finite_difference.__convolutional_smoother__(dxdt_hat, kernel, 1)
x_hat = utility.integrate_dxdt_hat(dxdt_hat, dt)
x0 = utility.estimate_initial_condition(x, x_hat)
x_hat = x_hat + x0
return x_hat, dxdt_hat
def jerk_sliding(x, dt, params, options={'solver': 'MOSEK'}):
'''
Use convex optimization (cvxpy) to solve for the jerk total variation regularized derivative.
Default solver is MOSEK: https://www.mosek.com/
Inputs
------
x : (np.array of floats, 1xN) time series to differentiate
dt : (float) time step
Parameters
----------
params : (list) [gamma], where gamma (float) is the regularization parameter
or if 'iterate' in options: [gamma, num_iterations]
options : (dict) {'solver': SOLVER} SOLVER options include: 'MOSEK' and 'CVXOPT',
in testing, 'MOSEK' was the most robust.
Outputs
-------
x_hat : estimated (smoothed) x
dxdt_hat : estimated derivative of x
'''
if type(params) is list:
gamma = params[0]
else:
gamma = params
window_size = 1000
stride = 200
if len(x) < window_size:
return jerk(x, dt, params, options=options)
# slide the jerk
final_xsmooth = []
final_xdot_hat = []
first_idx = 0
final_idx = first_idx + window_size
last_loop = False
final_weighting = []
try:
while not last_loop:
xsmooth, xdot_hat = __total_variation_regularized_derivative__(x[first_idx:final_idx], dt, 3, gamma, solver=options['solver'])
xsmooth = np.hstack(([0]*first_idx, xsmooth, [0]*(len(x)-final_idx) ))
final_xsmooth.append(xsmooth)
xdot_hat = np.hstack(([0]*first_idx, xdot_hat, [0]*(len(x)-final_idx) ))
final_xdot_hat.append(xdot_hat)
# blending
w = np.hstack(( [0]*first_idx,
|
np.arange(1, 201)
|
numpy.arange
|
""" Curve classes: Continuation, EquilibriumCurve, FoldCurve, HopfCurveOne, HopfCurveTwo
<NAME>, 2006;
Last edited: December 2012
Continuation is the ancestral class of all curve classes and contains the continuation algorithms
(Moore-Penrose, etc.) It also contains all methods that are general to any curve found using
continuation.
TO DO/Notes:
* Why are there two BranchPointFold in BifPoint.py?
* Highlight not working in plot_cycles
* Branch point curve
* Phase plane stuff! (see PyCont_PredPreyI-III examples in Phage project)
* Symbolic jacobians not working!!! (see PyCont_PredPreyI.py)
* Modify bifurcation point locators to handle nonzero parts; check MATCONT again
* LPC detection problem in PyCont_Logistic.py
* Branch points on fold curve problem (PyCont_PredPrey.py)
* Need to check update in BorderMethod. Not currently working. Sticking with random for now.
* Implement pseudo-Newton method for branch point locator (BranchPoint.locate)
* Need to revisit alternate branch locator in BranchPoint.process
* Allow user to toggle update in BorderMethods (see BranchFold.py for example) -- how to make matrix as well-conditioned as possible?
(In AddTestFunction: Changed to "self.testfunc = TF_type(self.sysfunc, C, update=False)
* Removed BP in Continuation class
* Implement branch points for FixedPointCuspCurve [Networks/Global/dat/PyCont_Oscillator.py]
* FixedPointCuspCurve - Merge AddTestFunction_FixedPoint_Mult and AddTestFunction_FixedPoint
* FixedPointCuspCurve - Merge CP_Fold and CP_Fold2
* Create NSCurve (PyCont_DiscPredPrey2.py) [using NS_Bor]
* Allow for xdomain (see PyCont_DiscPredPrey2.py)
* Labels plot off screen when xlim or ylim
* Add cleanLabels to all children classes? (e.g., in FixedPointNSCurve)
* Rename FoldCurve to LimitPointCurve
* Allow for PCargs to include different parameters (e.g. initpars)
"""
# -----------------------------------------------------------------------------------------
from __future__ import absolute_import, print_function
from .misc import *
from .TestFunc import *
from .BifPoint import *
from .Plotting import *
from PyDSTool import Point, Pointset, PointInfo, args
from PyDSTool.common import pickle, sortedDictValues, sortedDictKeys
from PyDSTool.errors import *
from PyDSTool.Symbolic import QuantSpec
try:
from PyDSTool.matplotlib_import import *
except ImportError:
from PyDSTool.matplotlib_unavailable import *
print("Warning: matplotlib failed to import properly and so is not")
print(" providing a graphing interface")
from numpy.random import random
from numpy import dot as matrixmultiply
from scipy import optimize, linalg
from numpy import array, float, complex, int, float64, complex64, int32, \
zeros, divide, subtract, arange, all, any, argsort, reshape, nonzero, \
log10, Inf, NaN, isfinite, r_, c_, sign, mod, mat, log2, \
subtract, divide, transpose, eye, real, imag, isnan, resize
from numpy.linalg import cond # not present in scipy.linalg!
from copy import copy, deepcopy
from math import ceil
#####
_classes = ['Continuation', 'EquilibriumCurve', 'FoldCurve', 'HopfCurveOne',
'HopfCurveTwo', 'FixedPointCurve', 'LimitCycleCurve',
'UserDefinedCurve', 'FixedPointFoldCurve', 'FixedPointFlipCurve',
'FixedPointNSCurve', 'FixedPointCuspCurve']
_constants = ['cont_args_list', 'cont_bif_points', 'equilibrium_args_list',
'equilibrium_bif_points', 'fold_args_list', 'fold_bif_points',
'hopf_args_list', 'hopf_bif_points', 'limitcycle_args_list',
'limitcycle_bif_points', 'fixedpoint_args_list',
'fixedpoint_bif_points', 'flip_args_list', 'flip_bif_points',
'NS_args_list', 'NS_bif_points', 'userdefined_args_list',
'all_args_list', 'all_point_types', 'all_curve_types',
'bif_curve_colors', 'bif_point_colors',
'stab_line_styles','auto_point_types', 'other_special_points',
'solution_measures', 'solution_measures_list']
__all__ = _classes + _constants
#####
cont_args_list = ['name','force','freepars','MaxNumPoints','MaxCorrIters',
'MaxTestIters','MaxStepSize', 'MinStepSize', 'StepSize',
'VarTol','FuncTol','TestTol', 'description', 'uservars',
'LocBifPoints','verbosity','ClosedCurve','SaveJacobian',
'SaveEigen', 'Corrector', 'UseAuto', 'StopAtPoints',
'SPOut']
cont_bif_points = ['B', 'SP']
equilibrium_args_list = ['LocBifPoints']
equilibrium_bif_points = ['BP', 'LP', 'H']
fold_args_list = ['LocBifPoints']
fold_bif_points = ['BT', 'ZH', 'CP']
#fold_bif_points = ['BT', 'ZH', 'CP', 'BP'] # Disabling BP for now.
hopf_args_list = ['LocBifPoints']
hopf_bif_points = ['BT', 'ZH', 'GH', 'DH']
fixedpoint_args_list = ['LocBifPoints', 'period']
fixedpoint_bif_points = ['BP', 'PD', 'LPC', 'NS']
fold_map_args_list = ['LocBifPoints', 'period']
fold_map_bif_points = ['CP']
flip_args_list = ['LocBifPoints', 'period']
flip_bif_points = []
NS_args_list = ['LocBifPoints', 'period']
NS_bif_points = []
cusp_args_list = ['LocBifPoints', 'period']
cusp_bif_points = ['']
limitcycle_args_list = ['LocBifPoints', 'NumCollocation', 'NumIntervals',
'AdaptMesh', 'NumSPOut', 'DiagVerbosity',
'SolutionMeasures', 'SaveFlow']
limitcycle_bif_points = ['PD', 'LPC', 'NS']
userdefined_args_list = ['LocBifPoints']
other_special_points = ['RG', 'UZ', 'P', 'MX', 'B']
auto_point_types = {1: 'BP', 2: 'LP', 3: 'H', 4: 'RG', -4: 'UZ', 5: 'LPC',
6: 'BP', 7: 'PD', 8: 'NS', 9: 'P', -9: 'MX'}
solution_measures_list = ['max', 'min', 'avg', 'nm2'] # Ordering is important
solution_measures = dict(zip(solution_measures_list,[0, 0, 1, 2]))
all_args_list = unique(cont_args_list + equilibrium_args_list + fold_args_list +
hopf_args_list + fixedpoint_args_list + flip_args_list +
NS_args_list + limitcycle_args_list)
all_point_types = unique(other_special_points + cont_bif_points +
equilibrium_bif_points + fold_bif_points +
hopf_bif_points + fixedpoint_bif_points +
flip_bif_points + NS_bif_points +
limitcycle_bif_points)
all_curve_types = ['EP-C', 'LP-C', 'H-C1', 'H-C2', 'FP-C', 'LC-C', 'FD-C',
'FL-C', 'NS-C', 'CP-C']
bif_curve_colors = {'EP-C': 'k', 'LP-C': 'r', 'H-C1': 'b', 'H-C2': 'b',
'FP-C': 'k', 'LC-C': 'm', 'UD-C': 'k', 'FD-C': 'r',
'FL-C': 'g', 'NS-C': 'b', 'CP-C': 'c'}
bif_point_colors = {'P': 'ok', 'RG': 'ok', 'LP': 'or', 'BP': 'og',
'H': 'ob', 'B': 'dr', 'BT': 'sy', 'ZH': 'sk',
'CP': 'sr', 'GH': 'sb', 'DH': 'sg', 'LPC': 'Dr',
'PD': 'Dg', 'NS': 'Db', 'MX': 'xr', 'UZ': '^r',
'SP': '*b'}
stab_line_styles = {'S': '-', 'U': '--', 'N': '-.', 'X': ':'}
class Continuation(object):
"""Abstract continuation class
Children: EquilibriumCurve, FoldCurve, HopfCurveOne, HopfCurveTwo,
LimitCycleCurve
"""
def __init__(self, model, gen, automod, plot, args=None):
self.curvetype = args['type']
self._ptlabel = self.curvetype.split('-')[0]
self.model = model
self.gensys = gen
self._autoMod = automod
self.UseAuto = False
if 'description' not in args:
self.description = 'None'
else:
self.description = args['description']
if not hasattr(self, 'parsdict'):
self.parsdict = self.model.query('pars')
self.freepars = args['freepars']
self.auxpars = args['auxpars']
if hasattr(self, 'varslist'):
# varsindices refers to the indices in the full set of variables
# that are used in this subset
self.varslist.sort()
if self.curvetype == 'UD-C':
# unused, self._systemuser -> user-supplied func will be used directly
self.varsindices = array([])
else:
orig_vars = self.model.query('vars')
# will call self._system, selecting vars from possible ones
self.varsindices = array([orig_vars.index(v) for v in self.varslist])
else:
if 'uservars' in args and self.curvetype != 'UD-C':
self.varslist = args['uservars']
orig_vars = self.model.query('vars')
# will call self._system, selecting vars from possible ones
self.varsindices = array([orig_vars.index(v) for v in self.varslist])
else:
self.varslist = self.model.query('vars')
self.varsindices = arange(len(self.varslist))
if self.gensys.haveJacobian_pars():
fargs, fspecstr = self.gensys.funcspec._auxfnspecs['Jacobian_pars']
Jquant = QuantSpec('J', fspecstr)
if Jquant.dim == 0:
# dim of vars == 1 == dim of pars
assert len(self.varslist) == 1
assert len(self.freepars) == 1
# Supplied Jac w.r.t. params is custom-made for only the free params in this continuation
# (or there's only one parameter in system)
self.parsindices = array([0])
else:
assert len(self.varslist) == Jquant.dim
Jquant0 = Jquant.fromvector(0)
if Jquant0.dim == 0:
# dim of free pars == 1
assert len(self.freepars) == 1
# Supplied Jac w.r.t. params is custom-made for only the free params in this continuation
# (or there's only one parameter in system)
self.parsindices = array([0])
else:
if len(self.freepars) == Jquant0.dim:
# Supplied Jac w.r.t. params is custom-made for only the free params in this continuation
self.parsindices = arange(range(Jquant0.dim))
else:
# Assume supplied Jac w.r.t. params is for all params in the original system
# therefore there should be fewer free params than # system parameters
assert len(self.freepars) < Jquant0.dim
self.parsindices = array([list(self.parsdict.keys()).index(p) for p in self.freepars])
else:
self.parsindices = array([list(self.parsdict.keys()).index(p) for p in self.freepars])
self.varsdim = len(self.varslist)
self.freeparsdim = len(self.freepars)
self.auxparsdim = len(self.auxpars)
self.dim = self.varsdim + self.freeparsdim + self.auxparsdim
if (self.curvetype != 'UD-C'):
self.sysfunc = Function((self.dim, self.varsdim), self._system)
else:
self.sysfunc = Function((self.dim, self.varsdim), self._systemuser)
if (self.curvetype != 'UD-C' and self.gensys.haveJacobian()):
if self.gensys.haveJacobian_pars():
self.sysfunc.jac = Function((self.sysfunc.n,
(self.sysfunc.m,self.sysfunc.n)),
self._systemjac_withpars)
else:
self.sysfunc.jac = Function((self.sysfunc.n,
(self.sysfunc.m,self.sysfunc.n)),
self._systemjac)
elif (self.curvetype == 'UD-C' and hasattr(self, '_userjac')):
self.sysfunc.jac = Function((self.sysfunc.n,
(self.sysfunc.m,self.sysfunc.n)),
self._systemjacuser)
else:
self.sysfunc.jac = Function((self.sysfunc.n,
(self.sysfunc.m,self.sysfunc.n)),
self.sysfunc.diff)
self.coords = self.sysfunc.coords = arange(self.varsdim).tolist()
self.params = self.sysfunc.params = (arange(self.freeparsdim \
+ self.auxparsdim) \
+ self.varsdim).tolist()
self.allvars = self.sysfunc.allvars = self.coords + self.params
# Initialize vars and pars based on initpoint
self.initpoint = self.model.query('ics')
for k, v in args['initpoint'].items():
if k in self.varslist or k in args['auxpars']:
self.initpoint[k] = v
elif k in self.model.query('pars'):
self.parsdict[k] = v
for p in args['freepars']:
self.initpoint[p] = self.parsdict[p]
self.initpoint = tocoords(self, self.initpoint.copy())
if 'initdirec' not in args:
self.initdirec = None
else:
self.initdirec = tocoords(self, args['initdirec'])
if 'initcycle' not in args:
self.initcycle = None
else:
self.initcycle = args['initcycle']
if not hasattr(self, "SPOut"):
self.SPOut = None
if not hasattr(self, "NumSPOut"):
self.NumSPOut = 300
self.preTF = None
self.reset()
# Removes extra parameters (first time parameter initpoint, system
# parameter auxpars, and uneditable parameter type) before sending
# to update() method
args = dict(args)
[args.pop(i) for i in ['initpoint','initdirec','initcycle','auxpars',
'type'] if i in args]
self.update(args)
self.fig = None
self.text_handles = []
self.plot = plot
self._statuscodes = {0: 'Unrecognized error encountered (check stderr output). Stopping continuation...',
-1: 'Do over.'}
def __copy__(self):
pickledself = pickle.dumps(self)
return pickle.loads(pickledself)
def __deepcopy__(self, memo=None, _nil=[]):
pickledself = pickle.dumps(self)
return pickle.loads(pickledself)
def reset(self, args=None):
"""Resets curve by setting default parameters and deleting solution curve."""
self.MaxNumPoints = 300
self.MaxCorrIters = 5
self.MaxTestIters = 10
self.MaxStepSize = 0.1
self.MinStepSize = 1e-5
self.StepSize = 0.01
self.VarTol = self.FuncTol = 1e-6
self.TestTol = 1e-4
self.ClosedCurve = 50
self.verbosity = 1
self.SPOut = None
self.NumSPOut = 300
self.sol = None
# record of newly computed solution segment by
# forward or backward methods
self.new_sol_segment = None
self.LocBifPoints = []
self.StopAtPoints = []
self.TestFuncs = None
self.BifPoints = {}
self.CurveInfo = PointInfo()
self.SaveJacobian = False
self.SaveEigen = False
self.Corrector = self._MoorePenrose
if args is not None:
self.update(args)
def update(self, args):
"""Update parameters for Continuation."""
if args is not None:
for k, v in args.items():
if k in cont_args_list:
if k == 'LocBifPoints':
if isinstance(v, str):
if v.lower() == 'all':
v = cont_bif_points
else:
v = [v]
# Handle stopping points
w = []
if 'StopAtPoints' in args:
w = args['StopAtPoints']
if isinstance(w, str):
if w.lower() == 'all':
w = cont_bif_points
else:
w = [w]
self.LocBifPoints = [bftype for bftype in v \
if bftype in cont_bif_points]
self.StopAtPoints = [bftype for bftype in w \
if bftype in cont_bif_points]
elif k == 'Corrector':
self.Corrector = getattr(self, '_' + v)
elif k != 'StopAtPoints':
exec('self.' + k + ' = ' + repr(v))
elif k not in all_args_list:
print("Warning: " + k + " is either not a valid parameter or immutable.")
def _preTestFunc(self, X, V):
J = self.sysfunc.jac(X)
self.sysfunc.J_coords = J[:,self.coords[0]:(self.coords[-1]+1)]
self.sysfunc.J_params = J[:,self.params[0]:(self.params[-1]+1)]
if self.preTF is not None:
self.preTF(X, V)
def _createTestFuncs(self):
"""Creates processors and test functions for Continuation class.
Note: In the following list, processors are in PyCont.Bifpoint
and test functions are in PyCont.TestFunc.
Point type (Processor): Test Function(s)
----------------------------------------
BP (BranchPoint): Branch_Det
"""
self.TestFuncs = []
self.BifPoints = {}
for bftype in self.LocBifPoints:
if bftype in cont_bif_points:
stop = bftype in self.StopAtPoints # Set stopping flag
#if bftype is 'BP':
#method = Branch_Det(self.CorrFunc, self, save=True,
#numpoints=self.MaxNumPoints+1)
#self.TestFuncs.append(method)
#self.BifPoints['BP'] = BranchPoint(method, iszero,
#stop=stop)
if bftype is 'B':
method = B_Check(self.CorrFunc, self, save=True,
numpoints=self.MaxNumPoints+1)
self.TestFuncs.append(method)
self.BifPoints['B'] = BPoint(method, iszero, stop=stop)
if self.SPOut is not None:
# add simple "user"-defined function to catch parameter values
# during continuation
for par, par_vals in self.SPOut.items():
try:
par_ix = self.params[self.freepars.index(par)]
except IndexError:
raise ValueError("Invalid free parameter %s" % par)
for i, pval in enumerate(par_vals):
method = ParTestFunc(self.sysfunc.n,
self, par_ix, pval, save=True,
numpoints=self.NumSPOut+1)
self.TestFuncs.append(method)
self.BifPoints['SP-%s-%i' % (par, i)] = \
SPoint(method, iszero, stop=False)
def _system(self, X):
VARS = dict(zip(self.varslist, array(X)[self.coords]))
for i, par in enumerate(self.freepars):
self.parsdict[par] = X[self.params[i]]
try:
t = self.parsdict['time']
except KeyError:
# autonomous system, t doesn't matter
t = 0
return self.gensys.Rhs(t, VARS, self.parsdict, asarray=True)[self.varsindices]
def _systemjac(self, x0, ind=None):
VARS = dict(zip(self.varslist, array(x0)[self.coords]))
for i, par in enumerate(self.freepars):
self.parsdict[par] = x0[self.params[i]]
try:
t = self.parsdict['time']
except KeyError:
# autonomous system, t doesn't matter
t = 0
jacx = self.gensys.Jacobian(t, VARS, self.parsdict, asarray=True)[self.varsindices]
jacp = self.sysfunc.diff(x0, ind=self.params)
try:
return c_[jacx, jacp][:,ind[0]:(ind[-1]+1)]
except:
return c_[jacx, jacp]
def _systemjac_withpars(self, x0, ind=None):
VARS = dict(zip(self.varslist, array(x0)[self.coords]))
for i, par in enumerate(self.freepars):
self.parsdict[par] = x0[self.params[i]]
try:
t = self.parsdict['time']
except KeyError:
# autonomous system, t doesn't matter
t = 0
jacx = self.gensys.Jacobian(t, VARS, self.parsdict, asarray=True)[self.varsindices]
jacp = self.gensys.JacobianP(t, VARS, self.parsdict, asarray=True)[self.parsindices]
try:
return c_[jacx, jacp][:,ind[0]:(ind[-1]+1)]
except:
return c_[jacx, jacp]
def _systemuser(self, X):
"""Calls self._userfunc, which is assumed to return an array of RHS
values for the relevant (possibly subset of) variables."""
VARS = dict(zip(self.varslist, array(X)[self.coords]))
for i, par in enumerate(self.freepars):
self.parsdict[par] = X[self.params[i]]
return self._userfunc(self, VARS, self.parsdict)
def _systemjacuser(self, x0, ind=None):
"""Calls self._userjac, which is assumed to return an array of
[Jac_x, Jac_p]."""
VARS = dict(zip(self.varslist, array(X)[self.coords]))
for i, par in enumerate(self.freepars):
self.parsdict[par] = X[self.params[i]]
return self._userjac(self, VARS, self.parsdict)
def _checkForBifPoints(self):
# increase efficiency by preventing many self. references
loc = self.loc
# these declarations just make references
curve = self.curve
V = self.V
# store commonly referenced values for efficiency
V_loc = V[loc]
curve_loc = curve[loc]
for bftype, bfinfo in self.BifPoints.items():
bftype = bftype.split('-')[0]
flag_list = []
for i, testfunc in enumerate(bfinfo.testfuncs):
for k in range(testfunc.m):
flag_list.append(bfinfo.flagfuncs[i](testfunc[loc-1][k],
testfunc[loc][k]))
# if bftype == 'NS':
# print loc, bftype, flag_list, testfunc[loc] # DREW WUZ HERE 2012
bfpoint_found = all(flag_list)
if bfpoint_found:
# Locate bifurcation point
Xval, Vval = bfinfo.locate((curve[loc-1], V[loc-1]),
(curve_loc, V_loc), self)
found = bfinfo.process(Xval, Vval, self)
if found:
# Move information one more step forward
if not bfinfo.stop:
curve[loc+1] = curve_loc
V[loc+1] = V_loc
for testfunc in self.TestFuncs:
testfunc[loc+1] = testfunc[loc]
else:
startx = copy(curve_loc)
startv = copy(V_loc)
curve[loc] = Xval
V[loc] = Vval
self._savePointInfo(loc)
self.CurveInfo[loc] = (bftype,
{'data': bfinfo.found[-1],
'plot': args()})
if not bfinfo.stop:
self.loc += 1
loc += 1 # update in sync with self.loc
V_loc = V[loc]
curve_loc = curve[loc]
else:
self.CurveInfo[loc] = ('P',
{'data': args(V = todict(self, startv)),
'startx': todict(self, startx),
'plot': args()})
return True
# Do not stop computations
return False
def exportGeomview(self, coords=None, filename="geom.dat"):
if coords is not None and len(coords) == 3:
GeomviewOutput = "(progn (geometry " + self.model.name + \
" { LIST {: axes_" + self.model.name + "}"
# for cname, curve in self.curves.iteritems():
GeomviewOutput += " {: " + self.name + "}"
GeomviewOutput += "}))\n\n"
# Get axes limits
alim = [[Inf,-Inf],[Inf,-Inf],[Inf,-Inf]]
# for cname, curve in self.curves.iteritems():
for n in range(len(coords)):
alim[n][0] = min(alim[n][0], min(self.sol[coords[n]]))
alim[n][1] = max(alim[n][1], max(self.sol[coords[n]]))
GeomviewOutput += "(progn (hdefine geometry axes_" + \
self.model.name + " { appearance { linewidth 2 } SKEL 4 3 " +\
"0 0 0 1 0 0 0 1 0 0 0 1 " + \
"2 0 1 1 0 0 1 2 0 2 0 1 0 1 2 0 3 0 0 1 1})\n\n"
#for cname, curve in self.curves.iteritems():
cname = self.name
GeomviewOutput += "(hdefine geometry " + cname + \
" { LIST {: curve_" + cname + "} {: specpts_" + cname + "}})\n\n"
GeomviewOutput += "(hdefine geometry curve_" + cname + \
" { appearance { linewidth 2 } SKEL " + \
repr(len(self.sol)) + " " + repr(len(self.sol)-1)
for n in range(len(self.sol)):
GeomviewOutput += " " + repr((self.sol[n][coords[0]]-alim[0][0])/(alim[0][1]-alim[0][0])) + \
" " + repr((self.sol[n][coords[1]]-alim[1][0])/(alim[1][1]-alim[1][0])) + \
" " + repr((self.sol[n][coords[2]]-alim[2][0])/(alim[2][1]-alim[2][0]))
for n in range(len(self.sol)-1):
GeomviewOutput += " 2 " + repr(n) + " " + repr(n+1) + " 0 0 0 1"
GeomviewOutput += "})\n\n"
GeomviewOutput += ")\n"
f = open(filename, "w")
f.write(GeomviewOutput)
f.close()
else:
raise PyDSTool_ValueError("Coordinates not specified or not of correct dimension.")
def display(self, coords=None, dirs=None, origin=None, figure=None,
axes=None, stability=False, domain=False, init_display=True,
points=True, **plot_args):
"""Plot curve in coordinates specified by coords.
Inputs:
coords -- pair of coordinates (None defaults to the first free
parameter and the first state variable)
Use a 3-tuple to export to geomview.
dirs -- tuple of coordinate directions IF coord is not in regular coords
origin -- Useful if want affine coordinates
"""
# Take care of calling with state variable w/o max/min for LC
disp_args = copy(plot_args)
if self.sol is not None:
if coords is None:
coords = [self.freepars[0], self.varslist[0]]
if self.curvetype == 'LC-C':
coords = list(coords)
for n in range(2):
if coords[n] in self.varslist:
# Default to max of solution
coords[n] = coords[n]+'_max'
if len(coords) == 3:
self.exportGeomview(coords=coords)
return
if origin is not None:
clist = self.sol.coordnames
clen = len(clist)
aorigin = array([origin[k] for k in clist])
X = zeros((2,len(self.sol)), float)
for n in range(2):
if coords[n] in self.sol.coordnames:
X[n] = self.sol[coords[n]]
if origin is not None:
X[n] = X[n] - origin[coords[n]]
elif coords[n] in self.parsdict.keys():
X[n] = array([self.parsdict[coords[n]]]*len(self.sol))
if origin is not None:
X[n] = X[n] - origin[coords[n]]
elif dirs is not None and coords[n] in dirs.keys():
# Project curve onto plane spanned by coordinate directions
# spanning variables and free parameters
X[n] = array([matrixmultiply(x-aorigin, dirs[coords[n]]) \
for x in self.sol])
else:
raise KeyError('Coordinate ' + coords[n] + ' is not defined.')
if init_display:
initializeDisplay(self.plot, figure=figure, axes=axes)
cfl = self.plot._cfl
cal = self.plot._cal
## Prints curve
# Get unique name
name = self.name
if name in self.plot[cfl][cal]:
num = 0
for k, v in self.plot[cfl][cal].items():
if isinstance(v, pargs) and k.split('_')[0] == name:
num += 1
name = name + '_' + repr(num)
self.plot[cfl][cal][name] = pargs()
self.plot[cfl][cal][name].curve = []
label = self.curvetype.split('-')[0]
self.plot[cfl][cal][name].type = label
if stability and self.SaveEigen:
if 'linewidth' not in disp_args: # Default linewidth 1
disp_args['linewidth'] = 1
disp_args['label'] = '_nolegend_'
stabdict = partition([x.labels[label]['stab'] \
for x in self.sol],['S','U','N'])
for stabtype, stablist in stabdict.items():
for curve in stablist:
self.plot[cfl][cal][name].curve.extend(plt.plot(X[0][curve[0]:curve[1]], \
X[1][curve[0]:curve[1]], \
bif_curve_colors[self.curvetype]+stab_line_styles[stabtype], **disp_args))
else:
if 'label' not in disp_args:
disp_args['label'] = name
self.plot[cfl][cal][name].curve.extend(plt.plot(X[0], X[1], \
bif_curve_colors[self.curvetype], **disp_args))
# Take care of labels
xlab = coords[0]
ylab = coords[1]
if self.curvetype == 'LC-C':
for smtype in self.SolutionMeasures:
if xlab.rfind('_'+smtype) > 0:
xlab = xlab[0:xlab.rfind('_'+smtype)]
break
for smtype in self.SolutionMeasures:
if ylab.rfind('_'+smtype) > 0:
ylab = ylab[0:ylab.rfind('_'+smtype)]
break
plt.xlabel(xlab)
plt.ylabel(ylab)
# Prints special points
if points:
for bftype in all_point_types:
bflist = self.sol.bylabel(bftype)
if bflist is not None:
for point in bflist:
if 'name' in point.labels[bftype]:
X = zeros(2, float)
for n in range(2):
if coords[n] in self.sol.coordnames:
X[n] = point[coords[n]]
if origin is not None:
X[n] = X[n] - origin[coords[n]]
elif coords[n] in self.parsdict.keys():
X[n] = self.parsdict[coords[n]]
if origin is not None:
X[n] = X[n] - origin[coords[n]]
elif dirs is not None and coords[n] in dirs.keys():
# Project point onto plane spanned by coordinate directions
# spanning variables and free parameters
X[n] = matrixmultiply(point-aorigin,
dirs[coords[n]])
# Print point
ptname = point.labels[bftype]['name']
self.plot[cfl][cal][name][ptname] = pargs()
self.plot[cfl][cal][name][ptname].point = \
plt.plot([X[0]], [X[1]],
bif_point_colors[bftype],
label='_nolegend_')
# Print label
ha = 'left'
if self.curvetype in ['LP-C','H-C1','H-C2','LC-C']:
va = 'top'
else:
va = 'bottom'
self.plot[cfl][cal][name][ptname].text = \
plt.text(X[0], X[1], ' '+ ptname,
ha=ha, va=va)
def _savePointInfo(self, loc):
"""Created a function for this since it needs to be called
both in _compute and when a bifurcation point is found. It
will have conditional statements for saving of Jacobian and
eigenvalues, as well as other possible tidbits of
information."""
ptlabel = self._ptlabel
self.CurveInfo[loc] = (ptlabel, \
{'data': args(V = todict(self, self.V[loc]),
ds = self.StepSize)})
# Save domain information
if 'B' in self.LocBifPoints:
val = self.BifPoints['B'].testfuncs[0][loc][0]
# if val >= 0 set domain = 'inside' otherwise 'outside'
self.CurveInfo[loc][ptlabel]['domain'] = (val >= 0) \
and 'inside' or 'outside'
# Save eigenvalue information
if self.SaveEigen:
# May be able to use J_coords here
jac = self.sysfunc.jac(self.curve[loc])
jacx = jac[:,self.coords[0]:(self.coords[-1]+1)]
jacp = jac[:,self.params[0]:(self.params[-1]+1)]
w, vr = linalg.eig(jacx)
self.CurveInfo[loc][ptlabel]['data'].evals = w
self.CurveInfo[loc][ptlabel]['data'].evecs = vr
if ptlabel == 'FP':
inside = [abs(eig) < 1-1e-6 for eig in w]
outside = [abs(eig) > 1+1e-6 for eig in w]
if all(inside):
self.CurveInfo[loc][ptlabel]['stab'] = 'S'
elif
|
all(outside)
|
numpy.all
|
import numpy as np
class Softmax(object):
def __init__(self, dims=[10, 3073]):
self.init_weights(dims=dims)
def init_weights(self, dims):
"""
Initializes the weight matrix of the Softmax classifier.
Note that it has shape (C, D) where C is the number of
classes and D is the feature size.
"""
self.W = np.random.normal(size=dims) * 0.0001
def loss(self, X, y):
"""
Calculates the softmax loss.
Inputs have dimension D, there are C classes, and we operate on minibatches
of N examples.
Inputs:
- X: A numpy array of shape (N, D) containing a minibatch of data.
- y: A numpy array of shape (N,) containing training labels; y[i] = c means
that X[i] has label c, where 0 <= c < C.
Returns a tuple of:
- loss as single float
"""
# Initialize the loss to zero.
loss = 0.0
# ================================================================ #
# YOUR CODE HERE:
# Calculate the normalized softmax loss. Store it as the variable loss.
# (That is, calculate the sum of the losses of all the training
# set margins, and then normalize the loss by the number of
# training examples.)
# ================================================================ #
N = X.shape[0]
C = self.W.shape[0]
WTX = [email protected]
eWTX = np.exp(WTX)
for i in range(N):
loss = loss + np.log(np.ones((1,C))@eWTX[:,i]) - WTX[y[i],i]
loss = loss/N
# ================================================================ #
# END YOUR CODE HERE
# ================================================================ #
return loss
def loss_and_grad(self, X, y):
"""
Same as self.loss(X, y), except that it also returns the gradient.
Output: grad -- a matrix of the same dimensions as W containing
the gradient of the loss with respect to W.
"""
# Initialize the loss and gradient to zero.
loss = 0.0
grad = np.zeros_like(self.W)
# ================================================================ #
# YOUR CODE HERE:
# Calculate the softmax loss and the gradient. Store the gradient
# as the variable grad.
# ================================================================ #
N = X.shape[0]
C = self.W.shape[0]
WTX = [email protected]
eWTX = np.exp(WTX)
# Calculate Loss
for i in range(N):
loss = loss + np.log(np.ones((1,C))@eWTX[:,i]) - WTX[y[i],i]
loss = loss/N
# Calculate Gradient
for j in range(C):
for i in range(N):
grad[j,:] = grad[j,:] + ( (1/(np.ones((1,C))@eWTX[:,i]))*(eWTX[j,i]) - (y[i]==j) )*X[i,:].T
grad = grad/N
# ================================================================ #
# END YOUR CODE HERE
# ================================================================ #
return loss, grad
def grad_check_sparse(self, X, y, your_grad, num_checks=10, h=1e-5):
"""
sample a few random elements and only return numerical
in these dimensions.
"""
for i in np.arange(num_checks):
ix = tuple([np.random.randint(m) for m in self.W.shape])
oldval = self.W[ix]
self.W[ix] = oldval + h # increment by h
fxph = self.loss(X, y)
self.W[ix] = oldval - h # decrement by h
fxmh = self.loss(X,y) # evaluate f(x - h)
self.W[ix] = oldval # reset
grad_numerical = (fxph - fxmh) / (2 * h)
grad_analytic = your_grad[ix]
rel_error = abs(grad_numerical - grad_analytic) / (abs(grad_numerical) + abs(grad_analytic))
print('numerical: %f analytic: %f, relative error: %e' % (grad_numerical, grad_analytic, rel_error))
def fast_loss_and_grad(self, X, y):
"""
A vectorized implementation of loss_and_grad. It shares the same
inputs and ouptuts as loss_and_grad.
"""
loss = 0.0
grad = np.zeros(self.W.shape) # initialize the gradient as zero
# ================================================================ #
# YOUR CODE HERE:
# Calculate the softmax loss and gradient WITHOUT any for loops.
# ================================================================ #
N = X.shape[0]
C = self.W.shape[0]
WTX = [email protected]
eWTX = np.exp(WTX)
loss = np.ones((1,N))@(np.log([email protected]((C,))) - WTX[y,np.arange(N)])/N
scaling = eWTX/([email protected]((C,)))
scaling[y,np.arange(N)] -= 1
grad = scaling@X/N
# END YOUR CODE HERE
# ================================================================ #
return loss, grad
def train(self, X, y, learning_rate=1e-3, num_iters=100,
batch_size=200, verbose=False):
"""
Train this linear classifier using stochastic gradient descent.
Inputs:
- X: A numpy array of shape (N, D) containing training data; there are N
training samples each of dimension D.
- y: A numpy array of shape (N,) containing training labels; y[i] = c
means that X[i] has label 0 <= c < C for C classes.
- learning_rate: (float) learning rate for optimization.
- num_iters: (integer) number of steps to take when optimizing
- batch_size: (integer) number of training examples to use at each step.
- verbose: (boolean) If true, print progress during optimization.
Outputs:
A list containing the value of the loss function at each training iteration.
"""
num_train, dim = X.shape
num_classes =
|
np.max(y)
|
numpy.max
|
""" Functions that help with SKA simulations
"""
import logging
import astropy.units as units
import matplotlib.pyplot as plt
import numpy
from astropy.coordinates import SkyCoord
import astropy.constants as constants
from data_models.memory_data_models import Skycomponent, BlockVisibility
from data_models.polarisation import PolarisationFrame
from processing_library.image.operations import create_image
from processing_library.util.coordinate_support import hadec_to_azel
from wrappers.serial.image.operations import show_image
from wrappers.serial.imaging.primary_beams import create_pb
from wrappers.serial.skycomponent.base import copy_skycomponent
from wrappers.serial.skycomponent.operations import apply_beam_to_skycomponent
log = logging.getLogger(__name__)
def find_times_above_elevation_limit(start_times, end_times, location, phasecentre, elevation_limit):
""" Find all times for which a phasecentre is above the elevation limit
:param start_times:
:param end_times:
:param location:
:param phasecentre:
:param elevation_limit:
:return:
"""
assert len(start_times) == len(end_times)
def valid_elevation(time, location, phasecentre):
ha = numpy.pi * time / 43200.0
dec = phasecentre.dec.rad
az, el = hadec_to_azel(ha, dec, location.lat.rad)
return el > elevation_limit * numpy.pi / 180.0
number_valid_times = 0
valid_start_times = []
for it, t in enumerate(start_times):
if valid_elevation(start_times[it], location, phasecentre) or \
valid_elevation(end_times[it], location, phasecentre):
valid_start_times.append(t)
number_valid_times += 1
assert number_valid_times > 0, "No data above elevation limit"
log.info("find_times_above_elevation_limit: Start times for chunks above elevation limit:")
return valid_start_times
def plot_visibility(vis_list, ax=None, title='Visibility',
y='amp', x='uvdist', **kwargs):
""" Standard plot of visibility
:param vis_list:
:param plot_file:
:param kwargs:
:return:
"""
for ivis, vis in enumerate(vis_list):
if y == 'amp':
yvalue = numpy.abs(vis.vis[...,0,0].flat)
else:
yvalue = numpy.angle(vis.vis[...,0,0].flat)
xvalue = vis.uvdist.flat
ax.plot(xvalue[yvalue>0.0], yvalue[yvalue>0.0], '.', color='b', markersize=0.2)
ax.set_xlabel(x)
ax.set_ylabel(y)
ax.set_title(title)
def plot_uvcoverage(vis_list, ax=None, plot_file='uvcoverage.png', title='UV coverage', **kwargs):
""" Standard plot of uv coverage
:param vis_list:
:param plot_file:
:param kwargs:
:return:
"""
for ivis, vis in enumerate(vis_list):
u =
|
numpy.array(vis.u[...].flat)
|
numpy.array
|
''' Testing trackvis module '''
from StringIO import StringIO
import numpy as np
from .. import trackvis as tv
from nose.tools import assert_true, assert_false, assert_equal, assert_raises
from numpy.testing import assert_array_equal, assert_array_almost_equal
from ..testing import parametric
@parametric
def test_write():
streams = []
out_f = StringIO()
tv.write(out_f, [], {})
yield assert_equal(out_f.getvalue(), tv.empty_header().tostring())
out_f.truncate(0)
# Write something not-default
tv.write(out_f, [], {'id_string':'TRACKb'})
# read it back
out_f.seek(0)
streams, hdr = tv.read(out_f)
yield assert_equal(hdr['id_string'], 'TRACKb')
# check that we can pass none for the header
out_f.truncate(0)
tv.write(out_f, [])
out_f.truncate(0)
tv.write(out_f, [], None)
# check that we check input values
out_f.truncate(0)
yield assert_raises(tv.HeaderError,
tv.write, out_f, [],{'id_string':'not OK'})
yield assert_raises(tv.HeaderError,
tv.write, out_f, [],{'version': 3})
yield assert_raises(tv.HeaderError,
tv.write, out_f, [],{'hdr_size': 0})
def streams_equal(stream1, stream2):
if not np.all(stream1[0] == stream1[0]):
return False
if stream1[1] is None:
if not stream2[1] is None:
return false
if stream1[2] is None:
if not stream2[2] is None:
return false
if not np.all(stream1[1] == stream1[1]):
return False
if not np.all(stream1[2] == stream1[2]):
return False
return True
def streamlist_equal(streamlist1, streamlist2):
if len(streamlist1) != len(streamlist2):
return False
for s1, s2 in zip(streamlist1, streamlist2):
if not streams_equal(s1, s2):
return False
return True
def test_round_trip():
out_f = StringIO()
xyz0 = np.tile(np.arange(5).reshape(5,1), (1, 3))
xyz1 = np.tile(np.arange(5).reshape(5,1) + 10, (1, 3))
streams = [(xyz0, None, None), (xyz1, None, None)]
tv.write(out_f, streams, {})
out_f.seek(0)
streams2, hdr = tv.read(out_f)
assert_true(streamlist_equal(streams, streams2))
# test that we can get out and pass in generators
out_f.seek(0)
streams3, hdr = tv.read(out_f, as_generator=True)
# check this is a generator rather than a list
assert_true(hasattr(streams3, 'next'))
# but that it results in the same output
assert_true(streamlist_equal(streams, list(streams3)))
# write back in
out_f.seek(0)
streams3, hdr = tv.read(out_f, as_generator=True)
# Now we need a new file object, because we're still using the old one for
# our generator
out_f_write = StringIO()
tv.write(out_f_write, streams3, {})
# and re-read just to check
out_f_write.seek(0)
streams2, hdr = tv.read(out_f_write)
assert_true(streamlist_equal(streams, streams2))
@parametric
def test_empty_header():
for endian in '<>':
for version in (1, 2):
hdr = tv.empty_header(endian, version)
yield assert_equal(hdr['id_string'], 'TRACK')
yield assert_equal(hdr['version'], version)
yield assert_equal(hdr['hdr_size'], 1000)
yield assert_array_equal(
hdr['image_orientation_patient'],
[0,0,0,0,0,0])
hdr = tv.empty_header(version=2)
yield assert_array_equal(hdr['vox_to_ras'], np.zeros((4,4)))
hdr_endian = tv.endian_codes[tv.empty_header().dtype.byteorder]
yield assert_equal(hdr_endian, tv.native_code)
@parametric
def test_get_affine():
hdr = tv.empty_header()
# default header gives useless affine
yield assert_array_equal(tv.aff_from_hdr(hdr),
np.diag([0,0,0,1]))
hdr['voxel_size'] = 1
yield assert_array_equal(tv.aff_from_hdr(hdr),
np.diag([0,0,0,1]))
# DICOM direction cosines
hdr['image_orientation_patient'] = [1,0,0,0,1,0]
yield assert_array_equal(tv.aff_from_hdr(hdr),
np.diag([-1,-1,1,1]))
# RAS direction cosines
hdr['image_orientation_patient'] = [-1,0,0,0,-1,0]
yield assert_array_equal(tv.aff_from_hdr(hdr),
np.eye(4))
# translations
hdr['origin'] = [1,2,3]
exp_aff = np.eye(4)
exp_aff[:3,3] = [-1,-2,3]
yield assert_array_equal(tv.aff_from_hdr(hdr),
exp_aff)
# now use the easier vox_to_ras field
hdr = tv.empty_header()
aff = np.eye(4)
aff[:3,:] =
|
np.arange(12)
|
numpy.arange
|
#!/usr/bin/bash
# Author: GMFTBY, sfs, hyx
# Time: 2019.11.5
from nltk.translate.bleu_score import sentence_bleu, corpus_bleu
from nltk.translate.bleu_score import SmoothingFunction
from nltk.collocations import BigramCollocationFinder
from nltk.probability import FreqDist
import argparse
import codecs
import numpy as np
import math
# ========== fuck nlg-eval fuck ========== #
# ========== Our own embedding-based metric ========== #
def cal_vector_extrema(x, y, dic):
# x and y are the list of the words
def vecterize(p):
vectors = []
for w in p:
if w in dic:
vectors.append(dic[w])
else:
vectors.append(dic['<unk>'])
if not vectors:
vectors.append(dic['<unk>'])
return np.stack(vectors)
x = vecterize(x)
y = vecterize(y)
vec_x = np.max(x, axis=0)
vec_y = np.max(y, axis=0)
assert len(vec_x) == len(vec_y), "len(vec_x) != len(vec_y)"
zero_list = np.zeros(len(vec_x))
if vec_x.all() == zero_list.all() or vec_y.all() == zero_list.all():
return float(1) if vec_x.all() == vec_y.all() else float(0)
res = np.array([[vec_x[i] * vec_y[i], vec_x[i] * vec_x[i], vec_y[i] * vec_y[i]] for i in range(len(vec_x))])
cos = sum(res[:, 0]) / (np.sqrt(sum(res[:, 1])) * np.sqrt(sum(res[:, 2])))
return cos
def cal_embedding_average(x, y, dic):
# x and y are the list of the words
def vecterize(p):
vectors = []
for w in p:
if w in dic:
vectors.append(dic[w])
else:
vectors.append(dic['<unk>'])
if not vectors:
vectors.append(dic['<unk>'])
return np.stack(vectors)
x = vecterize(x)
y = vecterize(y)
vec_x = np.array([0 for _ in range(len(x[0]))]) # 存放句向量
for x_v in x:
x_v = np.array(x_v)
vec_x = np.add(x_v, vec_x)
vec_x = vec_x / math.sqrt(sum(
|
np.square(vec_x)
|
numpy.square
|
#!/usr/bin/env python3
# -*- coding: utf-8 -*-
"""
Created on Wed Jun 12 14:47:12 2019
@author: minghao
"""
default_filename = '/Users/mac/Documents/University/Github/data_road/training/calib/um_000000.txt'
import os
import numpy as np
import cv2
class BevParams(object):
def __init__(self, bev_res, bev_xLimits, bev_zLimits, imSize):
bev_size = (round((bev_zLimits[1] - bev_zLimits[0]) / bev_res), \
round((bev_xLimits[1] - bev_xLimits[0]) / bev_res))
self.bev_size = bev_size
self.bev_res = bev_res
self.bev_xLimits = bev_xLimits
self.bev_zLimits = bev_zLimits
self.imSize = imSize
def px2meter(self, px_in):
return px_in * self.bev_res
def meter2px(self, meter_in):
return meter_in / self.bev_res #to_decide?
def convertPositionMetric2Pixel(self, XZpointArrays):
allX = XZpointArrays[:,0]
allZ = XZpointArrays[:,1]
allZconverted = self.bev_size[0] - self.meter2px(allZ - self.bev_zLimits[0])
allXconverted = self.meter2px(allX - self.bev_xLimits[0])
return np.float32( [allXconverted, allZconverted] ).T
def convertPositionPixel2Metric(self, XYpointArrays):
allX = XYpointArrays[:,0]
allY = XYpointArrays[:,1]
allYconverted = self.px2meter(self.bev_size[0] - allY) + self.bev_zLimits[0]
allXconverted = self.px2meter(allX) + self.bev_xLimits[0]
return np.float32( [allXconverted, allYconverted] ).T
def convertPositionPixel2Metric2(self, inputTupleY, inputTupleX):
result_arr = self.convertPositionPixel2Metric(np.array( [[inputTupleY],[inputTupleX]] ))
return (result_arr[0,0], result_arr[0,1])
def readKittiCalib(filename, dtype = np.float32):
out_dict = {}
with open(filename,'rb') as f:
allcontent = f.readlines()
for contentRaw in allcontent:
content = contentRaw.strip()
if len(content) == 0:
continue
if content[0]!='#':
tmp = content.decode().split(':')
assert len(tmp)==2, 'wrong file format, only one : per line!'
var = tmp[0].strip()
values = np.array(tmp[-1].strip().split(' '),dtype)
out_dict[var] = values
return out_dict
class KittiCalibration(object):
def __init__(self):
pass
def readFromFile(self,filename = default_filename):
cur_calibStuff_dict = readKittiCalib(filename)
self.setup(cur_calibStuff_dict)
def setup(self, dictWithKittiStuff, useRect = False):
dtype = np.float32
self.P2 = np.matrix(dictWithKittiStuff['P2']).reshape((3,4))
if useRect:
R2_1 = self.P2
else:
R0_rect_raw = np.array(dictWithKittiStuff['R0_rect']).reshape((3,3))
self.R0_rect = np.matrix(np.hstack((np.vstack((R0_rect_raw, np.zeros((1,3), dtype))), np.zeros((4,1), dtype))))
self.R0_rect[3,3]=1.0
R2_1 = np.dot(self.P2, self.R0_rect)
Tr_cam_to_road_raw = np.array( dictWithKittiStuff['Tr_cam_to_road'] ).reshape(3,4)
self.Tr_cam_to_road_raw = np.matrix( np.vstack((Tr_cam_to_road_raw, np.zeros((1,4), dtype))) )
self.Tr_cam_to_road_raw[3,3] = 1.0
self.Tr = np.dot( R2_1, self.Tr_cam_to_road_raw.I )
self.Tr33 = self.Tr[:,[0,2,3]]
def get_matrix33(self):
assert not self.Tr33 is None
return self.Tr33
class BirdsEyeView(object):
imSize = None
def __init__(self, bev_res= 0.05, bev_xRange_minMax = (-10, 10), bev_zRange_minMax = (6, 46)):
self.calib = KittiCalibration()
bev_res = bev_res
self.bevParams = BevParams(bev_res, bev_xRange_minMax, bev_zRange_minMax, self.imSize)
self.srcPoints = np.float32([ [0,0], [200,0], [200,200], [0,200] ])
def world2image_uvMat(self, uv_mat):
if uv_mat.shape[0] == 2:
if len(uv_mat.shape) == 1:
uv_mat = uv_mat[:,np.newaxis]
uv_mat = np.vstack( (uv_mat, np.ones((1,uv_mat.shape[1]))))
result = np.dot( self.Tr33, uv_mat )
resultB = np.broadcast_arrays(result, result[-1, :])
return resultB[0] / resultB[1]
def image2world_uvMat(self, uv_mat):
if uv_mat.shape[0] == 2:
if len(uv_mat.shape)==1:
uv_mat = uv_mat[:,np.newaxis]
uv_mat = np.vstack((uv_mat, np.ones((1, uv_mat.shape[1]))))
result =
|
np.dot(self.Tr33.I, uv_mat)
|
numpy.dot
|
import autograd.numpy as anp
import numpy as np
from autograd import grad
from ezmodel.custom2.kernel import GaussianKernel, KernelFactory
from ezmodel.custom2.optimizer import Adam
from ezmodel.core.model import Model
from ezmodel.core.transformation import NoNormalization
from ezmodel.util.transformation.zero_to_one import ZeroToOneNormalization
def predict(params, fac, n_closest, X, Y, x):
kernel = kernel_from_params(fac, params)
d = kernel.D(x[None, :], X)[0]
closest = d.argsort()[:n_closest]
kernel.fit(X[closest], Y[closest])
out = kernel.predict(x[None, :])
y_hat = out["y"][0]
return y_hat, kernel
def mse(params, fac, n_closest, X, Y, x, y):
y_hat, kernel = predict(params, fac, n_closest, X, Y, x)
mse = (y_hat - y) ** 2
return mse[0]
def kernel_from_params(fac, params):
theta = anp.exp(params[0])
kernel = fac.create(theta=theta)
return kernel
def avg_mse(params, X, y, n_nearest, fac):
_mse = []
for j in range(len(X)):
m = np.full(len(X), True)
m[j] = False
__mse = mse(params, fac, n_nearest, X[m], y[m], X[j], y[j])
_mse.append(__mse)
return np.array(_mse).mean()
class LGP(Model):
def __init__(self,
n_nearest=None,
norm_X=NoNormalization(),
# norm_y=NoNormalization(),
kernel="gaussian",
n_max_iter=0,
# norm_X=Standardization(),
norm_y=ZeroToOneNormalization(),
verbose=False,
**kwargs):
super().__init__(norm_X=norm_X, norm_y=norm_y, **kwargs)
self.n_nearest = n_nearest
self.n_max_iter = n_max_iter
self.fac = KernelFactory(kernel, **kwargs)
self.verbose = verbose
def _fit(self, X, y, **kwargs):
n_nearest = self.n_nearest
if n_nearest is None:
params = np.array([anp.log(1.0)])
K = np.arange(3, 16).astype(int)
vals = np.array([avg_mse(params, X, y, k, self.fac) for k in K])
n_nearest = K[vals.argmin()]
self.n_nearest = n_nearest
thetas = np.linspace(0.01, 10, 50)
vals = np.array([avg_mse(np.array([anp.log(theta)]), X, y, n_nearest, self.fac) for theta in thetas])
theta = thetas[vals.argmin()]
params = np.array([anp.log(theta)])
optim = Adam(params)
# optim = SGD(params, alpha=0.0001)
for i in range(self.n_max_iter):
_grad = []
# for j in np.random.permutation(len(X)):
for j in range(len(X)):
m = np.full(len(X), True)
m[j] = False
# _mse = mse(optim.X, self.fac, n_nearest, X[m], y[m], X[j], y[j])
func_grad = grad(mse)
__grad = func_grad(optim.X, self.fac, n_nearest, X[m], y[m], X[j], y[j])
_grad.append(__grad)
if np.any(np.isnan(__grad)):
func_grad(optim.X, self.fac, n_nearest, X[m], y[m], X[j], y[j])
if self.verbose:
_mse = avg_mse(optim.X, X, y, n_nearest, self.fac)
print(i, np.exp(optim.X), _mse)
_grad = np.array(_grad).sum(axis=0)
if np.all(_grad < 1e-8) or np.any(np.isnan(_grad)):
break
optim.apply(_grad)
# print(np.array(_mse).mean())
self.params = optim.X
def _predict(self, X, out):
y = []
for j in range(len(X)):
y_hat, _ = predict(self.params, self.fac, self.n_nearest, self.X, self.y, X[j])
y.append(y_hat)
out["y"] =
|
np.array(y)
|
numpy.array
|
"""
2D–3D Geometric Fusion network using Multi-Neighbourhood Graph Convolution for RGB-D indoor scene classification
2021 <NAME> <<EMAIL>>
"""
import torch
from torch_geometric.data import Data
import math
import random
import numpy as np
import transforms3d
import os
import h5py
import torch_cluster
from Fusion2D3DMUNEGC.utilities import utils
def dropout(P, F, p):
idx = random.sample(range(P.shape[0]), int(math.ceil((1-p)*P.shape[0])))
return P[idx, :], F[idx, :] if F is not None else None
def random_crop_3D(P, F, factor):
npoints = P.shape[0]
n_points_after_crop = np.round(npoints*factor).astype(np.int)
points_max = (P.max(axis=0)*1000).astype(np.int)
points_min = (P.min(axis=0)*1000).astype(np.int)
centroid = np.asarray([np.random.randint(low=points_min[0], high=points_max[0], dtype=int),
np.random.randint(low=points_min[1], high=points_max[1], dtype=int),
np.random.randint(low=points_min[2], high=points_max[2], dtype=int)])
centroid = centroid.astype(np.float32)/1000
rad = 0.1
inc = 0.2
npoints_inside_sphere = 0
x = torch.from_numpy(P)
y = torch.from_numpy(centroid).unsqueeze(0)
while npoints_inside_sphere < n_points_after_crop:
_, crop = torch_cluster.radius(x, y, rad, max_num_neighbors=n_points_after_crop)
npoints_inside_sphere = len(crop)
rad = np.round(rad + inc, 1)
return P[crop.numpy()], F[crop.numpy()]
class PCH5Dataset(torch.utils.data.Dataset):
def __init__(self, root_path, h5_folder, split,
transform3d=None,
range01=False, pos_int16=False,
random_crop=False, factor_rand=False, factor=1):
self.root_path = root_path
self.h5_path = os.path.join(self.root_path, h5_folder)
self.split = utils.read_string_list(os.path.join(self.root_path, split))
self.h5_folder = h5_folder
self.transform3d = transform3d
self.range01 = range01
self.pos_int16 = pos_int16
self.random_crop = random_crop
self.factor_rand = factor_rand
self.factor = factor
def __getitem__(self, index):
h5_file = h5py.File(os.path.join(self.h5_path, self.split[index]+".h5"), 'r')
cls = int(np.asarray((h5_file["label"])))
P = np.asarray(h5_file["points"])
if len(P.shape) == 3:
P = P.reshape(-1, 3)
if self.pos_int16:
P = (P/1000).astype(np.float32)
F = None
if 'feat' in h5_file.keys():
F = np.asarray(h5_file["feat"], dtype=np.float32)
if len(F.shape) == 1:
F = np.transpose([F])
elif len(F.shape) == 3:
F = F.reshape(-1, 3)
if self.range01:
F = F/255
else:
raise RuntimeError('node feat do not exist')
if self.random_crop:
if self.factor_rand is True:
factor = np.random.randint(low=self.factor*100, high=100, dtype=np.int)/100
else:
factor = self.factor
P, F = random_crop_3D(P, F, factor)
if self.transform3d is not None:
if self.transform3d["dropout"] > 0:
P, F = dropout(P, F, self.transform3d["dropout"])
M = np.eye(3)
if self.transform3d["rot"]:
angle = random.uniform(0, 2*math.pi)
M = np.dot(transforms3d.axangles.axangle2mat([0, 1, 0], angle), M)
if self.transform3d["mirror"] > 0:
if random.random() < self.transform3d["mirror"]/2:
M = np.dot(transforms3d.zooms.zfdir2mat(-1, [1, 0, 0]), M)
if random.random() < self.transform3d["mirror"]/2:
M = np.dot(transforms3d.zooms.zfdir2mat(-1, [0, 0, 1]), M)
P =
|
np.dot(P, M.T)
|
numpy.dot
|
from __future__ import division, absolute_import, print_function
import warnings
import numpy as np
from numpy.testing import (
run_module_suite, TestCase, assert_, assert_equal, assert_almost_equal,
assert_no_warnings, assert_raises, assert_array_equal, suppress_warnings
)
# Test data
_ndat = np.array([[0.6244, np.nan, 0.2692, 0.0116, np.nan, 0.1170],
[0.5351, -0.9403, np.nan, 0.2100, 0.4759, 0.2833],
[np.nan, np.nan, np.nan, 0.1042, np.nan, -0.5954],
[0.1610, np.nan, np.nan, 0.1859, 0.3146, np.nan]])
# Rows of _ndat with nans removed
_rdat = [np.array([0.6244, 0.2692, 0.0116, 0.1170]),
np.array([0.5351, -0.9403, 0.2100, 0.4759, 0.2833]),
np.array([0.1042, -0.5954]),
np.array([0.1610, 0.1859, 0.3146])]
# Rows of _ndat with nans converted to ones
_ndat_ones = np.array([[0.6244, 1.0, 0.2692, 0.0116, 1.0, 0.1170],
[0.5351, -0.9403, 1.0, 0.2100, 0.4759, 0.2833],
[1.0, 1.0, 1.0, 0.1042, 1.0, -0.5954],
[0.1610, 1.0, 1.0, 0.1859, 0.3146, 1.0]])
# Rows of _ndat with nans converted to zeros
_ndat_zeros = np.array([[0.6244, 0.0, 0.2692, 0.0116, 0.0, 0.1170],
[0.5351, -0.9403, 0.0, 0.2100, 0.4759, 0.2833],
[0.0, 0.0, 0.0, 0.1042, 0.0, -0.5954],
[0.1610, 0.0, 0.0, 0.1859, 0.3146, 0.0]])
class TestNanFunctions_MinMax(TestCase):
nanfuncs = [np.nanmin, np.nanmax]
stdfuncs = [np.min, np.max]
def test_mutation(self):
# Check that passed array is not modified.
ndat = _ndat.copy()
for f in self.nanfuncs:
f(ndat)
assert_equal(ndat, _ndat)
def test_keepdims(self):
mat = np.eye(3)
for nf, rf in zip(self.nanfuncs, self.stdfuncs):
for axis in [None, 0, 1]:
tgt = rf(mat, axis=axis, keepdims=True)
res = nf(mat, axis=axis, keepdims=True)
assert_(res.ndim == tgt.ndim)
def test_out(self):
mat = np.eye(3)
for nf, rf in zip(self.nanfuncs, self.stdfuncs):
resout = np.zeros(3)
tgt = rf(mat, axis=1)
res = nf(mat, axis=1, out=resout)
assert_almost_equal(res, resout)
assert_almost_equal(res, tgt)
def test_dtype_from_input(self):
codes = 'efdgFDG'
for nf, rf in zip(self.nanfuncs, self.stdfuncs):
for c in codes:
mat = np.eye(3, dtype=c)
tgt = rf(mat, axis=1).dtype.type
res = nf(mat, axis=1).dtype.type
assert_(res is tgt)
# scalar case
tgt = rf(mat, axis=None).dtype.type
res = nf(mat, axis=None).dtype.type
assert_(res is tgt)
def test_result_values(self):
for nf, rf in zip(self.nanfuncs, self.stdfuncs):
tgt = [rf(d) for d in _rdat]
res = nf(_ndat, axis=1)
assert_almost_equal(res, tgt)
def test_allnans(self):
mat = np.array([np.nan]*9).reshape(3, 3)
for f in self.nanfuncs:
for axis in [None, 0, 1]:
with warnings.catch_warnings(record=True) as w:
warnings.simplefilter('always')
assert_(np.isnan(f(mat, axis=axis)).all())
assert_(len(w) == 1, 'no warning raised')
assert_(issubclass(w[0].category, RuntimeWarning))
# Check scalars
with warnings.catch_warnings(record=True) as w:
warnings.simplefilter('always')
assert_(np.isnan(f(np.nan)))
assert_(len(w) == 1, 'no warning raised')
assert_(issubclass(w[0].category, RuntimeWarning))
def test_masked(self):
mat = np.ma.fix_invalid(_ndat)
msk = mat._mask.copy()
for f in [np.nanmin]:
res = f(mat, axis=1)
tgt = f(_ndat, axis=1)
assert_equal(res, tgt)
assert_equal(mat._mask, msk)
assert_(not np.isinf(mat).any())
def test_scalar(self):
for f in self.nanfuncs:
assert_(f(0.) == 0.)
def test_matrices(self):
# Check that it works and that type and
# shape are preserved
mat = np.matrix(np.eye(3))
for f in self.nanfuncs:
res = f(mat, axis=0)
assert_(isinstance(res, np.matrix))
assert_(res.shape == (1, 3))
res = f(mat, axis=1)
assert_(isinstance(res, np.matrix))
assert_(res.shape == (3, 1))
res = f(mat)
assert_(np.isscalar(res))
# check that rows of nan are dealt with for subclasses (#4628)
mat[1] = np.nan
for f in self.nanfuncs:
with warnings.catch_warnings(record=True) as w:
warnings.simplefilter('always')
res = f(mat, axis=0)
assert_(isinstance(res, np.matrix))
assert_(not np.any(np.isnan(res)))
assert_(len(w) == 0)
with warnings.catch_warnings(record=True) as w:
warnings.simplefilter('always')
res = f(mat, axis=1)
assert_(isinstance(res, np.matrix))
assert_(np.isnan(res[1, 0]) and not np.isnan(res[0, 0])
and not np.isnan(res[2, 0]))
assert_(len(w) == 1, 'no warning raised')
assert_(issubclass(w[0].category, RuntimeWarning))
with warnings.catch_warnings(record=True) as w:
warnings.simplefilter('always')
res = f(mat)
assert_(np.isscalar(res))
assert_(res != np.nan)
assert_(len(w) == 0)
class TestNanFunctions_ArgminArgmax(TestCase):
nanfuncs = [np.nanargmin, np.nanargmax]
def test_mutation(self):
# Check that passed array is not modified.
ndat = _ndat.copy()
for f in self.nanfuncs:
f(ndat)
assert_equal(ndat, _ndat)
def test_result_values(self):
for f, fcmp in zip(self.nanfuncs, [np.greater, np.less]):
for row in _ndat:
with suppress_warnings() as sup:
sup.filter(RuntimeWarning, "invalid value encountered in")
ind = f(row)
val = row[ind]
# comparing with NaN is tricky as the result
# is always false except for NaN != NaN
assert_(not np.isnan(val))
assert_(not fcmp(val, row).any())
assert_(not np.equal(val, row[:ind]).any())
def test_allnans(self):
mat = np.array([np.nan]*9).reshape(3, 3)
for f in self.nanfuncs:
for axis in [None, 0, 1]:
assert_raises(ValueError, f, mat, axis=axis)
assert_raises(ValueError, f, np.nan)
def test_empty(self):
mat = np.zeros((0, 3))
for f in self.nanfuncs:
for axis in [0, None]:
assert_raises(ValueError, f, mat, axis=axis)
for axis in [1]:
res = f(mat, axis=axis)
assert_equal(res, np.zeros(0))
def test_scalar(self):
for f in self.nanfuncs:
assert_(f(0.) == 0.)
def test_matrices(self):
# Check that it works and that type and
# shape are preserved
mat = np.matrix(np.eye(3))
for f in self.nanfuncs:
res = f(mat, axis=0)
assert_(isinstance(res, np.matrix))
assert_(res.shape == (1, 3))
res = f(mat, axis=1)
assert_(isinstance(res, np.matrix))
assert_(res.shape == (3, 1))
res = f(mat)
assert_(np.isscalar(res))
class TestNanFunctions_IntTypes(TestCase):
int_types = (np.int8, np.int16, np.int32, np.int64, np.uint8,
np.uint16, np.uint32, np.uint64)
mat = np.array([127, 39, 93, 87, 46])
def integer_arrays(self):
for dtype in self.int_types:
yield self.mat.astype(dtype)
def test_nanmin(self):
tgt = np.min(self.mat)
for mat in self.integer_arrays():
assert_equal(np.nanmin(mat), tgt)
def test_nanmax(self):
tgt = np.max(self.mat)
for mat in self.integer_arrays():
assert_equal(np.nanmax(mat), tgt)
def test_nanargmin(self):
tgt = np.argmin(self.mat)
for mat in self.integer_arrays():
assert_equal(np.nanargmin(mat), tgt)
def test_nanargmax(self):
tgt = np.argmax(self.mat)
for mat in self.integer_arrays():
assert_equal(np.nanargmax(mat), tgt)
def test_nansum(self):
tgt = np.sum(self.mat)
for mat in self.integer_arrays():
assert_equal(np.nansum(mat), tgt)
def test_nanprod(self):
tgt = np.prod(self.mat)
for mat in self.integer_arrays():
assert_equal(np.nanprod(mat), tgt)
def test_nancumsum(self):
tgt = np.cumsum(self.mat)
for mat in self.integer_arrays():
assert_equal(np.nancumsum(mat), tgt)
def test_nancumprod(self):
tgt = np.cumprod(self.mat)
for mat in self.integer_arrays():
assert_equal(np.nancumprod(mat), tgt)
def test_nanmean(self):
tgt = np.mean(self.mat)
for mat in self.integer_arrays():
assert_equal(np.nanmean(mat), tgt)
def test_nanvar(self):
tgt = np.var(self.mat)
for mat in self.integer_arrays():
assert_equal(np.nanvar(mat), tgt)
tgt = np.var(mat, ddof=1)
for mat in self.integer_arrays():
assert_equal(np.nanvar(mat, ddof=1), tgt)
def test_nanstd(self):
tgt = np.std(self.mat)
for mat in self.integer_arrays():
assert_equal(np.nanstd(mat), tgt)
tgt = np.std(self.mat, ddof=1)
for mat in self.integer_arrays():
assert_equal(np.nanstd(mat, ddof=1), tgt)
class SharedNanFunctionsTestsMixin(object):
def test_mutation(self):
# Check that passed array is not modified.
ndat = _ndat.copy()
for f in self.nanfuncs:
f(ndat)
assert_equal(ndat, _ndat)
def test_keepdims(self):
mat = np.eye(3)
for nf, rf in zip(self.nanfuncs, self.stdfuncs):
for axis in [None, 0, 1]:
tgt = rf(mat, axis=axis, keepdims=True)
res = nf(mat, axis=axis, keepdims=True)
assert_(res.ndim == tgt.ndim)
def test_out(self):
mat = np.eye(3)
for nf, rf in zip(self.nanfuncs, self.stdfuncs):
resout = np.zeros(3)
tgt = rf(mat, axis=1)
res = nf(mat, axis=1, out=resout)
assert_almost_equal(res, resout)
assert_almost_equal(res, tgt)
def test_dtype_from_dtype(self):
mat = np.eye(3)
codes = 'efdgFDG'
for nf, rf in zip(self.nanfuncs, self.stdfuncs):
for c in codes:
with suppress_warnings() as sup:
if nf in {np.nanstd, np.nanvar} and c in 'FDG':
# Giving the warning is a small bug, see gh-8000
sup.filter(np.ComplexWarning)
tgt = rf(mat, dtype=np.dtype(c), axis=1).dtype.type
res = nf(mat, dtype=np.dtype(c), axis=1).dtype.type
assert_(res is tgt)
# scalar case
tgt = rf(mat, dtype=np.dtype(c), axis=None).dtype.type
res = nf(mat, dtype=np.dtype(c), axis=None).dtype.type
assert_(res is tgt)
def test_dtype_from_char(self):
mat = np.eye(3)
codes = 'efdgFDG'
for nf, rf in zip(self.nanfuncs, self.stdfuncs):
for c in codes:
with suppress_warnings() as sup:
if nf in {np.nanstd, np.nanvar} and c in 'FDG':
# Giving the warning is a small bug, see gh-8000
sup.filter(np.ComplexWarning)
tgt = rf(mat, dtype=c, axis=1).dtype.type
res = nf(mat, dtype=c, axis=1).dtype.type
assert_(res is tgt)
# scalar case
tgt = rf(mat, dtype=c, axis=None).dtype.type
res = nf(mat, dtype=c, axis=None).dtype.type
assert_(res is tgt)
def test_dtype_from_input(self):
codes = 'efdgFDG'
for nf, rf in zip(self.nanfuncs, self.stdfuncs):
for c in codes:
mat = np.eye(3, dtype=c)
tgt = rf(mat, axis=1).dtype.type
res = nf(mat, axis=1).dtype.type
assert_(res is tgt, "res %s, tgt %s" % (res, tgt))
# scalar case
tgt = rf(mat, axis=None).dtype.type
res = nf(mat, axis=None).dtype.type
assert_(res is tgt)
def test_result_values(self):
for nf, rf in zip(self.nanfuncs, self.stdfuncs):
tgt = [rf(d) for d in _rdat]
res = nf(_ndat, axis=1)
assert_almost_equal(res, tgt)
def test_scalar(self):
for f in self.nanfuncs:
assert_(f(0.) == 0.)
def test_matrices(self):
# Check that it works and that type and
# shape are preserved
mat = np.matrix(np.eye(3))
for f in self.nanfuncs:
res = f(mat, axis=0)
assert_(isinstance(res, np.matrix))
assert_(res.shape == (1, 3))
res = f(mat, axis=1)
assert_(isinstance(res, np.matrix))
assert_(res.shape == (3, 1))
res = f(mat)
assert_(np.isscalar(res))
class TestNanFunctions_SumProd(TestCase, SharedNanFunctionsTestsMixin):
nanfuncs = [np.nansum, np.nanprod]
stdfuncs = [np.sum, np.prod]
def test_allnans(self):
# Check for FutureWarning
with warnings.catch_warnings(record=True) as w:
warnings.simplefilter('always')
res = np.nansum([np.nan]*3, axis=None)
assert_(res == 0, 'result is not 0')
assert_(len(w) == 0, 'warning raised')
# Check scalar
res = np.nansum(np.nan)
assert_(res == 0, 'result is not 0')
assert_(len(w) == 0, 'warning raised')
# Check there is no warning for not all-nan
np.nansum([0]*3, axis=None)
assert_(len(w) == 0, 'unwanted warning raised')
def test_empty(self):
for f, tgt_value in zip([np.nansum, np.nanprod], [0, 1]):
mat = np.zeros((0, 3))
tgt = [tgt_value]*3
res = f(mat, axis=0)
assert_equal(res, tgt)
tgt = []
res = f(mat, axis=1)
assert_equal(res, tgt)
tgt = tgt_value
res = f(mat, axis=None)
assert_equal(res, tgt)
class TestNanFunctions_CumSumProd(TestCase, SharedNanFunctionsTestsMixin):
nanfuncs = [np.nancumsum, np.nancumprod]
stdfuncs = [np.cumsum, np.cumprod]
def test_allnans(self):
for f, tgt_value in zip(self.nanfuncs, [0, 1]):
# Unlike other nan-functions, sum/prod/cumsum/cumprod don't warn on all nan input
with assert_no_warnings():
res = f([np.nan]*3, axis=None)
tgt = tgt_value*np.ones((3))
assert_(np.array_equal(res, tgt), 'result is not %s * np.ones((3))' % (tgt_value))
# Check scalar
res = f(np.nan)
tgt = tgt_value*np.ones((1))
assert_(np.array_equal(res, tgt), 'result is not %s * np.ones((1))' % (tgt_value))
# Check there is no warning for not all-nan
f([0]*3, axis=None)
def test_empty(self):
for f, tgt_value in zip(self.nanfuncs, [0, 1]):
mat = np.zeros((0, 3))
tgt = tgt_value*np.ones((0, 3))
res = f(mat, axis=0)
assert_equal(res, tgt)
tgt = mat
res = f(mat, axis=1)
assert_equal(res, tgt)
tgt = np.zeros((0))
res = f(mat, axis=None)
assert_equal(res, tgt)
def test_keepdims(self):
for f, g in zip(self.nanfuncs, self.stdfuncs):
mat = np.eye(3)
for axis in [None, 0, 1]:
tgt = f(mat, axis=axis, out=None)
res = g(mat, axis=axis, out=None)
assert_(res.ndim == tgt.ndim)
for f in self.nanfuncs:
d = np.ones((3, 5, 7, 11))
# Randomly set some elements to NaN:
rs = np.random.RandomState(0)
d[rs.rand(*d.shape) < 0.5] = np.nan
res = f(d, axis=None)
assert_equal(res.shape, (1155,))
for axis in np.arange(4):
res = f(d, axis=axis)
assert_equal(res.shape, (3, 5, 7, 11))
def test_matrices(self):
# Check that it works and that type and
# shape are preserved
mat = np.matrix(np.eye(3))
for f in self.nanfuncs:
for axis in np.arange(2):
res = f(mat, axis=axis)
assert_(isinstance(res, np.matrix))
assert_(res.shape == (3, 3))
res = f(mat)
assert_(res.shape == (1, 3*3))
def test_result_values(self):
for axis in (-2, -1, 0, 1, None):
tgt = np.cumprod(_ndat_ones, axis=axis)
res = np.nancumprod(_ndat, axis=axis)
assert_almost_equal(res, tgt)
tgt = np.cumsum(_ndat_zeros,axis=axis)
res = np.nancumsum(_ndat, axis=axis)
assert_almost_equal(res, tgt)
def test_out(self):
mat = np.eye(3)
for nf, rf in zip(self.nanfuncs, self.stdfuncs):
resout = np.eye(3)
for axis in (-2, -1, 0, 1):
tgt = rf(mat, axis=axis)
res = nf(mat, axis=axis, out=resout)
assert_almost_equal(res, resout)
assert_almost_equal(res, tgt)
class TestNanFunctions_MeanVarStd(TestCase, SharedNanFunctionsTestsMixin):
nanfuncs = [np.nanmean, np.nanvar, np.nanstd]
stdfuncs = [np.mean, np.var, np.std]
def test_dtype_error(self):
for f in self.nanfuncs:
for dtype in [np.bool_, np.int_, np.object_]:
assert_raises(TypeError, f, _ndat, axis=1, dtype=dtype)
def test_out_dtype_error(self):
for f in self.nanfuncs:
for dtype in [np.bool_, np.int_, np.object_]:
out = np.empty(_ndat.shape[0], dtype=dtype)
assert_raises(TypeError, f, _ndat, axis=1, out=out)
def test_ddof(self):
nanfuncs = [np.nanvar, np.nanstd]
stdfuncs = [np.var, np.std]
for nf, rf in zip(nanfuncs, stdfuncs):
for ddof in [0, 1]:
tgt = [rf(d, ddof=ddof) for d in _rdat]
res = nf(_ndat, axis=1, ddof=ddof)
assert_almost_equal(res, tgt)
def test_ddof_too_big(self):
nanfuncs = [np.nanvar, np.nanstd]
stdfuncs = [np.var, np.std]
dsize = [len(d) for d in _rdat]
for nf, rf in zip(nanfuncs, stdfuncs):
for ddof in range(5):
with suppress_warnings() as sup:
sup.record(RuntimeWarning)
sup.filter(np.ComplexWarning)
tgt = [ddof >= d for d in dsize]
res = nf(_ndat, axis=1, ddof=ddof)
assert_equal(np.isnan(res), tgt)
if any(tgt):
assert_(len(sup.log) == 1)
else:
assert_(len(sup.log) == 0)
def test_allnans(self):
mat = np.array([np.nan]*9).reshape(3, 3)
for f in self.nanfuncs:
for axis in [None, 0, 1]:
with warnings.catch_warnings(record=True) as w:
warnings.simplefilter('always')
assert_(np.isnan(f(mat, axis=axis)).all())
assert_(len(w) == 1)
assert_(issubclass(w[0].category, RuntimeWarning))
# Check scalar
assert_(np.isnan(f(np.nan)))
assert_(len(w) == 2)
assert_(issubclass(w[0].category, RuntimeWarning))
def test_empty(self):
mat = np.zeros((0, 3))
for f in self.nanfuncs:
for axis in [0, None]:
with warnings.catch_warnings(record=True) as w:
warnings.simplefilter('always')
assert_(np.isnan(f(mat, axis=axis)).all())
assert_(len(w) == 1)
assert_(issubclass(w[0].category, RuntimeWarning))
for axis in [1]:
with warnings.catch_warnings(record=True) as w:
warnings.simplefilter('always')
assert_equal(f(mat, axis=axis), np.zeros([]))
assert_(len(w) == 0)
class TestNanFunctions_Median(TestCase):
def test_mutation(self):
# Check that passed array is not modified.
ndat = _ndat.copy()
np.nanmedian(ndat)
assert_equal(ndat, _ndat)
def test_keepdims(self):
mat = np.eye(3)
for axis in [None, 0, 1]:
tgt = np.median(mat, axis=axis, out=None, overwrite_input=False)
res = np.nanmedian(mat, axis=axis, out=None, overwrite_input=False)
assert_(res.ndim == tgt.ndim)
d = np.ones((3, 5, 7, 11))
# Randomly set some elements to NaN:
w = np.random.random((4, 200)) * np.array(d.shape)[:, None]
w = w.astype(np.intp)
d[tuple(w)] = np.nan
with suppress_warnings() as sup:
sup.filter(RuntimeWarning)
res = np.nanmedian(d, axis=None, keepdims=True)
assert_equal(res.shape, (1, 1, 1, 1))
res = np.nanmedian(d, axis=(0, 1), keepdims=True)
assert_equal(res.shape, (1, 1, 7, 11))
res = np.nanmedian(d, axis=(0, 3), keepdims=True)
assert_equal(res.shape, (1, 5, 7, 1))
res = np.nanmedian(d, axis=(1,), keepdims=True)
assert_equal(res.shape, (3, 1, 7, 11))
res = np.nanmedian(d, axis=(0, 1, 2, 3), keepdims=True)
assert_equal(res.shape, (1, 1, 1, 1))
res = np.nanmedian(d, axis=(0, 1, 3), keepdims=True)
assert_equal(res.shape, (1, 1, 7, 1))
def test_out(self):
mat = np.random.rand(3, 3)
nan_mat = np.insert(mat, [0, 2], np.nan, axis=1)
resout = np.zeros(3)
tgt = np.median(mat, axis=1)
res = np.nanmedian(nan_mat, axis=1, out=resout)
assert_almost_equal(res, resout)
assert_almost_equal(res, tgt)
# 0-d output:
resout = np.zeros(())
tgt = np.median(mat, axis=None)
res = np.nanmedian(nan_mat, axis=None, out=resout)
assert_almost_equal(res, resout)
assert_almost_equal(res, tgt)
res = np.nanmedian(nan_mat, axis=(0, 1), out=resout)
assert_almost_equal(res, resout)
assert_almost_equal(res, tgt)
def test_small_large(self):
# test the small and large code paths, current cutoff 400 elements
for s in [5, 20, 51, 200, 1000]:
d = np.random.randn(4, s)
# Randomly set some elements to NaN:
w = np.random.randint(0, d.size, size=d.size // 5)
d.ravel()[w] = np.nan
d[:,0] = 1. # ensure at least one good value
# use normal median without nans to compare
tgt = []
for x in d:
nonan = np.compress(~np.isnan(x), x)
tgt.append(np.median(nonan, overwrite_input=True))
assert_array_equal(np.nanmedian(d, axis=-1), tgt)
def test_result_values(self):
tgt = [np.median(d) for d in _rdat]
res = np.nanmedian(_ndat, axis=1)
assert_almost_equal(res, tgt)
def test_allnans(self):
mat = np.array([np.nan]*9).reshape(3, 3)
for axis in [None, 0, 1]:
with suppress_warnings() as sup:
sup.record(RuntimeWarning)
assert_(np.isnan(np.nanmedian(mat, axis=axis)).all())
if axis is None:
assert_(len(sup.log) == 1)
else:
assert_(len(sup.log) == 3)
# Check scalar
assert_(np.isnan(np.nanmedian(np.nan)))
if axis is None:
assert_(len(sup.log) == 2)
else:
assert_(len(sup.log) == 4)
def test_empty(self):
mat = np.zeros((0, 3))
for axis in [0, None]:
with warnings.catch_warnings(record=True) as w:
warnings.simplefilter('always')
assert_(np.isnan(np.nanmedian(mat, axis=axis)).all())
assert_(len(w) == 1)
assert_(issubclass(w[0].category, RuntimeWarning))
for axis in [1]:
with warnings.catch_warnings(record=True) as w:
warnings.simplefilter('always')
assert_equal(np.nanmedian(mat, axis=axis), np.zeros([]))
assert_(len(w) == 0)
def test_scalar(self):
assert_(np.nanmedian(0.) == 0.)
def test_extended_axis_invalid(self):
d = np.ones((3, 5, 7, 11))
assert_raises(np.AxisError, np.nanmedian, d, axis=-5)
assert_raises(np.AxisError, np.nanmedian, d, axis=(0, -5))
assert_raises(np.AxisError, np.nanmedian, d, axis=4)
assert_raises(np.AxisError, np.nanmedian, d, axis=(0, 4))
assert_raises(ValueError, np.nanmedian, d, axis=(1, 1))
def test_float_special(self):
with suppress_warnings() as sup:
sup.filter(RuntimeWarning)
for inf in [np.inf, -np.inf]:
a = np.array([[inf, np.nan], [np.nan, np.nan]])
assert_equal(np.nanmedian(a, axis=0), [inf, np.nan])
assert_equal(np.nanmedian(a, axis=1), [inf, np.nan])
assert_equal(np.nanmedian(a), inf)
# minimum fill value check
a = np.array([[np.nan, np.nan, inf],
[np.nan, np.nan, inf]])
assert_equal(np.nanmedian(a), inf)
assert_equal(np.nanmedian(a, axis=0), [np.nan, np.nan, inf])
assert_equal(np.nanmedian(a, axis=1), inf)
# no mask path
a = np.array([[inf, inf], [inf, inf]])
assert_equal(np.nanmedian(a, axis=1), inf)
a = np.array([[inf, 7, -inf, -9],
[-10, np.nan, np.nan, 5],
[4, np.nan, np.nan, inf]],
dtype=np.float32)
if inf > 0:
assert_equal(np.nanmedian(a, axis=0), [4., 7., -inf, 5.])
assert_equal(np.nanmedian(a), 4.5)
else:
assert_equal(np.nanmedian(a, axis=0), [-10., 7., -inf, -9.])
assert_equal(np.nanmedian(a), -2.5)
assert_equal(np.nanmedian(a, axis=-1), [-1., -2.5, inf])
for i in range(0, 10):
for j in range(1, 10):
a = np.array([([np.nan] * i) + ([inf] * j)] * 2)
assert_equal(
|
np.nanmedian(a)
|
numpy.nanmedian
|
"""
@author: <NAME> <<EMAIL>>
"""
import numpy
from .linear_chain_crf import LCRFModelRepresentation, LCRF
from .utilities import (
HOSemi_AStarSearcher,
vectorized_logsumexp,
generate_partitions,
generate_partition_boundaries,
)
class HOSemiCRFADModelRepresentation(LCRFModelRepresentation):
r"""Model representation that will hold data structures to be used in :class:`HOSemiCRF` class
Attributes:
P_codebook: set of proper prefixes of the elements in the set of patterns :attr:`Z_codebook`
e.g. {'':0, 'P':1, 'L':2, 'O':3, 'L|O':4, ...}
P_codebook_rev: reversed codebook of :attr:`P_codebook`
e.g. {0:'', 1:'P', 2:'L', 3:'O', 4:'L|O', ...}
P_len: dictionary comprising the length (i.e. number of elements) of elements in :attr:`P_codebook`
e.g. {'':0, 'P':1, 'L':1, 'O':1, 'L|O':2, ...}
P_elems: dictionary comprising the composing elements of every prefix in :attr:`P_codebook`
e.g. {'':('',), 'P':('P',), 'L':('L',), 'O':('O',), 'L|O':('L','O'), ...}
P_numchar: dictionary comprising the number of characters for every prefix in :attr:`P_codebook`
e.g. {'':0, 'P':1, 'L':1, 'O':1, 'L|O':3, ...}
f_transition: a dictionary representing forward transition data structure having the form:
{pi:{pky, (pk, y)}} where pi represents the longest prefix element in :attr:`P_codebook`
for pky (representing the concatenation of elements in :attr:`P_codebook` and :attr:`Y_codebook`)
pky_codebook: generate a codebook for the elements of the set PY (the product of set P and Y)
pi_pky_map: a map between P elements and PY elements
z_pky_map: a map between elements of the Z set and PY set
it has the form/template {ypattern:[pky_elements]}
z_pi_piy_map: a map between elements of the Z set and PY set
it has the form/template {ypattern:(pk, pky, pi)}
"""
def __init__(self):
# call super class
super().__init__()
self.P_codebook = None
self.P_codebook_rev = None
self.P_len = None
self.P_elems = None
self.P_numchar = None
self.f_transition = None
self.pky_codebook = None
self.pi_pky_map = None
self.z_pky_map = None
self.z_pi_piy_map = None
def setup_model(self, modelfeatures, states, L):
"""setup and create the model representation
Creates all maps and codebooks needed by the :class:`HOSemiCRFAD` class
Args:
modelfeatures: set of features defining the model
states: set of states (i.e. tags)
L: length of longest segment
"""
super().setup_model(modelfeatures, states, L)
def generate_instance_properties(self):
"""generate instance properties that will be later used by :class:`HOSemiCRFAD` class
"""
super().generate_instance_properties()
self.P_codebook = self.get_forward_states()
self.P_codebook_rev = self.get_P_codebook_rev()
self.P_len, self.P_elems, self.P_numchar = self.get_P_info()
self.f_transition = self.get_forward_transition()
self.pky_codebook = self.get_pky_codebook()
self.pi_pky_map = self.get_pi_pky_map()
self.z_pky_map, self.z_pi_piy_map = self.map_pky_z()
def get_forward_states(self):
"""create set of forward states (referred to set P) and map each element to unique code
P is set of proper prefixes of the elements in :attr:`Z_codebook` set
"""
Y_codebook = self.Y_codebook
Z_elems = self.Z_elems
Z_len = self.Z_len
P = {}
for z_patt in Z_elems:
elems = Z_elems[z_patt]
z_len = Z_len[z_patt]
for i in range(z_len - 1):
P["|".join(elems[: i + 1])] = 1
for y in Y_codebook:
P[y] = 1
# empty element
P[""] = 1
P_codebook = {s: i for (i, s) in enumerate(P)}
# print("P_codebook ", P_codebook)
return P_codebook
def get_P_codebook_rev(self):
"""generate reversed codebook of :attr:`P_codebook`
"""
P_codebook = self.P_codebook
P_codebook_rev = {code: pi for pi, code in P_codebook.items()}
return P_codebook_rev
def get_P_info(self):
"""get the properties of P set (proper prefixes)
"""
P_codebook = self.P_codebook
P_len = {}
P_numchar = {}
P_elems = {}
for pi in P_codebook:
elems = pi.split("|")
P_elems[pi] = elems
if pi == "":
P_len[pi] = 0
P_numchar[pi] = 0
else:
P_len[pi] = len(elems)
P_numchar[pi] = len(pi)
return (P_len, P_elems, P_numchar)
def get_forward_transition(self):
"""generate forward transition data structure
Main tasks:
- create a set PY from the product of P and Y sets
- for each element in PY, determine the longest suffix existing in set P
- include all this info in :attr:`f_transition` dictionary
"""
Y_codebook = self.Y_codebook
P_codebook = self.P_codebook
P_numchar = self.P_numchar
Z_numchar = self.Z_numchar
# pk_y= {}
# for p in P_codebook:
# for y in Y_codebook:
# pk_y[(p, y)] = 1
pk_y = {(p, y) for p in P_codebook for y in Y_codebook}
pk_y_suffix = {}
for p in P_codebook:
if p != "":
len_p = P_numchar[p]
for (pk, y) in pk_y:
ref_str = pk + "|" + y
if pk == "":
len_ref = Z_numchar[y] + 1
else:
len_ref = P_numchar[pk] + Z_numchar[y] + 1
start_pos = len_ref - len_p
if start_pos >= 0:
# check suffix relation
check = ref_str[start_pos:] == p
# check = self.check_suffix(p, ref_str)
if check:
if (pk, y) in pk_y_suffix:
pk_y_suffix[(pk, y)].append(p)
else:
pk_y_suffix[(pk, y)] = [p]
pk_y_suffix = self.keep_longest_elems(pk_y_suffix)
f_transition = {}
for (pk, y), pi in pk_y_suffix.items():
if pk == "":
elmkey = y
else:
elmkey = pk + "|" + y
if pi in f_transition:
f_transition[pi][elmkey] = (pk, y)
else:
f_transition[pi] = {elmkey: (pk, y)}
# print("f_transition ", f_transition)
return f_transition
def get_pky_codebook(self):
"""generate a codebook for the elements of the set PY (the product of set P and Y)
"""
f_transition = self.f_transition
pky_codebook = {}
counter = 0
for pi in f_transition:
for pky in f_transition[pi]:
pky_codebook[pky] = counter
counter += 1
return pky_codebook
def map_pky_z(self):
"""generate a map between elements of the Z set and PY set"""
f_transition = self.f_transition
Z_codebook = self.Z_codebook
# given that we demand to have a unigram label features then Z set will always contain Y elems
Z_numchar = self.Z_numchar
P_numchar = self.P_numchar
pky_codebook = self.pky_codebook
P_codebook = self.P_codebook
z_pi_piy = {}
z_pky = {}
for pi in f_transition:
for pky, pk_y_tup in f_transition[pi].items():
pk, y = pk_y_tup
# get number of characters in the pky
if pk == "":
len_pky = Z_numchar[y]
else:
# +1 is for the separator '|'
len_pky = P_numchar[pk] + Z_numchar[y] + 1
for z in Z_codebook:
len_z = Z_numchar[z]
# check suffix relation
start_pos = len_pky - len_z
if start_pos >= 0:
check = pky[start_pos:] == z
if check:
pky_c = pky_codebook[pky]
pk_c = P_codebook[pk]
if z in z_pky:
z_pky[z].append(pky_c)
z_pi_piy[z][0].append(pk_c)
z_pi_piy[z][1].append(pky_c)
z_pi_piy[z][2].append(P_codebook[pi])
else:
z_pky[z] = [pky_c]
z_pi_piy[z] = ([pk_c], [pky_c], [P_codebook[pi]])
return (z_pky, z_pi_piy)
def get_pi_pky_map(self):
""" generate map between P elements and PY elements
Main tasks:
- for every element in PY, determine the longest suffix in P
- determine the two components in PY (i.e. p and y element)
- represent this info in a dictionary that will be used for forward/alpha matrix
"""
f_transition = self.f_transition
pky_codebook = self.pky_codebook
P_codebook = self.P_codebook
pi_pky_map = {}
for pi in f_transition:
pi_pky_map[pi] = [[], []]
for pky, (pk, __) in f_transition[pi].items():
pi_pky_map[pi][0].append(pky_codebook[pky])
pi_pky_map[pi][1].append(P_codebook[pk])
# convert list to numpy arrays
# for i in range(2):
# pi_pky_map[pi][i] = numpy.array(pi_pky_map[pi][i])
# pi_pky_map[pi] = tuple(pi_pky_map[pi])
return pi_pky_map
def filter_activated_states(
self, activated_states, accum_active_states, curr_boundary
):
"""filter/prune states and y features
Args:
activaed_states: dictionary containing possible active states/y features
it has the form {patt_len:{patt_1, patt_2, ...}}
accum_active_states: dictionary of only possible active states by position
it has the form {pos_1:{state_1, state_2, ...}}
boundary: tuple (u,v) representing the current boundary in the sequence
"""
Z_elems = self.Z_elems
filtered_activestates = {}
# generate partition boundaries
depth_node_map = {}
generate_partitions(
curr_boundary, self.L, self.max_patt_len, {}, depth_node_map, None
)
partition_boundaries = generate_partition_boundaries(depth_node_map)
for z_len in activated_states:
if z_len == 1:
continue
if z_len in partition_boundaries:
partitions = partition_boundaries[z_len]
filtered_activestates[z_len] = set()
for partition in partitions:
for z_patt in activated_states[z_len]:
check = True
zelems = Z_elems[z_patt]
for i in range(z_len):
bound = partition[i]
if zelems[i] not in accum_active_states[bound]:
check = False
break
if check:
filtered_activestates[z_len].add(z_patt)
return filtered_activestates
class HOSemiCRFAD(LCRF):
"""higher-order semi-CRF model that uses algorithmic differentiation in gradient computation
Args:
model: an instance of :class:`HOSemiCRFADModelRepresentation` class
seqs_representer: an instance of :class:`SeqsRepresenter` class
seqs_info: dictionary holding sequences info
Keyword Arguments:
load_info_fromdisk: integer from 0 to 5 specifying number of cached data
to be kept in memory. 0 means keep everything while
5 means load everything from disk
Attributes:
model: an instance of :class:`HOSemiCRFADModelRepresentation` class
weights: a numpy vector representing feature weights
seqs_representer: an instance of :class:`pyseqlab.feature_extraction.SeqsRepresenter` class
seqs_info: dictionary holding sequences info
beam_size: determines the size of the beam for state pruning
fun_dict: a function map
def_cached_entities: a list of the names of cached entities sorted (descending)
based on estimated space required in memory
"""
def __init__(self, model, seqs_representer, seqs_info, load_info_fromdisk=5):
super().__init__(model, seqs_representer, seqs_info, load_info_fromdisk)
def cached_entitites(self, load_info_fromdisk):
"""construct list of names of cached entities in memory
"""
def_cached_entities = super().cached_entitites(load_info_fromdisk)
inmemory_info = ["alpha", "Z", "beta", "fpotential"]
def_cached_entities += inmemory_info
return def_cached_entities
def compute_fpotential(self, w, active_features):
"""compute the potential of active features in a specified boundary
Args:
w: weight vector (numpy vector)
active_features: dictionary of activated features in a specified boundary
"""
model = self.model
pky_codebook = model.pky_codebook
z_pky_map = model.z_pky_map
f_potential = numpy.zeros(len(pky_codebook))
# to consider caching the w_indx and fval as in cached_pf
for z in active_features:
w_indx, f_val = active_features[z]
potential = numpy.dot(w[w_indx], f_val)
# get all pky's in coded format where z maintains a suffix relation with them
pky_c_list = z_pky_map[z]
f_potential[pky_c_list] += potential
return f_potential
def compute_forward_vec(self, w, seq_id):
"""compute the forward matrix (alpha matrix)
Args:
w: weight vector (numpy vector)
seq_id: integer representing unique id assigned to the sequence
.. note::
activefeatures need to be loaded first in :attr:`seqs.info`
"""
model = self.model
pi_pky_map = model.pi_pky_map
P_len = model.P_len
P_codebook = model.P_codebook
T = self.seqs_info[seq_id]["T"]
L = self.model.L
activefeatures = self.seqs_info[seq_id]["activefeatures"]
alpha = numpy.ones((T + 1, len(P_codebook)), dtype="longdouble") * (-numpy.inf)
alpha[0, P_codebook[""]] = 0
fpotential_perboundary = {}
for j in range(1, T + 1):
accumulator = (
numpy.ones((len(P_codebook), L), dtype="longdouble") * -numpy.inf
)
for d in range(L):
u = j - d
if u <= 0:
break
v = j
f_potential = self.compute_fpotential(w, activefeatures[u, v])
fpotential_perboundary[u, v] = f_potential
for pi in pi_pky_map:
if j >= P_len[pi]:
pi_c = P_codebook[pi]
pky_c_list, pk_c_list = pi_pky_map[pi]
vec = f_potential[pky_c_list] + alpha[u - 1, pk_c_list]
accumulator[pi_c, d] = vectorized_logsumexp(vec)
for pi in pi_pky_map:
if j >= P_len[pi]:
pi_c = P_codebook[pi]
if L > 1:
alpha[j, pi_c] = vectorized_logsumexp(accumulator[pi_c, :])
else:
alpha[j, pi_c] = accumulator[pi_c, 0]
self.seqs_info[seq_id]["fpotential"] = fpotential_perboundary
return alpha
def compute_backward_vec(self, w, seq_id):
"""compute the backward matrix (beta matrix)
Args:
w: weight vector (numpy vector)
seq_id: integer representing unique id assigned to the sequence
.. note::
fpotential per boundary dictionary should be available in :attr:`seqs.info`
"""
model = self.model
pi_pky_map = model.pi_pky_map
P_codebook = model.P_codebook
len_P = len(P_codebook)
T = self.seqs_info[seq_id]["T"]
L = model.L
fpotential_perboundary = self.seqs_info[seq_id]["fpotential"]
beta = numpy.ones((T + 2, len(P_codebook)), dtype="longdouble") * (-numpy.inf)
beta[T + 1, :] = 0
for j in reversed(range(1, T + 1)):
accum_mat = numpy.ones((len_P, L), dtype="longdouble") * (-numpy.inf)
for d in range(L):
track_comp = numpy.ones((len_P, len_P), dtype="longdouble") * (
-numpy.inf
)
u = j
v = j + d
if v > T:
break
f_potential = fpotential_perboundary[u, v]
for pi in pi_pky_map:
pi_c = P_codebook[pi]
pky_c_list, pk_c_list = pi_pky_map[pi]
vec = f_potential[pky_c_list] + beta[v + 1, pi_c]
track_comp[pk_c_list, pi_c] = vec
for p_c in P_codebook.values():
accum_mat[p_c, d] = vectorized_logsumexp(track_comp[p_c, :])
for p_c in P_codebook.values():
beta[u, p_c] = vectorized_logsumexp(accum_mat[p_c, :])
return beta
def compute_marginals(self, seq_id):
""" compute the marginal (i.e. probability of each y pattern at each position)
Args:
seq_id: integer representing unique id assigned to the sequence
.. note::
- fpotential per boundary dictionary should be available in :attr:`seqs.info`
- alpha matrix should be available in :attr:`seqs.info`
- beta matrix should be available in :attr:`seqs.info`
- Z (i.e. P(x)) should be available in :attr:`seqs.info`
"""
model = self.model
Z_codebook = model.Z_codebook
z_pi_piy = model.z_pi_piy_map
T = self.seqs_info[seq_id]["T"]
L = self.model.L
alpha = self.seqs_info[seq_id]["alpha"]
beta = self.seqs_info[seq_id]["beta"]
Z = self.seqs_info[seq_id]["Z"]
fpotential_perboundary = self.seqs_info[seq_id]["fpotential"]
P_marginals = numpy.zeros(
(L, T + 1, len(self.model.Z_codebook)), dtype="longdouble"
)
for j in range(1, T + 1):
for d in range(L):
u = j
v = j + d
if v > T:
break
boundary = (u, v)
f_potential = fpotential_perboundary[boundary]
for z in Z_codebook:
pi_c, piy_c, pk_c = z_pi_piy[z]
numerator = (
alpha[u - 1, pi_c] + f_potential[piy_c] + beta[v + 1, pk_c]
)
P_marginals[d, j, Z_codebook[z]] = numpy.exp(
vectorized_logsumexp(numerator) - Z
)
return P_marginals
def compute_feature_expectation(self, seq_id, P_marginals, grad):
"""compute the features expectations (i.e. expected count of the feature based on learned model)
Args:
seq_id: integer representing unique id assigned to the sequence
P_marginals: probability matrix for y patterns at each position in time
grad: numpy vector with dimension equal to the weight vector. It represents the gradient
that will be computed using the feature expectation and the global features of the sequence
.. note::
- activefeatures (per boundary) dictionary should be available in :attr:`seqs.info`
- P_marginal (marginal probability matrix) should be available in :attr:`seqs.info`
"""
activefeatures = self.seqs_info[seq_id]["activefeatures"]
Z_codebook = self.model.Z_codebook
for boundary, features_dict in activefeatures.items():
u, v = boundary
d = v - u
for z_patt in features_dict:
w_indx, f_val = features_dict[z_patt]
grad[w_indx] += f_val * P_marginals[d, u, Z_codebook[z_patt]]
def prune_states(self, score_vec, beam_size):
"""prune states that fall off the specified beam size
Args:
score_vec: score matrix
beam_size: specified size of the beam (integer)
"""
P_codebook_rev = self.model.P_codebook_rev
P_elems = self.model.P_elems
# using argpartition as better alternative to argsort
indx_partitioned_pi = numpy.argpartition(-score_vec, beam_size)
# identify top-k states/pi
indx_topk_pi = indx_partitioned_pi[:beam_size]
# get topk states
topk_pi = {P_codebook_rev[indx] for indx in indx_topk_pi}
topk_states = {P_elems[pi][-1] for pi in topk_pi}
return topk_states
def viterbi(self, w, seq_id, beam_size, stop_off_beam=False, y_ref=[], K=1):
"""decode sequences using viterbi decoder
Args:
w: weight vector (numpy vector)
seq_id: integer representing unique id assigned to the sequence
beam_size: integer representing the size of the beam
Keyword Arguments:
stop_off_beam: boolean indicating if to stop when the reference state \
falls off the beam (used in perceptron/search based learning)
y_ref: reference sequence list of labels (used while learning)
K: integer indicating number of decoded sequences required (i.e. top-k list)
A* searcher with viterbi will be used to generate k-decoded list
"""
model = self.model
P_elems = model.P_elems
pi_pky_map = model.pi_pky_map
P_codebook = model.P_codebook
P_codebook_rev = model.P_codebook_rev
L = model.L
len_P = len(P_codebook)
num_states = model.num_states
T = self.seqs_info[seq_id]["T"]
# records max score at every time step
delta = numpy.ones((T + 1, len(P_codebook)), dtype="longdouble") * (-numpy.inf)
pi_mat = numpy.ones((len_P, L), dtype="longdouble") * (-numpy.inf)
# the score for the empty sequence at time 0 is 1
delta[0, P_codebook[""]] = 0
back_track = {}
# records where violation occurs -- it is 1-based indexing
viol_index = []
if beam_size == num_states:
# case of exact search and decoding
l = {}
l["activefeatures"] = (seq_id,)
self.check_cached_info(seq_id, l)
active_features = self.seqs_info[seq_id]["activefeatures"]
for j in range(1, T + 1):
# reset pi_mat at every loop
pi_mat.fill(-numpy.inf)
backpointer = {}
for d in range(L):
u = j - d
if u <= 0:
break
v = j
boundary = (u, v)
# vector of size len(pky)
f_potential = self.compute_fpotential(w, active_features[boundary])
for pi in pi_pky_map:
pi_c = P_codebook[pi]
pky_c_list, pk_c_list = pi_pky_map[pi]
vec = f_potential[pky_c_list] + delta[u - 1, pk_c_list]
# print("f_potential[pky_c_list] ", f_potential[pky_c_list])
# print("delta[u-1, pk_c_list] ", delta[u-1, pk_c_list])
# print("vec ", vec)
pi_mat[pi_c, d] = numpy.max(vec)
argmax_indx = numpy.argmax(vec)
# print("argmax chosen ", argmax_ind)
pk_c_max = pk_c_list[argmax_indx]
# print('pk_c ', pk_c)
pk = P_codebook_rev[pk_c_max]
y = P_elems[pk][-1]
backpointer[d, pi_c] = (pk_c_max, y)
# print("backpointer ")
# print(backpointer)
# print("pi_mat")
# print(pi_mat)
# get the max for each pi across all segment lengths
for pi in pi_pky_map:
pi_c = P_codebook[pi]
delta[j, pi_c] = numpy.max(pi_mat[pi_c, :])
argmax_indx =
|
numpy.argmax(pi_mat[pi_c, :])
|
numpy.argmax
|
import numpy as np
import numpy.random as npr
import time#, timer
from . import gelmanrubin as gr
#reload(gr)
#import python_models as mc
#import models_c as mc
import multiprocessing as mp
def calcModel(nchains, functype, myfuncs, pedit, nextp, iortholist, funcx, cummodels, numparams, j, iblock=None, chains=None):
'''
Compute model light curve by combining model components. Also returns correlated noise parameters.
'''
#Build final model from model components
ymodels = np.ones((nchains, fit[j].nobj))
noisepars = [[] for i in range(nchains)]
k = 0
if chains == None:
chains = range(nchains)
if iblock == None:
iblock = range(cummodels[j],cummodels[j+1])
for i in range(cummodels[j],cummodels[j+1]):
if iblock.__contains__(i):
for n in chains:
if functype[i] == 'ortho':
#MODIFY COPY OF nextp ONLY
pedit[n,iortholist] = myfuncs[i](pedit[n,iortholist], funcx[i], fit[j].etc[k])
elif (functype[i] == 'ipmap') or (functype[i] == 'spline'):
ymodels[n] *= myfuncs[i](pedit[n,numparams[i]:numparams[i+1]], funcx[i], ymodels[n])
elif functype[i] == 'posoffset':
# Record change in Position 0 => cannot orthogonalize position parameters
ymodels[n] *= myfuncs[i](nextp[n,numparams[i]:numparams[i+1]], funcx[i], fit[j].etc[k])
elif hasattr(fit[j], 'timebins') and (functype[i] == 'ecl/tr'
or functype[i] == 'ramp'
or functype[i] == 'sinusoidal'):
# Average over high-resolution model
hiresmodel = myfuncs[i](pedit[n,numparams[i]:numparams[i+1]], funcx[i], fit[j].etc[k])
if len(fit[j].timebins) == fit[j].nobj:
for tb in range(len(fit[j].timebins)):
ymodels[n,tb] *= np.mean(hiresmodel[fit[j].timebins[tb]])
else:
for tb in range(len(fit[j].timebinsuc)):
ymodels[n,tb] *= np.mean(hiresmodel[fit[j].timebinsuc[tb]])
elif functype[i] == 'noise':
noisepars[n] = pedit[n,numparams[i]:numparams[i+1]]
else:
ymodels[n] *= myfuncs[i](pedit[n,numparams[i]:numparams[i+1]], funcx[i], fit[j].etc[k])
k += 1
return ymodels, noisepars
# Calculate chi^2
def calcChisq(y, sigma, ymodels, nchains, nextp, j, noisepars, isrednoise, wavelet, noisefunc, chains=None):
'''
Compute chi-squared with priors.
'''
if chains == None:
chains = range(nchains)
chi2 = np.zeros(nchains)
for n in chains:
if isrednoise == False:
#chi2[n] = mc.chisq(ymodels[n], y, sigma)
chi2[n] += np.sum((ymodels[n] - y)**2 / sigma**2)
else:
chi2[n] = noisefunc(noisepars[n], ymodels[n]-y, wavelet)
# Apply prior, if one exists
if len(fit[j].ipriors) > 0:
pbar = fit[j].priorvals[:,0] #prior mean
psigma = np.zeros(len(pbar)) #prior standard deviation
# Determine psigma based on which side of asymmetric Gaussian nextp is on
for i in range(len(fit[j].ipriors)):
if nextp[n,fit[j].ipriors[i]] < pbar[i]:
psigma[i] = fit[j].priorvals[i,1]
else:
psigma[i] = fit[j].priorvals[i,2]
#chi2[n] += fit[j].nobj*((nextp[n,fit[j].ipriors[i]] - pbar[i])/psigma[i])**2
chi2[n] += ((nextp[n,fit[j].ipriors[i]] - pbar[i])/psigma[i])**2
return chi2
def demc_block(y, pars, pmin, pmax, stepsize, numit, sigma, numparams, cummodels, functype, myfuncs, funcx, iortholist, fits, gamma=None, isGR=True, ncpu=1):
"""
This function uses a differential evolution Markov chain with block updating to assess uncertainties.
PARAMETERS
----------
y: Array containing dependent data
Params: Array of initial guess for parameters
#Pmin: Array of parameter minimum values
#Pmax: Array of parameter maximum values
stepsize: Array of 1-sigma change in parameter per iteration
Numit: Number of iterations to perform
Sigma: Standard deviation of data noise in y
Numparams: Number of parameters for each model
Cummodels: Cumulative number of models used
Functype: Define function type (eclipse, ramp, ip, etc), see models.py
Myfuncs: Pointers to model functions
Funcx: Array of x-axis values for myfuncs
fit: List of fit objects
gamma: Multiplcation factor in parameter differential, establishes acceptance rate
OUTPUTS
-------
This function returns an array of the best fitting parameters,
an array of all parameters over all iterations, and numaccept.
REFERENCES
----------
<NAME>. <NAME>, "Genetic algorithms and Markov Chain Monte Carlo: Differential Evolution Markov Chain makes Bayesian computing easy," Biometrics, 2006.
HISTORY
-------
Adapted from mcmc.py
<NAME>, UChicago August 2012
"""
global fit
fit = fits
params = np.copy(pars)
nchains, nump = params.shape
nextp = np.copy(params) #Proposed parameters
bestp = np.copy(params[0]) #Best-fit parameters
pedit = np.copy(params) #Editable parameters
numaccept = 0
allparams = np.zeros((nump, nchains, numit))
inotfixed = np.where(stepsize != 0)[0]
ishare = np.where(stepsize < 0)[0]
#ifree = np.where(stepsize > 0)[0]
outside = np.zeros((nchains, nump))
numevents = len(fit)
intsteps = np.min((numit/5,1e5))
isrednoise = False
wavelet = None
noisefunc = None
#UPDATE PARAMTER(S) EQUAL TO OTHER PARAMETER(S)
if (ishare.size > 0):
for s in range(ishare.size):
params[:,ishare[s]] = params[:,int(abs(stepsize[ishare[s]])-1)]
#Define blocks
blocks = []
for j in range(numevents):
#Build list of blocks
blocks = np.concatenate((blocks, fit[j].blocks))
for i in range(cummodels[j],cummodels[j+1]):
if functype[i] == 'noise':
# Set up for modified chi-squared calculation using correlated noise
isrednoise = True
wavelet = fit[j].etc[k]
noisefunc = myfuncs[i]
blocks = blocks.astype(int)
iblocks = []
eps = []
numblocks = blocks.max() + 1
numbp = np.zeros(numblocks)
ifree = [[] for i in range(numblocks)]
for b in range(numblocks):
#Map block indices
whereb = np.where(blocks == b)[0]
iblocks.append(whereb)
#Locate indices of free parameters in each block
for w in whereb:
ifree[b] = np.concatenate((ifree[b],numparams[w]+np.where(stepsize[numparams[w]:numparams[w+1]] > 0)[0])).astype(int)
#Calculate number of free parameters per block
numbp[b] += len(ifree[b])
eps.append(npr.normal(0, stepsize[ifree[b]]/100., [numit,numbp[b]]))
print("Number of free parameters per block:")
print(numbp)
numa = np.zeros(numblocks)
if gamma == None:
gamma = 2.38/np.sqrt(2.*numbp)
print("gamma:")
print(gamma)
#Calc chi-squared for model type using current params
currchisq = np.zeros(nchains)
currmodel = [[] for i in range(numevents)]
for j in range(numevents):
currmodel[j], noisepars = calcModel(nchains, functype, myfuncs, pedit, params, iortholist[j],
funcx, cummodels, numparams, j)
currchisq += calcChisq(y[j], sigma[j], currmodel[j], nchains, params, j, noisepars, isrednoise, wavelet, noisefunc)
bestchisq = currchisq[0]
#GENERATE RANDOM NUMBERS FOR MCMC
numnotfixed = len(inotfixed)
unif = npr.rand(numit,nchains)
randchains = npr.randint(0,nchains,[numit,nchains,2])
#START TIMER
clock = timer.Timer(numit,progress = np.arange(0.05,1.01,0.05))
#Run Differential Evolution Monte Carlo algorithm 'numit' times
for m in range(numit):
#Select next event (block) to update
b = m % numblocks
#Remove model component(s) that are taking a step
pedit = np.copy(params)
nextmodel = currmodel[:]
for j in range(numevents):
ymodels, noisepars = calcModel(nchains, functype, myfuncs, pedit, params, iortholist[j],
funcx, cummodels, numparams, j, iblocks[b])
nextmodel[j] = np.divide(currmodel[j],ymodels)
#Generate next step using differential evolution
for n in range(nchains):
rand1, rand2 = randchains[m,n]
while rand1 == n or rand2 == n or rand1 == rand2:
rand1, rand2 = npr.randint(0,nchains,2)
nextp[n,ifree[b]] = params[n,ifree[b]] + gamma[b]*(params[rand1,ifree[b]]-params[rand2,ifree[b]]) + eps[b][m]
#CHECK FOR NEW STEPS OUTSIDE BOUNDARIES
ioutside = np.where(np.bitwise_or(nextp[n] < pmin, nextp[n] > pmax))[0]
if (len(ioutside) > 0):
nextp[n,ioutside] = np.copy(params[n,ioutside])
outside[n,ioutside] += 1
#UPDATE PARAMTER(S) EQUAL TO OTHER PARAMETER(S)
if (ishare.size > 0):
for s in range(ishare.size):
nextp[:,ishare[s]] = nextp[:,int(abs(stepsize[ishare[s]])-1)]
#COMPUTE NEXT CHI SQUARED AND ACCEPTANCE VALUES
pedit = np.copy(nextp)
nextchisq = np.zeros(nchains)
for j in range(numevents):
ymodels, noisepars = calcModel(nchains, functype, myfuncs, pedit, params, iortholist[j], funcx, cummodels, numparams, j, iblocks[b])
nextmodel[j] = np.multiply(nextmodel[j],ymodels)
nextchisq += calcChisq(y[j], sigma[j], nextmodel[j], nchains, params, j, noisepars, isrednoise, wavelet, noisefunc)
#CALCULATE ACCEPTANCE PROBABILITY
accept = np.exp(0.5 * (currchisq - nextchisq))
#print(b,currchisq[0], nextchisq[0], accept[0])
for n in range(nchains):
if accept[n] >= 1:
#ACCEPT BETTER STEP
numaccept += 1
numa[b] += 1
params[n] = np.copy(nextp[n])
currchisq[n] = np.copy(nextchisq[n])
if (currchisq[n] < bestchisq):
bestp = np.copy(params[n])
bestchisq = np.copy(currchisq[n])
elif unif[m,n] <= accept[n]:
#ACCEPT WORSE STEP
numaccept += 1
numa[b] += 1
params[n] = np.copy(nextp[n])
currchisq[n] = np.copy(nextchisq[n])
allparams[:,:,m] = params.T
#PRINT INTERMEDIATE INFO
if ((m+1) % intsteps == 0) and (m > 0):
print("\n" + time.ctime())
#print("Number of times parameter tries to step outside its prior:")
#print(outside)
print("Current Best Parameters: ")
print(bestp)
#Apply Gelman-Rubin statistic
if isGR:
#Check for no accepted steps in each chain
#stdev = np.std(allparams[inotfixed],axis=1)
#ichain = np.where(stdev > 0.)[0]
#Call test
#psrf, meanpsrf = gr.convergetest(allparams[inotfixed,ichain,:m+1], len(ichain))
psrf, meanpsrf = gr.convergetest(allparams[inotfixed,:,:m+1], nchains)
numconv = np.sum(np.bitwise_and(psrf < 1.01, psrf >= 1.00))
print("Gelman-Rubin statistic for free parameters:")
print(psrf)
if numconv == numnotfixed: #and m >= 1e4:
print("All parameters have converged to within 1% of unity. Halting MCMC.")
allparams = allparams[:,:,:m+1]
break
clock.check(m+1)
print("Acceptance rate per block (%):")
print(100.*numa*numblocks/numit/nchains)
allparams = np.reshape(allparams,(nump, (m+1)*nchains))
return allparams, bestp, numaccept, (m+1)*nchains
#****************************************************************
def calcChi2(nchains, functype, myfuncs, pedit, nextp, iortholist, funcx, cummodels, numparams, j, isrednoise, wavelet, noisefunc, systematics, chains=None):
'''
Compute model light curve by combining model components.
'''
#Build final model from model components
ymodels = np.ones((nchains, fit[j].nobj))
noisepars = [[] for i in range(nchains)]
k = 0
if chains == None:
chains = range(nchains)
for i in range(cummodels[j],cummodels[j+1]):
for n in chains:
if functype[i] == 'ortho':
#MODIFY COPY OF nextp ONLY
pedit[n,iortholist] = myfuncs[i](pedit[n,iortholist], funcx[i], fit[j].etc[k])
elif (functype[i] == 'ipmap') or (functype[i] == 'spline'):
ymodels[n] *= myfuncs[i](pedit[n,numparams[i]:numparams[i+1]], funcx[i], ymodels[n])
elif functype[i] == 'posoffset':
# Record change in Position 0 => cannot orthogonalize position parameters
ymodels[n] *= myfuncs[i](nextp[n,numparams[i]:numparams[i+1]], funcx[i], fit[j].etc[k])
elif hasattr(fit[j], 'timebins') and (functype[i] == 'ecl/tr'
or functype[i] == 'ramp'
or functype[i] == 'sinusoidal'):
# Average over high-resolution model
hiresmodel = myfuncs[i](pedit[n,numparams[i]:numparams[i+1]], funcx[i], fit[j].etc[k])
if len(fit[j].timebins) == fit[j].nobj:
for tb in range(len(fit[j].timebins)):
ymodels[n,tb] *= np.mean(hiresmodel[fit[j].timebins[tb]])
else:
for tb in range(len(fit[j].timebinsuc)):
ymodels[n,tb] *= np.mean(hiresmodel[fit[j].timebinsuc[tb]])
elif functype[i] == 'noise':
noisepars[n] = pedit[n,numparams[i]:numparams[i+1]]
else:
ymodels[n] *= myfuncs[i](pedit[n,numparams[i]:numparams[i+1]], funcx[i], fit[j].etc[k])
k += 1
# Calculate chi^2
chi2 = np.zeros(nchains)
for n in chains:
if isrednoise == False:
#chi2[n] = mc.chisq(ymodels[n]*systematics[n][j], data[j], unc[j])
chi2[n] = np.sum((ymodels[n]*systematics[n][j] - data[j])**2 / unc[j]**2)
else:
chi2[n] = noisefunc(noisepars[n], ymodels[n]*systematics[n][j]-data[j], wavelet)
# Apply prior, if one exists
if len(fit[j].ipriors) > 0:
pbar = fit[j].priorvals[:,0] #prior mean
psigma = np.zeros(len(pbar)) #prior standard deviation
# Determine psigma based on which side of asymmetric Gaussian nextp is on
for i in range(len(fit[j].ipriors)):
if nextp[n,fit[j].ipriors[i]] < pbar[i]:
psigma[i] = fit[j].priorvals[i,1]
else:
psigma[i] = fit[j].priorvals[i,2]
#chi2[n] += fit[j].nobj*((nextp[n,fit[j].ipriors[i]] - pbar[i])/psigma[i])**2
chi2[n] += ((nextp[n,fit[j].ipriors[i]] - pbar[i])/psigma[i])**2
return chi2
#
def writeChi2(chi2):
'''
Write models after multiprocessing.
'''
global nextchisq
nextchisq += chi2
return
def demc(y, pars, pmin, pmax, stepsize, numit, sigma, numparams, cummodels, functype, myfuncs, funcx, iortholist, nights, fits, gamma=None, isGR=True, ncpu=1):
"""
This function uses a differential evolution Markov chain to assess uncertainties.
PARAMETERS
----------
y: Array containing dependent data
Params: Array of initial guess for parameters
#Pmin: Array of parameter minimum values
#Pmax: Array of parameter maximum values
stepsize: Array of 1-sigma change in parameter per iteration
Numit: Number of iterations to perform
Sigma: Standard deviation of data noise in y
Numparams: Number of parameters for each model
Cummodels: Cumulative number of models used
Functype: Define function type (eclipse, ramp, ip, etc), see models.py
Myfuncs: Pointers to model functions
Funcx: Array of x-axis values for myfuncs
fit: List of fit objects
gamma: Multiplcation factor in parameter differential, establishes acceptance rate
OUTPUTS
-------
This function returns an array of the best fitting parameters,
an array of all parameters over all iterations, and numaccept.
REFERENCES
----------
<NAME>, "Genetic algorithms and Markov Chain Monte Carlo: Differential Evolution Markov Chain makes Bayesian computing easy," Biometrics, 2006.
HISTORY
-------
Adapted from mcmc.py
<NAME>, UChicago August 2012
Multiplied prior by number of points in fit
January 2014
"""
global nextchisq, fit, data, unc
fit = fits
data = y
unc = sigma
params = np.copy(pars)
nchains, nump = params.shape
nextp = np.copy(params) #Proposed parameters
bestp = np.copy(params[0]) #Best-fit parameters
pedit = np.copy(params) #Editable parameters
numaccept = 0
#allparams must be 64-bit!
allparams =
|
np.zeros((nump, nchains, numit))
|
numpy.zeros
|
import matplotlib.pyplot as plt
import pandas as pd
from keras.layers import TimeDistributed
from keras.layers import RepeatVector
from keras.layers import LSTM
from keras.layers import Dense
from keras.models import Sequential
import numpy as np
'''
LSTM(Long short-term memory) を用いて、時系列データの予測を行います。
PythonのKerasを使います。
'''
'''簡単な例'''
#まずは、簡単な例で試します。 3つの入力から、2つの値を出力(予測)することにします。
# 学習用データ。xが入力、yが出力(答え)です。
x = np.array([[10, 20, 30], [20, 30, 40], [30, 40, 50], [40, 50, 60]])#[50,60,70],[60,70,80]
y = np.array([[40, 50], [50, 60], [60, 70], [70, 80]])#[80,90],[90,100]
# 行列のフォーマット変更。
# LSTMは、入力フォーマットを[サンプルの数, 入力のステップ数(この場合は3), features]とする必要があるためです。
x = x.reshape((x.shape[0], x.shape[1], 1))
y = y.reshape((y.shape[0], y.shape[1], 1))
#次にネットワークを構築します。
m = Sequential()
m.add(LSTM(100, activation='relu', input_shape=(3, 1)))
m.add(RepeatVector(2))
m.add(LSTM(100, activation='relu', return_sequences=True))
m.add(TimeDistributed(Dense(1)))
m.compile(optimizer='adam', loss='mse')
'''
少し解説します。
1行目:Sequentialは、あるレイヤの全ノードと、次レイヤの全ノードをつなぐDNNのモデルです。
2行目:DNNの第一レイヤとして、LSTMを追加します。第一引数は出力の次元数です。ここでは100としています。activationは活性化関数で、ここではReLUを使うように設定しています。input_shapeは、入力データのフォーマットです。
3行目:RepeatVectorにより、入力を繰り返します。ここでの繰り返し回数は、予測範囲(今回は2データ)となります。
4行目:再びLSTM。ただし、ここではreturn_sequences = Trueを指定します。
5行目:TimeDistributedを指定し、かつ、Dense(1)で、出力の次元数を「1」に指定します。
6行目:最後にcompileメソッドで、学習時の最適化手法や、損失関数を指定します。
ここでは最適化手法としてAdamを、損失関数としてMSE(Mean Squared Error 平均二乗誤差)を指定します。
'''
#fitメソッドで学習を行います。 学習。時間が少しかかる可能性があります。
##m.fit(x, y, epochs=1000, verbose=0)
#学習済みのモデルに、[50, 60, 70]という入力を与えて、結果がどうなるかを見てみます。 理想では[80, 90]となればOKです。
##x_input = np.array([50, 60, 70])
##x_input = x_input.reshape((1, 3, 1))
##yhat = m.predict(x_input)
##print(yhat)
'''
結果は以下です。
[[[82.211136]
[93.43616]]]
少しズレがありますが、まあまあですね。
'''
'''AirPassengers.csvで実験'''
#次に、AirPassengers.csvのデータで試してみます。 このデータは色々なところで勉強用に使われている。
#まずはデータを読み込みます。
df = pd.read_csv('AirPassengers.csv', index_col='Month', dtype={1: 'float'})
ts = df['#Passengers']
#学習用データxと、回答データyを用意します。 AirPassengersのデータは、「毎月の乗客数」です。
# 10年分くらいのデータがあるので、その一部を学習用に使います。
# 2年間のデータ(24データ)を用いて、次に一年(12データ)を予測するように学習します。
x = [] # train
y = [] # test (answer)
for i in range(0, 72):
tmpX = []
for j in range(0, 24): # 2年間のデータ(24データ)
tmpX.append(ts[i+j])
x.append(tmpX)
tmpY = []
for j in range(0, 12): # 1年間のデータ(12データ)
tmpY.append(ts[24+i+j])
y.append(tmpY)
#学習用データxと、回答データyができたので、numpy配列にして、LSTM用にreshapeします。
x = np.array(x)
y = np.array(y)
x = x.reshape((x.shape[0], x.shape[1], 1))
y = y.reshape((y.shape[0], y.shape[1], 1))
#ネットワークを組み、学習します。
m = Sequential()
# 入力データ数が24なので、input_shapeの値が(24,1)です。
m.add(LSTM(100, activation='relu', input_shape=(24, 1)))
# 予測範囲は12ステップなので、RepeatVectoorに12を指定する必要があります。
m.add(RepeatVector(12))
m.add(LSTM(100, activation='relu', return_sequences=True))
m.add(TimeDistributed(Dense(1)))
m.compile(optimizer='adam', loss='mse')
m.fit(x, y, epochs=1000, verbose=0)
#いざ、予測してみます。
# データ60番~83番から、次の一年(84番~95番)を予測
input = np.array(ts[60:84])
input = input.reshape((1, 24, 1))
yhat = m.predict(input)
# 可視化用に、予測結果yhatを、配列predictに格納
predict = []
for i in range(0, 12):
predict.append(yhat[0][i])
# 比較するために実データをプロット
plt.plot(ts)
# 予測したデータをプロット
xdata = np.arange(84, 96, 1)
plt.plot(xdata, predict, 'r')
# 良い感じに予測できています。
# さらに次の一年を予測してみます。
input = np.array(ts[72:96])
input = input.reshape((1, 24, 1))
yhat = m.predict(input)
predict = []
for i in range(0, 12):
predict.append(yhat[0][i])
plt.plot(ts)
xdata =
|
np.arange(96, 108, 1)
|
numpy.arange
|
"""
stereo matching tools
Copyright (C) 2017-2018, <NAME> <<EMAIL>>
"""
from __future__ import print_function
import numpy as np
# in case numba jit is not installed
try:
from numba import jit
except:
print('WARNING: numba package is not installed')
def jit(x):
return x
# cost volume functions
def censustransform_64(img, cw=5, cp=None, sep=1):
'''
Efficiently compute the census transform (CT) of img
using windows limited to 8 * 8 (cw<=8)
Args:
img: numpy array containing the input image
cw: size of the census window cw*cw-1 <= 64
cp: optional image with centralpixel values of all pixels,
useful for implementing the modified census transform
sep: optional control the spacing of the CT samples (default 1)
Returns:
a numpy array containing the CT at each pixel packed as a uint64 image
derived from: http://stackoverflow.com/questions/38265364/census-transform-in-python-opencv
'''
if cw > 8:
printf('census window cannot be larger than 8x8')
cw = min(cw,8)
hcw = int(cw/2)
# Initialize 64bit output array
census = np.zeros(img.shape, dtype='uint64')
# Center pixels
if cp is None:
cp = img
# Offsets of non-central pixels
offsets = [(u-hcw, v-hcw) for v in range(cw)
for u in range(cw)
if not u == hcw == v]
# Fill census bitstring
for u,v in offsets:
census = (census << 1) | (np.roll(img,(-v*sep,-u*sep), axis=(0,1)) >= cp)
return census
def censustransform(img, cw=5, cp=None, sep=1):
'''
Efficiently compute the census transform (CT) of img
sing windows of size cw * cw
Args:
img: numpy array containing the input image
cw: size of the census window, the transform will have cw*cw-1 bits
cp: optional image with centralpixel values of all pixels,
useful for implementing the modified census transform
sep: optional control the spacing of the CT samples (default 1)
Returns:
a numpy array containing the CT at each pixel packed as as many
uint64 image planes as needed to represent the (cw*cw-1) bits
derived from: http://stackoverflow.com/questions/38265364/census-transform-in-python-opencv
'''
hcw = int(cw/2)
# Initialize 64bit output array
census = None
# Center pixel values
if cp is None:
cp = img
# Offsets of non-central pixels
offsets = [(u-hcw, v-hcw) for v in range(cw)
for u in range(cw)
if not u == hcw == v]
def chunks(l, n):
"""Yield successive n-sized chunks from l."""
for i in range(0, len(l), n):
yield l[i:i + n]
for Loffsets in chunks(offsets,64):
# Initialize 64bit output array
Lcensus = np.zeros(img.shape, dtype='uint64')
# Fill census bitstring
for u,v in Loffsets:
Lcensus = (Lcensus << 1) | (np.roll(img,(-v*sep,-u*sep), axis=(0,1)) >= cp)
if census is None:
census = Lcensus
else: # concatenate along third axis if more than 64 bits are needed
if Lcensus.ndim==2:
Lcensus = np.expand_dims(Lcensus,axis=2)
if census.ndim==2:
census = np.expand_dims(census,axis=2)
census = np.dstack((census,Lcensus))
return census
def countbits(n):
'''
Count the number of bits set for all the elements of the numpy array up to uint64
https://stackoverflow.com/questions/9829578/fast-way-of-counting-non-zero-bits-in-positive-integer
Args:
n: numpy array of integer type (interpreted as uint64)
Returns:
numpy array with the number of bits for each element of n
'''
import numpy as np
if type(n) == np.ndarray: # force type in case of np.uint32
n = n.astype(np.uint64)
else: # in case of python number
n = int(n)
n = (n & 0x5555555555555555) + ((n & 0xAAAAAAAAAAAAAAAA) >> 1)
n = (n & 0x3333333333333333) + ((n & 0xCCCCCCCCCCCCCCCC) >> 2)
n = (n & 0x0F0F0F0F0F0F0F0F) + ((n & 0xF0F0F0F0F0F0F0F0) >> 4)
n = (n & 0x00FF00FF00FF00FF) + ((n & 0xFF00FF00FF00FF00) >> 8)
n = (n & 0x0000FFFF0000FFFF) + ((n & 0xFFFF0000FFFF0000) >> 16)
n = (n & 0x00000000FFFFFFFF) + ((n & 0xFFFFFFFF00000000) >> 32) # This last & isn't strictly necessary.
return n
def costvolumeSD(im1, im2, dmin=-20, dmax=20):
'''
creates a Squared Difference stereo cost volume
Args:
im1,im2: numpy arrays containing the stereo pair (im1 is reference)
dmin,dmax: minimum and maximum disparity to be explored
Returns:
numpy array containing cost volume of size [im1.shape[0], im1.shape[0], dmax+1 - dmin]
'''
imshape = im1.shape
CV =
|
np.zeros((imshape[0], imshape[1], dmax+1-dmin))
|
numpy.zeros
|
"""
get data loaders
"""
from __future__ import print_function
import os
import socket
import numpy as np
from torch.utils.data import DataLoader
from torchvision import datasets
from torchvision import transforms
def get_data_folder():
"""
return server-dependent path to store the data
"""
hostname = socket.gethostname()
if hostname.startswith('visiongpu'):
data_folder = '/data/vision/phillipi/rep-learn/datasets/imagenet'
elif hostname.startswith('yonglong-home'):
data_folder = '/home/yonglong/Data/data/imagenet'
else:
data_folder = './data/imagenet2012'
if not os.path.isdir(data_folder):
os.makedirs(data_folder)
return data_folder
class ImageFolderInstance(datasets.ImageFolder):
""": Folder datasets which returns the index of the image as well::
"""
def __getitem__(self, index):
"""
Args:
index (int): Index
Returns:
tuple: (image, target) where target is class_index of the target class.
"""
path, target = self.imgs[index]
img = self.loader(path)
if self.transform is not None:
img = self.transform(img)
if self.target_transform is not None:
target = self.target_transform(target)
return img, target, index
class ImageFolderSample(datasets.ImageFolder):
""": Folder datasets which returns (img, label, index, contrast_index):
"""
def __init__(self, root, transform=None, target_transform=None,
is_sample=False, k=4096):
super().__init__(root=root, transform=transform, target_transform=target_transform)
self.k = k
self.is_sample = is_sample
print('stage1 finished!')
if self.is_sample:
num_classes = len(self.classes)
num_samples = len(self.samples)
label = np.zeros(num_samples, dtype=np.int32)
for i in range(num_samples):
path, target = self.imgs[i]
label[i] = target
self.cls_positive = [[] for i in range(num_classes)]
for i in range(num_samples):
self.cls_positive[label[i]].append(i)
self.cls_negative = [[] for i in range(num_classes)]
for i in range(num_classes):
for j in range(num_classes):
if j == i:
continue
self.cls_negative[i].extend(self.cls_positive[j])
self.cls_positive = [
|
np.asarray(self.cls_positive[i], dtype=np.int32)
|
numpy.asarray
|
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
# Deep Recurrent Reinforcement Learning: 1 capa LSTM y 4 capas Dense, Funcion de activacion tanh, 12 episodes, 50 iteraciones
drnnLSTMtanhMakespan0=[799, 798, 799, 799, 805, 806, 799, 805, 805, 800, 798, 798]
drnnLSTMtanhMakespan1=[800, 798, 796, 800, 796, 794, 795, 798, 800, 798, 805, 798]
drnnLSTMtanhMakespan2=[796, 800, 798, 804, 800, 798, 798, 798, 800, 800, 802, 797]
drnnLSTMtanhMakespan3=[805, 800, 800, 803, 794, 802, 800, 798, 799, 804, 799, 806]
drnnLSTMtanhMakespan4=[796, 798, 795, 798, 796, 799, 800, 796, 796, 798, 806, 800]
drnnLSTMtanhMakespan5=[798, 798, 799, 800, 800, 808, 798, 798, 801, 796, 799, 798]
drnnLSTMtanhMakespan6=[800, 796, 805, 798, 798, 796, 799, 800, 803, 800, 798, 800]
drnnLSTMtanhMakespan7=[799, 805, 802, 805, 800, 799, 800, 799, 805, 800, 794, 796]
drnnLSTMtanhMakespan8=[799, 798, 800, 798, 798, 800, 800, 800, 804, 799, 800, 804]
drnnLSTMtanhMakespan9=[795, 800, 795, 796, 798, 796, 797, 800, 797, 798, 796, 795]
drnnLSTMtanhMakespan10=[804, 799, 805, 798, 798, 798, 805, 800, 796, 804, 796, 799]
drnnLSTMtanhMakespan11=[795, 803, 805, 798, 795, 801, 798, 798, 804, 803, 799, 804]
drnnLSTMtanhMakespan12=[798, 798, 799, 800, 798, 798, 799, 799, 801, 796, 799, 798]
drnnLSTMtanhMakespan13=[798, 798, 799, 797, 796, 796, 800, 797, 805, 800, 800, 794]
drnnLSTMtanhMakespan14=[800, 798, 798, 796, 800, 800, 798, 798, 802, 798, 802, 798]
drnnLSTMtanhMakespan15=[796, 796, 800, 801, 800, 800, 796, 794, 796, 800, 796, 798]
drnnLSTMtanhMakespan16=[798, 798, 795, 797, 795, 799, 800, 796, 795, 796, 800, 800]
drnnLSTMtanhMakespan17=[794, 795, 800, 798, 795, 796, 798, 796, 795, 794, 798, 796]
drnnLSTMtanhMakespan18=[797, 795, 794, 794, 800, 796, 796, 795, 798, 795, 798, 794]
drnnLSTMtanhMakespan19=[797, 795, 795, 796, 798, 799, 795, 799, 795, 794, 795, 795]
drnnLSTMtanhMakespan20=[796, 794, 798, 797, 798, 799, 795, 795, 797, 795, 795, 792]
drnnLSTMtanhMakespan21=[797, 795, 797, 793, 794, 794, 800, 794, 798, 795, 797, 795]
drnnLSTMtanhMakespan22=[794, 800, 798, 795, 795, 796, 796, 799, 795, 794, 795, 795]
drnnLSTMtanhMakespan23=[795, 795, 794, 795, 794, 794, 797, 799, 796, 794, 794, 795]
drnnLSTMtanhMakespan24=[798, 795, 795, 795, 792, 794, 795, 794, 794, 795, 795, 795]
drnnLSTMtanhMakespan25=[794, 792, 794, 795, 795, 794, 794, 794, 794, 795, 794, 793]
drnnLSTMtanhMakespan26=[794, 794, 795, 796, 798, 795, 794, 794, 794, 794, 795, 794]
drnnLSTMtanhMakespan27=[795, 794, 795, 795, 795, 794, 794, 794, 794, 794, 795, 795]
drnnLSTMtanhMakespan28=[795, 794, 794, 795, 794, 795, 795, 795, 795, 794, 795, 794]
drnnLSTMtanhMakespan29=[792, 794, 795, 794, 794, 795, 794, 793, 795, 794, 795, 792]
drnnLSTMtanhMakespan30=[795, 794, 795, 795, 794, 794, 794, 795, 794, 794, 794, 794]
drnnLSTMtanhMakespan31=[794, 794, 795, 794, 795, 793, 795, 795, 795, 792, 794, 794]
drnnLSTMtanhMakespan32=[795, 795, 794, 793, 795, 795, 795, 795, 794, 794, 795, 794]
drnnLSTMtanhMakespan33=[793, 794, 795, 793, 792, 795, 794, 794, 794, 794, 794, 795]
drnnLSTMtanhMakespan34=[794, 795, 795, 794, 794, 794, 794, 793, 794, 794, 794, 794]
drnnLSTMtanhMakespan35=[794, 794, 797, 793, 792, 794, 793, 794, 795, 794, 795, 792]
drnnLSTMtanhMakespan36=[794, 794, 793, 794, 795, 797, 795, 795, 794, 795, 793, 794]
drnnLSTMtanhMakespan37=[795, 793, 795, 794, 795, 798, 795, 794, 795, 793, 795, 794]
drnnLSTMtanhMakespan38=[794, 795, 793, 795, 794, 794, 794, 794, 794, 794, 797, 795]
drnnLSTMtanhMakespan39=[794, 794, 795, 794, 795, 795, 794, 795, 794, 795, 798, 797]
drnnLSTMtanhMakespan40=[795, 795, 794, 795, 794, 795, 795, 794, 794, 794, 795, 795]
drnnLSTMtanhMakespan41=[794, 795, 792, 794, 794, 798, 795, 794, 794, 794, 793, 795]
drnnLSTMtanhMakespan42=[793, 795, 794, 793, 794, 794, 792, 794, 795, 794, 794, 793]
drnnLSTMtanhMakespan43=[793, 792, 793, 794, 794, 795, 792, 794, 795, 794, 795, 794]
drnnLSTMtanhMakespan44=[793, 794, 795, 795, 794, 794, 795, 798, 794, 792, 795, 794]
drnnLSTMtanhMakespan45=[795, 794, 794, 794, 794, 792, 794, 795, 794, 796, 795, 794]
drnnLSTMtanhMakespan46=[794, 793, 793, 795, 795, 794, 794, 794, 794, 796, 794, 794]
drnnLSTMtanhMakespan47=[794, 794, 795, 794, 794, 795, 792, 795, 794, 795, 795, 794]
drnnLSTMtanhMakespan48=[794, 795, 794, 794, 794, 792, 794, 795, 796, 794, 794, 795]
drnnLSTMtanhMakespan49=[794, 794, 794, 794, 794, 794, 792, 794, 793, 794, 795, 794]
drnnLSTMtanhRewards0=[-0.1759911894273128, -0.17580964970257765, -0.1759911894273128, -0.1759911894273128, -0.177078750549934, -0.17725973169122497, -0.1759911894273128, -0.177078750549934, -0.177078750549934, -0.17617264919621228, -0.17580964970257765, -0.17580964970257765]
drnnLSTMtanhRewards1=[-0.17617264919621228, -0.17580964970257765, -0.17544633017412387, -0.17617264919621228, -0.17544633017412387, -0.17508269018743108, -0.17526455026455026, -0.17580964970257765, -0.17617264919621228, -0.177078750549934, -0.17580964970257765, -0.17580964970257765]
drnnLSTMtanhRewards2=[-0.17544633017412387, -0.17617264919621228, -0.17580964970257765, -0.1768976897689769, -0.17617264919621228, -0.17580964970257765, -0.17580964970257765, -0.17580964970257765, -0.17617264919621228, -0.17617264919621228, -0.17653532907770195, -0.17562802996914942]
drnnLSTMtanhRewards3=[-0.177078750549934, -0.17617264919621228, -0.17617264919621228, -0.17671654929577466, -0.17508269018743108, -0.17653532907770195, -0.17617264919621228, -0.17580964970257765, -0.1759911894273128, -0.1768976897689769, -0.1759911894273128, -0.17725973169122497]
drnnLSTMtanhRewards4=[-0.17580964970257765, -0.17544633017412387, -0.17526455026455026, -0.17580964970257765, -0.17544633017412387, -0.1759911894273128, -0.17617264919621228, -0.17544633017412387, -0.17544633017412387, -0.17580964970257765, -0.17725973169122497, -0.17617264919621228]
drnnLSTMtanhRewards5=[-0.17580964970257765, -0.17580964970257765, -0.1759911894273128, -0.17617264919621228, -0.17617264919621228, -0.1776214552648934, -0.17580964970257765, -0.17580964970257765, -0.1763540290620872, -0.17544633017412387, -0.1759911894273128, -0.17580964970257765]
drnnLSTMtanhRewards6=[-0.17617264919621228, -0.17544633017412387, -0.177078750549934, -0.17580964970257765, -0.17580964970257765, -0.17544633017412387, -0.1759911894273128, -0.17617264919621228, -0.17671654929577466, -0.17617264919621228, -0.17580964970257765, -0.17617264919621228]
drnnLSTMtanhRewards7=[-0.1759911894273128, -0.177078750549934, -0.17653532907770195, -0.177078750549934, -0.17617264919621228, -0.1759911894273128, -0.17617264919621228, -0.1759911894273128, -0.177078750549934, -0.17617264919621228, -0.17508269018743108, -0.17544633017412387]
drnnLSTMtanhRewards8=[-0.1759911894273128, -0.17580964970257765, -0.17617264919621228, -0.17580964970257765, -0.17580964970257765, -0.17617264919621228, -0.17617264919621228, -0.17617264919621228, -0.1768976897689769, -0.1759911894273128, -0.17617264919621228, -0.1768976897689769]
drnnLSTMtanhRewards9=[-0.17526455026455026, -0.17617264919621228, -0.17526455026455026, -0.17544633017412387, -0.17580964970257765, -0.17544633017412387, -0.17562802996914942, -0.17617264919621228, -0.17562802996914942, -0.17580964970257765, -0.17544633017412387, -0.17526455026455026]
drnnLSTMtanhRewards10=[-0.1768976897689769, -0.1759911894273128, -0.177078750549934, -0.17580964970257765, -0.17580964970257765, -0.17580964970257765, -0.177078750549934, -0.17617264919621228, -0.17544633017412387, -0.1768976897689769, -0.17544633017412387, -0.1759911894273128]
drnnLSTMtanhRewards11=[-0.17526455026455026, -0.17671654929577466, -0.177078750549934, -0.17580964970257765, -0.17526455026455026, -0.1763540290620872, -0.17580964970257765, -0.17580964970257765, -0.1768976897689769, -0.17671654929577466, -0.1759911894273128, -0.1768976897689769]
drnnLSTMtanhRewards12=[-0.17580964970257765, -0.17580964970257765, -0.1759911894273128, -0.17617264919621228, -0.17580964970257765, -0.17580964970257765, -0.1759911894273128, -0.1759911894273128, -0.1763540290620872, -0.17544633017412387, -0.1759911894273128, -0.17580964970257765]
drnnLSTMtanhRewards13=[-0.17580964970257765, -0.17580964970257765, -0.1759911894273128, -0.17562802996914942, -0.17544633017412387, -0.17544633017412387, -0.17617264919621228, -0.17562802996914942, -0.177078750549934, -0.17617264919621228, -0.17617264919621228, -0.17508269018743108]
drnnLSTMtanhRewards14=[-0.17617264919621228, -0.17580964970257765, -0.17580964970257765, -0.17544633017412387, -0.17617264919621228, -0.17617264919621228, -0.17580964970257765, -0.17580964970257765, -0.17653532907770195, -0.17580964970257765, -0.17653532907770195, -0.17580964970257765]
drnnLSTMtanhRewards15=[-0.17544633017412387, -0.17544633017412387, -0.17617264919621228, -0.1763540290620872, -0.17617264919621228, -0.17617264919621228, -0.17544633017412387, -0.17508269018743108, -0.17544633017412387, -0.17617264919621228, -0.17544633017412387, -0.17580964970257765]
drnnLSTMtanhRewards16=[-0.17580964970257765, -0.17580964970257765, -0.17526455026455026, -0.17562802996914942, -0.17526455026455026, -0.1759911894273128, -0.17617264919621228, -0.17544633017412387, -0.17544633017412387, -0.17526455026455026, -0.17617264919621228, -0.17617264919621228]
drnnLSTMtanhRewards17=[-0.17508269018743108, -0.17526455026455026, -0.17580964970257765, -0.17617264919621228, -0.17526455026455026, -0.17544633017412387, -0.17580964970257765, -0.17544633017412387, -0.17526455026455026, -0.17508269018743108, -0.17580964970257765, -0.17544633017412387]
drnnLSTMtanhRewards18=[-0.17562802996914942, -0.17526455026455026, -0.17508269018743108, -0.17508269018743108, -0.17617264919621228, -0.17544633017412387, -0.17544633017412387, -0.17526455026455026, -0.17580964970257765, -0.17526455026455026, -0.17580964970257765, -0.17508269018743108]
drnnLSTMtanhRewards19=[-0.17562802996914942, -0.17526455026455026, -0.17526455026455026, -0.17544633017412387, -0.17580964970257765, -0.1759911894273128, -0.17526455026455026, -0.1759911894273128, -0.17526455026455026, -0.17508269018743108, -0.17526455026455026, -0.17526455026455026]
drnnLSTMtanhRewards20=[-0.17544633017412387, -0.17508269018743108, -0.17580964970257765, -0.17562802996914942, -0.17580964970257765, -0.1759911894273128, -0.17526455026455026, -0.17526455026455026, -0.17526455026455026, -0.17562802996914942, -0.17526455026455026, -0.17471872931833224]
drnnLSTMtanhRewards21=[-0.17562802996914942, -0.17526455026455026, -0.17562802996914942, -0.1749007498897221, -0.17508269018743108, -0.17508269018743108, -0.17617264919621228, -0.17508269018743108, -0.17580964970257765, -0.17526455026455026, -0.17562802996914942, -0.17526455026455026]
drnnLSTMtanhRewards22=[-0.17508269018743108, -0.17617264919621228, -0.17580964970257765, -0.17526455026455026, -0.17526455026455026, -0.17544633017412387, -0.17544633017412387, -0.1759911894273128, -0.17526455026455026, -0.17508269018743108, -0.17526455026455026, -0.17526455026455026]
drnnLSTMtanhRewards23=[-0.17526455026455026, -0.17526455026455026, -0.17508269018743108, -0.17526455026455026, -0.17508269018743108, -0.17508269018743108, -0.17562802996914942, -0.1759911894273128, -0.17544633017412387, -0.17508269018743108, -0.17508269018743108, -0.17526455026455026]
drnnLSTMtanhRewards24=[-0.17580964970257765, -0.17526455026455026, -0.17526455026455026, -0.17526455026455026, -0.17471872931833224, -0.17508269018743108, -0.17526455026455026, -0.17508269018743108, -0.17508269018743108, -0.17526455026455026, -0.17526455026455026, -0.17526455026455026]
drnnLSTMtanhRewards25=[-0.17508269018743108, -0.17471872931833224, -0.17508269018743108, -0.17526455026455026, -0.17526455026455026, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17526455026455026, -0.17508269018743108, -0.1749007498897221]
drnnLSTMtanhRewards26=[-0.17508269018743108, -0.17508269018743108, -0.17526455026455026, -0.17544633017412387, -0.17580964970257765, -0.17526455026455026, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17526455026455026, -0.17508269018743108]
drnnLSTMtanhRewards27=[-0.17526455026455026, -0.17508269018743108, -0.17526455026455026, -0.17526455026455026, -0.17526455026455026, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17526455026455026, -0.17526455026455026]
drnnLSTMtanhRewards28=[-0.17526455026455026, -0.17508269018743108, -0.17508269018743108, -0.17526455026455026, -0.17508269018743108, -0.17526455026455026, -0.17526455026455026, -0.17526455026455026, -0.17526455026455026, -0.17508269018743108, -0.17526455026455026, -0.17508269018743108]
drnnLSTMtanhRewards29=[-0.17471872931833224, -0.17508269018743108, -0.17526455026455026, -0.17508269018743108, -0.17508269018743108, -0.17526455026455026, -0.17508269018743108, -0.1749007498897221, -0.17526455026455026, -0.17508269018743108, -0.17526455026455026, -0.17471872931833224]
drnnLSTMtanhRewards30=[-0.17526455026455026, -0.17508269018743108, -0.17526455026455026, -0.17526455026455026, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17526455026455026, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108]
drnnLSTMtanhRewards31=[-0.17508269018743108, -0.17508269018743108, -0.17526455026455026, -0.17508269018743108, -0.17526455026455026, -0.1749007498897221, -0.17526455026455026, -0.17471872931833224, -0.17526455026455026, -0.17526455026455026, -0.17508269018743108, -0.17508269018743108]
drnnLSTMtanhRewards32=[-0.17526455026455026, -0.17526455026455026, -0.1749007498897221, -0.17508269018743108, -0.17526455026455026, -0.17526455026455026, -0.17526455026455026, -0.17526455026455026, -0.17508269018743108, -0.17508269018743108, -0.17526455026455026, -0.17508269018743108]
drnnLSTMtanhRewards33=[-0.1749007498897221, -0.17508269018743108, -0.17526455026455026, -0.1749007498897221, -0.17471872931833224, -0.17526455026455026, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17526455026455026]
drnnLSTMtanhRewards34=[-0.17508269018743108, -0.17526455026455026, -0.17526455026455026, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.1749007498897221, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108]
drnnLSTMtanhRewards35=[-0.17508269018743108, -0.17508269018743108, -0.17562802996914942, -0.17471872931833224, -0.1749007498897221, -0.17508269018743108, -0.1749007498897221, -0.17508269018743108, -0.17526455026455026, -0.17508269018743108, -0.17526455026455026, -0.17471872931833224]
drnnLSTMtanhRewards36=[-0.17508269018743108, -0.17508269018743108, -0.1749007498897221, -0.17508269018743108, -0.17526455026455026, -0.17562802996914942, -0.17526455026455026, -0.17526455026455026, -0.17508269018743108, -0.17526455026455026, -0.1749007498897221, -0.17508269018743108]
drnnLSTMtanhRewards37=[-0.17526455026455026, -0.1749007498897221, -0.17526455026455026, -0.17508269018743108, -0.17526455026455026, -0.17580964970257765, -0.17526455026455026, -0.17508269018743108, -0.17526455026455026, -0.1749007498897221, -0.17526455026455026, -0.17508269018743108]
drnnLSTMtanhRewards38=[-0.17508269018743108, -0.17526455026455026, -0.1749007498897221, -0.17526455026455026, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17562802996914942, -0.17526455026455026]
drnnLSTMtanhRewards39=[-0.17508269018743108, -0.17508269018743108, -0.17526455026455026, -0.17508269018743108, -0.17526455026455026, -0.17526455026455026, -0.17508269018743108, -0.17526455026455026, -0.17508269018743108, -0.17526455026455026, -0.17580964970257765, -0.17562802996914942]
drnnLSTMtanhRewards40=[-0.17526455026455026, -0.17526455026455026, -0.17508269018743108, -0.17526455026455026, -0.17508269018743108, -0.17526455026455026, -0.17508269018743108, -0.17526455026455026, -0.17508269018743108, -0.17508269018743108, -0.17526455026455026, -0.17526455026455026]
drnnLSTMtanhRewards41=[-0.17508269018743108, -0.17526455026455026, -0.17471872931833224, -0.17508269018743108, -0.17508269018743108, -0.17580964970257765, -0.17526455026455026, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.1749007498897221, -0.17526455026455026]
drnnLSTMtanhRewards42=[-0.1749007498897221, -0.17526455026455026, -0.17508269018743108, -0.1749007498897221, -0.17508269018743108, -0.17508269018743108, -0.17471872931833224, -0.17508269018743108, -0.17526455026455026, -0.17508269018743108, -0.17508269018743108, -0.1749007498897221]
drnnLSTMtanhRewards43=[-0.1749007498897221, -0.17471872931833224, -0.1749007498897221, -0.17508269018743108, -0.17508269018743108, -0.17526455026455026, -0.17471872931833224, -0.17508269018743108, -0.17526455026455026, -0.17508269018743108, -0.17526455026455026, -0.17508269018743108]
drnnLSTMtanhRewards44=[-0.1749007498897221, -0.17508269018743108, -0.17526455026455026, -0.17526455026455026, -0.17508269018743108, -0.17508269018743108, -0.17580964970257765, -0.17508269018743108, -0.17526455026455026, -0.17471872931833224, -0.17526455026455026, -0.17508269018743108]
drnnLSTMtanhRewards45=[-0.17526455026455026, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17471872931833224, -0.17508269018743108, -0.17526455026455026, -0.17508269018743108, -0.17544633017412387, -0.17526455026455026, -0.17508269018743108]
drnnLSTMtanhRewards46=[-0.17508269018743108, -0.1749007498897221, -0.1749007498897221, -0.17526455026455026, -0.17526455026455026, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17544633017412387, -0.17508269018743108, -0.17508269018743108]
drnnLSTMtanhRewards47=[-0.17508269018743108, -0.17508269018743108, -0.17526455026455026, -0.17508269018743108, -0.17508269018743108, -0.17526455026455026, -0.17471872931833224, -0.17508269018743108, -0.17526455026455026, -0.17526455026455026, -0.17526455026455026, -0.17508269018743108]
drnnLSTMtanhRewards48=[-0.17508269018743108, -0.17526455026455026, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17471872931833224, -0.17508269018743108, -0.17526455026455026, -0.17544633017412387, -0.17508269018743108, -0.17508269018743108, -0.17526455026455026]
drnnLSTMtanhRewards49=[-0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17471872931833224, -0.17508269018743108, -0.1749007498897221, -0.17508269018743108, -0.17526455026455026, -0.17508269018743108]
# Deep Recurrent Reinforcement Learning: 1 capa LSTM y 4 capas Dense, Funcion de activacion relu, 12 episodes, 50 iteraciones
drnnLSTMreluMakespan0=[805, 800, 800, 800, 794, 800, 798, 809, 795, 800, 798, 798]
drnnLSTMreluMakespan1=[798, 798, 796, 799, 800, 796, 796, 798, 798, 794, 798, 800]
drnnLSTMreluMakespan2=[805, 805, 798, 799, 806, 799, 806, 799, 800, 798, 805, 795]
drnnLSTMreluMakespan3=[800, 800, 800, 796, 800, 800, 799, 806, 808, 798, 797, 798]
drnnLSTMreluMakespan4=[805, 805, 795, 796, 799, 804, 798, 794, 798, 794, 796, 810]
drnnLSTMreluMakespan5=[798, 798, 798, 795, 800, 798, 796, 802, 800, 800, 805, 801]
drnnLSTMreluMakespan6=[800, 798, 798, 795, 800, 796, 800, 798, 799, 796, 805, 800]
drnnLSTMreluMakespan7=[800, 800, 800, 799, 798, 798, 800, 805, 800, 799, 800, 801]
drnnLSTMreluMakespan8=[799, 800, 800, 799, 795, 795, 805, 795, 798, 800, 798, 800]
drnnLSTMreluMakespan9=[800, 796, 805, 798, 798, 795, 805, 800, 799, 795, 800, 805]
drnnLSTMreluMakespan10=[805, 798, 805, 800, 801, 805, 799, 805, 798, 800, 800, 798]
drnnLSTMreluMakespan11=[798, 803, 800, 797, 795, 796, 794, 799, 800, 800, 800, 796]
drnnLSTMreluMakespan12=[799, 798, 799, 795, 798, 795, 798, 798, 798, 795, 798, 798]
drnnLSTMreluMakespan13=[798, 798, 799, 796, 798, 796, 800, 799, 796, 794, 796, 795]
drnnLSTMreluMakespan14=[796, 798, 806, 799, 804, 798, 805, 798, 800, 805, 794, 800]
drnnLSTMreluMakespan15=[806, 795, 800, 796, 798, 796, 810, 798, 799, 798, 800, 800]
drnnLSTMreluMakespan16=[799, 796, 798, 798, 798, 800, 798, 810, 796, 805, 800, 795]
drnnLSTMreluMakespan17=[798, 798, 798, 794, 798, 805, 801, 798, 800, 799, 798, 798]
drnnLSTMreluMakespan18=[795, 800, 794, 798, 797, 798, 794, 800, 797, 796, 794, 794]
drnnLSTMreluMakespan19=[798, 802, 794, 798, 799, 795, 797, 795, 800, 796, 797, 796]
drnnLSTMreluMakespan20=[794, 797, 795, 794, 799, 795, 795, 795, 800, 797, 794, 798]
drnnLSTMreluMakespan21=[799, 798, 796, 795, 794, 798, 795, 795, 798, 798, 795, 794]
drnnLSTMreluMakespan22=[794, 794, 795, 797, 795, 795, 795, 792, 794, 795, 794, 794]
drnnLSTMreluMakespan23=[794, 794, 794, 794, 795, 796, 793, 794, 795, 794, 797, 795]
drnnLSTMreluMakespan24=[794, 792, 792, 794, 796, 792, 794, 795, 794, 792, 796, 795]
drnnLSTMreluMakespan25=[794, 795, 795, 794, 794, 792, 795, 792, 795, 794, 794, 794]
drnnLSTMreluMakespan26=[795, 794, 794, 795, 794, 794, 793, 794, 797, 795, 794, 795]
drnnLSTMreluMakespan27=[794, 794, 795, 796, 795, 797, 794, 794, 795, 801, 794, 795]
drnnLSTMreluMakespan28=[795, 795, 795, 795, 794, 792, 794, 797, 794, 795, 795, 795]
drnnLSTMreluMakespan29=[794, 792, 798, 794, 797, 795, 793, 795, 795, 794, 795, 795]
drnnLSTMreluMakespan30=[795, 794, 798, 794, 794, 795, 792, 796, 794, 796, 794, 794]
drnnLSTMreluMakespan31=[794, 795, 795, 794, 795, 794, 795, 795, 794, 794, 795, 795]
drnnLSTMreluMakespan32=[798, 794, 794, 794, 798, 792, 795, 795, 795, 796, 794, 795]
drnnLSTMreluMakespan33=[794, 796, 794, 794, 794, 795, 794, 794, 797, 793, 793, 795]
drnnLSTMreluMakespan34=[794, 794, 795, 794, 794, 793, 794, 795, 793, 795, 795, 794]
drnnLSTMreluMakespan35=[798, 796, 795, 794, 795, 795, 795, 795, 794, 795, 797, 795]
drnnLSTMreluMakespan36=[794, 796, 794, 794, 794, 794, 795, 795, 797, 796, 795, 795]
drnnLSTMreluMakespan37=[795, 794, 796, 795, 795, 795, 795, 794, 792, 797, 794, 793]
drnnLSTMreluMakespan38=[794, 798, 794, 792, 794, 792, 795, 797, 793, 794, 794, 797]
drnnLSTMreluMakespan39=[792, 794, 794, 794, 792, 795, 795, 795, 794, 794, 795, 794]
drnnLSTMreluMakespan40=[792, 795, 795, 792, 795, 795, 794, 795, 794, 795, 794, 795]
drnnLSTMreluMakespan41=[794, 797, 795, 794, 795, 795, 798, 794, 795, 796, 796, 794]
drnnLSTMreluMakespan42=[794, 795, 795, 795, 794, 795, 795, 794, 794, 795, 793, 795]
drnnLSTMreluMakespan43=[795, 794, 795, 794, 795, 795, 792, 794, 794, 795, 794, 795]
drnnLSTMreluMakespan44=[795, 794, 792, 795, 794, 794, 795, 794, 796, 795, 796, 794]
drnnLSTMreluMakespan45=[795, 794, 793, 794, 793, 795, 794, 794, 795, 794, 795, 794]
drnnLSTMreluMakespan46=[794, 796, 793, 794, 794, 795, 799, 795, 794, 794, 794, 794]
drnnLSTMreluMakespan47=[794, 794, 794, 794, 795, 793, 795, 795, 794, 795, 795, 795]
drnnLSTMreluMakespan48=[794, 794, 795, 794, 795, 795, 795, 794, 794, 795, 795, 794]
drnnLSTMreluMakespan49=[795, 795, 795, 794, 795, 795, 794, 795, 793, 793, 792, 792]
drnnLSTMreluRewards0=[-0.177078750549934, -0.17617264919621228, -0.17617264919621228, -0.17617264919621228, -0.17508269018743108, -0.17617264919621228, -0.17580964970257765, -0.1778021978021978, -0.17526455026455026, -0.17617264919621228, -0.17580964970257765, -0.17580964970257765]
drnnLSTMreluRewards1=[-0.17580964970257765, -0.17580964970257765, -0.17544633017412387, -0.1759911894273128, -0.17617264919621228, -0.17544633017412387, -0.17544633017412387, -0.17580964970257765, -0.17580964970257765, -0.17508269018743108, -0.17580964970257765, -0.17617264919621228]
drnnLSTMreluRewards2=[-0.177078750549934, -0.177078750549934, -0.17580964970257765, -0.1759911894273128, -0.17725973169122497, -0.1759911894273128, -0.17725973169122497, -0.1759911894273128, -0.17617264919621228, -0.17580964970257765, -0.177078750549934, -0.17526455026455026]
drnnLSTMreluRewards3=[-0.17617264919621228, -0.17617264919621228, -0.17617264919621228, -0.17544633017412387, -0.17617264919621228, -0.17617264919621228, -0.1759911894273128, -0.17725973169122497, -0.1776214552648934, -0.17580964970257765, -0.17562802996914942, -0.17580964970257765]
drnnLSTMreluRewards4=[-0.177078750549934, -0.177078750549934, -0.17526455026455026, -0.17544633017412387, -0.1759911894273128, -0.1768976897689769, -0.17580964970257765, -0.17508269018743108, -0.17580964970257765, -0.17508269018743108, -0.17544633017412387, -0.17798286090969018]
drnnLSTMreluRewards5=[-0.17580964970257765, -0.17580964970257765, -0.17580964970257765, -0.17526455026455026, -0.17617264919621228, -0.17580964970257765, -0.17544633017412387, -0.17653532907770195, -0.17617264919621228, -0.17617264919621228, -0.177078750549934, -0.1763540290620872]
drnnLSTMreluRewards6=[-0.17617264919621228, -0.17580964970257765, -0.17580964970257765, -0.17526455026455026, -0.17617264919621228, -0.17544633017412387, -0.17617264919621228, -0.17580964970257765, -0.1759911894273128, -0.17544633017412387, -0.177078750549934, -0.17617264919621228]
drnnLSTMreluRewards7=[-0.17617264919621228, -0.17617264919621228, -0.17617264919621228, -0.1759911894273128, -0.17580964970257765, -0.17580964970257765, -0.17617264919621228, -0.177078750549934, -0.17617264919621228, -0.1759911894273128, -0.17617264919621228, -0.1763540290620872]
drnnLSTMreluRewards8=[-0.1759911894273128, -0.17617264919621228, -0.17617264919621228, -0.1759911894273128, -0.17526455026455026, -0.17526455026455026, -0.177078750549934, -0.17526455026455026, -0.17580964970257765, -0.17617264919621228, -0.17580964970257765, -0.17617264919621228]
drnnLSTMreluRewards9=[-0.17617264919621228, -0.17544633017412387, -0.177078750549934, -0.17580964970257765, -0.17580964970257765, -0.177078750549934, -0.17526455026455026, -0.17617264919621228, -0.1759911894273128, -0.17526455026455026, -0.17617264919621228, -0.177078750549934]
drnnLSTMreluRewards10=[-0.177078750549934, -0.17580964970257765, -0.177078750549934, -0.17617264919621228, -0.1763540290620872, -0.1759911894273128, -0.177078750549934, -0.177078750549934, -0.17580964970257765, -0.17617264919621228, -0.17617264919621228, -0.17580964970257765]
drnnLSTMreluRewards11=[-0.17580964970257765, -0.17671654929577466, -0.17617264919621228, -0.17562802996914942, -0.17526455026455026, -0.17544633017412387, -0.17508269018743108, -0.1759911894273128, -0.17617264919621228, -0.17617264919621228, -0.17617264919621228, -0.17544633017412387]
drnnLSTMreluRewards12=[-0.1759911894273128, -0.17580964970257765, -0.1759911894273128, -0.17526455026455026, -0.17580964970257765, -0.17526455026455026, -0.17580964970257765, -0.17580964970257765, -0.17580964970257765, -0.17526455026455026, -0.17580964970257765, -0.17580964970257765]
drnnLSTMreluRewards13=[-0.17580964970257765, -0.17580964970257765, -0.1759911894273128, -0.17544633017412387, -0.17580964970257765, -0.17544633017412387, -0.17617264919621228, -0.1759911894273128, -0.17544633017412387, -0.17508269018743108, -0.17544633017412387, -0.17526455026455026]
drnnLSTMreluRewards14=[-0.17544633017412387, -0.17580964970257765, -0.17725973169122497, -0.1759911894273128, -0.1768976897689769, -0.17580964970257765, -0.177078750549934, -0.17580964970257765, -0.17617264919621228, -0.177078750549934, -0.17508269018743108, -0.17617264919621228]
drnnLSTMreluRewards15=[-0.17725973169122497, -0.17526455026455026, -0.17617264919621228, -0.17544633017412387, -0.17580964970257765, -0.17544633017412387, -0.17798286090969018, -0.17580964970257765, -0.1759911894273128, -0.17580964970257765, -0.17617264919621228, -0.17617264919621228]
drnnLSTMreluRewards16=[-0.1759911894273128, -0.17544633017412387, -0.17580964970257765, -0.17580964970257765, -0.17580964970257765, -0.17617264919621228, -0.17580964970257765, -0.17798286090969018, -0.17544633017412387, -0.177078750549934, -0.17617264919621228, -0.17526455026455026]
drnnLSTMreluRewards17=[-0.17580964970257765, -0.17580964970257765, -0.17580964970257765, -0.17508269018743108, -0.17580964970257765, -0.177078750549934, -0.1763540290620872, -0.17580964970257765, -0.17617264919621228, -0.1759911894273128, -0.17580964970257765, -0.17580964970257765]
drnnLSTMreluRewards18=[-0.17526455026455026, -0.17617264919621228, -0.17508269018743108, -0.17580964970257765, -0.17562802996914942, -0.17580964970257765, -0.17508269018743108, -0.17617264919621228, -0.17562802996914942, -0.17544633017412387, -0.17508269018743108, -0.17508269018743108]
drnnLSTMreluRewards19=[-0.17580964970257765, -0.17653532907770195, -0.17508269018743108, -0.17580964970257765, -0.1759911894273128, -0.17526455026455026, -0.17562802996914942, -0.17526455026455026, -0.17617264919621228, -0.17544633017412387, -0.17562802996914942, -0.17544633017412387]
drnnLSTMreluRewards20=[-0.17508269018743108, -0.17562802996914942, -0.17526455026455026, -0.17508269018743108, -0.1759911894273128, -0.17526455026455026, -0.17526455026455026, -0.17526455026455026, -0.17617264919621228, -0.17562802996914942, -0.17508269018743108, -0.17580964970257765]
drnnLSTMreluRewards21=[-0.1759911894273128, -0.17580964970257765, -0.17544633017412387, -0.17526455026455026, -0.17508269018743108, -0.17580964970257765, -0.17526455026455026, -0.17526455026455026, -0.17580964970257765, -0.17580964970257765, -0.17526455026455026, -0.17508269018743108]
drnnLSTMreluRewards22=[-0.17508269018743108, -0.17508269018743108, -0.17526455026455026, -0.17562802996914942, -0.17526455026455026, -0.17526455026455026, -0.17526455026455026, -0.17471872931833224, -0.17508269018743108, -0.17526455026455026, -0.17508269018743108, -0.17508269018743108]
drnnLSTMreluRewards23=[-0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17526455026455026, -0.17544633017412387, -0.1749007498897221, -0.17508269018743108, -0.17526455026455026, -0.17508269018743108, -0.17562802996914942, -0.17526455026455026]
drnnLSTMreluRewards24=[-0.17508269018743108, -0.17471872931833224, -0.17471872931833224, -0.17508269018743108, -0.17544633017412387, -0.17471872931833224, -0.17508269018743108, -0.17526455026455026, -0.17508269018743108, -0.17471872931833224, -0.17544633017412387, -0.17526455026455026]
drnnLSTMreluRewards25=[-0.17508269018743108, -0.17526455026455026, -0.17526455026455026, -0.17508269018743108, -0.17508269018743108, -0.17471872931833224, -0.17526455026455026, -0.17471872931833224, -0.17526455026455026, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108]
drnnLSTMreluRewards26=[-0.17526455026455026, -0.17508269018743108, -0.17508269018743108, -0.17526455026455026, -0.17508269018743108, -0.17508269018743108, -0.1749007498897221, -0.17508269018743108, -0.17562802996914942, -0.17526455026455026, -0.17508269018743108, -0.17526455026455026]
drnnLSTMreluRewards27=[-0.17508269018743108, -0.17508269018743108, -0.17526455026455026, -0.17544633017412387, -0.17526455026455026, -0.17562802996914942, -0.17508269018743108, -0.17508269018743108, -0.17526455026455026, -0.1763540290620872, -0.17508269018743108, -0.17526455026455026]
drnnLSTMreluRewards28=[-0.17526455026455026, -0.17526455026455026, -0.17526455026455026, -0.17526455026455026, -0.17508269018743108, -0.17471872931833224, -0.17508269018743108, -0.17562802996914942, -0.17508269018743108, -0.17526455026455026, -0.17526455026455026, -0.17526455026455026]
drnnLSTMreluRewards29=[-0.17508269018743108, -0.17471872931833224, -0.17580964970257765, -0.17508269018743108, -0.17562802996914942, -0.17526455026455026, -0.1749007498897221, -0.17526455026455026, -0.17526455026455026, -0.17508269018743108, -0.17526455026455026, -0.17526455026455026]
drnnLSTMreluRewards30=[-0.17526455026455026, -0.17508269018743108, -0.17580964970257765, -0.17508269018743108, -0.17508269018743108, -0.17526455026455026, -0.17471872931833224, -0.17544633017412387, -0.17508269018743108, -0.17544633017412387, -0.17508269018743108, -0.17508269018743108]
drnnLSTMreluRewards31=[-0.17508269018743108, -0.17526455026455026, -0.17526455026455026, -0.17508269018743108, -0.17526455026455026, -0.17508269018743108, -0.17526455026455026, -0.17526455026455026, -0.17508269018743108, -0.17508269018743108, -0.17526455026455026, -0.17526455026455026]
drnnLSTMreluRewards32=[-0.17580964970257765, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17580964970257765, -0.17471872931833224, -0.17526455026455026, -0.17526455026455026, -0.17526455026455026, -0.17544633017412387, -0.17508269018743108, -0.17526455026455026]
drnnLSTMreluRewards33=[-0.17508269018743108, -0.17544633017412387, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17526455026455026, -0.17508269018743108, -0.17508269018743108, -0.17562802996914942, -0.1749007498897221, -0.1749007498897221, -0.17526455026455026]
drnnLSTMreluRewards34=[-0.17508269018743108, -0.17508269018743108, -0.17526455026455026, -0.17508269018743108, -0.17508269018743108, -0.1749007498897221, -0.17508269018743108, -0.17526455026455026, -0.1749007498897221, -0.17526455026455026, -0.17526455026455026, -0.17508269018743108]
drnnLSTMreluRewards35=[-0.17580964970257765, -0.17544633017412387, -0.17526455026455026, -0.17508269018743108, -0.17526455026455026, -0.17526455026455026, -0.17526455026455026, -0.17526455026455026, -0.17508269018743108, -0.17526455026455026, -0.17562802996914942, -0.17526455026455026]
drnnLSTMreluRewards36=[-0.17508269018743108, -0.17544633017412387, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17526455026455026, -0.17526455026455026, -0.17562802996914942, -0.17544633017412387, -0.17526455026455026, -0.17526455026455026]
drnnLSTMreluRewards37=[-0.17526455026455026, -0.17508269018743108, -0.17544633017412387, -0.17526455026455026, -0.17526455026455026, -0.17526455026455026, -0.17526455026455026, -0.17508269018743108, -0.17471872931833224, -0.17562802996914942, -0.17508269018743108, -0.1749007498897221]
drnnLSTMreluRewards38=[-0.17508269018743108, -0.17580964970257765, -0.17508269018743108, -0.17471872931833224, -0.17508269018743108, -0.17471872931833224, -0.17526455026455026, -0.17562802996914942, -0.1749007498897221, -0.17508269018743108, -0.17508269018743108, -0.17562802996914942]
drnnLSTMreluRewards39=[-0.17471872931833224, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17471872931833224, -0.17526455026455026, -0.17526455026455026, -0.17526455026455026, -0.17508269018743108, -0.17508269018743108, -0.17526455026455026, -0.17508269018743108]
drnnLSTMreluRewards40=[-0.17471872931833224, -0.17526455026455026, -0.17526455026455026, -0.17471872931833224, -0.17526455026455026, -0.17526455026455026, -0.17508269018743108, -0.17526455026455026, -0.17508269018743108, -0.17526455026455026, -0.17508269018743108, -0.17526455026455026]
drnnLSTMreluRewards41=[-0.17508269018743108, -0.17562802996914942, -0.17526455026455026, -0.17508269018743108, -0.17526455026455026, -0.17526455026455026, -0.17580964970257765, -0.17508269018743108, -0.17526455026455026, -0.17544633017412387, -0.17544633017412387, -0.17508269018743108]
drnnLSTMreluRewards42=[-0.17508269018743108, -0.17526455026455026, -0.17526455026455026, -0.17526455026455026, -0.17508269018743108, -0.17526455026455026, -0.17526455026455026, -0.17508269018743108, -0.17508269018743108, -0.17526455026455026, -0.1749007498897221, -0.17526455026455026]
drnnLSTMreluRewards43=[-0.17526455026455026, -0.17508269018743108, -0.17526455026455026, -0.17508269018743108, -0.17526455026455026, -0.17526455026455026, -0.17471872931833224, -0.17508269018743108, -0.17508269018743108, -0.17526455026455026, -0.17508269018743108, -0.17526455026455026]
drnnLSTMreluRewards44=[-0.17526455026455026, -0.17508269018743108, -0.17471872931833224, -0.17526455026455026, -0.17508269018743108, -0.17508269018743108, -0.17526455026455026, -0.17508269018743108, -0.17544633017412387, -0.17526455026455026, -0.17544633017412387, -0.17508269018743108]
drnnLSTMreluRewards45=[-0.17526455026455026, -0.17508269018743108, -0.1749007498897221, -0.17508269018743108, -0.1749007498897221, -0.17526455026455026, -0.17508269018743108, -0.17508269018743108, -0.17526455026455026, -0.17508269018743108, -0.17526455026455026, -0.17508269018743108]
drnnLSTMreluRewards46=[-0.17508269018743108, -0.17544633017412387, -0.1749007498897221, -0.17508269018743108, -0.17508269018743108, -0.17526455026455026, -0.1759911894273128, -0.17526455026455026, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108]
drnnLSTMreluRewards47=[-0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17526455026455026, -0.1749007498897221, -0.17526455026455026, -0.17526455026455026, -0.17508269018743108, -0.17526455026455026, -0.17526455026455026, -0.17526455026455026]
drnnLSTMreluRewards48=[-0.17508269018743108, -0.17508269018743108, -0.17526455026455026, -0.17508269018743108, -0.17526455026455026, -0.17526455026455026, -0.17526455026455026, -0.17508269018743108, -0.17508269018743108, -0.17526455026455026, -0.17526455026455026, -0.17508269018743108]
drnnLSTMreluRewards49=[-0.17526455026455026, -0.17526455026455026, -0.17526455026455026, -0.17508269018743108, -0.17526455026455026, -0.17526455026455026, -0.17508269018743108, -0.17526455026455026, -0.1749007498897221, -0.1749007498897221, -0.17471872931833224, -0.17471872931833224]
# Deep Recurrent Reinforcement Learning: 1 capa GRU y 4 capas Dense, Funcion de activacion tanh, 12 episodes, 50 iteraciones
drnnGRUtanhMakespan0 = [798, 799, 798, 804, 805, 799, 801, 801, 801, 799, 798, 796]
drnnGRUtanhMakespan1 = [800, 798, 798, 798, 798, 798, 801, 798, 795, 796, 800, 796]
drnnGRUtanhMakespan2 = [795, 804, 805, 800, 800, 796, 804, 800, 795, 798, 798, 801]
drnnGRUtanhMakespan3 = [806, 796, 794, 797, 798, 800, 800, 808, 805, 798, 800, 809]
drnnGRUtanhMakespan4 = [805, 801, 795, 798, 798, 800, 796, 796, 805, 798, 799, 798]
drnnGRUtanhMakespan5 = [804, 799, 798, 804, 796, 799, 798, 805, 796, 805, 798, 800]
drnnGRUtanhMakespan6 = [800, 799, 794, 801, 799, 796, 800, 804, 797, 796, 800, 798]
drnnGRUtanhMakespan7 = [798, 800, 810, 810, 805, 800, 795, 798, 800, 805, 799, 800]
drnnGRUtanhMakespan8 = [798, 797, 800, 800, 804, 805, 798, 798, 801, 795, 798, 809]
drnnGRUtanhMakespan9 = [803, 800, 800, 805, 805, 798, 804, 803, 805, 801, 810, 801]
drnnGRUtanhMakespan10 = [798, 799, 798, 798, 805, 804, 805, 798, 799, 798, 800, 800]
drnnGRUtanhMakespan11 = [796, 795, 805, 800, 800, 798, 795, 804, 805, 798, 800, 800]
drnnGRUtanhMakespan12 = [799, 799, 809, 800, 799, 799, 797, 805, 799, 800, 798, 795]
drnnGRUtanhMakespan13 = [805, 800, 800, 805, 800, 799, 798, 801, 798, 797, 805, 800]
drnnGRUtanhMakespan14 = [800, 798, 800, 800, 800, 804, 804, 799, 799, 800, 798, 798]
drnnGRUtanhMakespan15 = [805, 800, 795, 800, 804, 795, 800, 798, 799, 798, 800, 796]
drnnGRUtanhMakespan16 = [806, 795, 801, 799, 799, 796, 796, 794, 802, 796, 800, 802]
drnnGRUtanhMakespan17 = [796, 800, 798, 800, 794, 800, 804, 805, 798, 810, 800, 798]
drnnGRUtanhMakespan18 = [798, 800, 794, 794, 797, 798, 800, 805, 798, 798, 804, 798]
drnnGRUtanhMakespan19 = [796, 800, 806, 799, 796, 800, 798, 805, 798, 799, 797, 805]
drnnGRUtanhMakespan20 = [805, 800, 799, 796, 805, 805, 805, 794, 809, 796, 800, 797]
drnnGRUtanhMakespan21 = [798, 800, 800, 800, 798, 801, 796, 801, 801, 801, 795, 799]
drnnGRUtanhMakespan22 = [798, 801, 797, 800, 799, 795, 799, 799, 800, 801, 800, 799]
drnnGRUtanhMakespan23 = [800, 798, 799, 805, 794, 800, 798, 796, 796, 804, 800, 794]
drnnGRUtanhMakespan24 = [800, 800, 798, 805, 804, 799, 798, 801, 800, 798, 798, 798]
drnnGRUtanhMakespan25 = [798, 798, 798, 795, 800, 803, 798, 798, 800, 799, 796, 798]
drnnGRUtanhMakespan26 = [796, 798, 798, 798, 805, 796, 798, 798, 805, 795, 801, 796]
drnnGRUtanhMakespan27 = [794, 796, 796, 800, 800, 798, 800, 798, 802, 798, 797, 798]
drnnGRUtanhMakespan28 = [799, 799, 800, 800, 798, 802, 799, 798, 795, 795, 794, 798]
drnnGRUtanhMakespan29 = [798, 796, 796, 797, 796, 798, 800, 800, 796, 798, 800, 795]
drnnGRUtanhMakespan30 = [799, 798, 795, 795, 800, 795, 798, 798, 799, 798, 805, 799]
drnnGRUtanhMakespan31 = [795, 799, 794, 794, 796, 795, 795, 794, 798, 797, 798, 795]
drnnGRUtanhMakespan32 = [797, 798, 795, 796, 798, 795, 797, 798, 795, 794, 795, 796]
drnnGRUtanhMakespan33 = [799, 795, 794, 794, 798, 795, 798, 797, 800, 796, 795, 794]
drnnGRUtanhMakespan34 = [798, 795, 798, 796, 798, 794, 796, 798, 798, 798, 796, 797]
drnnGRUtanhMakespan35 = [795, 798, 796, 798, 794, 801, 795, 800, 795, 800, 794, 800]
drnnGRUtanhMakespan36 = [798, 799, 796, 797, 795, 794, 800, 795, 795, 794, 795, 795]
drnnGRUtanhMakespan37 = [799, 798, 795, 795, 794, 795, 795, 796, 805, 795, 798, 796]
drnnGRUtanhMakespan38 = [798, 794, 795, 795, 795, 796, 795, 796, 800, 798, 797, 796]
drnnGRUtanhMakespan39 = [794, 795, 795, 797, 795, 795, 794, 794, 798, 795, 794, 798]
drnnGRUtanhMakespan40 = [795, 795, 795, 795, 795, 795, 794, 794, 793, 797, 794, 795]
drnnGRUtanhMakespan41 = [794, 794, 795, 793, 795, 795, 792, 794, 795, 794, 794, 794]
drnnGRUtanhMakespan42 = [795, 795, 795, 796, 794, 797, 795, 795, 792, 795, 796, 793]
drnnGRUtanhMakespan43 = [794, 795, 795, 794, 795, 794, 798, 794, 797, 795, 794, 794]
drnnGRUtanhMakespan44 = [795, 795, 793, 794, 795, 794, 795, 795, 794, 794, 795, 794]
drnnGRUtanhMakespan45 = [794, 794, 794, 794, 794, 794, 795, 794, 794, 794, 796, 795]
drnnGRUtanhMakespan46 = [795, 794, 795, 794, 794, 794, 793, 794, 795, 795, 794, 797]
drnnGRUtanhMakespan47 = [794, 794, 794, 794, 795, 794, 795, 792, 794, 795, 794, 794]
drnnGRUtanhMakespan48 = [795, 794, 794, 794, 795, 798, 794, 794, 794, 795, 794, 794]
drnnGRUtanhMakespan49 = [795, 795, 794, 795, 793, 795, 796, 794, 795, 794, 794, 797]
drnnGRUtanhRewards0 = [-0.17580964970257765, -0.1759911894273128, -0.17580964970257765, -0.1768976897689769, -0.177078750549934, -0.1759911894273128, -0.1763540290620872, -0.1763540290620872, -0.1763540290620872, -0.1759911894273128, -0.17580964970257765, -0.17544633017412387]
drnnGRUtanhRewards1 = [-0.17617264919621228, -0.17580964970257765, -0.17580964970257765, -0.17580964970257765, -0.17580964970257765, -0.17580964970257765, -0.1763540290620872, -0.17580964970257765, -0.17526455026455026, -0.17544633017412387, -0.17617264919621228, -0.17544633017412387]
drnnGRUtanhRewards2 = [-0.17526455026455026, -0.1768976897689769, -0.177078750549934, -0.17617264919621228, -0.17617264919621228, -0.17544633017412387, -0.1768976897689769, -0.17617264919621228, -0.17526455026455026, -0.17580964970257765, -0.17580964970257765, -0.1763540290620872]
drnnGRUtanhRewards3 = [-0.17725973169122497, -0.17544633017412387, -0.17508269018743108, -0.17562802996914942, -0.17580964970257765, -0.17617264919621228, -0.17617264919621228, -0.1776214552648934, -0.177078750549934, -0.17580964970257765, -0.17617264919621228, -0.1778021978021978]
drnnGRUtanhRewards4 = [-0.177078750549934, -0.1763540290620872, -0.17526455026455026, -0.17580964970257765, -0.17580964970257765, -0.17617264919621228, -0.17544633017412387, -0.17544633017412387, -0.177078750549934, -0.17580964970257765, -0.1759911894273128, -0.17580964970257765]
drnnGRUtanhRewards5 = [-0.1768976897689769, -0.1759911894273128, -0.17580964970257765, -0.1768976897689769, -0.17544633017412387, -0.1759911894273128, -0.17580964970257765, -0.177078750549934, -0.17544633017412387, -0.177078750549934, -0.17580964970257765, -0.17617264919621228]
drnnGRUtanhRewards6 = [-0.17617264919621228, -0.1759911894273128, -0.17508269018743108, -0.1763540290620872, -0.1759911894273128, -0.17544633017412387, -0.17617264919621228, -0.1768976897689769, -0.17562802996914942, -0.17544633017412387, -0.17617264919621228, -0.17580964970257765]
drnnGRUtanhRewards7 = [-0.17580964970257765, -0.17617264919621228, -0.17798286090969018, -0.177078750549934, -0.17798286090969018, -0.17617264919621228, -0.17526455026455026, -0.17580964970257765, -0.17617264919621228, -0.177078750549934, -0.1759911894273128, -0.17617264919621228]
drnnGRUtanhRewards8 = [-0.17580964970257765, -0.17562802996914942, -0.17617264919621228, -0.17617264919621228, -0.1768976897689769, -0.177078750549934, -0.17580964970257765, -0.17580964970257765, -0.17526455026455026, -0.1763540290620872, -0.17580964970257765, -0.1778021978021978]
drnnGRUtanhRewards9 = [-0.17671654929577466, -0.17617264919621228, -0.17617264919621228, -0.177078750549934, -0.177078750549934, -0.17580964970257765, -0.1768976897689769, -0.17671654929577466, -0.177078750549934, -0.1763540290620872, -0.17798286090969018, -0.1763540290620872]
drnnGRUtanhRewards10 = [-0.17580964970257765, -0.1759911894273128, -0.17580964970257765, -0.17580964970257765, -0.177078750549934, -0.1768976897689769, -0.177078750549934, -0.17580964970257765, -0.1759911894273128, -0.17580964970257765, -0.17617264919621228, -0.17617264919621228]
drnnGRUtanhRewards11 = [-0.17544633017412387, -0.17526455026455026, -0.177078750549934, -0.17617264919621228, -0.17617264919621228, -0.17580964970257765, -0.17526455026455026, -0.1768976897689769, -0.177078750549934, -0.17580964970257765, -0.17617264919621228, -0.17617264919621228]
drnnGRUtanhRewards12 = [-0.1759911894273128, -0.1759911894273128, -0.1778021978021978, -0.17617264919621228, -0.1759911894273128, -0.1759911894273128, -0.17562802996914942, -0.177078750549934, -0.1759911894273128, -0.17617264919621228, -0.17580964970257765, -0.17526455026455026]
drnnGRUtanhRewards13 = [-0.177078750549934, -0.17617264919621228, -0.17617264919621228, -0.177078750549934, -0.17617264919621228, -0.1759911894273128, -0.17580964970257765, -0.1763540290620872, -0.17580964970257765, -0.17562802996914942, -0.177078750549934, -0.17617264919621228]
drnnGRUtanhRewards14 = [-0.17617264919621228, -0.17580964970257765, -0.17617264919621228, -0.17617264919621228, -0.17617264919621228, -0.1768976897689769, -0.1768976897689769, -0.1759911894273128, -0.1759911894273128, -0.17617264919621228, -0.17580964970257765, -0.17580964970257765]
drnnGRUtanhRewards15 = [-0.177078750549934, -0.17617264919621228, -0.17617264919621228, -0.17526455026455026, -0.1768976897689769, -0.17526455026455026, -0.17617264919621228, -0.17580964970257765, -0.1759911894273128, -0.17580964970257765, -0.17617264919621228, -0.17544633017412387]
drnnGRUtanhRewards16 = [-0.17725973169122497, -0.17526455026455026, -0.1763540290620872, -0.1759911894273128, -0.1759911894273128, -0.17544633017412387, -0.17544633017412387, -0.17508269018743108, -0.17653532907770195, -0.17544633017412387, -0.17617264919621228, -0.17653532907770195]
drnnGRUtanhRewards17 = [-0.17544633017412387, -0.17617264919621228, -0.17580964970257765, -0.17617264919621228, -0.17617264919621228, -0.17508269018743108, -0.1768976897689769, -0.177078750549934, -0.17580964970257765, -0.17798286090969018, -0.17617264919621228, -0.17580964970257765]
drnnGRUtanhRewards18 = [-0.17580964970257765, -0.17617264919621228, -0.17508269018743108, -0.17508269018743108, -0.17562802996914942, -0.17580964970257765, -0.17617264919621228, -0.177078750549934, -0.17580964970257765, -0.17580964970257765, -0.1768976897689769, -0.17580964970257765]
drnnGRUtanhRewards19 = [-0.17544633017412387, -0.17617264919621228, -0.17725973169122497, -0.1759911894273128, -0.17544633017412387, -0.17617264919621228, -0.17580964970257765, -0.177078750549934, -0.17580964970257765, -0.17562802996914942, -0.1759911894273128, -0.177078750549934]
drnnGRUtanhRewards20 = [-0.17617264919621228, -0.177078750549934, -0.1759911894273128, -0.17544633017412387, -0.177078750549934, -0.177078750549934, -0.177078750549934, -0.17508269018743108, -0.1778021978021978, -0.17544633017412387, -0.17617264919621228, -0.17562802996914942]
drnnGRUtanhRewards21 = [-0.17580964970257765, -0.17617264919621228, -0.17617264919621228, -0.17617264919621228, -0.17580964970257765, -0.1763540290620872, -0.17544633017412387, -0.1763540290620872, -0.1763540290620872, -0.1763540290620872, -0.17526455026455026, -0.1759911894273128]
drnnGRUtanhRewards22 = [-0.17580964970257765, -0.1763540290620872, -0.17562802996914942, -0.17617264919621228, -0.1759911894273128, -0.17526455026455026, -0.1759911894273128, -0.1759911894273128, -0.17617264919621228, -0.1763540290620872, -0.17617264919621228, -0.1759911894273128]
drnnGRUtanhRewards23 = [-0.17617264919621228, -0.17580964970257765, -0.1759911894273128, -0.177078750549934, -0.17508269018743108, -0.17617264919621228, -0.17580964970257765, -0.17544633017412387, -0.17544633017412387, -0.1768976897689769, -0.17617264919621228, -0.17508269018743108]
drnnGRUtanhRewards24 = [-0.17617264919621228, -0.17617264919621228, -0.17580964970257765, -0.1759911894273128, -0.177078750549934, -0.1768976897689769, -0.17580964970257765, -0.1763540290620872, -0.17617264919621228, -0.17580964970257765, -0.17580964970257765, -0.17580964970257765]
drnnGRUtanhRewards25 = [-0.17580964970257765, -0.17580964970257765, -0.17580964970257765, -0.17526455026455026, -0.17617264919621228, -0.17580964970257765, -0.17671654929577466, -0.17580964970257765, -0.17617264919621228, -0.1759911894273128, -0.17544633017412387, -0.17580964970257765]
drnnGRUtanhRewards26 = [-0.17544633017412387, -0.17580964970257765, -0.17580964970257765, -0.17580964970257765, -0.177078750549934, -0.17544633017412387, -0.17580964970257765, -0.17580964970257765, -0.177078750549934, -0.17526455026455026, -0.1763540290620872, -0.17544633017412387]
drnnGRUtanhRewards27 = [-0.17508269018743108, -0.17544633017412387, -0.17544633017412387, -0.17617264919621228, -0.17617264919621228, -0.17580964970257765, -0.17617264919621228, -0.17580964970257765, -0.17653532907770195, -0.17580964970257765, -0.17562802996914942, -0.17580964970257765]
drnnGRUtanhRewards28 = [-0.1759911894273128, -0.1759911894273128, -0.17617264919621228, -0.17617264919621228, -0.17580964970257765, -0.17653532907770195, -0.17580964970257765, -0.1759911894273128, -0.17526455026455026, -0.17526455026455026, -0.17508269018743108, -0.17580964970257765]
drnnGRUtanhRewards29 = [-0.17580964970257765, -0.17544633017412387, -0.17544633017412387, -0.17562802996914942, -0.17544633017412387, -0.17580964970257765, -0.17617264919621228, -0.17617264919621228, -0.17544633017412387, -0.17580964970257765, -0.17617264919621228, -0.17526455026455026]
drnnGRUtanhRewards30 = [-0.1759911894273128, -0.17580964970257765, -0.17526455026455026, -0.17526455026455026, -0.17617264919621228, -0.17526455026455026, -0.17580964970257765, -0.1759911894273128, -0.17580964970257765, -0.17580964970257765, -0.177078750549934, -0.1759911894273128]
drnnGRUtanhRewards31 = [-0.17526455026455026, -0.1759911894273128, -0.17508269018743108, -0.17508269018743108, -0.17544633017412387, -0.17526455026455026, -0.17526455026455026, -0.17508269018743108, -0.17580964970257765, -0.17562802996914942, -0.17580964970257765, -0.17526455026455026]
drnnGRUtanhRewards32 = [-0.17562802996914942, -0.17580964970257765, -0.17526455026455026, -0.17526455026455026, -0.17544633017412387, -0.17580964970257765, -0.17562802996914942, -0.17580964970257765, -0.17526455026455026, -0.17508269018743108, -0.17544633017412387, -0.17526455026455026]
drnnGRUtanhRewards33 = [-0.1759911894273128, -0.17526455026455026, -0.17508269018743108, -0.17508269018743108, -0.17526455026455026, -0.17580964970257765, -0.17580964970257765, -0.17562802996914942, -0.17617264919621228, -0.17544633017412387, -0.17526455026455026, -0.17508269018743108]
drnnGRUtanhRewards34 = [-0.17580964970257765, -0.17526455026455026, -0.17580964970257765, -0.17544633017412387, -0.17580964970257765, -0.17544633017412387, -0.17508269018743108, -0.17580964970257765, -0.17580964970257765, -0.17580964970257765, -0.17544633017412387, -0.17562802996914942]
drnnGRUtanhRewards35 = [-0.17526455026455026, -0.17580964970257765, -0.17544633017412387, -0.17580964970257765, -0.17508269018743108, -0.1763540290620872, -0.17526455026455026, -0.17617264919621228, -0.17526455026455026, -0.17617264919621228, -0.17508269018743108, -0.17617264919621228]
drnnGRUtanhRewards36 = [-0.17580964970257765, -0.1759911894273128, -0.17544633017412387, -0.17562802996914942, -0.17526455026455026, -0.17508269018743108, -0.17617264919621228, -0.17526455026455026, -0.17526455026455026, -0.17508269018743108, -0.17526455026455026, -0.17526455026455026]
drnnGRUtanhRewards37 = [-0.1759911894273128, -0.17580964970257765, -0.17526455026455026, -0.17526455026455026, -0.17508269018743108, -0.17526455026455026, -0.17526455026455026, -0.17544633017412387, -0.177078750549934, -0.17526455026455026, -0.17580964970257765, -0.17544633017412387]
drnnGRUtanhRewards38 = [-0.17580964970257765, -0.17508269018743108, -0.17526455026455026, -0.17526455026455026, -0.17526455026455026, -0.17544633017412387, -0.17526455026455026, -0.17544633017412387, -0.17617264919621228, -0.17580964970257765, -0.17562802996914942, -0.17544633017412387]
drnnGRUtanhRewards39 = [-0.17508269018743108, -0.17526455026455026, -0.17526455026455026, -0.17526455026455026, -0.17562802996914942, -0.17526455026455026, -0.17508269018743108, -0.17508269018743108, -0.17580964970257765, -0.17508269018743108, -0.17526455026455026, -0.17580964970257765]
drnnGRUtanhRewards40 = [-0.17526455026455026, -0.17526455026455026, -0.17526455026455026, -0.17526455026455026, -0.17526455026455026, -0.17526455026455026, -0.17508269018743108, -0.17508269018743108, -0.1749007498897221, -0.17562802996914942, -0.17508269018743108, -0.17526455026455026]
drnnGRUtanhRewards41 = [-0.17508269018743108, -0.17508269018743108, -0.17526455026455026, -0.1749007498897221, -0.17526455026455026, -0.17526455026455026, -0.17526455026455026, -0.17508269018743108, -0.17471872931833224, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108]
drnnGRUtanhRewards42 = [-0.17526455026455026, -0.17526455026455026, -0.17526455026455026, -0.17508269018743108, -0.17544633017412387, -0.17562802996914942, -0.17526455026455026, -0.17526455026455026, -0.17471872931833224, -0.17526455026455026, -0.17544633017412387, -0.1749007498897221]
drnnGRUtanhRewards43 = [-0.17508269018743108, -0.17526455026455026, -0.17526455026455026, -0.17508269018743108, -0.17526455026455026, -0.17508269018743108, -0.17580964970257765, -0.17508269018743108, -0.17562802996914942, -0.17526455026455026, -0.17508269018743108, -0.17508269018743108]
drnnGRUtanhRewards44 = [-0.17526455026455026, -0.17526455026455026, -0.1749007498897221, -0.17508269018743108, -0.17526455026455026, -0.17508269018743108, -0.17526455026455026, -0.17526455026455026, -0.17508269018743108, -0.17508269018743108, -0.17526455026455026, -0.17508269018743108]
drnnGRUtanhRewards45 = [-0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17526455026455026, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17544633017412387, -0.17526455026455026]
drnnGRUtanhRewards46 = [-0.17526455026455026, -0.17508269018743108, -0.17526455026455026, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.1749007498897221, -0.17508269018743108, -0.17526455026455026, -0.17526455026455026, -0.17508269018743108, -0.17562802996914942]
drnnGRUtanhRewards47 = [-0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17526455026455026, -0.17508269018743108, -0.17526455026455026, -0.17471872931833224, -0.17508269018743108, -0.17526455026455026, -0.17508269018743108, -0.17508269018743108]
drnnGRUtanhRewards48 = [-0.17526455026455026, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17526455026455026, -0.17580964970257765, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17526455026455026, -0.17508269018743108, -0.17508269018743108]
drnnGRUtanhRewards49 = [-0.17526455026455026, -0.17526455026455026, -0.17508269018743108, -0.17526455026455026, -0.1749007498897221, -0.17526455026455026, -0.17544633017412387, -0.17526455026455026, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17562802996914942]
# Deep Recurrent Reinforcement Learning: 1 capa GRU y 4 capas Dense, Funcion de activacion relu, 12 episodes, 50 iteraciones
drnnGRUreluMakespan0 = [800, 799, 798, 797, 798, 800, 800, 796, 800, 794, 800, 800]
drnnGRUreluMakespan1 = [798, 800, 805, 795, 799, 808, 795, 800, 796, 798, 799, 798]
drnnGRUreluMakespan2 = [799, 800, 806, 800, 800, 805, 805, 798, 799, 807, 800, 800]
drnnGRUreluMakespan3 = [798, 795, 799, 800, 800, 796, 798, 800, 800, 804, 805, 800]
drnnGRUreluMakespan4 = [811, 800, 799, 800, 805, 798, 798, 799, 796, 804, 805, 804]
drnnGRUreluMakespan5 = [799, 795, 797, 800, 798, 800, 800, 798, 800, 797, 800, 798]
drnnGRUreluMakespan6 = [798, 800, 798, 799, 797, 798, 800, 796, 801, 799, 795, 798]
drnnGRUreluMakespan7 = [800, 804, 795, 801, 796, 806, 805, 798, 800, 799, 799, 804]
drnnGRUreluMakespan8 = [800, 799, 799, 800, 805, 796, 800, 800, 810, 796, 800, 798]
drnnGRUreluMakespan9 = [794, 800, 799, 805, 800, 800, 798, 798, 796, 795, 798, 796]
drnnGRUreluMakespan10 = [798, 800, 798, 801, 795, 802, 796, 809, 800, 800, 798, 795]
drnnGRUreluMakespan11 = [804, 800, 799, 799, 798, 803, 798, 798, 805, 803, 800, 796]
drnnGRUreluMakespan12 = [800, 799, 805, 797, 798, 796, 799, 794, 799, 805, 799, 800]
drnnGRUreluMakespan13 = [796, 800, 798, 800, 795, 799, 800, 804, 800, 794, 805, 805]
drnnGRUreluMakespan14 = [800, 795, 796, 798, 798, 801, 805, 794, 800, 801, 801, 796]
drnnGRUreluMakespan15 = [798, 800, 796, 796, 798, 794, 797, 800, 796, 801, 795, 799]
drnnGRUreluMakespan16 = [800, 805, 794, 800, 799, 800, 805, 801, 798, 800, 801, 799]
drnnGRUreluMakespan17 = [797, 803, 801, 808, 794, 799, 799, 800, 805, 796, 801, 796]
drnnGRUreluMakespan18 = [805, 800, 800, 804, 799, 798, 800, 799, 804, 796, 800, 804]
drnnGRUreluMakespan19 = [804, 798, 800, 799, 799, 799, 805, 795, 801, 799, 799, 805]
drnnGRUreluMakespan20 = [799, 804, 796, 798, 796, 798, 800, 805, 799, 810, 800, 800]
drnnGRUreluMakespan21 = [798, 799, 799, 805, 798, 798, 805, 798, 794, 799, 798, 798]
drnnGRUreluMakespan22 = [799, 798, 798, 796, 798, 805, 799, 798, 798, 799, 796, 798]
drnnGRUreluMakespan23 = [798, 805, 808, 798, 798, 805, 810, 796, 804, 799, 800, 799]
drnnGRUreluMakespan24 = [798, 796, 798, 795, 800, 798, 799, 798, 797, 805, 798, 800]
drnnGRUreluMakespan25 = [799, 796, 799, 798, 805, 798, 798, 800, 796, 794, 810, 798]
drnnGRUreluMakespan26 = [799, 798, 805, 800, 802, 798, 799, 799, 799, 794, 802, 797]
drnnGRUreluMakespan27 = [798, 800, 805, 796, 798, 795, 802, 796, 798, 800, 798, 794]
drnnGRUreluMakespan28 = [796, 805, 798, 800, 800, 798, 810, 798, 798, 798, 796, 796]
drnnGRUreluMakespan29 = [800, 798, 798, 802, 794, 798, 796, 808, 800, 800, 798, 799]
drnnGRUreluMakespan30 = [798, 796, 798, 798, 794, 798, 794, 800, 796, 794, 800, 800]
drnnGRUreluMakespan31 = [794, 802, 797, 799, 798, 800, 799, 799, 796, 796, 798, 798]
drnnGRUreluMakespan32 = [799, 798, 794, 795, 798, 805, 804, 797, 795, 800, 796, 798]
drnnGRUreluMakespan33 = [803, 799, 805, 796, 794, 798, 797, 798, 798, 794, 794, 798]
drnnGRUreluMakespan34 = [810, 796, 795, 798, 799, 798, 796, 795, 795, 797, 798, 798]
drnnGRUreluMakespan35 = [799, 799, 799, 799, 795, 798, 795, 800, 796, 795, 795, 796]
drnnGRUreluMakespan36 = [795, 797, 798, 799, 799, 799, 800, 794, 796, 795, 798, 800]
drnnGRUreluMakespan37 = [800, 798, 799, 794, 800, 796, 798, 798, 797, 800, 794, 798]
drnnGRUreluMakespan38 = [800, 799, 794, 796, 795, 800, 796, 804, 800, 795, 800, 798]
drnnGRUreluMakespan39 = [794, 798, 795, 804, 805, 799, 798, 800, 796, 798, 795, 794]
drnnGRUreluMakespan40 = [799, 798, 796, 798, 798, 799, 800, 796, 798, 798, 799, 798]
drnnGRUreluMakespan41 = [796, 798, 800, 797, 799, 796, 797, 796, 799, 804, 805, 798]
drnnGRUreluMakespan42 = [798, 794, 795, 799, 799, 798, 797, 798, 798, 798, 798, 795]
drnnGRUreluMakespan43 = [799, 798, 794, 794, 795, 794, 795, 799, 799, 800, 799, 794]
drnnGRUreluMakespan44 = [795, 796, 795, 799, 794, 795, 794, 796, 795, 794, 795, 796]
drnnGRUreluMakespan45 = [794, 797, 794, 795, 796, 795, 794, 799, 795, 794, 798, 798]
drnnGRUreluMakespan46 = [795, 795, 794, 795, 794, 794, 792, 794, 795, 797, 794, 794]
drnnGRUreluMakespan47 = [798, 796, 797, 798, 794, 798, 794, 797, 794, 803, 798, 798]
drnnGRUreluMakespan48 = [795, 794, 796, 798, 795, 794, 796, 795, 796, 794, 796, 796]
drnnGRUreluMakespan49 = [798, 798, 796, 798, 798, 796, 796, 798, 798, 798, 796, 798]
drnnGRUreluRewards0 = [-0.17617264919621228, -0.1759911894273128, -0.17580964970257765, -0.17562802996914942, -0.17580964970257765, -0.17617264919621228, -0.17617264919621228, -0.17544633017412387, -0.17617264919621228, -0.17508269018743108, -0.17617264919621228, -0.17617264919621228]
drnnGRUreluRewards1 = [-0.17580964970257765, -0.17617264919621228, -0.177078750549934, -0.17526455026455026, -0.1759911894273128, -0.1776214552648934, -0.17526455026455026, -0.17617264919621228, -0.17544633017412387, -0.17580964970257765, -0.1759911894273128, -0.17580964970257765]
drnnGRUreluRewards2 = [-0.1759911894273128, -0.17617264919621228, -0.17725973169122497, -0.17617264919621228, -0.17617264919621228, -0.177078750549934, -0.177078750549934, -0.17580964970257765, -0.1759911894273128, -0.1774406332453826, -0.17617264919621228, -0.17617264919621228]
drnnGRUreluRewards3 = [-0.17580964970257765, -0.17526455026455026, -0.1759911894273128, -0.17617264919621228, -0.17617264919621228, -0.17544633017412387, -0.17580964970257765, -0.17617264919621228, -0.17617264919621228, -0.1768976897689769, -0.177078750549934, -0.17617264919621228]
drnnGRUreluRewards4 = [-0.1781634446397188, -0.17617264919621228, -0.1759911894273128, -0.17617264919621228, -0.177078750549934, -0.17580964970257765, -0.17580964970257765, -0.1759911894273128, -0.17544633017412387, -0.1768976897689769, -0.177078750549934, -0.1768976897689769]
drnnGRUreluRewards5 = [-0.1759911894273128, -0.17526455026455026, -0.17562802996914942, -0.17617264919621228, -0.17580964970257765, -0.17617264919621228, -0.17617264919621228, -0.17580964970257765, -0.17617264919621228, -0.17562802996914942, -0.17617264919621228, -0.17580964970257765]
drnnGRUreluRewards6 = [-0.17580964970257765, -0.17617264919621228, -0.17580964970257765, -0.1759911894273128, -0.17562802996914942, -0.17580964970257765, -0.17544633017412387, -0.17617264919621228, -0.1763540290620872, -0.1759911894273128, -0.17526455026455026, -0.17580964970257765]
drnnGRUreluRewards7 = [-0.17617264919621228, -0.1768976897689769, -0.17526455026455026, -0.1763540290620872, -0.17544633017412387, -0.17725973169122497, -0.177078750549934, -0.17580964970257765, -0.17617264919621228, -0.1759911894273128, -0.1759911894273128, -0.1768976897689769]
drnnGRUreluRewards8 = [-0.17617264919621228, -0.1759911894273128, -0.1759911894273128, -0.17617264919621228, -0.177078750549934, -0.17544633017412387, -0.17617264919621228, -0.17617264919621228, -0.17798286090969018, -0.17544633017412387, -0.17617264919621228, -0.17580964970257765]
drnnGRUreluRewards9 = [-0.17508269018743108, -0.17617264919621228, -0.1759911894273128, -0.177078750549934, -0.17617264919621228, -0.17617264919621228, -0.17580964970257765, -0.17580964970257765, -0.17544633017412387, -0.17526455026455026, -0.17580964970257765, -0.17544633017412387]
drnnGRUreluRewards10 = [-0.17580964970257765, -0.17617264919621228, -0.17580964970257765, -0.1763540290620872, -0.17526455026455026, -0.17653532907770195, -0.17544633017412387, -0.1778021978021978, -0.17617264919621228, -0.17617264919621228, -0.17580964970257765, -0.17526455026455026]
drnnGRUreluRewards11 = [-0.1768976897689769, -0.17617264919621228, -0.1759911894273128, -0.1759911894273128, -0.17580964970257765, -0.17671654929577466, -0.17580964970257765, -0.17580964970257765, -0.177078750549934, -0.17671654929577466, -0.17617264919621228, -0.17544633017412387]
drnnGRUreluRewards12 = [-0.17617264919621228, -0.1759911894273128, -0.177078750549934, -0.17562802996914942, -0.17580964970257765, -0.17544633017412387, -0.1759911894273128, -0.17508269018743108, -0.1759911894273128, -0.177078750549934, -0.1759911894273128, -0.17617264919621228]
drnnGRUreluRewards13 = [-0.17544633017412387, -0.17617264919621228, -0.17580964970257765, -0.17617264919621228, -0.17526455026455026, -0.1759911894273128, -0.17617264919621228, -0.1768976897689769, -0.17617264919621228, -0.17508269018743108, -0.177078750549934, -0.177078750549934]
drnnGRUreluRewards14 = [-0.17617264919621228, -0.17526455026455026, -0.17544633017412387, -0.17580964970257765, -0.17580964970257765, -0.1763540290620872, -0.177078750549934, -0.17508269018743108, -0.17617264919621228, -0.1763540290620872, -0.1763540290620872, -0.17544633017412387]
drnnGRUreluRewards15 = [-0.17580964970257765, -0.17617264919621228, -0.17544633017412387, -0.17544633017412387, -0.17580964970257765, -0.17508269018743108, -0.17562802996914942, -0.17617264919621228, -0.17544633017412387, -0.1763540290620872, -0.17526455026455026, -0.1759911894273128]
drnnGRUreluRewards16 = [-0.17617264919621228, -0.177078750549934, -0.17508269018743108, -0.17617264919621228, -0.1759911894273128, -0.17617264919621228, -0.177078750549934, -0.1763540290620872, -0.17580964970257765, -0.17617264919621228, -0.1763540290620872, -0.1759911894273128]
drnnGRUreluRewards17 = [-0.17562802996914942, -0.17671654929577466, -0.1763540290620872, -0.1776214552648934, -0.17508269018743108, -0.1759911894273128, -0.17617264919621228, -0.1759911894273128, -0.177078750549934, -0.17544633017412387, -0.1763540290620872, -0.17544633017412387]
drnnGRUreluRewards18 = [-0.177078750549934, -0.17617264919621228, -0.17617264919621228, -0.1768976897689769, -0.1759911894273128, -0.17580964970257765, -0.17617264919621228, -0.1759911894273128, -0.1768976897689769, -0.17544633017412387, -0.17617264919621228, -0.1768976897689769]
drnnGRUreluRewards19 = [-0.1768976897689769, -0.17580964970257765, -0.17617264919621228, -0.1759911894273128, -0.1759911894273128, -0.1759911894273128, -0.177078750549934, -0.17526455026455026, -0.1763540290620872, -0.1759911894273128, -0.1759911894273128, -0.177078750549934]
drnnGRUreluRewards20 = [-0.1759911894273128, -0.1768976897689769, -0.17544633017412387, -0.17580964970257765, -0.17544633017412387, -0.17580964970257765, -0.17617264919621228, -0.177078750549934, -0.1759911894273128, -0.17798286090969018, -0.17617264919621228, -0.17617264919621228]
drnnGRUreluRewards21 = [-0.17580964970257765, -0.1759911894273128, -0.1759911894273128, -0.177078750549934, -0.17580964970257765, -0.17580964970257765, -0.177078750549934, -0.17580964970257765, -0.17508269018743108, -0.1759911894273128, -0.17580964970257765, -0.17580964970257765]
drnnGRUreluRewards22 = [-0.1759911894273128, -0.17580964970257765, -0.17580964970257765, -0.17544633017412387, -0.17580964970257765, -0.177078750549934, -0.1759911894273128, -0.17580964970257765, -0.17580964970257765, -0.1759911894273128, -0.17544633017412387, -0.17580964970257765]
drnnGRUreluRewards23 = [-0.17580964970257765, -0.177078750549934, -0.1776214552648934, -0.17580964970257765, -0.17580964970257765, -0.177078750549934, -0.17798286090969018, -0.17544633017412387, -0.1768976897689769, -0.1759911894273128, -0.17617264919621228, -0.1759911894273128]
drnnGRUreluRewards24 = [-0.17580964970257765, -0.17544633017412387, -0.17580964970257765, -0.17526455026455026, -0.17617264919621228, -0.17580964970257765, -0.1759911894273128, -0.17580964970257765, -0.17562802996914942, -0.177078750549934, -0.17580964970257765, -0.17617264919621228]
drnnGRUreluRewards25 = [-0.1759911894273128, -0.17544633017412387, -0.1759911894273128, -0.17580964970257765, -0.177078750549934, -0.17580964970257765, -0.17580964970257765, -0.17617264919621228, -0.17544633017412387, -0.17508269018743108, -0.17798286090969018, -0.17580964970257765]
drnnGRUreluRewards26 = [-0.1759911894273128, -0.17580964970257765, -0.177078750549934, -0.17617264919621228, -0.17653532907770195, -0.17580964970257765, -0.1759911894273128, -0.1759911894273128, -0.1759911894273128, -0.17508269018743108, -0.17653532907770195, -0.17562802996914942]
drnnGRUreluRewards27 = [-0.17580964970257765, -0.17617264919621228, -0.177078750549934, -0.17544633017412387, -0.17580964970257765, -0.17526455026455026, -0.17653532907770195, -0.17544633017412387, -0.17580964970257765, -0.17617264919621228, -0.17580964970257765, -0.17508269018743108]
drnnGRUreluRewards28 = [-0.17544633017412387, -0.177078750549934, -0.17580964970257765, -0.17617264919621228, -0.17617264919621228, -0.17580964970257765, -0.17798286090969018, -0.17580964970257765, -0.17580964970257765, -0.17580964970257765, -0.17544633017412387, -0.17544633017412387]
drnnGRUreluRewards29 = [-0.17617264919621228, -0.17580964970257765, -0.17580964970257765, -0.17653532907770195, -0.17508269018743108, -0.17580964970257765, -0.17544633017412387, -0.1776214552648934, -0.17617264919621228, -0.17617264919621228, -0.17580964970257765, -0.1759911894273128]
drnnGRUreluRewards30 = [-0.17580964970257765, -0.17544633017412387, -0.17580964970257765, -0.17580964970257765, -0.17508269018743108, -0.17580964970257765, -0.17508269018743108, -0.17617264919621228, -0.17544633017412387, -0.17508269018743108, -0.17617264919621228, -0.17617264919621228]
drnnGRUreluRewards31 = [-0.17508269018743108, -0.17653532907770195, -0.17562802996914942, -0.1759911894273128, -0.17580964970257765, -0.17617264919621228, -0.1759911894273128, -0.1759911894273128, -0.17544633017412387, -0.17544633017412387, -0.17580964970257765, -0.17580964970257765]
drnnGRUreluRewards32 = [-0.1759911894273128, -0.17580964970257765, -0.17508269018743108, -0.17526455026455026, -0.17580964970257765, -0.1768976897689769, -0.177078750549934, -0.17562802996914942, -0.17526455026455026, -0.17617264919621228, -0.17544633017412387, -0.17580964970257765]
drnnGRUreluRewards33 = [-0.17671654929577466, -0.1759911894273128, -0.177078750549934, -0.17544633017412387, -0.17508269018743108, -0.17580964970257765, -0.17562802996914942, -0.17580964970257765, -0.17580964970257765, -0.17508269018743108, -0.17508269018743108, -0.17580964970257765]
drnnGRUreluRewards34 = [-0.17798286090969018, -0.17544633017412387, -0.17526455026455026, -0.17580964970257765, -0.1759911894273128, -0.17580964970257765, -0.17544633017412387, -0.17526455026455026, -0.17526455026455026, -0.17562802996914942, -0.17580964970257765, -0.17580964970257765]
drnnGRUreluRewards35 = [-0.1759911894273128, -0.1759911894273128, -0.1759911894273128, -0.1759911894273128, -0.17526455026455026, -0.17580964970257765, -0.17526455026455026, -0.17617264919621228, -0.17544633017412387, -0.17526455026455026, -0.17526455026455026, -0.17544633017412387]
drnnGRUreluRewards36 = [-0.17526455026455026, -0.17562802996914942, -0.17580964970257765, -0.1759911894273128, -0.1759911894273128, -0.1759911894273128, -0.17617264919621228, -0.17508269018743108, -0.17544633017412387, -0.17526455026455026, -0.17580964970257765, -0.17617264919621228]
drnnGRUreluRewards37 = [-0.17617264919621228, -0.17580964970257765, -0.1759911894273128, -0.17508269018743108, -0.17617264919621228, -0.17544633017412387, -0.17580964970257765, -0.17580964970257765, -0.17562802996914942, -0.17617264919621228, -0.17508269018743108, -0.17580964970257765]
drnnGRUreluRewards38 = [-0.17617264919621228, -0.1759911894273128, -0.17508269018743108, -0.17544633017412387, -0.17526455026455026, -0.17617264919621228, -0.17544633017412387, -0.1768976897689769, -0.17617264919621228, -0.17526455026455026, -0.17617264919621228, -0.17580964970257765]
drnnGRUreluRewards39 = [-0.17508269018743108, -0.17580964970257765, -0.17526455026455026, -0.1768976897689769, -0.177078750549934, -0.1759911894273128, -0.17580964970257765, -0.17617264919621228, -0.17544633017412387, -0.17580964970257765, -0.17526455026455026, -0.17508269018743108]
drnnGRUreluRewards40 = [-0.1759911894273128, -0.17580964970257765, -0.17544633017412387, -0.17580964970257765, -0.17580964970257765, -0.1759911894273128, -0.17617264919621228, -0.17544633017412387, -0.17580964970257765, -0.17580964970257765, -0.1759911894273128, -0.17580964970257765]
drnnGRUreluRewards41 = [-0.17544633017412387, -0.17580964970257765, -0.17617264919621228, -0.17562802996914942, -0.1759911894273128, -0.17544633017412387, -0.17562802996914942, -0.17544633017412387, -0.1759911894273128, -0.1768976897689769, -0.177078750549934, -0.17580964970257765]
drnnGRUreluRewards42 = [-0.17580964970257765, -0.17508269018743108, -0.17526455026455026, -0.1759911894273128, -0.1759911894273128, -0.17580964970257765, -0.17562802996914942, -0.17580964970257765, -0.17580964970257765, -0.17580964970257765, -0.17580964970257765, -0.17526455026455026]
drnnGRUreluRewards43 = [-0.1759911894273128, -0.17580964970257765, -0.17508269018743108, -0.17508269018743108, -0.17526455026455026, -0.17508269018743108, -0.17526455026455026, -0.1759911894273128, -0.1759911894273128, -0.17617264919621228, -0.1759911894273128, -0.17508269018743108]
drnnGRUreluRewards44 = [-0.17526455026455026, -0.17544633017412387, -0.17526455026455026, -0.1759911894273128, -0.17508269018743108, -0.17526455026455026, -0.17508269018743108, -0.17544633017412387, -0.17526455026455026, -0.17508269018743108, -0.17526455026455026, -0.17544633017412387]
drnnGRUreluRewards45 = [-0.17508269018743108, -0.17562802996914942, -0.17508269018743108, -0.17526455026455026, -0.17544633017412387, -0.17526455026455026, -0.17508269018743108, -0.1759911894273128, -0.17526455026455026, -0.17508269018743108, -0.17580964970257765, -0.17580964970257765]
drnnGRUreluRewards46 = [-0.17526455026455026, -0.17526455026455026, -0.17508269018743108, -0.17526455026455026, -0.17508269018743108, -0.17508269018743108, -0.17471872931833224, -0.17508269018743108, -0.17526455026455026, -0.17562802996914942, -0.17508269018743108, -0.17508269018743108]
drnnGRUreluRewards47 = [-0.17580964970257765, -0.17544633017412387, -0.17562802996914942, -0.17580964970257765, -0.17508269018743108, -0.17580964970257765, -0.17508269018743108, -0.17562802996914942, -0.17508269018743108, -0.17671654929577466, -0.17580964970257765, -0.17580964970257765]
drnnGRUreluRewards48 = [-0.17526455026455026, -0.17508269018743108, -0.17544633017412387, -0.17580964970257765, -0.17526455026455026, -0.17508269018743108, -0.17544633017412387, -0.17526455026455026, -0.17544633017412387, -0.17508269018743108, -0.17544633017412387, -0.17544633017412387]
drnnGRUreluRewards49 = [-0.17580964970257765, -0.17580964970257765, -0.17544633017412387, -0.17580964970257765, -0.17580964970257765, -0.17544633017412387, -0.17544633017412387, -0.17580964970257765, -0.17580964970257765, -0.17580964970257765, -0.17544633017412387, -0.17580964970257765]
# Deep Reinforcement Learning: 5 capas Dense, Funcion de activacion tanh, 12 episodios, 50 iteraciones
drlTanhMakespan0 = [794, 794, 805, 799, 810, 800, 794, 810, 804, 806, 812, 808]
drlTanhMakespan1 = [796, 795, 795, 798, 799, 800, 800, 795, 797, 796, 797, 799]
drlTanhMakespan2 = [800, 797, 798, 801, 799, 800, 796, 795, 797, 796, 794, 798]
drlTanhMakespan3 = [800, 795, 799, 796, 799, 798, 795, 799, 795, 799, 798, 796]
drlTanhMakespan4 = [809, 795, 795, 800, 797, 795, 798, 798, 799, 799, 798, 798]
drlTanhMakespan5 = [795, 795, 795, 799, 795, 798, 795, 800, 795, 796, 795, 805]
drlTanhMakespan6 = [794, 800, 795, 793, 798, 795, 794, 798, 795, 799, 795, 796]
drlTanhMakespan7 = [795, 795, 795, 795, 798, 795, 797, 797, 795, 795, 798, 797]
drlTanhMakespan8 = [795, 795, 795, 794, 800, 800, 794, 795, 794, 794, 797, 795]
drlTanhMakespan9 = [793, 794, 796, 795, 796, 800, 794, 797, 793, 795, 798, 795]
drlTanhMakespan10 = [795, 795, 797, 794, 795, 798, 797, 795, 798, 794, 794, 794]
drlTanhMakespan11 = [795, 795, 795, 795, 797, 795, 795, 794, 795, 795, 795, 794]
drlTanhMakespan12 = [794, 798, 795, 794, 795, 795, 795, 797, 799, 795, 795, 795]
drlTanhMakespan13 = [795, 797, 795, 800, 796, 795, 796, 795, 795, 795, 798, 794]
drlTanhMakespan14 = [795, 795, 796, 794, 794, 794, 797, 795, 798, 795, 795, 793]
drlTanhMakespan15 = [799, 794, 795, 795, 795, 796, 801, 797, 795, 794, 795, 799]
drlTanhMakespan16 = [795, 795, 796, 798, 795, 795, 795, 795, 795, 798, 798, 796]
drlTanhMakespan17 = [800, 798, 795, 795, 798, 794, 795, 795, 797, 795, 796, 794]
drlTanhMakespan18 = [797, 800, 798, 797, 796, 794, 799, 797, 795, 796, 799, 798]
drlTanhMakespan19 = [797, 800, 795, 794, 794, 796, 795, 798, 796, 798, 797, 795]
drlTanhMakespan20 = [794, 795, 795, 799, 798, 797, 795, 795, 798, 795, 798, 795]
drlTanhMakespan21 = [796, 795, 795, 795, 795, 797, 798, 794, 797, 795, 796, 794]
drlTanhMakespan22 = [799, 796, 795, 795, 795, 795, 796, 795, 796, 798, 796, 795]
drlTanhMakespan23 = [799, 799, 795, 796, 796, 799, 796, 797, 794, 794, 798, 796]
drlTanhMakespan24 = [795, 795, 797, 800, 797, 795, 795, 796, 795, 795, 798, 799]
drlTanhMakespan25 = [795, 797, 795, 795, 795, 795, 800, 796, 795, 797, 795, 795]
drlTanhMakespan26 = [795, 795, 799, 794, 797, 794, 794, 798, 794, 796, 795, 798]
drlTanhMakespan27 = [796, 796, 795, 796, 798, 797, 794, 795, 794, 794, 794, 798]
drlTanhMakespan28 = [795, 795, 794, 798, 796, 796, 800, 797, 797, 796, 795, 794]
drlTanhMakespan29 = [795, 795, 798, 800, 797, 794, 796, 794, 792, 794, 794, 795]
drlTanhMakespan30 = [798, 797, 795, 799, 797, 800, 798, 799, 797, 800, 794, 796]
drlTanhMakespan31 = [794, 795, 800, 798, 800, 794, 800, 798, 799, 798, 798, 798]
drlTanhMakespan32 = [795, 795, 795, 794, 794, 794, 793, 795, 794, 793, 794, 795]
drlTanhMakespan33 = [794, 797, 792, 794, 795, 795, 797, 795, 795, 794, 792, 795]
drlTanhMakespan34 = [795, 794, 795, 798, 795, 796, 794, 795, 794, 794, 795, 794]
drlTanhMakespan35 = [796, 794, 797, 793, 794, 798, 795, 794, 793, 793, 795, 794]
drlTanhMakespan36 = [795, 795, 794, 795, 795, 795, 794, 795, 795, 793, 795, 794]
drlTanhMakespan37 = [794, 794, 798, 794, 794, 796, 795, 794, 793, 795, 795, 792]
drlTanhMakespan38 = [794, 796, 795, 794, 798, 798, 795, 795, 794, 794, 795, 794]
drlTanhMakespan39 = [794, 795, 795, 796, 792, 794, 795, 794, 795, 794, 794, 795]
drlTanhMakespan40 = [798, 795, 794, 795, 794, 794, 793, 795, 794, 794, 797, 794]
drlTanhMakespan41 = [795, 792, 795, 794, 794, 795, 794, 795, 792, 797, 795, 795]
drlTanhMakespan42 = [792, 794, 794, 795, 794, 794, 795, 794, 792, 794, 794, 794]
drlTanhMakespan43 = [794, 796, 794, 793, 795, 795, 793, 798, 794, 794, 798, 794]
drlTanhMakespan44 = [794, 794, 794, 794, 795, 794, 793, 794, 794, 795, 795, 794]
drlTanhMakespan45 = [790, 794, 793, 794, 793, 794, 795, 794, 791, 795, 795, 794]
drlTanhMakespan46 = [792, 794, 794, 794, 794, 794, 794, 793, 794, 794, 794, 794]
drlTanhMakespan47 = [794, 794, 794, 794, 794, 794, 794, 794, 792, 795, 793, 795]
drlTanhMakespan48 = [794, 794, 792, 792, 797, 794, 792, 794, 794, 795, 794, 795]
drlTanhMakespan49 = [795, 794, 794, 796, 794, 797, 794, 794, 794, 794, 794, 794]
drlTanhMakespan50 = [794, 792, 795, 794, 794, 794, 794, 794, 795, 794, 795, 794]
drlTanhMakespan51 = [794, 792, 796, 795, 794, 794, 795, 794, 795, 795, 795, 794]
drlTanhMakespan52 = [794, 794, 795, 792, 795, 795, 795, 792, 794, 793, 795, 794]
drlTanhMakespan53 = [794, 792, 794, 792, 794, 794, 794, 795, 795, 794, 794, 792]
drlTanhMakespan54 = [795, 793, 794, 794, 794, 792, 795, 794, 794, 792, 794, 796]
drlTanhMakespan55 = [795, 794, 794, 795, 795, 793, 794, 795, 794, 797, 795, 792]
drlTanhMakespan56 = [795, 795, 792, 795, 794, 795, 794, 794, 794, 795, 795, 795]
drlTanhMakespan57 = [795, 792, 795, 794, 795, 795, 792, 795, 794, 797, 792, 792]
drlTanhMakespan58 = [795, 795, 794, 795, 792, 794, 794, 794, 792, 792, 792, 793]
drlTanhMakespan59 = [795, 794, 792, 794, 794, 794, 792, 794, 794, 794, 793, 795]
drlTanhMakespan60 = [794, 795, 795, 795, 798, 794, 794, 794, 794, 794, 794, 792]
drlTanhMakespan61 = [792, 795, 794, 794, 795, 794, 792, 795, 795, 794, 794, 795]
drlTanhMakespan62 = [795, 794, 794, 794, 799, 794, 792, 794, 795, 795, 794, 793]
drlTanhMakespan63 = [791, 795, 792, 796, 794, 794, 792, 795, 793, 794, 792, 794]
drlTanhRewards0 = [-0.17508269018743108, -0.17508269018743108, -0.177078750549934, -0.1759911894273128, -0.17798286090969018, -0.17617264919621228, -0.17508269018743108, -0.17798286090969018, -0.1768976897689769, -0.17725973169122497, -0.17834394904458598, -0.1776214552648934]
drlTanhRewards1 = [-0.17544633017412387, -0.17526455026455026, -0.17526455026455026, -0.17580964970257765, -0.17617264919621228, -0.1759911894273128, -0.17617264919621228, -0.17526455026455026, -0.17562802996914942, -0.17544633017412387, -0.17562802996914942, -0.1759911894273128]
drlTanhRewards2 = [-0.17617264919621228, -0.17562802996914942, -0.17580964970257765, -0.1763540290620872, -0.1759911894273128, -0.17617264919621228, -0.17544633017412387, -0.17526455026455026, -0.17562802996914942, -0.17508269018743108, -0.17544633017412387, -0.17580964970257765]
drlTanhRewards3 = [-0.17617264919621228, -0.1759911894273128, -0.17526455026455026, -0.17544633017412387, -0.1759911894273128, -0.17580964970257765, -0.17526455026455026, -0.1759911894273128, -0.17526455026455026, -0.1759911894273128, -0.17580964970257765, -0.17544633017412387]
drlTanhRewards4 = [-0.1778021978021978, -0.17526455026455026, -0.17526455026455026, -0.17617264919621228, -0.17562802996914942, -0.17580964970257765, -0.17526455026455026, -0.17580964970257765, -0.1759911894273128, -0.17580964970257765, -0.1759911894273128, -0.17580964970257765]
drlTanhRewards5 = [-0.17526455026455026, -0.17526455026455026, -0.17526455026455026, -0.1759911894273128, -0.17526455026455026, -0.17580964970257765, -0.17526455026455026, -0.17617264919621228, -0.17526455026455026, -0.17544633017412387, -0.17526455026455026, -0.177078750549934]
drlTanhRewards6 = [-0.17508269018743108, -0.17617264919621228, -0.17526455026455026, -0.1749007498897221, -0.17580964970257765, -0.17526455026455026, -0.17508269018743108, -0.1759911894273128, -0.17526455026455026, -0.17580964970257765, -0.17526455026455026, -0.17544633017412387]
drlTanhRewards7 = [-0.17526455026455026, -0.17526455026455026, -0.17526455026455026, -0.17526455026455026, -0.17580964970257765, -0.17562802996914942, -0.17526455026455026, -0.17562802996914942, -0.17526455026455026, -0.17526455026455026, -0.17580964970257765, -0.17562802996914942]
drlTanhRewards8 = [-0.17526455026455026, -0.17526455026455026, -0.17526455026455026, -0.17508269018743108, -0.17617264919621228, -0.17508269018743108, -0.17617264919621228, -0.17508269018743108, -0.17526455026455026, -0.17508269018743108, -0.17562802996914942, -0.17526455026455026]
drlTanhRewards9 = [-0.1749007498897221, -0.17508269018743108, -0.17544633017412387, -0.17526455026455026, -0.17544633017412387, -0.17617264919621228, -0.17508269018743108, -0.17562802996914942, -0.17526455026455026, -0.1749007498897221, -0.17580964970257765, -0.17526455026455026]
drlTanhRewards10 = [-0.17526455026455026, -0.17526455026455026, -0.17562802996914942, -0.17508269018743108, -0.17526455026455026, -0.17580964970257765, -0.17562802996914942, -0.17580964970257765, -0.17526455026455026, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108]
drlTanhRewards11 = [-0.17526455026455026, -0.17526455026455026, -0.17526455026455026, -0.17526455026455026, -0.17562802996914942, -0.17526455026455026, -0.17526455026455026, -0.17508269018743108, -0.17526455026455026, -0.17526455026455026, -0.17508269018743108, -0.17526455026455026]
drlTanhRewards12 = [-0.17508269018743108, -0.17580964970257765, -0.17508269018743108, -0.17526455026455026, -0.17526455026455026, -0.17526455026455026, -0.17526455026455026, -0.17562802996914942, -0.1759911894273128, -0.17526455026455026, -0.17526455026455026, -0.17526455026455026]
drlTanhRewards13 = [-0.17526455026455026, -0.17526455026455026, -0.17562802996914942, -0.17617264919621228, -0.17544633017412387, -0.17544633017412387, -0.17526455026455026, -0.17526455026455026, -0.17526455026455026, -0.17526455026455026, -0.17580964970257765, -0.17508269018743108]
drlTanhRewards14 = [-0.17526455026455026, -0.17526455026455026, -0.17544633017412387, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17526455026455026, -0.17562802996914942, -0.17526455026455026, -0.17580964970257765, -0.17526455026455026, -0.1749007498897221]
drlTanhRewards15 = [-0.1759911894273128, -0.17508269018743108, -0.17526455026455026, -0.17526455026455026, -0.17526455026455026, -0.17544633017412387, -0.17562802996914942, -0.1763540290620872, -0.17526455026455026, -0.17508269018743108, -0.17526455026455026, -0.1759911894273128]
drlTanhRewards16 = [-0.17526455026455026, -0.17526455026455026, -0.17544633017412387, -0.17580964970257765, -0.17526455026455026, -0.17526455026455026, -0.17526455026455026, -0.17526455026455026, -0.17580964970257765, -0.17526455026455026, -0.17580964970257765, -0.17544633017412387]
drlTanhRewards17 = [-0.17617264919621228, -0.17580964970257765, -0.17526455026455026, -0.17526455026455026, -0.17508269018743108, -0.17580964970257765, -0.17526455026455026, -0.17562802996914942, -0.17526455026455026, -0.17526455026455026, -0.17544633017412387, -0.17508269018743108]
drlTanhRewards18 = [-0.17562802996914942, -0.17617264919621228, -0.17580964970257765, -0.17562802996914942, -0.17544633017412387, -0.1759911894273128, -0.17508269018743108, -0.17562802996914942, -0.17526455026455026, -0.17544633017412387, -0.1759911894273128, -0.17580964970257765]
drlTanhRewards19 = [-0.17562802996914942, -0.17617264919621228, -0.17526455026455026, -0.17508269018743108, -0.17508269018743108, -0.17544633017412387, -0.17526455026455026, -0.17544633017412387, -0.17580964970257765, -0.17580964970257765, -0.17562802996914942, -0.17526455026455026]
drlTanhRewards20 = [-0.17508269018743108, -0.17526455026455026, -0.17526455026455026, -0.1759911894273128, -0.17580964970257765, -0.17562802996914942, -0.17526455026455026, -0.17526455026455026, -0.17580964970257765, -0.17526455026455026, -0.17580964970257765, -0.17526455026455026]
drlTanhRewards21 = [-0.17544633017412387, -0.17526455026455026, -0.17526455026455026, -0.17526455026455026, -0.17526455026455026, -0.17580964970257765, -0.17562802996914942, -0.17508269018743108, -0.17562802996914942, -0.17526455026455026, -0.17544633017412387, -0.17508269018743108]
drlTanhRewards22 = [-0.1759911894273128, -0.17544633017412387, -0.17526455026455026, -0.17526455026455026, -0.17526455026455026, -0.17526455026455026, -0.17526455026455026, -0.17544633017412387, -0.17544633017412387, -0.17580964970257765, -0.17544633017412387, -0.17526455026455026]
drlTanhRewards23 = [-0.1759911894273128, -0.1759911894273128, -0.17544633017412387, -0.17544633017412387, -0.17526455026455026, -0.1759911894273128, -0.17544633017412387, -0.17562802996914942, -0.17508269018743108, -0.17508269018743108, -0.17580964970257765, -0.17544633017412387]
drlTanhRewards24 = [-0.17526455026455026, -0.17526455026455026, -0.17562802996914942, -0.17617264919621228, -0.17562802996914942, -0.17526455026455026, -0.17526455026455026, -0.17526455026455026, -0.17544633017412387, -0.17526455026455026, -0.17580964970257765, -0.1759911894273128]
drlTanhRewards25 = [-0.17526455026455026, -0.17526455026455026, -0.17562802996914942, -0.17526455026455026, -0.17526455026455026, -0.17526455026455026, -0.17617264919621228, -0.17544633017412387, -0.17562802996914942, -0.17526455026455026, -0.17526455026455026, -0.17526455026455026]
drlTanhRewards26 = [-0.17526455026455026, -0.17526455026455026, -0.1759911894273128, -0.17508269018743108, -0.17562802996914942, -0.17508269018743108, -0.17580964970257765, -0.17508269018743108, -0.17508269018743108, -0.17544633017412387, -0.17526455026455026, -0.17580964970257765]
drlTanhRewards27 = [-0.17544633017412387, -0.17544633017412387, -0.17526455026455026, -0.17544633017412387, -0.17580964970257765, -0.17562802996914942, -0.17508269018743108, -0.17526455026455026, -0.17508269018743108, -0.17508269018743108, -0.17580964970257765, -0.17508269018743108]
drlTanhRewards28 = [-0.17526455026455026, -0.17526455026455026, -0.17508269018743108, -0.17580964970257765, -0.17544633017412387, -0.17562802996914942, -0.17544633017412387, -0.17617264919621228, -0.17562802996914942, -0.17544633017412387, -0.17526455026455026, -0.17508269018743108]
drlTanhRewards29 = [-0.17526455026455026, -0.17526455026455026, -0.17580964970257765, -0.17617264919621228, -0.17562802996914942, -0.17508269018743108, -0.17544633017412387, -0.17508269018743108, -0.17471872931833224, -0.17508269018743108, -0.17508269018743108, -0.17526455026455026]
drlTanhRewards30 = [-0.17580964970257765, -0.17562802996914942, -0.17526455026455026, -0.1759911894273128, -0.17562802996914942, -0.17617264919621228, -0.17580964970257765, -0.1759911894273128, -0.17562802996914942, -0.17617264919621228, -0.17508269018743108, -0.17544633017412387]
drlTanhRewards31 = [-0.17508269018743108, -0.17526455026455026, -0.17580964970257765, -0.17617264919621228, -0.17617264919621228, -0.17508269018743108, -0.17617264919621228, -0.17580964970257765, -0.17580964970257765, -0.1759911894273128, -0.17580964970257765, -0.17580964970257765]
drlTanhRewards32 = [-0.17526455026455026, -0.17526455026455026, -0.17526455026455026, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.1749007498897221, -0.17526455026455026, -0.17508269018743108, -0.1749007498897221, -0.17508269018743108, -0.17526455026455026]
drlTanhRewards33 = [-0.17508269018743108, -0.17562802996914942, -0.17471872931833224, -0.17508269018743108, -0.17526455026455026, -0.17526455026455026, -0.17562802996914942, -0.17526455026455026, -0.17526455026455026, -0.17508269018743108, -0.17471872931833224, -0.17526455026455026]
drlTanhRewards34 = [-0.17526455026455026, -0.17508269018743108, -0.17526455026455026, -0.17580964970257765, -0.17526455026455026, -0.17508269018743108, -0.17544633017412387, -0.17526455026455026, -0.17508269018743108, -0.17508269018743108, -0.17526455026455026, -0.17508269018743108]
drlTanhRewards35 = [-0.17544633017412387, -0.17508269018743108, -0.17562802996914942, -0.1749007498897221, -0.17508269018743108, -0.17580964970257765, -0.17526455026455026, -0.17508269018743108, -0.1749007498897221, -0.1749007498897221, -0.17526455026455026, -0.17508269018743108]
drlTanhRewards36 = [-0.17526455026455026, -0.17526455026455026, -0.17508269018743108, -0.17526455026455026, -0.17526455026455026, -0.17526455026455026, -0.17508269018743108, -0.17526455026455026, -0.17526455026455026, -0.1749007498897221, -0.17526455026455026, -0.17508269018743108]
drlTanhRewards37 = [-0.17508269018743108, -0.17508269018743108, -0.17580964970257765, -0.17508269018743108, -0.17508269018743108, -0.17544633017412387, -0.1749007498897221, -0.17508269018743108, -0.17526455026455026, -0.17526455026455026, -0.17526455026455026, -0.17471872931833224]
drlTanhRewards38 = [-0.17508269018743108, -0.17544633017412387, -0.17526455026455026, -0.17508269018743108, -0.17580964970257765, -0.17580964970257765, -0.17526455026455026, -0.17526455026455026, -0.17508269018743108, -0.17508269018743108, -0.17526455026455026, -0.17508269018743108]
drlTanhRewards39 = [-0.17508269018743108, -0.17526455026455026, -0.17526455026455026, -0.17544633017412387, -0.17471872931833224, -0.17526455026455026, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17526455026455026, -0.17508269018743108, -0.17526455026455026]
drlTanhRewards40 = [-0.17580964970257765, -0.17526455026455026, -0.17508269018743108, -0.17526455026455026, -0.17508269018743108, -0.17508269018743108, -0.1749007498897221, -0.17526455026455026, -0.17508269018743108, -0.17508269018743108, -0.17562802996914942, -0.17508269018743108]
drlTanhRewards41 = [-0.17526455026455026, -0.17471872931833224, -0.17526455026455026, -0.17508269018743108, -0.17508269018743108, -0.17526455026455026, -0.17508269018743108, -0.17526455026455026, -0.17471872931833224, -0.17562802996914942, -0.17526455026455026, -0.17526455026455026]
drlTanhRewards42 = [-0.17471872931833224, -0.17508269018743108, -0.17508269018743108, -0.17526455026455026, -0.17508269018743108, -0.17508269018743108, -0.17526455026455026, -0.17508269018743108, -0.17471872931833224, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108]
drlTanhRewards43 = [-0.17508269018743108, -0.17544633017412387, -0.17508269018743108, -0.1749007498897221, -0.17526455026455026, -0.17526455026455026, -0.17580964970257765, -0.1749007498897221, -0.17508269018743108, -0.17508269018743108, -0.17580964970257765, -0.17508269018743108]
drlTanhRewards44 = [-0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17526455026455026, -0.17508269018743108, -0.17508269018743108, -0.1749007498897221, -0.17508269018743108, -0.17526455026455026, -0.17526455026455026, -0.17508269018743108]
drlTanhRewards45 = [-0.1749007498897221, -0.17435444714191128, -0.17508269018743108, -0.17508269018743108, -0.1749007498897221, -0.17508269018743108, -0.17526455026455026, -0.17508269018743108, -0.17453662842012357, -0.17526455026455026, -0.17526455026455026, -0.17508269018743108]
drlTanhRewards46 = [-0.17471872931833224, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.1749007498897221, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108]
drlTanhRewards47 = [-0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17471872931833224, -0.17526455026455026, -0.1749007498897221, -0.17526455026455026]
drlTanhRewards48 = [-0.17508269018743108, -0.17508269018743108, -0.17471872931833224, -0.17471872931833224, -0.17508269018743108, -0.17562802996914942, -0.17471872931833224, -0.17508269018743108, -0.17508269018743108, -0.17526455026455026, -0.17508269018743108, -0.17526455026455026]
drlTanhRewards49 = [-0.17526455026455026, -0.17508269018743108, -0.17508269018743108, -0.17544633017412387, -0.17508269018743108, -0.17562802996914942, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108]
drlTanhRewards50 = [-0.17508269018743108, -0.17471872931833224, -0.17526455026455026, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17526455026455026, -0.17526455026455026, -0.17508269018743108, -0.17508269018743108]
drlTanhRewards51 = [-0.17508269018743108, -0.17471872931833224, -0.17544633017412387, -0.17526455026455026, -0.17508269018743108, -0.17508269018743108, -0.17526455026455026, -0.17508269018743108, -0.17526455026455026, -0.17526455026455026, -0.17526455026455026, -0.17508269018743108]
drlTanhRewards52 = [-0.17508269018743108, -0.17508269018743108, -0.17526455026455026, -0.17471872931833224, -0.17526455026455026, -0.17526455026455026, -0.17526455026455026, -0.17471872931833224, -0.17508269018743108, -0.1749007498897221, -0.17526455026455026, -0.17508269018743108]
drlTanhRewards53 = [-0.17508269018743108, -0.17471872931833224, -0.17508269018743108, -0.17471872931833224, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17526455026455026, -0.17526455026455026, -0.17508269018743108, -0.17508269018743108, -0.17471872931833224]
drlTanhRewards54 = [-0.17526455026455026, -0.1749007498897221, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17471872931833224, -0.17526455026455026, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17471872931833224, -0.17544633017412387]
drlTanhRewards55 = [-0.17526455026455026, -0.17508269018743108, -0.17508269018743108, -0.17526455026455026, -0.17526455026455026, -0.1749007498897221, -0.17508269018743108, -0.17526455026455026, -0.17508269018743108, -0.17562802996914942, -0.17526455026455026, -0.17471872931833224]
drlTanhRewards56 = [-0.17526455026455026, -0.17526455026455026, -0.17471872931833224, -0.17508269018743108, -0.17526455026455026, -0.17526455026455026, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17526455026455026, -0.17526455026455026, -0.17526455026455026]
drlTanhRewards57 = [-0.17526455026455026, -0.17471872931833224, -0.17526455026455026, -0.17508269018743108, -0.17526455026455026, -0.17526455026455026, -0.17471872931833224, -0.17526455026455026, -0.17508269018743108, -0.17562802996914942, -0.17471872931833224, -0.17471872931833224]
drlTanhRewards58 = [-0.17526455026455026, -0.17526455026455026, -0.17508269018743108, -0.17526455026455026, -0.17471872931833224, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17471872931833224, -0.17471872931833224, -0.17471872931833224, -0.1749007498897221]
drlTanhRewards59 = [-0.17526455026455026, -0.17508269018743108, -0.17471872931833224, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17471872931833224, -0.17508269018743108, -0.17508269018743108, -0.1749007498897221, -0.17526455026455026]
drlTanhRewards60 = [-0.17508269018743108, -0.17526455026455026, -0.17526455026455026, -0.17580964970257765, -0.17526455026455026, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17471872931833224]
drlTanhRewards61 = [-0.17471872931833224, -0.17508269018743108, -0.17526455026455026, -0.17508269018743108, -0.17508269018743108, -0.17526455026455026, -0.17526455026455026, -0.17471872931833224, -0.17526455026455026, -0.17508269018743108, -0.17508269018743108, -0.17526455026455026]
drlTanhRewards62 = [-0.17526455026455026, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.1759911894273128, -0.17471872931833224, -0.17508269018743108, -0.17508269018743108, -0.17526455026455026, -0.17526455026455026, -0.17508269018743108, -0.1749007498897221]
drlTanhRewards63 = [-0.17453662842012357, -0.17471872931833224, -0.17526455026455026, -0.17544633017412387, -0.17508269018743108, -0.17508269018743108, -0.17471872931833224, -0.17526455026455026, -0.1749007498897221, -0.17508269018743108, -0.17471872931833224, -0.17508269018743108]
# Deep Reinforcement Learning: 5 capas Dense, Funcion de activacion relu, 12 episodios, 50 iteraciones
drlReluMakespan0 = [796, 798, 809, 798, 796, 800, 798, 799, 800, 794, 800, 798]
drlReluMakespan1 = [800, 800, 801, 806, 804, 806, 808, 798, 796, 796, 798, 800]
drlReluMakespan2 = [805, 805, 798, 800, 800, 798, 801, 799, 800, 806, 800, 800]
drlReluMakespan3 = [798, 799, 798, 795, 798, 808, 803, 800, 798, 795, 799, 800]
drlReluMakespan4 = [805, 805, 799, 796, 798, 803, 799, 800, 800, 800, 795, 794]
drlReluMakespan5 = [799, 796, 795, 800, 801, 796, 800, 795, 803, 800, 800, 805]
drlReluMakespan6 = [799, 795, 798, 794, 805, 796, 795, 799, 798, 795, 804, 796]
drlReluMakespan7 = [795, 798, 799, 798, 798, 799, 795, 794, 796, 794, 795, 805]
drlReluMakespan8 = [805, 794, 794, 795, 798, 795, 798, 795, 799, 800, 796, 798]
drlReluMakespan9 = [797, 797, 797, 794, 795, 794, 794, 797, 796, 795, 801, 799]
drlReluMakespan10 = [799, 794, 797, 795, 794, 794, 795, 795, 795, 796, 797, 799]
drlReluMakespan11 = [796, 798, 800, 795, 805, 794, 798, 796, 795, 794, 798, 795]
drlReluMakespan12 = [800, 795, 794, 798, 800, 805, 800, 798, 804, 799, 794, 803]
drlReluMakespan13 = [796, 799, 798, 794, 800, 794, 795, 796, 798, 795, 794, 799]
drlReluMakespan14 = [795, 798, 798, 798, 805, 798, 798, 798, 795, 794, 800, 796]
drlReluMakespan15 = [795, 798, 795, 805, 798, 794, 795, 798, 796, 794, 795, 796]
drlReluMakespan16 = [798, 795, 796, 799, 796, 798, 798, 795, 795, 795, 795, 799]
drlReluMakespan17 = [794, 798, 796, 798, 795, 801, 794, 798, 797, 795, 796, 801]
drlReluMakespan18 = [798, 795, 798, 798, 801, 798, 795, 795, 797, 800, 794, 800]
drlReluMakespan19 = [795, 798, 794, 800, 796, 795, 798, 797, 795, 794, 796, 796]
drlReluMakespan20 = [794, 794, 795, 795, 795, 795, 796, 798, 799, 799, 799, 795]
drlReluMakespan21 = [802, 796, 794, 797, 797, 800, 794, 794, 804, 803, 798, 797]
drlReluMakespan22 = [794, 795, 795, 795, 798, 795, 794, 799, 794, 803, 795, 794]
drlReluMakespan23 = [794, 798, 799, 794, 795, 795, 799, 795, 796, 795, 797, 799]
drlReluMakespan24 = [795, 794, 797, 800, 794, 795, 795, 795, 795, 800, 800, 798]
drlReluMakespan25 = [795, 794, 797, 796, 798, 795, 795, 794, 799, 795, 794, 798]
drlReluMakespan26 = [801, 795, 800, 794, 794, 796, 800, 798, 798, 799, 794, 796]
drlReluMakespan27 = [796, 795, 796, 795, 796, 795, 795, 800, 794, 794, 794, 796]
drlReluMakespan28 = [794, 794, 795, 796, 794, 795, 795, 797, 794, 794, 796, 795]
drlReluMakespan29 = [793, 794, 795, 800, 795, 795, 794, 798, 798, 796, 795, 794]
drlReluMakespan30 = [802, 794, 794, 798, 794, 796, 805, 794, 800, 794, 796, 794]
drlReluMakespan31 = [797, 794, 794, 794, 800, 800, 794, 794, 798, 795, 794, 798]
drlReluMakespan32 = [794, 798, 794, 795, 794, 795, 798, 794, 794, 795, 794, 798]
drlReluMakespan33 = [798, 794, 798, 795, 794, 793, 797, 798, 794, 794, 801, 793]
drlReluMakespan34 = [794, 798, 794, 795, 794, 793, 798, 795, 794, 800, 794, 795]
drlReluMakespan35 = [794, 796, 794, 796, 806, 795, 795, 795, 796, 795, 795, 799]
drlReluMakespan36 = [795, 794, 794, 796, 796, 798, 794, 796, 794, 795, 794, 795]
drlReluMakespan37 = [795, 794, 795, 798, 794, 794, 794, 794, 794, 794, 795, 797]
drlReluMakespan38 = [794, 798, 794, 798, 797, 794, 794, 795, 795, 794, 795, 795]
drlReluMakespan39 = [797, 794, 795, 796, 796, 796, 798, 794, 794, 795, 794, 798]
drlReluMakespan40 = [798, 795, 795, 798, 792, 795, 795, 794, 795, 794, 798, 794]
drlReluMakespan41 = [795, 794, 794, 794, 794, 794, 798, 793, 794, 794, 794, 793]
drlReluMakespan42 = [794, 794, 794, 794, 799, 794, 795, 794, 796, 794, 794, 794]
drlReluMakespan43 = [794, 797, 795, 794, 795, 794, 794, 795, 794, 794, 793, 794]
drlReluMakespan44 = [794, 792, 793, 794, 794, 796, 794, 798, 795, 794, 794, 796]
drlReluMakespan45 = [795, 794, 799, 794, 794, 793, 794, 795, 795, 793, 796, 794]
drlReluMakespan46 = [794, 796, 794, 794, 794, 794, 794, 793, 799, 792, 794, 794]
drlReluMakespan47 = [795, 794, 793, 794, 796, 797, 794, 794, 795, 794, 794, 794]
drlReluMakespan48 = [794, 794, 794, 792, 794, 794, 795, 794, 794, 794, 794, 794]
drlReluMakespan49 = [794, 794, 795, 792, 797, 797, 794, 794, 792, 800, 795, 795]
drlReluRewards0 = [-0.17544633017412387, -0.17580964970257765, -0.1778021978021978, -0.17544633017412387, -0.17580964970257765, -0.17617264919621228, -0.17580964970257765, -0.1759911894273128, -0.17508269018743108, -0.17617264919621228, -0.17617264919621228, -0.17580964970257765]
drlReluRewards1 = [-0.17617264919621228, -0.17617264919621228, -0.1763540290620872, -0.17725973169122497, -0.1768976897689769, -0.17725973169122497, -0.1776214552648934, -0.17580964970257765, -0.17544633017412387, -0.17544633017412387, -0.17580964970257765, -0.17617264919621228]
drlReluRewards2 = [-0.177078750549934, -0.177078750549934, -0.17580964970257765, -0.17617264919621228, -0.17617264919621228, -0.17580964970257765, -0.1763540290620872, -0.1759911894273128, -0.17617264919621228, -0.17725973169122497, -0.17617264919621228, -0.17617264919621228]
drlReluRewards3 = [-0.17580964970257765, -0.1759911894273128, -0.17580964970257765, -0.17526455026455026, -0.17580964970257765, -0.1776214552648934, -0.17671654929577466, -0.17617264919621228, -0.17580964970257765, -0.17526455026455026, -0.1759911894273128, -0.17617264919621228]
drlReluRewards4 = [-0.177078750549934, -0.177078750549934, -0.1759911894273128, -0.17544633017412387, -0.17580964970257765, -0.17671654929577466, -0.1759911894273128, -0.17617264919621228, -0.17617264919621228, -0.17617264919621228, -0.17526455026455026, -0.17508269018743108]
drlReluRewards5 = [-0.1759911894273128, -0.17544633017412387, -0.17526455026455026, -0.17617264919621228, -0.1763540290620872, -0.17544633017412387, -0.17526455026455026, -0.17617264919621228, -0.17671654929577466, -0.17617264919621228, -0.17617264919621228, -0.177078750549934]
drlReluRewards6 = [-0.1759911894273128, -0.17526455026455026, -0.17580964970257765, -0.17508269018743108, -0.177078750549934, -0.17544633017412387, -0.17526455026455026, -0.1759911894273128, -0.17580964970257765, -0.17526455026455026, -0.1768976897689769, -0.17544633017412387]
drlReluRewards7 = [-0.17526455026455026, -0.1759911894273128, -0.17580964970257765, -0.17580964970257765, -0.17580964970257765, -0.1759911894273128, -0.17526455026455026, -0.17544633017412387, -0.17508269018743108, -0.17508269018743108, -0.17526455026455026, -0.177078750549934]
drlReluRewards8 = [-0.177078750549934, -0.17508269018743108, -0.17508269018743108, -0.17526455026455026, -0.17580964970257765, -0.17526455026455026, -0.17580964970257765, -0.17526455026455026, -0.1759911894273128, -0.17617264919621228, -0.17544633017412387, -0.17580964970257765]
drlReluRewards9 = [-0.17562802996914942, -0.17562802996914942, -0.17562802996914942, -0.17526455026455026, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17562802996914942, -0.17544633017412387, -0.17526455026455026, -0.1763540290620872, -0.1759911894273128]
drlReluRewards10 = [-0.1759911894273128, -0.17508269018743108, -0.17562802996914942, -0.17526455026455026, -0.17508269018743108, -0.17508269018743108, -0.17526455026455026, -0.17526455026455026, -0.17526455026455026, -0.17544633017412387, -0.17562802996914942, -0.1759911894273128]
drlReluRewards11 = [-0.17544633017412387, -0.17580964970257765, -0.17617264919621228, -0.17526455026455026, -0.177078750549934, -0.17508269018743108, -0.17580964970257765, -0.17544633017412387, -0.17526455026455026, -0.17508269018743108, -0.17580964970257765, -0.17526455026455026]
drlReluRewards12 = [-0.17617264919621228, -0.17526455026455026, -0.17508269018743108, -0.17580964970257765, -0.17617264919621228, -0.177078750549934, -0.17617264919621228, -0.17580964970257765, -0.1768976897689769, -0.1759911894273128, -0.17508269018743108, -0.17671654929577466]
drlReluRewards13 = [-0.17544633017412387, -0.1759911894273128, -0.17580964970257765, -0.17508269018743108, -0.17617264919621228, -0.17508269018743108, -0.17526455026455026, -0.17544633017412387, -0.17580964970257765, -0.17526455026455026, -0.17508269018743108, -0.1759911894273128]
drlReluRewards14 = [-0.17526455026455026, -0.17580964970257765, -0.17580964970257765, -0.17580964970257765, -0.177078750549934, -0.17580964970257765, -0.17580964970257765, -0.17580964970257765, -0.17526455026455026, -0.17508269018743108, -0.17617264919621228, -0.17544633017412387]
drlReluRewards15 = [-0.17526455026455026, -0.17580964970257765, -0.17526455026455026, -0.177078750549934, -0.17580964970257765, -0.17508269018743108, -0.17526455026455026, -0.17580964970257765, -0.17544633017412387, -0.17508269018743108, -0.17526455026455026, -0.17544633017412387]
drlReluRewards16 = [-0.17580964970257765, -0.17526455026455026, -0.17544633017412387, -0.1759911894273128, -0.17580964970257765, -0.17544633017412387, -0.17580964970257765, -0.17526455026455026, -0.17526455026455026, -0.17526455026455026, -0.17526455026455026, -0.1759911894273128]
drlReluRewards17 = [-0.17508269018743108, -0.17580964970257765, -0.17544633017412387, -0.17580964970257765, -0.17526455026455026, -0.1763540290620872, -0.17508269018743108, -0.17580964970257765, -0.17562802996914942, -0.17526455026455026, -0.17544633017412387, -0.1763540290620872]
drlReluRewards18 = [-0.17580964970257765, -0.17526455026455026, -0.17580964970257765, -0.17580964970257765, -0.1763540290620872, -0.17580964970257765, -0.17526455026455026, -0.17526455026455026, -0.17562802996914942, -0.17617264919621228, -0.17508269018743108, -0.17617264919621228]
drlReluRewards19 = [-0.17526455026455026, -0.17580964970257765, -0.17508269018743108, -0.17617264919621228, -0.17544633017412387, -0.17526455026455026, -0.17580964970257765, -0.17562802996914942, -0.17526455026455026, -0.17508269018743108, -0.17544633017412387, -0.17544633017412387]
drlReluRewards20 = [-0.17508269018743108, -0.17508269018743108, -0.17526455026455026, -0.17526455026455026, -0.17526455026455026, -0.17526455026455026, -0.17544633017412387, -0.17580964970257765, -0.1759911894273128, -0.1759911894273128, -0.1759911894273128, -0.17526455026455026]
drlReluRewards21 = [-0.17653532907770195, -0.17544633017412387, -0.17562802996914942, -0.17508269018743108, -0.17562802996914942, -0.17617264919621228, -0.17508269018743108, -0.17508269018743108, -0.17580964970257765, -0.1768976897689769, -0.17671654929577466, -0.17562802996914942]
drlReluRewards22 = [-0.17508269018743108, -0.17526455026455026, -0.17526455026455026, -0.17526455026455026, -0.17580964970257765, -0.17526455026455026, -0.17508269018743108, -0.1759911894273128, -0.17508269018743108, -0.17671654929577466, -0.17526455026455026, -0.17508269018743108]
drlReluRewards23 = [-0.17508269018743108, -0.17580964970257765, -0.17526455026455026, -0.1759911894273128, -0.17508269018743108, -0.17526455026455026, -0.1759911894273128, -0.17526455026455026, -0.17544633017412387, -0.17526455026455026, -0.17562802996914942, -0.1759911894273128]
drlReluRewards24 = [-0.17526455026455026, -0.17508269018743108, -0.17562802996914942, -0.17617264919621228, -0.17508269018743108, -0.17526455026455026, -0.17526455026455026, -0.17526455026455026, -0.17526455026455026, -0.17617264919621228, -0.17617264919621228, -0.17580964970257765]
drlReluRewards25 = [-0.17526455026455026, -0.17508269018743108, -0.17562802996914942, -0.17544633017412387, -0.17580964970257765, -0.17526455026455026, -0.17526455026455026, -0.17508269018743108, -0.1759911894273128, -0.17526455026455026, -0.17508269018743108, -0.17580964970257765]
drlReluRewards26 = [-0.1763540290620872, -0.17526455026455026, -0.17617264919621228, -0.17508269018743108, -0.17544633017412387, -0.17508269018743108, -0.17617264919621228, -0.17580964970257765, -0.17580964970257765, -0.1759911894273128, -0.17508269018743108, -0.17544633017412387]
drlReluRewards27 = [-0.17544633017412387, -0.17526455026455026, -0.17544633017412387, -0.17526455026455026, -0.17544633017412387, -0.17526455026455026, -0.17526455026455026, -0.17617264919621228, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17544633017412387]
drlReluRewards28 = [-0.17508269018743108, -0.17508269018743108, -0.17526455026455026, -0.17544633017412387, -0.17508269018743108, -0.17526455026455026, -0.17526455026455026, -0.17562802996914942, -0.17508269018743108, -0.17508269018743108, -0.17544633017412387, -0.17526455026455026]
drlReluRewards29 = [-0.1749007498897221, -0.17526455026455026, -0.17508269018743108, -0.17617264919621228, -0.17526455026455026, -0.17526455026455026, -0.17508269018743108, -0.17580964970257765, -0.17580964970257765, -0.17544633017412387, -0.17526455026455026, -0.17508269018743108]
drlReluRewards30 = [-0.17653532907770195, -0.17508269018743108, -0.17508269018743108, -0.17580964970257765, -0.17508269018743108, -0.17544633017412387, -0.177078750549934, -0.17508269018743108, -0.17617264919621228, -0.17508269018743108, -0.17544633017412387, -0.17508269018743108]
drlReluRewards31 = [-0.17562802996914942, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17617264919621228, -0.17617264919621228, -0.17508269018743108, -0.17508269018743108, -0.17526455026455026, -0.17580964970257765, -0.17508269018743108, -0.17580964970257765]
drlReluRewards32 = [-0.17508269018743108, -0.17580964970257765, -0.17508269018743108, -0.17526455026455026, -0.17508269018743108, -0.17526455026455026, -0.17580964970257765, -0.17508269018743108, -0.17508269018743108, -0.17526455026455026, -0.17508269018743108, -0.17580964970257765]
drlReluRewards33 = [-0.17580964970257765, -0.17508269018743108, -0.17580964970257765, -0.17526455026455026, -0.17508269018743108, -0.1749007498897221, -0.17562802996914942, -0.17580964970257765, -0.17508269018743108, -0.17508269018743108, -0.1763540290620872, -0.1749007498897221]
drlReluRewards34 = [-0.17508269018743108, -0.17580964970257765, -0.17508269018743108, -0.17526455026455026, -0.17508269018743108, -0.1749007498897221, -0.17580964970257765, -0.17526455026455026, -0.17508269018743108, -0.17617264919621228, -0.17508269018743108, -0.17526455026455026]
drlReluRewards35 = [-0.17508269018743108, -0.17544633017412387, -0.17725973169122497, -0.17508269018743108, -0.17544633017412387, -0.17526455026455026, -0.17526455026455026, -0.17526455026455026, -0.17544633017412387, -0.17526455026455026, -0.17526455026455026, -0.1759911894273128]
drlReluRewards36 = [-0.17526455026455026, -0.17508269018743108, -0.17508269018743108, -0.17544633017412387, -0.17544633017412387, -0.17580964970257765, -0.17508269018743108, -0.17544633017412387, -0.17508269018743108, -0.17526455026455026, -0.17508269018743108, -0.17526455026455026]
drlReluRewards37 = [-0.17526455026455026, -0.17508269018743108, -0.17526455026455026, -0.17580964970257765, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17526455026455026, -0.17562802996914942]
drlReluRewards38 = [-0.17508269018743108, -0.17580964970257765, -0.17508269018743108, -0.17580964970257765, -0.17562802996914942, -0.17508269018743108, -0.17508269018743108, -0.17526455026455026, -0.17526455026455026, -0.17508269018743108, -0.17526455026455026, -0.17526455026455026]
drlReluRewards39 = [-0.17562802996914942, -0.17508269018743108, -0.17526455026455026, -0.17544633017412387, -0.17544633017412387, -0.17544633017412387, -0.17580964970257765, -0.17508269018743108, -0.17508269018743108, -0.17526455026455026, -0.17508269018743108, -0.17580964970257765]
drlReluRewards40 = [-0.17580964970257765, -0.17526455026455026, -0.17526455026455026, -0.17580964970257765, -0.17471872931833224, -0.17526455026455026, -0.17526455026455026, -0.17508269018743108, -0.17526455026455026, -0.17508269018743108, -0.17580964970257765, -0.17508269018743108]
drlReluRewards41 = [-0.17526455026455026, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17580964970257765, -0.1749007498897221, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.1749007498897221]
drlReluRewards42 = [-0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.1759911894273128, -0.17508269018743108, -0.17526455026455026, -0.17508269018743108, -0.17544633017412387, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108]
drlReluRewards43 = [-0.17508269018743108, -0.17562802996914942, -0.17526455026455026, -0.17508269018743108, -0.17526455026455026, -0.17508269018743108, -0.17508269018743108, -0.17526455026455026, -0.17508269018743108, -0.17508269018743108, -0.1749007498897221, -0.17508269018743108]
drlReluRewards44 = [-0.17508269018743108, -0.17471872931833224, -0.1749007498897221, -0.17508269018743108, -0.17508269018743108, -0.17544633017412387, -0.17508269018743108, -0.17580964970257765, -0.17526455026455026, -0.17508269018743108, -0.17508269018743108, -0.17544633017412387]
drlReluRewards45 = [-0.17526455026455026, -0.17508269018743108, -0.1759911894273128, -0.17508269018743108, -0.17508269018743108, -0.1749007498897221, -0.17508269018743108, -0.17526455026455026, -0.17526455026455026, -0.1749007498897221, -0.17544633017412387, -0.17508269018743108]
drlReluRewards46 = [-0.17508269018743108, -0.17544633017412387, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.1749007498897221, -0.1759911894273128, -0.17471872931833224, -0.17508269018743108, -0.17508269018743108]
drlReluRewards47 = [-0.17526455026455026, -0.17508269018743108, -0.1749007498897221, -0.17508269018743108, -0.17544633017412387, -0.17562802996914942, -0.17508269018743108, -0.17508269018743108, -0.17526455026455026, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108]
drlReluRewards48 = [-0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17471872931833224, -0.17508269018743108, -0.17508269018743108, -0.17526455026455026, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108, -0.17508269018743108]
drlReluRewards49 = [-0.17508269018743108, -0.17508269018743108, -0.17526455026455026, -0.17471872931833224, -0.17562802996914942, -0.17562802996914942, -0.17508269018743108, -0.17508269018743108, -0.17471872931833224, -0.17617264919621228, -0.17526455026455026, -0.17526455026455026]
if __name__ == "__main__":
##############################################
##############################################
##############################################
# Deep Recurrent Reinforcement Learning with 1 GRU layer and 4 Dense layers
drnnGRUtanhMakespan = []
drnnGRUtanhRewards = []
drnnGRUtanhMakespanList = []
drnnGRUtanhRewardsList = []
drnnGRUtanhMakespanValues = []
drnnGRUtanhRewardsValues = []
drnnGRUtanhMakespan.append(np.mean(drnnGRUtanhMakespan0))
drnnGRUtanhMakespan.append(np.mean(drnnGRUtanhMakespan1))
drnnGRUtanhMakespan.append(np.mean(drnnGRUtanhMakespan2))
drnnGRUtanhMakespan.append(np.mean(drnnGRUtanhMakespan3))
drnnGRUtanhMakespan.append(np.mean(drnnGRUtanhMakespan4))
drnnGRUtanhMakespan.append(np.mean(drnnGRUtanhMakespan5))
drnnGRUtanhMakespan.append(np.mean(drnnGRUtanhMakespan6))
drnnGRUtanhMakespan.append(np.mean(drnnGRUtanhMakespan7))
drnnGRUtanhMakespan.append(np.mean(drnnGRUtanhMakespan8))
drnnGRUtanhMakespan.append(np.mean(drnnGRUtanhMakespan9))
drnnGRUtanhMakespan.append(np.mean(drnnGRUtanhMakespan10))
drnnGRUtanhMakespan.append(np.mean(drnnGRUtanhMakespan11))
drnnGRUtanhMakespan.append(np.mean(drnnGRUtanhMakespan12))
drnnGRUtanhMakespan.append(np.mean(drnnGRUtanhMakespan13))
drnnGRUtanhMakespan.append(np.mean(drnnGRUtanhMakespan14))
drnnGRUtanhMakespan.append(np.mean(drnnGRUtanhMakespan15))
drnnGRUtanhMakespan.append(np.mean(drnnGRUtanhMakespan16))
drnnGRUtanhMakespan.append(np.mean(drnnGRUtanhMakespan17))
drnnGRUtanhMakespan.append(np.mean(drnnGRUtanhMakespan18))
drnnGRUtanhMakespan.append(np.mean(drnnGRUtanhMakespan19))
drnnGRUtanhMakespan.append(np.mean(drnnGRUtanhMakespan20))
drnnGRUtanhMakespan.append(np.mean(drnnGRUtanhMakespan21))
drnnGRUtanhMakespan.append(np.mean(drnnGRUtanhMakespan22))
drnnGRUtanhMakespan.append(np.mean(drnnGRUtanhMakespan23))
drnnGRUtanhMakespan.append(np.mean(drnnGRUtanhMakespan24))
drnnGRUtanhMakespan.append(np.mean(drnnGRUtanhMakespan25))
drnnGRUtanhMakespan.append(np.mean(drnnGRUtanhMakespan26))
drnnGRUtanhMakespan.append(np.mean(drnnGRUtanhMakespan27))
drnnGRUtanhMakespan.append(np.mean(drnnGRUtanhMakespan28))
drnnGRUtanhMakespan.append(np.mean(drnnGRUtanhMakespan29))
drnnGRUtanhMakespan.append(np.mean(drnnGRUtanhMakespan30))
drnnGRUtanhMakespan.append(np.mean(drnnGRUtanhMakespan31))
drnnGRUtanhMakespan.append(np.mean(drnnGRUtanhMakespan32))
drnnGRUtanhMakespan.append(np.mean(drnnGRUtanhMakespan33))
drnnGRUtanhMakespan.append(np.mean(drnnGRUtanhMakespan34))
drnnGRUtanhMakespan.append(np.mean(drnnGRUtanhMakespan35))
drnnGRUtanhMakespan.append(np.mean(drnnGRUtanhMakespan36))
drnnGRUtanhMakespan.append(np.mean(drnnGRUtanhMakespan37))
drnnGRUtanhMakespan.append(np.mean(drnnGRUtanhMakespan38))
drnnGRUtanhMakespan.append(np.mean(drnnGRUtanhMakespan39))
drnnGRUtanhMakespan.append(np.mean(drnnGRUtanhMakespan40))
drnnGRUtanhMakespan.append(np.mean(drnnGRUtanhMakespan41))
drnnGRUtanhMakespan.append(np.mean(drnnGRUtanhMakespan42))
drnnGRUtanhMakespan.append(np.mean(drnnGRUtanhMakespan43))
drnnGRUtanhMakespan.append(np.mean(drnnGRUtanhMakespan44))
drnnGRUtanhMakespan.append(np.mean(drnnGRUtanhMakespan45))
drnnGRUtanhMakespan.append(np.mean(drnnGRUtanhMakespan46))
drnnGRUtanhMakespan.append(np.mean(drnnGRUtanhMakespan47))
drnnGRUtanhMakespan.append(np.mean(drnnGRUtanhMakespan48))
drnnGRUtanhMakespan.append(np.mean(drnnGRUtanhMakespan49))
drnnGRUtanhRewards.append(np.mean(drnnGRUtanhRewards0))
drnnGRUtanhRewards.append(np.mean(drnnGRUtanhRewards1))
drnnGRUtanhRewards.append(np.mean(drnnGRUtanhRewards2))
drnnGRUtanhRewards.append(np.mean(drnnGRUtanhRewards3))
drnnGRUtanhRewards.append(np.mean(drnnGRUtanhRewards4))
drnnGRUtanhRewards.append(np.mean(drnnGRUtanhRewards5))
drnnGRUtanhRewards.append(np.mean(drnnGRUtanhRewards6))
drnnGRUtanhRewards.append(np.mean(drnnGRUtanhRewards7))
drnnGRUtanhRewards.append(np.mean(drnnGRUtanhRewards8))
drnnGRUtanhRewards.append(np.mean(drnnGRUtanhRewards9))
drnnGRUtanhRewards.append(np.mean(drnnGRUtanhRewards10))
drnnGRUtanhRewards.append(np.mean(drnnGRUtanhRewards11))
drnnGRUtanhRewards.append(np.mean(drnnGRUtanhRewards12))
drnnGRUtanhRewards.append(np.mean(drnnGRUtanhRewards13))
drnnGRUtanhRewards.append(np.mean(drnnGRUtanhRewards14))
drnnGRUtanhRewards.append(np.mean(drnnGRUtanhRewards15))
drnnGRUtanhRewards.append(np.mean(drnnGRUtanhRewards16))
drnnGRUtanhRewards.append(np.mean(drnnGRUtanhRewards17))
drnnGRUtanhRewards.append(np.mean(drnnGRUtanhRewards18))
drnnGRUtanhRewards.append(np.mean(drnnGRUtanhRewards19))
drnnGRUtanhRewards.append(np.mean(drnnGRUtanhRewards20))
drnnGRUtanhRewards.append(np.mean(drnnGRUtanhRewards21))
drnnGRUtanhRewards.append(np.mean(drnnGRUtanhRewards22))
drnnGRUtanhRewards.append(np.mean(drnnGRUtanhRewards23))
drnnGRUtanhRewards.append(np.mean(drnnGRUtanhRewards24))
drnnGRUtanhRewards.append(np.mean(drnnGRUtanhRewards25))
drnnGRUtanhRewards.append(np.mean(drnnGRUtanhRewards26))
drnnGRUtanhRewards.append(np.mean(drnnGRUtanhRewards27))
drnnGRUtanhRewards.append(np.mean(drnnGRUtanhRewards28))
drnnGRUtanhRewards.append(np.mean(drnnGRUtanhRewards29))
drnnGRUtanhRewards.append(np.mean(drnnGRUtanhRewards30))
drnnGRUtanhRewards.append(np.mean(drnnGRUtanhRewards31))
drnnGRUtanhRewards.append(np.mean(drnnGRUtanhRewards32))
drnnGRUtanhRewards.append(np.mean(drnnGRUtanhRewards33))
drnnGRUtanhRewards.append(np.mean(drnnGRUtanhRewards34))
drnnGRUtanhRewards.append(np.mean(drnnGRUtanhRewards35))
drnnGRUtanhRewards.append(np.mean(drnnGRUtanhRewards36))
drnnGRUtanhRewards.append(np.mean(drnnGRUtanhRewards37))
drnnGRUtanhRewards.append(np.mean(drnnGRUtanhRewards38))
drnnGRUtanhRewards.append(np.mean(drnnGRUtanhRewards39))
drnnGRUtanhRewards.append(np.mean(drnnGRUtanhRewards40))
drnnGRUtanhRewards.append(np.mean(drnnGRUtanhRewards41))
drnnGRUtanhRewards.append(np.mean(drnnGRUtanhRewards42))
drnnGRUtanhRewards.append(np.mean(drnnGRUtanhRewards43))
drnnGRUtanhRewards.append(np.mean(drnnGRUtanhRewards44))
drnnGRUtanhRewards.append(np.mean(drnnGRUtanhRewards45))
drnnGRUtanhRewards.append(np.mean(drnnGRUtanhRewards46))
drnnGRUtanhRewards.append(np.mean(drnnGRUtanhRewards47))
drnnGRUtanhRewards.append(np.mean(drnnGRUtanhRewards48))
drnnGRUtanhRewards.append(np.mean(drnnGRUtanhRewards49))
drnnGRUtanhMakespanList.append(drnnGRUtanhMakespan0)
drnnGRUtanhMakespanList.append(drnnGRUtanhMakespan1)
drnnGRUtanhMakespanList.append(drnnGRUtanhMakespan2)
drnnGRUtanhMakespanList.append(drnnGRUtanhMakespan3)
drnnGRUtanhMakespanList.append(drnnGRUtanhMakespan4)
drnnGRUtanhMakespanList.append(drnnGRUtanhMakespan5)
drnnGRUtanhMakespanList.append(drnnGRUtanhMakespan6)
drnnGRUtanhMakespanList.append(drnnGRUtanhMakespan7)
drnnGRUtanhMakespanList.append(drnnGRUtanhMakespan8)
drnnGRUtanhMakespanList.append(drnnGRUtanhMakespan9)
drnnGRUtanhMakespanList.append(drnnGRUtanhMakespan10)
drnnGRUtanhMakespanList.append(drnnGRUtanhMakespan11)
drnnGRUtanhMakespanList.append(drnnGRUtanhMakespan12)
drnnGRUtanhMakespanList.append(drnnGRUtanhMakespan13)
drnnGRUtanhMakespanList.append(drnnGRUtanhMakespan14)
drnnGRUtanhMakespanList.append(drnnGRUtanhMakespan15)
drnnGRUtanhMakespanList.append(drnnGRUtanhMakespan16)
drnnGRUtanhMakespanList.append(drnnGRUtanhMakespan17)
drnnGRUtanhMakespanList.append(drnnGRUtanhMakespan18)
drnnGRUtanhMakespanList.append(drnnGRUtanhMakespan19)
drnnGRUtanhMakespanList.append(drnnGRUtanhMakespan20)
drnnGRUtanhMakespanList.append(drnnGRUtanhMakespan21)
drnnGRUtanhMakespanList.append(drnnGRUtanhMakespan22)
drnnGRUtanhMakespanList.append(drnnGRUtanhMakespan23)
drnnGRUtanhMakespanList.append(drnnGRUtanhMakespan24)
drnnGRUtanhMakespanList.append(drnnGRUtanhMakespan25)
drnnGRUtanhMakespanList.append(drnnGRUtanhMakespan26)
drnnGRUtanhMakespanList.append(drnnGRUtanhMakespan27)
drnnGRUtanhMakespanList.append(drnnGRUtanhMakespan28)
drnnGRUtanhMakespanList.append(drnnGRUtanhMakespan29)
drnnGRUtanhMakespanList.append(drnnGRUtanhMakespan30)
drnnGRUtanhMakespanList.append(drnnGRUtanhMakespan31)
drnnGRUtanhMakespanList.append(drnnGRUtanhMakespan32)
drnnGRUtanhMakespanList.append(drnnGRUtanhMakespan33)
drnnGRUtanhMakespanList.append(drnnGRUtanhMakespan34)
drnnGRUtanhMakespanList.append(drnnGRUtanhMakespan35)
drnnGRUtanhMakespanList.append(drnnGRUtanhMakespan36)
drnnGRUtanhMakespanList.append(drnnGRUtanhMakespan37)
drnnGRUtanhMakespanList.append(drnnGRUtanhMakespan38)
drnnGRUtanhMakespanList.append(drnnGRUtanhMakespan39)
drnnGRUtanhMakespanList.append(drnnGRUtanhMakespan40)
drnnGRUtanhMakespanList.append(drnnGRUtanhMakespan41)
drnnGRUtanhMakespanList.append(drnnGRUtanhMakespan42)
drnnGRUtanhMakespanList.append(drnnGRUtanhMakespan43)
drnnGRUtanhMakespanList.append(drnnGRUtanhMakespan44)
drnnGRUtanhMakespanList.append(drnnGRUtanhMakespan45)
drnnGRUtanhMakespanList.append(drnnGRUtanhMakespan46)
drnnGRUtanhMakespanList.append(drnnGRUtanhMakespan47)
drnnGRUtanhMakespanList.append(drnnGRUtanhMakespan48)
drnnGRUtanhMakespanList.append(drnnGRUtanhMakespan49)
drnnGRUtanhRewardsList.append(drnnGRUtanhRewards0)
drnnGRUtanhRewardsList.append(drnnGRUtanhRewards1)
drnnGRUtanhRewardsList.append(drnnGRUtanhRewards2)
drnnGRUtanhRewardsList.append(drnnGRUtanhRewards3)
drnnGRUtanhRewardsList.append(drnnGRUtanhRewards4)
drnnGRUtanhRewardsList.append(drnnGRUtanhRewards5)
drnnGRUtanhRewardsList.append(drnnGRUtanhRewards6)
drnnGRUtanhRewardsList.append(drnnGRUtanhRewards7)
drnnGRUtanhRewardsList.append(drnnGRUtanhRewards8)
drnnGRUtanhRewardsList.append(drnnGRUtanhRewards9)
drnnGRUtanhRewardsList.append(drnnGRUtanhRewards10)
drnnGRUtanhRewardsList.append(drnnGRUtanhRewards11)
drnnGRUtanhRewardsList.append(drnnGRUtanhRewards12)
drnnGRUtanhRewardsList.append(drnnGRUtanhRewards13)
drnnGRUtanhRewardsList.append(drnnGRUtanhRewards14)
drnnGRUtanhRewardsList.append(drnnGRUtanhRewards15)
drnnGRUtanhRewardsList.append(drnnGRUtanhRewards16)
drnnGRUtanhRewardsList.append(drnnGRUtanhRewards17)
drnnGRUtanhRewardsList.append(drnnGRUtanhRewards18)
drnnGRUtanhRewardsList.append(drnnGRUtanhRewards19)
drnnGRUtanhRewardsList.append(drnnGRUtanhRewards20)
drnnGRUtanhRewardsList.append(drnnGRUtanhRewards21)
drnnGRUtanhRewardsList.append(drnnGRUtanhRewards22)
drnnGRUtanhRewardsList.append(drnnGRUtanhRewards23)
drnnGRUtanhRewardsList.append(drnnGRUtanhRewards24)
drnnGRUtanhRewardsList.append(drnnGRUtanhRewards25)
drnnGRUtanhRewardsList.append(drnnGRUtanhRewards26)
drnnGRUtanhRewardsList.append(drnnGRUtanhRewards27)
drnnGRUtanhRewardsList.append(drnnGRUtanhRewards28)
drnnGRUtanhRewardsList.append(drnnGRUtanhRewards29)
drnnGRUtanhRewardsList.append(drnnGRUtanhRewards30)
drnnGRUtanhRewardsList.append(drnnGRUtanhRewards31)
drnnGRUtanhRewardsList.append(drnnGRUtanhRewards32)
drnnGRUtanhRewardsList.append(drnnGRUtanhRewards33)
drnnGRUtanhRewardsList.append(drnnGRUtanhRewards34)
drnnGRUtanhRewardsList.append(drnnGRUtanhRewards35)
drnnGRUtanhRewardsList.append(drnnGRUtanhRewards36)
drnnGRUtanhRewardsList.append(drnnGRUtanhRewards37)
drnnGRUtanhRewardsList.append(drnnGRUtanhRewards38)
drnnGRUtanhRewardsList.append(drnnGRUtanhRewards39)
drnnGRUtanhRewardsList.append(drnnGRUtanhRewards40)
drnnGRUtanhRewardsList.append(drnnGRUtanhRewards41)
drnnGRUtanhRewardsList.append(drnnGRUtanhRewards42)
drnnGRUtanhRewardsList.append(drnnGRUtanhRewards43)
drnnGRUtanhRewardsList.append(drnnGRUtanhRewards44)
drnnGRUtanhRewardsList.append(drnnGRUtanhRewards45)
drnnGRUtanhRewardsList.append(drnnGRUtanhRewards46)
drnnGRUtanhRewardsList.append(drnnGRUtanhRewards47)
drnnGRUtanhRewardsList.append(drnnGRUtanhRewards48)
drnnGRUtanhRewardsList.append(drnnGRUtanhRewards49)
drnnGRUreluMakespan = []
drnnGRUreluRewards = []
drnnGRUreluMakespanList = []
drnnGRUreluRewardsList = []
drnnGRUreluMakespanValues = []
drnnGRUreluRewardsValues = []
drnnGRUreluMakespan.append(np.mean(drnnGRUreluMakespan0))
drnnGRUreluMakespan.append(np.mean(drnnGRUreluMakespan1))
drnnGRUreluMakespan.append(np.mean(drnnGRUreluMakespan2))
drnnGRUreluMakespan.append(np.mean(drnnGRUreluMakespan3))
drnnGRUreluMakespan.append(np.mean(drnnGRUreluMakespan4))
drnnGRUreluMakespan.append(np.mean(drnnGRUreluMakespan5))
drnnGRUreluMakespan.append(np.mean(drnnGRUreluMakespan6))
drnnGRUreluMakespan.append(np.mean(drnnGRUreluMakespan7))
drnnGRUreluMakespan.append(np.mean(drnnGRUreluMakespan8))
drnnGRUreluMakespan.append(np.mean(drnnGRUreluMakespan9))
drnnGRUreluMakespan.append(np.mean(drnnGRUreluMakespan10))
drnnGRUreluMakespan.append(np.mean(drnnGRUreluMakespan11))
drnnGRUreluMakespan.append(np.mean(drnnGRUreluMakespan12))
drnnGRUreluMakespan.append(np.mean(drnnGRUreluMakespan13))
drnnGRUreluMakespan.append(np.mean(drnnGRUreluMakespan14))
drnnGRUreluMakespan.append(np.mean(drnnGRUreluMakespan15))
drnnGRUreluMakespan.append(np.mean(drnnGRUreluMakespan16))
drnnGRUreluMakespan.append(np.mean(drnnGRUreluMakespan17))
drnnGRUreluMakespan.append(np.mean(drnnGRUreluMakespan18))
drnnGRUreluMakespan.append(np.mean(drnnGRUreluMakespan19))
drnnGRUreluMakespan.append(np.mean(drnnGRUreluMakespan20))
drnnGRUreluMakespan.append(np.mean(drnnGRUreluMakespan21))
drnnGRUreluMakespan.append(np.mean(drnnGRUreluMakespan22))
drnnGRUreluMakespan.append(np.mean(drnnGRUreluMakespan23))
drnnGRUreluMakespan.append(np.mean(drnnGRUreluMakespan24))
drnnGRUreluMakespan.append(np.mean(drnnGRUreluMakespan25))
drnnGRUreluMakespan.append(np.mean(drnnGRUreluMakespan26))
drnnGRUreluMakespan.append(np.mean(drnnGRUreluMakespan27))
drnnGRUreluMakespan.append(np.mean(drnnGRUreluMakespan28))
drnnGRUreluMakespan.append(np.mean(drnnGRUreluMakespan29))
drnnGRUreluMakespan.append(np.mean(drnnGRUreluMakespan30))
drnnGRUreluMakespan.append(np.mean(drnnGRUreluMakespan31))
drnnGRUreluMakespan.append(np.mean(drnnGRUreluMakespan32))
drnnGRUreluMakespan.append(np.mean(drnnGRUreluMakespan33))
drnnGRUreluMakespan.append(np.mean(drnnGRUreluMakespan34))
drnnGRUreluMakespan.append(np.mean(drnnGRUreluMakespan35))
drnnGRUreluMakespan.append(np.mean(drnnGRUreluMakespan36))
drnnGRUreluMakespan.append(np.mean(drnnGRUreluMakespan37))
drnnGRUreluMakespan.append(np.mean(drnnGRUreluMakespan38))
drnnGRUreluMakespan.append(np.mean(drnnGRUreluMakespan39))
drnnGRUreluMakespan.append(np.mean(drnnGRUreluMakespan40))
drnnGRUreluMakespan.append(np.mean(drnnGRUreluMakespan41))
drnnGRUreluMakespan.append(np.mean(drnnGRUreluMakespan42))
drnnGRUreluMakespan.append(np.mean(drnnGRUreluMakespan43))
drnnGRUreluMakespan.append(np.mean(drnnGRUreluMakespan44))
drnnGRUreluMakespan.append(np.mean(drnnGRUreluMakespan45))
drnnGRUreluMakespan.append(np.mean(drnnGRUreluMakespan46))
drnnGRUreluMakespan.append(np.mean(drnnGRUreluMakespan47))
drnnGRUreluMakespan.append(np.mean(drnnGRUreluMakespan48))
drnnGRUreluMakespan.append(np.mean(drnnGRUreluMakespan49))
drnnGRUreluRewards.append(np.mean(drnnGRUreluRewards0))
drnnGRUreluRewards.append(np.mean(drnnGRUreluRewards1))
drnnGRUreluRewards.append(np.mean(drnnGRUreluRewards2))
drnnGRUreluRewards.append(np.mean(drnnGRUreluRewards3))
drnnGRUreluRewards.append(np.mean(drnnGRUreluRewards4))
drnnGRUreluRewards.append(np.mean(drnnGRUreluRewards5))
drnnGRUreluRewards.append(np.mean(drnnGRUreluRewards6))
drnnGRUreluRewards.append(np.mean(drnnGRUreluRewards7))
drnnGRUreluRewards.append(np.mean(drnnGRUreluRewards8))
drnnGRUreluRewards.append(np.mean(drnnGRUreluRewards9))
drnnGRUreluRewards.append(np.mean(drnnGRUreluRewards10))
drnnGRUreluRewards.append(np.mean(drnnGRUreluRewards11))
drnnGRUreluRewards.append(np.mean(drnnGRUreluRewards12))
drnnGRUreluRewards.append(np.mean(drnnGRUreluRewards13))
drnnGRUreluRewards.append(np.mean(drnnGRUreluRewards14))
drnnGRUreluRewards.append(np.mean(drnnGRUreluRewards15))
drnnGRUreluRewards.append(np.mean(drnnGRUreluRewards16))
drnnGRUreluRewards.append(np.mean(drnnGRUreluRewards17))
drnnGRUreluRewards.append(np.mean(drnnGRUreluRewards18))
drnnGRUreluRewards.append(np.mean(drnnGRUreluRewards19))
drnnGRUreluRewards.append(np.mean(drnnGRUreluRewards20))
drnnGRUreluRewards.append(np.mean(drnnGRUreluRewards21))
drnnGRUreluRewards.append(np.mean(drnnGRUreluRewards22))
drnnGRUreluRewards.append(np.mean(drnnGRUreluRewards23))
drnnGRUreluRewards.append(np.mean(drnnGRUreluRewards24))
drnnGRUreluRewards.append(np.mean(drnnGRUreluRewards25))
drnnGRUreluRewards.append(np.mean(drnnGRUreluRewards26))
drnnGRUreluRewards.append(np.mean(drnnGRUreluRewards27))
drnnGRUreluRewards.append(np.mean(drnnGRUreluRewards28))
drnnGRUreluRewards.append(np.mean(drnnGRUreluRewards29))
drnnGRUreluRewards.append(np.mean(drnnGRUreluRewards30))
drnnGRUreluRewards.append(np.mean(drnnGRUreluRewards31))
drnnGRUreluRewards.append(np.mean(drnnGRUreluRewards32))
drnnGRUreluRewards.append(np.mean(drnnGRUreluRewards33))
drnnGRUreluRewards.append(np.mean(drnnGRUreluRewards34))
drnnGRUreluRewards.append(np.mean(drnnGRUreluRewards35))
drnnGRUreluRewards.append(np.mean(drnnGRUreluRewards36))
drnnGRUreluRewards.append(np.mean(drnnGRUreluRewards37))
drnnGRUreluRewards.append(np.mean(drnnGRUreluRewards38))
drnnGRUreluRewards.append(np.mean(drnnGRUreluRewards39))
drnnGRUreluRewards.append(np.mean(drnnGRUreluRewards40))
drnnGRUreluRewards.append(np.mean(drnnGRUreluRewards41))
drnnGRUreluRewards.append(np.mean(drnnGRUreluRewards42))
drnnGRUreluRewards.append(np.mean(drnnGRUreluRewards43))
drnnGRUreluRewards.append(np.mean(drnnGRUreluRewards44))
drnnGRUreluRewards.append(np.mean(drnnGRUreluRewards45))
drnnGRUreluRewards.append(np.mean(drnnGRUreluRewards46))
drnnGRUreluRewards.append(np.mean(drnnGRUreluRewards47))
drnnGRUreluRewards.append(np.mean(drnnGRUreluRewards48))
drnnGRUreluRewards.append(np.mean(drnnGRUreluRewards49))
drnnGRUreluMakespanList.append(drnnGRUreluMakespan0)
drnnGRUreluMakespanList.append(drnnGRUreluMakespan1)
drnnGRUreluMakespanList.append(drnnGRUreluMakespan2)
drnnGRUreluMakespanList.append(drnnGRUreluMakespan3)
drnnGRUreluMakespanList.append(drnnGRUreluMakespan4)
drnnGRUreluMakespanList.append(drnnGRUreluMakespan5)
drnnGRUreluMakespanList.append(drnnGRUreluMakespan6)
drnnGRUreluMakespanList.append(drnnGRUreluMakespan7)
drnnGRUreluMakespanList.append(drnnGRUreluMakespan8)
drnnGRUreluMakespanList.append(drnnGRUreluMakespan9)
drnnGRUreluMakespanList.append(drnnGRUreluMakespan10)
drnnGRUreluMakespanList.append(drnnGRUreluMakespan11)
drnnGRUreluMakespanList.append(drnnGRUreluMakespan12)
drnnGRUreluMakespanList.append(drnnGRUreluMakespan13)
drnnGRUreluMakespanList.append(drnnGRUreluMakespan14)
drnnGRUreluMakespanList.append(drnnGRUreluMakespan15)
drnnGRUreluMakespanList.append(drnnGRUreluMakespan16)
drnnGRUreluMakespanList.append(drnnGRUreluMakespan17)
drnnGRUreluMakespanList.append(drnnGRUreluMakespan18)
drnnGRUreluMakespanList.append(drnnGRUreluMakespan19)
drnnGRUreluMakespanList.append(drnnGRUreluMakespan20)
drnnGRUreluMakespanList.append(drnnGRUreluMakespan21)
drnnGRUreluMakespanList.append(drnnGRUreluMakespan22)
drnnGRUreluMakespanList.append(drnnGRUreluMakespan23)
drnnGRUreluMakespanList.append(drnnGRUreluMakespan24)
drnnGRUreluMakespanList.append(drnnGRUreluMakespan25)
drnnGRUreluMakespanList.append(drnnGRUreluMakespan26)
drnnGRUreluMakespanList.append(drnnGRUreluMakespan27)
drnnGRUreluMakespanList.append(drnnGRUreluMakespan28)
drnnGRUreluMakespanList.append(drnnGRUreluMakespan29)
drnnGRUreluMakespanList.append(drnnGRUreluMakespan30)
drnnGRUreluMakespanList.append(drnnGRUreluMakespan31)
drnnGRUreluMakespanList.append(drnnGRUreluMakespan32)
drnnGRUreluMakespanList.append(drnnGRUreluMakespan33)
drnnGRUreluMakespanList.append(drnnGRUreluMakespan34)
drnnGRUreluMakespanList.append(drnnGRUreluMakespan35)
drnnGRUreluMakespanList.append(drnnGRUreluMakespan36)
drnnGRUreluMakespanList.append(drnnGRUreluMakespan37)
drnnGRUreluMakespanList.append(drnnGRUreluMakespan38)
drnnGRUreluMakespanList.append(drnnGRUreluMakespan39)
drnnGRUreluMakespanList.append(drnnGRUreluMakespan40)
drnnGRUreluMakespanList.append(drnnGRUreluMakespan41)
drnnGRUreluMakespanList.append(drnnGRUreluMakespan42)
drnnGRUreluMakespanList.append(drnnGRUreluMakespan43)
drnnGRUreluMakespanList.append(drnnGRUreluMakespan44)
drnnGRUreluMakespanList.append(drnnGRUreluMakespan45)
drnnGRUreluMakespanList.append(drnnGRUreluMakespan46)
drnnGRUreluMakespanList.append(drnnGRUreluMakespan47)
drnnGRUreluMakespanList.append(drnnGRUreluMakespan48)
drnnGRUreluMakespanList.append(drnnGRUreluMakespan49)
drnnGRUreluRewardsList.append(drnnGRUreluRewards0)
drnnGRUreluRewardsList.append(drnnGRUreluRewards1)
drnnGRUreluRewardsList.append(drnnGRUreluRewards2)
drnnGRUreluRewardsList.append(drnnGRUreluRewards3)
drnnGRUreluRewardsList.append(drnnGRUreluRewards4)
drnnGRUreluRewardsList.append(drnnGRUreluRewards5)
drnnGRUreluRewardsList.append(drnnGRUreluRewards6)
drnnGRUreluRewardsList.append(drnnGRUreluRewards7)
drnnGRUreluRewardsList.append(drnnGRUreluRewards8)
drnnGRUreluRewardsList.append(drnnGRUreluRewards9)
drnnGRUreluRewardsList.append(drnnGRUreluRewards10)
drnnGRUreluRewardsList.append(drnnGRUreluRewards11)
drnnGRUreluRewardsList.append(drnnGRUreluRewards12)
drnnGRUreluRewardsList.append(drnnGRUreluRewards13)
drnnGRUreluRewardsList.append(drnnGRUreluRewards14)
drnnGRUreluRewardsList.append(drnnGRUreluRewards15)
drnnGRUreluRewardsList.append(drnnGRUreluRewards16)
drnnGRUreluRewardsList.append(drnnGRUreluRewards17)
drnnGRUreluRewardsList.append(drnnGRUreluRewards18)
drnnGRUreluRewardsList.append(drnnGRUreluRewards19)
drnnGRUreluRewardsList.append(drnnGRUreluRewards20)
drnnGRUreluRewardsList.append(drnnGRUreluRewards21)
drnnGRUreluRewardsList.append(drnnGRUreluRewards22)
drnnGRUreluRewardsList.append(drnnGRUreluRewards23)
drnnGRUreluRewardsList.append(drnnGRUreluRewards24)
drnnGRUreluRewardsList.append(drnnGRUreluRewards25)
drnnGRUreluRewardsList.append(drnnGRUreluRewards26)
drnnGRUreluRewardsList.append(drnnGRUreluRewards27)
drnnGRUreluRewardsList.append(drnnGRUreluRewards28)
drnnGRUreluRewardsList.append(drnnGRUreluRewards29)
drnnGRUreluRewardsList.append(drnnGRUreluRewards30)
drnnGRUreluRewardsList.append(drnnGRUreluRewards31)
drnnGRUreluRewardsList.append(drnnGRUreluRewards32)
drnnGRUreluRewardsList.append(drnnGRUreluRewards33)
drnnGRUreluRewardsList.append(drnnGRUreluRewards34)
drnnGRUreluRewardsList.append(drnnGRUreluRewards35)
drnnGRUreluRewardsList.append(drnnGRUreluRewards36)
drnnGRUreluRewardsList.append(drnnGRUreluRewards37)
drnnGRUreluRewardsList.append(drnnGRUreluRewards38)
drnnGRUreluRewardsList.append(drnnGRUreluRewards39)
drnnGRUreluRewardsList.append(drnnGRUreluRewards40)
drnnGRUreluRewardsList.append(drnnGRUreluRewards41)
drnnGRUreluRewardsList.append(drnnGRUreluRewards42)
drnnGRUreluRewardsList.append(drnnGRUreluRewards43)
drnnGRUreluRewardsList.append(drnnGRUreluRewards44)
drnnGRUreluRewardsList.append(drnnGRUreluRewards45)
drnnGRUreluRewardsList.append(drnnGRUreluRewards46)
drnnGRUreluRewardsList.append(drnnGRUreluRewards47)
drnnGRUreluRewardsList.append(drnnGRUreluRewards48)
drnnGRUreluRewardsList.append(drnnGRUreluRewards49)
for vector in drnnGRUtanhMakespanList:
for element in vector:
drnnGRUtanhMakespanValues.append(element)
for vector in drnnGRUtanhRewardsList:
for element in vector:
drnnGRUtanhRewardsValues.append(element)
##################
for vector in drnnGRUreluMakespanList:
for element in vector:
drnnGRUreluMakespanValues.append(element)
for vector in drnnGRUreluRewardsList:
for element in vector:
drnnGRUreluRewardsValues.append(element)
#####################
smoothGRUtanhMakespanValues = pd.Series(drnnGRUtanhMakespanValues).rolling(12).mean()
plt.plot(smoothGRUtanhMakespanValues)
plt.xlabel("Episodios")
plt.ylabel("Segundos")
plt.title("'Makespan' con red neuronal profunda que incluye 1 capa GRU")
plt.show()
smoothGRUtanhRewardsValues = pd.Series(drnnGRUtanhRewardsValues).rolling(12).mean()
plt.plot(smoothGRUtanhRewardsValues)
plt.xlabel("Episodios")
plt.ylabel("Premio")
plt.title("'Reward' con red neuronal profunda que incluye 1 capa GRU")
plt.show()
#####################
smoothGRUreluMakespanValues = pd.Series(drnnGRUreluMakespanValues).rolling(12).mean()
plt.plot(smoothGRUreluMakespanValues)
plt.xlabel("Episodios")
plt.ylabel("Segundos")
plt.title("'Makespan' con red neuronal profunda que incluye 1 capa GRU y ReLU")
plt.show()
smoothGRUreluRewardsValues = pd.Series(drnnGRUreluRewardsValues).rolling(12).mean()
plt.plot(smoothGRUreluRewardsValues)
plt.xlabel("Episodios")
plt.ylabel("Premio")
plt.title("'Reward' con red neuronal profunda que incluye 1 capa GRU y ReLU")
plt.show()
###################
plt.plot(smoothGRUtanhMakespanValues, color='blue', label='tanh')
plt.plot(smoothGRUreluMakespanValues, color='orange', label='relu')
plt.xlabel("Episodios")
plt.ylabel("Segundos")
plt.title("'Makespan' con red neuronal profunda que incluye 1 capa GRU")
plt.legend()
plt.show()
###################
plt.plot(smoothGRUtanhRewardsValues, color='blue', label='tanh')
plt.plot(smoothGRUreluRewardsValues, color='orange', label='relu')
plt.xlabel("Episodios")
plt.ylabel("Premio")
plt.title("'Reward' con red neuronal profunda que incluye 1 capa GRU")
plt.legend()
plt.show()
###################
drnnLSTMtanhMakespan = []
drnnLSTMtanhRewards = []
drnnLSTMtanhMakespanList = []
drnnLSTMtanhRewardsList = []
drnnLSTMtanhMakespanValues = []
drnnLSTMtanhRewardsValues = []
drnnLSTMtanhMakespan.append(np.mean(drnnLSTMtanhMakespan0))
drnnLSTMtanhMakespan.append(np.mean(drnnLSTMtanhMakespan1))
drnnLSTMtanhMakespan.append(np.mean(drnnLSTMtanhMakespan2))
drnnLSTMtanhMakespan.append(np.mean(drnnLSTMtanhMakespan3))
drnnLSTMtanhMakespan.append(np.mean(drnnLSTMtanhMakespan4))
drnnLSTMtanhMakespan.append(np.mean(drnnLSTMtanhMakespan5))
drnnLSTMtanhMakespan.append(np.mean(drnnLSTMtanhMakespan6))
drnnLSTMtanhMakespan.append(np.mean(drnnLSTMtanhMakespan7))
drnnLSTMtanhMakespan.append(np.mean(drnnLSTMtanhMakespan8))
drnnLSTMtanhMakespan.append(np.mean(drnnLSTMtanhMakespan9))
drnnLSTMtanhMakespan.append(np.mean(drnnLSTMtanhMakespan10))
drnnLSTMtanhMakespan.append(np.mean(drnnLSTMtanhMakespan11))
drnnLSTMtanhMakespan.append(np.mean(drnnLSTMtanhMakespan12))
drnnLSTMtanhMakespan.append(np.mean(drnnLSTMtanhMakespan13))
drnnLSTMtanhMakespan.append(np.mean(drnnLSTMtanhMakespan14))
drnnLSTMtanhMakespan.append(np.mean(drnnLSTMtanhMakespan15))
drnnLSTMtanhMakespan.append(np.mean(drnnLSTMtanhMakespan16))
drnnLSTMtanhMakespan.append(np.mean(drnnLSTMtanhMakespan17))
drnnLSTMtanhMakespan.append(np.mean(drnnLSTMtanhMakespan18))
drnnLSTMtanhMakespan.append(np.mean(drnnLSTMtanhMakespan19))
drnnLSTMtanhMakespan.append(np.mean(drnnLSTMtanhMakespan20))
drnnLSTMtanhMakespan.append(np.mean(drnnLSTMtanhMakespan21))
drnnLSTMtanhMakespan.append(np.mean(drnnLSTMtanhMakespan22))
drnnLSTMtanhMakespan.append(np.mean(drnnLSTMtanhMakespan23))
drnnLSTMtanhMakespan.append(np.mean(drnnLSTMtanhMakespan24))
drnnLSTMtanhMakespan.append(np.mean(drnnLSTMtanhMakespan25))
drnnLSTMtanhMakespan.append(np.mean(drnnLSTMtanhMakespan26))
drnnLSTMtanhMakespan.append(np.mean(drnnLSTMtanhMakespan27))
drnnLSTMtanhMakespan.append(np.mean(drnnLSTMtanhMakespan28))
drnnLSTMtanhMakespan.append(np.mean(drnnLSTMtanhMakespan29))
drnnLSTMtanhMakespan.append(np.mean(drnnLSTMtanhMakespan30))
drnnLSTMtanhMakespan.append(np.mean(drnnLSTMtanhMakespan31))
drnnLSTMtanhMakespan.append(np.mean(drnnLSTMtanhMakespan32))
drnnLSTMtanhMakespan.append(np.mean(drnnLSTMtanhMakespan33))
drnnLSTMtanhMakespan.append(np.mean(drnnLSTMtanhMakespan34))
drnnLSTMtanhMakespan.append(np.mean(drnnLSTMtanhMakespan35))
drnnLSTMtanhMakespan.append(np.mean(drnnLSTMtanhMakespan36))
drnnLSTMtanhMakespan.append(np.mean(drnnLSTMtanhMakespan37))
drnnLSTMtanhMakespan.append(np.mean(drnnLSTMtanhMakespan38))
drnnLSTMtanhMakespan.append(np.mean(drnnLSTMtanhMakespan39))
drnnLSTMtanhMakespan.append(np.mean(drnnLSTMtanhMakespan40))
drnnLSTMtanhMakespan.append(np.mean(drnnLSTMtanhMakespan41))
drnnLSTMtanhMakespan.append(np.mean(drnnLSTMtanhMakespan42))
drnnLSTMtanhMakespan.append(np.mean(drnnLSTMtanhMakespan43))
drnnLSTMtanhMakespan.append(np.mean(drnnLSTMtanhMakespan44))
drnnLSTMtanhMakespan.append(np.mean(drnnLSTMtanhMakespan45))
drnnLSTMtanhMakespan.append(np.mean(drnnLSTMtanhMakespan46))
drnnLSTMtanhMakespan.append(np.mean(drnnLSTMtanhMakespan47))
drnnLSTMtanhMakespan.append(np.mean(drnnLSTMtanhMakespan48))
drnnLSTMtanhMakespan.append(np.mean(drnnLSTMtanhMakespan49))
drnnLSTMtanhRewards.append(np.mean(drnnLSTMtanhRewards0))
drnnLSTMtanhRewards.append(np.mean(drnnLSTMtanhRewards1))
drnnLSTMtanhRewards.append(np.mean(drnnLSTMtanhRewards2))
drnnLSTMtanhRewards.append(np.mean(drnnLSTMtanhRewards3))
drnnLSTMtanhRewards.append(np.mean(drnnLSTMtanhRewards4))
drnnLSTMtanhRewards.append(np.mean(drnnLSTMtanhRewards5))
drnnLSTMtanhRewards.append(np.mean(drnnLSTMtanhRewards6))
drnnLSTMtanhRewards.append(np.mean(drnnLSTMtanhRewards7))
drnnLSTMtanhRewards.append(np.mean(drnnLSTMtanhRewards8))
drnnLSTMtanhRewards.append(np.mean(drnnLSTMtanhRewards9))
drnnLSTMtanhRewards.append(np.mean(drnnLSTMtanhRewards10))
drnnLSTMtanhRewards.append(np.mean(drnnLSTMtanhRewards11))
drnnLSTMtanhRewards.append(np.mean(drnnLSTMtanhRewards12))
drnnLSTMtanhRewards.append(np.mean(drnnLSTMtanhRewards13))
drnnLSTMtanhRewards.append(np.mean(drnnLSTMtanhRewards14))
drnnLSTMtanhRewards.append(np.mean(drnnLSTMtanhRewards15))
drnnLSTMtanhRewards.append(np.mean(drnnLSTMtanhRewards16))
drnnLSTMtanhRewards.append(np.mean(drnnLSTMtanhRewards17))
drnnLSTMtanhRewards.append(np.mean(drnnLSTMtanhRewards18))
drnnLSTMtanhRewards.append(np.mean(drnnLSTMtanhRewards19))
drnnLSTMtanhRewards.append(np.mean(drnnLSTMtanhRewards20))
drnnLSTMtanhRewards.append(np.mean(drnnLSTMtanhRewards21))
drnnLSTMtanhRewards.append(np.mean(drnnLSTMtanhRewards22))
drnnLSTMtanhRewards.append(np.mean(drnnLSTMtanhRewards23))
drnnLSTMtanhRewards.append(np.mean(drnnLSTMtanhRewards24))
drnnLSTMtanhRewards.append(np.mean(drnnLSTMtanhRewards25))
drnnLSTMtanhRewards.append(np.mean(drnnLSTMtanhRewards26))
drnnLSTMtanhRewards.append(np.mean(drnnLSTMtanhRewards27))
drnnLSTMtanhRewards.append(np.mean(drnnLSTMtanhRewards28))
drnnLSTMtanhRewards.append(np.mean(drnnLSTMtanhRewards29))
drnnLSTMtanhRewards.append(np.mean(drnnLSTMtanhRewards30))
drnnLSTMtanhRewards.append(np.mean(drnnLSTMtanhRewards31))
drnnLSTMtanhRewards.append(np.mean(drnnLSTMtanhRewards32))
drnnLSTMtanhRewards.append(np.mean(drnnLSTMtanhRewards33))
drnnLSTMtanhRewards.append(np.mean(drnnLSTMtanhRewards34))
drnnLSTMtanhRewards.append(np.mean(drnnLSTMtanhRewards35))
drnnLSTMtanhRewards.append(np.mean(drnnLSTMtanhRewards36))
drnnLSTMtanhRewards.append(np.mean(drnnLSTMtanhRewards37))
drnnLSTMtanhRewards.append(np.mean(drnnLSTMtanhRewards38))
drnnLSTMtanhRewards.append(np.mean(drnnLSTMtanhRewards39))
drnnLSTMtanhRewards.append(np.mean(drnnLSTMtanhRewards40))
drnnLSTMtanhRewards.append(np.mean(drnnLSTMtanhRewards41))
drnnLSTMtanhRewards.append(np.mean(drnnLSTMtanhRewards42))
drnnLSTMtanhRewards.append(np.mean(drnnLSTMtanhRewards43))
drnnLSTMtanhRewards.append(np.mean(drnnLSTMtanhRewards44))
drnnLSTMtanhRewards.append(np.mean(drnnLSTMtanhRewards45))
drnnLSTMtanhRewards.append(np.mean(drnnLSTMtanhRewards46))
drnnLSTMtanhRewards.append(np.mean(drnnLSTMtanhRewards47))
drnnLSTMtanhRewards.append(np.mean(drnnLSTMtanhRewards48))
drnnLSTMtanhRewards.append(np.mean(drnnLSTMtanhRewards49))
drnnLSTMtanhMakespanList.append(drnnLSTMtanhMakespan0)
drnnLSTMtanhMakespanList.append(drnnLSTMtanhMakespan1)
drnnLSTMtanhMakespanList.append(drnnLSTMtanhMakespan2)
drnnLSTMtanhMakespanList.append(drnnLSTMtanhMakespan3)
drnnLSTMtanhMakespanList.append(drnnLSTMtanhMakespan4)
drnnLSTMtanhMakespanList.append(drnnLSTMtanhMakespan5)
drnnLSTMtanhMakespanList.append(drnnLSTMtanhMakespan6)
drnnLSTMtanhMakespanList.append(drnnLSTMtanhMakespan7)
drnnLSTMtanhMakespanList.append(drnnLSTMtanhMakespan8)
drnnLSTMtanhMakespanList.append(drnnLSTMtanhMakespan9)
drnnLSTMtanhMakespanList.append(drnnLSTMtanhMakespan10)
drnnLSTMtanhMakespanList.append(drnnLSTMtanhMakespan11)
drnnLSTMtanhMakespanList.append(drnnLSTMtanhMakespan12)
drnnLSTMtanhMakespanList.append(drnnLSTMtanhMakespan13)
drnnLSTMtanhMakespanList.append(drnnLSTMtanhMakespan14)
drnnLSTMtanhMakespanList.append(drnnLSTMtanhMakespan15)
drnnLSTMtanhMakespanList.append(drnnLSTMtanhMakespan16)
drnnLSTMtanhMakespanList.append(drnnLSTMtanhMakespan17)
drnnLSTMtanhMakespanList.append(drnnLSTMtanhMakespan18)
drnnLSTMtanhMakespanList.append(drnnLSTMtanhMakespan19)
drnnLSTMtanhMakespanList.append(drnnLSTMtanhMakespan20)
drnnLSTMtanhMakespanList.append(drnnLSTMtanhMakespan21)
drnnLSTMtanhMakespanList.append(drnnLSTMtanhMakespan22)
drnnLSTMtanhMakespanList.append(drnnLSTMtanhMakespan23)
drnnLSTMtanhMakespanList.append(drnnLSTMtanhMakespan24)
drnnLSTMtanhMakespanList.append(drnnLSTMtanhMakespan25)
drnnLSTMtanhMakespanList.append(drnnLSTMtanhMakespan26)
drnnLSTMtanhMakespanList.append(drnnLSTMtanhMakespan27)
drnnLSTMtanhMakespanList.append(drnnLSTMtanhMakespan28)
drnnLSTMtanhMakespanList.append(drnnLSTMtanhMakespan29)
drnnLSTMtanhMakespanList.append(drnnLSTMtanhMakespan30)
drnnLSTMtanhMakespanList.append(drnnLSTMtanhMakespan31)
drnnLSTMtanhMakespanList.append(drnnLSTMtanhMakespan32)
drnnLSTMtanhMakespanList.append(drnnLSTMtanhMakespan33)
drnnLSTMtanhMakespanList.append(drnnLSTMtanhMakespan34)
drnnLSTMtanhMakespanList.append(drnnLSTMtanhMakespan35)
drnnLSTMtanhMakespanList.append(drnnLSTMtanhMakespan36)
drnnLSTMtanhMakespanList.append(drnnLSTMtanhMakespan37)
drnnLSTMtanhMakespanList.append(drnnLSTMtanhMakespan38)
drnnLSTMtanhMakespanList.append(drnnLSTMtanhMakespan39)
drnnLSTMtanhMakespanList.append(drnnLSTMtanhMakespan40)
drnnLSTMtanhMakespanList.append(drnnLSTMtanhMakespan41)
drnnLSTMtanhMakespanList.append(drnnLSTMtanhMakespan42)
drnnLSTMtanhMakespanList.append(drnnLSTMtanhMakespan43)
drnnLSTMtanhMakespanList.append(drnnLSTMtanhMakespan44)
drnnLSTMtanhMakespanList.append(drnnLSTMtanhMakespan45)
drnnLSTMtanhMakespanList.append(drnnLSTMtanhMakespan46)
drnnLSTMtanhMakespanList.append(drnnLSTMtanhMakespan47)
drnnLSTMtanhMakespanList.append(drnnLSTMtanhMakespan48)
drnnLSTMtanhMakespanList.append(drnnLSTMtanhMakespan49)
drnnLSTMtanhRewardsList.append(drnnLSTMtanhRewards0)
drnnLSTMtanhRewardsList.append(drnnLSTMtanhRewards1)
drnnLSTMtanhRewardsList.append(drnnLSTMtanhRewards2)
drnnLSTMtanhRewardsList.append(drnnLSTMtanhRewards3)
drnnLSTMtanhRewardsList.append(drnnLSTMtanhRewards4)
drnnLSTMtanhRewardsList.append(drnnLSTMtanhRewards5)
drnnLSTMtanhRewardsList.append(drnnLSTMtanhRewards6)
drnnLSTMtanhRewardsList.append(drnnLSTMtanhRewards7)
drnnLSTMtanhRewardsList.append(drnnLSTMtanhRewards8)
drnnLSTMtanhRewardsList.append(drnnLSTMtanhRewards9)
drnnLSTMtanhRewardsList.append(drnnLSTMtanhRewards10)
drnnLSTMtanhRewardsList.append(drnnLSTMtanhRewards11)
drnnLSTMtanhRewardsList.append(drnnLSTMtanhRewards12)
drnnLSTMtanhRewardsList.append(drnnLSTMtanhRewards13)
drnnLSTMtanhRewardsList.append(drnnLSTMtanhRewards14)
drnnLSTMtanhRewardsList.append(drnnLSTMtanhRewards15)
drnnLSTMtanhRewardsList.append(drnnLSTMtanhRewards16)
drnnLSTMtanhRewardsList.append(drnnLSTMtanhRewards17)
drnnLSTMtanhRewardsList.append(drnnLSTMtanhRewards18)
drnnLSTMtanhRewardsList.append(drnnLSTMtanhRewards19)
drnnLSTMtanhRewardsList.append(drnnLSTMtanhRewards20)
drnnLSTMtanhRewardsList.append(drnnLSTMtanhRewards21)
drnnLSTMtanhRewardsList.append(drnnLSTMtanhRewards22)
drnnLSTMtanhRewardsList.append(drnnLSTMtanhRewards23)
drnnLSTMtanhRewardsList.append(drnnLSTMtanhRewards24)
drnnLSTMtanhRewardsList.append(drnnLSTMtanhRewards25)
drnnLSTMtanhRewardsList.append(drnnLSTMtanhRewards26)
drnnLSTMtanhRewardsList.append(drnnLSTMtanhRewards27)
drnnLSTMtanhRewardsList.append(drnnLSTMtanhRewards28)
drnnLSTMtanhRewardsList.append(drnnLSTMtanhRewards29)
drnnLSTMtanhRewardsList.append(drnnLSTMtanhRewards30)
drnnLSTMtanhRewardsList.append(drnnLSTMtanhRewards31)
drnnLSTMtanhRewardsList.append(drnnLSTMtanhRewards32)
drnnLSTMtanhRewardsList.append(drnnLSTMtanhRewards33)
drnnLSTMtanhRewardsList.append(drnnLSTMtanhRewards34)
drnnLSTMtanhRewardsList.append(drnnLSTMtanhRewards35)
drnnLSTMtanhRewardsList.append(drnnLSTMtanhRewards36)
drnnLSTMtanhRewardsList.append(drnnLSTMtanhRewards37)
drnnLSTMtanhRewardsList.append(drnnLSTMtanhRewards38)
drnnLSTMtanhRewardsList.append(drnnLSTMtanhRewards39)
drnnLSTMtanhRewardsList.append(drnnLSTMtanhRewards40)
drnnLSTMtanhRewardsList.append(drnnLSTMtanhRewards41)
drnnLSTMtanhRewardsList.append(drnnLSTMtanhRewards42)
drnnLSTMtanhRewardsList.append(drnnLSTMtanhRewards43)
drnnLSTMtanhRewardsList.append(drnnLSTMtanhRewards44)
drnnLSTMtanhRewardsList.append(drnnLSTMtanhRewards45)
drnnLSTMtanhRewardsList.append(drnnLSTMtanhRewards46)
drnnLSTMtanhRewardsList.append(drnnLSTMtanhRewards47)
drnnLSTMtanhRewardsList.append(drnnLSTMtanhRewards48)
drnnLSTMtanhRewardsList.append(drnnLSTMtanhRewards49)
for vector in drnnLSTMtanhMakespanList:
for element in vector:
drnnLSTMtanhMakespanValues.append(element)
for vector in drnnLSTMtanhRewardsList:
for element in vector:
drnnLSTMtanhRewardsValues.append(element)
smoothLSTMtanhMakespanValues = pd.Series(drnnLSTMtanhMakespanValues).rolling(12).mean()
plt.plot(smoothLSTMtanhMakespanValues)
plt.xlabel("Episodios")
plt.ylabel("Segundos")
plt.title("'Makespan' utilizando LSTM con tanh")
plt.show()
smoothLSTMtanhRewardsValues = pd.Series(drnnLSTMtanhRewardsValues).rolling(12).mean()
plt.plot(smoothLSTMtanhRewardsValues)
plt.xlabel("Episodios")
plt.ylabel("Premio")
plt.title("'Reward' utilizando LSTM con tanh")
plt.show()
####################
drnnLSTMreluMakespan = []
drnnLSTMreluRewards = []
drnnLSTMreluMakespanList = []
drnnLSTMreluRewardsList = []
drnnLSTMreluMakespanValues = []
drnnLSTMreluRewardsValues = []
drnnLSTMreluMakespan.append(np.mean(drnnLSTMreluMakespan0))
drnnLSTMreluMakespan.append(np.mean(drnnLSTMreluMakespan1))
drnnLSTMreluMakespan.append(np.mean(drnnLSTMreluMakespan2))
drnnLSTMreluMakespan.append(np.mean(drnnLSTMreluMakespan3))
drnnLSTMreluMakespan.append(np.mean(drnnLSTMreluMakespan4))
drnnLSTMreluMakespan.append(np.mean(drnnLSTMreluMakespan5))
drnnLSTMreluMakespan.append(np.mean(drnnLSTMreluMakespan6))
drnnLSTMreluMakespan.append(np.mean(drnnLSTMreluMakespan7))
drnnLSTMreluMakespan.append(np.mean(drnnLSTMreluMakespan8))
drnnLSTMreluMakespan.append(np.mean(drnnLSTMreluMakespan9))
drnnLSTMreluMakespan.append(np.mean(drnnLSTMreluMakespan10))
drnnLSTMreluMakespan.append(np.mean(drnnLSTMreluMakespan11))
drnnLSTMreluMakespan.append(np.mean(drnnLSTMreluMakespan12))
drnnLSTMreluMakespan.append(np.mean(drnnLSTMreluMakespan13))
drnnLSTMreluMakespan.append(np.mean(drnnLSTMreluMakespan14))
drnnLSTMreluMakespan.append(np.mean(drnnLSTMreluMakespan15))
drnnLSTMreluMakespan.append(np.mean(drnnLSTMreluMakespan16))
drnnLSTMreluMakespan.append(np.mean(drnnLSTMreluMakespan17))
drnnLSTMreluMakespan.append(np.mean(drnnLSTMreluMakespan18))
drnnLSTMreluMakespan.append(np.mean(drnnLSTMreluMakespan19))
drnnLSTMreluMakespan.append(np.mean(drnnLSTMreluMakespan20))
drnnLSTMreluMakespan.append(np.mean(drnnLSTMreluMakespan21))
drnnLSTMreluMakespan.append(np.mean(drnnLSTMreluMakespan22))
drnnLSTMreluMakespan.append(np.mean(drnnLSTMreluMakespan23))
drnnLSTMreluMakespan.append(np.mean(drnnLSTMreluMakespan24))
drnnLSTMreluMakespan.append(np.mean(drnnLSTMreluMakespan25))
drnnLSTMreluMakespan.append(np.mean(drnnLSTMreluMakespan26))
drnnLSTMreluMakespan.append(np.mean(drnnLSTMreluMakespan27))
drnnLSTMreluMakespan.append(np.mean(drnnLSTMreluMakespan28))
drnnLSTMreluMakespan.append(np.mean(drnnLSTMreluMakespan29))
drnnLSTMreluMakespan.append(np.mean(drnnLSTMreluMakespan30))
drnnLSTMreluMakespan.append(np.mean(drnnLSTMreluMakespan31))
drnnLSTMreluMakespan.append(np.mean(drnnLSTMreluMakespan32))
drnnLSTMreluMakespan.append(np.mean(drnnLSTMreluMakespan33))
drnnLSTMreluMakespan.append(np.mean(drnnLSTMreluMakespan34))
drnnLSTMreluMakespan.append(np.mean(drnnLSTMreluMakespan35))
drnnLSTMreluMakespan.append(np.mean(drnnLSTMreluMakespan36))
drnnLSTMreluMakespan.append(np.mean(drnnLSTMreluMakespan37))
drnnLSTMreluMakespan.append(np.mean(drnnLSTMreluMakespan38))
drnnLSTMreluMakespan.append(np.mean(drnnLSTMreluMakespan39))
drnnLSTMreluMakespan.append(np.mean(drnnLSTMreluMakespan40))
drnnLSTMreluMakespan.append(np.mean(drnnLSTMreluMakespan41))
drnnLSTMreluMakespan.append(np.mean(drnnLSTMreluMakespan42))
drnnLSTMreluMakespan.append(np.mean(drnnLSTMreluMakespan43))
drnnLSTMreluMakespan.append(np.mean(drnnLSTMreluMakespan44))
drnnLSTMreluMakespan.append(np.mean(drnnLSTMreluMakespan45))
drnnLSTMreluMakespan.append(np.mean(drnnLSTMreluMakespan46))
drnnLSTMreluMakespan.append(np.mean(drnnLSTMreluMakespan47))
drnnLSTMreluMakespan.append(np.mean(drnnLSTMreluMakespan48))
drnnLSTMreluMakespan.append(np.mean(drnnLSTMreluMakespan49))
drnnLSTMreluRewards.append(np.mean(drnnLSTMreluRewards0))
drnnLSTMreluRewards.append(np.mean(drnnLSTMreluRewards1))
drnnLSTMreluRewards.append(np.mean(drnnLSTMreluRewards2))
drnnLSTMreluRewards.append(np.mean(drnnLSTMreluRewards3))
drnnLSTMreluRewards.append(np.mean(drnnLSTMreluRewards4))
drnnLSTMreluRewards.append(np.mean(drnnLSTMreluRewards5))
drnnLSTMreluRewards.append(np.mean(drnnLSTMreluRewards6))
drnnLSTMreluRewards.append(np.mean(drnnLSTMreluRewards7))
drnnLSTMreluRewards.append(np.mean(drnnLSTMreluRewards8))
drnnLSTMreluRewards.append(np.mean(drnnLSTMreluRewards9))
drnnLSTMreluRewards.append(np.mean(drnnLSTMreluRewards10))
drnnLSTMreluRewards.append(np.mean(drnnLSTMreluRewards11))
drnnLSTMreluRewards.append(np.mean(drnnLSTMreluRewards12))
drnnLSTMreluRewards.append(np.mean(drnnLSTMreluRewards13))
drnnLSTMreluRewards.append(np.mean(drnnLSTMreluRewards14))
drnnLSTMreluRewards.append(np.mean(drnnLSTMreluRewards15))
drnnLSTMreluRewards.append(np.mean(drnnLSTMreluRewards16))
drnnLSTMreluRewards.append(np.mean(drnnLSTMreluRewards17))
drnnLSTMreluRewards.append(np.mean(drnnLSTMreluRewards18))
drnnLSTMreluRewards.append(np.mean(drnnLSTMreluRewards19))
drnnLSTMreluRewards.append(np.mean(drnnLSTMreluRewards20))
drnnLSTMreluRewards.append(np.mean(drnnLSTMreluRewards21))
drnnLSTMreluRewards.append(np.mean(drnnLSTMreluRewards22))
drnnLSTMreluRewards.append(np.mean(drnnLSTMreluRewards23))
drnnLSTMreluRewards.append(np.mean(drnnLSTMreluRewards24))
drnnLSTMreluRewards.append(np.mean(drnnLSTMreluRewards25))
drnnLSTMreluRewards.append(np.mean(drnnLSTMreluRewards26))
drnnLSTMreluRewards.append(np.mean(drnnLSTMreluRewards27))
drnnLSTMreluRewards.append(np.mean(drnnLSTMreluRewards28))
drnnLSTMreluRewards.append(np.mean(drnnLSTMreluRewards29))
drnnLSTMreluRewards.append(np.mean(drnnLSTMreluRewards30))
drnnLSTMreluRewards.append(np.mean(drnnLSTMreluRewards31))
drnnLSTMreluRewards.append(np.mean(drnnLSTMreluRewards32))
drnnLSTMreluRewards.append(np.mean(drnnLSTMreluRewards33))
drnnLSTMreluRewards.append(np.mean(drnnLSTMreluRewards34))
drnnLSTMreluRewards.append(np.mean(drnnLSTMreluRewards35))
drnnLSTMreluRewards.append(np.mean(drnnLSTMreluRewards36))
drnnLSTMreluRewards.append(np.mean(drnnLSTMreluRewards37))
drnnLSTMreluRewards.append(np.mean(drnnLSTMreluRewards38))
drnnLSTMreluRewards.append(np.mean(drnnLSTMreluRewards39))
drnnLSTMreluRewards.append(np.mean(drnnLSTMreluRewards40))
drnnLSTMreluRewards.append(np.mean(drnnLSTMreluRewards41))
drnnLSTMreluRewards.append(np.mean(drnnLSTMreluRewards42))
drnnLSTMreluRewards.append(np.mean(drnnLSTMreluRewards43))
drnnLSTMreluRewards.append(np.mean(drnnLSTMreluRewards44))
drnnLSTMreluRewards.append(np.mean(drnnLSTMreluRewards45))
drnnLSTMreluRewards.append(np.mean(drnnLSTMreluRewards46))
drnnLSTMreluRewards.append(np.mean(drnnLSTMreluRewards47))
drnnLSTMreluRewards.append(np.mean(drnnLSTMreluRewards48))
drnnLSTMreluRewards.append(np.mean(drnnLSTMreluRewards49))
drnnLSTMreluMakespanList.append(drnnLSTMreluMakespan0)
drnnLSTMreluMakespanList.append(drnnLSTMreluMakespan1)
drnnLSTMreluMakespanList.append(drnnLSTMreluMakespan2)
drnnLSTMreluMakespanList.append(drnnLSTMreluMakespan3)
drnnLSTMreluMakespanList.append(drnnLSTMreluMakespan4)
drnnLSTMreluMakespanList.append(drnnLSTMreluMakespan5)
drnnLSTMreluMakespanList.append(drnnLSTMreluMakespan6)
drnnLSTMreluMakespanList.append(drnnLSTMreluMakespan7)
drnnLSTMreluMakespanList.append(drnnLSTMreluMakespan8)
drnnLSTMreluMakespanList.append(drnnLSTMreluMakespan9)
drnnLSTMreluMakespanList.append(drnnLSTMreluMakespan10)
drnnLSTMreluMakespanList.append(drnnLSTMreluMakespan11)
drnnLSTMreluMakespanList.append(drnnLSTMreluMakespan12)
drnnLSTMreluMakespanList.append(drnnLSTMreluMakespan13)
drnnLSTMreluMakespanList.append(drnnLSTMreluMakespan14)
drnnLSTMreluMakespanList.append(drnnLSTMreluMakespan15)
drnnLSTMreluMakespanList.append(drnnLSTMreluMakespan16)
drnnLSTMreluMakespanList.append(drnnLSTMreluMakespan17)
drnnLSTMreluMakespanList.append(drnnLSTMreluMakespan18)
drnnLSTMreluMakespanList.append(drnnLSTMreluMakespan19)
drnnLSTMreluMakespanList.append(drnnLSTMreluMakespan20)
drnnLSTMreluMakespanList.append(drnnLSTMreluMakespan21)
drnnLSTMreluMakespanList.append(drnnLSTMreluMakespan22)
drnnLSTMreluMakespanList.append(drnnLSTMreluMakespan23)
drnnLSTMreluMakespanList.append(drnnLSTMreluMakespan24)
drnnLSTMreluMakespanList.append(drnnLSTMreluMakespan25)
drnnLSTMreluMakespanList.append(drnnLSTMreluMakespan26)
drnnLSTMreluMakespanList.append(drnnLSTMreluMakespan27)
drnnLSTMreluMakespanList.append(drnnLSTMreluMakespan28)
drnnLSTMreluMakespanList.append(drnnLSTMreluMakespan29)
drnnLSTMreluMakespanList.append(drnnLSTMreluMakespan30)
drnnLSTMreluMakespanList.append(drnnLSTMreluMakespan31)
drnnLSTMreluMakespanList.append(drnnLSTMreluMakespan32)
drnnLSTMreluMakespanList.append(drnnLSTMreluMakespan33)
drnnLSTMreluMakespanList.append(drnnLSTMreluMakespan34)
drnnLSTMreluMakespanList.append(drnnLSTMreluMakespan35)
drnnLSTMreluMakespanList.append(drnnLSTMreluMakespan36)
drnnLSTMreluMakespanList.append(drnnLSTMreluMakespan37)
drnnLSTMreluMakespanList.append(drnnLSTMreluMakespan38)
drnnLSTMreluMakespanList.append(drnnLSTMreluMakespan39)
drnnLSTMreluMakespanList.append(drnnLSTMreluMakespan40)
drnnLSTMreluMakespanList.append(drnnLSTMreluMakespan41)
drnnLSTMreluMakespanList.append(drnnLSTMreluMakespan42)
drnnLSTMreluMakespanList.append(drnnLSTMreluMakespan43)
drnnLSTMreluMakespanList.append(drnnLSTMreluMakespan44)
drnnLSTMreluMakespanList.append(drnnLSTMreluMakespan45)
drnnLSTMreluMakespanList.append(drnnLSTMreluMakespan46)
drnnLSTMreluMakespanList.append(drnnLSTMreluMakespan47)
drnnLSTMreluMakespanList.append(drnnLSTMreluMakespan48)
drnnLSTMreluMakespanList.append(drnnLSTMreluMakespan49)
drnnLSTMreluRewardsList.append(drnnLSTMreluRewards0)
drnnLSTMreluRewardsList.append(drnnLSTMreluRewards1)
drnnLSTMreluRewardsList.append(drnnLSTMreluRewards2)
drnnLSTMreluRewardsList.append(drnnLSTMreluRewards3)
drnnLSTMreluRewardsList.append(drnnLSTMreluRewards4)
drnnLSTMreluRewardsList.append(drnnLSTMreluRewards5)
drnnLSTMreluRewardsList.append(drnnLSTMreluRewards6)
drnnLSTMreluRewardsList.append(drnnLSTMreluRewards7)
drnnLSTMreluRewardsList.append(drnnLSTMreluRewards8)
drnnLSTMreluRewardsList.append(drnnLSTMreluRewards9)
drnnLSTMreluRewardsList.append(drnnLSTMreluRewards10)
drnnLSTMreluRewardsList.append(drnnLSTMreluRewards11)
drnnLSTMreluRewardsList.append(drnnLSTMreluRewards12)
drnnLSTMreluRewardsList.append(drnnLSTMreluRewards13)
drnnLSTMreluRewardsList.append(drnnLSTMreluRewards14)
drnnLSTMreluRewardsList.append(drnnLSTMreluRewards15)
drnnLSTMreluRewardsList.append(drnnLSTMreluRewards16)
drnnLSTMreluRewardsList.append(drnnLSTMreluRewards17)
drnnLSTMreluRewardsList.append(drnnLSTMreluRewards18)
drnnLSTMreluRewardsList.append(drnnLSTMreluRewards19)
drnnLSTMreluRewardsList.append(drnnLSTMreluRewards20)
drnnLSTMreluRewardsList.append(drnnLSTMreluRewards21)
drnnLSTMreluRewardsList.append(drnnLSTMreluRewards22)
drnnLSTMreluRewardsList.append(drnnLSTMreluRewards23)
drnnLSTMreluRewardsList.append(drnnLSTMreluRewards24)
drnnLSTMreluRewardsList.append(drnnLSTMreluRewards25)
drnnLSTMreluRewardsList.append(drnnLSTMreluRewards26)
drnnLSTMreluRewardsList.append(drnnLSTMreluRewards27)
drnnLSTMreluRewardsList.append(drnnLSTMreluRewards28)
drnnLSTMreluRewardsList.append(drnnLSTMreluRewards29)
drnnLSTMreluRewardsList.append(drnnLSTMreluRewards30)
drnnLSTMreluRewardsList.append(drnnLSTMreluRewards31)
drnnLSTMreluRewardsList.append(drnnLSTMreluRewards32)
drnnLSTMreluRewardsList.append(drnnLSTMreluRewards33)
drnnLSTMreluRewardsList.append(drnnLSTMreluRewards34)
drnnLSTMreluRewardsList.append(drnnLSTMreluRewards35)
drnnLSTMreluRewardsList.append(drnnLSTMreluRewards36)
drnnLSTMreluRewardsList.append(drnnLSTMreluRewards37)
drnnLSTMreluRewardsList.append(drnnLSTMreluRewards38)
drnnLSTMreluRewardsList.append(drnnLSTMreluRewards39)
drnnLSTMreluRewardsList.append(drnnLSTMreluRewards40)
drnnLSTMreluRewardsList.append(drnnLSTMreluRewards41)
drnnLSTMreluRewardsList.append(drnnLSTMreluRewards42)
drnnLSTMreluRewardsList.append(drnnLSTMreluRewards43)
drnnLSTMreluRewardsList.append(drnnLSTMreluRewards44)
drnnLSTMreluRewardsList.append(drnnLSTMreluRewards45)
drnnLSTMreluRewardsList.append(drnnLSTMreluRewards46)
drnnLSTMreluRewardsList.append(drnnLSTMreluRewards47)
drnnLSTMreluRewardsList.append(drnnLSTMreluRewards48)
drnnLSTMreluRewardsList.append(drnnLSTMreluRewards49)
for vector in drnnLSTMreluMakespanList:
for element in vector:
drnnLSTMreluMakespanValues.append(element)
for vector in drnnLSTMreluRewardsList:
for element in vector:
drnnLSTMreluRewardsValues.append(element)
smoothLSTMreluMakespanValues = pd.Series(drnnLSTMreluMakespanValues).rolling(12).mean()
plt.plot(smoothLSTMreluMakespanValues)
plt.xlabel("Episodios")
plt.ylabel("Segundos")
plt.title("'Makespan' utilizando LSTM con relu")
plt.show()
smoothLSTMreluRewardsValues = pd.Series(drnnLSTMreluRewardsValues).rolling(12).mean()
plt.plot(smoothLSTMreluRewardsValues)
plt.xlabel("Episodios")
plt.ylabel("Premio")
plt.title("'Reward' utilizando LSTM con relu")
plt.show()
##################
plt.plot(smoothLSTMtanhMakespanValues, color='blue', label='tanh')
plt.plot(smoothLSTMreluMakespanValues, color='orange', label='relu')
plt.xlabel("Episodios")
plt.ylabel("Segundos")
plt.title("'Makespan' con red neuronal profunda que incluye 1 capa LSTM")
plt.legend()
plt.show()
##################
plt.plot(smoothLSTMtanhRewardsValues, color='blue', label='tanh')
plt.plot(smoothLSTMreluRewardsValues, color='orange', label='relu')
plt.xlabel("Episodios")
plt.ylabel("Premio")
plt.title("'Reward' con red neuronal profunda que incluye 1 capa LSTM")
plt.legend()
plt.show()
##################
##################
##################
drlTanhMakespan = []
drlTanhRewards = []
drlTanhMakespanList = []
drlTanhRewardsList = []
drlTanhMakespanValues = []
drlTanhRewardsValues = []
drlTanhMakespan.append(np.mean(drlTanhMakespan0))
drlTanhMakespan.append(np.mean(drlTanhMakespan1))
drlTanhMakespan.append(np.mean(drlTanhMakespan2))
drlTanhMakespan.append(np.mean(drlTanhMakespan3))
drlTanhMakespan.append(np.mean(drlTanhMakespan4))
drlTanhMakespan.append(np.mean(drlTanhMakespan5))
drlTanhMakespan.append(np.mean(drlTanhMakespan6))
drlTanhMakespan.append(np.mean(drlTanhMakespan7))
drlTanhMakespan.append(np.mean(drlTanhMakespan8))
drlTanhMakespan.append(np.mean(drlTanhMakespan9))
drlTanhMakespan.append(np.mean(drlTanhMakespan10))
drlTanhMakespan.append(np.mean(drlTanhMakespan11))
drlTanhMakespan.append(np.mean(drlTanhMakespan12))
drlTanhMakespan.append(np.mean(drlTanhMakespan13))
drlTanhMakespan.append(np.mean(drlTanhMakespan14))
drlTanhMakespan.append(np.mean(drlTanhMakespan15))
drlTanhMakespan.append(np.mean(drlTanhMakespan16))
drlTanhMakespan.append(np.mean(drlTanhMakespan17))
drlTanhMakespan.append(np.mean(drlTanhMakespan18))
drlTanhMakespan.append(np.mean(drlTanhMakespan19))
drlTanhMakespan.append(np.mean(drlTanhMakespan20))
drlTanhMakespan.append(np.mean(drlTanhMakespan21))
drlTanhMakespan.append(np.mean(drlTanhMakespan22))
drlTanhMakespan.append(np.mean(drlTanhMakespan23))
drlTanhMakespan.append(np.mean(drlTanhMakespan24))
drlTanhMakespan.append(np.mean(drlTanhMakespan25))
drlTanhMakespan.append(np.mean(drlTanhMakespan26))
drlTanhMakespan.append(np.mean(drlTanhMakespan27))
drlTanhMakespan.append(np.mean(drlTanhMakespan28))
drlTanhMakespan.append(np.mean(drlTanhMakespan29))
drlTanhMakespan.append(np.mean(drlTanhMakespan30))
drlTanhMakespan.append(np.mean(drlTanhMakespan31))
drlTanhMakespan.append(np.mean(drlTanhMakespan32))
drlTanhMakespan.append(np.mean(drlTanhMakespan33))
drlTanhMakespan.append(np.mean(drlTanhMakespan34))
drlTanhMakespan.append(np.mean(drlTanhMakespan35))
drlTanhMakespan.append(np.mean(drlTanhMakespan36))
drlTanhMakespan.append(np.mean(drlTanhMakespan37))
drlTanhMakespan.append(np.mean(drlTanhMakespan38))
drlTanhMakespan.append(np.mean(drlTanhMakespan39))
drlTanhMakespan.append(np.mean(drlTanhMakespan40))
drlTanhMakespan.append(np.mean(drlTanhMakespan41))
drlTanhMakespan.append(np.mean(drlTanhMakespan42))
drlTanhMakespan.append(np.mean(drlTanhMakespan43))
drlTanhMakespan.append(np.mean(drlTanhMakespan44))
drlTanhMakespan.append(np.mean(drlTanhMakespan45))
drlTanhMakespan.append(np.mean(drlTanhMakespan46))
drlTanhMakespan.append(np.mean(drlTanhMakespan47))
drlTanhMakespan.append(np.mean(drlTanhMakespan48))
drlTanhMakespan.append(np.mean(drlTanhMakespan49))
drlTanhRewards.append(np.mean(drlTanhRewards0))
drlTanhRewards.append(np.mean(drlTanhRewards1))
drlTanhRewards.append(np.mean(drlTanhRewards2))
drlTanhRewards.append(np.mean(drlTanhRewards3))
drlTanhRewards.append(np.mean(drlTanhRewards4))
drlTanhRewards.append(np.mean(drlTanhRewards5))
drlTanhRewards.append(np.mean(drlTanhRewards6))
drlTanhRewards.append(np.mean(drlTanhRewards7))
drlTanhRewards.append(np.mean(drlTanhRewards8))
drlTanhRewards.append(np.mean(drlTanhRewards9))
drlTanhRewards.append(np.mean(drlTanhRewards10))
drlTanhRewards.append(np.mean(drlTanhRewards11))
drlTanhRewards.append(np.mean(drlTanhRewards12))
drlTanhRewards.append(np.mean(drlTanhRewards13))
drlTanhRewards.append(np.mean(drlTanhRewards14))
drlTanhRewards.append(np.mean(drlTanhRewards15))
drlTanhRewards.append(np.mean(drlTanhRewards16))
drlTanhRewards.append(np.mean(drlTanhRewards17))
drlTanhRewards.append(np.mean(drlTanhRewards18))
drlTanhRewards.append(np.mean(drlTanhRewards19))
drlTanhRewards.append(np.mean(drlTanhRewards20))
drlTanhRewards.append(np.mean(drlTanhRewards21))
drlTanhRewards.append(np.mean(drlTanhRewards22))
drlTanhRewards.append(np.mean(drlTanhRewards23))
drlTanhRewards.append(np.mean(drlTanhRewards24))
drlTanhRewards.append(np.mean(drlTanhRewards25))
drlTanhRewards.append(np.mean(drlTanhRewards26))
drlTanhRewards.append(np.mean(drlTanhRewards27))
drlTanhRewards.append(np.mean(drlTanhRewards28))
drlTanhRewards.append(np.mean(drlTanhRewards29))
drlTanhRewards.append(np.mean(drlTanhRewards30))
drlTanhRewards.append(np.mean(drlTanhRewards31))
drlTanhRewards.append(np.mean(drlTanhRewards32))
drlTanhRewards.append(np.mean(drlTanhRewards33))
drlTanhRewards.append(np.mean(drlTanhRewards34))
drlTanhRewards.append(np.mean(drlTanhRewards35))
drlTanhRewards.append(np.mean(drlTanhRewards36))
drlTanhRewards.append(np.mean(drlTanhRewards37))
drlTanhRewards.append(np.mean(drlTanhRewards38))
drlTanhRewards.append(np.mean(drlTanhRewards39))
drlTanhRewards.append(np.mean(drlTanhRewards40))
drlTanhRewards.append(np.mean(drlTanhRewards41))
drlTanhRewards.append(np.mean(drlTanhRewards42))
drlTanhRewards.append(np.mean(drlTanhRewards43))
drlTanhRewards.append(np.mean(drlTanhRewards44))
drlTanhRewards.append(np.mean(drlTanhRewards45))
drlTanhRewards.append(np.mean(drlTanhRewards46))
drlTanhRewards.append(np.mean(drlTanhRewards47))
drlTanhRewards.append(np.mean(drlTanhRewards48))
drlTanhRewards.append(np.mean(drlTanhRewards49))
drlTanhMakespanList.append(drlTanhMakespan0)
drlTanhMakespanList.append(drlTanhMakespan1)
drlTanhMakespanList.append(drlTanhMakespan2)
drlTanhMakespanList.append(drlTanhMakespan3)
drlTanhMakespanList.append(drlTanhMakespan4)
drlTanhMakespanList.append(drlTanhMakespan5)
drlTanhMakespanList.append(drlTanhMakespan6)
drlTanhMakespanList.append(drlTanhMakespan7)
drlTanhMakespanList.append(drlTanhMakespan8)
drlTanhMakespanList.append(drlTanhMakespan9)
drlTanhMakespanList.append(drlTanhMakespan10)
drlTanhMakespanList.append(drlTanhMakespan11)
drlTanhMakespanList.append(drlTanhMakespan12)
drlTanhMakespanList.append(drlTanhMakespan13)
drlTanhMakespanList.append(drlTanhMakespan14)
drlTanhMakespanList.append(drlTanhMakespan15)
drlTanhMakespanList.append(drlTanhMakespan16)
drlTanhMakespanList.append(drlTanhMakespan17)
drlTanhMakespanList.append(drlTanhMakespan18)
drlTanhMakespanList.append(drlTanhMakespan19)
drlTanhMakespanList.append(drlTanhMakespan20)
drlTanhMakespanList.append(drlTanhMakespan21)
drlTanhMakespanList.append(drlTanhMakespan22)
drlTanhMakespanList.append(drlTanhMakespan23)
drlTanhMakespanList.append(drlTanhMakespan24)
drlTanhMakespanList.append(drlTanhMakespan25)
drlTanhMakespanList.append(drlTanhMakespan26)
drlTanhMakespanList.append(drlTanhMakespan27)
drlTanhMakespanList.append(drlTanhMakespan28)
drlTanhMakespanList.append(drlTanhMakespan29)
drlTanhMakespanList.append(drlTanhMakespan30)
drlTanhMakespanList.append(drlTanhMakespan31)
drlTanhMakespanList.append(drlTanhMakespan32)
drlTanhMakespanList.append(drlTanhMakespan33)
drlTanhMakespanList.append(drlTanhMakespan34)
drlTanhMakespanList.append(drlTanhMakespan35)
drlTanhMakespanList.append(drlTanhMakespan36)
drlTanhMakespanList.append(drlTanhMakespan37)
drlTanhMakespanList.append(drlTanhMakespan38)
drlTanhMakespanList.append(drlTanhMakespan39)
drlTanhMakespanList.append(drlTanhMakespan40)
drlTanhMakespanList.append(drlTanhMakespan41)
drlTanhMakespanList.append(drlTanhMakespan42)
drlTanhMakespanList.append(drlTanhMakespan43)
drlTanhMakespanList.append(drlTanhMakespan44)
drlTanhMakespanList.append(drlTanhMakespan45)
drlTanhMakespanList.append(drlTanhMakespan46)
drlTanhMakespanList.append(drlTanhMakespan47)
drlTanhMakespanList.append(drlTanhMakespan48)
drlTanhMakespanList.append(drlTanhMakespan49)
drlTanhRewardsList.append(drlTanhRewards0)
drlTanhRewardsList.append(drlTanhRewards1)
drlTanhRewardsList.append(drlTanhRewards2)
drlTanhRewardsList.append(drlTanhRewards3)
drlTanhRewardsList.append(drlTanhRewards4)
drlTanhRewardsList.append(drlTanhRewards5)
drlTanhRewardsList.append(drlTanhRewards6)
drlTanhRewardsList.append(drlTanhRewards7)
drlTanhRewardsList.append(drlTanhRewards8)
drlTanhRewardsList.append(drlTanhRewards9)
drlTanhRewardsList.append(drlTanhRewards10)
drlTanhRewardsList.append(drlTanhRewards11)
drlTanhRewardsList.append(drlTanhRewards12)
drlTanhRewardsList.append(drlTanhRewards13)
drlTanhRewardsList.append(drlTanhRewards14)
drlTanhRewardsList.append(drlTanhRewards15)
drlTanhRewardsList.append(drlTanhRewards16)
drlTanhRewardsList.append(drlTanhRewards17)
drlTanhRewardsList.append(drlTanhRewards18)
drlTanhRewardsList.append(drlTanhRewards19)
drlTanhRewardsList.append(drlTanhRewards20)
drlTanhRewardsList.append(drlTanhRewards21)
drlTanhRewardsList.append(drlTanhRewards22)
drlTanhRewardsList.append(drlTanhRewards23)
drlTanhRewardsList.append(drlTanhRewards24)
drlTanhRewardsList.append(drlTanhRewards25)
drlTanhRewardsList.append(drlTanhRewards26)
drlTanhRewardsList.append(drlTanhRewards27)
drlTanhRewardsList.append(drlTanhRewards28)
drlTanhRewardsList.append(drlTanhRewards29)
drlTanhRewardsList.append(drlTanhRewards30)
drlTanhRewardsList.append(drlTanhRewards31)
drlTanhRewardsList.append(drlTanhRewards32)
drlTanhRewardsList.append(drlTanhRewards33)
drlTanhRewardsList.append(drlTanhRewards34)
drlTanhRewardsList.append(drlTanhRewards35)
drlTanhRewardsList.append(drlTanhRewards36)
drlTanhRewardsList.append(drlTanhRewards37)
drlTanhRewardsList.append(drlTanhRewards38)
drlTanhRewardsList.append(drlTanhRewards39)
drlTanhRewardsList.append(drlTanhRewards40)
drlTanhRewardsList.append(drlTanhRewards41)
drlTanhRewardsList.append(drlTanhRewards42)
drlTanhRewardsList.append(drlTanhRewards43)
drlTanhRewardsList.append(drlTanhRewards44)
drlTanhRewardsList.append(drlTanhRewards45)
drlTanhRewardsList.append(drlTanhRewards46)
drlTanhRewardsList.append(drlTanhRewards47)
drlTanhRewardsList.append(drlTanhRewards48)
drlTanhRewardsList.append(drlTanhRewards49)
for vector in drlTanhMakespanList:
for element in vector:
drlTanhMakespanValues.append(element)
for vector in drlTanhRewardsList:
for element in vector:
drlTanhRewardsValues.append(element)
smoothdrlTanhMakespanValues = pd.Series(drlTanhMakespanValues).rolling(12).mean()
plt.plot(smoothdrlTanhMakespanValues)
plt.xlabel("Episodios")
plt.ylabel("Segundos")
plt.title("'Makespan' utilizando feedforward con tanh")
plt.show()
smoothdrlTanhRewardsValues = pd.Series(drlTanhRewardsValues).rolling(12).mean()
plt.plot(smoothdrlTanhRewardsValues)
plt.xlabel("Episodios")
plt.ylabel("Premio")
plt.title("'Reward' utilizando feedforward con tanh")
plt.show()
####################
drlReluMakespan = []
drlReluRewards = []
drlReluMakespanList = []
drlReluRewardsList = []
drlReluMakespanValues = []
drlReluRewardsValues = []
drlReluMakespan.append(np.mean(drlReluMakespan0))
drlReluMakespan.append(np.mean(drlReluMakespan1))
drlReluMakespan.append(np.mean(drlReluMakespan2))
drlReluMakespan.append(np.mean(drlReluMakespan3))
drlReluMakespan.append(np.mean(drlReluMakespan4))
drlReluMakespan.append(np.mean(drlReluMakespan5))
drlReluMakespan.append(np.mean(drlReluMakespan6))
drlReluMakespan.append(np.mean(drlReluMakespan7))
drlReluMakespan.append(np.mean(drlReluMakespan8))
drlReluMakespan.append(np.mean(drlReluMakespan9))
drlReluMakespan.append(np.mean(drlReluMakespan10))
drlReluMakespan.append(np.mean(drlReluMakespan11))
drlReluMakespan.append(np.mean(drlReluMakespan12))
drlReluMakespan.append(np.mean(drlReluMakespan13))
drlReluMakespan.append(np.mean(drlReluMakespan14))
drlReluMakespan.append(np.mean(drlReluMakespan15))
drlReluMakespan.append(np.mean(drlReluMakespan16))
drlReluMakespan.append(np.mean(drlReluMakespan17))
drlReluMakespan.append(np.mean(drlReluMakespan18))
drlReluMakespan.append(np.mean(drlReluMakespan19))
drlReluMakespan.append(np.mean(drlReluMakespan20))
drlReluMakespan.append(np.mean(drlReluMakespan21))
drlReluMakespan.append(np.mean(drlReluMakespan22))
drlReluMakespan.append(np.mean(drlReluMakespan23))
drlReluMakespan.append(np.mean(drlReluMakespan24))
drlReluMakespan.append(np.mean(drlReluMakespan25))
drlReluMakespan.append(np.mean(drlReluMakespan26))
drlReluMakespan.append(np.mean(drlReluMakespan27))
drlReluMakespan.append(np.mean(drlReluMakespan28))
drlReluMakespan.append(np.mean(drlReluMakespan29))
drlReluMakespan.append(np.mean(drlReluMakespan30))
drlReluMakespan.append(np.mean(drlReluMakespan31))
drlReluMakespan.append(np.mean(drlReluMakespan32))
drlReluMakespan.append(np.mean(drlReluMakespan33))
drlReluMakespan.append(np.mean(drlReluMakespan34))
drlReluMakespan.append(np.mean(drlReluMakespan35))
drlReluMakespan.append(np.mean(drlReluMakespan36))
drlReluMakespan.append(np.mean(drlReluMakespan37))
drlReluMakespan.append(np.mean(drlReluMakespan38))
drlReluMakespan.append(np.mean(drlReluMakespan39))
drlReluMakespan.append(np.mean(drlReluMakespan40))
drlReluMakespan.append(np.mean(drlReluMakespan41))
drlReluMakespan.append(np.mean(drlReluMakespan42))
drlReluMakespan.append(np.mean(drlReluMakespan43))
drlReluMakespan.append(np.mean(drlReluMakespan44))
drlReluMakespan.append(np.mean(drlReluMakespan45))
drlReluMakespan.append(np.mean(drlReluMakespan46))
drlReluMakespan.append(
|
np.mean(drlReluMakespan47)
|
numpy.mean
|
# pylint: disable=C0413, E1133
from typing import Literal
import resource
import sys
from pathlib import Path
import multiprocessing
from unittest.mock import AsyncMockMixin
import warnings
from joblib import Parallel, delayed
import numpy as np
import numba
from numba import prange
import torch
from scipy.cluster.vq import kmeans2
from sklearn.cluster import KMeans, BisectingKMeans
from sklearn import tree
from sklearn.tree import _tree
from sklearn.model_selection import cross_val_score
from halutmatmul.functions import create_codebook_start_end_idxs
sys.path.append(
str(Path(__file__).parent) + "/../../../maddness/python/"
) # for maddness import
from maddness.util.least_squares import ( # type: ignore[attr-defined]
encoded_lstsq,
_XW_encoded,
)
class DecisionTreeOffset:
DIMS = 0
THRESHOLDS = 1
CLASSES = 2
TOTAL = 3
DEFAULT_NEG_VALUE = -4419
@numba.jit(nopython=True, parallel=False)
def apply_hash_function_decision_tree(
X: np.ndarray, decision_tree: np.ndarray
) -> np.ndarray:
N, _ = X.shape
group_ids = np.zeros(N, dtype=np.int64) # needs to be int64 because of index :-)
B = decision_tree.shape[0] // 3
n_decisions = int(np.log2(B))
for depth in range(n_decisions):
index_offet = 2**depth - 1
split_thresholds = decision_tree[group_ids + B + index_offet]
dims = decision_tree[group_ids + index_offet].astype(np.int64)
# x = X[np.arange(N), dims]
# make it numba compatible
x = np.zeros(group_ids.shape[0], np.float32)
for i in range(x.shape[0]):
x[i] = X[i, dims[i]]
indicators = x > split_thresholds
group_ids = (group_ids * 2) + indicators
group_ids = decision_tree[group_ids + 2 * B].astype(np.int32)
return group_ids
@numba.jit(nopython=True, parallel=True)
def halut_encode_decision_tree(X: np.ndarray, numpy_array: np.ndarray) -> np.ndarray:
N, _ = X.shape
C = numpy_array.shape[0]
A_enc = np.empty((C, N), dtype=np.int32) # column-major
for c in prange(C):
A_enc[c] = apply_hash_function_decision_tree(X, numpy_array[c])
return np.ascontiguousarray(A_enc.T)
def apply_hash_function_pq(X: np.ndarray, prototypes: np.ndarray) -> np.ndarray:
group_ids = np.argsort(
np.array([np.linalg.norm(X - x, axis=1) for x in prototypes]).T, axis=1
)[:, :1].flatten()
return group_ids
def apply_hash_function_pq_tensor(
X: torch.Tensor, prototypes: torch.Tensor
) -> torch.Tensor:
group_ids = torch.argsort(
torch.stack([torch.linalg.norm(X - x, axis=1) for x in prototypes]).T, dim=1
)[:, :1].flatten()
return group_ids
def halut_encode_pq(X: np.ndarray, prototypes: np.ndarray) -> np.ndarray:
N, _ = X.shape
C = prototypes.shape[0]
A_enc = np.empty((C, N), dtype=np.int32) # column-major
pq_idxs = create_codebook_start_end_idxs(X.shape[1], C, algo="start")
for c in prange(C):
start_idx, end_idx = pq_idxs[c]
idxs = np.arange(start_idx, end_idx)
X_cut = X[:, idxs]
A_enc[c] = apply_hash_function_pq(X_cut, prototypes[c][:, idxs])
return np.ascontiguousarray(A_enc.T)
def halut_encode_pq_tensor(X: torch.Tensor, prototypes: torch.Tensor) -> torch.Tensor:
N, _ = X.shape
C = prototypes.shape[0]
K = prototypes.shape[1]
A_enc = torch.empty((C, N), dtype=torch.int32, device=str(X.device)) # column-major
pq_idxs = create_codebook_start_end_idxs(X.shape[1], C, algo="start")
for c in prange(C):
start_idx, end_idx = pq_idxs[c]
idxs = torch.arange(start_idx, end_idx, device=str(X.device))
X_cut = X[:, idxs]
A_enc[c] = apply_hash_function_pq_tensor(X_cut, prototypes[c][:, idxs])
offsets = torch.arange(C, dtype=torch.int32, device=str(X.device)) * K
return torch.Tensor.contiguous(A_enc.T) + offsets
def tree_to_numpy(
decision_tree: tree.DecisionTreeClassifier, depth: int = 4
) -> np.ndarray:
tree_ = decision_tree.tree_
class_names = decision_tree.classes_
B = 2**depth
total_length = B * DecisionTreeOffset.TOTAL
numpy_array = np.ones(total_length, np.float32) * DEFAULT_NEG_VALUE
def _add_leaf(value: int, class_name: int, depth: int, tree_id: int) -> None:
if tree_id >= B:
numpy_array[tree_id - B + DecisionTreeOffset.CLASSES * B] = class_name
else:
_add_leaf(value, class_name, depth + 1, 2 * tree_id)
_add_leaf(value, class_name, depth + 1, 2 * tree_id + 1)
def recurse_tree(node: int, depth: int, tree_id: int) -> None:
value = None
if tree_.n_outputs == 1:
value = tree_.value[node][0]
else:
value = tree_.value[node].T[0]
class_name = np.argmax(value)
if tree_.n_classes[0] != 1 and tree_.n_outputs == 1:
class_name = class_names[class_name]
# pylint: disable=c-extension-no-member
if tree_.feature[node] != _tree.TREE_UNDEFINED:
dim = tree_.feature[node]
threshold = tree_.threshold[node]
numpy_array[tree_id - 1] = dim
numpy_array[tree_id - 1 + DecisionTreeOffset.THRESHOLDS * B] = threshold
recurse_tree(tree_.children_left[node], depth + 1, 2 * tree_id)
recurse_tree(tree_.children_right[node], depth + 1, 2 * tree_id + 1)
else:
_add_leaf(value, class_name, depth, tree_id) # type: ignore[arg-type]
recurse_tree(0, 1, 1)
for i in range(B):
assert numpy_array[DecisionTreeOffset.CLASSES * B + i] != DEFAULT_NEG_VALUE
if numpy_array[i] == DEFAULT_NEG_VALUE:
numpy_array[i] = 0 # adding default dimension TODO: optimize
return numpy_array
def learn_decision_tree(
X: np.ndarray, K: int = 16, depth: int = 4, iterations: int = 25
) -> tuple[np.ndarray, np.ndarray]:
X = X.copy().astype(np.float32)
decision_tree_args = {
"min_samples_split": 2,
"max_depth": depth,
"min_samples_leaf": 20,
"max_leaf_nodes": 2**depth,
"splitter": "best",
# "criterion": "log_loss",
# "class_weight": "balanced",
}
centroids_list = []
assignments_list = []
scores = []
warnings.filterwarnings(
"ignore", category=UserWarning
) # ignores empty cluster warning for kmeans
# pylint: disable=import-outside-toplevel
from timeit import default_timer as timer
for _ in range(iterations):
start = timer()
centroids_, assignments_ = kmeans2(X, K, minit="points", iter=5)
end = timer()
print(f"kmeans time {end - start}")
# kmeans = KMeans(n_clusters=K, n_init=1).fit(X)
# kmeans = BisectingKMeans(n_clusters=K, n_init=1).fit(X)
# centroids_, assignments_ = kmeans.cluster_centers_, kmeans.labels_
clf_ = tree.DecisionTreeClassifier(**decision_tree_args)
start = timer()
score_ = cross_val_score(clf_, X, assignments_, cv=2, n_jobs=2)
end = timer()
print(f"cross_val_score time {end - start}", score_)
centroids_list.append(centroids_)
assignments_list.append(assignments_)
scores.append(np.mean(score_))
best_score = np.argsort(scores)[::-1]
centroids = centroids_list[best_score[0]]
assignments = assignments_list[best_score[0]]
clf = tree.DecisionTreeClassifier(**decision_tree_args)
clf = clf.fit(X, assignments)
# additional Infos
PRINT_DEBUG = False
numpy_array = tree_to_numpy(clf, depth=depth)
prediction = clf.predict(X)
bincount_pred = np.bincount(prediction)
if PRINT_DEBUG:
r = tree.export_text(clf)
print(r)
hist = np.bincount(assignments)
print(hist)
print(bincount_pred)
l2_error = np.mean(np.sqrt((centroids[prediction] - X) ** 2))
l1_error = np.mean((centroids[prediction] - X))
score = cross_val_score(clf, X, assignments, cv=5)
print("L2 error: ", l2_error)
print("L1 error: ", l1_error)
# Rebase
for i in range(bincount_pred.shape[0]):
if bincount_pred[i] > 0:
prediction_where = prediction == i
select_rows = X[prediction_where]
new_centroid = np.mean(select_rows, axis=0)
centroids[i] = new_centroid
if PRINT_DEBUG:
l2_error = np.mean(np.sqrt((centroids[prediction] - X) ** 2))
l1_error = np.mean((centroids[prediction] - X))
score = cross_val_score(clf, X, assignments, cv=5)
scores_2 = clf.score(X, assignments)
print("L2 error after: ", l2_error)
print("L1 error after: ", l1_error)
print("Prediction score: ", scores_2, score)
return centroids, numpy_array
def decision_tree_per_codebook(
c: int, pq_idxs: np.ndarray, X: np.ndarray, K: int, depth: int, C: int, D: int
) -> tuple[np.ndarray, np.ndarray]:
start_idx, end_idx = pq_idxs[c]
idxs = np.arange(start_idx, end_idx)
X_cut = X[:, idxs]
centroids, tree = learn_decision_tree(X_cut, K=K, depth=depth, iterations=5)
for i in range(K):
tree[i] = idxs[int(tree[i])]
centroids_extended = np.zeros((K, D), np.float32)
centroids_extended[:, idxs] = centroids
ram_usage = resource.getrusage(resource.RUSAGE_SELF).ru_maxrss
print(
f"Learning progress {X.shape}-{C}-{K}: {c + 1}/{C} "
f"({(ram_usage / (1024 * 1024)):.3f} GB)"
)
return tree, centroids_extended
def init_and_learn_hash_function_decision_tree(
X: np.ndarray,
C: int,
pq_perm_algo: Literal["start", "end"] = "start",
K: int = 16,
depth: int = 4,
) -> tuple[np.ndarray, np.ndarray]:
D = X.shape[1]
depth = int(np.ceil(np.log2(K)))
B = 2**depth
X = X.astype(np.float32)
all_prototypes =
|
np.zeros((C, K, D), dtype=np.float32)
|
numpy.zeros
|
# Adapted for numpy/ma/cdms2 by convertcdms.py
import MV2
import cdms2
import genutil
import unidata
import vcs
import numpy
from vcs import VCS_validation_functions
thermo_objects = []
def Es(T, method=None):
"""Computes saturated pressure in Pa given T in K, using the method:
1: Hyland-Wexler formulation, polynomial coeff (absolute norm)
2: Wexler formulation
3: Hyland-Wexler formulation, polynomial coeff (relative error norm)
4: classic Goff Gratch equation
5: 6.112*numpy.ma.exp(17.67*tempc/(tempc+243.5))
Default is method 1
Note: 1 and 2 use method 3 where T is not : 173.15 < T < 473.15
ref for 1, 2 and 3:
<NAME>., Journal of Applied Met., Vol 31, Dec 1992
( http://ams.allenpress.com/perlserv/?request=get-document&\
doi=10.1175%2F1520-0450(1992)031%3C1507%3APFTSVP%3E2.0.CO%3B2&ct=1 )
"""
if method is None:
method = 1
if method == 1:
# Put in C
x = T - 273.15
# Water vapor
c0 = 0.611220713E03
c1 = 0.443944344E02
c2 = 0.143195336E01
c3 = 0.263350515E-01
c4 = 0.310636053E-03
c5 = 0.185218710E-05
c6 = 0.103440324E-07
c7 = -0.468258100E-10
c8 = 0.466533033E-13
eswat = c0 + x * (c1 + x * (c2 + x * (c3 + x * (c4 + x * (c5 + x * (c6 +
x * (c7 + x * c8)))))))
# ice
c0 = .611153246E03
c1 = .503261230E02
c2 = .188595709E01
c3 = .422115970E-01
c4 = .620376691E-03
c5 = .616082536E-05
c6 = .405172828E-07
c7 = .161492905E-09
c8 = .297886454E-12
esice = c0 + x * (c1 + x * (c2 + x * (c3 + x * (c4 + x * (c5 + x * (c6 +
x * (c7 + x * c8)))))))
# Combine
es = MV2.where(MV2.less(T, 273.15), esice, eswat)
# Overwrite values outside valid range with method 2
mn, mx = genutil.minmax(T)
if mn < 173.16 or mx > 473.15:
es2 = Es(T, method=2)
es = MV2.where(MV2.less(T, 173.16), es2, es)
es = MV2.where(MV2.greater(T, 473.15), es2, es)
elif method == 2:
# over water
g0 = -0.29912729E4
g1 = -0.60170128E4
g2 = 0.1887643854E2
g3 = -0.28354721E-1
g4 = 0.17838301E-4
g5 = -0.84150417E-9
g6 = 0.44412543E-12
g7 = 0.2858487E1
# over ice
k0 = -0.58653696e4
k1 = 0.2224103300E2
k2 = 0.13749042E-1
k3 = -0.34031775E-4
k4 = 0.26967687E-7
k5 = 0.6918651
esice = (k0 + (k1 + k5 * MV2.log(T) + (k2 + (
k3 + k4 * T) * T) * T) * T) / T # over ice
eswat = (g0 + (g1 + (g2 + g7 * MV2.log(T) + (g3 + (g4 + (
g5 + g6 * T) * T) * T) * T) * T) * T) / T ** 2 # over water
es = MV2.where(MV2.less(T, 273.15), esice, eswat)
es = MV2.exp(es)
elif method == 3:
# Put in C
x = T - 273.15
# Water vapor
c0 = 0.611213476E03
c1 = 0.444007856E02
c2 = 0.143064234E01
c3 = 0.264461437E-01
c4 = 0.305930558E-03
c5 = 0.196237241E-05
c6 = 0.892344772E-08
c7 = -0.373208410E-10
c8 = 0.209339997E-13
eswat = c0 + x * (c1 + x * (c2 + x * (c3 + x * (c4 + x * (
c5 + x * (c6 + x * (c7 + x * c8)))))))
# ice
c0 = .611123516E03
c1 = .503109514E02
c2 = .1888369801E01
c3 = .420547422E-01
c4 = .614396778E-03
c5 = .602780717E-05
c6 = .387940929E-07
c7 = .149436277E-09
c8 = .262655803E-12
esice = c0 + x * (c1 + x * (c2 + x * (c3 + x * (c4 + x * (
c5 + x * (c6 + x * (c7 + x * c8)))))))
# Combine
es = MV2.where(MV2.less(T, 273.15), esice, eswat)
# Overwrite values outside valid range with method 2
mn, mx = genutil.minmax(T)
if mn < 173.16 or mx > 473.15:
es2 = Es(T, method=2)
es = MV2.where(MV2.less(T, 173.16), es2, es)
es = MV2.where(MV2.greater(T, 473.15), es2, es)
elif method == 4:
est = 101324.6 # Pa
Ts = 373.16 / T
a = -7.90298
b = 5.02808
c = -1.3816E-7
d = 11.344
f = 8.1328E-3
h = -3.49149
maxexp = int(numpy.log10(numpy.finfo(numpy.float).max))
minexp = 1 - a
es = a * (Ts - 1.)
es = es + b *
|
numpy.ma.log10(Ts)
|
numpy.ma.log10
|
from mpi4py import MPI
import numpy as np
import sys
def matrix(value, M, N):
matrix = np.zeros((M, N), np.int64)
return matrix
comm = MPI.COMM_SELF.Spawn(sys.executable, args=['agent.py'], maxprocs=2)
rank = comm.Get_rank()
size = comm.Get_size()
A = [(2,2), (0,4), (2,6), (4,5), (4,7), (5,7)]
B = [(1,1), (0,6), (4,4), (4,5), (5,5)]
M = len(A)
N = len(B)
m_n = np.array((M,N))
print("About to send matrix dimensions (M x N): " + str(m_n))
comm.Bcast(m_n, root = MPI.ROOT)
# Manager Process initializes matrices.
cost_matrix = matrix(0, M, N) # Initialize with zeros
for i in range(M):
for j in range(N):
cost_matrix[i,j] = (A[i][0] - B[j][0])**2 + (A[i][1] - B[j][1])**2 # distance function
print(cost_matrix)
dtw_matrix = matrix(0, M, N) # Initialize with zeros
dtw_matrix[0, 0] = cost_matrix[0, 0] # Initialize top left cell
neighbor = dtw_matrix[0, 0]
# COLUMN
for i in range(1, M):
dtw_matrix[i, 0] = neighbor + cost_matrix[i, 0]
neighbor = dtw_matrix[i, 0]
# ROW
neighbor = dtw_matrix[0, 0]
for i in range(1, N):
dtw_matrix[0, i] = neighbor + cost_matrix[0, i]
neighbor = dtw_matrix[0, i]
print(dtw_matrix)
send_row, send_column = True, True
req_for_row, req_for_col = [], []
row_counter, col_counter = 1, 1
row_limit, col_limit = M, N
complete = False
req_for_agent0, req_for_agent1 = None, None
while not complete:
if send_row == True:
print(f"sending row {row_counter}")
# Stuff that we need to send to the row process
previous_row = np.array(dtw_matrix[row_counter - 1, col_counter - 1:], dtype=np.int64)
cost_row = np.array(cost_matrix[row_counter, col_counter:], dtype=np.int64)
left_neighbor = np.array(dtw_matrix[row_counter, col_counter - 1], dtype=np.int64)
req_for_row.append(comm.Isend(np.array(len(previous_row), dtype=np.int64), dest=0, tag=1))
req_for_row.append(comm.Isend(previous_row, dest=0, tag=2))
req_for_row.append(comm.Isend(left_neighbor, dest=0, tag=3))
new_row = np.zeros(len(previous_row) - 1, dtype = np.int64)
req_for_row.append(comm.Isend(cost_row, dest=0, tag=4))
send_row = False
if send_column == True:
print(f"sending col {col_counter}")
previous_col = np.array(dtw_matrix[row_counter - 1:, col_counter - 1], dtype=np.int64)
cost_col =
|
np.array(cost_matrix[row_counter:, col_counter], dtype=np.int64)
|
numpy.array
|
import csv
import numpy as np
import matplotlib.cm as cm
import matplotlib.pyplot as plt
import seaborn as sns
from scipy import special as sp
from sklearn.metrics import confusion_matrix
from sklearn.tree import DecisionTreeClassifier
def _qfunc(x):
return 0.5-0.5*sp.erf(x/np.sqrt(2))
def theoretical_ser(M, SNR_db):
""" Calculate the theoretical probability of error for SQ M-QAM.
"""
SNR_l = 10**(SNR_db/10)
Pe = 4*(1-(1/np.sqrt(M)))*_qfunc(np.sqrt(3*SNR_l/(M-1))) \
- 4*(1-(1/np.sqrt(M)))**2 * _qfunc(np.sqrt(3*SNR_l/(M-1)))**2
return Pe
def ser(clf, X, y):
""" Calculate the misclassification rate, which
coincides with the symbol error rate (SER) for QAM transmission.
"""
y_pred = clf.predict(X)
ser = np.sum(y != y_pred)/len(y)
return ser
def plot_confusion_matrix(clf, X, y, num_classes):
""" Plot the confusion matrix
"""
y_pred = clf.predict(X)
conf_mtx = confusion_matrix(y, y_pred)
plt.figure(figsize=(10,6))
sns.heatmap(conf_mtx, cmap=sns.cm.rocket_r, square=True, linewidths=0.1,
annot=True, fmt='d', annot_kws={"fontsize": 8})
plt.tick_params(axis='both', which='major', labelsize=10,
bottom=False, top=False, left=False,
labelbottom=False, labeltop=True)
plt.yticks(rotation=0)
plt.show()
def plot_decision_boundary(classifier, X, y, legend=False, plot_training=True):
""" Plot the classifier decision regions
"""
num_classes = int(np.max(y))+1 #e.g. 16 for QAM-16
axes = [np.min(X[:,0]),
|
np.max(X[:,0])
|
numpy.max
|
# -*- coding: utf-8 -*-
import tensorflow as tf
import numpy as np
from cvxopt import matrix
from cvxopt.solvers import qp
from cvxopt import solvers
import matplotlib.pyplot as plt
tf.reset_default_graph()
np.random.seed(10)
rand_x = np.random.randn(1500)/50
np.random.seed(8)
rand_y = np.random.randn(1500)/50
# rand_y = np.random.randn(1500)/50
solvers.options['show_progress'] = False
other = [(1.23, 3.01),(0.98, 3.32),(1.77, 3.92),(1.48, 4.52),(0.63, 2.89), (1.92, 5.0), (1.1, 2.8),(0.71, 3.17),
(1.64, 4.54),(1.26, 3.96),(1.22, 2.84), (0.77, 2.59),(1.89, 5.1),(1.13,3.17), (1.31, 2.91)]
u2 = np.zeros((1515,1))
v2 = np.zeros((1515,1))
for i in range(500):
u2[i],v2[i] = 0.16+rand_x[i], 1.22+rand_y[i]
for i in range(500):
u2[i+500],v2[i+500] = 0.43+rand_x[i+500],1.45+rand_y[i+500]
for i in range(500):
u2[i+1000],v2[i+1000] = 0.04+rand_x[i+1000],1.59+rand_y[i+1000]
for i in range(15):
u2[i+1500],v2[i+1500] = other[i][0],other[i][1]
# Separate dataset into two subgroups.
X1 = tf.constant(u2[:1500])
y1 = tf.constant(v2[:1500])
X2 = tf.constant(u2[1500:])
y2 = tf.constant(v2[1500:])
w = tf.get_variable("w", shape=[1, 1], initializer=tf.contrib.layers.xavier_initializer(),dtype='float64')
b = tf.Variable(tf.constant(0.1, shape=[1],dtype='float64'))
z1 = tf.reduce_mean(tf.square(tf.matmul(X1,w)+b - y1))
z2 = tf.reduce_mean(tf.square(tf.matmul(X2,w)+b - y2))
# Define the max_mean of each subgroup's loss
# according to equation (1).
z = tf.maximum(z1,z2)
z1_grad = tf.gradients(ys=z1,xs=w)
z2_grad = tf.gradients(ys=z2,xs=w)
z1_grad_b = tf.gradients(ys=z1,xs=b)
z2_grad_b = tf.gradients(ys=z2,xs=b)
# MER = []
# MSE = []
sess = tf.Session()
sess.run(tf.global_variables_initializer())
print('start...')
for i in range(300):
# Compute the gradient of 'w'.
GG = np.zeros([2,1])
hh = np.zeros(2)
g1 = sess.run(z1_grad)
g2 = sess.run(z2_grad)
GG[0,:] = g1[0].reshape(-1)
GG[1,:] = g2[0].reshape(-1)
hh[0],hh[1] = sess.run([z1,z2])
P = matrix(GG)*matrix(GG).T
q = -matrix(hh)
G = matrix(-
|
np.eye(2)
|
numpy.eye
|
import numpy as np
class AxisAngle(object):
def __init__(self,angle,axis):
if type(axis) == str:
if axis=="x":
axis = np.array([1,0,0])
elif axis=="y":
axis = np.array([0,1,0])
elif axis=="z":
axis =
|
np.array([0,0,1])
|
numpy.array
|
import numpy as np
from gmmmc.gmm import GMM
from gmmmc.proposals.proposals import Proposal
import pdb
import logging
class GaussianStepMeansProposal(Proposal):
"""Gaussian Proposal distribution for means of a GMM"""
def __init__(self, step_sizes=(0.001,)):
"""
Gaussian proposal distribution for the means. The multivariate Gaussian is centered at the means of the current
state in the Markov Chain and has covariance given by step_sizes. Multiple step sizes can be specified.
The proposal algorithm will take these steps in the sequence specified in step_sizes.
Parameters
----------
step_sizes : 1-D array_like
Iterable containing the sequence of step sizes (covariances of the Gaussian proposal distribution"
"""
super(GaussianStepMeansProposal, self).__init__()
self.step_sizes = step_sizes
self.count_accepted = np.zeros((len(step_sizes),))
self.count_illegal = np.zeros((len(step_sizes),))
self.count_proposed = np.zeros((len(step_sizes),))
def propose(self, X, gmm, target, n_jobs=1):
"""
Propose a new set of GMM means.
Parameters
----------
X : 2-D array like of shape (n_samples, n_features)
The observed data or evidence.
gmm : GMM object
The current state (set of gmm parameters) in the Markov Chain
target : GMMPosteriorTarget object
The target distribution to be found.
n_jobs : int
Number of cpu cores to use in the calculation of log probabilities.
Returns
-------
: GMM
A new GMM object initialised with new mean parameters.
"""
new_means = np.array(gmm.means)
beta = target.beta
prior = target.prior
steps = [np.random.multivariate_normal(np.zeros(gmm.n_features),
step_size * np.eye(gmm.n_features),
size=gmm.n_mixtures)
for step_size in self.step_sizes]
# calculation of prior probabilities of only the means, since only means will change
log_priors = np.array([prior.means_prior.log_prob_single(gmm.means[mixture], mixture) for mixture in xrange(gmm.n_mixtures)])
log_prob_priors = np.sum(log_priors)
previous_prob = beta * gmm.log_likelihood(X, n_jobs) + np.sum(log_priors)
for i, step in enumerate(steps):
for mixture in xrange(gmm.n_mixtures):
self.count_proposed[i] += 1
# propose new means
new_mixture_means = gmm.means[mixture] + step[mixture]
# try out the new means
proposed_means = np.array(new_means)
proposed_means[mixture] = new_mixture_means
proposed_gmm = GMM(proposed_means, np.array(gmm.covars), np.array(gmm.weights))
# calculate new prior
new_log_prob_mixture = prior.means_prior.log_prob_single(new_mixture_means, mixture)
new_log_prob_priors = log_prob_priors - log_priors[mixture] + new_log_prob_mixture
# priors
proposed_prob = beta * proposed_gmm.log_likelihood(X, n_jobs) + new_log_prob_priors
# ratio
ratio = proposed_prob - previous_prob
if ratio > 0 or ratio > np.log(np.random.uniform()):
# accept proposal
new_means = proposed_means
previous_prob = proposed_prob
# update prior probability calculation
log_prob_priors = new_log_prob_priors
log_priors[mixture] = new_log_prob_mixture
self.count_accepted[i] += 1
return GMM(new_means, np.array(gmm.covars), np.array(gmm.weights))
class GaussianStepCovarProposal(Proposal):
def __init__(self, step_sizes=(0.001,)):
"""
Gaussian proposal function for the covariances of the GMM.
Parameters
----------
step_sizes : array_like
Array of covariance values for the Gaussian proposal.
"""
super(GaussianStepCovarProposal, self).__init__()
self.step_sizes = step_sizes
self.count_accepted = np.zeros((len(step_sizes),))
self.count_illegal = np.zeros((len(step_sizes),))
self.count_proposed = np.zeros((len(step_sizes),))
def propose(self, X, gmm, target, n_jobs=1):
"""
Propose a new set of GMM covariances (diagonal only).
Parameters
----------
X : 2-D array like of shape (n_samples, n_features)
The observed data or evidence.
gmm : GMM object
The current state (set of gmm parameters) in the Markov Chain
target : GMMPosteriorTarget object
The target distribution to be found.
n_jobs : int
Number of cpu cores to use in the calculation of log probabilities.
Returns
-------
: GMM
A new GMM object initialised with new covariance parameters.
"""
new_covars = np.array(gmm.covars)
beta = target.beta
prior = target.prior
previous_prob = beta * gmm.log_likelihood(X, n_jobs) + prior.log_prob(gmm)
steps = [np.random.multivariate_normal(np.zeros(gmm.n_features),
step_size * np.eye(gmm.n_features),
size=gmm.n_mixtures) for step_size in self.step_sizes]
log_priors = np.array([prior.means_prior.log_prob_single(gmm.means[mixture], mixture) for mixture in xrange(gmm.n_mixtures)])
log_prob_priors = np.sum(log_priors)
for i, step in enumerate(steps):
for mixture in xrange(gmm.n_mixtures):
self.count_proposed[i] += 1
# propose new covars
new_mixture_covars = gmm.covars[mixture] + step[mixture]
if (new_mixture_covars > 0).all(): # check covariances are valid
# try out the new covars
proposed_covars = np.array(new_covars)
proposed_covars[mixture] = new_mixture_covars
proposed_gmm = GMM(np.array(gmm.means), proposed_covars, np.array(gmm.weights))
# calculate desired distribution
new_log_prob_mixture = prior.covars_prior.log_prob_single(new_mixture_covars, mixture)
new_log_prob_priors = log_prob_priors - log_priors[mixture] + new_log_prob_mixture
proposed_prob = beta * proposed_gmm.log_likelihood(X, n_jobs) + new_log_prob_priors
# ratio
ratio = proposed_prob - previous_prob
if ratio > 0 or ratio > np.log(np.random.uniform()):
# accept proposal
new_covars = proposed_covars
previous_prob = proposed_prob
log_prob_priors = new_log_prob_priors
log_priors[mixture] = new_log_prob_mixture
self.count_accepted[i] += 1
else:
self.count_illegal[i] += 1
return GMM(np.array(gmm.means), np.array(new_covars), np.array(gmm.weights))
class GaussianStepWeightsProposal(Proposal):
def __init__(self, n_mixtures, step_sizes=(0.001,), threshold=0.001):
"""
Gaussian proposal function for the weights of a GMM.
Parameters
----------
n_mixtures
step_sizes
Notes
----------
The proposal function works by projecting the weight vector w onto the simplex defined by
w_1 + w_2 + ..... w_n = 1 , 0<=w_i<=1. The change of basis matrix is found by finding n-1 vectors lying on the plane
and using gramm schmidt to get an orthonormal basis. A Gaussian proposal function in (n-1)-d space is
used to find the next point on the simplex.
"""
super(GaussianStepWeightsProposal, self).__init__()
self.step_sizes = step_sizes
self.n_mixtures = n_mixtures
self.count_accepted = np.zeros((len(step_sizes),))
self.count_illegal = np.zeros((len(step_sizes),))
self.count_proposed = np.zeros((len(step_sizes),))
self.threshold = threshold
if n_mixtures > 1:
# get change of basis matrix mapping n dim coodinates to n-1 dim coordinates on simplex
# x1 + x2 + x3 ..... =1
points = np.random.dirichlet([1 for i in xrange(n_mixtures)], size=n_mixtures - 1)
points = points.T
self.plane_origin = np.ones((n_mixtures)) / float(n_mixtures)
# get vectors parallel to plane from its center (1/n,1/n,....)
parallel = points - np.ones(points.shape) / float(n_mixtures)
# do gramm schmidt to get mutually orthonormal vectors (basis)
self.e, _ = np.linalg.qr(parallel)
def transformSimplex(self, weights):
"""
Project weight vector onto the normal simplex.
Parameters
----------
weights : array_like of shape (n_mixtures,)
vector of weights for each gaussian component
Returns
-------
: array_like of shape (n_mixtures-1,)
vector of weights projected onto the simplex plane
"""
# project onto the simplex
return np.dot(self.e.T, weights - self.plane_origin)
def invTransformSimplex(self, simplex_coords):
"""
Transforms a point on the simplex to the original vector space.
Parameters
----------
simplex_coords : array_like of shape (n_mixtures - 1,)
Coordinates of a weight vector on the simplex.
Returns
-------
: array_like of shape(n_mixtures,)
vector of weights.
"""
return self.plane_origin + np.dot(self.e, simplex_coords)
def propose(self, X, gmm, target, n_jobs=1):
"""
Propose a new set of weight vectors.
Parameters
----------
X : 2-D array like of shape (n_samples, n_features)
The observed data or evidence.
gmm : GMM object
The current state (set of gmm parameters) in the Markov Chain
target : GMMPosteriorTarget object
The target distribution to be found.
n_jobs : int
Number of cpu cores to use in the calculation of log probabilities.
Returns
-------
: GMM
A new GMM object initialised with new covariance parameters.
"""
accepted = False
cur_gmm = gmm
if gmm.n_mixtures > 1:
for i, step_size in enumerate(self.step_sizes):
self.count_proposed[i] += 1
current_weights_transformed = self.transformSimplex(cur_gmm.weights)
proposed_weights_transformed = np.random.multivariate_normal(current_weights_transformed,
np.eye(self.n_mixtures - 1) * step_size)
proposed_weights = self.invTransformSimplex(proposed_weights_transformed)
if np.logical_and(0 <= proposed_weights, proposed_weights <= 1).all()\
and np.isclose(np.sum(proposed_weights), 1.0) and (proposed_weights>self.threshold).all():
previous_prob = target.log_prob(X, cur_gmm, n_jobs)
proposed_gmm = GMM(np.array(cur_gmm.means), np.array(cur_gmm.covars), proposed_weights)
proposed_prob = target.log_prob(X, proposed_gmm, n_jobs)
ratio = proposed_prob - previous_prob
if ratio > 0 or ratio > np.log(np.random.uniform()):
# accept proposal
self.count_accepted[i] += 1
accepted = True
cur_gmm = proposed_gmm
else:
self.count_illegal[i] += 1
if accepted is True:
return cur_gmm
else:
return GMM(np.array(gmm.means), np.array(gmm.covars),
|
np.array(gmm.weights)
|
numpy.array
|
from __future__ import division
from future.utils import viewitems
from builtins import int, zip
import concurrent.futures
import os
import itertools
from ._adaptive_threshold import threshold as athreshold
from .pool import pooler
from ._moving_window import moving_window
# from mpglue.raster_tools import create_raster
# from mpglue import moving_window
import numpy as np
import cv2
# SciPy
from scipy.ndimage.measurements import label as nd_label
from scipy.ndimage.measurements import mean as nd_mean
import scipy.stats as sci_stats
from scipy.stats import mode as sci_mode
from sklearn.preprocessing import StandardScaler
# Scikit-image
from skimage.exposure import rescale_intensity
from skimage.filters import threshold_local
from skimage.morphology import remove_small_objects, skeletonize
from skimage.morphology import thin as sk_thin
from skimage.feature import peak_local_max
from skimage.measure import regionprops
from skimage.measure import label as sk_label
import pymorph
from mahotas import thin as mthin
from mahotas.morph import hitmiss as mhitmiss
# from tqdm import tqdm
# from joblib import Parallel, delayed
def local_straightness(arr, kernel_filter, w, sigma_color, sigma_space):
"""
https://ieeexplore-ieee-org.ezproxy.library.uq.edu.au/document/1334256
https://docs.opencv.org/master/d4/d70/tutorial_anisotropic_image_segmentation_by_a_gst.html
Example:
>>> conv_kernels = set_kernel_pairs(methods=['compass'])
>>> kernel_filter = conv_kernels['compass']['kernels']
>>> local_straightness(array, kernel_filter, 3, 1, 1)
"""
diff_x = cv2.filter2D(np.float32(arr),
cv2.CV_32F,
kernel_filter[1],
borderType=cv2.BORDER_CONSTANT)
diff_y = cv2.filter2D(np.float32(arr),
cv2.CV_32F,
kernel_filter[0],
borderType=cv2.BORDER_CONSTANT)
diff_xy = diff_x * diff_y
diff_xx = diff_x * diff_x
diff_yy = diff_y * diff_y
c11 = cv2.boxFilter(np.float32(diff_xx), cv2.CV_32F, (w, w))
c22 = cv2.boxFilter(np.float32(diff_yy), cv2.CV_32F, (w, w))
c12 = cv2.boxFilter(np.float32(diff_xy), cv2.CV_32F, (w, w))
# c11 = cv2.bilateralFilter(np.float32(diff_xx), w, sigma_color, sigma_space)
# c22 = cv2.bilateralFilter(np.float32(diff_yy), w, sigma_color, sigma_space)
# c12 = cv2.bilateralFilter(np.float32(diff_xy), w, sigma_color, sigma_space)
gamma_max = (c11 + c22 + np.sqrt((c11 - c22)**2 + 4*c12**2)) / 2.0
gamma_min = (c11 + c22 - np.sqrt((c11 - c22)**2 + 4*c12**2)) / 2.0
s = 1.0 - (gamma_min / gamma_max)
return s
def logistic(x, **params):
return sci_stats.logistic.cdf(x, **params)
def sigmoid(x, a, b):
return 1.0 / (1.0 + np.exp(-b * (x - a)))
def log_transform(egm, scale=1e-6, logistic_alpha=1.6, logistic_beta=0.5):
"""
Transforms an EGM to probabilities
Args:
egm (2d array)
scale (Optional[float]): The scaling factor
logistic_alpha (Optional[float])
logistic_beta (Optional[float])
Returns:
Probabilities (2d array)
"""
# Mask
egm[egm == 0] = np.nan
log_min = np.nanpercentile(np.log(egm * scale), 2)
egm[np.isnan(egm)] = 0
# Log transform
egm_proba = np.where(egm > 0, np.log(egm * scale), log_min)
# Scale and clip
r, c = egm_proba.shape
zegm = np.where(egm_proba.ravel() > log_min)[0]
scaler = StandardScaler().fit(egm_proba.ravel()[zegm][:, np.newaxis])
egm_proba = scaler.transform(egm_proba.ravel()[:, np.newaxis]).reshape(r, c)
egm_proba = rescale_intensity(egm_proba, in_range=(-3, 3), out_range=(-3, 3))
# CDF
return logistic(egm_proba,
loc=logistic_alpha,
scale=logistic_beta)
def bayes(prior_a, prior_b, likelihood):
"""
Bayes rule
Args:
prior_a (float): The class prior probability.
prior_b (float): The class prior probability.
likelihood (float)
"""
posterior = (likelihood * prior_a) / (likelihood * prior_a + prior_b * (1.0 - prior_a))
posterior[np.isnan(posterior)] = 0
return posterior
class Params(object):
def __init__(self, **kwargs):
for k, v in viewitems(kwargs):
setattr(self, k, v)
def mopen(array2morph, se, iters=1):
return cv2.morphologyEx(np.uint8(array2morph),
cv2.MORPH_OPEN,
se,
iterations=iters)
def mclose(array2morph, se, iters=1):
return cv2.morphologyEx(np.uint8(array2morph),
cv2.MORPH_CLOSE,
se,
iterations=iters)
def merode(array2morph, se, iters=1):
return cv2.morphologyEx(np.uint8(array2morph),
cv2.MORPH_ERODE,
se,
iterations=iters)
def mdilate(array2morph, se, iters=1):
return cv2.morphologyEx(np.uint8(array2morph),
cv2.MORPH_DILATE,
se,
iterations=iters)
def closerec(array2morph, se, r=3, iters=5):
"""
Close by reconstruction
Args:
array2morph (2d array)
se (str)
r (Optional[int])
iters (Optional[int])
"""
if se == 'disk':
se = np.uint8(pymorph.sedisk(r=r))
elif se == 'cross':
se = np.uint8(pymorph.secross(r=r))
evi2_dist = np.float32(cv2.distanceTransform(np.uint8(np.where(array2morph >= 20, 1, 0)), cv2.DIST_L2, 3))
seed = np.uint8(np.where(evi2_dist >= 2,
cv2.morphologyEx(np.uint8(array2morph),
cv2.MORPH_OPEN,
se,
iterations=1),
0))
im_result = seed.copy()
for iter in range(0, iters):
im_dilated = cv2.morphologyEx(np.uint8(im_result),
cv2.MORPH_DILATE,
se,
iterations=1)
im_rec = np.minimum(im_dilated, array2morph)
im_result = im_rec.copy()
if np.allclose(seed, im_rec):
break
return im_result
def openrec(array2morph, se, iters=5):
"""
Open by reconstruction
Args:
array2morph (2d array)
se (2d array)
iters (Optional[int])
"""
evi2_dist = np.float32(cv2.distanceTransform(np.uint8(np.where(array2morph >= 20, 1, 0)), cv2.DIST_L2, 3))
seed = np.uint8(np.where(evi2_dist >= 2,
cv2.morphologyEx(np.uint8(array2morph),
cv2.MORPH_OPEN,
se,
iterations=1),
0))
im_result = seed.copy()
for iter in range(0, iters):
im_dilated = merode(im_result, se, iters=1)
im_rec = np.minimum(im_dilated, array2morph)
im_result = im_rec.copy()
if np.allclose(seed, im_rec):
break
return im_result
def set_kernel_pairs(methods=None):
"""
Creates 2d convolution kernels
Args:
methods (Optional[str list]): Choices are ['compass', 'kirsch', 'prewitt', 'roberts', 'scharr', 'sobel'].
Returns:
List of kernel filters
"""
returned_filters = dict()
if methods:
returned_filters['custom'] = dict(kernels=methods,
compass=True)
methods = ['compass', 'kirsch', 'prewitt', 'roberts', 'sobel']
# Prewitt compass
compass_filters = np.array([[[-1, -1, -1],
[1, -2, 1],
[1, 1, 1]],
[[-1, -1, 1],
[-1, -2, 1],
[1, 1, 1]],
[[-1, 1, 1],
[-1, -2, 1],
[-1, 1, 1]],
[[1, 1, 1],
[-1, -2, 1],
[-1, -1, 1]],
[[1, 1, 1],
[1, -2, 1],
[-1, -1, -1]],
[[1, 1, 1],
[1, -2, -1],
[1, -1, -1]],
[[1, 1, -1],
[1, -2, -1],
[1, 1, -1]]], dtype='float32')
# Sobel
sobel_filters = np.array([[[1, 2, 0],
[2, 0, -2],
[0, -2, -1]],
[[-1, -2, 0],
[-2, 0, 2],
[0, 2, 1]],
[[0, 2, 1],
[-2, 0, 2],
[-1, -2, 0]],
[[0, -2, -1],
[2, 0, -2],
[1, 2, 0]],
[[-1, 0, 1],
[-2, 0, 2],
[-1, 0, 1]],
[[1, 0, -1],
[2, 0, -2],
[1, 0, -1]],
[[-1, -2, -1],
[0, 0, 0],
[1, 2, 1]],
[[1, 2, 1],
[0, 0, 0],
[-1, -2, -1]]], dtype='float32')
# Scharr
scharr_filters = np.array([[[10, 3, 0],
[3, 0, -3],
[0, -3, -10]],
[[-10, -3, 0],
[-3, 0, 3],
[0, 3, 10]],
[[0, 3, 10],
[-3, 0, 3],
[-10, -3, 0]],
[[0, -3, -10],
[3, 0, -3],
[10, 3, 0]],
[[-10, 0, 10],
[-3, 0, 3],
[-10, 0, 10]],
[[10, 0, -10],
[3, 0, -3],
[10, 0, -10]],
[[-10, -3, -10],
[0, 0, 0],
[10, 3, 10]],
[[10, 3, 10],
[0, 0, 0],
[-10, -3, -10]]], dtype='float32')
# Roberts cross
roberts_filters = np.array([[[0, -1],
[1, 0]],
[[0, 1],
[-1, 0]],
[[-1, 0],
[0, 1]],
[[1, 0],
[0, -1]]], dtype='float32')
# Prewitt
prewitt_filters = np.array([[[1, 1, 1],
[0, 0, 0],
[-1, -1, -1]],
[[-1, -1, -1],
[0, 0, 0],
[1, 1, 1]],
[[1, 1, 0],
[1, 0, -1],
[0, -1, -1]],
[[-1, -1, 0],
[-1, 0, 1],
[0, 1, 1]],
[[1, 0, -1],
[1, 0, -1],
[1, 0, -1]],
[[-1, 0, 1],
[-1, 0, 1],
[-1, 0, 1]],
[[0, 1, 1],
[-1, 0, 1],
[-1, -1, 0]],
[[0, -1, -1],
[1, 0, -1],
[1, 1, 0]]], dtype='float32')
# Kirsch compass
kirsch_filters = np.array([[[5, 5, 5],
[-3, 0, -3],
[-3, -3, -3]],
[[5, 5, -3],
[5, 0, -3],
[-3, -3, -3]],
[[5, -3, -3],
[5, 0, -3],
[5, -3, -3]],
[[-3, -3, -3],
[5, 0, -3],
[5, 5, -3]],
[[-3, -3, -3],
[-3, 0, -3],
[5, 5, 5]],
[[-3, -3, -3],
[-3, 0, 5],
[-3, 5, 5]],
[[-3, -3, 5],
[-3, 0, 5],
[-3, -3, 5]]], dtype='float32')
if 'compass' in methods:
returned_filters['compass'] = dict(kernels=compass_filters,
compass=True)
if 'kirsch' in methods:
returned_filters['kirsch'] = dict(kernels=kirsch_filters,
compass=True)
if 'prewitt' in methods:
returned_filters['prewitt'] = dict(kernels=prewitt_filters,
compass=False)
if 'roberts' in methods:
returned_filters['roberts'] = dict(kernels=roberts_filters,
compass=False)
if 'scharr' in methods:
returned_filters['scharr'] = dict(kernels=scharr_filters,
compass=False)
if 'sobel' in methods:
returned_filters['sobel'] = dict(kernels=sobel_filters,
compass=False)
return returned_filters
def find_circles(intensity_array, kernel_size):
"""
Finds circles
Args:
intensity_array (2d array)
kernel_size (int)
"""
kernel_radius = int(kernel_size / 2.0)
kernel_circle = np.uint8(pymorph.sedisk(r=kernel_radius,
dim=2,
metric='euclidean',
flat=True,
h=0) * 1)
kernel_square = np.uint8(pymorph.sebox(r=kernel_radius) * 1)
circles = cv2.filter2D(np.float32(intensity_array),
cv2.CV_32F,
kernel_circle,
borderType=cv2.BORDER_CONSTANT)
squares = cv2.filter2D(np.float32(intensity_array),
cv2.CV_32F,
kernel_square,
borderType=cv2.BORDER_CONSTANT)
diff = circles - squares
local_max_coords = peak_local_max(diff,
min_distance=kernel_size,
indices=True)
local_max = np.zeros(intensity_array.shape, dtype='uint8')
for local_coord in local_max_coords:
local_coord[0] -= kernel_radius
local_coord[1] -= kernel_radius
local_max[local_coord[0]:local_coord[0]+kernel_size,
local_coord[1]:local_coord[1]+kernel_size] = kernel_circle
se = np.array([[0, 1, 0],
[1, 1, 1],
[0, 1, 0]], dtype='uint8')
return cv2.morphologyEx(local_max,
cv2.MORPH_GRADIENT,
se)
def _get_magnitude(image2convolve, kernel_filter):
"""
Calculates the Edge Gradient Magnitude from x and y derivatives
Args:
image2convolve (2d array)
kernel_filter (tuple)
Returns:
EGM as 2d array
"""
return cv2.magnitude(cv2.filter2D(np.float32(image2convolve),
cv2.CV_32F,
kernel_filter[1],
borderType=cv2.BORDER_CONSTANT),
cv2.filter2D(np.float32(image2convolve),
cv2.CV_32F,
kernel_filter[0],
borderType=cv2.BORDER_CONSTANT))
def get_magnitude(im, kernels=None, pad=15):
"""
Gets the Edge Gradient Magnitude (EGM) over multiple edge kernels
Args:
im (2d array)
kernels (Optional[list]
pad (Optional[int])
Returns:
Gradient edge magnitude as 2d array.
[Mean EGM] * [Max EGM]
"""
n_rows, n_cols = im.shape
# Pad image edges.
if pad > 0:
im = np.float32(cv2.copyMakeBorder(im, pad, pad, pad, pad, cv2.BORDER_REFLECT))
# The convolution kernel pairs
conv_kernels = set_kernel_pairs(methods=kernels)
# Mean EGM
# mag_p = np.zeros((len(conv_kernels), im.shape[0], im.shape[1]), dtype='float32')
mag_p = np.zeros(im.shape, dtype='float32')
for kernel_name, kernel_dict in viewitems(conv_kernels):
kernel_filters = kernel_dict['kernels']
mag_c = np.zeros(im.shape, dtype='float32')
if kernel_dict['compass']:
if isinstance(kernel_filters, list):
kiter = len(kernel_filters)
# Get the maximum EGM over all kernel pairs.
for ki in range(0, kiter):
for kw in range(0, 2):
# Image convolution
temp_egm = cv2.filter2D(np.float32(im),
cv2.CV_32F,
np.array(kernel_filters[ki], dtype='float32')[kw],
borderType=cv2.BORDER_CONSTANT)
mag_c = np.maximum(mag_c, temp_egm)
else:
# Get the maximum EGM over all kernels.
for ki in range(0, kernel_filters.shape[0]):
# Image convolution
temp_egm = cv2.filter2D(np.float32(im),
cv2.CV_32F,
kernel_filters[ki],
borderType=cv2.BORDER_CONSTANT)
mag_c = np.maximum(mag_c, temp_egm)
else:
if isinstance(kernel_filters, list):
kiter = len(kernel_filters)
# Get the maximum EGM over all kernel pairs.
for ki in range(0, kiter):
# EGM
temp_egm = _get_magnitude(im, np.array(kernel_filters[ki], dtype='float32'))
mag_c = np.maximum(mag_c, temp_egm)
else:
kiter = kernel_filters.shape[0]
# Get the maximum EGM over all kernel pairs.
for ki in range(0, kiter, 2):
# EGM
temp_egm = _get_magnitude(im, kernel_filters[ki:ki+2])
mag_c = np.maximum(mag_c, temp_egm)
mag_p += mag_c
if pad > 0:
# mag_p = mag_p.mean(axis=0)[pad:n_rows+pad, pad:n_cols+pad] * mag_p.max(axis=0)[pad:n_rows+pad, pad:n_cols+pad]
mag_p = mag_p[pad:n_rows+pad, pad:n_cols+pad] / len(conv_kernels)
else:
# mag_p = mag_p.mean(axis=0) * mag_p.max(axis=0)
mag_p = mag_p / len(conv_kernels)
mag_p[np.isnan(mag_p) | np.isinf(mag_p)] = 0.0
return mag_p
def get_mag_egm(ts_array, ts_r, ts_c, kernels):
# EGM holder
mag_egm = np.zeros((ts_array.shape[0], ts_r, ts_c), dtype='float32')
se = np.array([[0, 1, 0],
[1, 1, 1],
[0, 1, 0]], dtype='uint8')
# count = np.zeros((ts_r, ts_c), dtype='uint8')
# Get the EGM from each day.
for ti in range(0, ts_array.shape[0]):
mask = mdilate(np.where(ts_array[ti] == 0, 1, 0), se, iters=10)
# count[mask == 0] += 1
# Get the EGM over all 'kernels'.
magg_ = get_magnitude(ts_array[ti],
kernels=kernels,
pad=0)
# magg_[mask == 1] = 0
magg_[mask == 1] = np.nan
mag_egm[ti] = magg_
# Get the mean EGM over all layers
# mag_egm_mean = mag_egm.sum(axis=0) / np.float32(count)
mag_egm_mean = np.nanmean(mag_egm, axis=0)
mag_egm_med = np.nanmedian(mag_egm, axis=0)
mag_egm_cv = np.nanstd(mag_egm, axis=0) / mag_egm_med
mag_egm_cv = ((mag_egm_cv + mag_egm_med) / 2.0) * 10000.0
return mag_egm_mean, mag_egm_cv
def get_mag_dist(ts_array, ts_r, ts_c, cvm):
# EGM holder
mag_dist = np.zeros((ts_r, ts_c), dtype='float32')
# Get the edge distance from each day.
for ti in range(0, ts_array.shape[0]-3):
mag_dist_ = moving_window(ts_array[ti:ti+3],
statistic='distance',
window_size=3,
weights=cvm)
mag_dist += mag_dist_
return mag_dist / float(ts_array.shape[0]-3)
def _do_clahe(image2adjust, clip_perc, grid_tile):
"""
Contrast Limited Adaptive Histogram Equalization (CLAHE)
Args:
image2adjust (2d array)
clip_perc (float)
grid_tile (int)
Returns:
CLAHE adjusted 2d array
"""
clahe = cv2.createCLAHE(clipLimit=clip_perc, tileGridSize=grid_tile)
return clahe.apply(image2adjust)
def local_hist_eq(image2adjust, clip_percentages=None, grid_tiles=None, method='mean'):
"""
Computes multi-scale Contrast Limited Adaptive Histogram Equalization (CLAHE)
Args:
image2adjust (ndarray): The edge gradient magnitude array to adjust. Should be uint8 data type.
clip_percentages (Optional[float list]): A list of clip percentages for CLAHE. Default is [1.].
grid_tiles (Optional[tuple list]): A list of grid tuples for CLAHE. Default is [(16, 16)].
method (Optional[str]): The aggregation method.
Returns:
Adjusted image as 2d array.
"""
if not clip_percentages:
clip_percentages = [1.]
if grid_tiles:
grid_tiles = [(gk, gk) for gk in grid_tiles]
else:
grid_tiles = [(16, 16)]
rws, cls = image2adjust.shape
if method == 'mean':
temp_arr_eq = np.zeros((rws, cls), dtype='uint64')
elif method == 'median' or method == 'min':
temp_arr_eq = np.zeros((len(clip_percentages) * len(grid_tiles), rws, cls), dtype='uint64')
counter = 0
# Iterate over each clip percentage.
for clip_perc in clip_percentages:
# Iterate over each grid tile.
for grid_tile in grid_tiles:
# Compute CLAHE and add it to the output array.
if method == 'mean':
temp_arr_eq += _do_clahe(image2adjust, clip_perc, grid_tile)
# temp_arr_eq += rescale_intensity(exposure.equalize_adapthist(image2adjust,
# kernel_size=grid_tile[0],
# clip_limit=clip_perc),
# in_range=(0., 1.), out_range=(0, 255))
elif method == 'median' or method == 'min':
temp_arr_eq[counter] = _do_clahe(image2adjust, clip_perc, grid_tile)
counter += 1
# Return the mean CLAHE-adjusted edge gradient magnitude
if method == 'mean':
return np.float32(temp_arr_eq / float(len(clip_percentages) * len(grid_tiles))) / 255.0
# return np.uint8(np.divide(temp_arr_eq, float(len(clip_percentages) * len(grid_tiles))) / 255.)
elif method == 'median':
return np.float32(np.median(temp_arr_eq, axis=0) / 255.0)
elif method == 'min':
return np.float32(temp_arr_eq.min(axis=0) / 255.0)
def locate_endpoints(edge_image, locations='all'):
"""
Locates edge endpoints
Args:
edge_image (2d array)
locations (Optional[str]): Choices are ['all', 'small', 'broken'].
Returns:
Image endpoints, where endpoints = 1.
"""
# Setup the endpoint structuring elements for
# hit or miss morphology.
if locations == 'all':
endpoints = [np.array([[0, 0, 0], [0, 1, 0], [2, 1, 2]], dtype='uint8'),
np.array([[0, 0, 0], [0, 1, 2], [0, 2, 1]], dtype='uint8'),
np.array([[0, 0, 2], [0, 1, 1], [0, 0, 2]], dtype='uint8'),
np.array([[0, 2, 1], [0, 1, 2], [0, 0, 0]], dtype='uint8'),
np.array([[2, 1, 2], [0, 1, 0], [0, 0, 0]], dtype='uint8'),
np.array([[1, 2, 0], [2, 1, 0], [0, 0, 0]], dtype='uint8'),
np.array([[2, 0, 0], [1, 1, 0], [2, 0, 0]], dtype='uint8'),
np.array([[0, 0, 0], [2, 1, 0], [1, 2, 0]], dtype='uint8'),
np.array([[0, 0, 0], [0, 1, 0], [1, 2, 1]], dtype='uint8'),
np.array([[0, 0, 1], [0, 1, 2], [0, 0, 1]], dtype='uint8'),
np.array([[1, 2, 1], [0, 1, 0], [0, 0, 0]], dtype='uint8')]
elif locations == 'small':
endpoints = [np.array([[0, 0, 0], [0, 1, 0], [1, 1, 1]], dtype='uint8'),
np.array([[0, 0, 0], [0, 1, 1], [0, 1, 1]], dtype='uint8'),
np.array([[0, 0, 1], [0, 1, 1], [0, 0, 1]], dtype='uint8'),
np.array([[0, 1, 1], [0, 1, 1], [0, 0, 0]], dtype='uint8'),
np.array([[1, 1, 1], [0, 1, 0], [0, 0, 0]], dtype='uint8'),
np.array([[1, 1, 0], [1, 1, 0], [0, 0, 0]], dtype='uint8'),
np.array([[1, 0, 0], [1, 1, 0], [1, 0, 0]], dtype='uint8'),
np.array([[0, 0, 0], [1, 1, 0], [1, 1, 0]], dtype='uint8'),
np.array([[0, 0, 0], [0, 1, 0], [1, 1, 1]], dtype='uint8'),
np.array([[0, 0, 1], [0, 1, 1], [0, 0, 1]], dtype='uint8'),
np.array([[1, 1, 1], [0, 1, 0], [0, 0, 0]], dtype='uint8'),
np.array([[1, 0, 0], [1, 1, 0], [1, 0, 0]], dtype='uint8')]
elif locations == 'broken':
endpoints = [np.array([[0, 0, 0], [0, 1, 0], [1, 0, 1]], dtype='uint8'),
np.array([[0, 0, 1], [0, 1, 0], [0, 0, 1]], dtype='uint8'),
np.array([[1, 0, 1], [0, 1, 0], [0, 0, 0]], dtype='uint8'),
np.array([[1, 0, 0], [0, 1, 0], [1, 0, 0]], dtype='uint8')]
end_points = np.zeros(edge_image.shape, dtype='uint8')
# Find the endpoints.
for endpoint in endpoints:
end_points += mhitmiss(np.uint8(edge_image), endpoint)
end_points[end_points > 1] = 1
return end_points
def _locate_islands(edge_image):
"""
Locates single pixel islands
Args:
edge_image (2d array)
Returns:
Image endpoint islands, where islands = 1.
"""
# Setup the endpoint structuring elements for
# hit or miss morphology.
endpoint = np.array([[0, 0, 0], [0, 1, 0], [0, 0, 0]], dtype='uint8')
end_points = np.zeros(edge_image.shape, dtype='uint8')
end_points += mhitmiss(edge_image, endpoint)
end_points[end_points > 1] = 1
return end_points
def _trim_endpoints(edge_image,
iterations,
locations='all',
filter=False,
filter_ws=15,
filter_pct=.1,
skeleton=False):
"""
Trims unconnected lines, starting from endpoints
Args:
edge_image (2d array)
iterations (int)
locations (str)
filter (bool)
filter_ws (int)
filter_pct (float)
skeleton (bool)
"""
if filter:
edge_image_sum = moving_window(edge_image, statistic='sum', window_size=filter_ws)
for iter in range(0, iterations):
# Locate the endpoints
ep = locate_endpoints(edge_image, locations=locations)
# Filter high density areas.
if filter:
ep[edge_image_sum >= int((filter_ws * filter_ws) * filter_pct)] = 0
# Remove the endpoints from the edge image.
edge_image[ep == 1] = 0
# Fill small gaps after the first iteration.
if iter == 0:
edge_image = moving_window(edge_image, statistic='fill', window_size=3, n_neighbors=2)
# Remove remaining single pixels.
ep = _locate_islands(edge_image)
edge_image[ep == 1] = 0
if skeleton:
return _do_skeleton(edge_image)
else:
return edge_image
def _link_edge_endpoints(cr, max_gap, mag_image, **kwargs):
"""
Links edge endpoints
Args:
cr (2d array)
max_gap (int)
mag_image (2d array)
"""
# Link endpoints
cr = moving_window(np.uint8(cr*1),
statistic='link',
window_size=max_gap,
endpoint_array=locate_endpoints(np.uint8(cr*1)),
gradient_array=mag_image,
**kwargs)
# Fill broken links
# __--__ to
# ______
cr = _trim_endpoints(cr, 1, locations='broken')
cr = moving_window(cr, statistic='fill', window_size=3)
# A little cleanup before linking.
cr = _trim_endpoints(cr, 1, locations='all', filter=True, filter_ws=15, filter_pct=.1)
# Link endpoints.
return moving_window(cr * 1,
statistic='link',
window_size=max_gap,
endpoint_array=locate_endpoints(cr * 1),
gradient_array=mag_image,
**kwargs)
def canny_morphology(value_array, egm_array, l1, l2, k_size, l_egm, link_window):
"""
Args:
value_array (2d array): Float32 0-1
egm_array (2d array): Float32 0-1
l1 (int): Canny lower threshold.
l2 (int): Canny upper threshold.
k_size (int): Canny aperture size.
l_egm (float): The EGM lower threshold.
link_window (int): The link window size.
"""
canny_edge = cv2.Canny(np.uint8(value_array * 255.),
l1,
l2,
apertureSize=k_size,
L2gradient=True)
# canny_edge = moving_window(egm_array,
# window_size=3,
# weights=egd,
# statistic='suppression')
canny_edge[canny_edge > 0] = 1
canny_edge = _trim_endpoints(canny_edge, 1, locations='broken')
# Remove small edge objects.
# canny_edge = nd_label(canny_edge)[0]
canny_edge = sk_label(np.uint8(canny_edge), connectivity=2)
# canny_edge = np.uint64(remove_small_objects(canny_edge, min_size=5, connectivity=1))
# Remove objects with low EGM.
props = regionprops(canny_edge, intensity_image=egm_array)
canny_edge = np.float32(canny_edge)
for prop in props:
canny_edge[canny_edge == prop.label] = prop.mean_intensity
canny_edge[canny_edge <= l_egm] = 0
canny_edge[canny_edge > 0] = 1
# Link endpoints
canny_edge = _trim_endpoints(np.uint8(canny_edge), 1, locations='broken')
canny_edge = moving_window(np.uint8(canny_edge),
statistic='link',
window_size=link_window,
endpoint_array=locate_endpoints(np.uint8(canny_edge)),
gradient_array=egm_array,
smallest_allowed_gap=5)
# Remove small objects.
# canny_edge = nd_label(np.uint8(canny_edge))[0]
canny_edge = sk_label(np.uint8(canny_edge), connectivity=2)
canny_edge = np.uint64(remove_small_objects(canny_edge, min_size=10, connectivity=1))
# props = regionprops(canny_edge, intensity_image=egm_array)
# canny_edge = np.float32(canny_edge)
canny_edge[canny_edge > 0] = 1
return _trim_endpoints(canny_edge, 1)
# for prop in props:
#
# if (prop.eccentricity < .4) and (prop.area < 100):
# canny_edge[canny_edge == prop.label] = 0
#
# # if ((prop.major_axis_length + .00001) / (prop.minor_axis_length + .00001) < 2) and (prop.area < 100):
# # canny_edge[canny_edge == prop.label] = 0
#
# canny_edge[canny_edge > 0] = 1
# cannycv_r = cv2.threshold(np.uint8(canny_edge), 0, 1, cv2.THRESH_BINARY_INV)[1]
#
# dist = cv2.distanceTransform(np.uint8(cannycv_r), cv2.DIST_L2, 3)
#
# canny_edge = moving_window(dist, statistic='seg-dist', window_size=3)
#
# canny_edge = moving_window(np.uint8(canny_edge),
# statistic='link',
# window_size=link_window,
# endpoint_array=locate_endpoints(np.uint8(canny_edge)),
# gradient_array=egm_array,
# smallest_allowed_gap=5)
return canny_edge
def _do_skeleton(cr):
"""
Computes the morphological skeleton
Args:
cr (2d array)
Returns:
Image skeleton as 2d array
"""
# Fill holes to keep straighter skeleton lines.
return np.uint8(skeletonize(moving_window(np.uint8(cr), statistic='fill', window_size=3)))
def morphological_cleanup(cr,
min_line_size,
theta_45_iters=0,
theta_90_iters=0,
theta_180_iters=0,
pre_thin=False,
endpoint_iterations=0,
skeleton=False,
link_ends=False,
egm_array=None,
extend_endpoints=False,
max_gap=25,
min_egm=25,
smallest_allowed_gap=3,
medium_allowed_gap=7,
link_iters=1,
link_window_size=7,
extend_iters=1,
value_array=None):
"""
A function to morphologically clean binary edges
Args:
cr (2d array)
min_line_size (int)
theta_45_iters (Optional[int])
theta_90_iters (Optional[int])
theta_180_iters (Optional[int])
pre_thin (Optional[bool])
endpoint_iterations (Optional[int])
skeleton (Optional[bool])
link_ends (Optional[bool])
egm_array (Optional[2d array]): Edge gradient magnitude
extend_endpoints (Optional[bool])
max_gap (Optional[int])
min_egm (Optional[int])
smallest_allowed_gap (Optional[int])
medium_allowed_gap (Optional[int])
link_iters (Optional[int])
link_window_size (Optional[int])
extend_iters (Optional[int])
value_array (Optional[2d array])
Returns:
Morphologically cleaned edges as 2d array
"""
if isinstance(value_array, np.ndarray):
low_value_edge_idx = np.where((cr == 1) & (value_array < 0.2))
if pre_thin:
# Thin edges with 1 iteration
cr = pymorph.thin(pymorph.binary(cr), n=1, Iab=pymorph.endpoints())
# Remove small edge objects.
# cr = nd_label(cr)[0]
cr = sk_label(np.uint8(cr), connectivity=2)
cr = np.uint64(remove_small_objects(cr, min_size=min_line_size, connectivity=1))
cr[cr > 0] = 1
# Extend endpoints along
# the same gradient
# orientation.
if extend_endpoints:
# The edge gradient direction
egd_array = moving_window(egm_array,
window_size=link_window_size,
statistic='edge-direction')
for iter in range(0, extend_iters):
cr = moving_window(cr,
statistic='extend-endpoints',
window_size=3,
endpoint_array=locate_endpoints(cr),
gradient_array=egm_array*255.,
weights=egd_array)
# Thin edges
if (theta_180_iters > 0) and (theta_90_iters > 0) and (theta_45_iters > 0):
# cr = np.uint8(pymorph.thin(pymorph.binary(np.uint8(cr)), theta=180, n=theta_180_iters))
# cr2 = np.uint8(pymorph.thin(pymorph.binary(np.uint8(cr)), theta=90, n=theta_90_iters))
# cr3 = np.uint8(pymorph.thin(pymorph.binary(np.uint8(cr)), n=theta_45_iters))
#
# cr[(cr2 == 1) | (cr3 == 1)] = 1
cr = sk_thin(np.uint8(cr), max_iter=1)
else:
if theta_180_iters > 0:
cr = np.uint8(pymorph.thin(pymorph.binary(np.uint8(cr)), theta=180, n=theta_180_iters))
if theta_90_iters > 0:
cr = np.uint8(pymorph.thin(pymorph.binary(np.uint8(cr)), theta=90, n=theta_90_iters))
if theta_45_iters > 0:
cr = np.uint8(mthin(np.uint8(cr), max_iter=theta_45_iters))
# Remove small objects again after
# thinning and trimming.
if min_line_size > 0:
# cr, __ = nd_label(cr)
cr = sk_label(np.uint8(cr), connectivity=2)
cr = np.uint64(remove_small_objects(cr, min_size=min_line_size, connectivity=1))
cr[cr > 0] = 1
# if skeleton:
# crc = _do_skeleton(cr.copy())
# Link endpoints with small gaps.
if link_ends:
for link_iter in range(0, link_iters):
cr = _link_edge_endpoints(cr,
max_gap,
egm_array,
min_egm=min_egm,
smallest_allowed_gap=smallest_allowed_gap,
medium_allowed_gap=medium_allowed_gap)
cr = _trim_endpoints(cr, 1)
# import matplotlib.pyplot as plt
# cr = _do_skeleton(cr)
# plt.subplot(121)
# plt.imshow(crc)
# plt.subplot(122)
# plt.imshow(cr)
# plt.show()
# import sys
# sys.exit()
# Compute the morphological skeleton.
# The skeleton is morphological thinning with
# infinite iterations.
if skeleton:
cr = _do_skeleton(cr)
# Trim endpoints with ``endpoint_iterations`` iterations.
if endpoint_iterations > 0:
cr = _trim_endpoints(cr, endpoint_iterations, skeleton=True)
# Fill small holes
if isinstance(value_array, np.ndarray):
cr[low_value_edge_idx] = 1
cr = moving_window(cr, statistic='fill', window_size=3, n_neighbors=2)
# Fill broken links
# __--__ to
# ______
cr = _trim_endpoints(cr, 1, locations='broken')
return moving_window(cr, statistic='fill', window_size=3)
def init_distance(egm_array, threshold):
"""
Initializes a euclidean distance transform array
Args:
egm_array (2d array)
threshold (float or int)
"""
# Threshold the EGM into a binary edge/no edge array.
binary_array = np.uint8(np.where(egm_array < threshold, 1, 0))
# Get the euclidean distance from edge pixels.
dist = np.float32(cv2.distanceTransform(binary_array, cv2.DIST_L2, 3))
dist[dist < 0] = 0
dist /= dist.max()
return dist
def init_level_set(egm_array, threshold):
"""
Initializes a level set array
Args:
egm_array (2d array)
threshold (float or int)
"""
# Threshold the EGM into a binary edge/no edge array.
binary_array = np.uint8(np.where(egm_array < threshold, 1, 0))
# Get the euclidean distance from edge pixels.
dist = np.float32(cv2.distanceTransform(binary_array, cv2.DIST_L2, 3))
dist = np.where((binary_array == 1) & (dist > 1), dist, 0)
binary_array_r = np.uint8(cv2.threshold(binary_array, 0, 1, cv2.THRESH_BINARY_INV)[1])
dist_r = cv2.distanceTransform(binary_array_r, cv2.DIST_L2, 3)
return np.where(dist == 0, dist_r * -1., dist)
def multiscale_threshold(egm_array,
min_object_size,
windows=None,
link_ends=False,
theta_180_iters=1,
theta_90_iters=1,
theta_45_iters=1,
skeleton=False,
endpoint_iterations=1,
method='wmean',
ignore_thresh=15.0,
inverse_dist=True,
n_jobs=-1):
"""
Computes multi-scale adaptive threshold and morphological "cleaning"
Args:
egm_array (ndarray):
min_object_size (int):
windows (Optional[int list]):
link_ends (Optional[bool]):
theta_180_iters (Optional[int]):
theta_90_iters (Optional[int]):
theta_45_iters (Optional[int]):
skeleton (Optional[bool]):
endpoint_iterations (Optional[int]):
method (Optional[str]): Choices area ['gaussian', 'mean', 'median', 'weighted'].
ignore_thresh (Optional[float])
inverse_dist (Optional[bool])
n_jobs (Optional[int])
Returns:
Binary edges as 2d array
"""
if not isinstance(windows, list):
windows = [11, 21, 31, 41, 51, 61, 71]
# Get the image shape.
im_rows, im_cols = egm_array.shape
# Setup the output binary edge array holder.
thresholded_edges = np.zeros((im_rows, im_cols), dtype='uint8')
wp = 64
egm_array = cv2.copyMakeBorder(egm_array, wp, wp, wp, wp, cv2.BORDER_REFLECT)
for w in windows:
# Threshold the array with the current window size.
if method == 'gaussian':
# The gaussian threshold is a weighted sum of the window,
# where the weights are a gaussian window.
binary_adaptive_m = cv2.adaptiveThreshold(egm_array, 1,
cv2.ADAPTIVE_THRESH_GAUSSIAN_C,
cv2.THRESH_BINARY, w, 15.)
elif method == 'mean-c':
binary_adaptive_m = cv2.adaptiveThreshold(egm_array, 1,
cv2.ADAPTIVE_THRESH_MEAN_C,
cv2.THRESH_BINARY, w, 15.)
elif method == 'median':
binary_adaptive_m = threshold_local(egm_array, w, method=method)
elif method == 'wmean':
dist_transform = np.float64(init_distance(egm_array, 30))
dist_transform = np.float64(closerec(np.uint8(dist_transform*255.0), 'disk', r=3, iters=5))
dist_transform /= dist_transform.max()
binary_adaptive_m = athreshold(np.ascontiguousarray(egm_array, dtype='float64'),
w,
ignore_thresh=ignore_thresh,
rt=-25.0,
n_jobs=n_jobs,
method=method,
inverse_dist=inverse_dist,
edge_direction_array=None,
edge_distance_array=dist_transform)
elif method == 'bernson':
binary_adaptive_m = athreshold(np.ascontiguousarray(egm_array, dtype='float64'),
w,
ignore_thresh=15.,
rt=-10.,
n_jobs=n_jobs,
method=method)
elif method == 'niblack':
binary_adaptive_m = athreshold(np.ascontiguousarray(egm_array, dtype='float64'),
w,
ignore_thresh=15.,
rt=-10.,
k=-.01,
n_jobs=n_jobs,
method=method)
elif method == 'sauvola':
binary_adaptive_m = athreshold(np.ascontiguousarray(egm_array, dtype='float64'),
w,
ignore_thresh=15.,
rt=-10.,
k=-.01,
n_jobs=n_jobs,
method=method)
elif method == 'bradley':
binary_adaptive_m = athreshold(np.ascontiguousarray(egm_array, dtype='float64'),
w,
ignore_thresh=15.,
rt=1.,
n_jobs=n_jobs,
method=method)
elif method == 'otsu':
binary_adaptive_m = athreshold(np.ascontiguousarray(egm_array, dtype='float64'),
w,
ignore_thresh=15.,
rt=1.,
n_jobs=n_jobs,
method=method)
elif method == '60':
func = lambda arr: np.percentile(arr, 60)
binary_adaptive_m = threshold_local(egm_array, w, 'generic', param=func)
else:
raise ValueError('The method was not recognized.')
# Cleanup the binary edges with image morphology.
thresholded_edges += morphological_cleanup(binary_adaptive_m[wp:-wp, wp:-wp],
min_object_size,
theta_180_iters=theta_180_iters,
theta_90_iters=theta_90_iters,
theta_45_iters=theta_45_iters,
skeleton=skeleton,
endpoint_iterations=endpoint_iterations,
link_ends=link_ends,
egm_array=egm_array)
thresholded_edges[thresholded_edges > 1] = 1
return thresholded_edges
# def _remove_interior_islands(prop,
# min_area_int_,
# mean_threshold_,
# boundary_mean_,
# prop_area_weight_,
# bbox_pad,
# arows,
# acols,
# segments_g,
# original_binary_edge_g,
# se_cross):
def _remove_interior_islands(*args):
"""
Gets indices to remove interior island objects
"""
prop, min_area_int_, mean_threshold_, boundary_mean_, prop_area_weight_, bbox_pad, arows, acols, segments_g, original_binary_edge_g, se_cross = list(itertools.chain(*args))
# mean_threshold_ = 0.2 # The minimum EVI2 threshold allowed for objects
# boundary_mean_ = 0.25 # The maximum EVI2 threshold allowed for boundaries
# min_area_int_ = 222 # The minimum pixel count allowed for interior objects
# Get the bounding box of the current segment.
min_row, min_col, max_row, max_col = prop.bbox
# Expand the box.
min_row = min_row - bbox_pad if (min_row - bbox_pad) > 0 else 0
max_row = max_row + bbox_pad if (max_row + bbox_pad) < (arows - 1) else arows - 1
min_col = min_col - bbox_pad if (min_col - bbox_pad) > 0 else 0
max_col = max_col + bbox_pad if (max_col + bbox_pad) < (acols - 1) else acols - 1
# Get a subset of the current object.
labels_sub = segments_g[min_row:max_row, min_col:max_col]
# Get a subset of the pre-cleaned edges
if isinstance(original_binary_edge_g, np.ndarray):
binary_sub = original_binary_edge_g[min_row:max_row, min_col:max_col]
# Get the count of pre-cleaned
# edges in the object.
binary_edge_count = ((binary_sub == 1) & (labels_sub == prop.label)).sum()
# Don't include objects half covered by pre-cleaned edges.
if binary_edge_count >= int(prop.area * prop_area_weight_):
idx = list(np.where(labels_sub == prop.label))
idx[0] = idx[0] + min_row
idx[1] = idx[1] + min_col
return list(idx[0]), list(idx[1])
# Don't include objects with low EVI2.
if hasattr(prop, 'mean_intensity'):
if prop.mean_intensity < mean_threshold_:
idx = list(np.where(labels_sub == prop.label))
idx[0] = idx[0] + min_row
idx[1] = idx[1] + min_col
return list(idx[0]), list(idx[1])
# Get the current object.
labels_sub_center = np.uint8(np.where(labels_sub == prop.label, 1, 0))
# Get the boundary labels.
label_boundary = cv2.morphologyEx(labels_sub_center,
cv2.MORPH_DILATE,
se_cross,
iterations=2) - labels_sub_center
boundary_idx = np.where(label_boundary == 1)
# Check if the current object is completely
# surrounded by 1-2 other objects.
if np.any(boundary_idx):
boundary_values = labels_sub[boundary_idx]
# The parcel should be surrounded
# by other vegetation.
if boundary_values.mean() >= boundary_mean_:
unique_boundary_values = list(np.unique(boundary_values))
if (0 in unique_boundary_values) and (0 < len(unique_boundary_values) <= 2) and (prop.area < min_area_int_):
idx = list(np.where(labels_sub_center == 1))
idx[0] = idx[0] + min_row
idx[1] = idx[1] + min_col
return list(idx[0]), list(idx[1])
else:
return list(), list()
else:
return list(), list()
else:
return list(), list()
# def _clean_objects(prop,
# min_area_,
# min_area_int_,
# mean_threshold_,
# boundary_mean_,
# bbox_pad,
# arows,
# acols,
# segments_g,
# morphed_sep,
# morphed,
# se_cross,
# se_square):
def _clean_objects(*args):
"""
Area:
15m:
0.1 ha / [(15m x 15m) x 0.0001] = 5 pixels
5 ha / [(15m x 15m) x 0.0001] = 222 pixels
10 ha / [(15m x 15m) x 0.0001] = 444 pixels
20 ha / [(15m x 15m) x 0.0001] = 888 pixels
5,000 ha / [(15m x 15m) x 0.0001] = 222,222 pixels
10,000 ha / [(15m x 15m) x 0.0001] = 444,444 pixels
20,000 ha / [(15m x 15m) x 0.0001] = 888,888 pixels
"""
prop, min_area_, min_area_int_, mean_threshold_, boundary_mean_, bbox_pad, arows, acols, segments_g, morphed_sep, morphed, se_cross, se_square = list(itertools.chain(*args))
el_ = []
# mean_threshold_ = 0.2 # The minimum EVI2 threshold allowed for objects
# boundary_mean_ = 0.25 # The maximum EVI2 threshold allowed for boundaries
# min_area_ = 5 # The minimum pixel count allowed for any object
# max_area_ = 250000 # The maximum pixel count allowed for any object
# min_area_int_ = 222 # The minimum pixel count allowed for interior objects
# if prop.area > 10000:
# return el_, el_, el_, el_, el_
if hasattr(prop, 'mean_intensity'):
if prop.mean_intensity < mean_threshold_:
return el_, el_, el_, el_, el_
# Get the bounding box of the current segment.
min_row, min_col, max_row, max_col = prop.bbox
# Expand the box.
min_row = min_row - bbox_pad if (min_row - bbox_pad) > 0 else 0
max_row = max_row + bbox_pad if (max_row + bbox_pad) < (arows - 1) else arows - 1
min_col = min_col - bbox_pad if (min_col - bbox_pad) > 0 else 0
max_col = max_col + bbox_pad if (max_col + bbox_pad) < (acols - 1) else acols - 1
# Get a subset of the current object.
labels_sub = segments_g[min_row:max_row, min_col:max_col]
morphed_sep_sub = morphed_sep[min_row:max_row, min_col:max_col]
morphed_sub = morphed[min_row:max_row, min_col:max_col]
# Get the current object.
labels_sub_center = np.uint8(np.where(labels_sub == prop.label, 1, 0))
# Get the boundary labels.
label_boundary = cv2.morphologyEx(labels_sub_center,
cv2.MORPH_DILATE,
se_cross,
iterations=2) - labels_sub_center
boundary_idx = np.where(label_boundary == 1)
# Check if the current object is completely
# surrounded by 1-3 other objects.
if np.any(boundary_idx):
boundary_values = labels_sub[boundary_idx]
# The parcel should be surrounded
# by other vegetation.
if boundary_values.mean() >= boundary_mean_:
unique_boundary_values = list(np.unique(boundary_values))
if (0 in unique_boundary_values) and (0 < len(unique_boundary_values) <= 2) and (prop.area < min_area_int_):
return el_, el_, el_, el_, el_
# Morphological closing by reconstruction
closerec_sub = pymorph.closerec(pymorph.binary(labels_sub_center))
closerec_sub = merode(closerec_sub, se_cross)
closerec_sub = mopen(closerec_sub, se_square)
if (closerec_sub == 1).sum() < min_area_:
return el_, el_, el_, el_, el_
else:
idxs = list(np.where((morphed_sep_sub == 0) & (closerec_sub == 1)))
idxs[0] = idxs[0] + min_row
idxs[1] = idxs[1] + min_col
# Decrease the gaps between land cover
closerec_sub = cv2.morphologyEx(np.uint8(closerec_sub),
cv2.MORPH_DILATE,
se_cross,
iterations=2)
idx = list(np.where((morphed_sub == 0) & (closerec_sub == 1)))
idx[0] = idx[0] + min_row
idx[1] = idx[1] + min_col
return list(idxs[0]), list(idxs[1]), \
list(np.zeros(len(idx[0]), dtype='uint64') + prop.label), \
list(idx[0]), list(idx[1])
def clean_objects(segments,
intensity_array=None,
original_binary_edge=None,
binary=True,
min_object_area=5,
min_interior_count=222,
mean_threshold=0.2,
boundary_mean=0.25,
prop_area_weight=0.9,
bbox_pad=10,
chunk_size=100000,
n_jobs=1):
"""
Cleans objects with morphological operations
Args:
segments (2d array): The segmented objects array to be cleaned.
intensity_array (2d array): The intensity values.
original_binary_edge (2d array): The original edges as binary.
binary (Optional[bool]): Whether the input segments are binary (True) or labelled (False). Default is True.
min_object_area (Optional[int]): The minimum object area.
min_interior_count (Optional[int]): The minimum pixel count of interior pixels.
mean_threshold (float): The vegetation index mean threshold.
boundary_mean (float): The vegetation index boundary threshold.
prop_area_weight (float): The object property area weighting.
bbox_pad (Optional[int]): The `regionprops bbox` padding. Default is 10.
chunk_size (Optional[int]): The chunk size for multiprocessing. Default is 100,000.
n_jobs (Optional[int]):
Returns:
Segments dilated, Segments with separate boundaries
"""
# global morphed, segments_g, morphed_sep, se_cross, se_square, original_binary_edge_g
# segments_g = segments
# original_binary_edge_g = original_binary_edge
arows, acols = segments.shape
if binary:
segments = nd_label(segments)[0]
# segments = sk_label(np.uint8(segments), connectivity=1)
morphed_sep = np.zeros((arows, acols), dtype='uint8')
morphed = np.zeros((arows, acols), dtype='uint64')
se_cross = np.array([[0, 1, 0],
[1, 1, 1],
[0, 1, 0]], dtype='uint8')
se_square = np.array([[1, 1],
[1, 1]], dtype='uint8')
props = regionprops(segments,
intensity_image=intensity_array)
# Clean parcels
for chi in range(0, len(props), chunk_size):
data_gen = ((prop_,
min_object_area,
min_interior_count,
mean_threshold,
boundary_mean,
bbox_pad,
arows,
acols,
segments,
morphed_sep,
morphed,
se_cross,
se_square) for prop_ in props[chi:chi+chunk_size])
cleaned_parcels = []
with concurrent.futures.ThreadPoolExecutor(n_jobs) as executor:
for res in executor.map(_clean_objects, data_gen):
cleaned_parcels.append(res)
# cleaned_parcels = Parallel(backend='multiprocessing',
# n_jobs=n_jobs)(delayed(_clean_objects)(prop_,
# min_object_area,
# min_interior_count,
# mean_threshold,
# boundary_mean,
# bbox_pad,
# arows,
# acols,
# segments,
# morphed_sep,
# morphed,
# se_cross,
# se_square) for prop_ in props[chi:chi+chunk_size])
rowidx_sep_list, colidx_sep_list, labels_list, rowidx_list, colidx_list = zip(*cleaned_parcels)
labels_list = np.array(list(itertools.chain.from_iterable(labels_list)), dtype='uint64')
# piece together the parcels
if np.any(labels_list):
# Objects with separate boundaries
rowidx_sep_list = np.array(list(itertools.chain.from_iterable(rowidx_sep_list)), dtype='uint64')
colidx_sep_list = np.array(list(itertools.chain.from_iterable(colidx_sep_list)), dtype='uint64')
morphed_sep[(rowidx_sep_list, colidx_sep_list)] = 1
# Objects with dilated boundaries
rowidx_list = np.array(list(itertools.chain.from_iterable(rowidx_list)), dtype='uint64')
colidx_list = np.array(list(itertools.chain.from_iterable(colidx_list)), dtype='uint64')
morphed[(rowidx_list, colidx_list)] = labels_list
# One last check for interior islands
props = regionprops(morphed,
intensity_image=intensity_array)
# segments_g = morphed
for chi in range(0, len(props), chunk_size):
data_gen = ((prop_,
min_interior_count,
mean_threshold,
boundary_mean,
prop_area_weight,
bbox_pad,
arows,
acols,
morphed,
original_binary_edge,
se_cross) for prop_ in props[chi:chi+chunk_size])
cleaned_parcels = []
with concurrent.futures.ThreadPoolExecutor(n_jobs) as executor:
for res in executor.map(_remove_interior_islands, data_gen):
cleaned_parcels.append(res)
# cleaned_parcels = Parallel(backend='multiprocessing',
# n_jobs=n_jobs)(delayed(_remove_interior_islands)(prop_,
# min_interior_count,
# mean_threshold,
# boundary_mean,
# prop_area_weight,
# bbox_pad,
# arows,
# acols,
# morphed,
# original_binary_edge,
# se_cross) for prop_ in props[chi:chi+chunk_size])
rowidx_list, colidx_list = zip(*cleaned_parcels)
rowidx_list = np.array(list(itertools.chain.from_iterable(rowidx_list)), dtype='uint64')
colidx_list = np.array(list(itertools.chain.from_iterable(colidx_list)), dtype='uint64')
# piece together the parcels
if np.any(rowidx_list):
morphed[(rowidx_list, colidx_list)] = 0
morphed_sep[morphed == 0] = 0
return morphed, morphed_sep
def invert_size_check(im_edges, min_size_object, binary=True):
"""
Inverts binary edges and checks sizes
Args:
im_edges (2d array): Edge array, where edges = 1.
min_size_object (int): The minimum size of line to be retained.
binary (Optional[bool]): Whether to recode the output labelled edges to binary. Default is True.
Returns:
Image objects as 2d array
"""
# Invert edges to objects
im_objects = cv2.threshold(np.uint8(im_edges), 0, 1, cv2.THRESH_BINARY_INV)[1]
# Remove potential field objects smaller
# than size threshold.
im_objects = nd_label(im_objects)[0]
# im_objects = sk_label(np.uint8(im_objects), connectivity=1)
im_objects = np.uint64(remove_small_objects(im_objects,
min_size=min_size_object,
connectivity=1))
if binary:
im_objects[im_objects > 0] = 1
return im_objects
def _intersect_objects(prop):
lc_values = list()
area_values = list()
orient_values = list()
solidity_values = list()
eccentricity_values = list()
if prop.label == 0:
return lc_values, area_values, orient_values, solidity_values, eccentricity_values, list(), list()
# Get the bounding box of the current segment.
min_row, min_col, max_row, max_col = prop.bbox
# Get the label sub-array
# labels_sub = lc_objects_g[min_row:max_row, min_col:max_col]
# Get the indices of the current object.
# labels_sub_idx = list(np.where(labels_sub == prop.label))
labels_sub_idx = (prop.coords[:, 0], prop.coords[:, 1])
labels_sub_idx_object = (prop.coords[:, 0] - min_row, prop.coords[:, 1] - min_col)
n_samples = len(labels_sub_idx[0])
# labels_sub_idx[0] = labels_sub_idx[0] + min_row
# labels_sub_idx[1] = labels_sub_idx[1] + min_col
##################################
# LAND COVER CLASS ID INTERSECTION
##################################
if get_class_id_g:
# Get the land cover
# class for the object.
lc_array_sub = lc_array_g[min_row:max_row, min_col:max_col]
# Get the majority land cover class
lc_mode_object = sci_mode(lc_array_sub[tuple(labels_sub_idx_object)])
lc_mode = int(lc_mode_object.mode)
lc_count = int(lc_mode_object.count)
# Check that the land cover count
# is above the required threshold.
# The pixel count needed
# to meet the threshold
pix_count = int(prop.area * object_fraction_g)
# There must be at least
# `object_fraction_g` of the
# target class in the object.
if lc_count >= pix_count:
# Return the majority class
lc_values = list(np.zeros(n_samples, dtype='uint8') + lc_mode)
else:
# Return empty
lc_values = list(np.zeros(n_samples, dtype='uint8'))
# Get the current object.
# labels_sub_center = np.uint8(np.where(labels_sub == idx, 1, 0))
##########################
# OBJECT AREA INTERSECTION
##########################
if get_object_area_g:
object_area = round(prop.area * pixel_ha, 2)
area_values = list(np.zeros(n_samples, dtype='float32') + object_area)
########################
# OBJECT ID INTERSECTION
########################
if get_orientation_g:
orient_values = list(np.zeros(n_samples, dtype='float32') + prop.orientation)
if get_solidity_g:
solidity_values = list(np.zeros(n_samples, dtype='float32') + prop.solidity)
if get_eccentricity_g:
eccentricity_values = list(np.zeros(n_samples, dtype='float32') + prop.eccentricity)
# Return the object value
return lc_values, \
area_values, \
orient_values, \
solidity_values, \
eccentricity_values, \
list(labels_sub_idx[0]), \
list(labels_sub_idx[1])
def intersect_objects(lc_objects,
lc_objects_sep=None,
lc_array=None,
var_array=None,
objects_are_unique=False,
object_fraction=0.5,
get_object_area=True,
get_object_id=False,
get_class_id=False,
get_orientation=False,
get_solidity=False,
get_eccentricity=False,
cell_size=30.0,
n_jobs=1,
chunk_size=100000):
""""
Intersects land cover objects with a thematic land cover map
Args:
lc_objects (2d array): The segmented objects.
lc_objects_sep (2d array): The eroded segmented objects.
lc_array (Optional[2d array]): The land cover array, needed if `get_object_area = False`. Default is None.
var_array (Optional[2d array]): The image variables array. Default is None.
objects_are_unique (Optional[bool]): Whether the land cover objects of `lc_objects` are unique.
Default is False.
object_fraction (Optional[float]): The fraction of an object in `lc_objects` to be considered for intersection.
Default is 0.5.
get_object_area (Optional[bool]): Whether to return the object area. Default is True.
get_object_id (Optional[bool]): Whether to return the object id from `lc_objects`. Default is False.
get_class_id (Optional[bool]): Whether to return the land cover class id from `lc_array`. Default is False.
get_orientation (Optional[bool]): Whether to return the object orientation. Default is False.
get_solidity (Optional[bool]): Whether to return the object solidity. Default is False.
get_eccentricity (Optional[bool]): Whether to return the object eccentricity. Default is False.
cell_size (Optional[float]): The cell size, used when `get_object_area = True`. Default is 30.
n_jobs (Optional[int]): The number of parallel jobs. Default is 1.
chunk_size (Optional[int]): The chunk size for Pool. Default is 100,000.
"""
global object_fraction_g, get_object_area_g, get_object_id_g, \
get_class_id_g, get_orientation_g, get_solidity_g, get_eccentricity_g, \
pixel_ha, lc_objects_g, lc_array_g
object_fraction_g = object_fraction
get_object_area_g = get_object_area
get_object_id_g = get_object_id
get_class_id_g = get_class_id
get_orientation_g = get_orientation
get_solidity_g = get_solidity
get_eccentricity_g = get_eccentricity
lc_objects_g = lc_objects
lc_array_g = lc_array
# Square meters to hectares
pixel_ha = (cell_size * cell_size) * 0.0001
out_array = np.zeros((5, lc_objects.shape[0], lc_objects.shape[1]), dtype='float32')
# Get unique object ids.
if not objects_are_unique:
lc_objects[lc_objects > 0] = 1
lc_objects, n_objects = nd_label(lc_objects)
# Get object properties.
# zo = prop.area
props_int = regionprops(lc_objects)
for chi in range(0, len(props_int), chunk_size):
with pooler(processes=n_jobs) as pool:
# Get object statistics
intersected_objects = pool.map(_intersect_objects,
props_int[chi:chi+chunk_size],
chunk_size)
lc_values_, area_values_, ori_values_, sol_values_, ecc_values_, rowidx_list, colidx_list = zip(*intersected_objects)
# Join the lists
lc_values_ = np.array(list(itertools.chain.from_iterable(lc_values_)), dtype='uint8')
area_values_ = np.array(list(itertools.chain.from_iterable(area_values_)), dtype='float32')
ori_values_ = np.array(list(itertools.chain.from_iterable(ori_values_)), dtype='float32')
sol_values_ = np.array(list(itertools.chain.from_iterable(sol_values_)), dtype='float32')
ecc_values_ = np.array(list(itertools.chain.from_iterable(ecc_values_)), dtype='float32')
rowidx_list = np.array(list(itertools.chain.from_iterable(rowidx_list)), dtype='uint64')
colidx_list = np.array(list(itertools.chain.from_iterable(colidx_list)), dtype='uint64')
# Piece together the parcels
# land cover
if lc_values_.shape[0] > 0:
out_array[0, rowidx_list, colidx_list] = lc_values_
# area
if area_values_.shape[0] > 0:
out_array[1, rowidx_list, colidx_list] = area_values_
# orientation
if ori_values_.shape[0] > 0:
out_array[2, rowidx_list, colidx_list] = ori_values_
# solidarity
if sol_values_.shape[0] > 0:
out_array[3, rowidx_list, colidx_list] = sol_values_
# eccentricity
if ecc_values_.shape[0] > 0:
out_array[4, rowidx_list, colidx_list] = ecc_values_
if isinstance(lc_objects_sep, np.ndarray):
# Swap the land cover with the eroded objects
out_array[0] = np.where(lc_objects_sep > 0, out_array[0], 0)
if isinstance(var_array, np.ndarray):
# Give the objects unique labels
lc_objects_labs_ = sk_label(
|
np.uint8(lc_objects_sep)
|
numpy.uint8
|
# Copyright 2019 Image Analysis Lab, German Center for Neurodegenerative Diseases (DZNE), Bonn
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# IMPORTS
import os
import torch
import time
import matplotlib.pyplot as plt
import numpy as np
import itertools
import glob
from torch.autograd import Variable
from torch.optim import lr_scheduler
from torchvision import utils
from skimage import color
from models.losses import CombinedLoss
##
# Helper functions
##
def create_exp_directory(exp_dir_name):
"""
Function to create a directory if it does not exist yet.
:param str exp_dir_name: name of directory to create.
:return:
"""
if not os.path.exists(exp_dir_name):
try:
os.makedirs(exp_dir_name)
print("Successfully Created Directory @ {}".format(exp_dir_name))
except:
print("Directory Creation Failed - Check Path")
else:
print("Directory {} Exists ".format(exp_dir_name))
def dice_confusion_matrix(batch_output, labels_batch, num_classes):
"""
Function to compute the dice confusion matrix.
:param batch_output:
:param labels_batch:
:param num_classes:
:return:
"""
dice_cm = torch.zeros(num_classes, num_classes)
for i in range(num_classes):
gt = (labels_batch == i).float()
for j in range(num_classes):
pred = (batch_output == j).float()
inter = torch.sum(torch.mul(gt, pred)) + 0.0001
union = torch.sum(gt) + torch.sum(pred) + 0.0001
dice_cm[i, j] = 2 * torch.div(inter, union)
avg_dice = torch.mean(torch.diagflat(dice_cm))
return avg_dice, dice_cm
def iou_score(pred_cls, true_cls, nclass=79):
"""
compute the intersection-over-union score
both inputs should be categorical (as opposed to one-hot)
"""
intersect_ = []
union_ = []
for i in range(1, nclass):
intersect = ((pred_cls == i).float() + (true_cls == i).float()).eq(2).sum().item()
union = ((pred_cls == i).float() + (true_cls == i).float()).ge(1).sum().item()
intersect_.append(intersect)
union_.append(union)
return np.array(intersect_), np.array(union_)
def precision_recall(pred_cls, true_cls, nclass=79):
"""
Function to calculate recall (TP/(TP + FN) and precision (TP/(TP+FP) per class
:param pytorch.Tensor pred_cls: network prediction (categorical)
:param pytorch.Tensor true_cls: ground truth (categorical)
:param int nclass: number of classes
:return:
"""
tpos_fneg = []
tpos_fpos = []
tpos = []
for i in range(1, nclass):
all_pred = (pred_cls == i).float()
all_gt = (true_cls == i).float()
tpos.append((all_pred + all_gt).eq(2).sum().item())
tpos_fpos.append(all_pred.sum().item())
tpos_fneg.append(all_gt.sum().item())
return np.array(tpos), np.array(tpos_fneg), np.array(tpos_fpos)
##
# Plotting functions
##
def plot_predictions(images_batch, labels_batch, batch_output, plt_title, file_save_name):
"""
Function to plot predictions from validation set.
:param images_batch:
:param labels_batch:
:param batch_output:
:param plt_title:
:param file_save_name:
:return:
"""
f = plt.figure(figsize=(20, 20))
n, c, h, w = images_batch.shape
mid_slice = c // 2
images_batch = torch.unsqueeze(images_batch[:, mid_slice, :, :], 1)
grid = utils.make_grid(images_batch.cpu(), nrow=4)
plt.subplot(131)
plt.imshow(grid.numpy().transpose((1, 2, 0)))
plt.title('Slices')
grid = utils.make_grid(labels_batch.unsqueeze_(1).cpu(), nrow=4)[0]
color_grid = color.label2rgb(grid.numpy(), bg_label=0)
plt.subplot(132)
plt.imshow(color_grid)
plt.title('Ground Truth')
grid = utils.make_grid(batch_output.unsqueeze_(1).cpu(), nrow=4)[0]
color_grid = color.label2rgb(grid.numpy(), bg_label=0)
plt.subplot(133)
plt.imshow(color_grid)
plt.title('Prediction')
plt.suptitle(plt_title)
plt.tight_layout()
f.savefig(file_save_name, bbox_inches='tight')
plt.close(f)
plt.gcf().clear()
def plot_confusion_matrix(cm, classes, title='Confusion matrix', cmap=plt.cm.Blues, file_save_name="temp.pdf"):
"""
This function prints and plots the confusion matrix.
Normalization can be applied by setting `normalize=True`.
:param cm:
:param classes:
:param title:
:param cmap:
:param file_save_name:
:return:
"""
f = plt.figure(figsize=(35, 35))
plt.imshow(cm, interpolation='nearest', cmap=cmap)
plt.title(title)
plt.colorbar()
tick_marks = np.arange(len(classes))
plt.xticks(tick_marks, classes, rotation=45)
plt.yticks(tick_marks, classes)
fmt = '.2f'
thresh = cm.max() / 2.
for i, j in itertools.product(range(cm.shape[0]), range(cm.shape[1])):
plt.text(j, i, format(cm[i, j], fmt),
horizontalalignment="center",
color="white" if cm[i, j] > thresh else "black")
plt.tight_layout()
plt.ylabel('True label')
plt.xlabel('Predicted label')
f.savefig(file_save_name, bbox_inches='tight')
plt.close(f)
plt.gcf().clear()
##
# Training routine
##
class Solver(object):
"""
Class for training neural networks
"""
# gamma is the factor for lowering the lr and step_size is when it gets lowered
default_lr_scheduler_args = {"gamma": 0.05,
"step_size": 5}
def __init__(self, num_classes, optimizer=torch.optim.Adam, optimizer_args={}, loss_func=CombinedLoss(),
lr_scheduler_args={}):
# Merge and update the default arguments - optimizer
self.optimizer_args = optimizer_args
lr_scheduler_args_merged = Solver.default_lr_scheduler_args.copy()
lr_scheduler_args_merged.update(lr_scheduler_args)
# Merge and update the default arguments - lr scheduler
self.lr_scheduler_args = lr_scheduler_args_merged
self.optimizer = optimizer
self.loss_func = loss_func
self.num_classes = num_classes
self.classes = list(range(self.num_classes))
def train(self, model, train_loader, validation_loader, class_names, num_epochs,
log_params, expdir, scheduler_type, torch_v11, resume=True):
"""
Train Model with provided parameters for optimization
Inputs:
-- model - model to be trained
-- train_loader - training DataLoader Object
-- validation_loader - validation DataLoader Object
-- num_epochs = total number of epochs
-- log_params - parameters for logging the progress
-- expdir --directory to save check points
"""
create_exp_directory(expdir) # Experimental directory
create_exp_directory(log_params["logdir"]) # Logging Directory
# Instantiate the optimizer class
optimizer = self.optimizer(model.parameters(), **self.optimizer_args)
# Instantiate the scheduler class
if scheduler_type == "StepLR":
scheduler = lr_scheduler.StepLR(optimizer, step_size=self.lr_scheduler_args["step_size"],
gamma=self.lr_scheduler_args["gamma"])
else:
scheduler = None
# Set up logger format
a = "{}\t" * (self.num_classes - 2) + "{}"
epoch = -1 # To allow for restoration
print('-------> Starting to train')
# Code for restoring model
if resume:
try:
prior_model_paths = sorted(glob.glob(os.path.join(expdir, 'Epoch_*')), key=os.path.getmtime)
if prior_model_paths:
current_model = prior_model_paths.pop()
state = torch.load(current_model)
# Restore model dictionary
model.load_state_dict(state["model_state_dict"])
optimizer.load_state_dict(state["optimizer_state_dict"])
scheduler.load_state_dict(state["scheduler_state_dict"])
epoch = state["epoch"]
print("Successfully restored the model state. Resuming training from Epoch {}".format(epoch + 1))
except Exception as e:
print("No model to restore. Resuming training from Epoch 0. {}".format(e))
log_params["logger"].info("{} parameters in total".format(sum(x.numel() for x in model.parameters())))
while epoch < num_epochs:
epoch = epoch + 1
epoch_start = time.time()
# Update learning rate based on epoch number (only for pytorch version <1.2)
if torch_v11 and scheduler is not None:
scheduler.step()
loss_batch = np.zeros(1)
for batch_idx, sample_batch in enumerate(train_loader):
# Assign data
images_batch, labels_batch, weights_batch = sample_batch['image'], sample_batch['label'], \
sample_batch['weight']
# Map to variables
images_batch = Variable(images_batch)
labels_batch = Variable(labels_batch)
weights_batch = Variable(weights_batch)
if torch.cuda.is_available():
images_batch, labels_batch, weights_batch = images_batch.cuda(), labels_batch.cuda(), \
weights_batch.type(torch.FloatTensor).cuda()
model.train() # Set to training mode!
optimizer.zero_grad()
predictions = model(images_batch)
loss_total, loss_dice, loss_ce = self.loss_func(predictions, labels_batch, weights_batch)
loss_total.backward()
optimizer.step()
loss_batch += loss_total.item()
if batch_idx % (len(train_loader) // 2) == 0 or batch_idx == len(train_loader) - 1:
log_params["logger"].info("Train Epoch: {} [{}/{}] ({:.0f}%)] "
"with loss: {}".format(epoch, batch_idx,
len(train_loader),
100. * batch_idx / len(train_loader),
loss_batch / (batch_idx + 1)))
del images_batch, labels_batch, weights_batch, predictions, loss_total, loss_dice, loss_ce
# Update learning rate at the end based on epoch number (only for pytorch version > 1.1)
if not torch_v11 and scheduler is not None:
scheduler.step()
epoch_finish = time.time() - epoch_start
log_params["logger"].info("Train Epoch {} finished in {:.04f} seconds.".format(epoch, epoch_finish))
# End of Training, time to accumulate results
# Testing Loop on Training Data
# Set evaluation mode on the model
model.eval()
val_loss_total = 0
val_loss_dice = 0
val_loss_ce = 0
ints_ = np.zeros(self.num_classes - 1)
unis_ = np.zeros(self.num_classes - 1)
per_cls_counts_gt = np.zeros(self.num_classes - 1)
per_cls_counts_pred = np.zeros(self.num_classes - 1)
accs = np.zeros(self.num_classes - 1) # -1 to exclude background (still included in val loss)
with torch.no_grad():
if validation_loader is not None:
val_start = time.time()
cnf_matrix_validation = torch.zeros(self.num_classes, self.num_classes)
for batch_idx, sample_batch in enumerate(validation_loader):
images_batch, labels_batch, weights_batch = sample_batch['image'], sample_batch['label'], \
sample_batch['weight']
# Map to variables (no longer necessary after pytorch 0.40)
images_batch = Variable(images_batch)
labels_batch = Variable(labels_batch)
weights_batch = Variable(weights_batch)
if torch.cuda.is_available():
images_batch, labels_batch, weights_batch = images_batch.cuda(), labels_batch.cuda(), \
weights_batch.type(torch.FloatTensor).cuda()
# Get logits, sum up batch loss and get final predictions (argmax)
predictions = model(images_batch)
loss_total, loss_dice, loss_ce = self.loss_func(predictions, labels_batch, weights_batch)
val_loss_total += loss_total.item()
val_loss_dice += loss_dice.item()
val_loss_ce += loss_ce.item()
_, batch_output = torch.max(predictions, dim=1)
# Calculate iou_scores, accuracy and dice confusion matrix + sum over previous batches
int_, uni_ = iou_score(batch_output, labels_batch, self.num_classes)
ints_ += int_
unis_ += uni_
tpos, pcc_gt, pcc_pred = precision_recall(batch_output, labels_batch, self.num_classes)
accs += tpos
per_cls_counts_gt += pcc_gt
per_cls_counts_pred += pcc_pred
_, cm_batch = dice_confusion_matrix(batch_output, labels_batch, self.num_classes)
cnf_matrix_validation += cm_batch.cpu()
# Plot sample predictions
if batch_idx == 0:
plt_title = 'Validation Results Epoch ' + str(epoch)
file_save_name = os.path.join(log_params["logdir"],
'Epoch_' + str(epoch) + '_Validations_Predictions.pdf')
plot_predictions(images_batch, labels_batch, batch_output, plt_title, file_save_name)
del images_batch, labels_batch, weights_batch, predictions, batch_output, \
int_, uni_, tpos, pcc_gt, pcc_pred, loss_total, loss_dice, loss_ce # cm_batch,
# Get final measures and log them
ious = ints_ / unis_
val_loss_total /= (batch_idx + 1)
val_loss_dice /= (batch_idx + 1)
val_loss_ce /= (batch_idx + 1)
cnf_matrix_validation = cnf_matrix_validation / (batch_idx + 1)
val_end = time.time() - val_start
print("Completed Validation Dataset in {:0.4f} s".format(val_end))
save_name = os.path.join(log_params["logdir"], 'Epoch_' + str(epoch) + '_Validation_Dice_CM.pdf')
plot_confusion_matrix(cnf_matrix_validation.cpu().numpy(), self.classes, file_save_name=save_name)
# Log metrics
log_params["logger"].info("[Epoch {} stats]: MIoU: {:.4f}; "
"Mean Recall: {:.4f}; "
"Mean Precision: {:.4f}; "
"Avg loss total: {:.4f}; "
"Avg loss dice: {:.4f}; "
"Avg loss ce: {:.4f}".format(epoch, np.mean(ious),
np.mean(accs / per_cls_counts_gt),
|
np.mean(accs / per_cls_counts_pred)
|
numpy.mean
|
import numpy as np
from scipy.optimize import curve_fit
from sklearn.decomposition import PCA
from sklearn.linear_model import RidgeClassifier
from sklearn.multiclass import OneVsOneClassifier
from sklearn.model_selection import KFold
class NestedXval():
'''A generator for nested cross-validation that ensures that there is the
same number of trials for each class in training.
It is necessary to have the same number of trials in each category to
vectorize the training of the decoder so that the training of all 6
decoders (in one vs one scheme) is done simultaneously.
'''
def __init__(self, n_outer_splits=None):
'''Nested crossvalidation to get the same number of trials in each class for training'''
self.nouter = n_outer_splits
self.ninner = 2
self.outerxval = KFold(n_splits=n_outer_splits)
def split(self, targets):
'''Returns a generator that splits data in test, train and subspace
with the same number of trials in each category
'''
labels, counts = np.unique(targets, return_counts=True)
nclasses = len(labels)
if not np.all(counts[0] == counts) and max(counts) - min(counts) > 1:
raise ValueError("The number of trials in each class in not consistant")
interleaved_outer = np.concatenate(list(zip(*[np.where(targets == label)[0] for label in labels])))
leftovers = []
for iclass in np.where(np.min(counts) < counts)[0]:
leftovers.append(np.where(targets == labels[iclass])[0][-1])
interleaved_outer = np.concatenate((interleaved_outer, np.array(leftovers))).astype(int)
targets_ = targets[interleaved_outer]
outersplit = self.outerxval.split(targets)
for ioutsplit in range(self.nouter):
restinds, testinds = next(outersplit)
ntrain_per_class = np.ceil(len(restinds) / 2 / nclasses).astype(int)
inner_inds_by_class = [np.where(targets_[restinds] == label)[0] for label in labels]
traininds = np.concatenate(list(zip(*[restinds[classinds[:ntrain_per_class]] for classinds in inner_inds_by_class])))
subinds = np.concatenate([restinds[classinds[ntrain_per_class:]] for classinds in inner_inds_by_class])
testinds = interleaved_outer[testinds]
traininds = interleaved_outer[traininds]
subinds = interleaved_outer[subinds]
yield np.sort(testinds), np.sort(traininds), np.sort(subinds)
traininds = np.concatenate(list(zip(*[restinds[classinds[:-ntrain_per_class:-1]] for classinds in inner_inds_by_class])))
subinds = np.concatenate([restinds[classinds[-ntrain_per_class::-1]] for classinds in inner_inds_by_class])
testinds = interleaved_outer[testinds]
traininds = interleaved_outer[traininds]
subinds = interleaved_outer[subinds]
yield np.sort(testinds), np.sort(traininds), np.sort(subinds)
def sub_split(targets, trainind):
'''Cross-validation generator for the decoder and subspace trials
Function to split training trials in training and subspace trials, ensuring
that there is the same number of trials in each class for training.
Parameters
----------
targets : np.array - The targets (or y values)
trainind : np.array - The indices of the training trials
Returns
-------
Generator for each fold. Yields a tuple of np.array, one array for the
training trials and one array for the subspace
'''
targets = targets[trainind]
labels = np.unique(targets)
nclasses = len(labels)
ntrain_per_class = np.ceil(len(targets) / 2 / nclasses).astype(int)
inner_inds_by_class = [np.where(targets == label)[0] for label in labels]
ridgeind = np.concatenate(list(zip(*[classinds[:ntrain_per_class] for classinds in inner_inds_by_class])))
subind = np.concatenate([classinds[ntrain_per_class:] for classinds in inner_inds_by_class])
yield np.sort(trainind[ridgeind]), np.sort(trainind[subind])
ridgeind = np.concatenate(list(zip(*[classinds[:-ntrain_per_class:-1] for classinds in inner_inds_by_class])))
subind = np.concatenate([classinds[-ntrain_per_class::-1] for classinds in inner_inds_by_class])
yield np.sort(trainind[ridgeind]), np.sort(trainind[subind])
def combine_xval_folds(acc_fold):
'''Combine CTD cross-validation accuracies by averaging them
Parameters
----------
acc_fold : list of np.array<bins * bins> - The CTD accuracy matrices of all
the cross-validation folds
Returns
-------
np.array<bins * bins> - The averaged CTD accuracy
'''
return np.stack(acc_fold).mean(0)
def get_acc_mean(acc):
'''Averages all accuracy points in a CTD accuracy matrix
Parameters
----------
acc : np.array<bins * bins> - The CTD accuracy matrix
Returns
-------
float - The stability score
'''
return acc.mean()
def gaussian_func(x, mu, sigma, a):
'''A gaussian function
Parameters
----------
x : np.array - the x values to feed the function
mu : float - the mean of the gaussian
sigma : float - the standard deviation of the gaussian
a : float - a scaling coefficient
Returns
-------
The transformed values in a np.array for each value in x.
'''
b = .25
return a * np.exp(-(x-mu)**2/(2*sigma**2)) + b
def get_CT_score(acc, bounds, dstraining=None):
'''Get the "locality" score from a CTD accuracy
Fits a gaussian on each vector formed by training at a time bin and testing
at all time bins. Calculates the ratio between the maximum of the gaussian
divided by its standard deviation. Then averages all the ratios to get a
locality score.
Parameters
----------
acc : np.array<bins * bins> - The accuracy matrix of a CTD
bounds : a 2-element tuple of 3-element np.array - the bounds for
gaussian fitting for the locality score, e.g.
# mu sigma a
np.array([0, 2, 0]), # lower bounds
np.array([0, np.inf, 1])) # upper bounds
dstraining : int - if the CTD was trained on a subset of the time bins.
Every 'dstraining' bins have been selected for a down sampled training.
Returns
-------
Locality score
'''
if dstraining is None:
dstraining = 1
opted = []
nbinstrain, nbinstest = acc.shape
x = np.arange(nbinstest)
scores = np.empty(nbinstrain)
for ibintrain in range(nbinstrain):
data = acc[ibintrain, :]
data[data < .25] = .25
ibintest = dstraining - 1 + ibintrain * dstraining
params0 = [ibintest, 10, .5]
bounds[0][0] = np.max((ibintest - 5, 0))
bounds[1][0] = np.min((ibintest + 5, nbinstest))
try:
optparams = curve_fit(gaussian_func, x, data, params0, bounds=bounds)[0]
except RuntimeError:
optparams = [0, 0, 0]
scores[ibintrain] = 0
else:
max_val = np.max(gaussian_func(x, *optparams))
scores[ibintrain] = (max_val - .25) / optparams[1] * 1000
opted.append(optparams)
return np.mean(scores)
###############################################################################
################################## VECTORIZED #################################
###############################################################################
def vectorized_xval_CTD(X, y, population=None, permseed=None, subspace=True,
alpha=1, mask=None, dstraining=None):
'''Cross-validation of vectorized cross-temporal decoding
Cross-validation using a custom generator to ensure that the number of
trials in each class is identical.
Parameters
----------
X : np.array<trials * bins * neurons> - data
y : np.array<ntrials> - targets
population : np.array of int - the indices of the neurons included
permseed : int - a seed for permutation testing
subspace : bool - whether to use a subspace or not
alpha : float - the ridge parameter
mask : np.array<nbins> of bool - which bins to take to build the subspace
dstraining : int - Every 'dstraining' bins will be selected for a down
sampled training
Returns
-------
accuracy : np.array<bins * bins> - the CTD accuracy averaged across folds
of the cross-validation
'''
acc_fold = []
if subspace:
nestedxval = NestedXval(n_outer_splits=5)
for testind, trainind, subind in nestedxval.split(y):
acc_fold.append(vectorized_CTD_job(X, y, trainind, testind, subind=subind, population=population,
permseed=permseed, alpha=alpha, mask=mask, dstraining=dstraining)[0])
else:
kfold = KFold(n_splits=5)
for trainind, testind in kfold.split(y):
acc_fold.append(vectorized_CTD_job(X, y, trainind, testind, population=population, permseed=permseed,
alpha=alpha, mask=mask, dstraining=dstraining)[0])
accuracy = combine_xval_folds(acc_fold)
return accuracy
def vectorized_sub_split(X, y, restind, testind,
population=None, permseed=None, subspace=True, alpha=1, mask=None):
'''Cross-validation of vectorized cross-temporal decoding (for testing)
Cross-validation of CTD with pre-defined testing trials. Used for testing,
when the testing trials have already been set aside and only the remaining
trials must be split into training and subspace trials
Parameters
----------
X : np.array<trials * bins * neurons> - data
y : np.array<ntrials> - targets
restind : np.array - The indices of all the trials except the testing trials
testind : np.array - The indices of the testing trials
population : np.array of int - the indices of the neurons included
permseed : int - a seed for permutation testing
subspace : bool - whether to use a subspace or not
alpha : float - the ridge parameter
mask : np.array<nbins> of bool - which bins to take to build the subspace
Returns
-------
accuracy : np.array<bins * bins> - the CTD accuracy averaged across folds
of the cross-validation
'''
if subspace:
acc_split = []
for ridgeind, subind in sub_split(y, restind):
acc_split.append(vectorized_CTD_job(X, y, ridgeind, testind, subind=subind,
population=population, permseed=permseed, alpha=alpha, mask=mask)[0])
accuracy = combine_xval_folds(acc_split)
else:
accuracy = vectorized_CTD_job(X, y, restind, testind,
population=population, permseed=permseed, alpha=alpha, mask=mask)[0]
return accuracy
def vectorized_CTD_job(X, y, trainind, testind,
population=None, **kwargs):
'''Calling vectorized cross-temporal decoding with a given ensemble
Parameters
----------
X : np.array<trials * bins * neurons> - data
y : np.array<ntrials> - targets
trainind : np.array - The indices of the training trials
testind : np.array - The indices of the testing trials
population : np.array of int - the indices of the neurons included
**kwargs : keyword arguments for function 'vectorized_CTD'
Returns
-------
accuracy : np.array<bins * bins> - The CTD matrix accuracy
testout : np.array<bins * bins * test trials> - The output of the
classifier for each pair of train and test bins, for each trial
'''
if population is None:
population = np.arange(X.shape[-1])
newX = X[..., population]
return vectorized_CTD(newX, y, trainind, testind, **kwargs)
def vectorized_CTD(X, y, trainind, testind,
alpha=1, subind=None, mask=None, permseed=None, dstraining=None):
'''Vectorized cross-temporal decoding
This is a vectorized version of the cross-temporal decoding algorithm. The
six decoders (in a one vs one scheme) are trained simultaneously thanks to
vectorization which considerably speeds up computations. Unfortunately it
makes the code less readable. The decoding algorithm was inspired by scikit
learn's implementation of ridge regression. Note that to be able to
vectorize training and testing, each class must have the same number of
training and testing trials.
Parameters
----------
X : np.array<trials * bins * neurons> - data
y : np.array<ntrials> - targets
trainind : np.array
The indices of the training trials
testind : np.array
The indices of the testing trials
alpha : float - the ridge parameter
subind : np.array
The indices of trials used to define the subspace. If not None, a
subspace will be defined
mask : np.array<nbins> of bool - which bins to take to build the subspace
permseed : int - a seed for permutation testing, only the training trials
are shuffled
dstraining : int
Every 'dstraining' bins will be selected for a down sampled training
Returns
-------
accuracy : np.array<bins * bins> - The CTD matrix accuracy
testout : np.array<bins * bins * test trials> - The output of the
classifier for each pair of train and test bins, for each trial
'''
subspace = bool(subind is not None)
if dstraining is None:
dstraining = 1
ntrials, nbins, _ = X.shape
if mask is None:
mask = range(nbins)
labels = np.unique(y)
nclasses = len(labels)
Xtrain, Xtest = X[trainind], X[testind]
ytrain, ytest = y[trainind], y[testind]
if permseed is not None:
np.random.seed(permseed)
np.random.shuffle(ytrain)
if dstraining is not None:
Xtrain = Xtrain[:, dstraining-1::dstraining]
nbinstrain = Xtrain.shape[1]
else:
nbinstrain = nbins
Xtrain = Xtrain.transpose((1, 0, 2))
if subspace:
ysub = y[subind]
Xsub = X[:, mask][subind].mean(1) # Averaging over time bins
Xsub = np.stack([Xsub[ysub == label].mean(0) for label in labels])
subspace = PCA()
subspace.fit(Xsub)
Xtrain = (Xtrain - subspace.mean_[None, None, :]) @ subspace.components_.T[None, ...]
# We need to have the exact same number of trials for each class
_, traincounts = np.unique(ytrain, return_counts=True)
mintrials = np.min(traincounts)
if not np.all(traincounts[0] == traincounts):
mintrials = np.min(traincounts)
keptind = []
for iclass, count in enumerate(traincounts):
if count > mintrials:
inds = np.where(ytrain == labels[iclass])[0][:-(count-mintrials)]
else:
inds = np.where(ytrain == labels[iclass])[0]
keptind.append(inds)
keptind = np.concatenate(keptind)
else:
keptind = np.arange(len(ytrain))
ytrain_cut = ytrain[keptind]
Xtrain_cut = Xtrain[:, keptind]
nestimators = (nclasses * (nclasses - 1)) // 2
nsamples = mintrials * 2
nfeatures = Xtrain_cut.shape[-1]
ytrain_ =
|
np.empty((nestimators, nsamples))
|
numpy.empty
|
import unittest
import numpy as np
class PerceptronTest(unittest.TestCase):
def AND(self, x1, x2):
w1, w2, theta = .5, .5, .7
tmp = x1 * w1 + x2 * w2
if tmp <= theta:
return 0
elif tmp > theta:
return 1
def AND2(self, x1, x2):
x = np.array([x1, x2])
w = np.array([.5, .5])
b = -.7
tmp = np.sum(x * w) + b
if tmp <= 0:
return 0
elif tmp > 0:
return 1
def OR(self, x1, x2):
x = np.array([x1, x2])
w = np.array([.5, .5])
b = -.1
tmp = np.sum(x * w) + b
if tmp <= 0:
return 0
elif tmp > 0:
return 1
def NAND(self, x1, x2):
x = np.array([x1, x2])
w = np.array([-.5, -.5])
b = .7
tmp = np.sum(x * w) + b
if tmp <= 0:
return 0
elif tmp > 0:
return 1
def XOR(self, x1, x2):
s1 = self.OR(x1, x2)
s2 = self.NAND(x1, x2)
return self.AND(s1, s2)
def test_gate(self):
self.assertEqual(0, self.AND(0, 0))
self.assertEqual(0, self.AND(0, 1))
self.assertEqual(0, self.AND(1, 0))
self.assertEqual(1, self.AND(1, 1))
self.assertEqual(0, self.AND2(0, 0))
self.assertEqual(0, self.AND2(0, 1))
self.assertEqual(0, self.AND2(1, 0))
self.assertEqual(1, self.AND2(1, 1))
self.assertEqual(0, self.OR(0, 0))
self.assertEqual(1, self.OR(0, 1))
self.assertEqual(1, self.OR(1, 0))
self.assertEqual(1, self.OR(1, 1))
self.assertEqual(1, self.NAND(0, 0))
self.assertEqual(1, self.NAND(0, 1))
self.assertEqual(1, self.NAND(1, 0))
self.assertEqual(0, self.NAND(1, 1))
self.assertEqual(0, self.XOR(0, 0))
self.assertEqual(1, self.XOR(0, 1))
self.assertEqual(1, self.XOR(1, 0))
self.assertEqual(0, self.XOR(1, 1))
def test_wb(self):
x = np.array([0, 1])
w = np.array([.5, .5])
b = -.7
print(w * x)
print(
|
np.sum(w * x)
|
numpy.sum
|
#!/usr/bin/env python3
# -*- coding: utf-8 -*-
###
# Name: <NAME>
# Student ID: 002270716
# Email: <EMAIL>
# Course: PHYS220/MATH220/CPSC220 Fall 2018
# Assignment: CW09
###
###
# General Commends:
###
import numpy as np
import matplotlib.pyplot as plt
def gradient(x):
"""
Computes the differential operator for a given set of points x. This formulation uses the definition of gradient as a matrix, using matrix multiplication. The program starts by initializing the array,
filling all squares with 0's. The rows are then defined based on where in x they fall. The front and back edges use the forward and backward difference respectively, while points in the middle are defined
by the center difference.
"""
dx = x[1] - x[0]
newGrad = (((np.tri(len(x), len(x), 0) - np.tri(len(x), len(x), 1))) + (np.tri(len(x), len(x), -1) - np.tri(len(x), len(x), -2))) / (2*dx)
newGrad[0][0] = -1 / dx
newGrad[1][0] = 1 / dx
newGrad[-1][-1] = 1 / dx
newGrad[-2][-1] = -1 / dx
return newGrad
def plot(x, func, funcName, gradName):
"""
Graphs the given function as well as its derivative, using the gradient defined above. Functions are graphed in black, while their derivative is graphed in red. Both have their functions written on a
legend in the top right hand corner. Both functions are graphed over some range "x". For the titles, .format was not used as it appears to cause issues with brackets used in latex formatting.
"""
f = plt.figure(figsize=(16,12))
a = plt.axes()
funcGen =
|
np.vectorize(func)
|
numpy.vectorize
|
import numpy as np
class HMC():
def __init__(self, log_prob, grad_log_prob, invmetric_diag=None):
self.log_prob, self.grad_log_prob = log_prob, grad_log_prob
self.V = lambda x : self.log_prob(x)*-1.
#self.V_g = lambda x : self.grad_log_prob(x)*-1.
self.leapcount, self.Vgcount, self.Hcount = 0, 0, 0
if invmetric_diag is None: self.invmetric_diag = 1.
else: self.invmetric_diag = invmetric_diag
self.metricstd = self.invmetric_diag**-0.5
self.KE = lambda p: 0.5*(p**2 * self.invmetric_diag).sum()
self.KE_g = lambda p: p * self.invmetric_diag
def V_g(self, x):
self.Vgcount += 1
return self.grad_log_prob(x)*-1.
def unit_norm_KE(self, p):
return 0.5 * (p**2).sum()
def unit_norm_KE_g(self, p):
return p
def H(self, q,p):
self.Hcount += 1
return self.V(q) + self.KE(p)
def leapfrog(self, q, p, N, step_size):
self.leapcount += 1
q0, p0 = q, p
try:
p = p - 0.5*step_size * self.V_g(q)
for i in range(N-1):
q = q + step_size * self.KE_g(p)
p = p - step_size * self.V_g(q)
q = q + step_size * self.KE_g(p)
p = p - 0.5*step_size * self.V_g(q)
return q, p
except Exception as e:
print(e)
return q0, p0
def leapfrog1(self, q, p, step_size, Vgq=None): #This needs to be optimized to not estimate V_g again and again
self.leapcount += 1
q0, p0 = q, p
try:
if Vgq is None: Vgq = self.V_g(q)
p = p - 0.5*step_size * Vgq
q = q + step_size * self.KE_g(p)
p = p - 0.5*step_size * self.V_g(q)
return q, p, Vgq
except Exception as e:
print(e)
return q0, p0, Vgq
def metropolis(self, qp0, qp1):
q0, p0 = qp0
q1, p1 = qp1
H0 = self.H(q0, p0)
H1 = self.H(q1, p1)
prob = np.exp(H0 - H1)
#prob = min(1., np.exp(H0 - H1))
if np.isnan(prob) or np.isinf(prob) or (q0-q1).sum()==0:
return q0, p0, 2., [H0, H1]
elif np.random.uniform(0., 1., size=1) > min(1., prob):
return q0, p0, 0., [H0, H1]
else: return q1, p1, 1., [H0, H1]
def hmc_step(self, q, N, step_size):
'''Single hmc iteration
Parameters:
----------
q: initial position
N: number of leapfrog steps
step_size: step size for leapfrog iteration
Returns:
--------
A tuple of-
q
p
accepted (0/1/2)
acceptance probability
list of [Hcounts, Vcounts, nleapfrogs]
'''
self.leapcount, self.Vgcount, self.Hcount = 0, 0, 0
p = np.random.normal(size=q.size).reshape(q.shape) * self.metricstd
q1, p1 = self.leapfrog(q, p, N, step_size)
q, p, accepted, prob = self.metropolis([q, p], [q1, p1])
return q, p, accepted, prob, [self.Hcount, self.Vgcount, self.leapcount]
######################
class AdHMC_eps0(HMC):
def __init__(self, log_prob, grad_log_prob, invmetric_diag=None):
super().__init__(log_prob, grad_log_prob, invmetric_diag)
def get_stepsize(self, q0, p0, smin=0.01, smax=1.0, ntry=20, logspace=True, nsteps=1, eps=None):
H0 = self.H(q0, p0)
Hs = np.zeros(ntry)
if logspace: steps = np.logspace(np.log10(smin), np.log10(smax), ntry)
else: steps = np.linspace(smin, smax, ntry)
pwts = steps.copy()**0.5 #np.linspace(0.9, 1.1, steps.size)
for iss, ss in enumerate(steps):
#nsteps = int(steps.max()/ss)+1
q1, p1 = self.leapfrog(q0, p0, nsteps, ss)
Hs[iss] = self.H(q1, p1)
pp = np.exp(H0 - Hs) * pwts
pp[np.isnan(pp)] = 0
pp[np.isinf(pp)] = 0
pp /= pp.sum()
cdf = np.cumsum(pp)
if eps is None:
sx = np.random.uniform(low=cdf.min())
isx = np.where(sx > cdf)[0][-1]
sx2 = np.random.uniform(steps[isx], steps[isx+1])
prob = pp[isx+1] # * 1/(steps[isx+1]-steps[isx+1])
return sx2, pp[isx+1]
else:
prob = pp[np.where(steps > eps)[0][0]]
return prob
def hmc_step(self, q0, Nleap, smin=0.01, smax=1.0, Tint=0, ntry=10, nsteps=1):
'''Single hmc iteration
Parameters:
----------
q: initial position
N: number of leapfrog steps
step_size: step size for leapfrog iteration
smin: Minimum allowed step size
smin: Maximum allowed step size
Tint: Time of integration
ntry: Number of points to try for estimating first step size
nsteps: Number of steps per try for estimating first step size
Returns:
--------
A tuple of-
q
p
accepted (0/1/2)
acceptance probability
array of [pfactor denominator, pfactor numberator, stepsize]
list of [Hcounts, Vcounts, nleapfrogs]
'''
self.leapcount, self.Vgcount, self.Hcount = 0, 0, 0
p0 = np.random.normal(size=q0.size).reshape(q0.shape) * self.metricstd
H0 = self.H(q0, p0)
if (Tint == 0) and (Nleap == 0):
print("Tint and Nleap cannot be both zeros")
import sys
sys.exit()
elif (Tint != 0) and (Nleap != 0):
print("Tint and Nleap both given and are inconsistent")
import sys
sys.exit()
#First step is drawn from a distribution
ss, pf_den = self.get_stepsize(q0, p0, smin, smax, ntry=ntry, nsteps=nsteps)
eps = ss
if Tint == 0: N = Nleap
else: N = int(Tint/eps) + 1
#print("Steps size is %0.2f, and number of steps is %d"%(eps, N))
q1, p1 = self.leapfrog(q0, p0, N, ss)
H1 = self.H(q1, p1)
pb_num = self.get_stepsize(q1, -p1, smin=smin, smax=smax, eps=ss, ntry=ntry, nsteps=nsteps)
hastings_factor = pb_num/pf_den
prob = np.exp(H0 - H1) * hastings_factor
#print("prb, fac, metrop : ", prob, adfac, prob/adfac, pb_num, pf_den)
toret = [[prob, prob/hastings_factor, hastings_factor], np.stack([pf_den, pb_num, eps]), [self.Hcount, self.Vgcount, self.leapcount]]
if np.isnan(prob) or np.isinf(prob) or (q0-q1).sum()==0:
return q0, p0, 2., *toret
elif np.random.uniform(0., 1., size=1) > min(1., prob):
return q0, p0, 0., *toret
else: return q1, p1, 1., *toret
##
######################
class AdHMC(HMC):
def __init__(self, log_prob, grad_log_prob, invmetric_diag=None):
super().__init__(log_prob, grad_log_prob, invmetric_diag)
def get_stepsize(self, q0, p0, smin=0.01, smax=1.0, ntry=10, logspace=True, nsteps=1, eps=None):
H0 = self.H(q0, p0)
Hs = np.zeros(ntry)
if logspace: steps = np.logspace(np.log10(smin), np.log10(smax), ntry)
else: steps = np.linspace(smin, smax, ntry)
pwts = steps.copy()**0.5 #np.linspace(0.9, 1.1, steps.size)
for iss, ss in enumerate(steps):
#nsteps = int(steps.max()/ss)+1
q1, p1 = self.leapfrog(q0, p0, nsteps, ss)
Hs[iss] = self.H(q1, p1)
pp = np.exp(H0 - Hs) * pwts
pp[np.isnan(pp)] = 0
pp[np.isinf(pp)] = 0
pp /= pp.sum()
cdf = np.cumsum(pp)
if eps is None:
sx = np.random.uniform(low=cdf.min())
isx = np.where(sx > cdf)[0][-1]
sx2 = np.random.uniform(steps[isx], steps[isx+1])
prob = pp[isx+1] # * 1/(steps[isx+1]-steps[isx+1])
return sx2, pp[isx+1]
else:
prob = pp[np.where(steps > eps)[0][0]]
return prob
def hmc_step(self, q0, Nleap=100, nleap=10, ratios= [1/np.sqrt(2), np.sqrt(2)], pwts0 = [1., 1.], smin=0.01, smax=1.0, ntry_eps0=10, nsteps_eps0=1, logeps=True, verbose=False):
'''
Parameters:
----------
q: initial position
Nleap: number of leapfrog steps
nleap: number of leapfrog steps to adapt step size
smin: Minimum allowed step size
smin: Maximum allowed step size
ratios: ratio to change step size with after nleap steps- expected in INCREASING order
ntry_eps0: Number of points to try for estimating first step size
nsteps_eps0: Number of steps per try for estimating first step size
Returns:
--------
A tuple of-
q
p
accepted (0/1/2)
list of probabiliies [acc_prob, acc_prob/hastings_factor, hastings_factor]
array of checks [pfactor denominator, pfactor numberator, stepsize]
list of counts [Hcounts, Vcounts, nleapfrogs]
'''
#normprob is not implemented
self.leapcount, self.Vgcount, self.Hcount = 0, 0, 0
p0 = np.random.normal(size=q0.size).reshape(q0.shape) * self.metricstd
N = int(Nleap//nleap)
#First step is drawn from a distribution
eps, pf_den, pb_num =
|
np.zeros(N)
|
numpy.zeros
|
# Copyright (c) 2020-2021 by Fraunhofer Institute for Energy Economics
# and Energy System Technology (IEE), Kassel, and University of Kassel. All rights reserved.
# Use of this source code is governed by a BSD-style license that can be found in the LICENSE file.
import os
import numpy as np
import pandas as pd
from scipy.interpolate import interp1d
from pandapipes import pp_dir
from pandapower.io_utils import JSONSerializableClass
try:
import pandaplan.core.pplog as logging
except ImportError:
import logging
logger = logging.getLogger(__name__)
class Fluid(JSONSerializableClass):
"""
"""
def __init__(self, name, fluid_type, **kwargs):
"""
:param name:
:type name:
:param fluid_type:
:type fluid_type:
:param kwargs:
:type kwargs:
"""
super(Fluid, self).__init__()
self.name = name
if not isinstance(fluid_type, str) or fluid_type.lower() not in ["gas", "liquid"]:
logger.warning("The fluid %s has the fluid type %s which might cause problems in the "
"pipeflow calculation, as it expects either 'gas' or 'liquid'."
% (name, fluid_type))
self.fluid_type = fluid_type.lower()
self.is_gas = self.fluid_type == "gas"
self.all_properties = kwargs
for prop_name, prop in self.all_properties.items():
if not isinstance(prop, FluidProperty):
logger.warning("The property %s was not defined as a fluid property. This might "
"cause problems when trying to ask for values." % prop_name)
def __repr__(self):
"""
Definition of fluid representation in the console.
:return: representation of fluid in the console
:rtype: str
"""
r = "Fluid %s (%s) with properties:" % (self.name, self.fluid_type)
for key in self.all_properties.keys():
r += "\n - %s (%s)" % (key, self.all_properties[key].__class__.__name__[13:])
return r
def add_property(self, property_name, prop, overwrite=True, warn_on_duplicates=True):
"""
This function adds a new property.
:param property_name: Name of the new property
:type property_name: str
:param prop: Values for the property, for example a curve or just a constant value
:type prop: pandapipes.FluidProperty
:param overwrite: True if existing property with the same name shall be overwritten
:type overwrite: bool
:param warn_on_duplicates: True, if a warning of properties with the same name should be
returned
:type warn_on_duplicates: bool
:Example:
>>> fluid.add_property('water_density', pandapipes.FluidPropertyConstant(998.2061),
overwrite=True, warn_on_duplicates=False)
"""
if property_name in self.all_properties:
if warn_on_duplicates:
ow_string = "It will be overwritten." if overwrite else "It will not be replaced."
logger.warning("The property %s already exists. %s" % (property_name, ow_string))
if not overwrite:
return
self.all_properties[property_name] = prop
def get_property(self, property_name, *at_values):
"""
This function returns the value of the requested property.
:param property_name: Name of the searched property
:type property_name: str
:param at_values: Value for which the property should be returned
:type at_values:
:return: Returns property at the certain value
:rtype: pandapipes.FluidProperty
"""
if property_name not in self.all_properties:
raise UserWarning("The property %s was not defined for the fluid %s"
% (property_name, self.name))
return self.all_properties[property_name].get_at_value(*at_values)
def get_density(self, temperature):
"""
This function returns the density at a certain temperature.
:param temperature: Temperature at which the density is queried
:type temperature: float
:return: Density at the required temperature
"""
return self.get_property("density", temperature)
def get_viscosity(self, temperature):
"""
This function returns the viscosity at a certain temperature.
:param temperature: Temperature at which the viscosity is queried
:type temperature: float
:return: Viscosity at the required temperature
"""
return self.get_property("viscosity", temperature)
def get_heat_capacity(self, temperature):
"""
This function returns the heat capacity at a certain temperature.
:param temperature: Temperature at which the heat capacity is queried
:type temperature: float
:return: Heat capacity at the required temperature
"""
return self.get_property("heat_capacity", temperature)
def get_molar_mass(self):
"""
This function returns the molar mass.
:return: molar mass
"""
return self.get_property("molar_mass")
def get_compressibility(self, p_bar):
"""
This function returns the compressibility at a certain pressure.
:param p_bar: pressure at which the compressibility is queried
:type p_bar: float or array of floats
:return: compressibility at the required pressure
"""
return self.get_property("compressibility", p_bar)
def get_der_compressibility(self):
"""
This function returns the derivative of the compressibility with respect to pressure.
:return: derivative of the compressibility
"""
return self.get_property("der_compressibility")
class FluidProperty(JSONSerializableClass):
"""
Property Base Class
"""
def __init__(self):
"""
"""
super().__init__()
def get_at_value(self, *args):
"""
:param args:
:type args:
:return:
:rtype:
"""
raise NotImplementedError("Please implement a proper fluid property!")
def get_at_integral_value(self, *args):
"""
:param args:
:type args:
:return:
:rtype:
"""
raise NotImplementedError("Please implement a proper fluid property!")
class FluidPropertyInterExtra(FluidProperty):
"""
Creates Property with interpolated or extrapolated values.
"""
json_excludes = JSONSerializableClass.json_excludes + ["prop_getter"]
prop_getter_entries = {"x": "x", "y": "y", "_fill_value_orig": "fill_value"}
def __init__(self, x_values, y_values, method="interpolate_extrapolate"):
"""
:param x_values:
:type x_values:
:param y_values:
:type y_values:
:param method:
:type method:
"""
super(FluidPropertyInterExtra, self).__init__()
if method.lower() == "interpolate_extrapolate":
self.prop_getter = interp1d(x_values, y_values, fill_value="extrapolate")
else:
self.prop_getter = interp1d(x_values, y_values)
def get_at_value(self, arg):
"""
:param arg: Name of the property and one or more values (x-values) for which the y-values \
of the property are to be displayed
:type arg: str, float or array
:return: y-value/s
:rtype: float, array
"""
return self.prop_getter(arg)
def get_at_integral_value(self, upper_limit_arg, lower_limit_arg):
"""
:param arg: one or more values of upper and lower limit values for which the function \
of the property should calculate the integral for
:type arg: float or list-like objects
:return: integral between the limits
:rtype: float, array
:Example:
>>> comp_fact = get_fluid(net).all_properties["heat_capacity"].get_at_integral_value(t_upper_k, t_lower_k)
"""
mean = (self.prop_getter(upper_limit_arg) + self.prop_getter(upper_limit_arg)) / 2
return mean * (upper_limit_arg-lower_limit_arg)
@classmethod
def from_path(cls, path, method="interpolate_extrapolate"):
"""
Reads a text file with temperature values in the first column and property values in
second column.
:param path: Target path of the txt file
:type path: str
:param method: Method with which the values are to be interpolated
:type method: str
:return: interpolated values
:rtype: pandapipes.FluidProperty
"""
values = np.loadtxt(path)
return cls(values[:, 0], values[:, 1], method=method)
def to_dict(self):
d = super(FluidPropertyInterExtra, self).to_dict()
d.update({k: self.prop_getter.__dict__[k] for k in self.prop_getter_entries.keys()})
# d.update({"x_values": self.prop_getter.x, "y_values": self.prop_getter.y,
# "method": "interpolate_extrapolate"
# if self.prop_getter.fill_value == "extrapolate" else None})
return d
@classmethod
def from_dict(cls, d):
obj = JSONSerializableClass.__new__(cls)
d2 = {cls.prop_getter_entries[k]: v for k, v in d.items()
if k in cls.prop_getter_entries.keys()}
d3 = {k: v for k, v in d.items() if k not in cls.prop_getter_entries.keys()}
d3["prop_getter"] = interp1d(**d2)
obj.__dict__.update(d3)
return obj
class FluidPropertyConstant(FluidProperty):
"""
Creates Property with a constant value.
"""
def __init__(self, value, warn_dependent_variables=False):
"""
:param value:
:type value:
"""
super(FluidPropertyConstant, self).__init__()
self.value = value
self.warn_dependent_variables = warn_dependent_variables
def get_at_value(self, *args):
"""
:param args: Name of the property
:type args: str
:return: Value of the property
:rtype: float
:Example:
>>> heat_capacity = get_fluid(net).all_properties["heat_capacity"].get_at_value(293.15)
"""
if len(args) > 1:
raise UserWarning('Please define either none or an array-like argument')
elif len(args) == 1:
if self.warn_dependent_variables:
logger.warning('Constant property received several input variables, although it is'
'independent of these')
output = np.array([self.value]) * np.ones(len(args[0]))
else:
output = np.array([self.value])
return output
def get_at_integral_value(self, upper_limit_arg, lower_limit_arg):
"""
:param arg: one or more values of upper and lower limit values for which the function \
of the property should calculate the integral for
:type arg: float or list-like objects
:return: integral between the limits
:rtype: float, array
:Example:
>>> comp_fact = get_fluid(net).all_properties["heat_capacity"].get_at_integral_value(t_upper_k, t_lower_k)
"""
if isinstance(upper_limit_arg, pd.Series):
ul = self.value * upper_limit_arg.values
else:
ul = self.value * np.array(upper_limit_arg)
if isinstance(lower_limit_arg, pd.Series):
ll = self.value * lower_limit_arg.values
else:
ll = self.value * np.array(lower_limit_arg)
return ul - ll
@classmethod
def from_path(cls, path):
"""
Reads a text file with temperature values in the first column and property values in
second column.
:param path:
:type path:
:param method:
:type method:
:return:
:rtype:
"""
value = np.loadtxt(path).item()
return cls(value)
@classmethod
def from_dict(cls, d):
obj = super().from_dict(d)
if "warn_dependent_variables" not in obj.__dict__.keys():
obj.__dict__["warn_dependent_variables"] = False
return obj
class FluidPropertyLinear(FluidProperty):
"""
Creates Property with a linear course.
"""
def __init__(self, slope, offset):
"""
:param slope:
:type slope:
:param offset:
:type offset:
"""
super(FluidPropertyLinear, self).__init__()
self.slope = slope
self.offset = offset
def get_at_value(self, arg):
"""
:param arg: Name of the property and one or more values (x-values) for which the function \
of the property should be calculated
:type arg: str, float or array
:return: y-value or function values
:rtype: float, array
:Example:
>>> comp_fact = get_fluid(net).all_properties["compressibility"].get_at_value(p_bar)
"""
if isinstance(arg, pd.Series):
return self.offset + self.slope * arg.values
else:
return self.offset + self.slope * np.array(arg)
def get_at_integral_value(self, upper_limit_arg, lower_limit_arg):
"""
:param arg: one or more values of upper and lower limit values for which the function \
of the property should calculate the integral for
:type arg: float or list-like objects
:return: integral between the limits
:rtype: float, array
:Example:
>>> comp_fact = get_fluid(net).all_properties["heat_capacity"].get_at_integral_value(t_upper_k, t_lower_k)
"""
if isinstance(upper_limit_arg, pd.Series):
ul = self.offset * upper_limit_arg.values + 0.5 * self.slope * np.power(upper_limit_arg.values, 2)
else:
ul = self.offset *
|
np.array(upper_limit_arg)
|
numpy.array
|
#!/usr/bin/env python3
# -*- coding: utf-8 -*-
"""
Created on Sun May 20 18:40:04 2018
@author: alejandro
"""
import sys
sys.path.append('/usr/local/lib/python2.7/dist-packages/')
import pygame
import numpy as np
import sys
from freenect import sync_get_depth as get_depth
#funcion asigna la superficie de colores
def make_gamma():
num_pix = 2048 # valores posibles de profundidad NO JUGAR
npf = float(num_pix)
_gamma = np.empty((num_pix, 3), dtype=np.uint16)
for i in xrange(num_pix):
v = i / npf
v = pow(v, 3) * 6
pval = int(v * 35 * 256-500) # cambiar profundidad -----------40--------------
lb = pval & 0xf1
pval >>= 8
if pval == 0:
a = np.array([255, 255 - lb, 255 - lb], dtype=np.uint8)
elif pval == 1:
a = np.array([255, lb, 0], dtype=np.uint8)
elif pval == 2:
a = np.array([255 - lb, lb, 0], dtype=np.uint8)
elif pval == 3:
a =
|
np.array([255 - lb, 255, 0], dtype=np.uint8)
|
numpy.array
|
#!/usr/bin/env python
# just testing basic parallelization that will be used in the actual project
from __future__ import division,unicode_literals
from future.builtins import map,zip, range
import numpy as np
import itertools as it
# setup proper logging
import logging
logger = logging.getLogger('psnobfit')
logger.setLevel(logging.INFO)
# logger.setLevel(logging.DEBUG)
log_stream_handler = logging.StreamHandler()
log_stream_handler.setLevel(logging.DEBUG)
log_formatter = logging.Formatter('%(asctime)s %(name)s/%(levelname)-9s %(message)s')
log_stream_handler.setFormatter(log_formatter)
logger.addHandler(log_stream_handler)
del logging, log_stream_handler, log_formatter
# read cmd line options
from optparse import OptionParser
opt_parser = OptionParser()
opt_parser.add_option("-p", "--profile", dest="client_profile", default="unissh", action="store_const",
help="the profile to use for ipython.parallel")
options, args = opt_parser.parse_args()
# START: create remote evaluators and a few (or one) special one for #
# generating new points
logger.info("init")
from IPython.parallel import Client, require
c = Client(profile=options.client_profile)
c.clear() # clears remote engines
c.purge_results('all') # all results are memorized in the hub
if len(c.ids) < 2:
raise Exception('I need at least 2 clients.')
nbGens = min(1, len(c.ids) - 1)
generators = c.load_balanced_view(c.ids[:nbGens])
evaluators = c.load_balanced_view(c.ids[nbGens:])
# MAX number of tasks in total
MAX = 5000
# length of test data, sent over the wire
DIMSIZE = 10
# when adding machines, this is the number of additional tasks
# beyond the number of free machines
new_extra = DIMSIZE
# import some packages (also locally)
with c[:].sync_imports():
from IPython.utils.timing import time # time.time & time.clock for cpu time
#import time
from random import random
from numpy import pi, sum
import numpy
import math
# the actual function
def func_one(tid, data):
return tid, 1
def func_sum(tid, data):
'x is either a number or a list/vector of numbers'
time.sleep(math.log(1 + random()))
return tid, numpy.sum(data)
def func_eval(tid, data):
np = numpy
data = data * numpy.pi / 2
v = np.multiply(np.cos(numpy.pi + data), np.sin(data + numpy.pi / 2))
v = np.exp(np.linalg.norm(v - 1, 1) / len(data))
#s = np.sin(data[::2] + numpy.pi / 2)
#c = np.cos(data[1::2])
# v += np.sum(s) + np.sum(c) #np.append(s,c))
# time.sleep(1e-3)
#time.sleep(1e-2 + math.log(1 + random()))
return tid, v
func = func_eval
# some stats
added = 0
queue_size = 0
added = 0
nb_finished = 0
nb_generated = 0
loops = 0
tasks_added = 0
cum_sum = 0
best_x = None
best_obj = numpy.infty
last_best = best_obj
def status():
global last_best
s = '*' if last_best != best_obj else ' '
logger.info(
"pend %4d | + %2d | tot: %4d | finished: %4d | gen: %3d | best_obj: %.10f %s" %
(queue_size, new, added, nb_finished, nb_generated, best_obj, s))
last_best = best_obj
logger.info("start")
start_time = time.time()
# pending is the set of jobs we are expecting in each loop
pending = set([])
pending_generators = set([])
new_points = []
# collects all returns
results = []
allx = dict() # store all x vectors
def gen_points(new, DIMSIZE, cur_best_res=None, cur_best_x=None):
'''
generates @new new points, depends on results and allx
'''
np = numpy
#lambda rp : 10 * (np.random.rand(DIMSIZE) )
FACT = 3
OFF = 0
if np.random.random() < .2 or not cur_best_res:
return np.array([FACT * (np.random.rand(DIMSIZE) + OFF) for _ in range(new)])
# better local value new best point
ret = []
for i in range(new):
rv = (np.random.rand(DIMSIZE) - .5) / 5
# make it sparse
sp = np.random.rand(DIMSIZE) < .9
rv[sp] = 0
#import scipy
#rv = scipy.sparse.rand(DIMSIZE, 1, 0.1)
ret.append(np.minimum(2,
|
np.maximum(0, rv + cur_best_x)
|
numpy.maximum
|
# Copyright 2021-2022 NVIDIA Corporation
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
#
from __future__ import annotations
from typing import TYPE_CHECKING, Any, Optional
import numpy as np
from .. import legion
if TYPE_CHECKING:
import numpy.typing as npt
from . import Point
class Transform:
trans: npt.NDArray[np.int64]
def __init__(self, M: int, N: int, eye: bool = True):
"""
A Transform wraps an `legion_transform_{m}x{n}_t` in the Legion C API.
A transform is simply an MxN matrix that can be used to convert Point
objects from one coordinate space to another.
"""
self.M = M
self.N = N
if eye:
self.trans =
|
np.eye(M, N, dtype=np.int64)
|
numpy.eye
|
# Python 3.5
# Script written by <NAME> (<EMAIL>), <NAME> (<EMAIL>), and <NAME> (<EMAIL>)
# VERSION 0.1 - JUNE 2020
#--------TURN OFF MAGMASAT WARNING--------#
import warnings
warnings.filterwarnings("ignore", message="rubicon.objc.ctypes_patch has only been tested ")
warnings.filterwarnings("ignore", message="The handle")
#-----------------IMPORTS-----------------#
import pandas as pd
import numpy as np
import matplotlib as mpl
import matplotlib.pyplot as plt
import matplotlib.font_manager as font_manager
from cycler import cycler
from abc import ABC, abstractmethod
from scipy.optimize import root_scalar
from scipy.optimize import root
from scipy.optimize import minimize
import sys
import sympy
from copy import copy
# import anvil_server
#--------------MELTS preamble---------------#
from thermoengine import equilibrate
# instantiate thermoengine equilibrate MELTS instance
melts = equilibrate.MELTSmodel('1.2.0')
# Suppress phases not required in the melts simulation
phases = melts.get_phase_names()
for phase in phases:
melts.set_phase_inclusion_status({phase: False})
melts.set_phase_inclusion_status({'Fluid': True, 'Liquid': True})
#----------DEFINE SOME CONSTANTS-------------#
oxides = ['SiO2', 'TiO2', 'Al2O3', 'Fe2O3', 'Cr2O3', 'FeO', 'MnO', 'MgO', 'NiO', 'CoO', 'CaO', 'Na2O', 'K2O', 'P2O5',
'H2O', 'CO2']
anhydrous_oxides = ['SiO2', 'TiO2', 'Al2O3', 'Fe2O3', 'Cr2O3', 'FeO', 'MnO', 'MgO', 'NiO', 'CoO', 'CaO', 'Na2O', 'K2O', 'P2O5']
volatiles = ['H2O', 'CO2']
oxideMass = {'SiO2': 28.085+32, 'MgO': 24.305+16, 'FeO': 55.845+16, 'CaO': 40.078+16, 'Al2O3': 2*26.982+16*3, 'Na2O': 22.99*2+16,
'K2O': 39.098*2+16, 'MnO': 54.938+16, 'TiO2': 47.867+32, 'P2O5': 2*30.974+5*16, 'Cr2O3': 51.996*2+3*16,
'NiO': 58.693+16, 'CoO': 28.01+16, 'Fe2O3': 55.845*2+16*3,
'H2O': 18.02, 'CO2': 44.01}
CationNum = {'SiO2': 1, 'MgO': 1, 'FeO': 1, 'CaO': 1, 'Al2O3': 2, 'Na2O': 2,
'K2O': 2, 'MnO': 1, 'TiO2': 1, 'P2O5': 2, 'Cr2O3': 2,
'NiO': 1, 'CoO': 1, 'Fe2O3': 2, 'H2O': 2, 'CO2': 1}
OxygenNum = {'SiO2': 2, 'MgO': 1, 'FeO': 1, 'CaO': 1, 'Al2O3': 3, 'Na2O': 1,
'K2O': 1, 'MnO': 1, 'TiO2': 2, 'P2O5': 5, 'Cr2O3': 3,
'NiO': 1, 'CoO': 1, 'Fe2O3': 3, 'H2O': 1, 'CO2': 2}
CationCharge = {'SiO2': 4, 'MgO': 2, 'FeO': 2, 'CaO': 2, 'Al2O3': 3, 'Na2O': 1,
'K2O': 1, 'MnO': 2, 'TiO2': 4, 'P2O5': 5, 'Cr2O3': 3,
'NiO': 2, 'CoO': 2, 'Fe2O3': 3, 'H2O': 1, 'CO2': 4}
CationMass = {'SiO2': 28.085, 'MgO': 24.305, 'FeO': 55.845, 'CaO': 40.078, 'Al2O3': 26.982, 'Na2O': 22.990,
'K2O': 39.098, 'MnO': 54.938, 'TiO2': 47.867, 'P2O5': 30.974, 'Cr2O3': 51.996,
'NiO': 58.693, 'CoO': 28.01, 'Fe2O3': 55.845, 'H2O': 2, 'CO2': 12.01}
oxides_to_cations = {'SiO2': 'Si', 'MgO': 'Mg', 'FeO': 'Fe', 'CaO': 'Ca', 'Al2O3': 'Al', 'Na2O': 'Na',
'K2O': 'K', 'MnO': 'Mn', 'TiO2': 'Ti', 'P2O5': 'P', 'Cr2O3': 'Cr',
'NiO': 'Ni', 'CoO': 'Co', 'Fe2O3': 'Fe3', 'H2O': 'H', 'CO2': 'C'}
cations_to_oxides = {'Si': 'SiO2', 'Mg': 'MgO', 'Fe': 'FeO', 'Ca': 'CaO', 'Al': 'Al2O3', 'Na': 'Na2O',
'K': 'K2O', 'Mn': 'MnO', 'Ti': 'TiO2', 'P': 'P2O5', 'Cr': 'Cr2O3',
'Ni': 'NiO', 'Co': 'CoO', 'Fe3': 'Fe2O3', 'H': 'H2O', 'C': 'CO2'}
#----------DEFINE SOME EXCEPTIONS--------------#
class Error(Exception):
"""Base class for exceptions in this module."""
pass
class InputError(Error):
"""Exception raised for errors in the input.
Attributes:
expression -- input expression in which the error occurred
message -- explanation of the error
"""
def __init__(self, message):
self.message = message
class SaturationError(Error):
"""Exception raised for errors thrown when a sample does not reach saturation.
Attributes:
expression -- input expression in which the error occurred
message -- explanation of the error
"""
def __init__(self, message):
self.message = message
#----------DEFINE CUSTOM PLOTTING FORMATTING------------#
style = "seaborn-colorblind"
plt.style.use(style)
plt.rcParams["mathtext.default"] = "regular"
plt.rcParams["mathtext.fontset"] = "dejavusans"
mpl.rcParams['patch.linewidth'] = 1
mpl.rcParams['axes.linewidth'] = 1
plt.rcParams['axes.titlesize'] = 20
plt.rcParams['axes.labelsize'] = 18
plt.rcParams['xtick.labelsize'] = 14
plt.rcParams['ytick.labelsize'] = 14
plt.rcParams['legend.fontsize'] = 14
#Define color cycler based on plot style set here
the_rc = plt.style.library[style] #get style formatting set by plt.style.use()
color_list = the_rc['axes.prop_cycle'].by_key()['color'] #list of colors by hex code
color_cyler = the_rc['axes.prop_cycle'] #get the cycler
def printTable(myDict):
""" Pretty print a dictionary (as pandas DataFrame)
Parameters
----------
myDict: dict
A dictionary
Returns
-------
pandas DataFrame
The input dictionary converted to a pandas DataFrame
"""
try:
oxidesum = sum(myDict[oxide] for oxide in oxides)
myDict.update({"Sum oxides": oxidesum})
except:
pass
table = pd.DataFrame([v for v in myDict.values()], columns = ['value'],
index = [k for k in myDict.keys()])
return table
#----------DEFINE SOME UNIVERSAL INFORMATIVE METHODS--------------#
def get_model_names():
"""
Returns all available model names as a list of strings.
"""
model_names = []
for key, value in default_models.items():
model_names.append(key)
return model_names
#----------DEFINE SOME BASIC DATA TRANSFORMATION METHODS-----------#
def mol_to_wtpercent(sample):
"""
Takes in a pandas DataFrame containing multi-sample input or a dictionary containing single-sample input
and returns a pandas DataFrame object with oxide values converted from mole percent to wt percent.
Parameters
----------
oxides: pandas DataFrame object or dictionary
Variable name referring to the pandas DataFrame object that contains user-imported data or a dictionary
for single-sample input.
"""
data = sample
if isinstance(sample, pd.DataFrame):
for key, value in oxideMass.items():
data.loc[:, key] *= value
data["MPOSum"] = sum([data[oxide] for oxide in oxides])
for oxide in oxides:
data.loc[:, oxide] /= data['MPOSum']
data.loc[:, oxide] *= 100
del data['MPOSum']
elif isinstance(sample, dict):
for oxide in oxides:
if oxide in data.keys():
pass
else:
data[oxide] = 0.0
data = {oxide: data[oxide] for oxide in oxides}
for key, value in oxideMass.items():
data.update({key: (data[key] * value)})
MPOSum = sum(data.values())
for key, value in data.items():
data.update({key: 100 * value / MPOSum})
return data
def wtpercentOxides_to_molCations(oxides):
"""Takes in a pandas Series containing major element oxides in wt%, and converts it
to molar proportions of cations (normalised to 1).
Parameters
----------
oxides dict or pandas Series
Major element oxides in wt%.
Returns
-------
dict or pandas Series
Molar proportions of cations, normalised to 1.
"""
molCations = {}
_oxides = oxides.copy()
if type(oxides) == dict:
oxideslist = list(_oxides.keys())
elif type(oxides) == pd.core.series.Series:
oxideslist = list(_oxides.index)
else:
raise InputError("The composition input must be a pandas Series or dictionary.")
for ox in oxideslist:
cation = oxides_to_cations[ox]
molCations[cation] = CationNum[ox]*_oxides[ox]/oxideMass[ox]
if type(oxides) == pd.core.series.Series:
molCations = pd.Series(molCations)
molCations = molCations/molCations.sum()
else:
total = np.sum(list(molCations.values()))
for ox in oxideslist:
cation = oxides_to_cations[ox]
molCations[cation] = molCations[cation]/total
return molCations
def wtpercentOxides_to_molOxides(oxides):
""" Takes in a pandas Series or dict containing major element oxides in wt%, and converts it
to molar proportions (normalised to 1).
Parameters
----------
oxides dict or pandas Series
Major element oxides in wt%
Returns
-------
dict or pandas Series
Molar proportions of major element oxides, normalised to 1.
"""
molOxides = {}
_oxides = oxides.copy()
if type(oxides) == dict or type(oxides) == pd.core.series.Series:
if type(oxides) == dict:
oxideslist = list(oxides.keys())
elif type(oxides) == pd.core.series.Series:
oxideslist = list(oxides.index)
for ox in oxideslist:
molOxides[ox] = _oxides[ox]/oxideMass[ox]
if type(oxides) == pd.core.series.Series:
molOxides = pd.Series(molOxides)
molOxides = molOxides/molOxides.sum()
else:
total = np.sum(list(molOxides.values()))
for ox in oxideslist:
molOxides[ox] = molOxides[ox]/total
return molOxides
elif isinstance(sample, pd.DataFrame):
data = sample
for key, value in oxideMass.items():
data.loc[:, key] /= value
data["MPOSum"] = sum([data[oxide] for oxide in oxides])
for oxide in oxides:
data.loc[:, oxide] /= data['MPOSum']
del data['MPOSum']
return data
else:
raise InputError("The composition input must be a pandas Series or dictionary.")
def wtpercentOxides_to_molSingleO(oxides,exclude_volatiles=False):
""" Takes in a pandas Series containing major element oxides in wt%, and constructs
the chemical formula, on a single oxygen basis.
Parameters
----------
oxides dict or pandas Series
Major element oxides in wt%
Returns
-------
dict or pandas Series
The chemical formula of the composition, on a single oxygen basis. Each element is
a separate entry in the Series.
"""
molCations = {}
_oxides = oxides.copy()
if type(oxides) == dict:
oxideslist = list(oxides.keys())
elif type(oxides) == pd.core.series.Series:
oxideslist = list(oxides.index)
else:
raise InputError("The composition input must be a pandas Series or dictionary.")
total_O = 0.0
for ox in oxideslist:
if exclude_volatiles == False or (ox != 'H2O' and ox != 'CO2'):
cation = oxides_to_cations[ox]
molCations[cation] = CationNum[ox]*oxides[ox]/oxideMass[ox]
total_O += OxygenNum[ox]*oxides[ox]/oxideMass[ox]
if type(oxides) == pd.core.series.Series:
molCations = pd.Series(molCations)
molCations = molCations/total_O
else:
# total = np.sum(list(molCations.values()))
for ox in oxideslist:
if exclude_volatiles == False or (ox != 'H2O' and ox != 'CO2'):
cation = oxides_to_cations[ox]
molCations[cation] = molCations[cation]/total_O
return molCations
def wtpercentOxides_to_formulaWeight(sample,exclude_volatiles=False):
""" Converts major element oxides in wt% to the formula weight (on a 1 oxygen basis).
Parameters
----------
sample dict or pandas Series
Major element oxides in wt%.
exclude_volatiles bool
If True H2O and CO2 will be excluded from the formula weight calculation.
Returns
-------
float
The formula weight of the composition, on a one oxygen basis.
"""
if type(sample) == dict:
_sample = pd.Series(sample.copy())
elif type(sample) != pd.core.series.Series:
raise InputError("The composition input must be a pandas Series or dictionary.")
else:
_sample = sample.copy()
cations = wtpercentOxides_to_molSingleO(_sample,exclude_volatiles=exclude_volatiles)
if type(cations) != dict:
cations = dict(cations)
# if exclude_volatiles == True:
# if 'C' in cations:
# cations.pop('C')
# if 'H' in cations:
# cations.pop('H')
# newsum = 0
# for cation in cations:
# newsum += OxygenNum[cations_to_oxides[cation]]
# for cation in cations:
# cations[cation] = cations[cation]/newsum
FW = 15.999
for cation in list(cations.keys()):
FW += cations[cation]*CationMass[cations_to_oxides[cation]]
return FW
#----------DATA TRANSFORMATION FOR PANDAS DATAFRAMES---------#
def fluid_molfrac_to_wt(data, H2O_colname='XH2O_fl_VESIcal', CO2_colname='XCO2_fl_VESIcal'):
"""
Takes in a pandas dataframe object and converts only the fluid composition from mole fraction to wt%, leaving the melt composition
in tact. The user must specify the names of the XH2O_fl and XCO2_fl columns.
Parameters
----------
data: pandas DataFrame
Sample composition(s) containing columns for H2O and CO2 concentrations in the fluid.
H2O_colname: str
OPTIONAL. The default value is 'XH2O_fl', which is what is returned by ExcelFile() core calculations.
String containing the name of the column corresponding to the H2O concentration in the fluid, in mol fraction.
CO2_colname: str
OPTIONAL. The default value is 'XCO2_fl', which is what is returned by ExcelFile() core calculations.
String containing the name of the column corresponding to the CO2 concentration in the fluid, in mol fraction.
Returns
-------
pandas DataFrame
Original data passed plus newly calculated values are returned.
"""
convData = data.copy()
MPO_H2O_list = []
MPO_CO2_list = []
for index, row in convData.iterrows():
MPO_H2O_list.append(row[H2O_colname] * oxideMass["H2O"])
MPO_CO2_list.append(row[CO2_colname] * oxideMass["CO2"])
convData["MPO_H2O"] = MPO_H2O_list
convData["MPO_CO2"] = MPO_CO2_list
convData["H2O_fl_wt"] = 100 * convData["MPO_H2O"] / (convData["MPO_H2O"] + convData["MPO_CO2"])
convData["CO2_fl_wt"] = 100 * convData["MPO_CO2"] / (convData["MPO_H2O"] + convData["MPO_CO2"])
del convData["MPO_H2O"]
del convData["MPO_CO2"]
return convData
def fluid_wt_to_molfrac(data, H2O_colname='H2O_fl_wt', CO2_colname='CO2_fl_wt'):
"""
Takes in a pandas dataframe object and converts only the fluid composition from wt% to mole fraction, leaving the melt composition
in tact. The user must specify the names of the H2O_fl_wt and CO2_fl_wt columns.
Parameters
----------
data: pandas DataFrame
DataFrame containing columns for H2O and CO2 concentrations in the fluid.
H2O_colname: str
OPTIONAL. The default value is 'H2O_fl_wt', which is what is returned by ExcelFile() core calculations.
String containing the name of the column corresponding to the H2O concentration in the fluid, in wt%.
CO2_colname: str
OPTIONAL. The default value is 'CO2_fl_wt', which is what is returned by ExcelFile() core calculations.
String containing the name of the column corresponding to the CO2 concentration in the fluid, in wt%.
Returns
-------
pandas DataFrame
Original data passed plus newly calculated values are returned.
"""
convData = data.copy()
MPO_H2O_list = []
MPO_CO2_list = []
for index, row in convData.iterrows():
MPO_H2O_list.append(row[H2O_colname] / oxideMass["H2O"])
MPO_CO2_list.append(row[CO2_colname] / oxideMass["CO2"])
convData["MPO_H2O"] = MPO_H2O_list
convData["MPO_CO2"] = MPO_CO2_list
convData["XH2O_fl"] = convData["MPO_H2O"] / (convData["MPO_H2O"] + convData["MPO_CO2"])
convData["XCO2_fl"] = convData["MPO_CO2"] / (convData["MPO_H2O"] + convData["MPO_CO2"])
del convData["MPO_H2O"]
del convData["MPO_CO2"]
return convData
#----------DEFINE SOME NORMALIZATION METHODS-----------#
def normalize(sample):
"""Normalizes an input composition to 100%. This is the 'standard' normalization routine.
Parameters
----------
sample: pandas Series, dictionary, pandas DataFrame, or ExcelFile object
A single composition can be passed as a dictionary. Multiple compositions can be passed either as
a pandas DataFrame or an ExcelFile object. Compositional information as oxides must be present.
Returns
-------
Sample passed as > Returned as
pandas Series > pandas Series
dictionary > dictionary
pandas DataFrame > pandas DataFrame
ExcelFile object > pandas DataFrame
Normalized major element oxides.
"""
def single_normalize(sample):
single_sample = sample
return {k: 100.0 * v / sum(single_sample.values()) for k, v in single_sample.items()}
def multi_normalize(sample):
multi_sample = sample.copy()
multi_sample["Sum"] = sum([multi_sample[oxide] for oxide in oxides])
for column in multi_sample:
if column in oxides:
multi_sample[column] = 100.0*multi_sample[column]/multi_sample["Sum"]
del multi_sample["Sum"]
return multi_sample
if isinstance(sample, dict):
_sample = sample.copy()
return single_normalize(_sample)
elif isinstance(sample, pd.core.series.Series):
_sample = pd.Series(sample.copy())
sample_dict = sample.to_dict()
return pd.Series(single_normalize(sample_dict))
elif isinstance(sample, ExcelFile):
_sample = sample
data = _sample.data
return multi_normalize(data)
elif isinstance(sample, pd.DataFrame):
return multi_normalize(sample)
def normalize_FixedVolatiles(sample):
""" Normalizes major element oxides to 100 wt%, including volatiles. The volatile
wt% will remain fixed, whilst the other major element oxides are reduced proportionally
so that the total is 100 wt%.
Parameters
----------
sample: pandas Series, dictionary, pandas DataFrame, or ExcelFile object
Major element oxides in wt%
Returns
-------
Sample passed as > Returned as
pandas Series > pandas Series
dictionary > dictionary
pandas DataFrame > pandas DataFrame
ExcelFile object > pandas DataFrame
Normalized major element oxides.
"""
def single_FixedVolatiles(sample):
normalized = pd.Series({},dtype=float)
volatiles = 0
if 'CO2' in list(_sample.index):
volatiles += _sample['CO2']
if 'H2O' in list(_sample.index):
volatiles += _sample['H2O']
for ox in list(_sample.index):
if ox != 'H2O' and ox != 'CO2':
normalized[ox] = _sample[ox]
normalized = normalized/np.sum(normalized)*(100-volatiles)
if 'CO2' in list(_sample.index):
normalized['CO2'] = _sample['CO2']
if 'H2O' in list(_sample.index):
normalized['H2O'] = _sample['H2O']
return normalized
def multi_FixedVolatiles(sample):
multi_sample = sample.copy()
multi_sample["Sum_anhy"] = sum([multi_sample[oxide] for oxide in anhydrous_oxides])
multi_sample["Sum_vols"] = sum([multi_sample[vol] for vol in volatiles])
for column in multi_sample:
if column in anhydrous_oxides:
multi_sample[column] = 100.0*multi_sample[column]/multi_sample["Sum_anhy"]
multi_sample[column] = multi_sample[column] / (100.0/(100.0-multi_sample["Sum_vols"]))
del multi_sample["Sum_anhy"]
del multi_sample["Sum_vols"]
return multi_sample
if isinstance(sample, dict):
_sample = pd.Series(sample.copy())
return single_FixedVolatiles(_sample).to_dict()
elif isinstance(sample, pd.core.series.Series):
_sample = pd.Series(sample.copy())
return single_FixedVolatiles(_sample)
elif isinstance(sample, ExcelFile):
_sample = sample
data = _sample.data
return multi_FixedVolatiles(data)
elif isinstance(sample, pd.DataFrame):
return multi_FixedVolatiles(sample)
else:
raise InputError("The composition input must be a pandas Series or dictionary for single sample \
or a pandas DataFrame or ExcelFile object for multi-sample.")
def normalize_AdditionalVolatiles(sample):
"""Normalises major element oxide wt% to 100%, assuming it is volatile-free. If
H2O or CO2 are passed to the function, their un-normalized values will be retained
in addition to the normalized non-volatile oxides, summing to >100%.
Parameters
----------
sample: pandas Series, dictionary, pandas DataFrame, or ExcelFile object
Major element oxides in wt%
Returns
-------
Sample passed as > Returned as
pandas Series > pandas Series
dictionary > dictionary
pandas DataFrame > pandas DataFrame
ExcelFile object > pandas DataFrame
Normalized major element oxides.
"""
def single_AdditionalVolatiles(sample):
normalized = pd.Series({})
for ox in list(_sample.index):
if ox != 'H2O' and ox != 'CO2':
normalized[ox] = _sample[ox]
normalized = normalized/np.sum(normalized)*100
if 'H2O' in _sample.index:
normalized['H2O'] = _sample['H2O']
if 'CO2' in _sample.index:
normalized['CO2'] = _sample['CO2']
return normalized
def multi_AdditionalVolatiles(sample):
multi_sample = sample.copy()
multi_sample["Sum"] = sum([multi_sample[oxide] for oxide in anhydrous_oxides])
for column in multi_sample:
if column in anhydrous_oxides:
multi_sample[column] = 100.0*multi_sample[column]/multi_sample["Sum"]
del multi_sample["Sum"]
return multi_sample
if isinstance(sample, dict):
_sample = pd.Series(sample.copy())
return single_AdditionalVolatiles(_sample).to_dict()
elif isinstance(sample, pd.core.series.Series):
_sample = pd.Series(sample.copy())
return single_AdditionalVolatiles(sample)
elif isinstance(sample, ExcelFile):
_sample = sample
data = _sample.data
return multi_AdditionalVolatiles(data)
elif isinstance(sample, pd.DataFrame):
return multi_AdditionalVolatiles(sample)
else:
raise InputError("The composition input must be a pandas Series or dictionary for single sample \
or a pandas DataFrame or ExcelFile object for multi-sample.")
#------------DEFINE MAJOR CLASSES-------------------#
class ExcelFile(object):
"""An excel file with sample names and oxide compositions
Attributes
----------
filename: str
Path to the excel file, e.g., "my_file.xlsx"
sheet_name: str
OPTIONAL. Default value is 0 which gets the first sheet in the excel spreadsheet file. This implements the pandas.
read_excel() sheet_name parameter. But functionality to read in more than one sheet at a time (e.g., pandas.read_excel(sheet_name=None))
is not yet imlpemented in VESIcal. From the pandas 1.0.4 documentation:
Available cases:
- Defaults to 0: 1st sheet as a DataFrame
- 1: 2nd sheet as a DataFrame
- "Sheet1": Load sheet with name “Sheet1”
input_type: str or int
OPTIONAL. Default is 'wtpercent'. String defining whether the oxide composition is given in wt percent
("wtpercent", which is the default), mole percent ("molpercent"), or mole fraction ("molfrac").
label: str
OPTIONAL. Default is 'Label'. Name of the column within the passed Excel file referring to sample names.
"""
def __init__(self, filename, sheet_name=0, input_type='wtpercent', label='Label', **kwargs):
"""Return an ExcelFile object whoes parameters are defined here."""
if isinstance(sheet_name, str) or isinstance(sheet_name, int):
pass
else:
raise InputError("If sheet_name is passed, it must be of type str or int. Currently, VESIcal cannot import more than one sheet at a time.")
self.input_type = input_type
data = pd.read_excel(filename, sheet_name=sheet_name)
data = data.fillna(0)
try:
data = data.set_index(label)
except:
raise InputError(
"Imported file must contain a column of sample names. If this column is not titled 'Label' (the default value), you must pass the column name to arg label. For example: ExcelFile('myfile.xslx', label='SampleNames')") #TODO test
if 'model' in kwargs:
warnings.warn("You don't need to pass a model here, so it will be ignored. You can specify a model when performing calculations on your dataset (e.g., calculate_dissolved_volatiles())",RuntimeWarning)
total_iron_columns = ["FeOt", "FeOT", "FeOtot", "FeOtotal", "FeOstar", "FeO*"]
for name in total_iron_columns:
if name in data.columns:
if 'FeO' in data.columns:
warnings.warn("Both " + str(name) + " and FeO columns were passed. " + str(name) + " column will be ignored.",RuntimeWarning)
else:
warnings.warn("Total iron column " + str(name) + " detected. This column will be treated as FeO. If Fe2O3 data are not given, Fe2O3 will be 0.0.",RuntimeWarning)
data['FeO'] = data[name]
for oxide in oxides:
if oxide in data.columns:
pass
else:
data[oxide] = 0.0
# TODO test all input types produce correct values
if input_type == "wtpercent":
pass
if input_type == "molpercent":
data = mol_to_wtpercent(data)
if input_type == "molfrac":
data = mol_to_wtpercent(data)
self.data = data
def preprocess_sample(self,sample):
"""
Adds 0.0 values to any oxide data not passed.
Parameters
----------
sample: pandas DataFrame
self.data composition of samples in wt% oxides
Returns
-------
pandas DataFrame
"""
for oxide in oxides:
if oxide in self.data.columns:
pass
else:
self.data[oxide] = 0.0
return sample
def get_sample_oxide_comp(self, sample, norm='none'):
"""
Returns oxide composition of a single sample from a user-imported excel file as a dictionary
Parameters
----------
sample: string
Name of the desired sample
norm_style: string
OPTIONAL. Default value is 'standard'. This specifies the style of normalization applied to the sample.
'standard' normalizes the entire input composition (including any volatiles) to 100%.
'fixedvolatiles' normalizes oxides to 100%, including volatiles. The volatile
wt% will remain fixed, whilst the other major element oxides are reduced proportionally
so that the total is 100 wt%.
'additionalvolatiles' normalizes oxides to 100%, assuming it is volatile-free. If
H2O or CO2 are passed to the function, their un-normalized values will be retained
in addition to the normalized non-volatile oxides, summing to >100%.
'none' returns the value-for-value un-normalized composition.
Returns
-------
dictionary
Composition of the sample as oxides
"""
if norm == 'none' or norm == 'standard' or norm == 'fixedvolatiles' or norm == 'additionalvolatiles':
pass
else:
raise InputError('norm must be either none, standard, fixedvolatiles, or additionalvolatiles.')
data = self.data
my_sample = pd.DataFrame(data.loc[sample])
sample_dict = (my_sample.to_dict()[sample])
sample_oxides = {}
for item, value in sample_dict.items():
if item in oxides:
sample_oxides.update({item: value})
if norm == 'standard':
return normalize(sample_oxides)
if norm == 'fixedvolatiles':
return normalize_FixedVolatiles(sample_oxides)
if norm == 'additionalvolatiles':
return normalize_AdditionalVolatiles(sample_oxides)
if norm == 'none':
return sample_oxides
def get_XH2O_fluid(self, sample, temperature, pressure, H2O, CO2):
"""An internally used function to calculate fluid composition.
Parameters
----------
sample: dictionary
Sample composition in wt% oxides
temperature: float
Temperature in degrees C.
pressure: float
Pressure in bars
H2O: float
wt% H2O in the system
CO2: float
wt% CO2 in the system
Returns
-------
float
Mole fraction of H2O in the H2O-CO2 fluid
"""
pressureMPa = pressure / 10.0
bulk_comp = {oxide: sample[oxide] for oxide in oxides}
bulk_comp["H2O"] = H2O
bulk_comp["CO2"] = CO2
feasible = melts.set_bulk_composition(bulk_comp)
output = melts.equilibrate_tp(temperature, pressureMPa, initialize=True)
(status, temperature, pressureMPa, xmlout) = output[0]
fluid_comp = melts.get_composition_of_phase(xmlout, phase_name='Fluid', mode='component')
#NOTE mode='component' returns endmember component keys with values in mol fraction.
if "Water" in fluid_comp:
H2O_fl = fluid_comp["Water"]
else:
H2O_fl = 0.0
# if H2O_fl == 0:
# raise SaturationError("Composition not fluid saturated.")
return H2O_fl
def save_excelfile(self, filename, calculations, sheet_name=None): #TODO how to handle if user just wants to normalize data?
"""
Saves data calculated by the user in batch processing mode (using the ExcelFile class methods) to an organized
excel file, with the original user data plus any calculated data.
Parameters
----------
filename: string
Name of the file. Extension (.xlsx) should be passed along with the name itself, all in quotes (e.g., 'myfile.xlsx').
calculations: list
List of variables containing calculated outputs from any of the core ExcelFile functions: calculate_dissolved_volatiles,
calculate_equilibrium_fluid_comp, and calculate_saturation_pressure.
sheet_name: None or list
OPTIONAL. Default value is None. Allows user to set the name of the sheet or sheets written to the Excel file.
Returns
-------
Excel File
Creates and saves an Excel file with data from each calculation saved to its own sheet.
"""
if isinstance(calculations, list):
if isinstance(sheet_name, list) or sheet_name is None:
pass
else:
raise InputError("calculations and sheet_name must be type list. If you only have one calculation or sheet_name to pass, make sure they are passed in square brackets []")
with pd.ExcelWriter(filename) as writer:
self.data.to_excel(writer, 'Original_User_Data')
if sheet_name is None:
for n, df in enumerate(calculations):
df.to_excel(writer, 'Calc%s' % n)
elif isinstance(sheet_name, list):
if len(sheet_name) == len(calculations):
pass
else:
raise InputError("calculations and sheet_name must have the same length")
for i in range(len(calculations)):
if isinstance(sheet_name[i], str):
calculations[i].to_excel(writer, sheet_name[i])
else:
raise InputError("if sheet_name is passed, it must be list of strings")
else:
raise InputError("sheet_name must be type list")
return print("Saved " + str(filename))
def calculate_dissolved_volatiles(self, temperature, pressure, X_fluid=1, print_status=True, model='MagmaSat', record_errors=False, **kwargs):
"""
Calculates the amount of H2O and CO2 dissolved in a magma at the given P/T conditions and fluid composition. Fluid composition
will be matched to within 0.0001 mole fraction.
Parameters
----------
temperature: float, int, or str
Temperature, in degrees C. Can be passed as float, in which case the
passed value is used as the temperature for all samples. Alternatively, temperature information for each individual
sample may already be present in the ExcelFile object. If so, pass the str value corresponding to the column
title in the ExcelFile object.
presure: float, int, or str
Pressure, in bars. Can be passed as float or int, in which case the
passed value is used as the pressure for all samples. Alternatively, pressure information for each individual
sample may already be present in the ExcelFile object. If so, pass the str value corresponding to the column
title in the ExcelFile object.
X_fluid: float, int, or str
OPTIONAL: Default value is 1. The mole fraction of H2O in the H2O-CO2 fluid. X_fluid=1 is a pure H2O fluid. X_fluid=0 is a
pure CO2 fluid. Can be passed as a float or int, in which case the passed value is used as the X_fluid for all samples.
Alternatively, X_fluid information for each individual sample may already be present in the ExcelFile object. If so, pass
the str value corresponding to the column title in the ExcelFile object.
print_status: bool
OPTIONAL: The default value is True, in which case the progress of the calculation will be printed to the terminal.
If set to False, nothing will be printed. MagmaSat calculations tend to be slow, and so a value of True is recommended
for most use cases.
model: string
The default value is 'MagmaSat'. Any other model name can be passed here as a string (in single quotes).
record_errors: bool
OPTIONAL: If True, any errors arising during the calculation will be recorded as a column.
Returns
-------
pandas DataFrame
Original data passed plus newly calculated values are returned.
"""
data = self.preprocess_sample(self.data)
dissolved_data = data.copy()
if isinstance(temperature, str):
file_has_temp = True
temp_name = temperature
elif isinstance(temperature, float) or isinstance(temperature, int):
file_has_temp = False
else:
raise InputError("temp must be type str or float or int")
if isinstance(pressure, str):
file_has_press = True
press_name = pressure
elif isinstance(pressure, float) or isinstance(pressure, int):
file_has_press = False
else:
raise InputError("pressure must be type str or float or int")
if isinstance(X_fluid, str):
file_has_X = True
X_name = X_fluid
elif isinstance(X_fluid, float) or isinstance(X_fluid, int):
file_has_X = False
if X_fluid != 0 and X_fluid !=1:
if X_fluid < 0.001 or X_fluid > 0.999:
raise InputError("X_fluid is calculated to a precision of 0.0001 mole fraction. \
Value for X_fluid must be between 0.0001 and 0.9999.")
else:
raise InputError("X_fluid must be type str or float or int")
H2Ovals = []
CO2vals = []
warnings = []
errors = []
if model in get_models(models='mixed'):
for index, row in dissolved_data.iterrows():
try:
if file_has_temp == True:
temperature = row[temp_name]
if file_has_press == True:
pressure = row[press_name]
if file_has_X == True:
X_fluid = row[X_name]
bulk_comp = {oxide: row[oxide] for oxide in oxides}
calc = calculate_dissolved_volatiles(sample=bulk_comp, pressure=pressure, temperature=temperature,
X_fluid=(X_fluid, 1-X_fluid), model=model,
silence_warnings=True, **kwargs)
H2Ovals.append(calc.result['H2O_liq'])
CO2vals.append(calc.result['CO2_liq'])
warnings.append(calc.calib_check)
errors.append('')
except Exception as inst:
H2Ovals.append(np.nan)
CO2vals.append(np.nan)
warnings.append('Calculation Failed.')
errors.append(sys.exc_info()[0])
dissolved_data["H2O_liq_VESIcal"] = H2Ovals
dissolved_data["CO2_liq_VESIcal"] = CO2vals
if file_has_temp == False:
dissolved_data["Temperature_C_VESIcal"] = temperature
if file_has_press == False:
dissolved_data["Pressure_bars_VESIcal"] = pressure
if file_has_X == False:
dissolved_data["X_fluid_input_VESIcal"] = X_fluid
dissolved_data["Model"] = model
dissolved_data["Warnings"] = warnings
if record_errors == True:
dissolved_data["Errors"] = errors
return dissolved_data
elif model == 'MagmaSat':
XH2Ovals = []
XCO2vals = []
FluidProportionvals = []
for index, row in dissolved_data.iterrows():
if print_status == True:
print("Calculating sample " + str(index))
try:
if file_has_temp == True:
temperature = row[temp_name]
if file_has_press == True:
pressure = row[press_name]
if file_has_X == True:
X_fluid = row[X_name]
bulk_comp = {oxide: row[oxide] for oxide in oxides}
calc = calculate_dissolved_volatiles(sample=bulk_comp, pressure=pressure, temperature=temperature,
X_fluid=X_fluid, model=model, silence_warnings=True,
verbose=True)
H2Ovals.append(calc.result['H2O_liq'])
CO2vals.append(calc.result['CO2_liq'])
XH2Ovals.append(calc.result['XH2O_fl'])
XCO2vals.append(calc.result['XCO2_fl'])
FluidProportionvals.append(calc.result['FluidProportion_wt'])
warnings.append(calc.calib_check)
errors.append('')
except Exception as inst:
H2Ovals.append(np.nan)
CO2vals.append(np.nan)
XH2Ovals.append(np.nan)
XCO2vals.append(np.nan)
FluidProportionvals.append(np.nan)
warnings.append('Calculation Failed.')
errors.append(sys.exc_info()[0])
dissolved_data["H2O_liq_VESIcal"] = H2Ovals
dissolved_data["CO2_liq_VESIcal"] = CO2vals
if file_has_temp == False:
dissolved_data["Temperature_C_VESIcal"] = temperature
if file_has_press == False:
dissolved_data["Pressure_bars_VESIcal"] = pressure
if file_has_X == False:
dissolved_data["X_fluid_input_VESIcal"] = X_fluid
dissolved_data["Model"] = model
dissolved_data["Warnings"] = warnings
if record_errors == True:
dissolved_data["Errors"] = errors
return dissolved_data
else:
XH2Ovals = []
XCO2vals = []
FluidProportionvals = []
for index, row in dissolved_data.iterrows():
if file_has_temp == True:
temperature = row[temp_name]
if file_has_press == True:
pressure = row[press_name]
if file_has_X == True:
X_fluid = row[X_name]
bulk_comp = {oxide: row[oxide] for oxide in oxides}
if 'Water' in model:
try:
calc = calculate_dissolved_volatiles(sample=bulk_comp, pressure=pressure, temperature=temperature,
X_fluid=X_fluid, model=model, silence_warnings=True)
H2Ovals.append(calc.result)
warnings.append(calc.calib_check)
except:
H2Ovals.append(0)
warnings.append('Calculation Failed #001')
if 'Carbon' in model:
try:
calc = calculate_dissolved_volatiles(sample=bulk_comp, pressure=pressure, temperature=temperature,
X_fluid=X_fluid, model=model, silence_warnings=True)
CO2vals.append(calc.result)
warnings.append(calc.calib_check)
except:
CO2vals.append(0)
warnings.append('Calculation Failed #002')
if 'Water' in model:
dissolved_data["H2O_liq_VESIcal"] = H2Ovals
if 'Carbon' in model:
dissolved_data["CO2_liq_VESIcal"] = CO2vals
if file_has_temp == False:
dissolved_data["Temperature_C_VESIcal"] = temperature
if file_has_press == False:
dissolved_data["Pressure_bars_VESIcal"] = pressure
if file_has_X == False:
dissolved_data["X_fluid_input_VESIcal"] = X_fluid
dissolved_data["Model"] = model
dissolved_data["Warnings"] = warnings
return dissolved_data
def calculate_equilibrium_fluid_comp(self, temperature, pressure, print_status=False, model='MagmaSat', **kwargs):
#TODO make molfrac the default
"""
Returns H2O and CO2 concentrations in wt% or mole fraction in a fluid in equilibrium with the given sample(s) at the given P/T condition.
Parameters
----------
sample: ExcelFile object
Compositional information on samples in oxides.
temperature: float, int, or str
Temperature, in degrees C. Can be passed as float, in which case the
passed value is used as the temperature for all samples. Alternatively, temperature information for each individual
sample may already be present in the ExcelFile object. If so, pass the str value corresponding to the column
title in the ExcelFile object.
presure: float, int, or str
Pressure, in bars. Can be passed as float or int, in which case the
passed value is used as the pressure for all samples. Alternatively, pressure information for each individual
sample may already be present in the ExcelFile object. If so, pass the str value corresponding to the column
title in the ExcelFile object.
model: string
OPTIONAL: Default is 'MagmaSat'. Any other model name can be passed here.
Returns
-------
pandas DataFrame
Original data passed plus newly calculated values are returned.
"""
data = self.preprocess_sample(self.data)
fluid_data = data.copy()
if isinstance(temperature, str):
file_has_temp = True
temp_name = temperature
elif isinstance(temperature, float) or isinstance(temperature, int):
file_has_temp = False
else:
raise InputError("temp must be type str or float or int")
if isinstance(pressure, str):
file_has_press = True
press_name = pressure
elif isinstance(pressure, float) or isinstance(pressure, int):
file_has_press = False
else:
raise InputError("pressure must be type str or float or int")
H2Ovals = []
CO2vals = []
warnings = []
if model in get_models(models='mixed') or model == "MooreWater":
for index, row in fluid_data.iterrows():
try:
if file_has_temp == True:
temperature = row[temp_name]
if file_has_press == True:
pressure = row[press_name]
bulk_comp = {oxide: row[oxide] for oxide in oxides}
calc = calculate_equilibrium_fluid_comp(sample=bulk_comp, pressure=pressure, temperature=temperature,
model=model, silence_warnings=True, **kwargs)
H2Ovals.append(calc.result['H2O'])
CO2vals.append(calc.result['CO2'])
warnings.append(calc.calib_check)
except:
H2Ovals.append(np.nan)
CO2vals.append(np.nan)
warnings.append("Calculation Failed.")
fluid_data["XH2O_fl_VESIcal"] = H2Ovals
fluid_data["XCO2_fl_VESIcal"] = CO2vals
if file_has_temp == False:
fluid_data["Temperature_C_VESIcal"] = temperature
if file_has_press == False:
fluid_data["Pressure_bars_VESIcal"] = pressure
fluid_data["Model"] = model
fluid_data["Warnings"] = warnings
return fluid_data
elif model == 'MagmaSat':
for index, row in fluid_data.iterrows():
if print_status == True:
print("Calculating sample " + str(index))
try:
if file_has_temp == True:
temperature = row[temp_name]
if file_has_press == True:
pressure = row[press_name]
bulk_comp = {oxide: row[oxide] for oxide in oxides}
calc = calculate_equilibrium_fluid_comp(sample=bulk_comp, pressure=pressure, temperature=temperature, model=model, silence_warnings=True)
H2Ovals.append(calc.result['H2O'])
CO2vals.append(calc.result['CO2'])
warnings.append(calc.calib_check)
except:
H2Ovals.append(np.nan)
CO2vals.append(np.nan)
warnings.append("Calculation Failed.")
fluid_data["XH2O_fl_VESIcal"] = H2Ovals
fluid_data["XCO2_fl_VESIcal"] = CO2vals
if file_has_temp == False:
fluid_data["Temperature_C_VESIcal"] = temperature
if file_has_press == False:
fluid_data["Pressure_bars_VESIcal"] = pressure
fluid_data["Model"] = model
fluid_data["Warnings"] = warnings
return fluid_data
else:
saturated = []
for index, row in fluid_data.iterrows():
try:
if file_has_temp == True:
temperature = row[temp_name]
if file_has_press == True:
pressure = row[press_name]
bulk_comp = {oxide: row[oxide] for oxide in oxides}
calc = calculate_equilibrium_fluid_comp(sample=bulk_comp, pressure=pressure, temperature=temperature, model=model, silence_warnings=True)
saturated.append(calc.result)
warnings.append(calc.calib_check)
except:
saturated.append(np.nan)
warnings.append("Calculation Failed.")
fluid_data["Saturated_VESIcal"] = saturated
if file_has_temp == False:
fluid_data["Temperature_C_VESIcal"] = temperature
if file_has_press == False:
fluid_data["Pressure_bars_VESIcal"] = pressure
fluid_data["Model"] = model
fluid_data["Warnings"] = warnings
return fluid_data
def calculate_saturation_pressure(self, temperature, print_status=True, model='MagmaSat', **kwargs): #TODO fix weird printing
"""
Calculates the saturation pressure of multiple sample compositions in the ExcelFile.
Parameters
----------
temperature: float, int, or str
Temperature at which to calculate saturation pressures, in degrees C. Can be passed as float or int, in which case the
passed value is used as the temperature for all samples. Alternatively, temperature information for each individual
sample may already be present in the passed ExcelFile object. If so, pass the str value corresponding to the column
title in the passed ExcelFile object.
print_status: bool
OPTIONAL: The default value is True, in which case the progress of the calculation will be printed to the terminal.
If set to False, nothing will be printed. MagmaSat calculations tend to be slow, and so a value of True is recommended
more most use cases.
model: string
OPTIONAL: Default is 'MagmaSat'. Any other model name can be passed here.
Returns
-------
pandas DataFrame object
Values returned are saturation pressure in bars, the mass of fluid present, and the composition of the
fluid present.
"""
data = self.preprocess_sample(self.data)
satp_data = data.copy()
if isinstance(temperature, str):
file_has_temp = True
temp_name = temperature
elif isinstance(temperature, float) or isinstance(temperature, int):
file_has_temp = False
else:
raise InputError("temperature must be type str or float or int")
if model != 'MagmaSat':
satP = []
warnings = []
for index, row in satp_data.iterrows():
try:
if file_has_temp == True:
temperature = row[temp_name]
bulk_comp = {oxide: row[oxide] for oxide in oxides}
calc = calculate_saturation_pressure(sample=bulk_comp, temperature=temperature,
model=model, silence_warnings=True, **kwargs)
satP.append(calc.result)
warnings.append(calc.calib_check)
except:
satP.append(np.nan)
warnings.append("Calculation Failed")
satp_data["SaturationP_bars_VESIcal"] = satP
if file_has_temp == False:
satp_data["Temperature_C_VESIcal"] = temperature
satp_data["Model"] = model
satp_data["Warnings"] = warnings
return satp_data
else:
satP = []
flmass = []
flH2O = []
flCO2 = []
flsystem_wtper = []
warnings = []
for index, row in satp_data.iterrows():
if print_status == True:
print("Calculating sample " + str(index))
try:
if file_has_temp == True:
temperature = row[temp_name]
bulk_comp = {oxide: row[oxide] for oxide in oxides}
calc = calculate_saturation_pressure(sample=bulk_comp, temperature=temperature, model=model, verbose=True, silence_warnings=True)
satP.append(calc.result["SaturationP_bars"])
flmass.append(calc.result["FluidMass_grams"])
flsystem_wtper.append(calc.result["FluidProportion_wt"])
flH2O.append(calc.result["XH2O_fl"])
flCO2.append(calc.result["XCO2_fl"])
warnings.append(calc.calib_check)
except:
satP.append(np.nan)
flmass.append(np.nan)
flsystem_wtper.append(np.nan)
flH2O.append(np.nan)
flCO2.append(np.nan)
warnings.append("Calculation Failed")
satp_data["SaturationP_bars_VESIcal"] = satP
if file_has_temp == False:
satp_data["Temperature_C_VESIcal"] = temperature
satp_data["XH2O_fl_VESIcal"] = flH2O
satp_data["XCO2_fl_VESIcal"] = flCO2
satp_data["FluidMass_grams_VESIcal"] = flmass
satp_data["FluidSystem_wt_VESIcal"] = flsystem_wtper
satp_data["Model"] = model
satp_data["Warnings"] = warnings
if print_status == True:
print("Done!")
return satp_data
class CalibrationRange(object):
""" The CalibrationRange object allows the range of allowable parameters to be specified and
used in checking and reporting of the results.
"""
def __init__(self, parameter_name, value, checkfunction=None, units='', model_name='',
fail_msg='',fail_dict={}, pass_msg='', pass_dict={}, description_msg='', description_dict={}):
self.parameter_name = parameter_name
self.value = value
self.checkfunction = checkfunction
self.units = units
self.model_name = model_name
self.fail_msg = (copy(fail_msg), copy(fail_dict))
self.pass_msg = (copy(pass_msg), copy(pass_dict))
self.description_msg = (copy(description_msg), copy(description_dict))
def check(self,parameters):
"""Method for checking whether parameters satisfy the calibration range."""
if self.parameter_name in parameters:
return self.checkfunction(self.value,parameters[self.parameter_name])
else:
return None
def string(self,parameters,report_nonexistance=True):
"""Returns a string statement of the calibration check"""
if type(parameters) == type(None):
msgdict = self.description_msg[1]
if type(self.value) == float or type(self.value) == int:
msgdict['calib_val'] = self.value
elif type(self.value) == list or type(self.value) == tuple or type(self.value) == np.ndarray:
for i in range(len(self.value)):
msgdict['calib_val'+str(i)] = self.value[i]
if 'param_name' not in msgdict:
msgdict['param_name'] = self.parameter_name
if 'units' not in msgdict:
msgdict['units'] = self.units
if 'model_name' not in msgdict:
msgdict['model_name'] = self.model_name
return self.description_msg[0].format(**msgdict)
else:
check = self.check(parameters)
if check == True:
msgdict = self.pass_msg[1]
msgdict['param_val'] = parameters[self.parameter_name]
if type(self.value) == float or type(self.value) == int:
msgdict['calib_val'] = self.value
elif type(self.value) == list or type(self.value) == tuple or type(self.value) == np.ndarray:
for i in range(len(self.value)):
msgdict['calib_val'+str(i)] = self.value[i]
if 'param_name' not in msgdict:
msgdict['param_name'] = self.parameter_name
if 'units' not in msgdict:
msgdict['units'] = self.units
if 'model_name' not in msgdict:
msgdict['model_name'] = self.model_name
return self.pass_msg[0].format(**msgdict)
elif check == False:
msgdict = self.fail_msg[1]
msgdict['param_val'] = parameters[self.parameter_name]
if type(self.value) == float or type(self.value) == int:
msgdict['calib_val'] = self.value
elif type(self.value) == list or type(self.value) == tuple or type(self.value) == np.ndarray:
for i in range(len(self.value)):
msgdict['calib_val'+str(i)] = self.value[i]
if 'param_name' not in msgdict:
msgdict['param_name'] = self.parameter_name
if 'units' not in msgdict:
msgdict['units'] = self.units
if 'model_name' not in msgdict:
msgdict['model_name'] = self.model_name
return self.fail_msg[0].format(**msgdict)
else:
if report_nonexistance == True:
return "A value for {} was not provided.".format(self.parameter_name)
else:
return ''
# class old_CalibrationRange(object):
# """ The CalibrationRange object allows the range of allowable parameters to be specified and
# used in checking and reporting of the results.
# """
# def __init__(self,parameter_name,value,unit='',modelname='',explanation_string=None,
# parameter_string=None,value_fmt="{:.1f}"):
# self.parameter_name = parameter_name
# self.value = value
# self.value_fmt = value_fmt
# self.model_name = modelname
# self.unit = unit
# self.explanation_string = explanation_string
# if parameter_string is not None:
# self.parameter_string = parameter_string
# else:
# self.parameter_string = parameter_name
#
# @abstractmethod
# def check(self,parameters):
# """Method for checking whether parameters satisfy the calibration range."""
# return True
#
# @abstractmethod
# def string(self,parameters):
# """Returns a string statement of the calibration check"""
# return 'No string return defined. '
class Model(object):
"""The model object implements a volatile solubility model. It is composed
of the methods needed to evaluate :func:`VESIcal.calculate_dissolved_volatiles`,
:func:`VESIcal.calculate_equilibrium_fluid_comp`, and :func:`calculate_saturation_pressure`. The
fugacity and activity models for the volatiles species must be specified,
defaulting to ideal.
"""
def __init__(self):
self.set_volatile_species(None)
self.set_fugacity_model(fugacity_idealgas())
self.set_activity_model(activity_idealsolution())
self.set_calibration_ranges([])
self.set_solubility_dependence(False)
def set_volatile_species(self,volatile_species):
if type(volatile_species) == str:
volatile_species = [volatile_species]
elif type(volatile_species) != list:
raise InputError("volatile_species must be a str or list.")
self.volatile_species = volatile_species
def set_fugacity_model(self,fugacity_model):
self.fugacity_model = fugacity_model
def set_activity_model(self,activity_model):
self.activity_model = activity_model
def set_calibration_ranges(self,calibration_ranges):
self.calibration_ranges = calibration_ranges
def set_solubility_dependence(self,solubility_dependence):
self.solubility_dependence = solubility_dependence
@abstractmethod
def calculate_dissolved_volatiles(self,**kwargs):
pass
@abstractmethod
def calculate_equilibrium_fluid_comp(self,**kwargs):
pass
@abstractmethod
def calculate_saturation_pressure(self,**kwargs):
pass
@abstractmethod
def preprocess_sample(self,**kwargs):
pass
# @abstractmethod
def check_calibration_range(self,parameters,report_nonexistance=True):
""" Checks whether the given parameters are within the ranges defined by the
CalibrationRange objects for the model and its fugacity and activity models. An empty
string will be returned if all parameters are within the calibration range. If a
parameter is not within the calibration range, a description of the problem will be
returned in the string.
Parameters
----------
parameters dict
Dictionary keys are the names of the parameters to be checked, e.g., pressure
temperature, SiO2, etc. Values are the values of each parameter. A complete set
need not be given.
Returns
-------
str
String description of any parameters falling outside of the calibration range.
"""
s = ''
for cr in self.calibration_ranges:
if cr.check(parameters) == False:
s += cr.string(parameters,report_nonexistance)
for cr in self.fugacity_model.calibration_ranges:
if cr.check(parameters) == False:
s += cr.string(parameters,report_nonexistance)
for cr in self.activity_model.calibration_ranges:
if cr.check(parameters) == False:
s += cr.string(parameters,report_nonexistance)
return s
def get_calibration_range(self):
""" Returns a string describing the calibration ranges defined by the CalibrationRange
objects for each model, and its associated fugacity and activity models.
Returns
-------
str
String description of the calibration range objects."""
s = ''
for cr in self.calibration_ranges:
s += cr.string(None)
for cr in self.fugacity_model.calibration_ranges:
s += cr.string(None)
for cr in self.activity_model.calibration_ranges:
s += cr.string(None)
return s
class FugacityModel(object):
""" The fugacity model object is for implementations of fugacity models
for individual volatile species, though it may depend on the mole
fraction of other volatile species. It contains all the methods required
to calculate the fugacity at a given pressure and mole fraction.
"""
def __init__(self):
self.set_calibration_ranges([])
def set_calibration_ranges(self,calibration_ranges):
self.calibration_ranges = calibration_ranges
@abstractmethod
def fugacity(self,pressure,**kwargs):
"""
"""
# @abstractmethod
def check_calibration_range(self,parameters,report_nonexistance=True):
s = ''
for cr in self.calibration_ranges:
if cr.check(parameters) == False:
s += cr.string(parameters,report_nonexistance)
return s
class activity_model(object):
""" The activity model object is for implementing activity models
for volatile species in melts. It contains all the methods required to
evaluate the activity.
"""
def __init__(self):
self.set_calibration_ranges([])
def set_calibration_ranges(self,calibration_ranges):
self.calibration_ranges = calibration_ranges
@abstractmethod
def activity(self,X,**kwargs):
"""
"""
# @abstractmethod
def check_calibration_range(self,parameters,report_nonexistance=True):
s = ''
for cr in self.calibration_ranges:
if cr.check(parameters) == False:
s += cr.string(parameters,report_nonexistance)
return s
class Calculate(object):
""" The Calculate object is a template for implementing user-friendly methods for
running calculations using the volatile solubility models. All Calculate methods
have a common workflow- sample is read in, preprocessed, the calculation is performed,
the calibration range is checked, and the results stored.
"""
def __init__(self,sample,model='MagmaSat',silence_warnings=False,preprocess_sample=False,**kwargs):
if model == 'MagmaSat':
self.model = MagmaSat()
elif type(model) == str:
self.model = default_models[model]
else:
self.model = model
self.sample = sample.copy()
if preprocess_sample == True:
self.sample = self.model.preprocess_sample(self.sample)
self.result = self.calculate(sample=self.sample,**kwargs)
self.calib_check = self.check_calibration_range(sample=self.sample,**kwargs)
if self.calib_check is not None and silence_warnings == False:
if self.calib_check != '':
warnings.warn(self.calib_check,RuntimeWarning)
@abstractmethod
def calculate(self):
""" """
@abstractmethod
def check_calibration_range(self):
""" """
#-------------DEFAULT CALIBRATIONRANGE OBJECTS---------------#
def crf_EqualTo(calibval,paramval):
return calibval == paramval
crmsg_EqualTo_pass = "The {param_name} ({param_val:.1f} {units}) is equal to {calib_val:.1f} {units} as required by the calibration range of the {model_name} model. "
crmsg_EqualTo_fail = "The {param_name} is outside the calibration range of the {model_name} model, as {param_val:.1f} {units} is not equal to {calib_val:.1f} {units}. "
crmsg_EqualTo_description = "The {model_name} model is calibrated for {param_name} equal to {calib_val:.1f} {units}. "
def crf_GreaterThan(calibval,paramval):
return paramval > calibval
crmsg_GreaterThan_pass = "The {param_name} ({param_val:.1f} {units}) is greater than {calib_val:.1f} {units} as required by the calibration range of the {model_name} model. "
crmsg_GreaterThan_fail = "The {param_name} is outside the calibration range of the {model_name} model, as {param_val:.1f} {units} is not greater than {calib_val:.1f} {units}. "
crmsg_GreaterThan_description = "The {model_name} model is calibrated for {param_name} greater than {calib_val:.1f} {units}. "
def crf_LessThan(calibval,paramval):
return paramval < calibval
crmsg_LessThan_pass = "The {param_name} ({param_val:.1f} {units}) is less than {calib_val:.1f} {units} as required by the calibration range of the {model_name} model. "
crmsg_LessThan_fail = "The {param_name} is outside the calibration range of the {model_name} model, as {param_val:.1f} {units} is not less than {calib_val:.1f} {units}. "
crmsg_LessThan_description = "The {model_name} model is calibrated for {param_name} less than {calib_val:.1f} {units}. "
def crf_Between(calibval,paramval):
return paramval > calibval[0] and paramval < calibval[1]
crmsg_Between_pass = "The {param_name} ({param_val:.1f} {units}) is between {calib_val0:.1f} and {calib_val1:.1f} {units} as required by the calibration range of the {model_name} model. "
crmsg_Between_fail = "The {param_name} is outside the calibration range of the {model_name} model, as {param_val:.1f} {units} is not between {calib_val0:.1f} and {calib_val1:.1f} {units}. "
crmsg_Between_description = "The {model_name} model is calibrated for {param_name} between {calib_val0:.1f} and {calib_val1:.1f} {units}. "
def crf_LiuComp(calibval=None,sample={}):
SiTest = sample['SiO2'] >= 75.0 and sample['SiO2'] <= 77.0
NaTest = sample['Na2O'] >= 3.4 and sample['Na2O'] <= 4.7
KTest = sample['K2O'] >= 3.6 and sample['K2O'] <= 5.7
AlTest = sample['Al2O3'] >= 12.1 and sample['Al2O3'] <= 13.5
return all([SiTest, NaTest, KTest, AlTest])
crmsg_LiuComp_pass = "The sample appears to be similar in composition to the rhyolites and haplogranites used to calibrate the Liu et al. model."
crmsg_LiuComp_fail = "As the Liu et al. model incorperates no term for compositional dependence, users must take extreme care when extrapolating this model to compositions which differ significantly from the haplogranites and rhyolites in the calibration dataset. These warnings are simply a guide; we suggest that users carefully compare their major element data to the calibration dataset to check for suitability."
crmsg_LiuComp_description = "The Liu et al. model is suitable for haplogranites and rhyolites."
#-------------FUGACITY MODELS--------------------------------#
class fugacity_idealgas(FugacityModel):
""" An instance of FugacityModel for an ideal gas.
"""
def fugacity(self,pressure,X_fluid=1.0,**kwargs):
""" Returns the fugacity of an ideal gas, i.e., the partial pressure.
Parameters
----------
pressure float
Total pressure of the system, in bars.
X_fluid float
The mole fraction of the species in the vapour phase.
Returns
-------
float
Fugacity (partial pressure) in bars
"""
return pressure*X_fluid
class fugacity_KJ81_co2(FugacityModel):
""" Implementation of the Kerrick and Jacobs (1981) EOS for mixed fluids. This class
will return the properties of the CO2 component of the mixed fluid.
"""
def __init__(self):
self.set_calibration_ranges([CalibrationRange('pressure',20000.0,crf_LessThan,'bar','Kerrick and Jacobs (1981) EOS',
fail_msg=crmsg_LessThan_fail, pass_msg=crmsg_LessThan_pass, description_msg=crmsg_LessThan_description),
CalibrationRange('temperature',1050,crf_LessThan,'oC','Kerrick and Jacobs (1981) EOS',
fail_msg=crmsg_LessThan_fail, pass_msg=crmsg_LessThan_pass, description_msg=crmsg_LessThan_description)])
def fugacity(self,pressure,temperature,X_fluid,**kwargs):
""" Calculates the fugacity of CO2 in a mixed CO2-H2O fluid. Above 1050C,
it assumes H2O and CO2 do not interact, as the equations are not defined
beyond this point.
Parameters
----------
pressure float
Total pressure of the system in bars.
temperature float
Temperature in degC
X_fluid float
Mole fraction of CO2 in the fluid.
Returns
-------
float
fugacity of CO2 in bars
"""
if X_fluid == 0:
return 0
elif temperature >= 1050.0:
return pressure*np.exp(self.lnPhi_mix(pressure,temperature,1.0))*X_fluid
else:
return pressure*np.exp(self.lnPhi_mix(pressure,temperature,X_fluid))*X_fluid
def volume(self,P,T,X_fluid):
""" Calculates the volume of the mixed fluid, by solving Eq (28) of Kerrick and
Jacobs (1981) using scipy.root_scalar.
Parameters
----------
P float
Total pressure of the system, in bars.
T float
Temperature in degC
X_fluid float
Mole fraction of CO2 in the fluid
Returns
-------
float
Volume of the mixed fluid.
"""
if X_fluid != 1.0:
# x0 = self.volume(P,T,1.0)*X_fluid + self.volume_h(P,T)*(1-X_fluid)
# print(x0)
if P >= 20000 and T<800-273.15:
x0 = (X_fluid*25+(1-X_fluid)*15)
else:
x0 = (X_fluid*35+(1-X_fluid)*15)
else:
if P >= 20000 and T<800-273.15:
x0 = 25
else:
x0=35
return root_scalar(self.root_volume,x0=x0,x1=x0*0.9,args=(P,T,X_fluid)).root
def root_volume(self,v,P,T,X_fluid):
""" Returns the difference between the lhs and rhs of Eq (28) of Kerrick and Jacobs (1981).
For use with a root finder to obtain the volume of the mixed fluid.
Parameters
----------
v float
Guess for the volume
P float
Total system pressure in bars.
T float
Temperature in degC
X_fluid float
Mole fraction of CO2 in the fluid.
Returns
-------
float
Difference between lhs and rhs of Eq (28) of Kerrick and Jacobs (1981), in bars.
"""
T = T + 273.15
c = {}
h = {}
c['b'] = 58.0
c['c'] = (28.31 + 0.10721*T - 8.81e-6*T**2)*1e6
c['d'] = (9380.0 - 8.53*T + 1.189e-3*T**2)*1e6
c['e'] = (-368654.0 + 715.9*T + 0.1534*T**2)*1e6
h['b'] = 29.0
h['c'] = (290.78 - 0.30276*T + 1.4774e-4*T**2)*1e6#3
h['d'] = (-8374.0 + 19.437*T - 8.148e-3*T**2)*1e6
h['e'] = (76600.0 - 133.9*T + 0.1071*T**2)*1e6
if X_fluid == 1:
bm = c['b']
cm = c['c']
c12= c['c']
dm = c['d']
d12= c['d']
em = c['e']
e12 =c['e']
else:
bm = X_fluid*c['b'] + (1-X_fluid)*h['b']
c12 = (c['c']*h['c'])**0.5
cm = c['c']*X_fluid**2 + h['c']*(1-X_fluid)**2 + 2*X_fluid*(1-X_fluid)*c12
d12 = (c['d']*h['d'])**0.5
dm = c['d']*X_fluid**2 + h['d']*(1-X_fluid)**2 + 2*X_fluid*(1-X_fluid)*d12
e12 = (c['e']*h['e'])**0.5
em = c['e']*X_fluid**2 + h['e']*(1-X_fluid)**2 + 2*X_fluid*(1-X_fluid)*e12
am = cm + dm/v + em/v**2
y = bm/(4*v)
pt1 = (83.14 * T * (1 + y + y**2 - y**3)) / (v*(1-y)**3)
pt2 = - am / (T**0.5 * v * (v+bm))
return -(P - pt1 - pt2)
def volume_h(self,P,T):
""" Calculates the volume of a pure H2O fluid, by solving Eq (14) of
Kerrick and Jacobs (1981).
Parameters
----------
P float
Total pressure in bars.
T float
Temperature in degC.
Returns
-------
Difference between lhs and rhs of Eq (14) of Kerrick and Jacobs (1981), in bars.
"""
return root_scalar(self.root_volume_h,x0=15,x1=35,args=(P,T)).root
def root_volume_h(self,v,P,T):
""" Returns the difference between the lhs and rhs of Eq (14) of
Kerrick and Jacobs (1981). For use with a root solver to identify the
volume of a pure H2O fluid.
Parameters
----------
v float
Guess for the volume
P float
Total pressure in bars.
T float
Temperature in degC.
Returns
-------
float
The difference between the lhs and rhs of Eq (14) of Kerrick and Jacobs (1981),
in bars.
"""
T = T + 273.15
h = {}
h['b'] = 29.0
h['c'] = (290.78 - 0.30276*T + 1.4774e-4*T**2)*1e6#3
h['d'] = (-8374.0 + 19.437*T - 8.148e-3*T**2)*1e6
h['e'] = (76600.0 - 133.9*T + 0.1071*T**2)*1e6
h['a'] = h['c'] + h['d']/v + h['e']/v**2
y = h['b']/(4*v)
pt1 = (83.14 * T * (1 + y + y**2 - y**3)) / (v*(1-y)**3)
pt2 = - h['a'] / (T**0.5 * v * (v+h['b']))
return -(P - pt1 - pt2)
def lnPhi_mix(self,P,T,X_fluid):
""" Calculates the natural log of the fugacity coefficient for CO2 in a
mixed CO2-H2O fluid. Uses Eq (27) of Kerrick and Jacobs (1981).
Parameters
----------
P float
Total pressure in bars.
T float
Temperature in degC
X_fluid float
The mole fraction of CO2 in the fluid.
Returns
-------
float
The natural log of the fugacity coefficient for CO2 in a mixed fluid.
"""
T = T + 273.15
v = self.volume(P,T-273.15,X_fluid)
c = {}
h = {}
c['b'] = 58.0
c['c'] = (28.31 + 0.10721*T - 8.81e-6*T**2)*1e6
c['d'] = (9380.0 - 8.53*T + 1.189e-3*T**2)*1e6
c['e'] = (-368654.0 + 715.9*T + 0.1534*T**2)*1e6
h['b'] = 29.0
h['c'] = (290.78 - 0.30276*T + 1.4774e-4*T**2)*1e6#3
h['d'] = (-8374.0 + 19.437*T - 8.148e-3*T**2)*1e6
h['e'] = (76600.0 - 133.9*T + 0.1071*T**2)*1e6
if X_fluid == 1:
bm = c['b']
cm = c['c']
c12= c['c']
dm = c['d']
d12= c['d']
em = c['e']
e12 =c['e']
else:
bm = X_fluid*c['b'] + (1-X_fluid)*h['b']
c12 = (c['c']*h['c'])**0.5
cm = c['c']*X_fluid**2 + h['c']*(1-X_fluid)**2 + 2*X_fluid*(1-X_fluid)*c12
d12 = (c['d']*h['d'])**0.5
dm = c['d']*X_fluid**2 + h['d']*(1-X_fluid)**2 + 2*X_fluid*(1-X_fluid)*d12
e12 = (c['e']*h['e'])**0.5
em = c['e']*X_fluid**2 + h['e']*(1-X_fluid)**2 + 2*X_fluid*(1-X_fluid)*e12
am = cm + dm/v + em/v**2
y = bm/(4*v)
# Z = (1+y+y**2-y**3)/(1-y)**2 - am/(83.14*T**1.5*(v+bm))
Z = v*P/(83.14*T)
lnPhi = 0
lnPhi += (4*y-3*y**2)/(1-y)**2 + (c['b']/bm * (4*y-2*y**2)/(1-y)**3)
lnPhi += - (2*c['c']*X_fluid+2*(1-X_fluid)*c12)/(83.14*T**1.5*bm)*np.log((v+bm)/v)
lnPhi += - cm*c['b']/(83.14*T**1.5*bm*(v+bm))
lnPhi += cm*c['b']/(83.14*T**1.5*bm**2)*np.log((v+bm)/v)
lnPhi += - (2*c['d']*X_fluid+2*d12*(1-X_fluid)+dm)/(83.14*T**1.5*bm*v)
lnPhi += (2*c['d']*X_fluid+2*(1-X_fluid)*d12+dm)/(83.14*T**1.5*bm**2)*np.log((v+bm)/v)
lnPhi += c['b']*dm/(83.14*T**1.5*v*bm*(v+bm)) + 2*c['b']*dm/(83.14*T**1.5*bm**2*(v+bm))
lnPhi += - 2*c['b']*dm/(83.14*T**1.5*bm**3)*np.log((v+bm)/v)
lnPhi += - (2*c['e']*X_fluid + 2*(1-X_fluid)*e12+2*em)/(83.14*T**1.5*2*bm*v**2)
lnPhi += (2*c['e']*X_fluid+2*e12*(1-X_fluid)+2*em)/(83.14*T**1.5*bm**2*v)
lnPhi += - (2*c['e']*X_fluid+2*e12*(1-X_fluid)+2*em)/(83.14*T**1.5*bm**3)*np.log((v+bm)/v)
lnPhi += em*c['b']/(83.14*T**1.5*2*bm*v**2*(v+bm)) - 3*em*c['b']/(83.14*T**1.5*2*bm**2*v*(v+bm))
lnPhi += 3*em*c['b']/(83.14*T**1.5*bm**4)*np.log((v+bm)/v) - 3*em*c['b']/(83.14*T**1.5*bm**3*(v+bm))
lnPhi += - np.log(Z)
return lnPhi
class fugacity_KJ81_h2o(FugacityModel):
"""Implementation of the Kerrick and Jacobs (1981) EOS for mixed fluids. This class
will return the properties of the H2O component of the mixed fluid.
"""
def __init__(self):
self.set_calibration_ranges([CalibrationRange('pressure',20000.0,crf_LessThan,'bar','Kerrick and Jacobs (1981) EOS',
fail_msg=crmsg_LessThan_fail, pass_msg=crmsg_LessThan_pass, description_msg=crmsg_LessThan_description),
CalibrationRange('temperature',1050,crf_LessThan,'oC','Kerrick and Jacobs (1981) EOS',
fail_msg=crmsg_LessThan_fail, pass_msg=crmsg_LessThan_pass, description_msg=crmsg_LessThan_description)])
def fugacity(self,pressure,temperature,X_fluid,**kwargs):
""" Calculates the fugacity of H2O in a mixed CO2-H2O fluid. Above 1050C,
it assumes H2O and CO2 do not interact, as the equations are not defined
beyond this point.
Parameters
----------
pressure float
Total pressure of the system in bars.
temperature float
Temperature in degC
X_fluid float
Mole fraction of H2O in the fluid.
Returns
-------
float
fugacity of H2O in bars
"""
if X_fluid == 0:
return 0
elif temperature >= 1050:
return pressure*np.exp(self.lnPhi_mix(pressure,temperature,1.0))*X_fluid
else:
return pressure*np.exp(self.lnPhi_mix(pressure,temperature,X_fluid))*X_fluid
def volume(self,P,T,X_fluid):
""" Calculates the volume of the mixed fluid, by solving Eq (28) of Kerrick and
Jacobs (1981) using scipy.root_scalar.
Parameters
----------
P float
Total pressure of the system, in bars.
T float
Temperature in degC
X_fluid float
Mole fraction of H2O in the fluid
Returns
-------
float
Volume of the mixed fluid.
"""
if X_fluid != 1.0:
# x0 = self.volume(P,T,1.0)*X_fluid + self.volume_h(P,T)*(1-X_fluid)
# print(x0)
if P >= 20000 and T<800-273.15:
x0 = ((1-X_fluid)*25+X_fluid*15)
else:
x0 = ((1-X_fluid)*35+X_fluid*15)
else:
if P >= 20000 and T<800-273.15:
x0 = 10
else:
x0=15
return root_scalar(self.root_volume,x0=x0,x1=x0*0.9,args=(P,T,X_fluid)).root
def root_volume(self,v,P,T,X_fluid):
""" Returns the difference between the lhs and rhs of Eq (28) of Kerrick and Jacobs (1981).
For use with a root finder to obtain the volume of the mixed fluid.
Parameters
----------
v float
Guess for the volume
P float
Total system pressure in bars.
T float
Temperature in degC
X_fluid float
Mole fraction of H2O in the fluid.
Returns
-------
float
Difference between lhs and rhs of Eq (28) of Kerrick and Jacobs (1981), in bars.
"""
T = T + 273.15
c = {}
h = {}
c['b'] = 58.0
c['c'] = (28.31 + 0.10721*T - 8.81e-6*T**2)*1e6
c['d'] = (9380.0 - 8.53*T + 1.189e-3*T**2)*1e6
c['e'] = (-368654.0 + 715.9*T + 0.1534*T**2)*1e6
h['b'] = 29.0
h['c'] = (290.78 - 0.30276*T + 1.4774e-4*T**2)*1e6#3
h['d'] = (-8374.0 + 19.437*T - 8.148e-3*T**2)*1e6
h['e'] = (76600.0 - 133.9*T + 0.1071*T**2)*1e6
if X_fluid == 1:
bm = h['b']
cm = h['c']
dm = h['d']
em = h['e']
c12= h['c']
d12= h['d']
e12= h['e']
else:
bm = X_fluid*h['b'] + (1-X_fluid)*c['b']
c12 = (c['c']*h['c'])**0.5
cm = h['c']*X_fluid**2 + c['c']*(1-X_fluid)**2 + 2*X_fluid*(1-X_fluid)*c12
d12 = (c['d']*h['d'])**0.5
dm = h['d']*X_fluid**2 + c['d']*(1-X_fluid)**2 + 2*X_fluid*(1-X_fluid)*d12
e12 = (c['e']*h['e'])**0.5
em = h['e']*X_fluid**2 + c['e']*(1-X_fluid)**2 + 2*X_fluid*(1-X_fluid)*e12
am = cm + dm/v + em/v**2
y = bm/(4*v)
pt1 = (83.14 * T * (1 + y + y**2 - y**3)) / (v*(1-y)**3)
pt2 = - am / (T**0.5 * v * (v+bm))
return -(P - pt1 - pt2)
def volume_c(self,P,T):
""" Calculates the volume of a pure CO2 fluid, by solving Eq (14) of
Kerrick and Jacobs (1981).
Parameters
----------
P float
Total pressure in bars.
T float
Temperature in degC.
Returns
-------
Difference between lhs and rhs of Eq (14) of Kerrick and Jacobs (1981), in bars.
"""
return root_scalar(self.root_volume_c,x0=15,x1=35,args=(P,T)).root
def root_volume_c(self,v,P,T):
""" Returns the difference between the lhs and rhs of Eq (14) of
Kerrick and Jacobs (1981). For use with a root solver to identify the
volume of a pure H2O fluid.
Parameters
----------
v float
Guess for the volume
P float
Total pressure in bars.
T float
Temperature in degC.
Returns
-------
float
The difference between the lhs and rhs of Eq (14) of Kerrick and Jacobs (1981),
in bars.
"""
T = T + 273.15
c = {}
c['b'] = 58.0
c['c'] = (28.31 + 0.10721*T - 8.81e-6*T**2)*1e6
c['d'] = (9380.0 - 8.53*T + 1.189e-3*T**2)*1e6
c['e'] = (-368654.0 + 715.9*T + 0.1534*T**2)*1e6
c['a'] = c['c'] + c['d']/v + c['e']/v**2
y = c['b']/(4*v)
pt1 = (83.14 * T * (1 + y + y**2 - y**3)) / (v*(1-y)**3)
pt2 = - c['a'] / (T**0.5 * v * (v+c['b']))
return -(P - pt1 - pt2)
def lnPhi_mix(self,P,T,X_fluid):
""" Calculates the natural log of the fugacity coefficient for H2O in a
mixed CO2-H2O fluid. Uses Eq (27) of Kerrick and Jacobs (1981).
Parameters
----------
P float
Total pressure in bars.
T float
Temperature in degC
X_fluid float
The mole fraction of H2O in the fluid.
Returns
-------
float
The natural log of the fugacity coefficient for H2O in a mixed fluid.
"""
T = T + 273.15
v = self.volume(P,T-273.15,X_fluid)
c = {}
h = {}
c['b'] = 58.0
c['c'] = (28.31 + 0.10721*T - 8.81e-6*T**2)*1e6
c['d'] = (9380.0 - 8.53*T + 1.189e-3*T**2)*1e6
c['e'] = (-368654.0 + 715.9*T + 0.1534*T**2)*1e6
h['b'] = 29.0
h['c'] = (290.78 - 0.30276*T + 1.4774e-4*T**2)*1e6#3
h['d'] = (-8374.0 + 19.437*T - 8.148e-3*T**2)*1e6
h['e'] = (76600.0 - 133.9*T + 0.1071*T**2)*1e6
if X_fluid == 1:
bm = h['b']
cm = h['c']
dm = h['d']
em = h['e']
c12= h['c']
d12= h['d']
e12= h['e']
else:
bm = X_fluid*h['b'] + (1-X_fluid)*c['b']
c12 = (c['c']*h['c'])**0.5
cm = h['c']*X_fluid**2 + c['c']*(1-X_fluid)**2 + 2*X_fluid*(1-X_fluid)*c12
d12 = (c['d']*h['d'])**0.5
dm = h['d']*X_fluid**2 + c['d']*(1-X_fluid)**2 + 2*X_fluid*(1-X_fluid)*d12
e12 = (c['e']*h['e'])**0.5
em = h['e']*X_fluid**2 + c['e']*(1-X_fluid)**2 + 2*X_fluid*(1-X_fluid)*e12
am = cm + dm/v + em/v**2
y = bm/(4*v)
# Z = (1+y+y**2-y**3)/(1-y)**2 - am/(83.14*T**1.5*(v+bm))
Z = v*P/(83.14*T)
lnPhi = 0
lnPhi += (4*y-3*y**2)/(1-y)**2 + (h['b']/bm * (4*y-2*y**2)/(1-y)**3)
lnPhi += - (2*h['c']*X_fluid+2*(1-X_fluid)*c12)/(83.14*T**1.5*bm)*np.log((v+bm)/v)
lnPhi += - cm*h['b']/(83.14*T**1.5*bm*(v+bm))
lnPhi += cm*h['b']/(83.14*T**1.5*bm**2)*np.log((v+bm)/v)
lnPhi += - (2*h['d']*X_fluid+2*d12*(1-X_fluid)+dm)/(83.14*T**1.5*bm*v)
lnPhi += (2*h['d']*X_fluid+2*(1-X_fluid)*d12+dm)/(83.14*T**1.5*bm**2)*np.log((v+bm)/v)
lnPhi += h['b']*dm/(83.14*T**1.5*v*bm*(v+bm)) + 2*h['b']*dm/(83.14*T**1.5*bm**2*(v+bm))
lnPhi += - 2*h['b']*dm/(83.14*T**1.5*bm**3)*np.log((v+bm)/v)
lnPhi += - (2*h['e']*X_fluid + 2*(1-X_fluid)*e12+2*em)/(83.14*T**1.5*2*bm*v**2)
lnPhi += (2*h['e']*X_fluid+2*e12*(1-X_fluid)+2*em)/(83.14*T**1.5*bm**2*v)
lnPhi += - (2*h['e']*X_fluid+2*e12*(1-X_fluid)+2*em)/(83.14*T**1.5*bm**3)*np.log((v+bm)/v)
lnPhi += em*h['b']/(83.14*T**1.5*2*bm*v**2*(v+bm)) - 3*em*h['b']/(83.14*T**1.5*2*bm**2*v*(v+bm))
lnPhi += 3*em*h['b']/(83.14*T**1.5*bm**4)*np.log((v+bm)/v) - 3*em*h['b']/(83.14*T**1.5*bm**3*(v+bm))
lnPhi += - np.log(Z)
return lnPhi
class fugacity_ZD09_co2(FugacityModel):
""" Implementation of the Zhang and Duan (2009) fugacity model for pure CO2
fluids."""
def __init__(self):
self.set_calibration_ranges([CalibrationRange('pressure',[1,1e5],crf_Between,'bar','Zhang and Duan (2009) EOS',
fail_msg=crmsg_Between_fail, pass_msg=crmsg_Between_pass, description_msg=crmsg_Between_description),
CalibrationRange('temperature',[200,2300],crf_Between,'oC','Zhang and Duan (2009) EOS',
fail_msg=crmsg_Between_fail, pass_msg=crmsg_Between_pass, description_msg=crmsg_Between_description)])
def fugacity(self,pressure,temperature,X_fluid=1.0,**kwargs):
""" Calculates the fugacity of a pure CO2 fluid, or a mixed fluid assuming
ideal mixing. Implements eqn (14) of Zhang and Duan (2009).
Paramters
---------
pressure float
Pressure in bars
temperature float
Temperature in degC
X_fluid float
Mole fraction of CO2 in the fluid. Default is 1.0.
Returns
-------
float
Fugacity of CO2, standard state 1 bar.
"""
P = pressure/10
T = temperature + 273.15
a = np.array([0.0,
2.95177298930e-2,
-6.33756452413e3,
-2.75265428882e5,
1.29128089283e-3,
-1.45797416153e2,
7.65938947237e4,
2.58661493537e-6,
0.52126532146,
-1.39839523753e2,
-2.36335007175e-8,
5.35026383543e-3,
-0.27110649951,
2.50387836486e4,
0.73226726041,
1.5483335997e-2])
e = 235.0
s = 3.79
Pm = 3.0636*P*s**3/e
Tm = 154*T/e
Vm = root_scalar(self.Vm,x0=200,x1=100,args=(P,T)).root
S1 = ((a[1]+a[2]/Tm**2+a[3]/Tm**3)/Vm+
(a[4]+a[5]/Tm**2+a[6]/Tm**3)/(2*Vm**2)+
(a[7]+a[8]/Tm**2+a[9]/Tm**3)/(4*Vm**4)+
(a[10]+a[11]/Tm**2+a[12]/Tm**3)/(5*Vm**5)+
(a[13]/(2*a[15]*Tm**3)*(a[14]+1-(a[14]+1+a[15]/Vm**2)*
np.exp(-a[15]/Vm**2)))
)
Z = Pm*Vm/(8.314*Tm)
lnfc = Z - 1 - np.log(Z) + S1
return P*np.exp(lnfc)*10
def Vm(self,Vm,P,T):
""" Function to use for solving for the parameter Vm, defined by eqn (8) of
Zhang and Duan (2009). Called by scipy.fsolve in the fugacity method.
Parameters
----------
Vm float
Guessed value of Vm
P float
Pressure in MPa
T float
Temperature in K
Returns
-------
float
Difference between (rearranged) LHS and RHS of eqn (8) of Zhang and Duan (2009).
"""
Pm = 3.0636*P*3.79**3/235.0
Tm = 154*T/235.0
a = np.array([0.0,
2.95177298930e-2,
-6.33756452413e3,
-2.75265428882e5,
1.29128089283e-3,
-1.45797416153e2,
7.65938947237e4,
2.58661493537e-6,
0.52126532146,
-1.39839523753e2,
-2.36335007175e-8,
5.35026383543e-3,
-0.27110649951,
2.50387836486e4,
0.73226726041,
1.5483335997e-2])
return ((1+(a[1]+a[2]/Tm**2+a[3]/Tm**3)/Vm+
(a[4]+a[5]/Tm**2+a[6]/Tm**3)/Vm**2+
(a[7]+a[8]/Tm**2+a[9]/Tm**3)/Vm**4)*0.08314*Tm/Pm - Vm
)
class fugacity_MRK_co2(FugacityModel):
""" Modified Redlick Kwong fugacity model as used by VolatileCalc. Python implementation by
<NAME> (github.com/DJRgeoscience/VolatileCalcForPython), based on VB code by Newman &
Lowenstern.
"""
def __init__(self):
self.set_calibration_ranges([])
def fugacity(self,pressure,temperature,X_fluid=1.0,**kwargs):
""" Calculates the fugacity of CO2 in a pure or mixed H2O-CO2 fluid (assuming ideal mixing).
Parameters
----------
pressure float
Total pressure of the system in bars.
temperature float
Temperature in degC
X_fluid float
Mole fraction of CO2 in the fluid.
Returns
-------
float
fugacity of CO2 in bars
"""
fug = self.MRK(pressure,temperature+273.15)
return fug*X_fluid
def FNA(self,TK):
return (166800000 - 193080 * (TK - 273.15) + 186.4 * (TK - 273.15)**2 - 0.071288 * ((TK - 273.15)**3)) * 1.01325
def FNB(self,TK):
return 1.01325 * (73030000 - 71400 * (TK - 273.15) + 21.57 * (TK - 273.15)**2)
def FNC(self,TK):
R = 83.14321
return 1.01325 * (np.exp(-11.071 + 5953 / TK - 2746000 / TK**2 + 464600000 / TK**3) * 0.5 * R * R * TK**2.5 / 1.02668 + 40123800)
def FNF(self,V,TK,A,B,P):
R = 83.14321
return R * TK / (V - B) - A / ((V * V + B * V) * TK**0.5) - P
def MRK(self,P,TK): #Redlich-Kwong routine to estimate endmember H2O and CO2 fugacities
R = 83.14321
B_1 = 14.6
B_2 = 29.7
for X_1 in [0,1]:
B = X_1 * B_1 + (1 - X_1) * B_2
A = X_1**2 * self.FNA(TK) + 2 * X_1 * (1 - X_1) * self.FNC(TK) + (1 - X_1)**2 * self.FNB(TK)
Temp2 = B + 5
Q = 1
Temp1 = 0
while abs(Temp2 - Temp1) >= 0.00001:
Temp1 = Temp2
F_1 = (self.FNF(Temp1 + 0.01, TK, A, B, P) - self.FNF(Temp1, TK, A, B, P)) / 0.01
Temp2 = Temp1 - Q * self.FNF(Temp1, TK, A, B, P) / F_1
F_2 = (self.FNF(Temp2 + 0.01, TK, A, B, P) - self.FNF(Temp2, TK, A, B, P)) / 0.01
if F_2 * F_1 <= 0:
Q = Q / 2.
if abs(Temp2 - Temp1) > 0.00001:
F_1 = F_2
V = Temp2
G_1 = np.log(V / (V - B)) + B_1 / (V - B) - 2 * (X_1 * self.FNA(TK) + (1 - X_1) * self.FNC(TK)) * np.log((V + B) / V) / (R * TK**1.5 * B)
G_1 = G_1 + (np.log((V + B) / V) - B / (V + B)) * A * B_1 / (R * TK**1.5 * B**2) - np.log(P * V / (R * TK))
G_1 = np.exp(G_1)
G_2 = np.log(V / (V - B)) + B_2 / (V - B) - 2 * (X_1 * self.FNC(TK) + (1 - X_1) * self.FNB(TK)) * np.log((V + B) / V) / (R * TK**1.5 * B)
G_2 = G_2 + (np.log((V + B) / V) - B / (V + B)) * A * B_2 / (R * TK**1.5 * B**2) - np.log(P * V / (R * TK))
G_2 = np.exp(G_2)
if X_1 == 0:
fCO2o = G_2 * P #The fugacity of CO2
# return fCO2o
if X_1 == 1:
fH2Oo = G_1 * P #The fugacity of H2O
# return fH2Oo
return fCO2o
class fugacity_MRK_h2o(FugacityModel):
""" Modified Redlick Kwong fugacity model as used by VolatileCalc. Python implementation by
<NAME> (github.com/DJRgeoscience/VolatileCalcForPython), based on VB code by Newman &
Lowenstern.
"""
def __init__(self):
self.set_calibration_ranges([])
def fugacity(self,pressure,temperature,X_fluid=1.0,**kwargs):
""" Calculates the fugacity of H2O in a pure or mixed H2O-CO2 fluid (assuming ideal mixing).
Parameters
----------
pressure float
Total pressure of the system in bars.
temperature float
Temperature in degC
X_fluid float
Mole fraction of H2O in the fluid.
Returns
-------
float
fugacity of CO2 in bars
"""
fug = self.MRK(pressure,temperature+273.15)
return fug*X_fluid
def FNA(self,TK):
return (166800000 - 193080 * (TK - 273.15) + 186.4 * (TK - 273.15)**2 - 0.071288 * ((TK - 273.15)**3)) * 1.01325
def FNB(self,TK):
return 1.01325 * (73030000 - 71400 * (TK - 273.15) + 21.57 * (TK - 273.15)**2)
def FNC(self,TK):
R = 83.14321
return 1.01325 * (np.exp(-11.071 + 5953 / TK - 2746000 / TK**2 + 464600000 / TK**3) * 0.5 * R * R * TK**2.5 / 1.02668 + 40123800)
def FNF(self,V,TK,A,B,P):
R = 83.14321
return R * TK / (V - B) - A / ((V * V + B * V) * TK**0.5) - P
def MRK(self,P,TK): #Redlich-Kwong routine to estimate endmember H2O and CO2 fugacities
R = 83.14321
B_1 = 14.6
B_2 = 29.7
# X_1 = 1
for X_1 in [0,1]:
B = X_1 * B_1 + (1 - X_1) * B_2
A = X_1**2 * self.FNA(TK) + 2 * X_1 * (1 - X_1) * self.FNC(TK) + (1 - X_1)**2 * self.FNB(TK)
Temp2 = B + 5
Q = 1
Temp1 = 0
while abs(Temp2 - Temp1) >= 0.00001:
Temp1 = Temp2
F_1 = (self.FNF(Temp1 + 0.01, TK, A, B, P) - self.FNF(Temp1, TK, A, B, P)) / 0.01
Temp2 = Temp1 - Q * self.FNF(Temp1, TK, A, B, P) / F_1
F_2 = (self.FNF(Temp2 + 0.01, TK, A, B, P) - self.FNF(Temp2, TK, A, B, P)) / 0.01
if F_2 * F_1 <= 0:
Q = Q / 2.
if abs(Temp2 - Temp1) > 0.00001:
F_1 = F_2
V = Temp2
G_1 = np.log(V / (V - B)) + B_1 / (V - B) - 2 * (X_1 * self.FNA(TK) + (1 - X_1) * self.FNC(TK)) * np.log((V + B) / V) / (R * TK**1.5 * B)
G_1 = G_1 + (np.log((V + B) / V) - B / (V + B)) * A * B_1 / (R * TK**1.5 * B**2) - np.log(P * V / (R * TK))
G_1 = np.exp(G_1)
G_2 = np.log(V / (V - B)) + B_2 / (V - B) - 2 * (X_1 * self.FNC(TK) + (1 - X_1) * self.FNB(TK)) * np.log((V + B) / V) / (R * TK**1.5 * B)
G_2 = G_2 + (np.log((V + B) / V) - B / (V + B)) * A * B_2 / (R * TK**1.5 * B**2) - np.log(P * V / (R * TK))
G_2 = np.exp(G_2)
if X_1 == 0:
fCO2o = G_2 * P #The fugacity of CO2
# return fCO2o
if X_1 == 1:
fH2Oo = G_1 * P #The fugacity of H2O
# return fH2Oo
return fH2Oo
class fugacity_HB_co2(FugacityModel):
"""
Implementation of the Holloway and Blank (1994) Modified Redlich Kwong EoS for CO2.
"""
def __init__(self):
self.set_calibration_ranges([CalibrationRange('pressure',[1,1e5],crf_Between,'bar','Redlich Kwong EOS',
fail_msg=crmsg_Between_fail, pass_msg=crmsg_Between_pass, description_msg=crmsg_Between_description),
CalibrationRange('temperature',500.0,crf_GreaterThan,'oC','Redlich Kwong EOS',
fail_msg=crmsg_GreaterThan_fail, pass_msg=crmsg_GreaterThan_pass, description_msg=crmsg_GreaterThan_description)])
self.HBmodel = fugacity_HollowayBlank()
def fugacity(self,pressure,temperature,X_fluid=1.0,**kwargs):
return self.HBmodel.fugacity(pressure=pressure, temperature=temperature, species='CO2')*X_fluid
class fugacity_HB_h2o(FugacityModel):
"""
Implementation of the Holloway and Blank (1994) Modified Redlich Kwong EoS for H2O.
"""
def __init__(self):
self.set_calibration_ranges([CalibrationRange('pressure',[1,1e5],crf_Between,'bar','Redlich Kwong EOS',
fail_msg=crmsg_Between_fail, pass_msg=crmsg_Between_pass, description_msg=crmsg_Between_description),
CalibrationRange('temperature',500.0,crf_GreaterThan,'oC','Redlich Kwong EOS',
fail_msg=crmsg_GreaterThan_fail, pass_msg=crmsg_GreaterThan_pass, description_msg=crmsg_GreaterThan_description)])
self.HBmodel = fugacity_HollowayBlank()
def fugacity(self,pressure,temperature,X_fluid=1.0,**kwargs):
return self.HBmodel.fugacity(pressure=pressure, temperature=temperature, species='H2O')*X_fluid
class fugacity_HollowayBlank(FugacityModel):
"""
Implementation of the Modified Redlich Kwong presented in Holloway and Blank (1994) Reviews
in Mineralogy and Geochemistry vol. 30. Originally written in Quickbasic. CO2 calculations
translated to Matlab by <NAME> and translated to python by <NAME> for VESIcal.
H2O calculations translated to VisualBasic by <NAME> and translated to python by
<NAME> for VESIcal.
"""
def __init__(self):
self.set_calibration_ranges([CalibrationRange('pressure',[1,1e5],crf_Between,'bar','MRK EOS (Holloway and Blank, 1994)',
fail_msg=crmsg_Between_fail, pass_msg=crmsg_Between_pass, description_msg=crmsg_Between_description),
CalibrationRange('temperature',500,crf_GreaterThan,'oC','MRK EOS (Holloway and Blank, 1994)',
fail_msg=crmsg_GreaterThan_fail, pass_msg=crmsg_GreaterThan_pass, description_msg=crmsg_GreaterThan_description)])
def REDKW(self, BP, A2B):
"""
The RK routine. A routine to calculate compressibility factor and fugacity coefficient
with the Redlich-Kwong equation following Edmister (1968). This solution for supercritical
fluid.
Parameters
----------
BP: float
B parameter sum from RKCALC
A2B: float
A parameter sum from RKCALC
Returns
-------
float
XLNFP (fugacity coefficient?)
"""
if A2B < 1*10**(-10):
A2B = 0.001
#Define constants
TH = 0.333333
RR = -A2B*BP**2
QQ = BP*(A2B-BP-1)
XN = QQ*TH+RR-0.074074
XM = QQ-TH
XNN = XN*XN*0.25
XMM = XM**3 / 27.0
ARG = XNN+XMM
if ARG > 0:
X = np.sqrt(ARG)
F = 1
XN2 = -XN*0.5
iXMM = XN2+X
if iXMM < 0:
F = -1
XMM = F*((F*iXMM)**TH)
F = 1
iXNN = XN2 - X
if iXNN < 0:
F = -1
XNN = F*((F*iXNN)**TH)
Z = XMM+XNN+TH
ZBP = Z-BP
if ZBP < 0.000001:
ZBP = 0.000001
BPZ = 1+BP/Z
FP = Z-1-np.log(ZBP)-A2B*np.log(BPZ)
if FP < -37 or FP > 37:
FP = 0.000001
elif ARG <0:
COSPHI = np.sqrt(-XNN/XMM)
if XN > 0:
COSPHI = -COSPHI
TANPHI = np.sqrt(1-COSPHI**2)/COSPHI
PHI = np.arctan(TANPHI)*TH
FAC = 2*np.sqrt(-XM*TH)
#sort for largest root
R1 = np.cos(PHI)
R2 = np.cos(PHI+2.0944)
R3 = np.cos(PHI+4.18879)
RH = R2
if R1 > R2:
RH = R1
if R3 > RH:
RH = R3
Z = RH*FAC+TH
ZBP = Z-BP
if ZBP < 0.000001:
ZBP = 0.000001
BPZ = 1+BP/Z
FP = Z-1-np.log(ZBP)-A2B*np.log(BPZ)
if FP < -37 or FP > 37:
FP = 0.000001
else:
FP = 1
Z = 1
XLNFP = FP
return XLNFP
def Saxena(self, TK, pb):
"""
High pressure corresponding states routines from Saxena and Fei (1987) GCA
vol. 51, 783-791.
Parameters
----------
TK: float
Temperature in K.
pb: float
Pressure in bars.
Returns
-------
float
XLNF, Natural log of the ratio F(P)/F(4000 bar)
"""
#Define integration limit
PO = 4000
#Critical temperatures and pressures for CO2
TR = TK/304.2
PR = pb/73.9
PC = 73.9
#Virial coeficients
A = 2.0614-2.2351/TR**2 - 0.39411*np.log(TR)
B = 0.055125/TR + 0.039344/TR**2
C = -1.8935*10**(-6)/TR - 1.1092*10**(-5)/TR**2 - 2.1892*10**(-5)/TR**3
D = 5.0527*10**(-11)/TR - 6.3033*10**(-21)/TR**3
#Calculate molar volume
Z = A+B*PR+C*PR**2+D*PR**3
V = Z*83.0117*TK/pb
#integrate from PO (4000 bars) to P to calculate ln fugacity
LNF = A*np.log(pb/PO)+(B/PC)*(pb-PO)+(C/(2*PC**2))*(pb**2-PO**2)
LNF = LNF+(D/(3*PC**3))*(pb**3-PO**3)
XLNF = LNF
return XLNF
def RKCALC(self, temperature, pressure, species):
"""
Calculation of pure gas MRK properties following Holloway 1981, 1987
Parameters
----------
temperature: float
Temperature in degrees K.
pressure: float
Pressure in atmospheres.
Returns
-------
float
Natural log of the fugacity of a pure gas.
"""
#Define constants
R = 82.05736
RR = 6732.2
pb = 1.013*pressure
PBLN = np.log(pb)
TCEL = temperature-273.15
RXT = R*temperature
RT = R*temperature**1.5 * 10**(-6)
if species == 'CO2':
#Calculate T-dependent MRK A parameter CO2
ACO2M = 73.03 - 0.0714*TCEL + 2.157*10**(-5)*TCEL**2
#Define MRK B parameter for CO2
BSUM = 29.7
ASUM = ACO2M / (BSUM*RT)
elif species == 'H2O':
#Calculate T-dependent MRK A parameter H2O
AH2OM = 115.98 - np.double(0.0016295)*temperature - 1.4984*10**(-5)*temperature**2
#Define MRK B parameter for H2O
BSUM = 14.5
ASUM = AH2OM / (BSUM*RT)
BSUM = pressure*BSUM/RXT
XLNFP = self.REDKW(BSUM, ASUM)
#Convert to ln(fugacity)
PUREG = XLNFP + PBLN
return PUREG
def fugacity(self, pressure, temperature, species, **kwargs):
"""
Calculates fugacity.
Parameters
----------
temperature: float
Temperature in degrees C.
pressure: float
Pressure in bars.
species: str
Choose which species to calculate. Options are 'H2O' and 'CO2'.
Returns
-------
float
Fugacity coefficient for passed species
"""
#convert temp and press to atmospheres and Kelvin
pressureAtmo = pressure/1.013
temperatureK = temperature + 273.15
PO = 4000/1.013
#Use the MRK below 4,000 bars, Saxena above 4,000 bars
if pressure > 4000 and species=='CO2':
iPUREG = self.RKCALC(temperatureK, PO, species)
XLNF = self.Saxena(temperatureK, pressure)
PUREG = iPUREG + XLNF
else:
PUREG = self.RKCALC(temperatureK, pressureAtmo, species)
#Convert from ln(fugacity) to fugacity
stdf = np.exp(PUREG)
return stdf
class fugacity_RK_co2(FugacityModel):
"""
Implementation of the Redlich Kwong EoS for CO2.
Code derived from http://people.ds.cam.ac.uk/pjb10/thermo/pure.html - <NAME> 30 October 2003.
"""
def __init__(self):
self.set_calibration_ranges([CalibrationRange('pressure',[1,1e5],crf_Between,'bar','Redlich Kwong EOS',
fail_msg=crmsg_Between_fail, pass_msg=crmsg_Between_pass, description_msg=crmsg_Between_description),
CalibrationRange('temperature',[500],crf_GreaterThan,'oC','Redlich Kwong EOS',
fail_msg=crmsg_GreaterThan_fail, pass_msg=crmsg_GreaterThan_pass, description_msg=crmsg_GreaterThan_description)])
# self.set_calibration_ranges([cr_Between('pressure',[1.0,1e5],'bar','Redlich Kwong EOS'),
# cr_GreaterThan('temperature',500,'oC','Redlich Kwong EOS')])
self.RKmodel = fugacity_RedlichKwong()
def fugacity(self,pressure,temperature,X_fluid,**kwargs):
return self.RKmodel.fugacity(pressure, temperature, X_fluid, 'CO2')
class fugacity_RK_h2o(FugacityModel):
"""
Implementation of the Redlich Kwong EoS for H2O.
Code derived from http://people.ds.cam.ac.uk/pjb10/thermo/pure.html - <NAME> 30 October 2003.
"""
def __init__(self):
self.set_calibration_ranges([CalibrationRange('pressure',[1,1e5],crf_Between,'bar','Redlich Kwong EOS',
fail_msg=crmsg_Between_fail, pass_msg=crmsg_Between_pass, description_msg=crmsg_Between_description),
CalibrationRange('temperature',500,crf_GreaterThan,'oC','Redlich Kwong EOS',
fail_msg=crmsg_GreaterThan_fail, pass_msg=crmsg_GreaterThan_pass, description_msg=crmsg_GreaterThan_description)])
self.RKmodel = fugacity_RedlichKwong()
def fugacity(self,pressure,temperature,X_fluid,**kwargs):
return self.RKmodel.fugacity(pressure, temperature, X_fluid, 'H2O')
class fugacity_RedlichKwong(FugacityModel):
"""
Implementation of the Redlich Kwong EoS
Code derived from http://people.ds.cam.ac.uk/pjb10/thermo/pure.html - <NAME> 30 October 2003.
"""
def __init__(self):
self.set_calibration_ranges([CalibrationRange('pressure',[1,1e5],crf_Between,'bar','Redlich Kwong EOS',
fail_msg=crmsg_Between_fail, pass_msg=crmsg_Between_pass, description_msg=crmsg_Between_description),
CalibrationRange('temperature',500,crf_GreaterThan,'oC','Redlich Kwong EOS',
fail_msg=crmsg_GreaterThan_fail, pass_msg=crmsg_GreaterThan_pass, description_msg=crmsg_GreaterThan_description)])
def gamma(self, pressure, temperature, species):
"""
Calculates fugacity coefficients.
Parameters
----------
temperature: fload
Temperature in degrees C.
pressure: float
Pressure in bars.
species: str
Choose which species to calculate. Options are 'H2O' and 'CO2'.
Returns
-------
float
Fugacity coefficient for passed species.
"""
temperatureK = temperature + 273.15
R = 8.3145
fluid_species_names = ['CO2', 'H2O']
critical_params = {'CO2':{ "cT": 304.15,
"cP": 73.8659,
"o": 0.225
},
'H2O':{ "cT": 647.25,
"cP": 221.1925,
"o": 0.334
}
}
#Calculate a and b parameters (depend only on critical parameters)...
a = 0.42748 * R**2.0 * critical_params[species]["cT"]**(2.5) / (critical_params[species]["cP"] * 10.0**5)
b = 0.08664 * R * critical_params[species]["cT"] / (critical_params[species]["cP"] * 10.0**5)
kappa = 0.0
#Calculate coefficients in the cubic equation of state...
#coeffs: (C0, C1, C2, A, B)
A = a * pressure * 10.0**5 / (np.sqrt(temperatureK) * (R * temperatureK)**2.0)
B = b * pressure * 10.0**5 / (R * temperatureK)
C2 = -1.0
C1 = A - B - B * B
C0 = -A * B
#Solve the cubic equation for Z0 - Z2, D...
Q1 = C2 * C1 / 6.0 - C0 / 2.0 - C2**3.0 / 27.0
P1 = C2**2.0 / 9.0 - C1 / 3.0
D = Q1**2.0 - P1**3.0
if D >= 0:
kOneThird = 1.0 / 3.0
absQ1PSqrtD = np.fabs(Q1 + np.sqrt(D))
temp1 = absQ1PSqrtD**kOneThird
temp1 *= (Q1 + np.sqrt(D)) / absQ1PSqrtD
absQ1MSqrtD = np.fabs(Q1 - np.sqrt(D))
temp2 = absQ1MSqrtD**kOneThird
temp2 *= (Q1 - np.sqrt(D)) / absQ1MSqrtD
Z0 = temp1 + temp2 - C2 / 3.0
else:
temp1 = Q1**2.0 / (P1**3.0)
temp2 = np.sqrt(1.0 - temp1) / np.sqrt(temp1)
temp2 *= Q1 / np.fabs(Q1)
gamma = np.arctan(temp2)
if gamma < 0:
gamma = gamma + np.pi
Z0 = 2.0 * np.sqrt(P1) * np.cos(gamma/3.0) - C2 / 3.0
Z1 = 2.0 * np.sqrt(P1) * np.cos((gamma + 2.0 * np.pi) / 3.0) - C2/3.0
Z2 = 2.0 * np.sqrt(P1) * np.cos((gamma + 4.0 * np.pi) / 3.0) - C2/3.0
if Z0 < Z1:
temp0 = Z0
Z0 = Z1
Z1 = temp0
if Z1 < Z2:
temp0 = Z1
Z1 = Z2
Z2 = temp0
if Z0 < Z1:
temp0 = Z0
Z0 = Z1
Z1 = temp0
#Calculate Departure Functions
gamma = np.exp(Z0 - 1.0 - np.log(Z0-B) - A * np.log(1.0+B/Z0)/B)
Hdep = R * temperatureK * (Z0 - 1.0 - 1.5*A*np.log(1.0+B/Z0)/B)
Sdep = R * (np.log(Z0-B) - 0.5*A*np.log(1.0+B/Z0)/B)
return gamma
def fugacity(self, pressure, temperature, X_fluid=1.0, species='H2O', **kwargs):
"""
Calculates the fugacity of H2O in a mixed H2O-CO2 fluid using the universal relationships:
P_i = f_i/gamma_i = (fpure_i * Xfluid_i) / gamma_i
See Iacovino (2015) EPSL for further explanation.
"""
gammaH2O = self.gamma(pressure, temperature, 'H2O')
gammaCO2 = self.gamma(pressure, temperature, 'CO2')
fugacityH2Opure = pressure * gammaH2O
fugacityCO2pure = pressure * gammaCO2
if species == 'H2O':
return fugacityH2Opure * X_fluid
elif species == 'CO2':
return fugacityCO2pure * X_fluid
else:
raise InputError("Species must be H2O or CO2.")
#---------------ACTVITY MODELS-------------------------------#
class activity_idealsolution(activity_model):
""" Implements an ideal solution activity model, i.e. it
will always return the mole fraction.
"""
def activity(self,X):
""" The activity of the component in an ideal solution, i.e., it
will return the mole fraction.
Parameters
----------
X float
The mole fraction of the species in the solution.
Returns
-------
float
The activity of the species in the solution, i.e., the mole fraction.
"""
return X
#------------PURE FLUID MODELS-------------------------------#
class ShishkinaCarbon(Model):
""" Implementation of the Shishkina et al. (2014) carbon solubility model, as a Model class.
"""
def __init__(self):
self.set_volatile_species(['CO2'])
self.set_fugacity_model(fugacity_idealgas())
self.set_activity_model(activity_idealsolution())
self.set_solubility_dependence(False)
self.set_calibration_ranges([CalibrationRange('pressure',[500.0,5000.0],crf_Between,'bar','Shishkina et al. carbon',
fail_msg=crmsg_Between_fail, pass_msg=crmsg_Between_pass, description_msg=crmsg_Between_description),
CalibrationRange('temperature',[1200.0,1250.0],crf_Between,'oC','Shishkina et al. carbon',
fail_msg=crmsg_Between_fail, pass_msg=crmsg_Between_pass, description_msg=crmsg_Between_description)])
def preprocess_sample(self,sample):
""" Returns sample, unmodified. The Pi* compositional parameter is a ratio of cations,
therefore the value is not affected by the normalization of the sample. Shishkina et al.
imply the accuracy of the calculations are little affected whether Fe(tot) or Fe2+ is
used.
Parameters
----------
sample: dict or pandas Series
The major element oxides in wt%.
Returns
-------
dict or pandas Series
The major element oxides in wt%.
"""
if type(sample) != dict and type(sample) != pd.core.series.Series:
raise InputError("sample must be a dict or a pandas Series.")
return sample
def PiStar(self,sample):
"""Shishkina et al. (2014) Eq (11)
Calculates the Pi* parameter for use in calculating CO2 solubility.
Parameters
----------
sample: pandas Series or dict
Major element oxides in wt%.
Returns
-------
float
The value of the Pi* compositional parameter.
"""
_mols = wtpercentOxides_to_molCations(sample)
if all(cation in _mols for cation in ['Ca','K','Na','Mg','Fe','Si','Al']) == False:
raise InputError("To calculate PiStar, values for CaO, K2O, Na2O, MgO, FeO, SiO2, and Al2O3\
must be provided in sample.")
_pi = (_mols['Ca'] + 0.8*_mols['K'] + 0.7*_mols['Na'] + 0.4*_mols['Mg'] + 0.4*_mols['Fe'])/\
(_mols['Si']+_mols['Al'])
return _pi
def calculate_dissolved_volatiles(self,pressure,sample,X_fluid=1,**kwargs):
""" Calculates the dissolved CO2 concentration in wt%, using equation (13) of Shishkina et al. (2014).
Parameters
----------
pressure: float
(Total) pressure in bars.
sample: dict or pandas Series
Major element concentrations in wt%. Normalization does not matter.
X_fluid: float
The mol-fraction of the fluid that is CO2. Default is 1, i.e. a pure CO2 fluid.
Returns
-------
float
The dissolved CO2 concentration in wt%.
"""
if X_fluid < 0 or X_fluid > 1:
raise InputError("X_fluid must have a value between 0 and 1.")
if pressure < 0:
raise InputError("pressure must be a positive value.")
PiStar = self.PiStar(sample)
fugacity = self.fugacity_model.fugacity(pressure=pressure,X_fluid=X_fluid,**kwargs)
A = 1.150
B = 6.71
C= -1.345
if fugacity == 0:
return 0
else:
return np.exp(A*np.log(fugacity/10)+B*PiStar+C)/1e4
def calculate_equilibrium_fluid_comp(self,pressure,sample,**kwargs):
""" Returns 1.0 if a pure CO2 fluid is saturated.
Returns 0.0 if a pure CO2 fluid is undersaturated.
Parameters
----------
pressure float
The total pressure of the system in bars.
sample dict or pandas Series
Major element oxides in wt%
Returns
-------
float
1.0 if CO2-fluid saturated, 0.0 otherwise.
"""
if self.calculate_saturation_pressure(sample=sample,**kwargs) < pressure:
return 0.0
else:
return 1.0
def calculate_saturation_pressure(self,sample,**kwargs):
""" Calculates the pressure at which a pure CO2 fluid is saturated, for the given
sample composition and CO2 concentration. Calls the scipy.root_scalar routine, which makes
repeated calls to the calculate_dissolved_volatiles method.
Parameters
----------
sample dict or pandas Series
Major elements in wt%, including CO2 (also in wt%).
Returns
-------
float
Saturation pressure in bar
"""
if 'CO2' not in sample:
raise InputError("sample must contain CO2.")
if sample['CO2'] < 0:
raise InputError("CO2 concentration must be greater than 0 wt%.")
try:
satP = root_scalar(self.root_saturation_pressure,bracket=[1e-15,1e5],args=(sample,kwargs)).root
except:
warnings.warn("Saturation pressure not found.",RuntimeWarning)
satP = np.nan
return satP
def root_saturation_pressure(self,pressure,sample,kwargs):
""" Function called by scipy.root_scalar when finding the saturation pressure using
calculate_saturation_pressure.
Parameters
----------
pressure float
Pressure guess in bars
sample dict or pandas Series
Major element oxides in wt%, including CO2 (also in wt%).
kwargs dictionary
Additional keyword arguments supplied to calculate_saturation_pressure. Might be required for
the fugacity or activity models.
Returns
-------
float
The differece between the dissolved CO2 at the pressure guessed, and the CO2 concentration
passed in the sample variable.
"""
return self.calculate_dissolved_volatiles(pressure=pressure,sample=sample,**kwargs)-sample['CO2']
class ShishkinaWater(Model):
""" Implementation of the Shishkina et al. (2014) H2O solubility model as a Model class.
"""
def __init__(self):
self.set_volatile_species(['H2O'])
self.set_fugacity_model(fugacity_idealgas())
self.set_activity_model(activity_idealsolution())
self.set_solubility_dependence(False)
self.set_calibration_ranges([CalibrationRange('pressure',[500.0,5000.0],crf_Between,'bar','Shishkina et al. water',
fail_msg=crmsg_Between_fail, pass_msg=crmsg_Between_pass, description_msg=crmsg_Between_description),
CalibrationRange('temperature',[1200.0,1250.0],crf_Between,'oC','Shishkina et al. water',
fail_msg=crmsg_Between_fail, pass_msg=crmsg_Between_pass, description_msg=crmsg_Between_description)])
def preprocess_sample(self,sample):
""" Returns sample, renormlized so that the major element oxides (excluding volatiles) sum to 100%.
Normalization must be done this way as the compositional dependence of the solubility takes the
mole fractions of Na2O and K2O as inputs, presumably assuming no volatiles in the bulk composition.
Volatile concentrations are left unchanged.
Parameters
----------
sample: dict or pandas Series
The major element oxides in wt%.
Returns
-------
dict or pandas Series
The major element oxides in wt%.
"""
return normalize_AdditionalVolatiles(sample)
def calculate_dissolved_volatiles(self,pressure,sample,X_fluid=1.0,**kwargs):
"""Calculates the dissolved H2O concentration using Eqn (9) of Shishkina et al. (2014).
Parameters
----------
pressure float
Total pressure in bars
sample pandas Series or dict
Major element oxides in wt%. Normalized to zero-volatiles so that the total-alkalis
mol fraction can be determined accurately.
X_fluid float
The mol fraction of H2O in the fluid
Returns
-------
float
The H2O concentration in wt%
"""
if type(sample) != dict and type(sample) != pd.core.series.Series:
raise InputError("sample must be a dict or pandas Series.")
if all(ox in sample for ox in ['Na2O','K2O']) == False:
raise InputError("Na2O and K2O must be present in sample.")
if pressure < 0:
raise InputError("Pressure must be positive.")
_mols = wtpercentOxides_to_molCations(sample)
_mol_volatiles = 0
if 'H' in _mols:
_mol_volatiles += _mols['H']
if 'C' in _mols:
_mol_volatiles += _mols['C']
total_alkalis = (_mols['Na'] + _mols['K'])/(1-_mol_volatiles)
fugacity = self.fugacity_model.fugacity(pressure,X_fluid=X_fluid,**kwargs)
a = 3.36e-7 * (fugacity/10)**3 - 2.33e-4*(fugacity/10)**2 + 0.0711*(fugacity/10) - 1.1309
b = -1.2e-5*(fugacity/10)**2 + 0.0196*(fugacity/10)+1.1297
return a*total_alkalis + b
def calculate_equilibrium_fluid_comp(self,pressure,sample,**kwargs):
""" Returns 1.0 if a pure H2O fluid is saturated.
Returns 0.0 if a pure H2O fluid is undersaturated.
Parameters
----------
pressure float
The total pressure of the system in bars.
sample pandas Series or dict
Major element oxides in wt%, normalized on the basis of
no volatiles.
Returns
-------
float
1.0 if H2O-fluid saturated, 0.0 otherwise.
"""
if self.calculate_saturation_pressure(sample=sample,**kwargs) < pressure:
return 0.0
else:
return 1.0
def calculate_saturation_pressure(self,sample,**kwargs):
""" Calculates the pressure at which a pure H2O fluid is saturated, for the given
sample composition and H2O concentration. Calls the scipy.root_scalar routine, which makes
repeated calls to the calculate_dissolved_volatiles method.
Parameters
----------
sample pandas Series or dict
Major elements in wt% (normalized to 100%), including H2O (also in wt%, not included
in normalization).
Returns
-------
float
Saturation pressure in bar
"""
if 'H2O' not in sample:
raise InputError("sample must contain H2O")
if sample['H2O'] < 0:
raise InputError("H2O concentration must be greater than 0 wt%.")
if sample['H2O'] < self.calculate_dissolved_volatiles(sample=sample,pressure=0,**kwargs):
return np.nan
try:
satP = root_scalar(self.root_saturation_pressure,bracket=[1e-15,1e5],args=(sample,kwargs)).root
except:
warnings.warn("Saturation pressure not found.",RuntimeWarning)
satP = np.nan
return satP
def root_saturation_pressure(self,pressure,sample,kwargs):
""" Function called by scipy.root_scalar when finding the saturation pressure using
calculate_saturation_pressure.
Parameters
----------
pressure float
Pressure guess in bars
sample pandas Series or dict
Major elements in wt% (normalized to 100%), including H2O (also in wt%, not included
in normalization).
kwargs dictionary
Additional keyword arguments supplied to calculate_saturation_pressure. Might be required for
the fugacity or activity models.
Returns
-------
float
The differece between the dissolved H2O at the pressure guessed, and the H2O concentration
passed in the sample variable.
"""
return self.calculate_dissolved_volatiles(pressure=pressure,sample=sample,**kwargs)-sample['H2O']
class DixonCarbon(Model):
"""
Implementation of the Dixon (1997) carbon solubility model, as a Model class.
"""
def __init__(self):
self.set_volatile_species(['CO2'])
self.set_fugacity_model(fugacity_MRK_co2())
self.set_activity_model(activity_idealsolution())
self.set_calibration_ranges([])
self.set_solubility_dependence(False)
def preprocess_sample(self,sample):
""" Returns sample, normalized, keep volatiles unchanged.
Parameters
----------
sample: pandas Series or dict
The major element oxides in wt%.
Returns
-------
pandas Series or dict
The major element oxides in wt%.
"""
return normalize_FixedVolatiles(sample)
def calculate_dissolved_volatiles(self,pressure,sample,X_fluid=1.0,**kwargs):
"""Calculates the dissolved CO2 concentration using Eqn (3) of Dixon (1997).
Parameters
----------
pressure float
Total pressure in bars.
sample pandas Series or dict
Major element oxides in wt%.
X_fluid float
The mol fraction of CO2 in the fluid.
Returns
-------
float
The CO2 concentration in wt%.
"""
if X_fluid < 0 or X_fluid > 1:
raise InputError("X_fluid must have a value between 0 and 1.")
if pressure < 0:
raise InputError("Pressure must be positive.")
if type(sample) != dict and type(sample) != pd.core.series.Series:
raise InputError("sample must be a dict or pandas Series")
if 'SiO2' not in sample:
raise InputError("sample must contain SiO2.")
if pressure == 0:
return 0
Mr = wtpercentOxides_to_formulaWeight(sample)
XCO3 = self.molfrac_molecular(pressure=pressure,sample=sample,X_fluid=X_fluid,**kwargs)
# return (4400 * XCO3) / (36.6 - 44*XCO3)
return (4400 * XCO3) / (Mr - 44*XCO3)
def calculate_equilibrium_fluid_comp(self,pressure,sample,**kwargs):
""" Returns 1.0 if a pure H2O fluid is saturated.
Returns 0.0 if a pure H2O fluid is undersaturated.
Parameters
----------
pressure float
The total pressure of the system in bars.
sample pandas Series or dict
Major element oxides in wt% (including CO2).
Returns
-------
float
1.0 if CO2-fluid saturated, 0.0 otherwise.
"""
if self.calculate_saturation_pressure(sample=sample,**kwargs) < pressure:
return 0.0
else:
return 1.0
def calculate_saturation_pressure(self,sample,X_fluid=1.0,**kwargs):
"""
Calculates the pressure at which a pure CO2 fluid is saturated, for the given sample
composition and CO2 concentration. Calls the scipy.root_scalar routine, which makes
repeated called to the calculate_dissolved_volatiles method.
Parameters
----------
sample pandas Series or dict
Major element oxides in wt% (including CO2).
X_fluid float
The mole fraction of CO2 in the fluid. Default is 1.0.
Returns
-------
float
Calculated saturation pressure in bars.
"""
if type(sample) != dict and type(sample) != pd.core.series.Series:
raise InputError("sample must be a dict or pandas Series.")
if 'CO2' not in sample:
raise InputError("sample must contain CO2.")
if sample['CO2'] < 0:
raise InputError("Dissolved CO2 concentration must be greater than 0 wt%.")
try:
satP = root_scalar(self.root_saturation_pressure,x0=100.0,x1=1000.0,args=(sample,kwargs)).root
except:
warnings.warn("Saturation pressure not found.",RuntimeWarning)
satP = np.nan
return np.real(satP)
def molfrac_molecular(self,pressure,sample,X_fluid=1.0,**kwargs):
"""Calculates the mole fraction of CO3(-2) dissolved when in equilibrium with
a pure CO2 fluid at 1200C, using Eqn (1) of Dixon (1997).
Parameters
----------
pressure float
Total pressure in bars.
sample pandas Series or dict
Major element oxides in wt%.
X_fluid float
Mole fraction of CO2 in the fluid.
Returns
-------
float
Mole fraction of CO3(2-) dissolved."""
DeltaVr = 23.14 #cm3 mole-1
P0 = 1
R = 83.15
T0 = 1473.15
fugacity = self.fugacity_model.fugacity(pressure=pressure,X_fluid=X_fluid,**kwargs)
XCO3Std = self.XCO3_Std(sample)
return XCO3Std * fugacity * np.exp(-DeltaVr * (pressure-P0)/(R*T0))
def XCO3_Std(self,sample):
""" Calculates the mole fraction of CO3(2-) dissolved when in equilibrium with pure
CO2 vapour at 1200C and 1 bar, using Eq (8) of Dixon (1997).
Parameters
----------
sample pandas Series or dict
Major element oxides in wt%.
Returns
-------
float
Mole fraction of CO3(2-) dissolved at 1 bar and 1200C.
"""
if sample['SiO2'] > 48.9:
return 3.817e-7
else:
return 8.697e-6 - 1.697e-7*sample['SiO2']
def root_saturation_pressure(self,pressure,sample,kwargs):
""" The function called by scipy.root_scalar when finding the saturation pressure using
calculate_saturation_pressure.
Parameters
----------
pressure float
Total pressure in bars.
sample pandas Series or dict
Major element oxides in wt% (including CO2).
Returns
-------
float
The difference between the dissolved CO2 the pressure guessed, and the CO2 concentration
passed in the sample variable.
"""
return self.calculate_dissolved_volatiles(pressure=pressure,sample=sample,**kwargs) - sample['CO2']
class DixonWater(Model):
"""
Implementation of the Dixon (1997) water solubility model, as a Model class.
"""
def __init__(self):
self.set_volatile_species(['H2O'])
self.set_fugacity_model(fugacity_MRK_h2o())
self.set_activity_model(activity_idealsolution())
self.set_calibration_ranges([])
self.set_solubility_dependence(False)
def preprocess_sample(self,sample):
""" Returns sample, normalized, holding volatile concentrations constant.
Parameters
----------
sample: pandas Series or dict
The major element oxides in wt%.
Returns
-------
pandas Series or dict
The major element oxides in wt%.
"""
return normalize_FixedVolatiles(sample)
def calculate_dissolved_volatiles(self,pressure,sample,X_fluid=1.0,**kwargs):
"""Calculates the dissolved H2O concentration using Eqns (5) and (6) of Dixon (1997).
Parameters
----------
pressure float
Total pressure in bars.
sample pandas Series or dict
Major element oxides in wt%.
X_fluid float
The mol fraction of H2O in the fluid.
Returns
-------
float
The H2O concentration in wt%.
"""
if type(sample) != dict and type(sample) != pd.core.series.Series:
raise InputError("sample must be a dict or a pandas Series.")
if 'SiO2' not in sample:
raise InputError("sample must contain SiO2.")
if pressure < 0:
raise InputError("Pressure must be positive")
if X_fluid < 0 or X_fluid > 1:
raise InputError("X_fluid must have a value between 0 and 1.")
if pressure == 0:
return 0
XH2O = self.molfrac_molecular(pressure=pressure,sample=sample,X_fluid=X_fluid,**kwargs)
XOH = self.XOH(pressure=pressure,sample=sample,X_fluid=X_fluid,**kwargs)
Mr = wtpercentOxides_to_formulaWeight(sample)
XB = XH2O + 0.5*XOH
# return 1801.5*XB/(36.6-18.6*XB)
return 1801.5*XB/(Mr-18.6*XB)
def calculate_equilibrium_fluid_comp(self,pressure,sample,**kwargs):
""" Returns 1.0 if a pure H2O fluid is saturated.
Returns 0.0 if a pure H2O fluid is undersaturated.
Parameters
----------
pressure float
The total pressure of the system in bars.
sample pandas Series or dict
Major element oxides in wt% (including H2O).
Returns
-------
float
1.0 if H2O-fluid saturated, 0.0 otherwise.
"""
if self.calculate_saturation_pressure(sample=sample,**kwargs) < pressure:
return 0.0
else:
return 1.0
def calculate_saturation_pressure(self,sample,X_fluid=1.0,**kwargs):
"""
Calculates the pressure at which a pure H2O fluid is saturated, for the given sample
composition and H2O concentration. Calls the scipy.root_scalar routine, which makes
repeated called to the calculate_dissolved_volatiles method.
Parameters
----------
sample pandas Series or dict
Major element oxides in wt% (including H2O).
X_fluid float
The mole fraction of H2O in the fluid. Default is 1.0.
Returns
-------
float
Calculated saturation pressure in bars.
"""
if 'H2O' not in sample:
raise InputError("sample must contain H2O")
if sample['H2O'] < 0:
raise InputError("H2O concentration must be greater than 0 wt%.")
try:
satP = root_scalar(self.root_saturation_pressure,x0=100.0,x1=1000.0,args=(sample,kwargs)).root
except:
warnings.warn("Saturation pressure not found.",RuntimeWarning)
satP = np.nan
return np.real(satP)
def molfrac_molecular(self,pressure,sample,X_fluid=1.0,**kwargs):
"""Calculates the mole fraction of molecular H2O dissolved when in equilibrium with
a pure H2O fluid at 1200C, using Eqn (2) of Dixon (1997).
Parameters
----------
pressure float
Total pressure in bars.
sample pandas Series or dict
Major element oxides in wt%.
X_fluid float
Mole fraction of H2O in the fluid.
Returns
-------
float
Mole fraction of molecular H2O dissolved.
"""
VH2O = 12 #cm3 mole-1
P0 = 1
R = 83.15
T0 = 1473.15
XH2OStd = self.XH2O_Std(sample)
fugacity = self.fugacity_model.fugacity(pressure=pressure,X_fluid=X_fluid,**kwargs)
return XH2OStd * fugacity * np.exp(-VH2O * (pressure-P0)/(R*T0))
def XH2O_Std(self,sample):
""" Calculates the mole fraction of molecular H2O dissolved when in equilibrium with pure
H2O vapour at 1200C and 1 bar, using Eq (9) of Dixon (1997).
Parameters
----------
sample pandas Series or dict
Major element oxides in wt%.
Returns
-------
float
Mole fraction of molecular water dissolved at 1 bar and 1200C.
"""
if sample['SiO2'] > 48.9:
return 3.28e-5
else:
return -3.04e-5 + 1.29e-6*sample['SiO2']
def XOH(self,pressure,sample,X_fluid=1.0,**kwargs):
"""
Calculates the mole fraction of hydroxyl groups dissolved by solving Eq (4) of
Dixon (1997). Calls scipy.root_scalar to find the root of the XOH_root method.
Parameters
----------
pressure float
Total pressure in bars.
sample pandas Series or dict
Major element oxides in wt%.
X_fluid float
Mole fraction of H2O in the fluid.
Returns
-------
float
Mole fraction of hydroxyl groups dissolved.
"""
XH2O = self.molfrac_molecular(pressure=pressure,sample=sample,X_fluid=X_fluid,**kwargs)
if XH2O < 1e-14:
return 0
return np.exp(root_scalar(self.XOH_root,x0=np.log(0.5),x1=np.log(0.1),args=(XH2O)).root)
def XOH_root(self,XOH,XH2O):
"""
Method called by scipy.root_scalar when finding the saturation pressure using the
calculate_saturation_pressure method. Implements Eq (4) of Dixon (1997).
Parameters
----------
XOH float
Guess for the mole fraction of hydroxyl groups dissolved in melt.
XH2O float
Mole fraction of molecular water dissolved in melt.
Returns
-------
float
The difference between the RHS and LHS of Eq (4) of Dixon (1997) for the
guessed value of XOH.
"""
A = 0.403
B = 15.333
C = 10.894
XOH = np.exp(XOH)
term = (XOH)**2.0/(XH2O*(1.0-XOH-XH2O))
lhs = - np.log(term)
rhs = A + B*XOH + C*XH2O
return rhs - lhs
def root_saturation_pressure(self,pressure,sample,kwargs):
""" Function called by scipy.root_scalar when finding the saturation pressure using
calculate_saturation_pressure.
Parameters
----------
pressure float
Pressure guess in bars
sample pandas Series or dict
Major elements in wt% (normalized to 100%), including H2O.
kwargs dictionary
Additional keyword arguments supplied to calculate_saturation_pressure. Might be required for
the fugacity or activity models.
Returns
-------
float
The differece between the dissolved H2O at the pressure guessed, and the H2O concentration
passed in the sample variable.
"""
return self.calculate_dissolved_volatiles(pressure=pressure,sample=sample,**kwargs) - sample['H2O']
class IaconoMarzianoWater(Model):
"""
Implementation of the Iacono-Marziano et al. (2012) water solubility model, as a Model class. Two
calibrations are provided- the one incorporating the H2O content as a parameter (hydrous), and the
one that does not (anhydrous). Specify which should be used when initialising the model, with the
bool variable hydrous.
"""
def __init__(self,hydrous=True):
"""
Initialise the model.
Parameters
----------
hydrous bool
Whether to use the hydrous parameterization, or not.
"""
self.set_volatile_species(['H2O'])
self.set_fugacity_model(fugacity_idealgas())
self.set_activity_model(activity_idealsolution())
self.hydrous = hydrous
self.set_calibration_ranges([])
self.set_solubility_dependence(False) #Not dependent on CO2 conc, H2O dependence dealt with within model.
def preprocess_sample(self,sample):
"""
Returns sample, normalized to 100 wt%, without changing the wt% of H2O and CO2 if the
hydrous parameterization is being used (default). If the anhydrous parameterization is
used, it will normalize without including H2O and CO2.
Parameters
----------
sample pandas Series or dict
Major element oxides in wt%.
Returns
-------
pandas Series or dict
Major element oxides normalized to wt%.
"""
if self.hydrous == True:
return normalize_FixedVolatiles(sample)
else:
return normalize_AdditionalVolatiles(sample)
def calculate_dissolved_volatiles(self,pressure,temperature,sample,X_fluid=1.0,
hydrous_coeffs=True,webapp_coeffs=False,**kwargs):
"""
Calculates the dissolved H2O concentration, using Eq (13) of Iacono-Marziano et al. (2012).
If using the hydrous parameterization, it will use the scipy.root_scalar routine to find the
root of the root_dissolved_volatiles method.
Parameters
----------
pressure float
Total pressure in bars.
temperature float
Temperature in C
sample pandas Series or dict
Major element oxides in wt%.
X_fluid float
Mole fraction of H2O in the fluid. Default is 1.0.
hydrous_coeffs bool
Use the hydrous or anhydrous NBO/O paramterisation (True for hydrous). Default is True.
webapp_coeffs bool
If True, use the pre-review hydrous coefficients, as implemented in the IM webapp.
Default is False.
Returns
-------
float
Dissolved H2O concentration in wt%.
"""
temperature = temperature + 273.15 #translate T from C to K
if type(sample) != dict and type(sample) != pd.core.series.Series:
raise InputError("sample must be a dict or a pandas Series.")
if pressure < 0:
raise InputError("Pressure must be positive.")
if X_fluid < 0 or X_fluid > 1:
raise InputError("X_fluid must have a value between 0 and 1.")
if pressure == 0:
return 0
if hydrous_coeffs == True:
if X_fluid==0:
return 0
H2O = root_scalar(self.root_dissolved_volatiles,args=(pressure,temperature,sample,X_fluid,hydrous_coeffs,kwargs),
x0=1.0,x1=2.0).root
return H2O
else:
a = 0.54
b = 1.24
B = -2.95
C = 0.02
fugacity = self.fugacity_model.fugacity(pressure=pressure,X_fluid=X_fluid,temperature=temperature-273.15,**kwargs)
if fugacity == 0:
return 0
NBO_O = self.NBO_O(sample=sample,hydrous_coeffs=False)
H2O = np.exp(a*np.log(fugacity) + b*NBO_O + B + C*pressure/temperature)
return H2O
def calculate_equilibrium_fluid_comp(self,pressure,temperature,sample,**kwargs):
""" Returns 1.0 if a pure H2O fluid is saturated.
Returns 0.0 if a pure H2O fluid is undersaturated.
Parameters
----------
pressure float
The total pressure of the system in bars.
temperature float
The temperature of the system in C.
sample pandas Series or dict
Major element oxides in wt% (including H2O).
Returns
-------
float
1.0 if H2O-fluid saturated, 0.0 otherwise.
"""
if pressure > self.calculate_saturation_pressure(temperature=temperature,sample=sample,**kwargs):
return 0.0
else:
return 1.0
def calculate_saturation_pressure(self,temperature,sample,**kwargs):
"""
Calculates the pressure at which a pure H2O fluid is saturated, for the given sample
composition and H2O concentration. Calls the scipy.root_scalar routine, which makes
repeated called to the calculate_dissolved_volatiles method.
Parameters
----------
temperature float
The temperature of the system in C.
sample pandas Series or dict
Major element oxides in wt% (including H2O).
X_fluid float
The mole fraction of H2O in the fluid. Default is 1.0.
Returns
-------
float
Calculated saturation pressure in bars.
"""
if type(sample) != dict and type(sample) != pd.core.series.Series:
raise InputError("sample must be a dict or a pandas Series.")
if 'H2O' not in sample:
raise InputError("sample must contain H2O.")
if sample['H2O'] < 0.0:
raise InputError("Dissolved H2O must be greater than 0 wt%.")
try:
satP = root_scalar(self.root_saturation_pressure,args=(temperature,sample,kwargs),
bracket=[1e-15,1e5]).root
except:
warnings.warn("Saturation pressure not found.",RuntimeWarning)
satP = np.nan
return satP
def root_saturation_pressure(self,pressure,temperature,sample,kwargs):
""" Function called by scipy.root_scalar when finding the saturation pressure using
calculate_saturation_pressure.
Parameters
----------
pressure float
Pressure guess in bars
temperature float
The temperature of the system in C.
sample pandas Series or dict
Major elements in wt% (normalized to 100%), including H2O.
kwargs dictionary
Additional keyword arguments supplied to calculate_saturation_pressure. Might be required for
the fugacity or activity models.
Returns
-------
float
The differece between the dissolved H2O at the pressure guessed, and the H2O concentration
passed in the sample variable.
"""
return sample['H2O'] - self.calculate_dissolved_volatiles(pressure=pressure,temperature=temperature,sample=sample,**kwargs)
def root_dissolved_volatiles(self,h2o,pressure,temperature,sample,X_fluid,webapp_coeffs,kwargs):
""" Function called by calculate_dissolved_volatiles method when the hydrous parameterization is
being used.
Parameters
----------
h2o float
Guess for the H2O concentration in wt%.
pressure float
Total pressure in bars.
temperature float
Temperature in K.
sample pandas Series or dict
Major element oxides in wt%.
X_fluid float
Mole fraction of H2O in the fluid.
kwargs dictionary
Keyword arguments
Returns
-------
float
Difference between H2O guessed and the H2O calculated.
"""
if webapp_coeffs == False:
a = 0.53
b = 2.35
B = -3.37
C = -0.02
else:
a = 0.52096846
b = 2.11575907
B = -3.24443335
C = -0.02238884
sample_copy = sample.copy()
sample_copy['H2O'] = h2o
NBO_O = self.NBO_O(sample=sample_copy,hydrous_coeffs=True)
fugacity = self.fugacity_model.fugacity(pressure=pressure,X_fluid=X_fluid,temperature=temperature-273.15,**kwargs)
return h2o - np.exp(a*np.log(fugacity) + b*NBO_O + B + C*pressure/temperature)
def NBO_O(self,sample,hydrous_coeffs=True):
"""
Calculates NBO/O according to Appendix A.1. of Iacono-Marziano et al. (2012). NBO/O
is calculated on either a hydrous or anhyrous basis, as set when initialising the
Model class.
Parameters
----------
sample pandas Series or dict
Major element oxides in wt% (including H2O if using the hydrous parameterization).
Returns
-------
float
NBO/O.
"""
if all(ox in sample for ox in ['K2O','Na2O','CaO','MgO','FeO','Al2O3','SiO2','TiO2','Al2O3']) == False:
raise InputError("sample must contain K2O, Na2O, CaO, MgO, FeO, Al2O3, SiO2, TiO2 and Al2O3.")
X = wtpercentOxides_to_molOxides(sample)
NBO = 2*(X['K2O']+X['Na2O']+X['CaO']+X['MgO']+X['FeO']-X['Al2O3'])
O = 2*X['SiO2']+2*X['TiO2']+3*X['Al2O3']+X['MgO']+X['FeO']+X['CaO']+X['Na2O']+X['K2O']
if hydrous_coeffs == True:
if 'H2O' not in sample:
raise InputError("sample must contain H2O.")
NBO = NBO + 2*X['H2O']
O = O + X['H2O']
return NBO/O
class IaconoMarzianoCarbon(Model):
"""
Implementation of the Iacono-Marziano et al. (2012) carbon solubility model, as a Model class. Two
calibrations are provided- the one incorporating the H2O content as a parameter (hydrous), and the
one that does not (anhydrous). Specify which should be used when initialising the model, with the
bool variable hydrous.
"""
def __init__(self):
"""
Initialise the model.
Parameters
----------
hydrous bool
Whether to use the hydrous parameterization, or not.
"""
self.set_volatile_species(['CO2'])
self.set_fugacity_model(fugacity_idealgas())
self.set_activity_model(activity_idealsolution())
self.set_calibration_ranges([])
self.set_solubility_dependence(True)
def preprocess_sample(self,sample):
"""
Returns sample, normalized to 100 wt%, without changing the wt% of H2O and CO2 if the
hydrous parameterization is being used (default). If the anhydrous parameterization is
used, it will normalize without including H2O and CO2.
Parameters
----------
sample pandas Series or dict
Major element oxides in wt%.
Returns
-------
pandas Series or dict
Major element oxides normalized to wt%.
"""
if self.hydrous == True:
return normalize_FixedVolatiles(sample)
else:
return normalize_AdditionalVolatiles(sample)
def calculate_dissolved_volatiles(self,pressure,temperature,sample,X_fluid=1,
hydrous_coeffs=True, **kwargs):
"""
Calculates the dissolved CO2 concentration, using Eq (12) of Iacono-Marziano et al. (2012).
If using the hydrous parameterization, it will use the scipy.root_scalar routine to find the
root of the root_dissolved_volatiles method.
Parameters
----------
pressure float
Total pressure in bars.
temperature float
Temperature in C
sample pandas Series or dict
Major element oxides in wt%.
X_fluid float
Mole fraction of H2O in the fluid. Default is 1.0.
hydrous_coeffs bool
Use the hydrous or anhydrous NBO/O paramterisation (True for hydrous). Default is True.
Returns
-------
float
Dissolved H2O concentration in wt%.
"""
temperature = temperature + 273.15 #translate T from C to K
if type(sample) != dict and type(sample) != pd.core.series.Series:
raise InputError("sample must be a dict or a pandas Series.")
if pressure < 0:
raise InputError("Pressure must be positive.")
if temperature <= 0:
raise InputError("Temperature must be greater than 0K.")
if X_fluid < 0 or X_fluid > 1:
raise InputError("X_fluid must have a value between 0 and 1.")
if pressure == 0:
return 0
if hydrous_coeffs == True:
if 'H2O' not in sample:
raise InputError("sample must contain H2O if using the hydrous parameterization.")
if sample['H2O'] < 0:
raise InputError("Dissolved H2O must be positive.")
im_h2o_model = IaconoMarzianoWater()
h2o = im_h2o_model.calculate_dissolved_volatiles(pressure=pressure,temperature=temperature-273.15,
sample=sample,X_fluid=1-X_fluid,**kwargs)
sample_h2o = sample.copy()
sample_h2o['H2O'] = h2o
d = np.array([-16.4,4.4,-17.1,22.8])
a = 1.0
b = 17.3
B = -6.0
C = 0.12
NBO_O = self.NBO_O(sample=sample_h2o,hydrous_coeffs=True)
molarProps = wtpercentOxides_to_molOxides(sample_h2o)
else:
d = np.array([2.3,3.8,-16.3,20.1])
a = 1.0
b = 15.8
B = -5.3
C = 0.14
NBO_O = self.NBO_O(sample=sample,hydrous_coeffs=False)
molarProps = wtpercentOxides_to_molOxides(sample)
fugacity = self.fugacity_model.fugacity(pressure=pressure,X_fluid=X_fluid,temperature=temperature-273.15,**kwargs)
if fugacity == 0:
return 0
if all(ox in molarProps for ox in ['Al2O3','CaO','K2O','Na2O','FeO','MgO','Na2O','K2O']) == False:
raise InputError("sample must contain Al2O3, CaO, K2O, Na2O, FeO, MgO, Na2O, and K2O.")
x = list()
if 'H2O' in molarProps:
x.append(molarProps['H2O'])
else:
x.append(0.0)
x.append(molarProps['Al2O3']/(molarProps['CaO']+molarProps['K2O']+molarProps['Na2O']))
x.append((molarProps['FeO']+molarProps['MgO']))
x.append((molarProps['Na2O']+molarProps['K2O']))
x = np.array(x)
CO3 = np.exp(np.sum(x*d) + a*np.log(fugacity) + b*NBO_O + B + C*pressure/temperature)
CO2 = CO3/1e4#/(12+16*3)*(12+16*2)/1e4
return CO2
def calculate_equilibrium_fluid_comp(self,pressure,temperature,sample,**kwargs):
""" Returns 1.0 if a pure CO2 fluid is saturated.
Returns 0.0 if a pure CO2 fluid is undersaturated.
Parameters
----------
pressure float
The total pressure of the system in bars.
temperature float
The temperature of the system in C.
sample pandas Series or dict
Major element oxides in wt% (including H2O).
Returns
-------
float
1.0 if CO2-fluid saturated, 0.0 otherwise.
"""
if pressure > self.calculate_saturation_pressure(temperature=temperature,sample=sample,**kwargs):
return 0.0
else:
return 1.0
def calculate_saturation_pressure(self,temperature,sample,**kwargs):
"""
Calculates the pressure at which a pure CO2 fluid is saturated, for the given sample
composition and CO2 concentration. Calls the scipy.root_scalar routine, which makes
repeated called to the calculate_dissolved_volatiles method.
Parameters
----------
temperature float
The temperature of the system in C.
sample pandas Series or dict
Major element oxides in wt% (including CO2).
Returns
-------
float
Calculated saturation pressure in bars.
"""
if temperature <= 0:
raise InputError("Temperature must be greater than 0K.")
if type(sample) != dict and type(sample) != pd.core.series.Series:
raise InputError("sample must be a dict or a pandas Series.")
if 'CO2' not in sample:
raise InputError("sample must contain CO2")
if sample['CO2'] < 0:
raise InputError("Dissolved CO2 must be greater than 0 wt%.")
try:
satP = root_scalar(self.root_saturation_pressure,args=(temperature,sample,kwargs),
bracket=[1e-15,1e5]).root
except:
warnings.warn("Saturation pressure not found.",RuntimeWarning)
satP = np.nan
return satP
def root_saturation_pressure(self,pressure,temperature,sample,kwargs):
""" Function called by scipy.root_scalar when finding the saturation pressure using
calculate_saturation_pressure.
Parameters
----------
pressure float
Pressure guess in bars
temperature float
The temperature of the system in C.
sample pandas Series or dict
Major elements in wt% (normalized to 100%), including CO2.
kwargs dictionary
Additional keyword arguments supplied to calculate_saturation_pressure. Might be required for
the fugacity or activity models.
Returns
-------
float
The differece between the dissolved CO2 at the pressure guessed, and the CO2 concentration
passed in the sample variable.
"""
return sample['CO2'] - self.calculate_dissolved_volatiles(pressure=pressure,temperature=temperature,sample=sample,**kwargs)
def NBO_O(self,sample,hydrous_coeffs=True):
"""
Calculates NBO/O according to Appendix A.1. of Iacono-Marziano et al. (2012). NBO/O
is calculated on either a hydrous or anhyrous basis, as set when initialising the
Model class.
Parameters
----------
sample pandas Series or dict
Major element oxides in wt% (including H2O if using the hydrous parameterization).
Returns
-------
float
NBO/O.
"""
if type(sample) != dict and type(sample) != pd.core.series.Series:
raise InputError("sample must be a dict or a pandas Series,")
if all(ox in sample for ox in ['K2O','Na2O','CaO','MgO','FeO','Al2O3','SiO2','TiO2']) == False:
raise InputError("sample must contain K2O, Na2O, CaO, MgO, FeO, Al2O3, SiO2, and TiO2.")
X = wtpercentOxides_to_molOxides(sample)
NBO = 2*(X['K2O']+X['Na2O']+X['CaO']+X['MgO']+X['FeO']-X['Al2O3'])
O = 2*X['SiO2']+2*X['TiO2']+3*X['Al2O3']+X['MgO']+X['FeO']+X['CaO']+X['Na2O']+X['K2O']
if hydrous_coeffs == True:
if 'H2O' not in X:
raise InputError("sample must contain H2O if using the hydrous parameterization.")
NBO = NBO + 2*X['H2O']
O = O + X['H2O']
return NBO/O
class EguchiCarbon(Model):
"""
Implementation of the Eguchi and Dasgupta (2018) CO2 solubility model for andesitic melts.
Uses the Zhang and Duan (2009) CO2 EOS for fugacity calculations, assuming a pure CO2 fluid,
or ideal mixing for mixed fluids.
"""
def __init__(self):
warnings.warn("Eguchi model is not working correctly. Do not use any results calculated.")
self.set_volatile_species(['CO2'])
self.set_fugacity_model(fugacity_ZD09_co2())
self.set_activity_model(activity_idealsolution())
self.set_solubility_dependence(False)
self.set_calibration_ranges([CalibrationRange('pressure',[500.0,50000.0],crf_Between,'bar','Eguchi & Dasgupta (2018) carbon',
fail_msg=crmsg_Between_fail, pass_msg=crmsg_Between_pass, description_msg=crmsg_Between_description),
CalibrationRange('temperature',[950.0,1600],crf_Between,'oC','Eguchi & Dasgupta (2018) carbon',
fail_msg=crmsg_Between_fail, pass_msg=crmsg_Between_pass, description_msg=crmsg_Between_description)])
def preprocess_sample(self,sample,ferric_total=0.15):
""" Returns normalized sample composition, with ferric iron. Where a sample
already contains ferric iron, the composition will be normalized to 100 wt%
(excluding H2O and CO2). Where a sample contains only FeO, ferric iron will
be calculated using the ferric/total iron ratio provided.
Parameters
----------
sample pandas Series or dict
Major element oxides in wt%.
ferric_total float
Mole ratio of ferric to total iron to be used
for calculating Fe2O3 and FeO when only FeO is
provided. Default is 0.15.
Returns
-------
pandas Series or dict
Normalized major element oxides in wt%.
"""
if type(sample) != dict and type(sample) != pd.core.series.Series:
raise InputError("sample must be a dict or a pandas Series.")
if 'FeO' not in sample:
raise InputError("sample must contain FeO.")
_sample = sample.copy()
for ox in ['TiO2','P2O5']:
if ox not in _sample:
_sample[ox] = 0.0
if 'Fe2O3' not in _sample:
Fe_t = _sample['FeO']/oxideMass['FeO']
Fe3 = ferric_total*Fe_t
Fe2 = Fe_t - Fe3
_sample['FeO'] = Fe2*oxideMass['FeO']
_sample['Fe2O3'] = Fe3*oxideMass['Fe2O3']/2
return normalize_AdditionalVolatiles(_sample)
def calculate_dissolved_volatiles(self,pressure,temperature,sample,X_fluid=1.0,**kwargs):
"""
Calculates the dissolved (total) CO2 using eqs (9) and (10) of Eguchi and Dasgupta (2018).
Parameters
----------
pressure float
Pressure in bars
temperature float
Temperature in C
sample pandas Series or dict
Major element oxides in wt%.
X_fluid float
The mole fraction of CO2 in the fluid.
Returns
-------
float
Dissolved CO2 concentration.
"""
if pressure < 0:
raise InputError("Pressure must be greater than 0 bar.")
if pressure == 0:
return 0
XCO3 = self.Xi_melt(pressure=pressure,temperature=temperature,sample=sample,X_fluid=X_fluid,species='CO3')
XCO2 = self.Xi_melt(pressure=pressure,temperature=temperature,sample=sample,X_fluid=X_fluid,species='CO2')
FW_one = wtpercentOxides_to_formulaWeight(sample)
CO2_CO2 = ((44.01*XCO2)/(44.01*XCO2+(1-(XCO2+XCO3))*FW_one))*100
CO2_CO3 = ((44.01*XCO3)/(44.01*XCO3+(1-(XCO2+XCO3))*FW_one))*100
return CO2_CO2 + CO2_CO3
def calculate_equilibrium_fluid_comp(self,pressure,temperature,sample,**kwargs):
""" Returns 1.0 if a pure CO2 fluid is saturated.
Returns 0.0 if a pure CO2 fluid is undersaturated.
Parameters
----------
pressure float
The total pressure of the system in bars.
temperature float
The temperature of the system in C.
sample pandas Series or dict
Major element oxides in wt% (including H2O).
Returns
-------
float
1.0 if CO2-fluid saturated, 0.0 otherwise.
"""
satP = self.calculate_saturation_pressure(temperature=temperature,sample=sample,X_fluid=1.0,**kwargs)
if pressure < satP:
return 1.0
else:
return 0.0
def calculate_saturation_pressure(self,temperature,sample,X_fluid=1.0,**kwargs):
"""
Calculates the pressure at which a pure CO2 fluid is saturated, for the given sample
composition and CO2 concentration. Calls the scipy.root_scalar routine, which makes
repeated called to the calculate_dissolved_volatiles method.
Parameters
----------
temperature float
The temperature of the system in C.
sample pandas Series or dict
Major element oxides in wt% (including CO2).
X_fluid float
The mole fraction of H2O in the fluid. Default is 1.0.
Returns
-------
float
Calculated saturation pressure in bars.
"""
if 'CO2' not in sample:
raise InputError("sample must contain CO2.")
if sample['CO2'] < 0.0:
raise InputError("Concentration of CO2 must be greater than 0 wt%.")
try:
satP = root_scalar(self.root_saturation_pressure,x0=1000.0,x1=2000.0,
args=(temperature,sample,X_fluid,kwargs)).root
except:
warnings.warn("Saturation pressure not found.",RuntimeWarning)
satP = np.nan
return satP
def root_saturation_pressure(self,pressure,temperature,sample,X_fluid,kwargs):
""" Function called by scipy.root_scalar when finding the saturation pressure using
calculate_saturation_pressure.
Parameters
----------
pressure float
Pressure guess in bars
temperature float
The temperature of the system in C.
sample pandas Series or dict
Major elements in wt% (normalized to 100%), including CO2.
kwargs dictionary
Additional keyword arguments supplied to calculate_saturation_pressure. Might be required for
the fugacity or activity models.
Returns
-------
float
The differece between the dissolved CO2 at the pressure guessed, and the CO2 concentration
passed in the sample variable.
"""
return sample['CO2'] - self.calculate_dissolved_volatiles(pressure=pressure,temperature=temperature,sample=sample,X_fluid=X_fluid,**kwargs)
def Xi_melt(self,pressure,temperature,sample,species,X_fluid=1.0,**kwargs):
"""
Calculates the mole fraction of dissolved molecular CO2 or carbonate CO3(2-), using
eqn (9) of Eguchi and Dasgupta (2018).
Parameters
----------
pressure float
Pressure in bars.
temperature float
Temperature in C.
sample pandas Series or dict
Major element oxides in wt%.
species str
Which species to calculate, molecular CO2 'CO2' or carbonate ion 'CO3'.
X_fluid float
The mole fraction of CO2 in the fluid. Default is 1.0.
Returns
-------
float
Mole fraction of selected species in the melt
"""
temperature = temperature + 273.15 #translate T from C to K
if all(ox in sample for ox in ['MgO','CaO','FeO','Na2O','K2O','MnO','Al2O3','Fe2O3','SiO2','TiO2','P2O5']) == False:
raise InputError("sample must contain MgO, CaO, FeO, Na2O, K2O, MnO, Al2O3, Fe2O3, SiO3, TiO2, and P2O5.")
if X_fluid < 0 or X_fluid > 1:
raise InputError("X_fluid must have a value between 0 and 1.")
if pressure < 0:
raise InputError("Pressure must be positive.")
if temperature <= 0:
raise InputError("Temperature must be greater than 0K.")
if species == 'CO3':
DH = -1.65e5
DV = 2.38e-5
DS = -43.64
B = 1.47e3
yNBO = 3.29
A_CaO = 1.68e5
A_Na2O = 1.76e5
A_K2O = 2.11e5
elif species == 'CO2':
DH = -9.02e4
DV = 1.92e-5
DS = -43.08
B = 1.12e3
yNBO = -7.09
A_CaO = 0
A_Na2O = 0
A_K2O = 0
else:
raise InputError("species variable must be either 'CO2' or 'CO3'.")
R = 8.314
# Calculate NBO term
cations = wtpercentOxides_to_molSingleO(sample)
oxides = wtpercentOxides_to_molOxides(sample)
NM = (cations['Mg'] + cations['Ca'] + cations['Fe'] + cations['Na'] +
cations['K'] + cations['Mn'])
Al = cations['Al'] - NM
if Al > 0:
Al = NM
else:
Al = cations['Al']
Fe = cations['Fe3'] + Al
if Al > 0:
Fe = 0
if Al < 0 and Fe > 0:
Fe = - Al
if Al < 0 and Fe < 0:
Fe = cations['Fe3']
Tet = cations['Si'] + cations['Ti'] + cations['P'] + Al + Fe
NBO = 2 - 4*Tet
lnfCO2 = np.log(self.fugacity_model.fugacity(pressure=pressure,temperature=temperature-273.15,X_fluid=X_fluid))
lnXi = ((DH/(R*temperature)-(pressure*1e5*DV)/(R*temperature)+DS/R) +
(A_CaO*oxides['CaO']+A_Na2O*oxides['Na2O']+A_K2O*oxides['K2O'])/(R*temperature) +
(B*lnfCO2/temperature) + yNBO*NBO
)
return np.exp(lnXi)
class MooreWater(Model):
"""
Implementation of the Moore et al. (1998) H2O solubility model for magmas up to 3,000 bars.
"""
def __init__(self):
"""
Initialize the model.
"""
self.set_volatile_species(['H2O'])
self.set_fugacity_model(fugacity_HB_h2o())
self.set_activity_model(activity_idealsolution())
self.set_solubility_dependence(False)
self.set_calibration_ranges([CalibrationRange('pressure',[1.0,3000.0],crf_Between,'bar','Moore et al. (1998) water',
fail_msg=crmsg_Between_fail, pass_msg=crmsg_Between_pass, description_msg=crmsg_Between_description),
CalibrationRange('temperature',[700.0,1200],crf_Between,'oC','Moore et al. (1998) water',
fail_msg=crmsg_Between_fail, pass_msg=crmsg_Between_pass, description_msg=crmsg_Between_description)])
# self.set_calibration_ranges([cr_Between('pressure',[1.0,3000.0],'bar','Moore et al. (1998) water'),
# cr_Between('temperature',[700.0+273.15,1200+273.15],'oC','Moore et al. (1998) water')])
def preprocess_sample(self, sample):
"""
Returns sample with extranneous (non oxide) information removed and any missing oxides given a value of 0.0.
"""
for oxide in oxides:
if oxide in sample.keys():
pass
else:
sample[oxide] = 0.0
bulk_comp = {oxide: sample[oxide] for oxide in oxides}
self.bulk_comp_orig = sample
return bulk_comp
def calculate_dissolved_volatiles(self, sample, pressure, temperature, X_fluid=1.0, **kwargs):
"""
Parameters
----------
sample dict
Composition of sample in wt% oxides.
pressure float
Pressure in bars.
temperature float
Temperature in degrees C.
X_fluid float
OPTIONAL. Default is 1.0. Mole fraction of H2O in the H2O-CO2 fluid.
Returns
-------
float
Calculated dissolved H2O concentration in wt%.
"""
_sample = sample.copy()
_sample['H2O'] = 0.0
_sample['CO2'] = 0.0
_sample = normalize(_sample)
fH2O = self.fugacity_model.fugacity(pressure=pressure,temperature=temperature,X_fluid=X_fluid,**kwargs)
aParam = 2565.0
bParam_Al2O3 = -1.997
bParam_FeOt = -0.9275
bParam_Na2O = 2.736
cParam = 1.171
dParam = -14.21
temperatureK = temperature + 273.15
sample_molfrac = wtpercentOxides_to_molOxides(_sample)
FeOtot = sample_molfrac['FeO'] + sample_molfrac['Fe2O3']*0.8998
b_x_sum = (bParam_Al2O3 * sample_molfrac['Al2O3']) + (bParam_FeOt * FeOtot) + (bParam_Na2O * sample_molfrac['Na2O'])
two_ln_XH2Omelt = (aParam / temperatureK) + b_x_sum * (pressure/temperatureK) + cParam * np.log(fH2O) + dParam
ln_XH2Omelt = two_ln_XH2Omelt / 2.0
XH2Omelt = np.exp(ln_XH2Omelt)
sample_molfrac['H2O'] = XH2Omelt
#Normalize mol fractions to sum to 1, while preserving XH2O
for key, value in sample_molfrac.items():
if key != 'H2O':
sample_molfrac.update({key: value/((1/(1-sample_molfrac['H2O'])))})
sample_wtper = mol_to_wtpercent(sample_molfrac)
return sample_wtper['H2O']
def calculate_equilibrium_fluid_comp(self, sample, pressure, temperature, **kwargs):
"""
Parameters
----------
sample dict
Composition of sample in wt% oxides.
pressure float
Pressure in bars.
temperature float
Temperature in degrees C.
Returns
-------
float
Calculated equilibrium fluid concentration in XH2Ofluid mole fraction.
"""
_sample = sample.copy()
sample_anhy = sample.copy()
sample_anhy["H2O"] = 0.0
sample_anhy["CO2"] = 0.0
aParam = 2565.0
bParam_Al2O3 = -1.997
bParam_FeOt = -0.9275
bParam_Na2O = 2.736
cParam = 1.171
dParam = -14.21
temperatureK = temperature + 273.15
sample_molfrac_anhy = wtpercentOxides_to_molOxides(sample_anhy)
sample_molfrac_hy = wtpercentOxides_to_molOxides(_sample)
FeOtot = sample_molfrac_anhy['FeO'] + sample_molfrac_anhy['Fe2O3']*0.8998
b_x_sum = (bParam_Al2O3 * sample_molfrac_anhy['Al2O3']) + (bParam_FeOt * FeOtot) + (bParam_Na2O * sample_molfrac_anhy['Na2O'])
ln_fH2O = (2 * np.log(sample_molfrac_hy['H2O']) - (aParam/temperatureK) - b_x_sum * (pressure/temperatureK) - dParam) / cParam
fH2O = np.exp(ln_fH2O)
XH2O_fl = fH2O / pressure
# SM: I've changed this to return X_H2O only, as otherwise it doesn't conform to other single-volatile
# models. I'm not sure this is the best solution though.
# return (XCO2_fl, XH2O_fl)
return XH2O_fl
def calculate_saturation_pressure(self,temperature,sample,X_fluid=1.0,**kwargs):
"""
Calculates the pressure at which a an H2O-bearing fluid is saturated. Calls the scipy.root_scalar
routine, which makes repeated called to the calculate_dissolved_volatiles method.
Parameters
----------
sample dict
Composition of sample in wt% oxides.
temperature float
Temperature in degrees C.
X_fluid float
OPTIONAL. Default is 1.0. Mole fraction of H2O in the H2O-CO2 fluid.
Returns
-------
float
Calculated saturation pressure in bars.
"""
_sample = sample.copy()
temperatureK = temperature + 273.15
if temperatureK <= 0.0:
raise InputError("Temperature must be greater than 0K.")
if X_fluid < 0 or X_fluid > 1:
raise InputError("X_fluid must have a value between 0 and 1.")
if type(sample) != dict and type(sample) != pd.core.series.Series:
raise InputError("sample must be a dict or a pandas Series.")
if 'H2O' not in sample:
raise InputError("sample must contain H2O.")
if sample['H2O'] < 0.0:
raise InputError("Dissolved H2O concentration must be greater than 0 wt%.")
try:
satP = root_scalar(self.root_saturation_pressure,args=(temperature,_sample,X_fluid,kwargs),
x0=100.0,x1=2000.0).root
except:
warnings.warn("Saturation pressure not found.",RuntimeWarning)
satP = np.nan
return np.real(satP)
def root_saturation_pressure(self,pressure,temperature,sample,X_fluid,kwargs):
""" Function called by scipy.root_scalar when finding the saturation pressure using
calculate_saturation_pressure.
Parameters
----------
pressure float
Pressure guess in bars
temperature float
The temperature of the system in C.
sample pandas Series or dict
Major elements in wt% (normalized to 100%), including H2O.
kwargs dictionary
Additional keyword arguments supplied to calculate_saturation_pressure. Might be required for
the fugacity or activity models.
Returns
-------
float
The differece between the dissolved H2O at the pressure guessed, and the H2O concentration
passed in the sample variable.
"""
return self.calculate_dissolved_volatiles(pressure=pressure,temperature=temperature,sample=sample,X_fluid=X_fluid,**kwargs) - sample['H2O']
class LiuWater(Model):
"""
Implementation of the Liu et al. (2005) H2O solubility model for metaluminous high-silica rhyolitic melts.
"""
def __init__(self):
"""
Initialize the model.
"""
self.set_volatile_species(['H2O'])
self.set_fugacity_model(fugacity_idealgas())
self.set_activity_model(activity_idealsolution())
self.set_solubility_dependence(False)
self.set_calibration_ranges([CalibrationRange('pressure',[1.0,5000.0],crf_Between,'bar','Liu et al. (2005) water',
fail_msg=crmsg_Between_fail, pass_msg=crmsg_Between_pass, description_msg=crmsg_Between_description),
CalibrationRange('temperature',[700.0,1200],crf_Between,'oC','Liu et al. (2005) water',
fail_msg=crmsg_Between_fail, pass_msg=crmsg_Between_pass, description_msg=crmsg_Between_description),
CalibrationRange('sample',None,crf_LiuComp,None,None,
fail_msg=crmsg_LiuComp_fail, pass_msg=crmsg_LiuComp_pass, description_msg=crmsg_LiuComp_description)])
# self.set_calibration_ranges([cr_Between('pressure',[1.0,5000.0],'bar','Liu et al. (2005) water'),
# cr_Between('temperature',[700.0,1200],'oC','Liu et al. (2005) water')])
def preprocess_sample(self, sample):
"""
Returns sample with extranneous (non oxide) information removed and any missing oxides given a value of 0.0.
"""
for oxide in oxides:
if oxide in sample.keys():
pass
else:
sample[oxide] = 0.0
bulk_comp = {oxide: sample[oxide] for oxide in oxides}
self.bulk_comp_orig = sample
return bulk_comp
def calculate_dissolved_volatiles(self, sample, pressure, temperature, X_fluid=1.0, **kwargs):
"""
Parameters
----------
sample dict
Composition of sample in wt% oxides.
pressure float
Pressure in bars.
temperature float
Temperature in degrees C.
X_fluid float
OPTIONAL. Default is 1.0. Mole fraction of H2O in the H2O-CO2 fluid.
Returns
-------
float
Calculated dissolved H2O concentration in wt%.
"""
pressureMPa = pressure / 10.0
Pw = pressureMPa * X_fluid
PCO2 = pressureMPa * (1 - X_fluid)
temperatureK = temperature + 273.15
H2Ot = ((354.94*Pw**(0.5) + 9.623*Pw - 1.5223*Pw**(1.5)) / temperatureK +
0.0012439*Pw**(1.5) + PCO2*(-1.084*10**(-4)*Pw**(0.5) - 1.362*10**(-5)*Pw))
return H2Ot
def calculate_equilibrium_fluid_comp(self, sample, pressure, temperature, **kwargs):
"""
Parameters
----------
sample dict
Composition of sample in wt% oxides.
pressure float
Pressure in bars.
temperature float
Temperature in degrees C.
Returns
-------
float
Calculated equilibrium fluid concentration in XH2Ofluid mole fraction.
"""
temperatureK = temperature + 273.15
pressureMPa = pressure / 10.0
_sample = sample.copy()
H2Ot = _sample["H2O"]
#calculate saturation pressure and assert that input P <= SatP
satP = self.calculate_saturation_pressure(temperature,sample)
is_saturated = satP - pressure
if is_saturated >= 0:
pass
else:
warnings.warn("{:.1f} bars is above the saturation pressure ({:.1f} bars) for this sample. Results from this calculation may be nonsensical.".format(pressure,satP))
#Use sympy to solve solubility equation for XH2Ofluid
XH2Ofluid = sympy.symbols('XH2Ofluid') #XH2Ofluid is the variable to solve for
equation = ((354.94*(XH2Ofluid*pressureMPa)**(0.5) + 9.623*(XH2Ofluid*pressureMPa)
- 1.5223*(XH2Ofluid*pressureMPa)**(1.5)) / temperatureK
+ 0.0012439*(XH2Ofluid*pressureMPa)**(1.5)
+ pressureMPa*(1-XH2Ofluid)*(-1.084*10**(-4)*(XH2Ofluid*pressureMPa)**(0.5)
- 1.362*10**(-5)*(XH2Ofluid*pressureMPa)) - H2Ot)
XH2Ofluid = sympy.solve(equation, XH2Ofluid)[0]
if XH2Ofluid > 1:
XH2Ofluid = 1
if XH2Ofluid < 0:
XH2Ofluid = 0
return XH2Ofluid
def calculate_saturation_pressure(self,temperature,sample,X_fluid=1.0,**kwargs):
"""
Calculates the pressure at which a an H2O-bearing fluid is saturated. Calls the scipy.root_scalar
routine, which makes repeated called to the calculate_dissolved_volatiles method.
Parameters
----------
sample dict
Composition of sample in wt% oxides.
temperature float
Temperature in degrees C.
X_fluid float
OPTIONAL. Default is 1.0. Mole fraction of H2O in the H2O-CO2 fluid.
Returns
-------
float
Calculated saturation pressure in bars.
"""
_sample = sample.copy()
temperatureK = temperature + 273.15
if temperatureK <= 0.0:
raise InputError("Temperature must be greater than 0K.")
if X_fluid < 0 or X_fluid > 1:
raise InputError("X_fluid must have a value between 0 and 1.")
if type(sample) != dict and type(sample) != pd.core.series.Series:
raise InputError("sample must be a dict or a pandas Series.")
if 'H2O' not in sample:
raise InputError("sample must contain H2O.")
if sample['H2O'] < 0.0:
raise InputError("Dissolved H2O concentration must be greater than 0 wt%.")
try:
satP = root_scalar(self.root_saturation_pressure,args=(temperature,_sample,X_fluid,kwargs),
x0=10.0,x1=200.0).root
except:
warnings.warn("Saturation pressure not found.",RuntimeWarning)
satP = np.nan
return np.real(satP)
def root_saturation_pressure(self,pressure,temperature,sample,X_fluid,kwargs):
""" Function called by scipy.root_scalar when finding the saturation pressure using
calculate_saturation_pressure.
Parameters
----------
pressure float
Pressure guess in bars
temperature float
The temperature of the system in C.
sample pandas Series or dict
Major elements in wt% (normalized to 100%), including H2O.
kwargs dictionary
Additional keyword arguments supplied to calculate_saturation_pressure. Might be required for
the fugacity or activity models.
Returns
-------
float
The differece between the dissolved H2O at the pressure guessed, and the H2O concentration
passed in the sample variable.
"""
return self.calculate_dissolved_volatiles(pressure=pressure,temperature=temperature,sample=sample,X_fluid=X_fluid,**kwargs) - sample['H2O']
class LiuCarbon(Model):
"""
Implementation of the Liu et al. (2005) H2O-CO2 solubility model for metaluminous high-silica rhyolitic melts.
"""
def __init__(self):
"""
Initialize the model.
"""
self.set_volatile_species(['CO2'])
self.set_fugacity_model(fugacity_idealgas())
self.set_activity_model(activity_idealsolution())
self.set_solubility_dependence(False)
self.set_calibration_ranges([CalibrationRange('pressure',[1.0,5000.0],crf_Between,'bar','Liu et al. (2005) carbon',
fail_msg=crmsg_Between_fail, pass_msg=crmsg_Between_pass, description_msg=crmsg_Between_description),
CalibrationRange('temperature',[700.0,1200],crf_Between,'oC','Liu et al. (2005) carbon',
fail_msg=crmsg_Between_fail, pass_msg=crmsg_Between_pass, description_msg=crmsg_Between_description),
CalibrationRange('sample',None,crf_LiuComp,None,None,
fail_msg=crmsg_LiuComp_fail, pass_msg=crmsg_LiuComp_pass, description_msg=crmsg_LiuComp_description)])
def preprocess_sample(self, sample):
"""
Returns sample with extranneous (non oxide) information removed and any missing oxides given a value of 0.0.
"""
for oxide in oxides:
if oxide in sample.keys():
pass
else:
sample[oxide] = 0.0
bulk_comp = {oxide: sample[oxide] for oxide in oxides}
self.bulk_comp_orig = sample
return bulk_comp
def calculate_dissolved_volatiles(self, sample, pressure, temperature, X_fluid=1, **kwargs):
"""
Parameters
----------
sample dict
Composition of sample in wt% oxides.
pressure float
Pressure in bars.
temperature float
Temperature in degrees C.
X_fluid float
OPTIONAL. Default is 1. Mole fraction of CO2 in the H2O-CO2 fluid.
Returns
-------
float
Calculated dissolved CO2 concentration in wt%.
"""
pressureMPa = pressure / 10.0
Pw = pressureMPa * (1 - X_fluid)
PCO2 = pressureMPa * X_fluid #(1 - X_fluid)
temperatureK = temperature + 273.15
CO2melt_ppm = (PCO2*(5668 - 55.99*Pw)/temperatureK
+ PCO2*(0.4133*Pw**(0.5) + 2.041*10**(-3)*Pw**(1.5)))
CO2melt = CO2melt_ppm / 10000
return CO2melt
def calculate_equilibrium_fluid_comp(self, sample, pressure, temperature, **kwargs):
"""
Parameters
----------
sample dict
Composition of sample in wt% oxides.
pressure float
Pressure in bars.
temperature float
Temperature in degrees C.
Returns
-------
float
Calculated equilibrium fluid concentration in XCO2fluid mole fraction.
"""
temperatureK = temperature + 273.15
pressureMPa = pressure / 10.0
if temperatureK <= 0.0:
raise InputError("Temperature must be greater than 0K.")
if type(sample) != dict and type(sample) != pd.core.series.Series:
raise InputError("sample must be a dict or a pandas Series.")
if 'CO2' not in sample:
raise InputError("sample must contain CO2.")
if sample['CO2'] < 0.0:
raise InputError("Dissolved CO2 concentration must be greater than 0 wt%.")
_sample = sample.copy()
CO2melt_wt = _sample["CO2"]
CO2melt_ppm = CO2melt_wt * 10000
#calculate saturation pressure and assert that input P <= SatP
satP = self.calculate_saturation_pressure(temperature,sample)
is_saturated = satP - pressure
if is_saturated >= 0:
pass
else:
warnings.warn(str(pressure) + " bars is above the saturation pressure (" + str(satP) + " bars) for this sample. Results from this calculation may be nonsensical.")
#Use sympy to solve solubility equation for XH2Ofluid
XCO2fluid = sympy.symbols('XCO2fluid') #XCO2fluid is the variable to solve for
equation = (((XCO2fluid*pressureMPa)*(5668 - 55.99*(pressureMPa*(1-XCO2fluid)))/temperatureK
+ (XCO2fluid*pressureMPa)*(0.4133*(pressureMPa*(1-XCO2fluid))**(0.5)
+ 2.041*10**(-3)*(pressureMPa*(1-XCO2fluid))**(1.5))) - CO2melt_ppm)
XCO2fluid = sympy.solve(equation, XCO2fluid)[0]
if XCO2fluid > 1:
XCO2fluid = 1
if XCO2fluid < 0:
XCO2fluid = 0
return XCO2fluid #1 - XCO2fluid
def calculate_saturation_pressure(self,temperature,sample,X_fluid=1.0,**kwargs):
"""
Calculates the pressure at which a an CO2-bearing fluid is saturated. Calls the scipy.root_scalar
routine, which makes repeated called to the calculate_dissolved_volatiles method.
Parameters
----------
sample dict
Composition of sample in wt% oxides.
temperature float
Temperature in degrees C.
X_fluid float
OPTIONAL. Default is 0. Mole fraction of CO2 in the H2O-CO2 fluid.
Returns
-------
float
Calculated saturation pressure in bars.
"""
_sample = sample.copy()
temperatureK = temperature + 273.15
if temperatureK <= 0.0:
raise InputError("Temperature must be greater than 0K.")
if X_fluid < 0 or X_fluid > 1:
raise InputError("X_fluid must have a value between 0 and 1.")
if type(sample) != dict and type(sample) != pd.core.series.Series:
raise InputError("sample must be a dict or a pandas Series.")
if 'CO2' not in sample:
raise InputError("sample must contain CO2.")
if sample['CO2'] < 0.0:
raise InputError("Dissolved CO2 concentration must be greater than 0 wt%.")
try:
satP = root_scalar(self.root_saturation_pressure,args=(temperature,_sample,X_fluid,kwargs),
x0=10.0,x1=2000.0).root
except:
warnings.warn("Saturation pressure not found.",RuntimeWarning)
satP = np.nan
return np.real(satP)
def root_saturation_pressure(self,pressure,temperature,sample,X_fluid,kwargs):
""" Function called by scipy.root_scalar when finding the saturation pressure using
calculate_saturation_pressure.
Parameters
----------
pressure float
Pressure guess in bars
temperature float
The temperature of the system in C.
sample pandas Series or dict
Major elements in wt% (normalized to 100%), including H2O.
kwargs dictionary
Additional keyword arguments supplied to calculate_saturation_pressure. Might be required for
the fugacity or activity models.
Returns
-------
float
The differece between the dissolved H2O at the pressure guessed, and the H2O concentration
passed in the sample variable.
"""
return self.calculate_dissolved_volatiles(pressure=pressure,temperature=temperature,sample=sample,X_fluid=X_fluid,**kwargs) - sample['CO2']
class AllisonCarbon(Model):
"""
Implementation of the Allison et al. (2019) CO2 solubility model. Which type of fit, and
which composition must be selected when the Model is initialized. The fit may be either
thermodynamic or power-law. The composition may be chosen from sunset, sfvf, erebus, vesuvius,
etna, or stromboli. Default is the power-law fit to sunset.
"""
def __init__(self):
"""
Initialize the model.
"""
self.set_volatile_species(['CO2'])
self.set_fugacity_model(fugacity_HB_co2())
self.set_activity_model(activity_idealsolution())
self.set_calibration_ranges([CalibrationRange('pressure',[0.0,6000.0],crf_Between,'bar','Allison et al. (2019) carbon',
fail_msg=crmsg_Between_fail, pass_msg=crmsg_Between_pass, description_msg=crmsg_Between_description),
CalibrationRange('temperature',1200,crf_EqualTo,'oC','Allison et al. (2019) carbon',
fail_msg=crmsg_EqualTo_fail, pass_msg=crmsg_EqualTo_pass, description_msg=crmsg_EqualTo_description)])
self.set_solubility_dependence(False)
def preprocess_sample(self,sample):
"""
Returns sample normalized to 100wt%, keeping the concentrations of H2O and CO2 constant.
Parameters
----------
sample pandas Series or dict
Major element oxides in wt%.
Returns
-------
pandas Series
Normalized major element oxides in wt%.
"""
return normalize_AdditionalVolatiles(sample)
def calculate_dissolved_volatiles(self,pressure,temperature,sample=None,X_fluid=1.0,
model_loc='sunset',model_fit='thermodynamic',**kwargs):
"""
Calclates the dissolved CO2 concentration using (Eqns) 2-7 or 10-11 from Allison et al. (2019).
Parameters
----------
pressure float
Pressure in bars.
temperature float
Temperature in C.
sample pandas Series, dict or None
Major element oxides in wt%. Required if using the thermodynamic fits, need not be
provided if using the power law fits. Default is None.
X_fluid float
The mole fraction of CO2 in the fluid. Default is 1.0.
model_fit str
Either 'power' for the power-law fits, or 'thermodynamic' for the
thermodynamic fits.
model_loc str
One of 'sunset', 'sfvf', 'erebus', 'vesuvius', 'etna', 'stromboli'.
Returns
-------
float
Dissolved CO2 concentration in wt%.
"""
temperature = temperature + 273.15 #translate T from C to K
if temperature <= 0.0:
raise InputError("Temperature must be greater than 0K.")
if pressure < 0.0:
raise InputError("Pressure must be positive.")
if X_fluid < 0 or X_fluid > 1:
raise InputError("X_fluid must have a value between 0 and 1.")
if model_fit not in ['power','thermodynamic']:
raise InputError("model_fit must be one of 'power', or 'thermodynamic'.")
if model_loc not in ['sunset','sfvf','erebus','vesuvius','etna','stromboli']:
raise InputError("model_loc must be one of 'sunset', 'sfvf', 'erebus', 'vesuvius', 'etna', or 'stromboli'.")
if pressure == 0:
return 0
if model_fit == 'thermodynamic':
if type(sample) != dict and type(sample) != pd.core.series.Series:
raise InputError("Thermodynamic fit requires sample to be a dict or a pandas Series.")
P0 = 1000 # bar
params = dict({'sunset':[16.4,-14.67],
'sfvf':[15.02,-14.87],
'erebus':[15.83,-14.65],
'vesuvius':[24.42,-14.04],
'etna':[21.59,-14.28],
'stromboli':[14.93,-14.68]})
DV = params[model_loc][0]
lnK0 = params[model_loc][1]
lnK = lnK0 - (pressure-P0)*DV/(10*8.3141*temperature)
fCO2 = self.fugacity_model.fugacity(pressure=pressure,temperature=temperature-273.15,X_fluid=X_fluid,**kwargs)
Kf = np.exp(lnK)*fCO2
XCO3 = Kf/(1-Kf)
# FWone = wtpercentOxides_to_formulaWeight(sample)#,exclude_volatiles=True)
FWone = 36.594
wtCO2 = (44.01*XCO3)/((44.01*XCO3)+(1-XCO3)*FWone)*100
return wtCO2
if model_fit == 'power':
params = dict({'stromboli':[1.05,0.883],
'etna':[2.831,0.797],
'vesuvius':[4.796,0.754],
'sfvf':[3.273,0.74],
'sunset':[4.32,0.728],
'erebus':[5.145,0.713]})
fCO2 = self.fugacity_model.fugacity(pressure=pressure,temperature=temperature-273.15,X_fluid=X_fluid,**kwargs)
return params[model_loc][0]*fCO2**params[model_loc][1]/1e4
def calculate_equilibrium_fluid_comp(self,pressure,temperature,sample,**kwargs):
""" Returns 1.0 if a pure CO2 fluid is saturated.
Returns 0.0 if a pure CO2 fluid is undersaturated.
Parameters
----------
pressure float
The total pressure of the system in bars.
temperature float
The temperature of the system in C.
sample pandas Series or dict
Major element oxides in wt% (including H2O).
Returns
-------
float
1.0 if CO2-fluid saturated, 0.0 otherwise.
"""
satP = self.calculate_saturation_pressure(temperature=temperature,sample=sample,X_fluid=1.0,**kwargs)
if pressure < satP:
return 1.0
else:
return 0.0
def calculate_saturation_pressure(self,temperature,sample,X_fluid=1.0,**kwargs):
"""
Calculates the pressure at which a pure CO2 fluid is saturated, for the given sample
composition and CO2 concentration. Calls the scipy.root_scalar routine, which makes
repeated called to the calculate_dissolved_volatiles method.
Parameters
----------
temperature float
The temperature of the system in C.
sample pandas Series
Major element oxides in wt% (including CO2).
X_fluid float
The mole fraction of H2O in the fluid. Default is 1.0.
Returns
-------
float
Calculated saturation pressure in bars.
"""
if temperature <= 0.0:
raise InputError("Temperature must be greater than 0K.")
if X_fluid < 0 or X_fluid > 1:
raise InputError("X_fluid must have a value between 0 and 1.")
if type(sample) != dict and type(sample) != pd.core.series.Series:
raise InputError("sample must be a dict or a pandas Series.")
if 'CO2' not in sample:
raise InputError("sample must contain CO2.")
if sample['CO2'] < 0.0:
raise InputError("Dissolved CO2 concentration must be greater than 0 wt%.")
try:
satP = root_scalar(self.root_saturation_pressure,args=(temperature,sample,X_fluid,kwargs),
x0=1000.0,x1=2000.0).root
except:
warnings.warn("Saturation pressure not found.",RuntimeWarning)
satP = np.nan
return satP
def root_saturation_pressure(self,pressure,temperature,sample,X_fluid,kwargs):
""" Function called by scipy.root_scalar when finding the saturation pressure using
calculate_saturation_pressure.
Parameters
----------
pressure float
Pressure guess in bars
temperature float
The temperature of the system in C.
sample pandas Series or dict
Major elements in wt% (normalized to 100%), including CO2.
kwargs dictionary
Additional keyword arguments supplied to calculate_saturation_pressure. Might be required for
the fugacity or activity models.
Returns
-------
float
The differece between the dissolved CO2 at the pressure guessed, and the CO2 concentration
passed in the sample variable.
"""
return sample['CO2'] - self.calculate_dissolved_volatiles(pressure=pressure,temperature=temperature,sample=sample,X_fluid=X_fluid,**kwargs)
#------------MIXED FLUID MODELS-------------------------------#
class MixedFluid(Model):
"""
Implements the generic framework for mixed fluid solubility. Any set of pure fluid solubility
models may be specified.
"""
def __init__(self,models):
"""
Initializes the mixed fluid model.
Parameters
----------
models dictionary
Dictionary with names of volatile species as keys, and the model objects as values.
"""
self.models = tuple(model for model in models.values())
self.set_volatile_species(list(models.keys()))
def preprocess_sample(self,sample):
""" Returns sample, unmodified.
Parameters
----------
sample pandas Series or dict
Major element oxides in wt%.
Returns
-------
pandas Series or dict
Major element oxides in wt%.
"""
if type(sample) != dict and type(sample) != pd.core.series.Series:
raise InputError("sample must be a dict or a pandas Series.")
_sample = sample.copy()
_sample = self.models[0].preprocess_sample(_sample)
return _sample
def calculate_dissolved_volatiles(self,pressure,X_fluid,returndict=False,**kwargs):
"""
Calculates the dissolved volatile concentrations in wt%, using each model's
calculate_dissolved_volatiles method. At present the volatile concentrations are
not propagated through.
Parameters
----------
pressure float
The total pressure in bars.
X_fluid float, numpy.ndarry, dict, pandas Series
The mole fraction of each species in the fluid. If the mixed fluid model
contains only two species (e.g. CO2 and H2O), the value of the first species in
self.volatile_species may be passed on its own as a float.
returndict bool
If True, the results will be returned in a dict, otherwise they will be returned
as a tuple.
Returns
-------
tuple
Dissolved volatile concentrations of each species in the model, in the order set
by self.volatile_species.
"""
if (type(X_fluid) == float or type(X_fluid) == int) and len(self.volatile_species) == 2:
X_fluid = (X_fluid,1-X_fluid)
elif len(X_fluid) != len(self.volatile_species):
raise InputError("X_fluid must have the same length as the number of volatile species\
in the MixedFluids Model class, or it may have length 1 if two species are present\
in the MixedFluids Model class.")
if np.sum(X_fluid) != 1.0:
raise InputError("X_fluid must sum to 1.0")
if any(val<0 for val in X_fluid) or any(val>1 for val in X_fluid):
raise InputError("Each mole fraction in X_fluid must have a value between 0 and 1.")
if type(X_fluid) == dict or type(X_fluid) == pd.core.series.Series:
X_fluid = tuple(X_fluid[species] for species in self.volatile_species)
# If the models don't depend on the concentration of volatiles, themselves.
if all(model.solubility_dependence == False for model in self.models):
result = tuple(model.calculate_dissolved_volatiles(pressure=pressure,X_fluid=Xi,**kwargs) for model, Xi in zip(self.models,X_fluid))
# If one of the models depends on the other volatile concentration
elif len(self.models) == 2 and self.models[0].solubility_dependence == False and 'sample' in kwargs:
result0 = self.models[0].calculate_dissolved_volatiles(pressure=pressure,X_fluid=X_fluid[0],**kwargs)
samplecopy = kwargs['sample'].copy()
samplecopy[self.volatile_species[0]] = result0
kwargs['sample'] = samplecopy
result1 = self.models[1].calculate_dissolved_volatiles(pressure=pressure,X_fluid=X_fluid[1],**kwargs)
result = (result0,result1)
elif len(self.models) == 2 and self.models[1].solubility_dependence == False and 'sample' in kwargs:
result1 = self.models[1].calculate_dissolved_volatiles(pressure=pressure,X_fluid=X_fluid[1],**kwargs)
samplecopy = kwargs['sample'].copy()
samplecopy[self.volatile_species[1]] = result1
kwargs['sample'] = samplecopy
result0 = self.models[0].calculate_dissolved_volatiles(pressure=pressure,X_fluid=X_fluid[0],**kwargs)
result = (result0,result1)
else:
raise InputError("The solubility dependence of the models is not currently supported by the MixedFluid model.")
if returndict == True:
resultsdict = {}
for i,v in zip(range(len(self.volatile_species)),self.volatile_species):
resultsdict.update({v+'_liq':result[i]})
return resultsdict
else:
return result
def calculate_equilibrium_fluid_comp(self,pressure,sample,return_dict=True,**kwargs):
""" Calculates the composition of the fluid in equilibrium with the dissolved volatile
concentrations passed. If a fluid phase is undersaturated at the chosen pressure (0,0) will
be returned. Note, this currently assumes the given H2O and CO2 concentrations are
the system total, not the total dissolved. If one of the volatile species has a zero or
negative concentration, the pure fluid model for the other volatile species will be used.
Parameters
----------
pressure float
The total pressure in bars.
sample pandas Series or dict
Major element oxides in wt% (including volatiles).
return_dict bool
Set the return type, if true a dict will be returned, if False two floats will be
returned. Default is True.
Returns
-------
dict or floats
Mole fractions of the volatile species in the fluid, in the order given by
self.volatile_species if floats.
"""
if len(self.volatile_species) != 2:
raise InputError("Currently equilibrium fluid compositions can only be calculated when\
two volatile species are present.")
dissolved_at_0bar = [self.models[0].calculate_dissolved_volatiles(sample=sample,pressure=0.0,**kwargs),
self.models[1].calculate_dissolved_volatiles(sample=sample,pressure=0.0,**kwargs)]
if sample[self.volatile_species[0]] <= 0.0 or sample[self.volatile_species[0]] <= dissolved_at_0bar[0]:
Xv0 = 0.0
Xv1 = self.models[1].calculate_equilibrium_fluid_comp(pressure=pressure,sample=sample,**kwargs)
elif sample[self.volatile_species[1]] <= 0.0 or sample[self.volatile_species[1]] <= dissolved_at_0bar[1]:
Xv1 = 0.0
Xv0 = self.models[0].calculate_equilibrium_fluid_comp(pressure=pressure,sample=sample,**kwargs)
else:
satP = self.calculate_saturation_pressure(sample,**kwargs)
if satP < pressure:
if return_dict == True:
return {self.volatile_species[0]:0,self.volatile_species[1]:0}
else:
return (0,0)
molfracs = wtpercentOxides_to_molOxides(sample)
(Xt0, Xt1) = (molfracs[self.volatile_species[0]],molfracs[self.volatile_species[1]])
try:
Xv0 = root_scalar(self.root_for_fluid_comp,bracket=[1e-15,1-1e-15],args=(pressure,Xt0,Xt1,sample,kwargs)).root
Xv1 = 1 - Xv0
except:
try:
Xv0 = root_scalar(self.root_for_fluid_comp,x0=0.5,x1=0.1,args=(pressure,Xt0,Xt1,sample,kwargs)).root
Xv1 = 1 - Xv0
except:
raise SaturationError("Equilibrium fluid not found. Likely an issue with the numerical solver.")
if return_dict == True:
return {self.volatile_species[0]:Xv0,self.volatile_species[1]:Xv1}
else:
return Xv0, Xv1
def calculate_saturation_pressure(self,sample,**kwargs):
"""
Calculates the pressure at which a fluid will be saturated, given the dissolved volatile
concentrations. If one of the volatile species has a zero or negative concentration the
pure fluid model for the other species will be used. If one of the volatile species has a
concentration lower than the concentration dissolved at 0 bar, the pure fluid model for the
other species will be used.
Parameters
----------
sample pandas Series or dict
Major element oxides in wt% (including volatiles).
Returns
-------
float
The saturation pressure in bars.
"""
dissolved_at_0bar = [self.models[0].calculate_dissolved_volatiles(sample=sample,pressure=0.0,**kwargs),
self.models[1].calculate_dissolved_volatiles(sample=sample,pressure=0.0,**kwargs)]
if sample[self.volatile_species[0]] <= 0.0 or sample[self.volatile_species[0]] <= dissolved_at_0bar[0]:
satP = self.models[1].calculate_saturation_pressure(sample=sample,**kwargs)
elif sample[self.volatile_species[1]] <= 0.0 or sample[self.volatile_species[1]] <= dissolved_at_0bar[1]:
satP = self.models[0].calculate_saturation_pressure(sample=sample,**kwargs)
else:
volatile_concs = np.array(tuple(sample[species] for species in self.volatile_species))
x0 = 0
for model in self.models:
xx0 = model.calculate_saturation_pressure(sample=sample,**kwargs)
if np.isnan(xx0) == False:
x0 += xx0
try:
satP = root(self.root_saturation_pressure,x0=[x0,0.5],args=(volatile_concs,sample,kwargs)).x[0]
except:
warnings.warn("Saturation pressure not found.",RuntimeWarning)
satP = np.nan
return satP
def calculate_isobars_and_isopleths(self,pressure_list,isopleth_list=[0,1],points=51,
return_dfs=True,extend_to_zero=True,**kwargs):
"""
Calculates isobars and isopleths. Isobars can be calculated for any number of pressures. Variables
required by each of the pure fluid models must be passed, e.g. sample, temperature, etc.
Parameters
----------
pressure_list list
List of all pressure values at which to calculate isobars, in bars.
isopleth_list list
Default value is None, in which case only isobars will be calculated. List of all
fluid compositions in mole fraction (of the first species in self.volatile_species) at which
to calcualte isopleths. Values can range from 0 to 1.
points int
The number of points in each isobar and isopleth. Default value is 101.
return_dfs bool
If True, the results will be returned as two pandas DataFrames, as produced by the MagmaSat
method. If False the results will be returned as lists of numpy arrays.
Returns
-------
pandas DataFrame object(s) or list(s)
If isopleth_list is not None, two objects will be returned, one with the isobars and the second with
the isopleths. If return_dfs is True, two pandas DataFrames will be returned with column names
'Pressure' or 'XH2O_fl', 'H2O_liq', and 'CO2_liq'. If return_dfs is False, two lists of numpy arrays
will be returned. Each array is an individual isobar or isopleth, in the order passed via pressure_list
or isopleth_list. The arrays are the concentrations of H2O and CO2 in the liquid, in the order of the
species in self.volatile_species.
"""
if len(self.volatile_species) != 2 or 'H2O' not in self.volatile_species or 'CO2' not in self.volatile_species:
raise InputError("calculate_isobars_and_isopleths may only be used with a H2O-CO2 fluid.")
H2O_id = self.volatile_species.index('H2O')
CO2_id = self.volatile_species.index('CO2')
has_isopleths = True
if isopleth_list is None:
has_isopleths = False
isobars_df = pd.DataFrame(columns=['Pressure','H2O_liq','CO2_liq'])
isobars = []
for pressure in pressure_list:
dissolved = np.zeros([2,points])
Xv0 = np.linspace(0.0,1.0,points)
for i in range(points):
dissolved[:,i] = self.calculate_dissolved_volatiles(pressure=pressure,X_fluid=(Xv0[i],1-Xv0[i]),**kwargs)
isobars_df = isobars_df.append({'Pressure':pressure,'H2O_liq':dissolved[H2O_id,i],'CO2_liq':dissolved[CO2_id,i]},ignore_index=True)
isobars.append(dissolved)
if has_isopleths == True:
isopleths_df = pd.DataFrame(columns=['XH2O_fl','H2O_liq','CO2_liq'])
isopleths = []
for isopleth in isopleth_list:
dissolved = np.zeros([2,points])
pmin = np.nanmin(pressure_list)
pmax = np.nanmax(pressure_list)
if pmin == pmax:
pmin = 0.0
pressure = np.linspace(pmin,pmax,points)
for i in range(points):
dissolved[:,i] = self.calculate_dissolved_volatiles(pressure=pressure[i],X_fluid=(isopleth,1-isopleth),**kwargs)
isopleths_df = isopleths_df.append({'XH2O_fl':[isopleth,1-isopleth][H2O_id],'H2O_liq':dissolved[H2O_id,i],'CO2_liq':dissolved[CO2_id,i]},ignore_index=True)
isopleths.append(dissolved)
if return_dfs == True:
if has_isopleths == True:
return (isobars_df, isopleths_df)
else:
return isobars_df
else:
if has_isopleths == True:
return (isobars, isopleths)
else:
return isobars
def calculate_degassing_path(self,sample,pressure='saturation',fractionate_vapor=0.0,final_pressure=100.0,
steps=101,return_dfs=True,round_to_zero=True,**kwargs):
"""
Calculates the dissolved volatiles in a progressively degassing sample.
Parameters
----------
sample pandas Series or dict
Major element oxides in wt% (including volatiles).
pressure string, float, int, list, or numpy array
Defaults to 'saturation', the calculation will begin at the saturation pressure. If a number is passed
as either a float or int, this will be the starting pressure. If a list of numpy array is passed, the
pressure values in the list or array will define the degassing path, i.e. final_pressure and steps
variables will be ignored. Units are bars.
fractionate_vapor float
What proportion of vapor should be removed at each step. If 0.0 (default), the degassing path will
correspond to closed-system degassing. If 1.0, the degassing path will correspond to open-system
degassing.
final_pressure float
The final pressure on the degassing path, in bars. Ignored if a list or numpy array is passed as the
pressure variable. Default is 1 bar.
steps int
The number of steps in the degassing path. Ignored if a list or numpy array are passed as the pressure
variable.
return_dfs bool
If True, the results will be returned in a pandas DataFrame, if False, two numpy arrays will be returned.
round_to_zero bool
If True, the first entry of FluidProportion_wt will be rounded to zero, rather than being a value
within numerical error of zero. Default is True.
Returns
-------
pandas DataFrame or numpy arrays
If return_dfs is True (default), a DataFrame with columns 'Pressure_bars', 'H2O_liq', 'CO2_liq',
'H2O_fl', 'CO2_fl', and 'FluidProportion_wt', is returned. Dissolved volatiles are in wt%,
the proportions of volatiles in the fluid are in mole fraction. Otherwise a numpy array containing
the dissolved volatile concentrations, and a numpy array containing the mole fractions of
volatiles in the fluid is returned. The columns are in the order of the volatiles in
self.volatile_species.
"""
# if 'model' in kwargs and model=='Liu':
# final_pressure = 1.0
wtptoxides = sample.copy()
wtptoxides = normalize_FixedVolatiles(wtptoxides)
wtm0s, wtm1s = (wtptoxides[self.volatile_species[0]],wtptoxides[self.volatile_species[1]])
if pressure == 'saturation':
p0 = self.calculate_saturation_pressure(wtptoxides,**kwargs)
pressures = np.linspace(p0,final_pressure,steps)
elif type(pressure) == float or type(pressure) == int:
pressures = np.linspace(pressure,final_pressure,steps)
elif type(pressure) == list or type(pressure) == np.ndarray:
pressures = pressure
Xv = np.zeros([2,len(pressures)])
wtm = np.zeros([2,len(pressures)])
for i in range(len(pressures)):
try:
X_fluid = self.calculate_equilibrium_fluid_comp(pressure=pressures[i],sample=wtptoxides,return_dict=False,**kwargs)
Xv[:,i] = X_fluid
if X_fluid == (0,0):
wtm[:,i] = (wtptoxides[self.volatile_species[0]],wtptoxides[self.volatile_species[1]])
else:
if X_fluid[0] == 0:
wtm[0,i] = wtptoxides[self.volatile_species[0]]
wtm[1,i] = self.calculate_dissolved_volatiles(pressure=pressures[i],sample=wtptoxides,X_fluid=X_fluid,**kwargs)[1]
elif X_fluid[1] == 0:
wtm[1,i] = wtptoxides[self.volatile_species[1]]
wtm[0,i] = self.calculate_dissolved_volatiles(pressure=pressures[i],sample=wtptoxides,X_fluid=X_fluid,**kwargs)[0]
else:
wtm[:,i] = self.calculate_dissolved_volatiles(pressure=pressures[i],sample=wtptoxides,X_fluid=X_fluid,**kwargs)
wtptoxides[self.volatile_species[0]] = wtm[0,i] + (1-fractionate_vapor)*(wtm0s-wtm[0,i])
wtptoxides[self.volatile_species[1]] = wtm[1,i] + (1-fractionate_vapor)*(wtm1s-wtm[1,i])
# wtptoxides = normalize_FixedVolatiles(wtptoxides)
except:
Xv[:,i] = [np.nan]*np.shape(Xv)[0]
wtm[:,i] = wtm[:,i-1]
if return_dfs == True:
exsolved_degassing_df = pd.DataFrame()
exsolved_degassing_df['Pressure_bars'] = pressures
exsolved_degassing_df['H2O_liq'] = wtm[self.volatile_species.index('H2O'),:]
exsolved_degassing_df['CO2_liq'] = wtm[self.volatile_species.index('CO2'),:]
exsolved_degassing_df['H2O_fl'] = Xv[self.volatile_species.index('H2O'),:]
exsolved_degassing_df['CO2_fl'] = Xv[self.volatile_species.index('CO2'),:]
exsolved_degassing_df['FluidProportion_wt'] = (wtm0s+wtm1s)-exsolved_degassing_df['H2O_liq']-exsolved_degassing_df['CO2_liq']
if round_to_zero == True and np.round(exsolved_degassing_df.loc[0,'FluidProportion_wt'],2)==0:
exsolved_degassing_df.loc[0,'FluidProportion_wt'] = 0.0
return exsolved_degassing_df
else:
return (wtm, Xv)
def root_saturation_pressure(self,x,volatile_concs,sample,kwargs):
""" Function called by scipy.root when finding the saturation pressure using
calculate_saturation_pressure.
Parameters
----------
x numpy array
The guessed value for the root. x[0] is the pressure (in bars) and x[1] is the
mole fraction of the first volatile in self.volatile_species.
volatile_concs numpy array
The dissolved volatile concentrations, in the same order as self.volatile_species.
sample pandas Series or dict
Major element oxides in wt% (including volatiles).
kwargs dictionary
Dictionary of keyword arguments, which may be required by the pure-fluid models.
Returns
-------
numpy array
The difference in the dissolved volatile concentrations, and those predicted with the
pressure and fluid composition specified by x.
"""
if x[1] < 0:
x[1] = 0
elif x[1] > 1:
x[1] = 1
if x[0] <= 0:
x[0] = 1e-15
misfit = np.array(self.calculate_dissolved_volatiles(pressure=x[0],X_fluid=(x[1],1-x[1]),sample=sample,**kwargs)) - volatile_concs
return misfit
def root_for_fluid_comp(self,Xv0,pressure,Xt0,Xt1,sample,kwargs):
""" Function called by scipy.root_scalar when calculating the composition of equilibrium fluid
in the calculate_equilibrium_fluid_comp method.
Parameters
----------
Xv0 float
The guessed mole fraction of the first volatile species in self.volatile_species.
pressure float
The total pressure in bars.
Xt0 float
The total mole fraction of the first volatile species in self.volatile_species.
Xt1 float
The total mole fraction of the second volatile species in self.volatile_species.
sample pandas Series
Major element oxides in wt%
kwargs dictionary
A dictionary of keyword arguments that may be required by the pure fluid models.
Returns
-------
float
The differene in the LHS and RHS of the mass balance equation. Eq X in manuscript.
"""
wtt0 = sample[self.volatile_species[0]]
wtt1 = sample[self.volatile_species[1]]
wtm0, wtm1 = self.calculate_dissolved_volatiles(pressure=pressure,X_fluid=(Xv0,1-Xv0),sample=sample,**kwargs)
Xm0 = Xt0/wtt0*wtm0
Xm1 = Xt1/wtt1*wtm1
if self.volatile_species[0] == 'CO2' and Xv0 != Xm0:
f = (Xt0-Xm0)/(Xv0-Xm0)
return (1-f)*Xm1 + f*(1-Xv0) - Xt1
else:
f = (Xt1-Xm1)/((1-Xv0)-Xm1)
return (1-f)*Xm0 + f*Xv0 - Xt0
def check_calibration_range(self,parameters,report_nonexistance=True):
""" Checks whether the given parameters are within the ranges defined by the
CalibrationRange objects for each model and its fugacity and activity models. An empty
string will be returned if all parameters are within the calibration range. If a
parameter is not within the calibration range, a description of the problem will be
returned in the string.
Parameters
----------
parameters dict
Dictionary keys are the names of the parameters to be checked, e.g., pressure
temperature, SiO2, etc. Values are the values of each parameter. A complete set
need not be given.
Returns
-------
str
String description of any parameters falling outside of the calibration range.
"""
s = ''
for model in self.models:
for cr in model.calibration_ranges:
if cr.check(parameters) == False:
s += cr.string(parameters,report_nonexistance)
for cr in model.fugacity_model.calibration_ranges:
if cr.check(parameters) == False:
s += cr.string(parameters,report_nonexistance)
for cr in model.activity_model.calibration_ranges:
if cr.check(parameters) == False:
s += cr.string(parameters,report_nonexistance)
return s
def get_calibration_range(self):
""" Returns a string describing the calibration ranges defined by the CalibrationRange
objects for each model, and its associated fugacity and activity models.
Returns
-------
str
String description of the calibration range objects."""
s = ''
for model in self.models:
for cr in model.calibration_ranges:
s += cr.string(None)
for cr in model.fugacity_model.calibration_ranges:
s += cr.string(None)
for cr in model.activity_model.calibration_ranges:
s += cr.string(None)
return s
class MagmaSat(Model):
"""
An object to instantiate a thermoengine equilibrate class
"""
def __init__(self):
self.melts_version = '1.2.0' #just here so users can see which version is being used
self.set_volatile_species(['H2O', 'CO2'])
self.set_calibration_ranges([CalibrationRange('pressure',[0.0,30000.0],crf_Between,'bar','MagmaSat',
fail_msg=crmsg_Between_fail, pass_msg=crmsg_Between_pass, description_msg=crmsg_Between_description),
CalibrationRange('temperature',[550,1730],crf_Between,'oC','MagmaSat',
fail_msg=crmsg_Between_fail, pass_msg=crmsg_Between_pass, description_msg=crmsg_Between_description)])
def preprocess_sample(self,sample): #TODO test this by passing weird shit to sample
"""
Returns sample with 0.0 values for any oxides not passed.
Parameters
----------
sample: dictionary
Sample composition in wt% oxides
Returns
-------
dictionary
Sample composition in wt% oxides
"""
for oxide in oxides:
if oxide in sample.keys():
pass
else:
sample[oxide] = 0.0
self.bulk_comp_orig = sample
return sample
def check_calibration_range(self,parameters,**kwargs):
""" Checks whether supplied parameters and calculated results are within the calibration range
of the model, defined by the CalibrationRange objects. An empty string will be returned if all
parameters are within the calibration range. If a parameter is not within the calibration range,
a description of the problem will be returned in the string.
Parameters
----------
parameters dict
Dictionary keys are the names of the parameters to be checked, e.g., pressure
temperature, SiO2, etc. Values are the values of each parameter. A complete set
need not be given.
Returns
-------
str
String description of any parameters falling outside of the calibration range.
"""
s = ''
for cr in self.calibration_ranges:
if cr.check(parameters) == False:
s += cr.string(parameters,report_nonexistance=False)
return s
def get_calibration_range(self):
""" Returns a string describing the calibration ranges defined by the CalibrationRange
objects for the model.
Returns
-------
str
String description of the calibration range objects."""
s = ''
for cr in self.calibration_ranges:
s += cr.string(None)
return s
def get_fluid_mass(self, sample, temperature, pressure, H2O, CO2):
"""An internally used function to calculate fluid mass.
Parameters
----------
sample: dictionary
Sample composition in wt% oxides
temperature: float
Temperature in degrees C.
pressure: float
Pressure in bars
H2O: float
wt% H2O in the system
CO2: float
wt% CO2 in the system
Returns
-------
float
mass of the fluid in grams
"""
pressureMPa = pressure / 10.0
bulk_comp = {oxide: sample[oxide] for oxide in oxides}
bulk_comp["H2O"] = H2O
bulk_comp["CO2"] = CO2
feasible = melts.set_bulk_composition(bulk_comp)
output = melts.equilibrate_tp(temperature, pressureMPa, initialize=True)
(status, temperature, pressureMPa, xmlout) = output[0]
fluid_mass = melts.get_mass_of_phase(xmlout, phase_name='Fluid')
return fluid_mass
def get_XH2O_fluid(self, sample, temperature, pressure, H2O, CO2):
"""An internally used function to calculate fluid composition.
Parameters
----------
sample: dictionary
Sample composition in wt% oxides
temperature: float
Temperature in degrees C.
pressure: float
Pressure in bars
H2O: float
wt% H2O in the system
CO2: float
wt% CO2 in the system
Returns
-------
float
Mole fraction of H2O in the H2O-CO2 fluid
"""
pressureMPa = pressure / 10.0
bulk_comp = {oxide: sample[oxide] for oxide in oxides}
bulk_comp["H2O"] = H2O
bulk_comp["CO2"] = CO2
feasible = melts.set_bulk_composition(bulk_comp)
output = melts.equilibrate_tp(temperature, pressureMPa, initialize=True)
(status, temperature, pressureMPa, xmlout) = output[0]
fluid_comp = melts.get_composition_of_phase(xmlout, phase_name='Fluid', mode='component')
#NOTE mode='component' returns endmember component keys with values in mol fraction.
if "Water" in fluid_comp:
H2O_fl = fluid_comp["Water"]
else:
H2O_fl = 0.0
# if H2O_fl == 0:
# raise SaturationError("Composition not fluid saturated.")
return H2O_fl
def calculate_dissolved_volatiles(self, sample, temperature, pressure, X_fluid=1, H2O_guess=0.0, verbose=False, **kwargs):
#TODO make better initial guess at higher XH2Ofl
#TODO make refinements faster
"""
Calculates the amount of H2O and CO2 dissolved in a magma at saturation at the given P/T conditions and fluid composition.
Fluid composition will be matched to within 0.0001 mole fraction.
Parameters
----------
sample: dict or pandas Series
Compositional information on one sample in oxides.
temperature: float or int
Temperature, in degrees C.
presure: float or int
Pressure, in bars.
X_fluid: float or int
The default value is 1. The mole fraction of H2O in the H2O-CO2 fluid. X_fluid=1 is a pure H2O fluid. X_fluid=0 is a pure CO2 fluid.
verbose: bool
OPTIONAL: Default is False. If set to True, returns H2O and CO2 concentration in the melt, H2O and CO2 concentration in
the fluid, mass of the fluid in grams, and proportion of fluid in the system in wt%.
Returns
-------
dict
A dictionary of dissolved volatile concentrations in wt% with keys H2O and CO2.
"""
sample = self.preprocess_sample(sample)
if isinstance(X_fluid, int) or isinstance(X_fluid, float):
pass
else:
raise InputError("X_fluid must be type int or float")
if isinstance(H2O_guess, int) or isinstance(H2O_guess, float):
pass
else:
raise InputError("H2O_guess must be type int or float")
pressureMPa = pressure / 10.0
bulk_comp = {oxide: sample[oxide] for oxide in oxides}
if X_fluid != 0 and X_fluid !=1:
if X_fluid < 0.001 or X_fluid > 0.999:
raise InputError("X_fluid is calculated to a precision of 0.0001 mole fraction. \
Value for X_fluid must be between 0.0001 and 0.9999.")
H2O_val = H2O_guess
CO2_val = 0.0
fluid_mass = 0.0
while fluid_mass <= 0:
if X_fluid == 0:
CO2_val += 0.1
elif X_fluid >= 0.5:
H2O_val += 0.2
CO2_val = (H2O_val / X_fluid) - H2O_val #NOTE this is setting XH2Owt of the system (not of the fluid) to X_fluid
#TODO this is what needs to be higher for higher XH2O. Slows down computation by a second or two
else:
H2O_val += 0.1
CO2_val = (H2O_val / X_fluid) - H2O_val #NOTE this is setting XH2Owt of the system (not of the fluid) to X_fluid
#TODO this is what needs to be higher for higher XH2O. Slows down computation by a second or two
fluid_mass = self.get_fluid_mass(sample, temperature, pressure, H2O_val, CO2_val)
bulk_comp["H2O"] = H2O_val
bulk_comp["CO2"] = CO2_val
feasible = melts.set_bulk_composition(bulk_comp)
output = melts.equilibrate_tp(temperature, pressureMPa, initialize=True)
(status, temperature, pressureMPa, xmlout) = output[0]
liquid_comp = melts.get_composition_of_phase(xmlout, phase_name='Liquid', mode='oxide_wt')
fluid_comp = melts.get_composition_of_phase(xmlout, phase_name='Fluid', mode='component')
if "Water" in fluid_comp:
H2O_fl = fluid_comp["Water"]
else:
H2O_fl = 0.0
XH2O_fluid = H2O_fl
#------Coarse Check------#
while XH2O_fluid < X_fluid - 0.1: #too low coarse check
H2O_val += 0.2
XH2O_fluid = self.get_XH2O_fluid(sample, temperature, pressure, H2O_val, CO2_val)
while XH2O_fluid > X_fluid + 0.1: #too high coarse check
CO2_val += 0.1
XH2O_fluid = self.get_XH2O_fluid(sample, temperature, pressure, H2O_val, CO2_val)
#------Refinement 1------#
while XH2O_fluid < X_fluid - 0.01: #too low refinement 1
H2O_val += 0.05
XH2O_fluid = self.get_XH2O_fluid(sample, temperature, pressure, H2O_val, CO2_val)
while XH2O_fluid > X_fluid + 0.01: #too high refinement 1
CO2_val += 0.01
XH2O_fluid = self.get_XH2O_fluid(sample, temperature, pressure, H2O_val, CO2_val)
#------Refinement 2------#
while XH2O_fluid < X_fluid - 0.001: #too low refinement 2
H2O_val += 0.005
XH2O_fluid = self.get_XH2O_fluid(sample, temperature, pressure, H2O_val, CO2_val)
while XH2O_fluid > X_fluid + 0.001: #too high refinement 2
CO2_val += 0.001
XH2O_fluid = self.get_XH2O_fluid(sample, temperature, pressure, H2O_val, CO2_val)
#------Final refinement------#
while XH2O_fluid < X_fluid - 0.0001: #too low final refinement
H2O_val += 0.001
XH2O_fluid = self.get_XH2O_fluid(sample, temperature, pressure, H2O_val, CO2_val)
while XH2O_fluid > X_fluid + 0.0001: #too high final refinement
CO2_val += 0.0001
XH2O_fluid = self.get_XH2O_fluid(sample, temperature, pressure, H2O_val, CO2_val)
#------Get calculated values------#
bulk_comp["H2O"] = H2O_val
bulk_comp["CO2"] = CO2_val
feasible = melts.set_bulk_composition(bulk_comp)
output = melts.equilibrate_tp(temperature, pressureMPa, initialize=True)
(status, temperature, pressureMPa, xmlout) = output[0]
fluid_mass = melts.get_mass_of_phase(xmlout, phase_name='Fluid')
system_mass = melts.get_mass_of_phase(xmlout, phase_name='System')
liquid_comp = melts.get_composition_of_phase(xmlout, phase_name='Liquid', mode='oxide_wt')
fluid_comp = melts.get_composition_of_phase(xmlout, phase_name='Fluid', mode='component')
if "H2O" in liquid_comp:
H2O_liq = liquid_comp["H2O"]
else:
H2O_liq = 0
if "CO2" in liquid_comp:
CO2_liq = liquid_comp["CO2"]
else:
CO2_liq = 0
if "Water" in fluid_comp:
H2O_fl = fluid_comp["Water"]
else:
H2O_fl = 0.0
if "Carbon Dioxide" in fluid_comp:
CO2_fl = fluid_comp["Carbon Dioxide"]
else:
CO2_fl = 0.0
XH2O_fluid = H2O_fl
if verbose == True:
return {"temperature": temperature, "pressure": pressure,
"H2O_liq": H2O_liq, "CO2_liq": CO2_liq,
"XH2O_fl": H2O_fl, "XCO2_fl": CO2_fl,
"FluidProportion_wt": 100*fluid_mass/system_mass}
if verbose == False:
return {"CO2": CO2_liq, "H2O": H2O_liq}
def calculate_equilibrium_fluid_comp(self, sample, temperature, pressure, verbose=False, **kwargs): #TODO fix weird printing
"""
Returns H2O and CO2 concentrations in wt% in a fluid in equilibrium with the given sample at the given P/T condition.
Parameters
----------
sample: dict or pandas Series
Compositional information on one sample in oxides.
temperature: float or int
Temperature, in degrees C.
presure: float or int
Pressure, in bars. #TODO check units
verbose: bool
OPTIONAL: Default is False. If set to True, returns H2O and CO2 concentration in the fluid, mass of the fluid in grams,
and proportion of fluid in the system in wt%.
Returns
-------
dict
A dictionary of fluid composition in wt% with keys 'H2O' and 'CO2' is returned. #TODO make list?
"""
sample = self.preprocess_sample(sample)
if isinstance(temperature, float) or isinstance(temperature, int):
pass
else:
raise InputError("temp must be type float or int")
if isinstance(pressure, float) or isinstance(pressure, int):
pass
else:
raise InputError("presure must be type float or int")
pressureMPa = pressure / 10.0
bulk_comp = {oxide: sample[oxide] for oxide in oxides}
feasible = melts.set_bulk_composition(bulk_comp)
output = melts.equilibrate_tp(temperature, pressureMPa, initialize=True)
(status, temperature, pressureMPa, xmlout) = output[0]
fluid_mass = melts.get_mass_of_phase(xmlout, phase_name='Fluid')
flsystem_wtper = 100 * fluid_mass / (fluid_mass + melts.get_mass_of_phase(xmlout, phase_name='Liquid'))
if fluid_mass > 0.0:
fluid_comp = melts.get_composition_of_phase(xmlout, phase_name='Fluid', mode='component')
fluid_comp_H2O = fluid_comp['Water']
fluid_comp_CO2 = fluid_comp['Carbon Dioxide']
else:
fluid_comp_H2O = 0
fluid_comp_CO2 = 0
feasible = melts.set_bulk_composition(bulk_comp) #reset
if verbose == False:
return {'CO2': fluid_comp_CO2, 'H2O': fluid_comp_H2O}
if verbose == True:
return {'CO2': fluid_comp_CO2, 'H2O': fluid_comp_H2O, 'FluidMass_grams': fluid_mass, 'FluidProportion_wt': flsystem_wtper}
def calculate_saturation_pressure(self, sample, temperature, verbose=False, **kwargs):
"""
Calculates the saturation pressure of a sample composition.
Parameters
----------
sample: dict, pandas Series
Compositional information on one sample. A single sample can be passed as a dict or pandas Series.
temperature: flaot or int
Temperature of the sample in degrees C.
verbose: bool
OPTIONAL: Default is False. If set to False, only the saturation pressure is returned. If set to True,
the saturation pressure, mass of fluid in grams, proportion of fluid in wt%, and H2O and CO2 concentrations
in the fluid in mole fraction are all returned in a dict.
Returns
-------
float or dict
If verbose is set to False: Saturation pressure in bars.
If verbose is set to True: dict of all calculated values.
"""
sample = self.preprocess_sample(sample)
bulk_comp_orig = sample
bulk_comp = {oxide: sample[oxide] for oxide in oxides}
feasible = melts.set_bulk_composition(bulk_comp)
#Coarse search
fluid_mass = 0
pressureMPa = 2000 #NOTE that pressure is in MPa for MagmaSat calculations but reported in bars.
while fluid_mass <= 0:
pressureMPa -= 100
if pressureMPa <= 0:
break
output = melts.equilibrate_tp(temperature, pressureMPa, initialize=True)
(status, temperature, pressureMPa, xmlout) = output[0]
fluid_mass = melts.get_mass_of_phase(xmlout, phase_name='Fluid')
pressureMPa+=100
#Refined search 1
feasible = melts.set_bulk_composition(bulk_comp)
fluid_mass = 0
while fluid_mass <= 0:
pressureMPa -= 10
if pressureMPa <= 0:
break
output = melts.equilibrate_tp(temperature, pressureMPa, initialize=True)
(status, temperature, pressureMPa, xmlout) = output[0]
fluid_mass = melts.get_mass_of_phase(xmlout, phase_name='Fluid')
pressureMPa += 10
#Refined search 2
feasible = melts.set_bulk_composition(bulk_comp)
fluid_mass = 0
while fluid_mass <= 0:
pressureMPa -= 1
if pressureMPa <= 0:
break
output = melts.equilibrate_tp(temperature, pressureMPa, initialize=True)
(status, temperature, pressureMPa, xmlout) = output[0]
fluid_mass = melts.get_mass_of_phase(xmlout, phase_name='Fluid')
if pressureMPa != np.nan:
satP = pressureMPa*10 #convert pressure to bars
flmass = fluid_mass
flsystem_wtper = 100 * fluid_mass / (fluid_mass + melts.get_mass_of_phase(xmlout, phase_name='Liquid'))
flcomp = melts.get_composition_of_phase(xmlout, phase_name='Fluid', mode='component')
try:
flH2O = flcomp['Water']
except:
flH2O = 0.0
try:
flCO2 = flcomp['Carbon Dioxide']
except:
flCO2 = 0.0
else:
flmass = np.nan
flsystem_wtper = np.nan
flH2O = np.nan
flCO2 = np.nan
warnmessage = 'Calculation failed.'
feasible = melts.set_bulk_composition(bulk_comp_orig) #this needs to be reset always!
if verbose == False:
try:
warnings.warn(warnmessage)
except:
pass
return satP
elif verbose == True:
try:
warnings.warn(warnmessage)
except:
pass
return {"SaturationP_bars": satP, "FluidMass_grams": flmass, "FluidProportion_wt": flsystem_wtper,
"XH2O_fl": flH2O, "XCO2_fl": flCO2}
def calculate_isobars_and_isopleths(self, sample, temperature, pressure_list, isopleth_list=None, print_status=False, **kwargs):
"""
Calculates isobars and isopleths at a constant temperature for a given sample. Isobars can be calculated
for any number of pressures.
Parameters
----------
sample: dict
Dictionary with values for sample composition as oxides in wt%.
temperature: float
Temperature in degrees C.
pressure_list: list
List of all pressure values at which to calculate isobars, in bars.
isopleth_list: list
OPTIONAL: Default value is None in which case only isobars will be calculated.
List of all fluid compositions in mole fraction H2O (XH2Ofluid) at which to calcualte isopleths. Values can range from 0-1.
print_status: bool
OPTIONAL: Default is False. If set to True, progress of the calculations will be printed to the terminal.
Returns
-------
pandas DataFrame objects
Two pandas DataFrames are returned; the first has isobar data, and the second has isopleth data. Columns in the
isobar dataframe are 'Pressure', 'H2Omelt', and 'CO2melt', correpsonding to pressure in bars and dissolved H2O
and CO2 in the liquid in wt%. Columns in the isopleth dataframe are 'Pressure', 'H2Ofl', and 'CO2fl',
corresponding to pressure in bars and H2O and CO2 concentration in the H2O-CO2 fluid, in wt%.
"""
sample = self.preprocess_sample(sample)
bulk_comp = sample
if isinstance(pressure_list, list):
P_vals = pressure_list
else:
raise InputError("pressure_list must be of type list")
if isopleth_list is None:
has_isopleths = False
iso_vals = [0, 0.25, 0.5, 0.75, 1]
elif isinstance(isopleth_list, list):
iso_vals = isopleth_list
has_isopleths = True
if 0 not in iso_vals:
iso_vals[0:0] = [0]
if 1 not in iso_vals:
iso_vals.append(1)
else:
raise InputError("isopleth_list must be of type list")
isobar_data = []
isopleth_data = []
for X in iso_vals:
isopleth_data.append([X, 0.0, 0.0])
H2O_val = 0.0
CO2_val = 0.0
fluid_mass = 0.0
# Calculate equilibrium phase assemblage for all P/T conditions, check if saturated in fluid...
for i in P_vals:
guess = 0.0
if print_status == True:
print("Calculating isobar at " + str(i) + " bars")
for X in iso_vals:
if print_status == True and has_isopleths == True:
print("Calculating isopleth at " + str(X))
saturated_vols = self.calculate_dissolved_volatiles(sample=sample, temperature=temperature, pressure=i, H2O_guess=guess, X_fluid=X)
isobar_data.append([i, saturated_vols['H2O'], saturated_vols['CO2']])
isopleth_data.append([X, saturated_vols['H2O'], saturated_vols['CO2']])
guess = saturated_vols['H2O']
if print_status == True:
print("Done!")
isobars_df = pd.DataFrame(isobar_data, columns=['Pressure', 'H2O_liq', 'CO2_liq'])
isopleths_df = pd.DataFrame(isopleth_data, columns=['XH2O_fl', 'H2O_liq', 'CO2_liq'])
feasible = melts.set_bulk_composition(self.bulk_comp_orig) #reset
if has_isopleths == True:
return isobars_df, isopleths_df
if has_isopleths == False:
return isobars_df, None #TODO should this just return isobars_df? Currently this requires two items to unpack, I think?
def calculate_degassing_path(self, sample, temperature, pressure='saturation', fractionate_vapor=0.0, init_vapor=0.0, **kwargs):
"""
Calculates degassing path for one sample
Parameters
----------
sample: dict
Dictionary with values for sample composition as oxides in wt%. If pulling from an uploaded file
with data for many samples, first call get_sample_oxide_comp() to get the sample desired. Then pass
the result into this function.
temperature: float
Temperature at which to calculate degassing paths, in degrees C.
pressure: float
OPTIONAL. The perssure at which to begin the degassing calculations. Default value is 'saturation', which runs the
calculation with the initial pressure at the saturation pressure. If a pressure greater than the saturation pressure
is input, the calculation will start at saturation, since this is the first pressure at which any degassing will
occur.
fractionate_vapor: float
OPTIONAL. Proportion of vapor removed at each pressure step.
Default value is 0.0 (completely closed-system degassing). Specifies the type of calculation performed, either
closed system (0.0) or open system (1.0) degassing. If any value between <1.0 is chosen, user can also specify the
'init_vapor' argument (see below). A value in between 0 and 1 will remove that proportion of vapor at each step.
For example, for a value of 0.2, the calculation will remove 20% of the vapor and retain 80% of the vapor at each
pressure step.
init_vapor: float
OPTIONAL. Default value is 0.0. Specifies the amount of vapor (in wt%) coexisting with the melt before
degassing.
Returns
-------
pandas DataFrame object
"""
sample = self.preprocess_sample(sample)
sample = normalize(sample)
bulk_comp_orig = sample
bulk_comp = {oxide: sample[oxide] for oxide in oxides}
feasible = melts.set_bulk_composition(bulk_comp)
# Get saturation pressure
data = self.calculate_saturation_pressure(sample=sample, temperature=temperature, verbose=True)
if pressure == 'saturation' or pressure >= data["SaturationP_bars"]:
SatP_MPa = data["SaturationP_bars"] / 10.0
else:
SatP_MPa = pressure / 10.0
#If pressure is low, use smaller P steps
if SatP_MPa >= 50:
MPa_step = 10
elif SatP_MPa < 50:
MPa_step = 1
P_array = np.arange(1.0, SatP_MPa, MPa_step)
P_array = -np.sort(-P_array)
fl_wtper = data["FluidProportion_wt"]
if fractionate_vapor == 0 or fractionate_vapor == 0.0: #closed-system
while fl_wtper <= init_vapor:
output = melts.equilibrate_tp(temperature, SatP_MPa)
(status, temperature, p, xmlout) = output[0]
fl_mass = melts.get_mass_of_phase(xmlout, phase_name='Fluid')
liq_mass = melts.get_mass_of_phase(xmlout, phase_name='Liquid')
fl_comp = melts.get_composition_of_phase(xmlout, phase_name='Fluid')
fl_wtper = 100 * fl_mass / (fl_mass+liq_mass)
try:
bulk_comp["H2O"] += fl_comp["H2O"]*0.0005
except:
bulk_comp["H2O"] = bulk_comp["H2O"] * 1.1
try:
bulk_comp["CO2"] += fl_comp["CO2"]*0.0005
except:
bulk_comp["CO2"] = bulk_comp["CO2"] * 1.1
bulk_comp = normalize(bulk_comp)
feasible = melts.set_bulk_composition(bulk_comp)
output = melts.equilibrate_tp(temperature, P_array)
pressure_list = []
H2Oliq = []
CO2liq = []
H2Ofl = []
CO2fl = []
fluid_wtper = []
for i in range(len(output)):
(status, temperature, p, xmlout) = output[i]
liq_comp = melts.get_composition_of_phase(xmlout, phase_name='Liquid')
fl_comp = melts.get_composition_of_phase(xmlout, phase_name='Fluid')
liq_mass = melts.get_mass_of_phase(xmlout, phase_name='Liquid')
fl_mass = melts.get_mass_of_phase(xmlout, phase_name='Fluid')
fl_wtper = 100 * fl_mass / (fl_mass+liq_mass)
pressure_list.append(p * 10.0)
try:
H2Oliq.append(liq_comp["H2O"])
except:
H2Oliq.append(0)
try:
CO2liq.append(liq_comp["CO2"])
except:
CO2liq.append(0)
try:
H2Ofl.append(fl_comp["H2O"])
except:
H2Ofl.append(0)
try:
CO2fl.append(fl_comp["CO2"])
except:
CO2fl.append(0)
fluid_wtper.append(fl_wtper)
try:
bulk_comp["H2O"] = liq_comp["H2O"]
except:
bulk_comp["H2O"] = 0
try:
bulk_comp["CO2"] = liq_comp["CO2"]
except:
bulk_comp["CO2"] = 0
fluid_wtper.append(fl_wtper)
feasible = melts.set_bulk_composition(bulk_comp_orig)
fl_wtper = data["FluidProportion_wt"]
exsolved_degassing_df = pd.DataFrame(list(zip(pressure_list, H2Oliq, CO2liq, H2Ofl, CO2fl, fluid_wtper)),
columns =['Pressure_bars', 'H2O_liq', 'CO2_liq', 'H2O_fl', 'CO2_fl', 'FluidProportion_wt'])
return exsolved_degassing_df
else:
pressure = []
H2Oliq = []
CO2liq = []
H2Ofl = []
CO2fl = []
fluid_wtper = []
for i in P_array:
fl_mass = 0.0
feasible = melts.set_bulk_composition(bulk_comp)
output = melts.equilibrate_tp(temperature, i)
(status, temperature, p, xmlout) = output[0]
liq_comp = melts.get_composition_of_phase(xmlout, phase_name='Liquid')
fl_comp = melts.get_composition_of_phase(xmlout, phase_name='Fluid', mode='component')
liq_mass = melts.get_mass_of_phase(xmlout, phase_name='Liquid')
fl_mass = melts.get_mass_of_phase(xmlout, phase_name='Fluid')
fl_wtper = 100 * fl_mass / (fl_mass+liq_mass)
if fl_mass > 0:
pressure.append(p * 10.0)
try:
H2Oliq.append(liq_comp["H2O"])
except:
H2Oliq.append(0)
try:
CO2liq.append(liq_comp["CO2"])
except:
CO2liq.append(0)
try:
H2Ofl.append(fl_comp["Water"])
except:
H2Ofl.append(0)
try:
CO2fl.append(fl_comp["Carbon Dioxide"])
except:
CO2fl.append(0)
fluid_wtper.append(fl_wtper)
try:
bulk_comp["H2O"] = liq_comp["H2O"] + (bulk_comp["H2O"] - liq_comp["H2O"]) * (1-fractionate_vapor)
except:
bulk_comp["H2O"] = 0
try:
bulk_comp["CO2"] = liq_comp["CO2"] + (bulk_comp["CO2"] - liq_comp["CO2"]) * (1-fractionate_vapor)
except:
bulk_comp["CO2"] = 0
bulk_comp = normalize(bulk_comp)
feasible = melts.set_bulk_composition(bulk_comp_orig) #this needs to be reset always!
open_degassing_df = pd.DataFrame(list(zip(pressure, H2Oliq, CO2liq, H2Ofl, CO2fl, fluid_wtper)),
columns =['Pressure_bars', 'H2O_liq', 'CO2_liq', 'XH2O_fl', 'XCO2_fl', 'FluidProportion_wt'])
return open_degassing_df
#-----------MAGMASAT PLOTTING FUNCTIONS-----------#
def smooth_isobars_and_isopleths(isobars=None, isopleths=None):
"""
Takes in a dataframe with calculated isobar and isopleth information (e.g., output from calculate_isobars_and_isopleths)
and smooths the data for plotting.
Parameters
----------
isobars: pandas DataFrame
OPTIONAL. DataFrame object containing isobar information as calculated by calculate_isobars_and_isopleths.
isopleths: pandas DataFrame
OPTIONAL. DataFrame object containing isopleth information as calculated by calculate_isobars_and_isopleths.
Returns
-------
pandas DataFrame
DataFrame with x and y values for all isobars and all isopleths. Useful if a user wishes to do custom plotting
with isobar and isopleth data rather than using the built-in `plot_isobars_and_isopleths()` function.
"""
if isobars is not None:
P_vals = isobars.Pressure.unique()
isobars_lists = isobars.values.tolist()
# add zero values to volatiles list
isobars_lists.append([0.0, 0.0, 0.0, 0.0])
isobars = {}
# do some data smoothing
for pressure in P_vals:
Pxs = [item[1] for item in isobars_lists if item[0] == pressure]
Pys = [item[2] for item in isobars_lists if item[0] == pressure]
try:
np.seterr(divide='ignore', invalid='ignore') #turn off numpy warning
## calcualte polynomial
Pz = np.polyfit(Pxs, Pys, 3)
Pf = np.poly1d(Pz)
## calculate new x's and y's
Px_new = np.linspace(Pxs[0], Pxs[-1], 50)
Py_new = Pf(Px_new)
# Save x's and y's
isobars.update({str(pressure)+"xvals": Px_new})
isobars.update({str(pressure)+"yvals": Py_new})
except:
isobars.update({str(pressure)+"xvals": Pxs})
isobars.update({str(pressure)+"yvals": Pys})
if isopleths is not None:
XH2O_vals = isopleths.XH2O_fl.unique()
isopleths_lists = isopleths.values.tolist()
isopleths = {}
for Xfl in XH2O_vals:
Xxs = [item[1] for item in isopleths_lists if item[0] == Xfl]
Xys = [item[2] for item in isopleths_lists if item[0] == Xfl]
try:
## calcualte polynomial
Xz = np.polyfit(Xxs, Xys, 2)
Xf = np.poly1d(Xz)
## calculate new x's and y's
Xx_new = np.linspace(Xxs[0], Xxs[-1], 50)
Xy_new = Xf(Xx_new)
# Save x's and y's
isopleths.update({str(Xfl)+"xvals":Xx_new})
isopleths.update({str(Xfl)+"yvals":Xy_new})
except:
isopleths.update({str(Xfl)+"xvals":Xxs})
isopleths.update({str(Xfl)+"yvals":Xys})
np.seterr(divide='warn', invalid='warn') #turn numpy warning back on
if isobars is not None:
if isopleths is not None:
return pd.DataFrame(isobars), pd.DataFrame(isopleths)
else:
return pd.DataFrame(isobars)
else:
if isopleths is not None:
isopleth_frame = pd.DataFrame.from_dict(isopleths, orient='index')
isopleth_frame = isopleth_frame.transpose()
print(isopleth_frame)
return pd.DataFrame(isopleth_frame)
def plot(isobars=None, isopleths=None, degassing_paths=None, custom_H2O=None, custom_CO2=None,
isobar_labels=None, isopleth_labels=None, degassing_path_labels=None, custom_labels=None,
extend_isobars_to_zero=True, **kwargs):
"""
Custom automatic plotting of model calculations in VESIcal.
Isobars, isopleths, and degassing paths can be plotted. Labels can be specified for each.
Any combination of isobars, isopleths, and degassing paths can be plotted.
Parameters
----------
isobars: pandas DataFrame or list
OPTIONAL. DataFrame object containing isobar information as calculated by calculate_isobars_and_isopleths. Or a list
of DataFrame objects.
isopleths: pandas DataFrame or list
OPTIONAL. DataFrame object containing isopleth information as calculated by calculate_isobars_and_isopleths. Or a list
of DataFrame objects.
degassing_paths: list
OPTIONAL. List of DataFrames with degassing information as generated by calculate_degassing_path().
custom_H2O: list
OPTIONAL. List of groups of H2O values to plot as points. For example myfile.data['H2O'] is one group of H2O values.
Must be passed with custom_CO2 and must be same length as custom_CO2.
custom_CO2: list
OPTIONAL. List of groups of CO2 values to plot as points.For example myfile.data['CO2'] is one group of CO2 values.
Must be passed with custom_H2O and must be same length as custom_H2O.
isobar_labels: list
OPTIONAL. Labels for the plot legend. Default is None, in which case each plotted line will be given the generic
legend name of "Isobars n", with n referring to the nth isobars passed. Isobar pressure is given in parentheses.
The user can pass their own labels as a list of strings. If more than one set of isobars is passed, the labels should
refer to each set of isobars, not each pressure.
isopleth_labels: list
OPTIONAL. Labels for the plot legend. Default is None, in which case each plotted isopleth will be given the generic
legend name of "Isopleth n", with n referring to the nth isopleths passed. Isopleth XH2O values are given in
parentheses. The user can pass their own labels as a list of strings. If more than one set of isopleths is passed,
the labels should refer to each set of isopleths, not each XH2O value.
degassing_path_labels: list
OPTIONAL. Labels for the plot legend. Default is None, in which case each plotted line will be given the generic
legend name of "Pathn", with n referring to the nth degassing path passed. The user can pass their own labels
as a list of strings.
custom_labels: list
OPTIONAL. Labels for the plot legend. Default is None, in which case each group of custom points will be given the
generic legend name of "Customn", with n referring to the nth degassing path passed. The user can pass their own labels
as a list of strings.
extend_isobars_to_zero bool
If True (default), isobars will be extended to zero, even if there is a finite solubility at zero partial pressure.
Returns
-------
matplotlib object
Plot with x-axis as H2O wt% in the melt and y-axis as CO2 wt% in the melt. Isobars, or lines of
constant pressure at which the sample magma composition is saturated, and isopleths, or lines of constant
fluid composition at which the sample magma composition is saturated, are plotted if passed. Degassing
paths, or the concentration of dissolved H2O and CO2 in a melt equilibrated along a path of decreasing
pressure, is plotted if passed.
"""
if custom_H2O is not None:
if custom_CO2 is None:
raise InputError("If x data is passed, y data must also be passed.")
else:
if len(custom_H2O) == len(custom_CO2):
pass
else:
raise InputError("x and y data must be same length")
if custom_CO2 is not None:
if custom_H2O is None:
raise InputError("If y data is passed, x data must also be passed.")
plt.figure(figsize=(12,8))
plt.xlabel('H$_2$O wt%')
plt.ylabel('CO$_2$ wt%')
labels = []
if isobars is not None:
if isinstance(isobars, pd.DataFrame):
isobars = [isobars]
for i in range(len(isobars)):
P_vals = isobars[i].Pressure.unique()
isobars_lists = isobars[i].values.tolist()
# add zero values to volatiles list
isobars_lists.append([0.0, 0.0, 0.0, 0.0])
np.seterr(divide='ignore', invalid='ignore') #turn off numpy warning
warnings.filterwarnings("ignore", message="Polyfit may be poorly conditioned")
# do some data smoothing
P_iter = 0
for pressure in P_vals:
P_iter += 1
Pxs = [item[1] for item in isobars_lists if item[0] == pressure]
Pys = [item[2] for item in isobars_lists if item[0] == pressure]
if len(isobars) > 1:
if P_iter == 1:
P_list = [int(i) for i in P_vals]
if isinstance(isobar_labels, list):
labels.append(str(isobar_labels[i]) + ' (' + ', '.join(map(str, P_list)) + " bars)")
else:
labels.append('Isobars ' + str(i+1) + ' (' + ', '.join(map(str, P_list)) + " bars)")
else:
labels.append('_nolegend_')
try:
np.seterr(divide='ignore', invalid='ignore') #turn off numpy warning
## calcualte polynomial
Pz = np.polyfit(Pxs, Pys, 3)
Pf = np.poly1d(Pz)
## calculate new x's and y's
Px_new = np.linspace(Pxs[0], Pxs[-1], 50)
Py_new = Pf(Px_new)
if extend_isobars_to_zero == True and Px_new[0]*Py_new[0] != 0.0:
if Px_new[0] > Py_new[0]:
Px_newer = np.zeros(np.shape(Px_new)[0]+1)
Px_newer[0] = 0
Px_newer[1:] = Px_new
Px_new = Px_newer
Py_newer = np.zeros(np.shape(Py_new)[0]+1)
Py_newer[0] = Py_new[0]
Py_newer[1:] = Py_new
Py_new = Py_newer
else:
Px_newer = np.zeros(np.shape(Px_new)[0]+1)
Px_newer[0] = Px_new[0]
Px_newer[1:] = Px_new
Px_new = Px_newer
Py_newer = np.zeros(np.shape(Py_new)[0]+1)
Py_newer[0] = 0
Py_newer[1:] = Py_new
Py_new = Py_newer
if extend_isobars_to_zero == True and Px_new[-1]*Py_new[-1] != 0.0:
if Px_new[-1] < Py_new[-1]:
Px_newer = np.zeros(np.shape(Px_new)[0]+1)
Px_newer[-1] = 0
Px_newer[:-1] = Px_new
Px_new = Px_newer
Py_newer = np.zeros(np.shape(Py_new)[0]+1)
Py_newer[-1] = Py_new[-1]
Py_newer[:-1] = Py_new
Py_new = Py_newer
else:
Px_newer = np.zeros(np.shape(Px_new)[0]+1)
Px_newer[-1] = Px_new[-1]
Px_newer[:-1] = Px_new
Px_new = Px_newer
Py_newer = np.zeros(np.shape(Py_new)[0]+1)
Py_newer[-1] = 0
Py_newer[:-1] = Py_new
Py_new = Py_newer
# Plot some stuff
if len(isobars) > 1:
plt.plot(Px_new, Py_new, color=color_list[i])
else:
plt.plot(Px_new, Py_new)
except:
if len(isobars) > 1:
plt.plot(Pxs, Pys, color=color_list[i])
else:
plt.plot(Pxs, Pys)
if len(isobars) == 1:
labels = [str(P_val) + " bars" for P_val in P_vals]
if isopleths is not None:
if isinstance(isopleths, pd.DataFrame):
isopleths = [isopleths]
for i in range(len(isopleths)):
XH2O_vals = isopleths[i].XH2O_fl.unique()
isopleths_lists = isopleths[i].values.tolist()
np.seterr(divide='ignore', invalid='ignore') #turn off numpy warning
warnings.filterwarnings("ignore", message="Polyfit may be poorly conditioned")
# do some data smoothing
H_iter = 0
for Xfl in XH2O_vals:
H_iter += 1
Xxs = [item[1] for item in isopleths_lists if item[0] == Xfl]
Xys = [item[2] for item in isopleths_lists if item[0] == Xfl]
if len(isopleths) > 1:
if H_iter == 1:
H_list = [i for i in XH2O_vals]
if isinstance(isopleth_labels, list):
labels.append(str(isopleth_labels[i]) + ' (' + ', '.join(map(str, H_list)) + " XH2Ofluid)")
else:
labels.append('Isopleths ' + str(i+1) + ' (' + ', '.join(map(str, H_list)) + " XH2Ofluid)")
else:
labels.append('_nolegend_')
try:
np.seterr(divide='ignore', invalid='ignore') #turn off numpy warning
## calcualte polynomial
Xz = np.polyfit(Xxs, Xys, 2)
Xf = np.poly1d(Xz)
## calculate new x's and y's
Xx_new = np.linspace(Xxs[0], Xxs[-1], 50)
Xy_new = Xf(Xx_new)
# Plot some stuff
if len(isopleths) == 1:
plt.plot(Xx_new, Xy_new, ls='dashed', color='k')
else:
plt.plot(Xx_new, Xy_new, ls='dashed', color=color_list[i])
except:
if len(isopleths) == 1:
plt.plot(Xxs, Xys, ls='dashed', color='k')
else:
plt.plot(Xxs, Xys, ls='dashed', color=color_list[i])
if len(isopleths) == 1:
H_list = [i for i in XH2O_vals]
iso_label_iter = 0
for i in XH2O_vals:
iso_label_iter += 1
if iso_label_iter == 1:
labels.append('Isopleths (' + ', '.join(map(str, H_list)) + " XH2Ofluid)")
else:
labels.append('_nolegend_')
if degassing_paths is not None:
if isinstance(degassing_paths, pd.DataFrame):
degassing_paths = [degassing_paths]
degassing_colors = color_list.copy()
degassing_colors.reverse()
iterno = 0
for i in range(len(degassing_paths)):
if degassing_path_labels == None:
iterno += 1
labels.append('Path%s' %iterno)
plt.plot(degassing_paths[i]["H2O_liq"], degassing_paths[i]["CO2_liq"], '-', color=degassing_colors[i])
else:
labels.append(degassing_path_labels[iterno])
plt.plot(degassing_paths[i]["H2O_liq"], degassing_paths[i]["CO2_liq"], '-', color=degassing_colors[i])
iterno += 1
for i in range(len(degassing_paths)):
plt.plot(degassing_paths[i]["H2O_liq"].max(), degassing_paths[i]["CO2_liq"].max(), 'o', color=degassing_colors[i])
labels.append('_nolegend_')
if custom_H2O is not None and custom_CO2 is not None:
if isinstance(custom_H2O, pd.DataFrame):
custom_H2O = [custom_H2O]
if isinstance(custom_CO2, pd.DataFrame):
custom_CO2 = [custom_CO2]
iterno = 0
for i in range(len(custom_H2O)):
if custom_labels == None:
iterno +=1
labels.append('Custom%s' %iterno)
plt.plot(custom_H2O[i], custom_CO2[i], 'o', color=color_list[i])
else:
labels.append(custom_labels[iterno])
plt.plot(custom_H2O[i], custom_CO2[i], 'o', color=color_list[i])
iterno += 1
plt.legend(labels)
plt.xlim(left=0)
plt.ylim(bottom=0)
|
np.seterr(divide='warn', invalid='warn')
|
numpy.seterr
|
import os
import anndata
import mudata
import numpy as np
import pandas as pd
import pytest
from scvi import REGISTRY_KEYS
from scvi.data import synthetic_iid
from .utils import generic_setup_mudata_manager
def test_setup_mudata():
adata = synthetic_iid()
adata.obs["cont1"] =
|
np.random.normal(size=(adata.shape[0],))
|
numpy.random.normal
|
"""
Some functions for working with the Abel habit model.
"""
import numpy as np
from numpy import sqrt, exp
from scipy.stats import norm
import quantecon as qe
inv_sqrt_2pi = 1 / sqrt(2 * np.pi)
class AbelModel:
"""
Represents the model.
"""
def __init__(self, β=0.99,
γ=2.5,
ρ=0.9,
σ=0.002,
x0=0.1,
α=1,
grid_size=60):
self.β, self.γ, self.ρ, self.σ = β, γ, ρ, σ
self.α, self.x0 = α, x0
# derived constants
self.b = x0 + σ**2 * (1 - γ)
self.k0 = β * exp(self.b * (1 - γ) + σ**2 * (1 - γ)**2 / 2)
self.k1 = (ρ - α) * (1 - γ)
# Parameters in the stationary distribution
self.svar = σ**2 / (1 - ρ**2)
self.ssd = sqrt(self.svar)
self.smean = self.b / (1 - ρ)
# A discrete approximation of the stationary dist
std_range, n = 3, 20
mc = qe.tauchen(0, 1, std_range, n)
w_vec = mc.state_values
self.sx_vec = self.smean + self.ssd * w_vec
self.sp_vec = mc.P[0, :] # Any row
# A grid of points for interpolation
a, b = self.smean + 3 * self.ssd, self.smean - 3 * self.ssd
self.x_grid = np.linspace(a, b, grid_size)
def sim_state(self, x0=None, num_paths=1000, ts_length=1000):
"""
Simulate the state process. If x0 is None, then
draw from the stationary distribution.
"""
ρ, b, σ = self.ρ, self.b, self.σ
X = np.ones((num_paths, ts_length))
W = np.random.randn(num_paths, ts_length)
if x0 is None:
X[:, 0] = self.smean
else:
X[:, 0] = x0
for t in range(ts_length-1):
X[:, t+1] = ρ * X[:, t] + b + σ * W[:, t+1]
return X
def A(self, g, Ag, std_range=3, shock_state_size=20):
"""
Apply A to g and return Ag. The argument g is a vector, which is
converted to a function by linear interpolation.
Integration uses Gaussian quadrature.
"""
# Unpack parameters
β, γ, ρ, σ, x0, α = self.β, self.γ, self.ρ, self.σ, self.x0, self.α
b, k0, k1 = self.b, self.k0, self.k1
# Extract state and probs for N(0, 1) shocks
mc = qe.tauchen(0, 1, std_range, shock_state_size)
w_vec = mc.state_values
p_vec = mc.P[0, :] # Any row, all columns
# Interpolate g and allocate memory for new g
g_func = lambda x: np.interp(x, self.x_grid, g)
# Apply the operator K to g, computing Kg and || Kg ||
for (i, x) in enumerate(self.x_grid):
mf = k0 * exp(k1 * x)
Ag[i] = mf * np.dot(g_func(ρ * x + b + w_vec), p_vec)
# Calculate the norm of Ag
Ag_func = lambda x: np.interp(x, self.x_grid, Ag)
r = np.sqrt(np.dot(Ag_func(self.sx_vec)**2, self.sp_vec))
return r
def local_spec_rad_iterative(self, tol=1e-7, max_iter=5000):
"""
Compute the spectral radius of the operator A associated with the Abel
model self via the local spectral radios
"""
n = len(self.x_grid)
g_in =
|
np.ones(n)
|
numpy.ones
|
# Masses of compact remnant from CO core masses
__author__ = "<NAME> (<EMAIL>)"
# for fit
import numpy as np
import scipy
from scipy.optimize import curve_fit
# for plot
import matplotlib as mpl
import matplotlib.pyplot as plt
import matplotlib.gridspec as gridspec
def linear(x, a, b):
return a * x + b
def fitting_func_Z(data, a, b, c, d):
"""
shifted cube plus square term, with the coefficient of the cubic
term linear function in log10(Z)
"""
mco = data[0]
Z = data[1]
return linear(
|
np.log10(Z)
|
numpy.log10
|
###
# pySuStaIn: a Python implementation of the Subtype and Stage Inference (SuStaIn) algorithm
#
# If you use pySuStaIn, please cite the following core papers:
# 1. The original SuStaIn paper: https://doi.org/10.1038/s41467-018-05892-0
# 2. The pySuStaIn software paper: https://doi.org/10.1016/j.softx.2021.100811
#
# Please also cite the corresponding progression pattern model you use:
# 1. The piece-wise linear z-score model (i.e. ZscoreSustain): https://doi.org/10.1038/s41467-018-05892-0
# 2. The event-based model (i.e. MixtureSustain): https://doi.org/10.1016/j.neuroimage.2012.01.062
# with Gaussian mixture modeling (i.e. 'mixture_gmm'): https://doi.org/10.1093/brain/awu176
# or kernel density estimation (i.e. 'mixture_kde'): https://doi.org/10.1002/alz.12083
# 3. The model for discrete ordinal data (i.e. OrdinalSustain): https://doi.org/10.3389/frai.2021.613261
#
# Thanks a lot for supporting this project.
#
# Authors: <NAME> (<EMAIL>) and <NAME> (<EMAIL>)
# Contributors: <NAME> (<EMAIL>), <NAME> (<EMAIL>), <NAME> (<EMAIL>)
###
import warnings
from tqdm.auto import tqdm
import numpy as np
import scipy.stats as stats
from matplotlib import pyplot as plt
from pySuStaIn.AbstractSustain import AbstractSustainData
from pySuStaIn.AbstractSustain import AbstractSustain
#*******************************************
#The data structure class for MixtureSustain. It holds the positive/negative likelihoods that get passed around and re-indexed in places.
class MixtureSustainData(AbstractSustainData):
def __init__(self, L_yes, L_no, numStages):
assert(L_yes.shape[0] == L_no.shape[0] and L_yes.shape[1] == L_no.shape[1])
self.L_yes = L_yes
self.L_no = L_no
self.__numStages = numStages
def getNumSamples(self):
return self.L_yes.shape[0]
def getNumBiomarkers(self):
return self.L_no.shape[1]
def getNumStages(self):
return self.__numStages
def reindex(self, index):
return MixtureSustainData(self.L_yes[index,], self.L_no[index,], self.__numStages)
#*******************************************
#An implementation of the AbstractSustain class with mixture model based events
class MixtureSustain(AbstractSustain):
def __init__(self,
L_yes,
L_no,
biomarker_labels,
N_startpoints,
N_S_max,
N_iterations_MCMC,
output_folder,
dataset_name,
use_parallel_startpoints,
seed=None):
# The initializer for the mixture model based events implementation of AbstractSustain
# Parameters:
# L_yes - probability of positive class for all subjects across all biomarkers (from mixture modelling)
# dim: number of subjects x number of biomarkers
# L_no - probability of negative class for all subjects across all biomarkers (from mixture modelling)
# dim: number of subjects x number of biomarkers
# biomarker_labels - the names of the biomarkers as a list of strings
# N_startpoints - number of startpoints to use in maximum likelihood step of SuStaIn, typically 25
# N_S_max - maximum number of subtypes, should be 1 or more
# N_iterations_MCMC - number of MCMC iterations, typically 1e5 or 1e6 but can be lower for debugging
# output_folder - where to save pickle files, etc.
# dataset_name - for naming pickle files
# use_parallel_startpoints - boolean for whether or not to parallelize the maximum likelihood loop
# seed - random number seed
N = L_yes.shape[1] # number of biomarkers
assert (len(biomarker_labels) == N), "number of labels should match number of biomarkers"
self.biomarker_labels = biomarker_labels
numStages = L_yes.shape[1] #number of stages == number of biomarkers here
self.__sustainData = MixtureSustainData(L_yes, L_no, numStages)
super().__init__(self.__sustainData,
N_startpoints,
N_S_max,
N_iterations_MCMC,
output_folder,
dataset_name,
use_parallel_startpoints,
seed)
def _initialise_sequence(self, sustainData, rng):
# Randomly initialises a sequence
S = rng.permutation(sustainData.getNumStages()) #np.random.permutation(sustainData.getNumStages())
S = S.reshape(1, len(S))
return S
def _calculate_likelihood_stage(self, sustainData, S):
'''
Computes the likelihood of a single event based model
Inputs:
=======
sustainData - a MixtureData type that contains:
L_yes - likelihood an event has occurred in each subject
dim: number of subjects x number of biomarkers
L_no - likelihood an event has not occurred in each subject
dim: number of subjects x number of biomarkers
S - the current ordering of the z-score stages for a particular subtype
dim: 1 x number of events
Outputs:
========
p_perm_k - the probability of each subjects data at each stage of a particular subtype
in the SuStaIn model
'''
M = sustainData.getNumSamples()
N = sustainData.getNumStages()
S_int = S.astype(int)
arange_Np1 = np.arange(0, N+1)
p_perm_k = np.zeros((M, N+1))
#**** THIS VERSION IS ROUGHLY 10x FASTER THAN THE ONE BELOW
cp_yes = np.cumprod(sustainData.L_yes[:, S_int], 1)
cp_no = np.cumprod(sustainData.L_no[:, S_int[::-1]], 1) #do the cumulative product from the end of the sequence
# Even faster version to avoid loops
p_perm_k[:, 0] = cp_no[:, -1]
p_perm_k[:, 1:-1] = cp_no[:, :-1][:, ::-1] * cp_yes[:, :-1]
p_perm_k[:, -1] = cp_yes[:, -1]
p_perm_k *= 1 / (N + 1)
return p_perm_k
def _optimise_parameters(self, sustainData, S_init, f_init, rng):
# Optimise the parameters of the SuStaIn model
M = sustainData.getNumSamples()
N_S = S_init.shape[0]
N = sustainData.getNumStages()
S_opt = S_init.copy() # have to copy or changes will be passed to S_init
f_opt = np.array(f_init).reshape(N_S, 1, 1)
f_val_mat = np.tile(f_opt, (1, N + 1, M))
f_val_mat = np.transpose(f_val_mat, (2, 1, 0))
p_perm_k = np.zeros((M, N + 1, N_S))
for s in range(N_S):
p_perm_k[:, :, s] = self._calculate_likelihood_stage(sustainData, S_opt[s])
p_perm_k_weighted = p_perm_k * f_val_mat
# the second summation axis is different to Matlab version
#p_perm_k_norm = p_perm_k_weighted / np.tile(np.sum(np.sum(p_perm_k_weighted, 1), 1).reshape(M, 1, 1), (1, N + 1, N_S))
# adding 1e-250 fixes divide by zero problem that happens rarely
p_perm_k_norm = p_perm_k_weighted / np.sum(p_perm_k_weighted + 1e-250, axis=(1, 2), keepdims=True)
f_opt = (np.squeeze(sum(sum(p_perm_k_norm))) / sum(sum(sum(p_perm_k_norm)))).reshape(N_S, 1, 1)
f_val_mat = np.tile(f_opt, (1, N + 1, M))
f_val_mat = np.transpose(f_val_mat, (2, 1, 0))
order_seq = rng.permutation(N_S) #np.random.permutation(N_S) # this will produce different random numbers to Matlab
for s in order_seq:
order_bio = rng.permutation(N) #np.random.permutation(N) # this will produce different random numbers to Matlab
for i in order_bio:
current_sequence = S_opt[s]
assert(len(current_sequence)==N)
current_location = np.array([0] * N)
current_location[current_sequence.astype(int)] = np.arange(N)
selected_event = i
move_event_from = current_location[selected_event]
possible_positions = np.arange(N)
possible_sequences = np.zeros((len(possible_positions), N))
possible_likelihood = np.zeros((len(possible_positions), 1))
possible_p_perm_k = np.zeros((M, N + 1, len(possible_positions)))
for index in range(len(possible_positions)):
current_sequence = S_opt[s]
#choose a position in the sequence to move an event to
move_event_to = possible_positions[index]
#move this event in its new position
current_sequence = np.delete(current_sequence, move_event_from, 0) # this is different to the Matlab version, which call current_sequence(move_event_from) = []
new_sequence = np.concatenate([current_sequence[
|
np.arange(move_event_to)
|
numpy.arange
|
#!/usr/bin/env python3
# -*- coding:utf-8 -*-
# Copyright (c) Megvii, Inc. and its affiliates.
import os
from loguru import logger
import cv2
import copy
import json
import torch
import numpy as np
from pycocotools.coco import COCO
from .. import RandomShapeSingle, YOLOXResizeImage
from ..dataloading import get_yolox_datadir
from .datasets_wrapper import Dataset
class COCODataset(Dataset):
"""
COCO dataset class.
"""
def __init__(
self,
data_dir=None,
json_file="instances_train2017.json",
ann_folder="annotations",
name="train2017",
img_size=(416, 416),
preproc=None,
cache=False,
):
"""
COCO dataset initialization. Annotation data are read into memory by COCO API.
Args:
data_dir (str): dataset root directory
json_file (str): COCO json file name
ann_folder (str): COCO annotations folder name (e.g. 'annotations')
name (str): COCO data name (e.g. 'train2017' or 'val2017')
img_size (int): target image size after pre-processing
preproc: data augmentation strategy
"""
super().__init__(img_size)
if data_dir is None:
data_dir = os.path.join(get_yolox_datadir(), "COCO")
self.data_dir = data_dir
self.json_file = json_file
self.ann_folder = ann_folder
self.coco = COCO(os.path.join(self.data_dir, self.ann_folder, self.json_file))
self.ids = self.coco.getImgIds()
self.class_ids = sorted(self.coco.getCatIds())
cats = self.coco.loadCats(self.coco.getCatIds())
self._classes = tuple([c["name"] for c in cats])
self.imgs = None
self.name = name
self.img_size = img_size
self.preproc = preproc
self.annotations = self._load_coco_annotations()
if cache:
self._cache_images()
def __len__(self):
return len(self.ids)
def __del__(self):
del self.imgs
def _load_coco_annotations(self):
return [self.load_anno_from_ids(_ids) for _ids in self.ids]
def _cache_images(self):
logger.warning(
"\n********************************************************************************\n"
"You are using cached images in RAM to accelerate training.\n"
"This requires large system RAM.\n"
"Make sure you have 200G+ RAM and 136G available disk space for training COCO.\n"
"********************************************************************************\n"
)
max_h = self.img_size[0]
max_w = self.img_size[1]
cache_file = self.data_dir + "/img_resized_cache_" + self.name + ".array"
if not os.path.exists(cache_file):
logger.info(
"Caching images for the first time. This might take about 20 minutes for COCO"
)
self.imgs = np.memmap(
cache_file,
shape=(len(self.ids), max_h, max_w, 3),
dtype=np.uint8,
mode="w+",
)
from tqdm import tqdm
from multiprocessing.pool import ThreadPool
NUM_THREADs = min(8, os.cpu_count())
loaded_images = ThreadPool(NUM_THREADs).imap(
lambda x: self.load_resized_img(x),
range(len(self.annotations)),
)
pbar = tqdm(enumerate(loaded_images), total=len(self.annotations))
for k, out in pbar:
self.imgs[k][: out.shape[0], : out.shape[1], :] = out.copy()
self.imgs.flush()
pbar.close()
else:
logger.warning(
"You are using cached imgs! Make sure your dataset is not changed!!\n"
"Everytime the self.input_size is changed in your exp file, you need to delete\n"
"the cached data and re-generate them.\n"
)
logger.info("Loading cached imgs...")
self.imgs = np.memmap(
cache_file,
shape=(len(self.ids), max_h, max_w, 3),
dtype=np.uint8,
mode="r+",
)
def load_anno_from_ids(self, id_):
im_ann = self.coco.loadImgs(id_)[0]
width = im_ann["width"]
height = im_ann["height"]
anno_ids = self.coco.getAnnIds(imgIds=[int(id_)], iscrowd=False)
annotations = self.coco.loadAnns(anno_ids)
objs = []
for obj in annotations:
x1 = np.max((0, obj["bbox"][0]))
y1 = np.max((0, obj["bbox"][1]))
x2 = np.min((width, x1 + np.max((0, obj["bbox"][2]))))
y2 = np.min((height, y1 + np.max((0, obj["bbox"][3]))))
if obj["area"] > 0 and x2 >= x1 and y2 >= y1:
obj["clean_bbox"] = [x1, y1, x2, y2]
objs.append(obj)
num_objs = len(objs)
res = np.zeros((num_objs, 5))
for ix, obj in enumerate(objs):
cls = self.class_ids.index(obj["category_id"])
res[ix, 0:4] = obj["clean_bbox"]
res[ix, 4] = cls
r = min(self.img_size[0] / height, self.img_size[1] / width)
res[:, :4] *= r
img_info = (height, width)
resized_info = (int(height * r), int(width * r))
file_name = (
im_ann["file_name"]
if "file_name" in im_ann
else "{:012}".format(id_) + ".jpg"
)
return (res, img_info, resized_info, file_name)
def load_anno(self, index):
return self.annotations[index][0]
def load_resized_img(self, index):
img = self.load_image(index)
r = min(self.img_size[0] / img.shape[0], self.img_size[1] / img.shape[1])
resized_img = cv2.resize(
img,
(int(img.shape[1] * r), int(img.shape[0] * r)),
interpolation=cv2.INTER_LINEAR,
).astype(np.uint8)
return resized_img
def load_image(self, index):
file_name = self.annotations[index][3]
img_file = os.path.join(self.data_dir, self.name, file_name)
img = cv2.imread(img_file)
assert img is not None
return img
def pull_item(self, index):
id_ = self.ids[index]
res, img_info, resized_info, _ = self.annotations[index]
if self.imgs is not None:
pad_img = self.imgs[index]
img = pad_img[: resized_info[0], : resized_info[1], :].copy()
else:
img = self.load_resized_img(index)
return img, res.copy(), img_info, np.array([id_])
@Dataset.mosaic_getitem
def __getitem__(self, index):
"""
One image / label pair for the given index is picked up and pre-processed.
Args:
index (int): data index
Returns:
img (numpy.ndarray): pre-processed image
padded_labels (torch.Tensor): pre-processed label data.
The shape is :math:`[max_labels, 5]`.
each label consists of [class, xc, yc, w, h]:
class (float): class index.
xc, yc (float) : center of bbox whose values range from 0 to 1.
w, h (float) : size of bbox whose values range from 0 to 1.
info_img : tuple of h, w.
h, w (int): original shape of the image
img_id (int): same as the input index. Used for evaluation.
"""
img, target, img_info, img_id = self.pull_item(index)
if self.preproc is not None:
img, target = self.preproc(img, target, self.input_dim)
return img, target, img_info, img_id
# 数据清洗
def data_clean(coco, img_ids, catid2clsid, image_dir, type):
records = []
ct = 0
for img_id in img_ids:
img_anno = coco.loadImgs(img_id)[0]
im_fname = img_anno['file_name']
im_w = float(img_anno['width'])
im_h = float(img_anno['height'])
ins_anno_ids = coco.getAnnIds(imgIds=img_id, iscrowd=False) # 读取这张图片所有标注anno的id
instances = coco.loadAnns(ins_anno_ids) # 这张图片所有标注anno。每个标注有'segmentation'、'bbox'、...
bboxes = []
anno_id = [] # 注解id
for inst in instances:
x, y, box_w, box_h = inst['bbox'] # 读取物体的包围框
x1 = max(0, x)
y1 = max(0, y)
x2 = min(im_w - 1, x1 + max(0, box_w - 1))
y2 = min(im_h - 1, y1 + max(0, box_h - 1))
if inst['area'] > 0 and x2 >= x1 and y2 >= y1:
inst['clean_bbox'] = [x1, y1, x2, y2] # inst增加一个键值对
bboxes.append(inst) # 这张图片的这个物体标注保留
anno_id.append(inst['id'])
else:
logger.warn(
'Found an invalid bbox in annotations: im_id: {}, '
'area: {} x1: {}, y1: {}, x2: {}, y2: {}.'.format(
img_id, float(inst['area']), x1, y1, x2, y2))
num_bbox = len(bboxes) # 这张图片的物体数
# 左上角坐标+右下角坐标+类别id
gt_bbox = np.zeros((num_bbox, 4), dtype=np.float32)
gt_class = np.zeros((num_bbox, 1), dtype=np.int32)
gt_score = np.ones((num_bbox, 1), dtype=np.float32) # 得分的标注都是1
is_crowd = np.zeros((num_bbox, 1), dtype=np.int32)
difficult = np.zeros((num_bbox, 1), dtype=np.int32)
gt_poly = [None] * num_bbox
for i, box in enumerate(bboxes):
catid = box['category_id']
gt_class[i][0] = catid2clsid[catid]
gt_bbox[i, :] = box['clean_bbox']
is_crowd[i][0] = box['iscrowd']
if 'segmentation' in box:
gt_poly[i] = box['segmentation']
im_fname = os.path.join(image_dir,
im_fname) if image_dir else im_fname
coco_rec = {
'im_file': im_fname,
'im_id': np.array([img_id]),
'h': im_h,
'w': im_w,
'is_crowd': is_crowd,
'gt_class': gt_class,
'anno_id': anno_id,
'gt_bbox': gt_bbox,
'gt_score': gt_score,
'gt_poly': gt_poly,
}
# logger.debug('Load file: {}, im_id: {}, h: {}, w: {}.'.format(im_fname, img_id, im_h, im_w))
records.append(coco_rec) # 注解文件。
ct += 1
logger.info('{} samples in {} set.'.format(ct, type))
return records
def get_class_msg(anno_path):
_catid2clsid = {}
_clsid2catid = {}
_clsid2cname = {}
with open(anno_path, 'r', encoding='utf-8') as f2:
dataset_text = ''
for line in f2:
line = line.strip()
dataset_text += line
eval_dataset = json.loads(dataset_text)
categories = eval_dataset['categories']
for clsid, cate_dic in enumerate(categories):
catid = cate_dic['id']
cname = cate_dic['name']
_catid2clsid[catid] = clsid
_clsid2catid[clsid] = catid
_clsid2cname[clsid] = cname
class_names = []
num_classes = len(_clsid2cname.keys())
for clsid in range(num_classes):
class_names.append(_clsid2cname[clsid])
return _catid2clsid, _clsid2catid, _clsid2cname, class_names
class PPYOLO_COCOEvalDataset(torch.utils.data.Dataset):
def __init__(self, data_dir, json_file, ann_folder, name, cfg, transforms):
self.data_dir = data_dir
self.json_file = json_file
self.ann_folder = ann_folder
self.name = name
# 验证集
val_path = os.path.join(self.data_dir, self.ann_folder, self.json_file)
val_pre_path = os.path.join(self.data_dir, self.name)
# 种类id
_catid2clsid, _clsid2catid, _clsid2cname, class_names = get_class_msg(val_path)
val_dataset = COCO(val_path)
val_img_ids = val_dataset.getImgIds()
keep_img_ids = [] # 只跑有gt的图片,跟随PaddleDetection
for img_id in val_img_ids:
ins_anno_ids = val_dataset.getAnnIds(imgIds=img_id, iscrowd=False) # 读取这张图片所有标注anno的id
if len(ins_anno_ids) == 0:
continue
keep_img_ids.append(img_id)
val_img_ids = keep_img_ids
val_records = data_clean(val_dataset, val_img_ids, _catid2clsid, val_pre_path, 'val')
self.coco = val_dataset
self.records = val_records
self.context = cfg.context
self.transforms = transforms
self.catid2clsid = _catid2clsid
self.clsid2catid = _clsid2catid
self.num_record = len(val_records)
self.indexes = [i for i in range(self.num_record)]
def __len__(self):
return len(self.indexes)
def __getitem__(self, idx):
img_idx = self.indexes[idx]
sample = copy.deepcopy(self.records[img_idx])
# transforms
for transform in self.transforms:
sample = transform(sample, self.context)
# 取出感兴趣的项
pimage = sample['image']
im_size = np.array([sample['h'], sample['w']]).astype(np.float32)
id = sample['im_id']
return pimage, im_size, id
class FCOS_COCOEvalDataset(torch.utils.data.Dataset):
def __init__(self, data_dir, json_file, ann_folder, name, cfg, sample_transforms):
self.data_dir = data_dir
self.json_file = json_file
self.ann_folder = ann_folder
self.name = name
# 验证集
val_path = os.path.join(self.data_dir, self.ann_folder, self.json_file)
val_pre_path = os.path.join(self.data_dir, self.name)
# 种类id
_catid2clsid, _clsid2catid, _clsid2cname, class_names = get_class_msg(val_path)
val_dataset = COCO(val_path)
val_img_ids = val_dataset.getImgIds()
keep_img_ids = [] # 只跑有gt的图片,跟随PaddleDetection
for img_id in val_img_ids:
ins_anno_ids = val_dataset.getAnnIds(imgIds=img_id, iscrowd=False) # 读取这张图片所有标注anno的id
if len(ins_anno_ids) == 0:
continue
keep_img_ids.append(img_id)
val_img_ids = keep_img_ids
val_records = data_clean(val_dataset, val_img_ids, _catid2clsid, val_pre_path, 'val')
self.coco = val_dataset
self.records = val_records
self.context = cfg.context
self.sample_transforms = sample_transforms
self.catid2clsid = _catid2clsid
self.clsid2catid = _clsid2catid
self.num_record = len(val_records)
self.indexes = [i for i in range(self.num_record)]
def __len__(self):
return len(self.indexes)
def __getitem__(self, idx):
img_idx = self.indexes[idx]
sample = copy.deepcopy(self.records[img_idx])
# sample_transforms
for sample_transform in self.sample_transforms:
sample = sample_transform(sample, self.context)
# 取出感兴趣的项
pimage = sample['image']
im_info = sample['im_info']
im_id = sample['im_id']
return pimage, im_info, im_id
class PPYOLO_COCOTrainDataset(torch.utils.data.Dataset):
def __init__(self, data_dir, json_file, ann_folder, name, cfg, sample_transforms, batch_size):
self.data_dir = data_dir
self.json_file = json_file
self.ann_folder = ann_folder
self.name = name
# 训练集
train_path = os.path.join(self.data_dir, self.ann_folder, self.json_file)
train_pre_path = os.path.join(self.data_dir, self.name)
# 种类id
_catid2clsid, _clsid2catid, _clsid2cname, class_names = get_class_msg(train_path)
train_dataset = COCO(train_path)
train_img_ids = train_dataset.getImgIds()
train_records = data_clean(train_dataset, train_img_ids, _catid2clsid, train_pre_path, 'train')
self.coco = train_dataset
self.records = train_records
self.context = cfg.context
self.sample_transforms = sample_transforms
self.catid2clsid = _catid2clsid
self.clsid2catid = _clsid2catid
self.num_record = len(train_records)
self.with_mixup = cfg.decodeImage.get('with_mixup', False)
self.with_cutmix = cfg.decodeImage.get('with_cutmix', False)
self.with_mosaic = cfg.decodeImage.get('with_mosaic', False)
self.batch_size = batch_size
# 一轮的步数。丢弃最后几个样本。
self.train_steps = self.num_record // batch_size
# mixup、cutmix、mosaic数据增强的轮数
self.aug_epochs = cfg.aug_epochs
# 训练样本
self.indexes_ori = [i for i in range(self.num_record)]
self.indexes = copy.deepcopy(self.indexes_ori)
# 每个epoch之前洗乱
np.random.shuffle(self.indexes)
self.indexes = self.indexes[:self.train_steps * self.batch_size]
self._len = len(self.indexes)
# 多尺度训练
self.sizes = cfg.randomShape['sizes']
self.shapes = []
while len(self.shapes) < self.train_steps:
shape = np.random.choice(self.sizes)
self.shapes.append(shape)
# 输出特征图数量
self.n_layers = len(cfg.head['downsample'])
self._epoch = 0
def __len__(self):
return self._len
def set_epoch(self, epoch_id):
self._epoch = epoch_id
# 多尺度训练
self.shapes = []
while len(self.shapes) < self.train_steps:
shape = np.random.choice(self.sizes)
self.shapes.append(shape)
self.indexes = copy.deepcopy(self.indexes_ori)
# 每个epoch之前洗乱
np.random.shuffle(self.indexes)
self.indexes = self.indexes[:self._len]
def __getitem__(self, idx):
iter_id = idx // self.batch_size
img_idx = self.indexes[idx]
shape = self.shapes[iter_id]
sample = copy.deepcopy(self.records[img_idx])
sample["curr_iter"] = iter_id
# 为mixup数据增强做准备
if self.with_mixup and self._epoch <= self.aug_epochs:
num = len(self.records)
mix_idx = np.random.randint(0, num)
while mix_idx == img_idx: # 为了不选到自己
mix_idx = np.random.randint(0, num)
sample['mixup'] = copy.deepcopy(self.records[mix_idx])
sample['mixup']["curr_iter"] = iter_id
# 为cutmix数据增强做准备
if self.with_cutmix and self._epoch <= self.aug_epochs:
num = len(self.records)
mix_idx = np.random.randint(0, num)
while mix_idx == img_idx: # 为了不选到自己
mix_idx = np.random.randint(0, num)
sample['cutmix'] = copy.deepcopy(self.records[mix_idx])
sample['cutmix']["curr_iter"] = iter_id
# 为mosaic数据增强做准备
if self.with_mosaic and self._epoch <= self.aug_epochs:
num = len(self.records)
mix_idx =
|
np.random.randint(0, num)
|
numpy.random.randint
|
# Licensed under a 3-clause BSD style license - see LICENSE.rst
# coding=utf-8
"""
Classes and utilities for operating the wavefront sensors of the MMTO and analyzing the data they produce
"""
import warnings
import pathlib
import numpy as np
import photutils
import matplotlib.pyplot as plt
import matplotlib.cm as cm
from skimage import feature
from scipy import ndimage, optimize
from scipy.ndimage import rotate
from scipy.spatial import cKDTree
import lmfit
import astropy.units as u
from astropy.io import fits
from astropy.io import ascii
from astropy import stats, visualization, timeseries
from astropy.modeling.models import Gaussian2D, Polynomial2D
from astropy.modeling.fitting import LevMarLSQFitter
from astropy.table import conf as table_conf
from astroscrappy import detect_cosmics
from ccdproc.utils.slices import slice_from_string
from .config import recursive_subclasses, merge_config, mmtwfs_config
from .telescope import TelescopeFactory
from .f9topbox import CompMirror
from .zernike import ZernikeVector, zernike_slopes, cart2pol, pol2cart
from .custom_exceptions import WFSConfigException, WFSAnalysisFailed, WFSCommandException
import logging
import logging.handlers
log = logging.getLogger("WFS")
log.setLevel(logging.INFO)
warnings.simplefilter(action="ignore", category=FutureWarning)
table_conf.replace_warnings = ['attributes']
__all__ = ['SH_Reference', 'WFS', 'F9', 'NewF9', 'F5', 'Binospec', 'MMIRS', 'WFSFactory', 'wfs_norm', 'check_wfsdata',
'wfsfind', 'grid_spacing', 'center_pupil', 'get_apertures', 'match_apertures', 'aperture_distance', 'fit_apertures',
'get_slopes', 'make_init_pars', 'slope_diff', 'mk_wfs_mask']
def wfs_norm(data, interval=visualization.ZScaleInterval(contrast=0.05), stretch=visualization.LinearStretch()):
"""
Define default image normalization to use for WFS images
"""
norm = visualization.mpl_normalize.ImageNormalize(
data,
interval=interval,
stretch=stretch
)
return norm
def check_wfsdata(data, header=False):
"""
Utility to validate WFS data
Parameters
----------
data : FITS filename or 2D ndarray
WFS image
Returns
-------
data : 2D np.ndarray
Validated 2D WFS image
"""
hdr = None
if isinstance(data, (str, pathlib.PosixPath)):
# we're a fits file (hopefully)
try:
with fits.open(data, ignore_missing_simple=True) as h:
data = h[-1].data # binospec images put the image data into separate extension so always grab last available.
if header:
hdr = h[-1].header
except Exception as e:
msg = "Error reading FITS file, %s (%s)" % (data, repr(e))
raise WFSConfigException(value=msg)
if not isinstance(data, np.ndarray):
msg = "WFS image data in improper format, %s" % type(data)
raise WFSConfigException(value=msg)
if len(data.shape) != 2:
msg = "WFS image data has improper shape, %dD. Must be 2D image." % len(data.shape)
raise WFSConfigException(value=msg)
if header and hdr is not None:
return data, hdr
else:
return data
def mk_wfs_mask(data, thresh_factor=50., outfile="wfs_mask.fits"):
"""
Take a WFS image and mask/scale it so that it can be used as a reference for pupil centering
Parameters
----------
data : FITS filename or 2D ndarray
WFS image
thresh_factor : float (default: 50.)
Fraction of maximum value below which will be masked to 0.
outfile : string (default: wfs_mask.fits)
Output FITS file to write the resulting image to.
Returns
-------
scaled : 2D ndarray
Scaled and masked WFS image
"""
data = check_wfsdata(data)
mx = data.max()
thresh = mx / thresh_factor
data[data < thresh] = 0.
scaled = data / mx
if outfile is not None:
fits.writeto(outfile, scaled)
return scaled
def wfsfind(data, fwhm=7.0, threshold=5.0, plot=True, ap_radius=5.0, std=None):
"""
Use photutils.DAOStarFinder() to find and centroid spots in a Shack-Hartmann WFS image.
Parameters
----------
data : FITS filename or 2D ndarray
WFS image
fwhm : float (default: 5.)
FWHM in pixels of DAOfind convolution kernel
threshold : float
DAOfind threshold in units of the standard deviation of the image
plot: bool
Toggle plotting of the reference image and overlayed apertures
ap_radius : float
Radius of plotted apertures
"""
# data should be background subtracted first...
data = check_wfsdata(data)
if std is None:
mean, median, std = stats.sigma_clipped_stats(data, sigma=3.0, maxiters=5)
daofind = photutils.DAOStarFinder(fwhm=fwhm, threshold=threshold*std, sharphi=0.95)
sources = daofind(data)
if sources is None:
msg = "WFS spot detection failed or no spots detected."
raise WFSAnalysisFailed(value=msg)
# this may be redundant given the above check...
nsrcs = len(sources)
if nsrcs == 0:
msg = "No WFS spots detected."
raise WFSAnalysisFailed(value=msg)
# only keep spots more than 1/4 as bright as the max. need this for f/9 especially.
sources = sources[sources['flux'] > sources['flux'].max()/4.]
fig = None
if plot:
fig, ax = plt.subplots()
fig.set_label("WFSfind")
positions = list(zip(sources['xcentroid'], sources['ycentroid']))
apertures = photutils.CircularAperture(positions, r=ap_radius)
norm = wfs_norm(data)
ax.imshow(data, cmap='Greys', origin='lower', norm=norm, interpolation='None')
apertures.plot(color='red', lw=1.5, alpha=0.5, axes=ax)
return sources, fig
def grid_spacing(data, apertures):
"""
Measure the WFS grid spacing which changes with telescope focus.
Parameters
----------
data : WFS image (FITS or np.ndarray)
apertures : `~astropy.table.Table`
WFS aperture data to analyze
Returns
-------
xspacing, yspacing : float, float
Average grid spacing in X and Y axes
"""
data = check_wfsdata(data)
x = np.arange(data.shape[1])
y = np.arange(data.shape[0])
bx = np.arange(data.shape[1]+1)
by = np.arange(data.shape[0]+1)
# bin the spot positions along the axes and use Lomb-Scargle to measure the grid spacing in each direction
xsum = np.histogram(apertures['xcentroid'], bins=bx)
ysum = np.histogram(apertures['ycentroid'], bins=by)
k = np.linspace(10.0, 50., 1000) # look for spacings from 10 to 50 pixels (plenty of range, but not too small to alias)
f = 1.0 / k # convert spacing to frequency
xp = timeseries.LombScargle(x, xsum[0]).power(f)
yp = timeseries.LombScargle(y, ysum[0]).power(f)
# the peak of the power spectrum will coincide with the average spacing
xspacing = k[xp.argmax()]
yspacing = k[yp.argmax()]
return xspacing, yspacing
def center_pupil(input_data, pup_mask, threshold=0.8, sigma=10., plot=True):
"""
Find the center of the pupil in a WFS image using skimage.feature.match_template(). This generates
a correlation image and we centroid the peak of the correlation to determine the center.
Parameters
----------
data : str or 2D ndarray
WFS image to analyze, either FITS file or ndarray image data
pup_mask : str or 2D ndarray
Pupil model to use in the template matching
threshold : float (default: 0.0)
Sets image to 0 where it's below threshold * image.max()
sigma : float (default: 20.)
Sigma of gaussian smoothing kernel
plot : bool
Toggle plotting of the correlation image
Returns
-------
cen : tuple (float, float)
X and Y pixel coordinates of the pupil center
"""
data = np.copy(check_wfsdata(input_data))
pup_mask = check_wfsdata(pup_mask).astype(np.float64) # need to force float64 here to make scipy >= 1.4 happy...
# smooth the image to increae the S/N.
smo = ndimage.gaussian_filter(data, sigma)
# use skimage.feature.match_template() to do a fast cross-correlation between the WFS image and the pupil model.
# the location of the peak of the correlation will be the center of the WFS pattern.
match = feature.match_template(smo, pup_mask, pad_input=True)
find_thresh = threshold * match.max()
t = photutils.detection.find_peaks(match, find_thresh, box_size=5, centroid_func=photutils.centroids.centroid_com)
if t is None:
msg = "No valid pupil or spot pattern detected."
raise WFSAnalysisFailed(value=msg)
peak = t['peak_value'].max()
xps = []
yps = []
# if there are peaks that are very nearly correlated, average their positions
for p in t:
if p['peak_value'] >= 0.95*peak:
xps.append(p['x_centroid'])
yps.append(p['y_centroid'])
xp = np.mean(xps)
yp = np.mean(yps)
fig = None
if plot:
fig, ax = plt.subplots()
fig.set_label("Pupil Correlation Image (masked)")
ax.imshow(match, interpolation=None, cmap=cm.magma, origin='lower')
ax.scatter(xp, yp, marker="+", color="green")
return xp, yp, fig
def get_apertures(data, apsize, fwhm=5.0, thresh=7.0, plot=True, cen=None):
"""
Use wfsfind to locate and centroid spots. Measure their S/N ratios and the sigma of a 2D gaussian fit to
the co-added spot.
Parameters
----------
data : str or 2D ndarray
WFS image to analyze, either FITS file or ndarray image data
apsize : float
Diameter/width of the SH apertures
Returns
-------
srcs : astropy.table.Table
Detected WFS spot positions and properties
masks : list of photutils.ApertureMask objects
Masks used for aperture centroiding
snrs : 1D np.ndarray
S/N for each located spot
sigma : float
"""
data = check_wfsdata(data)
# set maxiters to None to let this clip all the way to convergence
if cen is None:
mean, median, stddev = stats.sigma_clipped_stats(data, sigma=3.0, maxiters=None)
else:
xcen, ycen = int(cen[0]), int(cen[1])
mean, median, stddev = stats.sigma_clipped_stats(data[ycen-50:ycen+50, xcen-50:ycen+50], sigma=3.0, maxiters=None)
# use wfsfind() and pass it the clipped stddev from here
with warnings.catch_warnings():
warnings.simplefilter("ignore")
srcs, wfsfind_fig = wfsfind(data, fwhm=fwhm, threshold=thresh, std=stddev, plot=plot)
# we use circular apertures here because they generate square masks of the appropriate size.
# rectangular apertures produced masks that were sqrt(2) too large.
# see https://github.com/astropy/photutils/issues/499 for details.
apers = photutils.CircularAperture(
list(zip(srcs['xcentroid'], srcs['ycentroid'])),
r=apsize/2.
)
masks = apers.to_mask(method='subpixel')
sigma = 0.0
snrs = []
if len(masks) >= 1:
spot = np.zeros(masks[0].shape)
for m in masks:
subim = m.cutout(data)
# make co-added spot image for use in calculating the seeing
if subim.shape == spot.shape:
spot += subim
signal = subim.sum()
noise = np.sqrt(stddev**2 * subim.shape[0] * subim.shape[1])
snr = signal / noise
snrs.append(snr)
snrs = np.array(snrs)
# set up 2D gaussian model plus constant background to fit to the coadded spot
with warnings.catch_warnings():
# ignore astropy warnings about issues with the fit...
warnings.simplefilter("ignore")
g2d = Gaussian2D(amplitude=spot.max(), x_mean=spot.shape[1]/2, y_mean=spot.shape[0]/2)
p2d = Polynomial2D(degree=0)
model = g2d + p2d
fitter = LevMarLSQFitter()
y, x = np.mgrid[:spot.shape[0], :spot.shape[1]]
fit = fitter(model, x, y, spot)
sigma = 0.5 * (fit.x_stddev_0.value + fit.y_stddev_0.value)
return srcs, masks, snrs, sigma, wfsfind_fig
def match_apertures(refx, refy, spotx, spoty, max_dist=25.):
"""
Given reference aperture and spot X/Y positions, loop through reference apertures and find closest spot. Use
max_dist to exclude matches that are too far from reference position. Return masks to use to denote validly
matched apertures.
"""
refs = np.array([refx, refy])
spots = np.array([spotx, spoty])
match = np.nan * np.ones(len(refx))
matched = []
for i in np.arange(len(refx)):
dists = np.sqrt((spots[0]-refs[0][i])**2 + (spots[1]-refs[1][i])**2)
min_i = np.argmin(dists)
if np.min(dists) < max_dist:
if min_i not in matched:
match[i] = min_i
matched.append(min_i)
else:
if min_i not in matched:
match[i] = np.nan
ref_mask = ~np.isnan(match)
src_mask = match[ref_mask]
return ref_mask, src_mask.astype(int)
def aperture_distance(refx, refy, spotx, spoty):
"""
Calculate the sum of the distances between each reference aperture and the closest measured spot position.
This total distance is the statistic to minimize when fitting the reference aperture grid to the data.
"""
refs = np.array([refx, refy]).transpose()
spots = np.array([spotx, spoty]).transpose()
tree = cKDTree(refs)
mindist, _ = tree.query(spots)
tot_dist = mindist.sum()
return np.log(tot_dist)
def fit_apertures(pars, ref, spots):
"""
Scale the reference positions by the fit parameters and calculate the total distance between the matches.
The parameters of the fit are:
``xc, yc = center positions``
``scale = magnification of the grid (focus)``
``xcoma, ycoma = linear change in magnification as a function of x/y (coma)``
'ref' and 'spots' are assumed to be dict-like and must have the keys 'xcentroid' and 'ycentroid'.
Parameters
----------
pars : list-like
The fit parameters passed in as a 5 element list: (xc, yc, scale, xcoma, ycoma)
ref : dict-like
Dict containing ``xcentroid`` and ``ycentroid`` keys that contain the reference X and Y
positions of the apertures.
spots : dict-like
Dict containing ``xcentroid`` and ``ycentroid`` keys that contain the measured X and Y
positions of the apertures.
Returns
-------
dist : float
The cumulative distance between the matched reference and measured aperture positions.
"""
xc = pars[0]
yc = pars[1]
scale = pars[2]
xcoma = pars[3]
ycoma = pars[4]
refx = ref['xcentroid'] * (scale + ref['xcentroid'] * xcoma) + xc
refy = ref['ycentroid'] * (scale + ref['ycentroid'] * ycoma) + yc
spotx = spots['xcentroid']
spoty = spots['ycentroid']
dist = aperture_distance(refx, refy, spotx, spoty)
return dist
def get_slopes(data, ref, pup_mask, fwhm=7., thresh=5., cen=[255, 255],
cen_thresh=0.8, cen_sigma=10., cen_tol=50., spot_snr_thresh=3.0, plot=True):
"""
Analyze a WFS image and produce pixel offsets between reference and observed spot positions.
Parameters
----------
data : str or 2D np.ndarray
FITS file or np.ndarray containing WFS observation
ref : `~astropy.table.Table`
Table of reference apertures
pup_mask : str or 2D np.ndarray
FITS file or np.ndarray containing mask used to register WFS spot pattern via cross-correlation
fwhm : float (default: 7.0)
FWHM of convolution kernel applied to image by the spot finding algorithm
thresh : float (default: 5.0)
Number of sigma above background for a spot to be considered detected
cen : list-like with 2 elements (default: [255, 255])
Expected position of the center of the WFS spot pattern in form [X_cen, Y_cen]
cen_thresh : float (default: 0.8)
Masking threshold as fraction of peak value used in `~photutils.detection.find_peaks`
cen_sigma : float (default: 10.0)
Width of gaussian filter applied to image by `~mmtwfs.wfs.center_pupil`
cen_tol : float (default: 50.0)
Tolerance for difference between expected and measureed pupil center
spot_snr_thresh : float (default: 3.0)
S/N tolerance for a WFS spot to be considered valid for analysis
plot : bool
Toggle plotting of image with aperture overlays
Returns
-------
results : dict
Results of the wavefront slopes measurement packaged into a dict with the following keys:
slopes - mask np.ndarry containing the slope values in pixel units
pup_coords - pupil coordinates for the position for each slope value
spots - `~astropy.table.Table` as returned by photutils star finder routines
src_aps - `~photutils.aperture.CircularAperture` for each detected spot
spacing - list-like of form (xspacing, yspacing) containing the mean spacing between rows and columns of spots
center - list-like of form (xcen, ycen) containing the center of the spot pattern
ref_mask - np.ndarray of matched spots in reference image
src_mask - np.ndarray of matched spots in the data image
spot_sigma - sigma of a gaussian fit to a co-addition of detected spots
figures - dict of figures that are optionally produced
grid_fit - dict of best-fit parameters of grid fit used to do fine registration between source and reference spots
"""
data = check_wfsdata(data)
pup_mask = check_wfsdata(pup_mask)
if ref.pup_outer is None:
raise WFSConfigException("No pupil information applied to SH reference.")
pup_outer = ref.pup_outer
pup_inner = ref.pup_inner
# input data should be background subtracted for best results. this initial guess of the center positions
# will be good enough to get the central obscuration, but will need to be fine-tuned for aperture association.
xcen, ycen, pupcen_fig = center_pupil(data, pup_mask, threshold=cen_thresh, sigma=cen_sigma, plot=plot)
if np.hypot(xcen-cen[0], ycen-cen[1]) > cen_tol:
msg = f"Measured pupil center [{round(xcen)}, {round(ycen)}] more than {cen_tol} pixels from {cen}."
raise WFSAnalysisFailed(value=msg)
# using the mean spacing is straightforward for square apertures and a reasonable underestimate for hexagonal ones
ref_spacing = np.mean([ref.xspacing, ref.yspacing])
apsize = ref_spacing
srcs, masks, snrs, sigma, wfsfind_fig = get_apertures(data, apsize, fwhm=fwhm, thresh=thresh, cen=(xcen, ycen))
# ignore low S/N spots
srcs = srcs[snrs > spot_snr_thresh]
# get grid spacing of the data
xspacing, yspacing = grid_spacing(data, srcs)
# find the scale difference between data and ref and use as init
init_scale = (xspacing/ref.xspacing + yspacing/ref.yspacing) / 2.
# apply masking to detected sources to avoid partially illuminated apertures at the edges
srcs['dist'] = np.sqrt((srcs['xcentroid'] - xcen)**2 + (srcs['ycentroid'] - ycen)**2)
srcs = srcs[(srcs['dist'] > pup_inner*init_scale) & (srcs['dist'] < pup_outer*init_scale)]
# if we don't detect spots in at least half of the reference apertures, we can't usually get a good wavefront measurement
if len(srcs) < 0.5 * len(ref.masked_apertures['xcentroid']):
msg = "Only %d spots detected out of %d apertures." % (len(srcs), len(ref.masked_apertures['xcentroid']))
raise WFSAnalysisFailed(value=msg)
src_aps = photutils.CircularAperture(
list(zip(srcs['xcentroid'], srcs['ycentroid'])),
r=apsize/2.
)
# set up to do a fit of the reference apertures to the spot positions with the center, scaling, and position-dependent
# scaling (coma) as free parameters
args = (ref.masked_apertures, srcs)
par_keys = ('xcen', 'ycen', 'scale', 'xcoma', 'ycoma')
pars = (xcen, ycen, init_scale, 0.0, 0.0)
coma_bound = 1e-4 # keep coma constrained by now since it can cause trouble
# scipy.optimize.minimize can do bounded minimization so leverage that to keep the solution within a reasonable range.
bounds = (
(xcen-15, xcen+15), # hopefully we're not too far off from true center...
(ycen-15, ycen+15),
(init_scale-0.05, init_scale+0.05), # reasonable range of expected focus difference...
(-coma_bound, coma_bound),
(-coma_bound, coma_bound)
)
try:
min_results = optimize.minimize(fit_apertures, pars, args=args, bounds=bounds, options={'ftol': 1e-13, 'gtol': 1e-7})
except Exception as e:
msg = f"Aperture grid matching failed: {e}"
raise WFSAnalysisFailed(value=msg)
fit_results = {}
for i, k in enumerate(par_keys):
fit_results[k] = min_results['x'][i]
# this is more reliably the center of the actual pupil image whereas fit_results shifts a bit depending on detected spots.
# the lenslet pattern can move around a bit on the pupil, but we need the center of the pupil to calculate their pupil
# coordinates.
pup_center = [xcen, ycen]
scale = fit_results['scale']
xcoma, ycoma = fit_results['xcoma'], fit_results['ycoma']
refx = ref.masked_apertures['xcentroid'] * (scale + ref.masked_apertures['xcentroid'] * xcoma) + fit_results['xcen']
refy = ref.masked_apertures['ycentroid'] * (scale + ref.masked_apertures['ycentroid'] * ycoma) + fit_results['ycen']
xspacing = scale * ref.xspacing
yspacing = scale * ref.yspacing
# coarse match reference apertures to spots
spacing = np.max([xspacing, yspacing])
ref_mask, src_mask = match_apertures(refx, refy, srcs['xcentroid'], srcs['ycentroid'], max_dist=spacing/2.)
# these are unscaled so that the slope includes defocus
trim_refx = ref.masked_apertures['xcentroid'][ref_mask] + fit_results['xcen']
trim_refy = ref.masked_apertures['ycentroid'][ref_mask] + fit_results['ycen']
ref_aps = photutils.CircularAperture(
list(zip(trim_refx, trim_refy)),
r=ref_spacing/2.
)
slope_x = srcs['xcentroid'][src_mask] - trim_refx
slope_y = srcs['ycentroid'][src_mask] - trim_refy
pup_coords = (ref_aps.positions - pup_center) / [pup_outer, pup_outer]
aps_fig = None
if plot:
norm = wfs_norm(data)
aps_fig, ax = plt.subplots()
aps_fig.set_label("Aperture Positions")
ax.imshow(data, cmap='Greys', origin='lower', norm=norm, interpolation='None')
ax.scatter(pup_center[0], pup_center[1])
src_aps.plot(color='blue', axes=ax)
# need full slopes array the size of the complete set of reference apertures and pre-filled with np.nan for masking
slopes = np.nan * np.ones((2, len(ref.masked_apertures['xcentroid'])))
slopes[0][ref_mask] = slope_x
slopes[1][ref_mask] = slope_y
figures = {}
figures['pupil_center'] = pupcen_fig
figures['slopes'] = aps_fig
results = {
"slopes": np.ma.masked_invalid(slopes),
"pup_coords": pup_coords.transpose(),
"spots": srcs,
"src_aps": src_aps,
"spacing": (xspacing, yspacing),
"center": pup_center,
"ref_mask": ref_mask,
"src_mask": src_mask,
"spot_sigma": sigma,
"figures": figures,
"grid_fit": fit_results
}
return results
def make_init_pars(nmodes=21, modestart=2, init_zv=None):
"""
Make a set of initial parameters that can be used with `~lmfit.minimize` to make a wavefront fit with
parameter names that are compatible with ZernikeVectors.
Parameters
----------
nmodes: int (default: 21)
Number of Zernike modes to fit.
modestart: int (default: 2)
First Zernike mode to be used.
init_zv: ZernikeVector (default: None)
ZernikeVector containing initial values for the fit.
Returns
-------
params: `~lmfit.Parameters` instance
Initial parameters in form that can be passed to `~lmfit.minimize`.
"""
pars = []
for i in range(modestart, modestart+nmodes, 1):
key = "Z{:02d}".format(i)
if init_zv is not None:
val = init_zv[key].value
if val < 2. * np.finfo(float).eps:
val = 0.0
else:
val = 0.0
zpar = (key, val)
pars.append(zpar)
params = lmfit.Parameters()
params.add_many(*pars)
return params
def slope_diff(pars, coords, slopes, norm=False):
"""
For a given set of wavefront fit parameters, calculate the "distance" between the predicted and measured wavefront
slopes. This function is used by `~lmfit.minimize` which expects the sqrt to be applied rather than a chi-squared,
"""
parsdict = pars.valuesdict()
rho, phi = cart2pol(coords)
xslope = slopes[0]
yslope = slopes[1]
pred_xslope, pred_yslope = zernike_slopes(parsdict, rho, phi, norm=norm)
dist = np.sqrt((xslope - pred_xslope)**2 + (yslope - pred_yslope)**2)
return dist
class SH_Reference(object):
"""
Class to handle Shack-Hartmann reference data
"""
def __init__(self, data, fwhm=4.5, threshold=20.0, plot=True):
"""
Read WFS reference image and generate reference magnifications (i.e. grid spacing) and
aperture positions.
Parameters
----------
data : FITS filename or 2D ndarray
WFS reference image
fwhm : float
FWHM in pixels of DAOfind convolution kernel
threshold : float
DAOfind threshold in units of the standard deviation of the image
plot : bool
Toggle plotting of the reference image and overlayed apertures
"""
self.data = check_wfsdata(data)
data = data - np.median(data)
self.apertures, self.figure = wfsfind(data, fwhm=fwhm, threshold=threshold, plot=plot)
if plot:
self.figure.set_label("Reference Image")
self.xcen = self.apertures['xcentroid'].mean()
self.ycen = self.apertures['ycentroid'].mean()
self.xspacing, self.yspacing = grid_spacing(data, self.apertures)
# make masks for each reference spot and fit a 2D gaussian to get its FWHM. the reference FWHM is subtracted in
# quadrature from the observed FWHM when calculating the seeing.
apsize = np.mean([self.xspacing, self.yspacing])
apers = photutils.CircularAperture(
list(zip(self.apertures['xcentroid'], self.apertures['ycentroid'])),
r=apsize/2.
)
masks = apers.to_mask(method='subpixel')
self.photapers = apers
self.spot = np.zeros(masks[0].shape)
for m in masks:
subim = m.cutout(data)
# make co-added spot image for use in calculating the seeing
if subim.shape == self.spot.shape:
self.spot += subim
self.apertures['xcentroid'] -= self.xcen
self.apertures['ycentroid'] -= self.ycen
self.apertures['dist'] = np.sqrt(self.apertures['xcentroid']**2 + self.apertures['ycentroid']**2)
self.masked_apertures = self.apertures
self.pup_inner = None
self.pup_outer = None
def adjust_center(self, x, y):
"""
Adjust reference center to new x, y position.
"""
self.apertures['xcentroid'] += self.xcen
self.apertures['ycentroid'] += self.ycen
self.apertures['xcentroid'] -= x
self.apertures['ycentroid'] -= y
self.apertures['dist'] = np.sqrt(self.apertures['xcentroid']**2 + self.apertures['ycentroid']**2)
self.xcen = x
self.ycen = y
self.apply_pupil(self.pup_inner, self.pup_outer)
def apply_pupil(self, pup_inner, pup_outer):
"""
Apply a pupil mask to the reference apertures
"""
if pup_inner is not None and pup_outer is not None:
self.masked_apertures = self.apertures[(self.apertures['dist'] > pup_inner) & (self.apertures['dist'] < pup_outer)]
self.pup_inner = pup_inner
self.pup_outer = pup_outer
def pup_coords(self, pup_outer):
"""
Take outer radius of pupil and calculate pupil coordinates for the masked apertures
"""
coords = (self.masked_apertures['xcentroid']/pup_outer, self.masked_apertures['ycentroid']/pup_outer)
return coords
def WFSFactory(wfs="f5", config={}, **kwargs):
"""
Build and return proper WFS sub-class instance based on the value of 'wfs'.
"""
config = merge_config(config, dict(**kwargs))
wfs = wfs.lower()
types = recursive_subclasses(WFS)
wfses = [t.__name__.lower() for t in types]
wfs_map = dict(list(zip(wfses, types)))
if wfs not in wfses:
raise WFSConfigException(value="Specified WFS, %s, not valid or not implemented." % wfs)
if 'plot' in config:
plot = config['plot']
else:
plot = True
wfs_cls = wfs_map[wfs](config=config, plot=plot)
return wfs_cls
class WFS(object):
"""
Defines configuration pattern and methods common to all WFS systems
"""
def __init__(self, config={}, plot=True, **kwargs):
key = self.__class__.__name__.lower()
self.__dict__.update(merge_config(mmtwfs_config['wfs'][key], config))
self.telescope = TelescopeFactory(telescope=self.telescope, secondary=self.secondary)
self.secondary = self.telescope.secondary
self.plot = plot
self.connected = False
self.ref_fwhm = self.ref_spot_fwhm()
# this factor calibrates spot motion in pixels to nm of wavefront error
self.tiltfactor = self.telescope.nmperasec * (self.pix_size.to(u.arcsec).value)
# if this is the same for all modes, load it once here
if hasattr(self, "reference_file"):
refdata, hdr = check_wfsdata(self.reference_file, header=True)
refdata = self.trim_overscan(refdata, hdr)
reference = SH_Reference(refdata, plot=self.plot)
# now assign 'reference' for each mode so that it can be accessed consistently in all cases
for mode in self.modes:
if 'reference_file' in self.modes[mode]:
refdata, hdr = check_wfsdata(self.modes[mode]['reference_file'], header=True)
refdata = self.trim_overscan(refdata, hdr)
self.modes[mode]['reference'] = SH_Reference(
refdata,
plot=self.plot
)
else:
self.modes[mode]['reference'] = reference
def ref_spot_fwhm(self):
"""
Calculate the Airy FWHM in pixels of a perfect WFS spot from the optical prescription and detector pixel size
"""
theta_fwhm = 1.028 * self.eff_wave / self.lenslet_pitch
det_fwhm = np.arctan(theta_fwhm).value * self.lenslet_fl
det_fwhm_pix = det_fwhm.to(u.um).value / self.pix_um.to(u.um).value
return det_fwhm_pix
def get_flipud(self, mode=None):
"""
Determine if the WFS image needs to be flipped up/down
"""
return False
def get_fliplr(self, mode=None):
"""
Determine if the WFS image needs to be flipped left/right
"""
return False
def ref_pupil_location(self, mode, hdr=None):
"""
Get the center of the pupil on the reference image
"""
ref = self.modes[mode]['reference']
x = ref.xcen
y = ref.ycen
return x, y
def seeing(self, mode, sigma, airmass=None):
"""
Given a sigma derived from a gaussian fit to a WFS spot, deconvolve the systematic width from the reference image
and relate the remainder to r_0 and thus a seeing FWHM.
"""
# the effective wavelength of the WFS imagers is about 600-700 nm. mmirs and the oldf9 system use blue-blocking filters
wave = self.eff_wave
wave = wave.to(u.m).value # r_0 equation expects meters so convert
refwave = 500 * u.nm # standard wavelength that seeing values are referenced to
refwave = refwave.to(u.m).value
# calculate the physical size of each aperture.
ref = self.modes[mode]['reference']
apsize_pix = np.max((ref.xspacing, ref.yspacing))
d = self.telescope.diameter * apsize_pix / self.pup_size
d = d.to(u.m).value # r_0 equation expects meters so convert
# we need to deconvolve the instrumental spot width from the measured one to get the portion of the width that
# is due to spot motion
ref_sigma = stats.funcs.gaussian_fwhm_to_sigma * self.ref_fwhm
if sigma > ref_sigma:
corr_sigma = np.sqrt(sigma**2 - ref_sigma**2)
else:
return 0.0 * u.arcsec, 0.0 * u.arcsec
corr_sigma *= self.pix_size.to(u.rad).value # r_0 equation expects radians so convert
# this equation relates the motion within a single aperture to the characteristic scale size of the
# turbulence, r_0.
r_0 = (0.179 * (wave**2) * (d**(-1/3))/corr_sigma**2)**0.6
# this equation relates the turbulence scale size to an expected image FWHM at the given wavelength.
raw_seeing = u.Quantity(u.rad * 0.98 * wave / r_0, u.arcsec)
# seeing scales as lambda^-1/5 so calculate factor to scale to reference lambda
wave_corr = refwave**-0.2 / wave**-0.2
raw_seeing *= wave_corr
# correct seeing to zenith
if airmass is not None:
seeing = raw_seeing / airmass**0.6
else:
seeing = raw_seeing
return seeing, raw_seeing
def pupil_mask(self, hdr=None):
"""
Load and return the WFS spot mask used to locate and register the pupil
"""
pup_mask = check_wfsdata(self.wfs_mask)
return pup_mask
def reference_aberrations(self, mode, **kwargs):
"""
Create reference ZernikeVector for 'mode'.
"""
z = ZernikeVector(**self.modes[mode]['ref_zern'])
return z
def get_mode(self, hdr):
"""
If mode is not specified, either set it to the default mode or figure out the mode from the header.
"""
mode = self.default_mode
return mode
def process_image(self, fitsfile):
"""
Process the image to make it suitable for accurate wavefront analysis. Steps include nuking cosmic rays,
subtracting background, handling overscan regions, etc.
"""
rawdata, hdr = check_wfsdata(fitsfile, header=True)
trimdata = self.trim_overscan(rawdata, hdr=hdr)
# MMIRS gets a lot of hot pixels/CRs so make a quick pass to nuke them
cr_mask, data = detect_cosmics(trimdata, sigclip=5., niter=5, cleantype='medmask', psffwhm=5.)
# calculate the background and subtract it
bkg_estimator = photutils.ModeEstimatorBackground()
mask = photutils.make_source_mask(data, nsigma=2, npixels=5, dilate_size=11)
bkg = photutils.Background2D(data, (10, 10), filter_size=(5, 5), bkg_estimator=bkg_estimator, mask=mask)
data -= bkg.background
return data, hdr
def trim_overscan(self, data, hdr=None):
"""
Use the DATASEC in the header to determine the region to trim out. If no header provided or if the header
doesn't contain DATASEC, return data unchanged.
"""
if hdr is None:
return data
if 'DATASEC' not in hdr:
# if no DATASEC in header, punt and return unchanged
return data
datasec = slice_from_string(hdr['DATASEC'], fits_convention=True)
return data[datasec]
def measure_slopes(self, fitsfile, mode=None, plot=True, flipud=False, fliplr=False):
"""
Take a WFS image in FITS format, perform background subtration, pupil centration, and then use get_slopes()
to perform the aperture placement and spot centroiding.
"""
data, hdr = self.process_image(fitsfile)
plot = plot and self.plot
# flip data up/down if we need to. only binospec needs to currently.
if flipud or self.get_flipud(mode=mode):
data = np.flipud(data)
# flip left/right if we need to. no mode currently does, but who knows what the future holds.
if fliplr or self.get_fliplr(mode=mode):
data = np.fliplr(data)
if mode is None:
mode = self.get_mode(hdr)
if mode not in self.modes:
msg = "Invalid mode, %s, for WFS system, %s." % (mode, self.__class__.__name__)
raise WFSConfigException(value=msg)
# if available, get the rotator angle out of the header
if 'ROT' in hdr:
rotator = hdr['ROT'] * u.deg
else:
rotator = 0.0 * u.deg
# if there's a ROTOFF in the image header, grab it and adjust the rotator angle accordingly
if 'ROTOFF' in hdr:
rotator -= hdr['ROTOFF'] * u.deg
# make mask for finding wfs spot pattern
pup_mask = self.pupil_mask(hdr=hdr)
# get adjusted reference center position and update the reference
xcen, ycen = self.ref_pupil_location(mode, hdr=hdr)
self.modes[mode]['reference'].adjust_center(xcen, ycen)
# apply pupil to the reference
self.modes[mode]['reference'].apply_pupil(self.pup_inner, self.pup_size/2.)
ref_zv = self.reference_aberrations(mode, hdr=hdr)
zref = ref_zv.array
if len(zref) < self.nzern:
pad = np.zeros(self.nzern - len(zref))
zref = np.hstack((zref, pad))
try:
slope_results = get_slopes(
data,
self.modes[mode]['reference'],
pup_mask,
fwhm=self.find_fwhm,
thresh=self.find_thresh,
cen=self.cor_coords,
cen_thresh=self.cen_thresh,
cen_sigma=self.cen_sigma,
cen_tol=self.cen_tol,
plot=plot
)
slopes = slope_results['slopes']
coords = slope_results['pup_coords']
ref_pup_coords = self.modes[mode]['reference'].pup_coords(self.pup_size/2.)
rho, phi = cart2pol(ref_pup_coords)
ref_slopes = -(1. / self.tiltfactor) * np.array(zernike_slopes(ref_zv, rho, phi))
aps = slope_results['src_aps']
ref_mask = slope_results['ref_mask']
src_mask = slope_results['src_mask']
figures = slope_results['figures']
except WFSAnalysisFailed as e:
log.warning(f"Wavefront slope measurement failed: {e}")
slope_fig = None
if plot:
slope_fig, ax = plt.subplots()
slope_fig.set_label("WFS Image")
norm = wfs_norm(data)
ax.imshow(data, cmap='Greys', origin='lower', norm=norm, interpolation='None')
results = {}
results['slopes'] = None
results['figures'] = {}
results['mode'] = mode
results['figures']['slopes'] = slope_fig
return results
except Exception as e:
raise WFSAnalysisFailed(value=str(e))
# use the average width of the spots to estimate the seeing and use the airmass to extrapolate to zenith seeing
if 'AIRMASS' in hdr:
airmass = hdr['AIRMASS']
else:
airmass = None
seeing, raw_seeing = self.seeing(mode=mode, sigma=slope_results['spot_sigma'], airmass=airmass)
if plot:
sub_slopes = slopes - ref_slopes
x = aps.positions.transpose()[0][src_mask]
y = aps.positions.transpose()[1][src_mask]
uu = sub_slopes[0][ref_mask]
vv = sub_slopes[1][ref_mask]
norm = wfs_norm(data)
figures['slopes'].set_label("Aperture Positions and Spot Movement")
ax = figures['slopes'].axes[0]
ax.imshow(data, cmap='Greys', origin='lower', norm=norm, interpolation='None')
aps.plot(color='blue', axes=ax)
ax.quiver(x, y, uu, vv, scale_units='xy', scale=0.2, pivot='tip', color='red')
xl = [0.1*data.shape[1]]
yl = [0.95*data.shape[0]]
ul = [1.0/self.pix_size.value]
vl = [0.0]
ax.quiver(xl, yl, ul, vl, scale_units='xy', scale=0.2, pivot='tip', color='red')
ax.scatter([slope_results['center'][0]], [slope_results['center'][1]])
ax.text(0.12*data.shape[1], 0.95*data.shape[0], "1{0:unicode}".format(u.arcsec), verticalalignment='center')
ax.set_title("Seeing: %.2f\" (%.2f\" @ zenith)" % (raw_seeing.value, seeing.value))
results = {}
results['seeing'] = seeing
results['raw_seeing'] = raw_seeing
results['slopes'] = slopes
results['ref_slopes'] = ref_slopes
results['ref_zv'] = ref_zv
results['spots'] = slope_results['spots']
results['pup_coords'] = coords
results['ref_pup_coords'] = ref_pup_coords
results['apertures'] = aps
results['xspacing'] = slope_results['spacing'][0]
results['yspacing'] = slope_results['spacing'][1]
results['xcen'] = slope_results['center'][0]
results['ycen'] = slope_results['center'][1]
results['pup_mask'] = pup_mask
results['data'] = data
results['header'] = hdr
results['rotator'] = rotator
results['mode'] = mode
results['ref_mask'] = ref_mask
results['src_mask'] = src_mask
results['fwhm'] = stats.funcs.gaussian_sigma_to_fwhm * slope_results['spot_sigma']
results['figures'] = figures
results['grid_fit'] = slope_results['grid_fit']
return results
def fit_wavefront(self, slope_results, plot=True):
"""
Use results from self.measure_slopes() to fit a set of zernike polynomials to the wavefront shape.
"""
plot = plot and self.plot
if slope_results['slopes'] is not None:
results = {}
slopes = -self.tiltfactor * slope_results['slopes']
coords = slope_results['ref_pup_coords']
rho, phi = cart2pol(coords)
zref = slope_results['ref_zv']
params = make_init_pars(nmodes=self.nzern, init_zv=zref)
results['fit_report'] = lmfit.minimize(slope_diff, params, args=(coords, slopes))
zfit = ZernikeVector(coeffs=results['fit_report'])
results['raw_zernike'] = zfit
# derotate the zernike solution to match the primary mirror coordinate system
total_rotation = self.rotation - slope_results['rotator']
zv_rot = ZernikeVector(coeffs=results['fit_report'])
zv_rot.rotate(angle=-total_rotation)
results['rot_zernike'] = zv_rot
# subtract the reference aberrations
zsub = zv_rot - zref
results['ref_zernike'] = zref
results['zernike'] = zsub
pred_slopes = np.array(zernike_slopes(zfit, rho, phi))
diff = slopes - pred_slopes
diff_pix = diff / self.tiltfactor
rms = np.sqrt((diff[0]**2 + diff[1]**2).mean())
results['residual_rms_asec'] = rms / self.telescope.nmperasec * u.arcsec
results['residual_rms'] = rms * zsub.units
results['zernike_rms'] = zsub.rms
results['zernike_p2v'] = zsub.peak2valley
fig = None
if plot:
ref_mask = slope_results['ref_mask']
src_mask = slope_results['src_mask']
im = slope_results['data']
gnorm = wfs_norm(im)
fig, ax = plt.subplots()
fig.set_label("Zernike Fit Residuals")
ax.imshow(im, cmap='Greys', origin='lower', norm=gnorm, interpolation='None')
x = slope_results['apertures'].positions.transpose()[0][src_mask]
y = slope_results['apertures'].positions.transpose()[1][src_mask]
ax.quiver(x, y, diff_pix[0][ref_mask], diff_pix[1][ref_mask], scale_units='xy',
scale=0.05, pivot='tip', color='red')
xl = [0.1*im.shape[1]]
yl = [0.95*im.shape[0]]
ul = [0.2/self.pix_size.value]
vl = [0.0]
ax.quiver(xl, yl, ul, vl, scale_units='xy', scale=0.05, pivot='tip', color='red')
ax.text(0.12*im.shape[1], 0.95*im.shape[0], "0.2{0:unicode}".format(u.arcsec), verticalalignment='center')
ax.text(
0.95*im.shape[1],
0.95*im.shape[0],
"Residual RMS: {0.value:0.2f}{0.unit:unicode}".format(results['residual_rms_asec']),
verticalalignment='center',
horizontalalignment='right'
)
iq = np.sqrt(results['residual_rms_asec']**2 +
(results['zernike_rms'].value / self.telescope.nmperasec * u.arcsec)**2)
ax.set_title("Image Quality: {0.value:0.2f}{0.unit:unicode}".format(iq))
results['resid_plot'] = fig
else:
results = None
return results
def calculate_primary(self, zv, threshold=0.0 * u.nm, mask=[]):
"""
Calculate force corrections to primary mirror and any required focus offsets. Use threshold to determine which
terms in 'zv' to use in the force calculations. Any terms with normalized amplitude less than threshold will
not be used in the force calculation. In addition, individual terms can be forced to be masked.
"""
zv.denormalize()
zv_masked = ZernikeVector()
zv_norm = zv.copy()
zv_norm.normalize()
log.debug(f"thresh: {threshold} mask {mask}")
for z in zv:
if abs(zv_norm[z]) >= threshold:
zv_masked[z] = zv[z]
log.debug(f"{z}: Good")
else:
log.debug(f"{z}: Bad")
zv_masked.denormalize() # need to assure we're using fringe coeffs
log.debug(f"\nInput masked: {zv_masked}")
# use any available error bars to mask down to 1 sigma below amplitude or 0 if error bars are larger than amplitude.
for z in zv_masked:
frac_err = 1. - min(zv_masked.frac_error(key=z), 1.)
zv_masked[z] *= frac_err
log.debug(f"\nErrorbar masked: {zv_masked}")
forces, m1focus, zv_allmasked = self.telescope.calculate_primary_corrections(
zv=zv_masked,
mask=mask,
gain=self.m1_gain
)
log.debug(f"\nAll masked: {zv_allmasked}")
return forces, m1focus, zv_allmasked
def calculate_focus(self, zv):
"""
Convert Zernike defocus to um of secondary offset.
"""
z_denorm = zv.copy()
z_denorm.denormalize() # need to assure we're using fringe coeffs
frac_err = 1. - min(z_denorm.frac_error(key='Z04'), 1.)
foc_corr = -self.m2_gain * frac_err * z_denorm['Z04'] / self.secondary.focus_trans
return foc_corr.round(2)
def calculate_cc(self, zv):
"""
Convert Zernike coma (Z07 and Z08) into arcsec of secondary center-of-curvature tilts.
"""
z_denorm = zv.copy()
z_denorm.denormalize() # need to assure we're using fringe coeffs
# fix coma using tilts around the M2 center of curvature.
y_frac_err = 1. - min(z_denorm.frac_error(key='Z07'), 1.)
x_frac_err = 1. - min(z_denorm.frac_error(key='Z08'), 1.)
cc_y_corr = -self.m2_gain * y_frac_err * z_denorm['Z07'] / self.secondary.theta_cc
cc_x_corr = -self.m2_gain * x_frac_err * z_denorm['Z08'] / self.secondary.theta_cc
return cc_x_corr.round(3), cc_y_corr.round(3)
def calculate_recenter(self, fit_results, defoc=1.0):
"""
Perform zero-coma hexapod tilts to align the pupil center to the center-of-rotation.
The location of the CoR is configured to be at self.cor_coords.
"""
xc = fit_results['xcen']
yc = fit_results['ycen']
xref = self.cor_coords[0]
yref = self.cor_coords[1]
dx = xc - xref
dy = yc - yref
total_rotation = u.Quantity(self.rotation - fit_results['rotator'], u.rad).value
dr, phi = cart2pol([dx, dy])
derot_phi = phi + total_rotation
az, el = pol2cart([dr, derot_phi])
az *= self.az_parity * self.pix_size * defoc # pix size scales with the pupil size as focus changes.
el *= self.el_parity * self.pix_size * defoc
return az.round(3), el.round(3)
def clear_m1_corrections(self):
"""
Clear corrections applied to the primary mirror. This includes the 'm1spherical' offsets sent to the secondary.
"""
log.info("Clearing WFS corrections from M1 and m1spherical offsets from M2.")
clear_forces, clear_m1focus = self.telescope.clear_forces()
return clear_forces, clear_m1focus
def clear_m2_corrections(self):
"""
Clear corrections sent to the secondary mirror, specifically the 'wfs' offsets.
"""
log.info("Clearing WFS offsets from M2's hexapod.")
cmds = self.secondary.clear_wfs()
return cmds
def clear_corrections(self):
"""
Clear all applied WFS corrections
"""
forces, m1focus = self.clear_m1_corrections()
cmds = self.clear_m2_corrections()
return forces, m1focus, cmds
def connect(self):
"""
Set state to connected
"""
self.telescope.connect()
self.secondary.connect()
if self.telescope.connected and self.secondary.connected:
self.connected = True
else:
self.connected = False
def disconnect(self):
"""
Set state to disconnected
"""
self.telescope.disconnect()
self.secondary.disconnect()
self.connected = False
class F9(WFS):
"""
Defines configuration and methods specific to the F/9 WFS system
"""
def __init__(self, config={}, plot=True):
super(F9, self).__init__(config=config, plot=plot)
self.connected = False
# set up CompMirror object
self.compmirror = CompMirror()
def connect(self):
"""
Run parent connect() method and then connect to the topbox if we can connect to the rest.
"""
super(F9, self).connect()
if self.connected:
self.compmirror.connect()
def disconnect(self):
"""
Run parent disconnect() method and then disconnect the topbox
"""
super(F9, self).disconnect()
self.compmirror.disconnect()
class NewF9(F9):
"""
Defines configuration and methods specific to the F/9 WFS system with the new SBIG CCD
"""
def process_image(self, fitsfile):
"""
Process the image to make it suitable for accurate wavefront analysis. Steps include nuking cosmic rays,
subtracting background, handling overscan regions, etc.
"""
rawdata, hdr = check_wfsdata(fitsfile, header=True)
cr_mask, data = detect_cosmics(rawdata, sigclip=15., niter=5, cleantype='medmask', psffwhm=10.)
# calculate the background and subtract it
bkg_estimator = photutils.ModeEstimatorBackground()
mask = photutils.make_source_mask(data, nsigma=2, npixels=7, dilate_size=13)
bkg = photutils.Background2D(data, (50, 50), filter_size=(15, 15), bkg_estimator=bkg_estimator, mask=mask)
data -= bkg.background
return data, hdr
class F5(WFS):
"""
Defines configuration and methods specific to the F/5 WFS systems
"""
def __init__(self, config={}, plot=True):
super(F5, self).__init__(config=config, plot=plot)
self.connected = False
self.sock = None
# load lookup table for off-axis aberrations
self.aberr_table = ascii.read(self.aberr_table_file)
def process_image(self, fitsfile):
"""
Process the image to make it suitable for accurate wavefront analysis. Steps include nuking cosmic rays,
subtracting background, handling overscan regions, etc.
"""
rawdata, hdr = check_wfsdata(fitsfile, header=True)
trimdata = self.trim_overscan(rawdata, hdr=hdr)
cr_mask, data = detect_cosmics(trimdata, sigclip=15., niter=5, cleantype='medmask', psffwhm=10.)
# calculate the background and subtract it
bkg_estimator = photutils.ModeEstimatorBackground()
mask = photutils.make_source_mask(data, nsigma=2, npixels=5, dilate_size=11)
bkg = photutils.Background2D(data, (20, 20), filter_size=(10, 10), bkg_estimator=bkg_estimator, mask=mask)
data -= bkg.background
return data, hdr
def ref_pupil_location(self, mode, hdr=None):
"""
For now we set the F/5 wfs center by hand based on engineering data. Should determine this more carefully.
"""
x = 262.0
y = 259.0
return x, y
def focal_plane_position(self, hdr):
"""
Need to fill this in for the hecto f/5 WFS system. For now will assume it's always on-axis.
"""
return 0.0 * u.deg, 0.0 * u.deg
def calculate_recenter(self, fit_results, defoc=1.0):
"""
Perform zero-coma hexapod tilts to align the pupil center to the center-of-rotation.
The location of the CoR is configured to be at self.cor_coords.
"""
xc = fit_results['xcen']
yc = fit_results['ycen']
xref = self.cor_coords[0]
yref = self.cor_coords[1]
dx = xc - xref
dy = yc - yref
cam_rotation = self.rotation - 90 * u.deg # pickoff plus fold mirror makes a 90 deg rotation
total_rotation = u.Quantity(cam_rotation - fit_results['rotator'], u.rad).value
dr, phi = cart2pol([dx, -dy]) # F/5 camera needs an up/down flip
derot_phi = phi + total_rotation
az, el = pol2cart([dr, derot_phi])
az *= self.az_parity * self.pix_size * defoc # pix size scales with the pupil size as focus changes.
el *= self.el_parity * self.pix_size * defoc
return az.round(3), el.round(3)
def reference_aberrations(self, mode, hdr=None):
"""
Create reference ZernikeVector for 'mode'. Pass 'hdr' to self.focal_plane_position() to get position of
the WFS when the data was acquired.
"""
# for most cases, this gets the reference focus
z_default = ZernikeVector(**self.modes[mode]['ref_zern'])
# now get the off-axis aberrations
z_offaxis = ZernikeVector()
if hdr is None:
log.warning("Missing WFS header. Assuming data is acquired on-axis.")
field_r = 0.0 * u.deg
field_phi = 0.0 * u.deg
else:
field_r, field_phi = self.focal_plane_position(hdr)
# ignore piston and x/y tilts
for i in range(4, 12):
k = "Z%02d" % i
z_offaxis[k] = np.interp(field_r.to(u.deg).value, self.aberr_table['field_r'], self.aberr_table[k]) * u.um
# remove the 90 degree offset between the MMT and zernike conventions and then rotate the offaxis aberrations
z_offaxis.rotate(angle=field_phi - 90. * u.deg)
z = z_default + z_offaxis
return z
class Binospec(F5):
"""
Defines configuration and methods specific to the Binospec WFS system. Binospec uses the same aberration table
as the F5 system so we inherit from that.
"""
def get_flipud(self, mode):
"""
Method to determine if the WFS image needs to be flipped up/down
During the first binospec commissioning run the images were flipped u/d as they came in. Since then, they are
left as-is and get flipped internally based on this flag. The reference file is already flipped.
"""
return True
def ref_pupil_location(self, mode, hdr=None):
"""
If a header is passed in, use Jan Kansky's linear relations to get the pupil center on the reference image.
Otherwise, use the default method.
"""
if hdr is None:
ref = self.modes[mode]['reference']
x = ref.xcen
y = ref.ycen
else:
for k in ['STARXMM', 'STARYMM']:
if k not in hdr:
# we'll be lenient for now with missing header info. if not provided, assume we're on-axis.
msg = f"Missing value, {k}, that is required to transform Binospec guider coordinates. Defaulting to 0.0."
log.warning(msg)
hdr[k] = 0.0
y = 232.771 + 0.17544 * hdr['STARXMM']
x = 265.438 + -0.20406 * hdr['STARYMM'] + 12.0
return x, y
def focal_plane_position(self, hdr):
"""
Transform from the Binospec guider coordinate system to MMTO focal plane coordinates.
"""
for k in ['ROT', 'STARXMM', 'STARYMM']:
if k not in hdr:
# we'll be lenient for now with missing header info. if not provided, assume we're on-axis.
msg = f"Missing value, {k}, that is required to transform Binospec guider coordinates. Defaulting to 0.0."
log.warning(msg)
hdr[k] = 0.0
guide_x = hdr['STARXMM']
guide_y = hdr['STARYMM']
rot = hdr['ROT']
guide_r = np.sqrt(guide_x**2 + guide_y**2) * u.mm
rot = u.Quantity(rot, u.deg) # make sure rotation is cast to degrees
# the MMTO focal plane coordinate convention has phi=0 aligned with +Y instead of +X
if guide_y != 0.0:
guide_phi = np.arctan2(guide_x, guide_y) * u.rad
else:
guide_phi = 90. * u.deg
# transform radius in guider coords to degrees in focal plane
focal_r = (guide_r / self.secondary.plate_scale).to(u.deg)
focal_phi = guide_phi + rot + self.rotation
log.debug(f"guide_phi: {guide_phi.to(u.rad)} rot: {rot}")
return focal_r, focal_phi
def in_wfs_region(self, xw, yw, x, y):
"""
Determine if a position is within the region available to Binospec's WFS
"""
return True # placekeeper until the optical prescription is implemented
def pupil_mask(self, hdr, npts=14):
"""
Generate a synthetic pupil mask
"""
if hdr is not None:
x_wfs = hdr.get('STARXMM', 150.0)
y_wfs = hdr.get('STARYMM', 0.0)
else:
x_wfs = 150.0
y_wfs = 0.0
log.warning("Header information not available for Binospec pupil mask. Assuming default position.")
good = []
center = self.pup_size / 2.
obsc = self.telescope.obscuration.value
spacing = 2.0 / npts
for x in np.arange(-1, 1, spacing):
for y in np.arange(-1, 1, spacing):
r = np.hypot(x, y)
if (r < 1 and np.hypot(x, y) >= obsc):
if self.in_wfs_region(x_wfs, y_wfs, x, y):
x_impos = center * (x + 1.)
y_impos = center * (y + 1.)
amp = 1.
# this is kind of a hacky way to dim spots near the edge, but easier than doing full calc
# of the aperture intersection with pupil. it also doesn't need to be that accurate for the
# purposes of the cross-correlation used to register the pupil.
if r > 1. - spacing:
amp = 1. - (r - (1. - spacing)) / spacing
if r - obsc < spacing:
amp = (r - obsc) / spacing
good.append((amp, x_impos, y_impos))
yi, xi = np.mgrid[0:self.pup_size, 0:self.pup_size]
im = np.zeros((self.pup_size, self.pup_size))
sigma = 3.
for g in good:
im += Gaussian2D(g[0], g[1], g[2], sigma, sigma)(xi, yi)
# Measured by hand from reference LED image
cam_rot = 0.595
im_rot = rotate(im, cam_rot, reshape=False)
im_rot[im_rot < 1e-2] = 0.0
return im_rot
class MMIRS(F5):
"""
Defines configuration and methods specific to the MMIRS WFS system
"""
def __init__(self, config={}, plot=True):
super(MMIRS, self).__init__(config=config, plot=plot)
# Parameters describing MMIRS pickoff mirror geometry
# Location and diameter of exit pupil
# Determined by tracing chief ray at 7.2' field angle with mmirs_asbuiltoptics_20110107_corronly.zmx
self.zp = 71.749 / 0.02714
self.dp = self.zp / 5.18661 # Working f/# from Zemax file
# Location of fold mirror
self.zm = 114.8
# Angle of fold mirror
self.am = 42 * u.deg
# Following dimensions from drawing MMIRS-1233_Rev1.pdf
# Diameter of pickoff mirror
self.pickoff_diam = (6.3 * u.imperial.inch).to(u.mm).value
# X size of opening in pickoff mirror
self.pickoff_xsize = (3.29 * u.imperial.inch).to(u.mm).value
# Y size of opening in pickoff mirror
self.pickoff_ysize = (3.53 * u.imperial.inch).to(u.mm).value
# radius of corner in pickoff mirror
self.pickoff_rcirc = (0.4 * u.imperial.inch).to(u.mm).value
def mirrorpoint(self, x0, y0, x, y):
"""
Compute intersection of ray with pickoff mirror.
The ray leaves the exit pupil at position x,y and hits the focal surface at x0,y0.
Math comes from http://geomalgorithms.com/a05-_intersect-1.html
"""
# Point in focal plane
P0 = np.array([x0, y0, 0])
# Point in exit pupil
P1 = np.array([x * self.dp / 2, y * self.dp / 2, self.zp])
# Pickoff mirror intesection with optical axis
V0 = np.array([0, 0, self.zm])
# normal to mirror
if (x0 < 0):
n = np.array([-np.sin(self.am), 0, np.cos(self.am)])
else:
n = np.array([np.sin(self.am), 0, np.cos(self.am)])
w = P0 - V0
# Vector connecting P0 to P1
u = P1 - P0
# Distance from P0 to intersection as a fraction of abs(u)
s = -n.dot(w) / n.dot(u)
# Intersection point on mirror
P = P0 + s * u
return (P[0], P[1])
def onmirror(self, x, y, side):
"""
Determine if a point is on the pickoff mirror surface:
x,y = coordinates of ray
side=1 means right face of the pickoff mirror, -1=left face
"""
if np.hypot(x, y) > self.pickoff_diam / 2.:
return False
if x * side < 0:
return False
x = abs(x)
y = abs(y)
if ((x > self.pickoff_xsize/2) or (y > self.pickoff_ysize/2)
or (x > self.pickoff_xsize/2 - self.pickoff_rcirc and y > self.pickoff_ysize/2 - self.pickoff_rcirc
and np.hypot(x - (self.pickoff_xsize/2 - self.pickoff_rcirc),
y - (self.pickoff_ysize/2 - self.pickoff_rcirc)) > self.pickoff_rcirc)):
return True
else:
return False
def drawoutline(self, ax):
"""
Draw outline of MMIRS pickoff mirror onto matplotlib axis, ax
"""
circ = np.arange(360) * u.deg
ax.plot(np.cos(circ) * self.pickoff_diam/2, np.sin(circ) * self.pickoff_diam/2, "b")
ax.set_aspect('equal', 'datalim')
ax.plot(
[-(self.pickoff_xsize/2 - self.pickoff_rcirc), (self.pickoff_xsize/2 - self.pickoff_rcirc)],
[self.pickoff_ysize/2, self.pickoff_ysize/2],
"b"
)
ax.plot(
[-(self.pickoff_xsize/2 - self.pickoff_rcirc), (self.pickoff_xsize/2 - self.pickoff_rcirc)],
[-self.pickoff_ysize/2, -self.pickoff_ysize/2],
"b"
)
ax.plot(
[-(self.pickoff_xsize/2), -(self.pickoff_xsize/2)],
[self.pickoff_ysize/2 - self.pickoff_rcirc, -(self.pickoff_ysize/2 - self.pickoff_rcirc)],
"b"
)
ax.plot(
[(self.pickoff_xsize/2), (self.pickoff_xsize/2)],
[self.pickoff_ysize/2 - self.pickoff_rcirc, -(self.pickoff_ysize/2 - self.pickoff_rcirc)],
"b"
)
ax.plot(
np.cos(circ[0:90]) * self.pickoff_rcirc + self.pickoff_xsize/2 - self.pickoff_rcirc,
np.sin(circ[0:90]) * self.pickoff_rcirc + self.pickoff_ysize/2 - self.pickoff_rcirc,
"b"
)
ax.plot(
np.cos(circ[90:180]) * self.pickoff_rcirc - self.pickoff_xsize/2 + self.pickoff_rcirc,
np.sin(circ[90:180]) * self.pickoff_rcirc + self.pickoff_ysize/2 - self.pickoff_rcirc,
"b"
)
ax.plot(
np.cos(circ[180:270]) * self.pickoff_rcirc - self.pickoff_xsize/2 + self.pickoff_rcirc,
np.sin(circ[180:270]) * self.pickoff_rcirc - self.pickoff_ysize/2 + self.pickoff_rcirc,
"b"
)
ax.plot(
np.cos(circ[270:360]) * self.pickoff_rcirc + self.pickoff_xsize/2 - self.pickoff_rcirc,
np.sin(circ[270:360]) * self.pickoff_rcirc - self.pickoff_ysize/2 + self.pickoff_rcirc,
"b"
)
ax.plot([0, 0], [self.pickoff_ysize/2, self.pickoff_diam/2], "b")
ax.plot([0, 0], [-self.pickoff_ysize/2, -self.pickoff_diam/2], "b")
def plotgrid(self, x0, y0, ax, npts=15):
"""
Plot a grid of points representing Shack-Hartmann apertures corresponding to wavefront sensor positioned at
a focal plane position of x0, y0 mm. This position is written in the FITS header keywords GUIDERX and GUIDERY.
"""
ngood = 0
for x in np.arange(-1, 1, 2.0 / npts):
for y in np.arange(-1, 1, 2.0 / npts):
if (np.hypot(x, y) < 1 and np.hypot(x, y) >= self.telescope.obscuration): # Only plot points w/in the pupil
xm, ym = self.mirrorpoint(x0, y0, x, y) # Get intersection with pickoff
if self.onmirror(xm, ym, x0/abs(x0)): # Find out if point is on the mirror surface
ax.scatter(xm, ym, 1, "g")
ngood += 1
else:
ax.scatter(xm, ym, 1, "r")
return ngood
def plotgrid_hdr(self, hdr, ax, npts=15):
"""
Wrap self.plotgrid() and get x0, y0 values from hdr.
"""
if 'GUIDERX' not in hdr or 'GUIDERY' not in hdr:
msg = "No MMIRS WFS position available in header."
raise WFSCommandException(value=msg)
x0 = hdr['GUIDERX']
y0 = hdr['GUIDERY']
ngood = self.plotgrid(x0, y0, ax=ax, npts=npts)
return ngood
def pupil_mask(self, hdr, npts=15):
"""
Use MMIRS pickoff mirror geometry to calculate the pupil mask
"""
if 'GUIDERX' not in hdr or 'GUIDERY' not in hdr:
msg = "No MMIRS WFS position available in header."
raise WFSCommandException(value=msg)
if 'CA' not in hdr:
msg = "No camera rotation angle available in header."
raise WFSCommandException(value=msg)
cam_rot = hdr['CA']
x0 = hdr['GUIDERX']
y0 = hdr['GUIDERY']
good = []
center = self.pup_size / 2.
obsc = self.telescope.obscuration.value
spacing = 2.0 / npts
for x in np.arange(-1, 1, spacing):
for y in np.arange(-1, 1, spacing):
r = np.hypot(x, y)
if (r < 1 and np.hypot(x, y) >= obsc):
xm, ym = self.mirrorpoint(x0, y0, x, y)
if self.onmirror(xm, ym, x0/abs(x0)):
x_impos = center * (x + 1.)
y_impos = center * (y + 1.)
amp = 1.
# this is kind of a hacky way to dim spots near the edge, but easier than doing full calc
# of the aperture intersection with pupil. it also doesn't need to be that accurate for the
# purposes of the cross-correlation used to register the pupil.
if r > 1. - spacing:
amp = 1. - (r - (1. - spacing)) / spacing
if r - obsc < spacing:
amp = (r - obsc) / spacing
good.append((amp, x_impos, y_impos))
yi, xi = np.mgrid[0:self.pup_size, 0:self.pup_size]
im =
|
np.zeros((self.pup_size, self.pup_size))
|
numpy.zeros
|
"""
Tests for the BNMF Gibbs sampler.
"""
import sys, os
project_location = os.path.dirname(__file__)+"/../../../"
sys.path.append(project_location)
import numpy, math, pytest, itertools
from BNMTF.code.models.bnmf_gibbs_optimised import bnmf_gibbs_optimised
""" Test constructor """
def test_init():
# Test getting an exception when R and M are different sizes, and when R is not a 2D array.
R1 = numpy.ones(3)
M = numpy.ones((2,3))
I,J,K = 5,3,1
lambdaU = numpy.ones((I,K))
lambdaV = numpy.ones((J,K))
alpha, beta = 3, 1
priors = { 'alpha':alpha, 'beta':beta, 'lambdaU':lambdaU, 'lambdaV':lambdaV }
with pytest.raises(AssertionError) as error:
bnmf_gibbs_optimised(R1,M,K,priors)
assert str(error.value) == "Input matrix R is not a two-dimensional array, but instead 1-dimensional."
R2 = numpy.ones((4,3,2))
with pytest.raises(AssertionError) as error:
bnmf_gibbs_optimised(R2,M,K,priors)
assert str(error.value) == "Input matrix R is not a two-dimensional array, but instead 3-dimensional."
R3 = numpy.ones((3,2))
with pytest.raises(AssertionError) as error:
bnmf_gibbs_optimised(R3,M,K,priors)
assert str(error.value) == "Input matrix R is not of the same size as the indicator matrix M: (3, 2) and (2, 3) respectively."
# Similarly for lambdaU, lambdaV
R4 = numpy.ones((2,3))
lambdaU = numpy.ones((2+1,1))
priors = { 'alpha':alpha, 'beta':beta, 'lambdaU':lambdaU, 'lambdaV':lambdaV }
with pytest.raises(AssertionError) as error:
bnmf_gibbs_optimised(R4,M,K,priors)
assert str(error.value) == "Prior matrix lambdaU has the wrong shape: (3, 1) instead of (2, 1)."
lambdaU = numpy.ones((2,1))
lambdaV = numpy.ones((3+1,1))
priors = { 'alpha':alpha, 'beta':beta, 'lambdaU':lambdaU, 'lambdaV':lambdaV }
with pytest.raises(AssertionError) as error:
bnmf_gibbs_optimised(R4,M,K,priors)
assert str(error.value) == "Prior matrix lambdaV has the wrong shape: (4, 1) instead of (3, 1)."
# Test getting an exception if a row or column is entirely unknown
lambdaU = numpy.ones((2,1))
lambdaV = numpy.ones((3,1))
M1 = [[1,1,1],[0,0,0]]
M2 = [[1,1,0],[1,0,0]]
priors = { 'alpha':alpha, 'beta':beta, 'lambdaU':lambdaU, 'lambdaV':lambdaV }
with pytest.raises(AssertionError) as error:
bnmf_gibbs_optimised(R4,M1,K,priors)
assert str(error.value) == "Fully unobserved row in R, row 1."
with pytest.raises(AssertionError) as error:
bnmf_gibbs_optimised(R4,M2,K,priors)
assert str(error.value) == "Fully unobserved column in R, column 2."
# Finally, a successful case
I,J,K = 3,2,2
R5 = 2*numpy.ones((I,J))
lambdaU = numpy.ones((I,K))
lambdaV = numpy.ones((J,K))
M = numpy.ones((I,J))
priors = { 'alpha':alpha, 'beta':beta, 'lambdaU':lambdaU, 'lambdaV':lambdaV }
BNMF = bnmf_gibbs_optimised(R5,M,K,priors)
assert numpy.array_equal(BNMF.R,R5)
assert numpy.array_equal(BNMF.M,M)
assert BNMF.I == I
assert BNMF.J == J
assert BNMF.K == K
assert BNMF.size_Omega == I*J
assert BNMF.alpha == alpha
assert BNMF.beta == beta
assert numpy.array_equal(BNMF.lambdaU,lambdaU)
assert numpy.array_equal(BNMF.lambdaV,lambdaV)
# And when lambdaU and lambdaV are integers
I,J,K = 3,2,2
R5 = 2*numpy.ones((I,J))
lambdaU = 3.
lambdaV = 4.
M = numpy.ones((I,J))
priors = { 'alpha':alpha, 'beta':beta, 'lambdaU':lambdaU, 'lambdaV':lambdaV }
BNMF = bnmf_gibbs_optimised(R5,M,K,priors)
assert numpy.array_equal(BNMF.R,R5)
assert numpy.array_equal(BNMF.M,M)
assert BNMF.I == I
assert BNMF.J == J
assert BNMF.K == K
assert BNMF.size_Omega == I*J
assert BNMF.alpha == alpha
assert BNMF.beta == beta
assert numpy.array_equal(BNMF.lambdaU,lambdaU*numpy.ones((I,K)))
assert numpy.array_equal(BNMF.lambdaV,lambdaV*numpy.ones((J,K)))
""" Test initialing parameters """
def test_initialise():
I,J,K = 5,3,2
R = numpy.ones((I,J))
M = numpy.ones((I,J))
lambdaU = 2*numpy.ones((I,K))
lambdaV = 3*numpy.ones((J,K))
alpha, beta = 3, 1
priors = { 'alpha':alpha, 'beta':beta, 'lambdaU':lambdaU, 'lambdaV':lambdaV }
# First do a random initialisation - we can then only check whether values are correctly initialised
init = 'random'
BNMF = bnmf_gibbs_optimised(R,M,K,priors)
BNMF.initialise(init)
assert BNMF.tau >= 0.0
for i,k in itertools.product(xrange(0,I),xrange(0,K)):
assert BNMF.U[i,k] >= 0.0
for j,k in itertools.product(xrange(0,J),xrange(0,K)):
assert BNMF.V[j,k] >= 0.0
# Then initialise with expectation values
init = 'exp'
BNMF = bnmf_gibbs_optimised(R,M,K,priors)
BNMF.initialise(init)
assert BNMF.tau >= 0.0
for i,k in itertools.product(xrange(0,I),xrange(0,K)):
assert BNMF.U[i,k] == 1./2.
for j,k in itertools.product(xrange(0,J),xrange(0,K)):
assert BNMF.V[j,k] == 1./3.
#assert BNMF.tau == 3./1.
""" Test computing values for alpha, beta, mu, tau. """
I,J,K = 5,3,2
R = numpy.ones((I,J))
M = numpy.ones((I,J))
M[0,0], M[2,2], M[3,1] = 0, 0, 0
lambdaU = 2*numpy.ones((I,K))
lambdaV = 3*numpy.ones((J,K))
alpha, beta = 3, 1
priors = { 'alpha':alpha, 'beta':beta, 'lambdaU':lambdaU, 'lambdaV':lambdaV }
init = 'exp' #U=1/2,V=1/3
def test_alpha_s():
BNMF = bnmf_gibbs_optimised(R,M,K,priors)
BNMF.initialise(init)
alpha_s = alpha + 6.
assert BNMF.alpha_s() == alpha_s
def test_beta_s():
BNMF = bnmf_gibbs_optimised(R,M,K,priors)
BNMF.initialise(init)
beta_s = beta + .5*(12*(2./3.)**2) #U*V.T = [[1/6+1/6,..]]
assert abs(BNMF.beta_s() - beta_s) < 0.000000000000001
def test_tauU():
BNMF = bnmf_gibbs_optimised(R,M,K,priors)
BNMF.initialise(init)
BNMF.tau = 3.
#V^2 = [[1/9,1/9],[1/9,1/9],[1/9,1/9]], sum_j V^2 = [2/9,1/3,2/9,2/9,1/3] (index=i)
tauU = 3.*numpy.array([[2./9.,2./9.],[1./3.,1./3.],[2./9.,2./9.],[2./9.,2./9.],[1./3.,1./3.]])
for i,k in itertools.product(xrange(0,I),xrange(0,K)):
assert BNMF.tauU(k)[i] == tauU[i,k]
def test_muU():
BNMF = bnmf_gibbs_optimised(R,M,K,priors)
BNMF.initialise(init)
BNMF.tau = 3.
#U*V^T - Uik*Vjk = [[1/6,..]], so Rij - Ui * Vj + Uik * Vjk = 5/6
tauU = 3.*numpy.array([[2./9.,2./9.],[1./3.,1./3.],[2./9.,2./9.],[2./9.,2./9.],[1./3.,1./3.]])
muU = 1./tauU * ( 3. * numpy.array([[2.*(5./6.)*(1./3.),10./18.],[15./18.,15./18.],[10./18.,10./18.],[10./18.,10./18.],[15./18.,15./18.]]) - lambdaU )
for i,k in itertools.product(xrange(0,I),xrange(0,K)):
assert abs(BNMF.muU(tauU[:,k],k)[i] - muU[i,k]) < 0.000000000000001
def test_tauV():
BNMF = bnmf_gibbs_optimised(R,M,K,priors)
BNMF.initialise(init)
BNMF.tau = 3.
#U^2 = [[1/4,1/4],[1/4,1/4],[1/4,1/4],[1/4,1/4],[1/4,1/4]], sum_i U^2 = [1,1,1] (index=j)
tauV = 3.*numpy.array([[1.,1.],[1.,1.],[1.,1.]])
for j,k in itertools.product(xrange(0,J),xrange(0,K)):
assert BNMF.tauV(k)[j] == tauV[j,k]
def test_muV():
BNMF = bnmf_gibbs_optimised(R,M,K,priors)
BNMF.initialise(init)
BNMF.tau = 3.
#U*V^T - Uik*Vjk = [[1/6,..]], so Rij - Ui * Vj + Uik * Vjk = 5/6
tauV = 3.*numpy.array([[1.,1.],[1.,1.],[1.,1.]])
muV = 1./tauV * ( 3. * numpy.array([[4.*(5./6.)*(1./2.),4.*(5./6.)*(1./2.)],[4.*(5./6.)*(1./2.),4.*(5./6.)*(1./2.)],[4.*(5./6.)*(1./2.),4.*(5./6.)*(1./2.)]]) - lambdaV )
for j,k in itertools.product(xrange(0,J),xrange(0,K)):
assert BNMF.muV(tauV[:,k],k)[j] == muV[j,k]
""" Test some iterations, and that the values have changed in U and V. """
def test_run():
I,J,K = 10,5,2
R = numpy.ones((I,J))
M = numpy.ones((I,J))
M[0,0], M[2,2], M[3,1] = 0, 0, 0
lambdaU = 2*numpy.ones((I,K))
lambdaV = 3*numpy.ones((J,K))
alpha, beta = 3, 1
priors = { 'alpha':alpha, 'beta':beta, 'lambdaU':lambdaU, 'lambdaV':lambdaV }
init = 'exp' #U=1/2,V=1/3
U_prior = numpy.ones((I,K))/2.
V_prior = numpy.ones((J,K))/3.
iterations = 15
BNMF = bnmf_gibbs_optimised(R,M,K,priors)
BNMF.initialise(init)
(Us,Vs,taus) = BNMF.run(iterations)
assert BNMF.all_U.shape == (iterations,I,K)
assert BNMF.all_V.shape == (iterations,J,K)
assert BNMF.all_tau.shape == (iterations,)
for i,k in itertools.product(xrange(0,I),xrange(0,K)):
assert Us[0,i,k] != U_prior[i,k]
for j,k in itertools.product(xrange(0,J),xrange(0,K)):
assert Vs[0,j,k] != V_prior[j,k]
assert taus[1] != alpha/float(beta)
""" Test approximating the expectations for U, V, tau """
def test_approx_expectation():
burn_in = 2
thinning = 3 # so index 2,5,8 -> m=3,m=6,m=9
(I,J,K) = (5,3,2)
Us = [numpy.ones((I,K)) * 3*m**2 for m in range(1,10+1)] #first is 1's, second is 4's, third is 9's, etc.
Vs = [numpy.ones((J,K)) * 2*m**2 for m in range(1,10+1)]
taus = [m**2 for m in range(1,10+1)]
expected_exp_tau = (9.+36.+81.)/3.
expected_exp_U = numpy.array([[9.+36.+81.,9.+36.+81.],[9.+36.+81.,9.+36.+81.],[9.+36.+81.,9.+36.+81.],[9.+36.+81.,9.+36.+81.],[9.+36.+81.,9.+36.+81.]])
expected_exp_V = numpy.array([[(9.+36.+81.)*(2./3.),(9.+36.+81.)*(2./3.)],[(9.+36.+81.)*(2./3.),(9.+36.+81.)*(2./3.)],[(9.+36.+81.)*(2./3.),(9.+36.+81.)*(2./3.)]])
R = numpy.ones((I,J))
M = numpy.ones((I,J))
lambdaU = 2*numpy.ones((I,K))
lambdaV = 3*numpy.ones((J,K))
alpha, beta = 3, 1
priors = { 'alpha':alpha, 'beta':beta, 'lambdaU':lambdaU, 'lambdaV':lambdaV }
BNMF = bnmf_gibbs_optimised(R,M,K,priors)
BNMF.all_U = Us
BNMF.all_V = Vs
BNMF.all_tau = taus
(exp_U, exp_V, exp_tau) = BNMF.approx_expectation(burn_in,thinning)
assert expected_exp_tau == exp_tau
assert numpy.array_equal(expected_exp_U,exp_U)
assert numpy.array_equal(expected_exp_V,exp_V)
""" Test computing the performance of the predictions using the expectations """
def test_predict():
burn_in = 2
thinning = 3 # so index 2,5,8 -> m=3,m=6,m=9
(I,J,K) = (5,3,2)
Us = [numpy.ones((I,K)) * 3*m**2 for m in range(1,10+1)] #first is 1's, second is 4's, third is 9's, etc.
Vs = [numpy.ones((J,K)) * 2*m**2 for m in range(1,10+1)]
Us[2][0,0] = 24 #instead of 27 - to ensure we do not get 0 variance in our predictions
taus = [m**2 for m in range(1,10+1)]
R = numpy.array([[1,2,3],[4,5,6],[7,8,9],[10,11,12],[13,14,15]],dtype=float)
M = numpy.ones((I,J))
lambdaU = 2*numpy.ones((I,K))
lambdaV = 3*numpy.ones((J,K))
alpha, beta = 3, 1
priors = { 'alpha':alpha, 'beta':beta, 'lambdaU':lambdaU, 'lambdaV':lambdaV }
#expected_exp_U = numpy.array([[125.,126.],[126.,126.],[126.,126.],[126.,126.],[126.,126.]])
#expected_exp_V = numpy.array([[84.,84.],[84.,84.],[84.,84.]])
#R_pred = numpy.array([[21084.,21084.,21084.],[ 21168.,21168.,21168.],[21168.,21168.,21168.],[21168.,21168.,21168.],[21168.,21168.,21168.]])
M_test =
|
numpy.array([[0,0,1],[0,1,0],[0,0,0],[1,1,0],[0,0,0]])
|
numpy.array
|
# Copyright (c) 2016, The Bifrost Authors. All rights reserved.
# Copyright (c) 2016, NVIDIA CORPORATION. All rights reserved.
#
# Redistribution and use in source and binary forms, with or without
# modification, are permitted provided that the following conditions
# are met:
# * Redistributions of source code must retain the above copyright
# notice, this list of conditions and the following disclaimer.
# * Redistributions in binary form must reproduce the above copyright
# notice, this list of conditions and the following disclaimer in the
# documentation and/or other materials provided with the distribution.
# * Neither the name of The Bifrost Authors nor the names of its
# contributors may be used to endorse or promote products derived
# from this software without specific prior written permission.
#
# THIS SOFTWARE IS PROVIDED BY THE COPYRIGHT HOLDERS ``AS IS'' AND ANY
# EXPRESS OR IMPLIED WARRANTIES, INCLUDING, BUT NOT LIMITED TO, THE
# IMPLIED WARRANTIES OF MERCHANTABILITY AND FITNESS FOR A PARTICULAR
# PURPOSE ARE DISCLAIMED. IN NO EVENT SHALL THE COPYRIGHT OWNER OR
# CONTRIBUTORS BE LIABLE FOR ANY DIRECT, INDIRECT, INCIDENTAL, SPECIAL,
# EXEMPLARY, OR CONSEQUENTIAL DAMAGES (INCLUDING, BUT NOT LIMITED TO,
# PROCUREMENT OF SUBSTITUTE GOODS OR SERVICES; LOSS OF USE, DATA, OR
# PROFITS; OR BUSINESS INTERRUPTION) HOWEVER CAUSED AND ON ANY THEORY
# OF LIABILITY, WHETHER IN CONTRACT, STRICT LIABILITY, OR TORT
# (INCLUDING NEGLIGENCE OR OTHERWISE) ARISING IN ANY WAY OUT OF THE USE
# OF THIS SOFTWARE, EVEN IF ADVISED OF THE POSSIBILITY OF SUCH DAMAGE.
import ctypes
import unittest
import numpy as np
import bifrost as bf
class TestMap(unittest.TestCase):
def setUp(self):
np.random.seed(1234)
def run_simple_test(self, x, funcstr, func):
x_orig = x
x = bf.asarray(x, 'cuda')
y = bf.empty_like(x)
x.flags['WRITEABLE'] = False
x.bf.immutable = True # TODO: Is this actually doing anything? (flags is, just not sure about bf.immutable)
bf.map(funcstr, x=x, y=y)
x = x.copy('system')
y = y.copy('system')
if isinstance(x_orig, bf.ndarray):
x_orig = x
# Note: Using func(x) is dangerous because bf.ndarray does things like
# lazy .conj(), which break when used as if it were np.ndarray.
np.testing.assert_equal(y, func(x_orig))
def run_simple_test_funcs(self, x):
self.run_simple_test(x, "y = x+1", lambda x: x+1)
self.run_simple_test(x, "y = x*3", lambda x: x*3)
# Note: Must use "f" suffix to avoid very slow double-precision math
self.run_simple_test(x, "y = rint(pow(x, 2.f))", lambda x: x**2)
self.run_simple_test(x, "auto tmp = x; y = tmp*tmp", lambda x: x*x)
self.run_simple_test(x, "y = x; y += x", lambda x: x+x)
def test_simple_1D(self):
n = 7919
x = np.random.randint(256, size=n)
self.run_simple_test_funcs(x)
def test_simple_2D(self):
n = 89
x = np.random.randint(256, size=(n,n))
self.run_simple_test_funcs(x)
def test_simple_2D_padded(self):
n = 89
x = np.random.randint(256, size=(n,n))
x = bf.asarray(x, space='cuda')
x = x[:,1:]
self.run_simple_test_funcs(x)
def test_simple_3D(self):
n = 23
x = np.random.randint(256, size=(n,n,n))
self.run_simple_test_funcs(x)
def test_simple_3D_padded(self):
n = 23
x =
|
np.random.randint(256, size=(n,n,n))
|
numpy.random.randint
|
from __future__ import absolute_import, division, print_function
from six.moves import range
import os
import h5py
import numpy as np
from xfel.euxfel.read_geom import read_geom
from libtbx.phil import parse
import six
from libtbx.utils import Sorry
import datetime
from xfel.util.jungfrau import pad_stacked_format
phil_scope = parse("""
unassembled_file = None
.type = path
.help = hdf5 file used to read in image data.
geom_file = None
.type = path
.help = geometry file to be read in for detector (.geom).
output_file = None
.type = path
.help = output file path
detector_distance = None
.type = float
.help = Detector distance
wavelength = None
.type = float
.help = If not provided, try to find wavelength in unassembled file.
beam_file = None
.type = path
.help = Overrides wavelength. Reads the pulse IDs in the provided file \
to get a list of wavelengths for the master.
include_spectra = False
.type = bool
.help = If true, 2D spectral data will be included in the master file, \
as read from the beam_file.
energy_offset = None
.type = float
.help = If set, add this offset (in eV) to the energy axis in the \
spectra in the beam file and to the per-shot wavelength.
mask_file = None
.type = str
.help = Path to file with external bad pixel mask.
split_modules_into_asics = True
.type = bool
.help = Whether to split the 4x2 modules into indivdual asics \
accounting for borders and gaps.
trusted_range = None
.type = floats(size=2)
.help = Set the trusted range
raw = False
.type = bool
.help = Whether the data being analyzed is raw data from the JF16M or has \
been corrected and padded.
unassembled_data_key = None
.type = str
.expert_level = 2
.help = Override hdf5 key name in unassembled file
pedestal_file = None
.type = str
.help = path to Jungfrau pedestal file
gain_file = None
.type = str
.help = path to Jungfrau gain file
raw_file = None
.type = str
.help = path to Jungfrau raw file
nexus_details {
instrument_name = SwissFEL ARAMIS BEAMLINE ESB
.type = str
.help = Name of instrument
instrument_short_name = ESB
.type = str
.help = short name for instrument, perhaps the acronym
source_name = SwissFEL ARAMIS
.type = str
.help = Name of the neutron or x-ray storage ring/facility
source_short_name = SwissFEL ARAMIS
.type = str
.help = short name for source, perhaps the acronym
start_time = None
.type = str
.help = ISO 8601 time/date of the first data point collected in UTC, \
using the Z suffix to avoid confusion with local time
end_time = None
.type = str
.help = ISO 8601 time/date of the last data point collected in UTC, \
using the Z suffix to avoid confusion with local time. \
This field should only be filled when the value is accurately \
observed. If the data collection aborts or otherwise prevents \
accurate recording of the end_time, this field should be omitted
end_time_estimated = None
.type = str
.help = ISO 8601 time/date of the last data point collected in UTC, \
using the Z suffix to avoid confusion with local time. \
This field may be filled with a value estimated before an \
observed value is avilable.
sample_name = None
.type = str
.help = Descriptive name of sample
total_flux = None
.type = float
.help = flux incident on beam plane in photons per second
}
""")
'''
This script creates a master nexus file by taking in as input a) an hdf5 file and b) a .geom file
The hd5f file is generated by the JF16M after processing the raw images and doing appropriate gain corrections
The assumed parameters for the detector can be seen in the __init__ function and should be changed
if they are modified at in the future
'''
class jf16m_cxigeom2nexus(object):
def __init__(self, args):
self.params_from_phil(args)
if self.params.detector_distance == None:
self.params.detector_distance = 100.0 # Set detector distance arbitrarily if nothing is provided
self.hierarchy = read_geom(self.params.geom_file)
self.n_quads = 4
self.n_modules = 8
def params_from_phil(self, args):
user_phil = []
for arg in args:
if os.path.isfile(arg):
user_phil.append(parse(file_name=arg))
else:
try:
user_phil.append(parse(arg))
except Exception as e:
raise Sorry("Unrecognized argument: %s"%arg)
self.params = phil_scope.fetch(sources=user_phil).extract()
def _create_scalar(self, handle,path,dtype,value):
dataset = handle.create_dataset(path, (),dtype=dtype)
dataset[()] = value
def create_vector(self,handle, name, value, **attributes):
handle.create_dataset(name, (1,), data = [value], dtype='f')
for key,attribute in six.iteritems(attributes):
handle[name].attrs[key] = attribute
def create_nexus_master_file(self):
'''
Hierarchical structure of master nexus file. Format information available here
http://download.nexusformat.org/sphinx/classes/base_classes/NXdetector_module.html#nxdetector-module
--> entry
--> data
--> definition (leaf)
--> instrument
--> sample
'''
output_file_name = self.params.output_file if self.params.output_file is not None else os.path.splitext(self.params.unassembled_file)[0]+'_master.h5'
f = h5py.File(output_file_name, 'w')
f.attrs['NX_class'] = 'NXroot'
f.attrs['file_name'] = os.path.basename(output_file_name)
f.attrs['file_time'] = datetime.datetime.utcnow().strftime("%Y-%m-%dT%H:%M:%SZ")
f.attrs['HDF5_Version'] = h5py.version.hdf5_version
entry = f.create_group('entry')
entry.attrs['NX_class'] = 'NXentry'
if self.params.nexus_details.start_time: entry['start_time'] = self.params.nexus_details.start_time
if self.params.nexus_details.end_time: entry['end_time'] = self.params.nexus_details.end_time
if self.params.nexus_details.end_time_estimated: entry['end_time_estimated'] = self.params.nexus_details.end_time_estimated
# --> definition
self._create_scalar(entry, 'definition', 'S4', np.string_('NXmx'))
# --> data
data = entry.create_group('data')
data.attrs['NX_class'] = 'NXdata'
data_key = 'data'
if self.params.unassembled_data_key:
unassembled_data_key = self.params.unassembled_data_key
else:
if self.params.raw:
unassembled_data_key = "data/JF07T32V01/data"
else:
unassembled_data_key = "data/data"
data[data_key] = h5py.ExternalLink(self.params.unassembled_file, unassembled_data_key)
if self.params.raw_file is not None:
assert not self.params.raw
with h5py.File(self.params.pedestal_file, "r") as pedh5:
print("Padding raw pedestal data")
mean_pedestal = [pad_stacked_format(raw) for raw in pedh5["gains"]]
print("Padding raw pedestal RMS data")
sigma_pedestal = [pad_stacked_format(raw) for raw in pedh5["gainsRMS"]]
data.create_dataset("pedestal", data=mean_pedestal, dtype=np.float32)
data.create_dataset('pedestalRMS', data=sigma_pedestal, dtype=np.float32)
with h5py.File(self.params.gain_file, "r") as gainh5:
print("Padding gains")
gains = [pad_stacked_format(raw) for raw in gainh5["gains"]]
data.create_dataset("gains", data=gains, dtype=np.float32)
data.attrs['signal'] = 'data'
raw_file_handle = h5py.File(self.params.raw_file, "r")
res_file_handle = h5py.File(self.params.unassembled_file, "r")
raw_dset = raw_file_handle["data/JF07T32V01/data"]
raw_shape = raw_dset.shape
_, raw_slowDim, raw_fastDim = raw_shape
raw_type = raw_dset.dtype
num_imgs = res_file_handle['data/data'].shape[0]
raw_layout = h5py.VirtualLayout(shape=(num_imgs, raw_slowDim, raw_fastDim), dtype=raw_type)
raw_pulses = raw_file_handle['data/JF07T32V01/pulse_id'][()][:, 0]
assert np.all(raw_pulses == np.sort(raw_pulses)) # NOTE; this is quick, however I think this is always the case
res_pulses = h5py.File(self.params.unassembled_file, 'r')['data/pulse_id'][()]
raw_source = h5py.VirtualSource(self.params.raw_file, 'data/JF07T32V01/data', shape=raw_shape)
for res_imgnum, raw_imgnum in enumerate(np.searchsorted(raw_pulses, res_pulses)):
raw_layout[res_imgnum] = raw_source[raw_imgnum]
data.create_virtual_dataset('raw', raw_layout)
if self.params.raw:
if self.params.pedestal_file:
# named gains instead of pedestal in JF data files
data['pedestal'] = h5py.ExternalLink(self.params.pedestal_file, 'gains')
data['pedestalRMS'] = h5py.ExternalLink(self.params.pedestal_file, 'gainsRMS')
if self.params.gain_file:
data['gains'] = h5py.ExternalLink(self.params.gain_file, 'gains')
if self.params.pedestal_file or self.params.gain_file:
data.attrs['signal'] = 'data'
#--> sample
sample = entry.create_group('sample')
sample.attrs['NX_class'] = 'NXsample'
if self.params.nexus_details.sample_name: sample['name'] = self.params.nexus_details.sample_name
sample['depends_on'] = '.' # This script does not support scans/gonios
# --> source
source = entry.create_group('source')
source.attrs['NX_class'] = 'NXsource'
source['name'] = self.params.nexus_details.source_name
source['name'].attrs['short_name'] = self.params.nexus_details.source_short_name
# --> instrument
instrument = entry.create_group('instrument')
instrument.attrs['NX_class'] = 'NXinstrument'
instrument["name"] = self.params.nexus_details.instrument_name
instrument["name"].attrs["short_name"] = self.params.nexus_details.instrument_short_name
beam = instrument.create_group('beam')
beam.attrs['NX_class'] = 'NXbeam'
if self.params.nexus_details.total_flux:
self._create_scalar(beam, 'total_flux', 'f', self.params.nexus_details.total_flux)
beam['total_flux'].attrs['units'] = 'Hz'
if self.params.wavelength is None and self.params.beam_file is None:
wavelengths = h5py.File(self.params.unassembled_file, 'r')['instrument/photon_wavelength_A']
beam.create_dataset('incident_wavelength', (1,), data=
|
np.mean(wavelengths)
|
numpy.mean
|
# -*- coding: utf-8 -*-
"""
Segments and curves intersections in n-dimensional Euclidean space
The module provides routines and methods for determining segments and curves intersections
in n-dimensional Euclidean space.
Currently, two methods are implemented out of the box:
- ``exact`` -- exact interesections by solving system of linear equations
- ``almost`` -- almost intersections by shortest connecting segments
``almost`` method can be useful for 3 and higher dimensions.
"""
import typing as ty
import typing_extensions as ty_ext
import abc
import enum
import warnings
import numpy as np
import networkx as nx
from scipy.spatial.distance import pdist, squareform
import skcurve._base
from skcurve import _geomalg
from skcurve._numeric import F_EPS
if ty.TYPE_CHECKING:
from skcurve._base import Point, Segment, Curve # noqa
_intersect_methods = {} # type: ty.Dict[str, ty.Type['IntersectionMethodBase']]
_default_intersect_method = None # type: ty.Optional[str]
class IntersectionWarning(UserWarning):
"""All intersection warnings
"""
class IntersectionError(Exception):
"""All intersection errors
"""
class IntersectionType(enum.Enum):
"""The types of intersection cases
"""
NONE = 0
EXACT = 1
OVERLAP = 2
ALMOST = 3
def __call__(self, intersect_data: ty.Optional[ty.Union['Point', 'Segment']] = None) -> 'IntersectionInfo':
if self == IntersectionType.NONE and intersect_data is not IntersectionType:
raise ValueError(f"'intersect_data' must be 'None' for type {self}")
if self == IntersectionType.EXACT and not isinstance(intersect_data, skcurve._base.Point):
raise ValueError(f"'intersect_data' must be 'Point' for type {self}")
if (self in (IntersectionType.OVERLAP, IntersectionType.ALMOST) and
not isinstance(intersect_data, skcurve._base.Segment)):
raise ValueError(f"'intersect_data' must be 'Segment' for type {self}")
return IntersectionInfo(intersect_data, self)
IntersectionInfo = ty.NamedTuple('IntersectionInfo', [
('data', ty.Optional[ty.Union['Point', 'Segment']]),
('type', IntersectionType),
])
NOT_INTERSECTED = IntersectionInfo(None, IntersectionType.NONE) # type: ty_ext.Final[IntersectionInfo]
"""The constant for cases when the intersection does not exist"""
class SegmentsIntersection:
"""The class represents the intersection of two segments
Parameters
----------
segment1 : Segment
The first segment object
segment2 : Segment
The second segment object
intersect_info : IntersectionInfo
The intersection info object
"""
__slots__ = ('_segment1', '_segment2', '_intersect_info')
def __init__(self, segment1: 'Segment', segment2: 'Segment',
intersect_info: IntersectionInfo) -> None:
self._segment1 = segment1
self._segment2 = segment2
self._intersect_info = intersect_info
def __repr__(self) -> str:
return '{}({}, {}, {}, type={})'.format(
type(self).__name__,
self.segment1,
self.segment2,
self.intersect_point,
self.intersect_type.name,
)
def __bool__(self) -> bool:
return self.intersect_type != IntersectionType.NONE
@property
def segment1(self) -> 'Segment':
"""The first segment
Returns
-------
segment : Segment
The first segment that intersects the second segment
"""
return self._segment1
@property
def segment2(self) -> 'Segment':
"""The second segment
Returns
-------
segment : Segment
The second segment that intersects the first segment
"""
return self._segment2
@property
def intersect_info(self) -> IntersectionInfo:
"""Returns the information about intersection
Returns
-------
info : IntersectionInfo
Intersection info named tuple ``(data, type)`` where:
- ``data`` is None or Point or Segment
- ``type`` intersection type (IntersectionType) NONE/EXACT/OVERLAP/ALMOST
"""
return self._intersect_info
@property
def intersect_type(self) -> IntersectionType:
"""Returns the type of intersection
Returns
-------
type : IntersectionType
Intersection type enum item (NONE/EXACT/OVERLAP/ALMOST)
"""
return self._intersect_info.type
@property
def intersect_point(self) -> ty.Optional['Point']:
"""Returns the intersection point
Returns
-------
point : Point, None
The intersection point or None if the intersection does not exist
Notes
-----
If the intersection type is OVERLAP or ALMOST will be returned
``intersect_segment.point(t=0.5)`` as intersection point.
See Also
--------
intersect_segment
"""
if not self:
return None
if self.intersect_type == IntersectionType.EXACT:
return self._intersect_info.data
else:
return self._intersect_info.data.point(0.5)
@property
def intersect_segment(self) -> ty.Optional['Segment']:
"""Returns the segment if the segments are overlapped or almost intersected
Returns
-------
segment : Segment, None
Overlapping or connecting segment if the segments are overlapped or
almost intersected. None if the intersection does not exist.
Notes
-----
If the intersection type is EXACT a singular segment will be returned.
See Also
--------
intersect_point
"""
if not self:
return None
if self.intersect_type == IntersectionType.EXACT:
return skcurve._base.Segment(self._intersect_info.data, self._intersect_info.data)
else:
return self._intersect_info.data
class IntersectionMethodBase(abc.ABC):
"""The base class for all intersection methods
"""
def __call__(self,
obj1: ty.Union['Segment', 'Curve'], obj2: ty.Union['Segment', 'Curve']) \
-> ty.Union[SegmentsIntersection, ty.List[SegmentsIntersection]]:
valid_obj_types = (skcurve._base.Segment, skcurve._base.Curve)
if not isinstance(obj1, valid_obj_types) or not isinstance(obj2, valid_obj_types):
raise TypeError('"obj1" and "obj2" arguments must be \'Segment\' or \'Curve\'.')
if obj1.ndim != obj2.ndim:
raise ValueError('The dimension of both objects must be equal.')
if isinstance(obj1, skcurve._base.Segment) and isinstance(obj2, skcurve._base.Segment):
intersect_info = self._intersect_segments(obj1, obj2)
return SegmentsIntersection(
segment1=obj1,
segment2=obj2,
intersect_info=intersect_info,
)
elif isinstance(obj1, skcurve._base.Curve) and isinstance(obj2, skcurve._base.Curve):
if obj1.size == 0 or obj2.size == 0:
return []
return self._intersect_curves(obj1, obj2)
else:
# Intersections between the curve and the segment
obj1_is_segment = isinstance(obj1, skcurve._base.Segment)
obj2_is_segment = isinstance(obj2, skcurve._base.Segment)
curve1 = ty.cast(skcurve._base.Segment, obj1).to_curve() if obj1_is_segment else obj1
curve2 = ty.cast(skcurve._base.Segment, obj2).to_curve() if obj2_is_segment else obj2
if curve1.size == 0 or curve2.size == 0:
return []
intersections = self._intersect_curves(curve1, curve2)
for i, intersection in enumerate(intersections):
intersections[i] = SegmentsIntersection(
segment1=ty.cast(skcurve._base.Segment, obj1) if obj1_is_segment else intersection.segment1,
segment2=ty.cast(skcurve._base.Segment, obj2) if obj2_is_segment else intersection.segment2,
intersect_info=intersection.intersect_info,
)
return intersections
@staticmethod
def _curves_intersect_indices(intersect_matrix: np.ndarray, self_intersect: bool) -> \
ty.Tuple[np.ndarray, np.ndarray]:
"""Computes the segments indices for curves intersection matrix
Parameters
----------
intersect_matrix : np.ndarray
The curves intersection boolean matrix: M1xM2 array (MxM array for self intersection)
self_intersect : bool
The flag defines the curve self-intersection
Returns
-------
s1 : np.ndarray
The segments indices for the first curve (or the first segments when self interesection)
s2 : np.ndarray
The segments indices for the second curve (or the second segments when self interesection)
"""
if self_intersect:
# Removing duplicate combinations of segments when self intersection.
i, j = np.tril_indices(intersect_matrix.shape[0], k=0)
intersect_matrix[i, j] = False
s1, s2 = np.nonzero(intersect_matrix)
if self_intersect:
# Removing coincident and adjacent segments
adjacent = np.abs(s1 - s2) < 2
s1 = s1[~adjacent]
s2 = s2[~adjacent]
return s1, s2
@abc.abstractmethod
def _intersect_segments(self, segment1: 'Segment', segment2: 'Segment') -> IntersectionInfo:
"""Should implement segments intersection algorithm
"""
@abc.abstractmethod
def _intersect_curves(self, curve1: 'Curve', curve2: 'Curve') -> ty.List[SegmentsIntersection]:
"""Should implement curves intersection algorithm
"""
def intersect_methods() -> ty.List[str]:
"""Returns the list of available intersect methods
Returns
-------
methods : List[str]
The list of available intersect methods
See Also
--------
get_intersect_method
register_intersect_method
"""
return list(_intersect_methods.keys())
def get_intersect_method(method: str, **params) -> 'IntersectionMethodBase':
"""Returns the intersection method callable for the given method name
Parameters
----------
method : str
Intersection method name
params : mapping
The method parameters
Returns
-------
intersect : IntersectionMethodBase
Intersection method class
See Also
--------
intersect_methods
register_intersect_method
Raises
------
NameError : If intersect method is unknown
"""
if method not in _intersect_methods:
raise NameError(
f"Unknown method '{method}'. The following methods are available: {intersect_methods()}")
return _intersect_methods[method](**params)
def default_intersect_method() -> str:
"""Returns default intersect method
Returns
-------
method : str
Default intersect method
"""
return _default_intersect_method
def set_default_intersect_method(method: str) -> None:
"""Sets the given intersect method as default
Parameters
----------
method : str
Method name
See Also
--------
default_intersect_method
register_intersect_method
"""
global _default_intersect_method
if method not in _intersect_methods:
raise NameError(
f"Unknown method '{method}'. The following methods are available: {intersect_methods()}")
_default_intersect_method = method
def register_intersect_method(method: str, default: bool = False):
"""Decorator for registering segment intersection methods
The decorator can be used for registering new intersection methods.
The intersection method should be callable and implement `IntersectionMethod` protocol.
Parameters
----------
method : str
Method name
default : bool
Makes registered method as default
See Also
--------
IntersectionMethod
intersect_methods
get_intersect_method
"""
def decorator(cls: ty.Type[IntersectionMethodBase]):
if method in _intersect_methods:
raise ValueError(f"'{method}' intersect method already registered for {_intersect_methods[method]}")
if not issubclass(cls, IntersectionMethodBase):
raise TypeError(f"{cls} is not a subclass of 'IntersectionMethodBase'")
_intersect_methods[method] = cls
if default:
set_default_intersect_method(method)
return decorator
@register_intersect_method(method='exact', default=True)
class ExactIntersectionMethod(IntersectionMethodBase):
"""The method to determine the exact segments and curves intersection
We should solve the linear system of the following equations:
x1 + t1 * (x2 - x1) = x3 + t2 * (x4 - x3)
y1 + t1 * (y2 - y1) = y3 + t2 * (y4 - y3)
...
n1 + t1 * (n2 - n3) = n3 + t2 * (n4 - n3)
The solution of this system is t1 and t2 parameter values.
If t1 and t2 in the range [0, 1], the segments are intersect.
If the coefficient matrix is non-symmetric (for n-dim > 2),
it requires a solver for over-determined system.
Parameters
----------
feps : float
Floating point epsilon. F_EPS by default
"""
def __init__(self, feps: float = F_EPS):
self._feps = feps
def _intersect_segments(self, segment1: 'Segment', segment2: 'Segment') -> IntersectionInfo:
# Firstly, we should check all corner cases (overlap, parallel, not coplanar, singular...).
if segment1.collinear(segment2):
# We return overlap segment because we do not know exactly what point needed in this case.
overlap_segment = segment1.overlap(segment2)
if overlap_segment is None:
return NOT_INTERSECTED
return IntersectionType.OVERLAP(overlap_segment)
if segment1.parallel(segment2):
return NOT_INTERSECTED
if not segment1.coplanar(segment2):
return NOT_INTERSECTED
if segment1.singular or segment2.singular:
return NOT_INTERSECTED
# After checking all corner cases we are sure that
# two segments (or lines) should intersected.
a = np.stack((segment1.direction.data,
-segment2.direction.data), axis=1)
b = (segment2.p1 - segment1.p1).data
if segment1.ndim == 2:
try:
t = np.linalg.solve(a, b)
except np.linalg.LinAlgError as err:
warnings.warn(f'Cannot solve system of equations: {err}', IntersectionWarning)
return NOT_INTERSECTED
else:
t, residuals, *_ = np.linalg.lstsq(a, b, rcond=None)
if residuals.size > 0 and residuals[0] > self._feps:
warnings.warn(
f"The 'lstsq' residuals are {residuals} > {self._feps}. Computation result might be wrong.",
IntersectionWarning)
if np.all(((t > 0) | np.isclose(t, 0)) &
((t < 1) | np.isclose(t, 1))):
intersect_point1 = segment1.point(t[0])
intersect_point2 = segment2.point(t[1])
if intersect_point1 != intersect_point2:
distance = intersect_point1.distance(intersect_point2)
if distance > self._feps:
warnings.warn(
f"Incorrect solution. The points for 't1' and 't2' are different (distance: {distance}).",
IntersectionWarning)
return NOT_INTERSECTED
return IntersectionType.EXACT(intersect_point1)
return NOT_INTERSECTED
def _intersect_curves(self, curve1: 'Curve', curve2: 'Curve') -> ty.List[SegmentsIntersection]:
s1, s2 = self._find_segments_bbox_intersection(curve1, curve2)
if s1.size == 0:
return []
intersections = []
for segment1, segment2 in zip(curve1.segments[s1], curve2.segments[s2]):
intersect_info = self._intersect_segments(segment1, segment2)
if intersect_info.type != IntersectionType.NONE:
intersections.append(SegmentsIntersection(
segment1=segment1,
segment2=segment2,
intersect_info=intersect_info,
))
return intersections
def _find_segments_bbox_intersection(self, curve1: 'Curve', curve2: 'Curve') \
-> ty.Tuple[np.ndarray, np.ndarray]:
"""Finds intersections between axis-aligned bounding boxes (AABB) of curves segments
`Curve` 1 and `Curve` 2 can be different objects or the same objects (self intersection).
"""
self_intersect = curve2 is curve1
if curve1.size == 0 or curve2.size == 0:
return (
np.array([], dtype=np.int64),
np.array([], dtype=np.int64),
)
# Get beginning and ending points of segments
p11 = curve1.data[:-1, :]
p12 = curve1.data[1:, :]
curve1_pmin = np.minimum(p11, p12)
curve1_pmax = np.maximum(p11, p12)
if self_intersect:
curve2_pmin = curve1_pmin
curve2_pmax = curve1_pmax
else:
p21 = curve2.data[:-1, :]
p22 = curve2.data[1:, :]
curve2_pmin =
|
np.minimum(p21, p22)
|
numpy.minimum
|
# Licensed to the Apache Software Foundation (ASF) under one
# or more contributor license agreements. See the NOTICE file
# distributed with this work for additional information
# regarding copyright ownership. The ASF licenses this file
# to you under the Apache License, Version 2.0 (the
# "License"); you may not use this file except in compliance
# with the License. You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing,
# software distributed under the License is distributed on an
# "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY
# KIND, either express or implied. See the License for the
# specific language governing permissions and limitations
# under the License.
"""Benchmark script for ImageNet models on mobile GPU.
see README.md for the usage and results of this script.
"""
import argparse
import numpy as np
import tvm
from tvm.contrib.util import tempdir
import tvm.contrib.graph_runtime as runtime
from tvm import relay
from util import get_network, print_progress
def evaluate_network(network, target, target_host, dtype, repeat):
# connect to remote device
tracker = tvm.rpc.connect_tracker(args.host, args.port)
remote = tracker.request(args.rpc_key)
print_progress(network)
net, params, input_shape, output_shape = get_network(network, batch_size=1, dtype=dtype)
print_progress("%-20s building..." % network)
with relay.build_config(opt_level=3):
graph, lib, params = relay.build(
net, target=target, target_host=target_host, params=params)
tmp = tempdir()
if 'android' in str(target) or 'android' in str(target_host):
from tvm.contrib import ndk
filename = "%s.so" % network
lib.export_library(tmp.relpath(filename), ndk.create_shared)
else:
filename = "%s.tar" % network
lib.export_library(tmp.relpath(filename))
# upload library and params
print_progress("%-20s uploading..." % network)
ctx = remote.context(str(target), 0)
remote.upload(tmp.relpath(filename))
rlib = remote.load_module(filename)
module = runtime.create(graph, rlib, ctx)
data_tvm = tvm.nd.array((
|
np.random.uniform(size=input_shape)
|
numpy.random.uniform
|
#!/usr/bin/env python
# -*- coding: utf-8 -*-
import numpy as np
from isochrones.dartmouth import Dartmouth_Isochrone
from isochrones import StarModel
import pandas as pd
import os
import kplr
import glob
from fgkcupid import teff2bv, age_model
from kepler_data import load_kepler_data
import acf
DATA_DIR = "data"
LC_DIR = "/Users/ruthangus/.kplr/data/lightcurves"
RESULTS_DIR = "results"
class star(object):
def __init__(self, id, mass=None, teff=None, logg=None, feh=None,
prot=None, BV=None, jmag=None, hmag=None, kmag=None,
Gmag=None, kepmag=None, parallax=None, gyro_age=None,
DATA_DIR=DATA_DIR, RESULTS_DIR=RESULTS_DIR, LC_DIR=LC_DIR,
download_lc=True):
"""
Routines for calculating the age of a single star. Currently only
suitable for Kepler targets.
PARAMETERS
----------
id: str
The id of the star. A kepler star will have a 9-digit integer.
mass: tuple (mass, mass_err) (optional)
Mass in Solar masses.
teff: tuple (teff, teff_err) (optional)
Effective temperature (K).
logg: tuple (logg, logg_err) (optional)
log (g).
feh: tuple (feh, feh_err) (optional)
Iron abundance.
prot: tuple (prot, prot_err) (optional)
The rotation period in days.
BV: tuple (B-V, B-V_err) (optional)
The B-V colour.
jmag: tuple (jmag, jmag_err) (optional)
The j-band magnitude.
hmag: tuple (hmag, hmag_err) (optional)
The h-band magnitude.
kmag: tuple (kmag, kmag_err) (optional)
The k-band magnitude.
Gmag: tuple (Gmag, Gmag_err) (optional)
The Gaia magnitude.
kepmag: tuple (kepmag, kepmag_err) (optional)
The Kepler magnitude.
parallax: tuple (parallax, parallax_err) (optional)
The Gaia parallax.
parallax: tuple (Gyro_age, gyro_age_err) (optional)
The gyro age.
DATA_DIR: str
The directory containing training data for the age model.
RESULTS_DIR: str
The directory for saving results.
LC_DIR: str
The directory containing the kepler light curves.
download_lc: bool
if True the Kepler light curve is downloaded and an ACF computed.
"""
assert type(id) == str, "ID must be a string"
assert len(id) < 10, "ID must be a 9-digit KIC id."
self.id = str(int(id)).zfill(9) # make sure id is in KIC format.
self.mass, self.teff, self.logg, self.feh = mass, teff, logg, feh
self.prot, self.BV, self.Gmag = prot, BV, Gmag
self.kepmag, self.parallax, self.gyro_age = kepmag, parallax, gyro_age
self.jmag, self.kmag, self.hmag = jmag, kmag, hmag
# KIC parameters.
if not self.teff or not self.logg or not self.feh or not self.kepmag:
print("Searching database for stellar parameters...")
data = pd.read_csv(os.path.join(DATA_DIR, "ruth_matched.csv"))
m = np.array(data["kepid"]) == int(self.id)
if len(np.array(data["kepid"][m])): # load from kepler-TGAS cat
id = np.array(data["kepid"])[m]
if not self.teff:
self.teff = (float(np.array(data["teff"])[m]),
float(.5*(np.array(data["teff_err1"])[m]) +
np.abs(float(np.array
(data["teff_err2"])[m]))))
if not self.feh:
self.feh = (float(np.array(data["feh"])[m]),
float(.5*(np.array(data["feh_err1"])[m]) +
np.abs(float(np.array
(data["feh_err2"])[m]))))
if not self.logg:
self.logg = (float(np.array(data["logg"])[m]),
float(.5*(np.array(data["logg_err1"])[m]) +
np.abs(float(np.array
(data["logg_err2"])[m]))))
if not self.kepmag:
self.kepmag = (float(np.array(data["kepmag"])[m]), .1)
if not self.jmag:
self.jmag = (float(np.array(data["jmag"])[m]),
float(np.array(data["jmag_err"])[m]))
if not self.hmag:
self.hmag = (float(
|
np.array(data["hmag"])
|
numpy.array
|
"""terrain.py
takes all terrain files (terrain/*) and converts them
into stickmanranger terrain objects.
NOTE: may want to run this code in a background thread,
as it will probably take a while and cause graphics
to crash.
"""
__author__ = '<NAME>'
__version__ = '0.0'
from pprint import pprint
import os
import sys
from pygame.surface import Surface
from pygame.transform import scale
from pygame.locals import QUIT
import numpy
try:
from _internal import *
from smr_error import SMRError
except ImportError:
from ._internal import *
from .smr_error import SMRError
VALID_COMMANDS = ('air', 'water', 'size')
# there once was a fellow named finn
# who threw all his legs in a bin
# he realized, at last
# he could not move so fast
# and punched himself right in the chin.
class Terrain:
built_image = None
top_water = PICS['Other']['top_water']
surface_symbol = '*'
ground_symbol = '#'
water_symbol = '-'
air_symbol = '~'
sign_symbol = '^'
pit_symbol = '_'
top_water_symbol = '+'
# alpha values. can be overriden with headers. (see flat.smr-terrain)
air = 200
water = 100
def_air = (0, 0, 0, 200)
def_water = (0, 50, 200, 100)
def __init__(self, image, template='flat', block_size=10, use_numpy=True):
self.image1 = PICS['terrain_templates'][image]['1']
self.image2 = PICS['terrain_templates'][image]['0']
self.template = template
self.size = block_size
self.use_numpy = use_numpy
try:
Terrain.terrain2dlist_texts
except AttributeError:
self.load_text()
def get_array(self):
"""find and return the proper array of terrain for the current object"""
return self.terrain2dlist_texts[self.template]['text']
def __iter__(self):
for i in self.terrain2dlist:
yield i
def __getitem__(self, pos):
arr = self.terrain2dlist_texts[self.template]['text']
return arr[pos[1]][pos[0]]
def __eq__(self, other):
return self.__dict__ == other.__dict__
def get_solid(self, pos):
"""return true if the block at pos is solid."""
return self.is_solid(self[pos])
@staticmethod
def is_solid(item):
return item in (Terrain.ground_symbol, Terrain.surface_symbol)
@staticmethod
def is_water(item):
return item in (Terrain.water_symbol, Terrain.top_water_symbol)
@staticmethod
def is_pit(item):
return item == Terrain.pit_symbol
@staticmethod
def is_air(item):
return item == Terrain.air_symbol
def load_text(self):
try:
Terrain.terrain2dlist_texts
except AttributeError:
Terrain.terrain2dlist_texts = {}
all_texts = Terrain.terrain2dlist_texts
terrain_texts = {}
terrain2dlist_texts = {}
for text in os.listdir(TDIR):
a = text.split('.')[0]
terrain_texts[a] = open(os.path.join(TDIR, text)).read()
for terrain, key in zip(terrain_texts.values(), terrain_texts.keys()):
main_dict = {
'size': self.size,
'air': self.def_air,
'water': self.def_water
}
if terrain.startswith('@'):
# remove @ symbol
header = terrain.split('\n')[0][1:]
terrain = '\n'.join(terrain.split('\n')[1:])
header = header.split('|')
# remove all whitespace
header = [part.strip().replace((' '), '')
.replace('\n', '')
.replace('\r', '')
.replace('\t', '')
for part in header]
for command in header:
parts = command.split('=')
print
if not parts[0] in ('air', 'water', 'size'):
raise SyntaxError(
'%a is not a valid command for header' % parts[0])
else:
main_dict[parts[0]] = eval(parts[1])
lines = []
for line in terrain.split('\n'):
if ';' in line:
line = line.split(';')[0].strip()
# dont append blank lines!
if line != '':
lines.append(line)
terrain2dlist = []
for line in lines:
chars = []
for char in line:
chars.append(char)
terrain2dlist.append(chars if not self.use_numpy
else numpy.array(chars))
main_dict['text'] = terrain2dlist if not self.use_numpy \
else
|
numpy.array(terrain2dlist)
|
numpy.array
|
import numpy as np
import string
import pickle
import os
def save_object(obj, path):
directory = os.path.dirname(path)
if not os.path.exists(directory):
os.makedirs(directory)
with open(path, "wb") as f:
pickle.dump(obj, f, protocol=pickle.HIGHEST_PROTOCOL)
with open("last_object_saved", "w") as f:
f.write(path)
def load_object(path):
with open(path, "rb") as f:
return pickle.load(f)
def flip_dict(map):
return {v: k for k, v in map.items()}
def encodings(text, ix_to_char):
if ix_to_char is None:
ix_to_char = dict(enumerate(set(text)))
char_to_ix = flip_dict(ix_to_char)
return char_to_ix, ix_to_char
def read_corpus(data_path):
with open(data_path, "r") as f:
return f.read()
def process_text(data_path, lower=True, remove_punctuation=False):
raw = read_corpus(data_path)
out = raw
if lower:
out = out.lower()
if remove_punctuation:
out = strip_punctuation(out)
return out
def prepare_numeric(processed_text, char_to_ix):
return [char_to_ix[ch] for ch in processed_text]
def data_split(numeric, num_sites, fraction):
num_phrases = len(numeric) // num_sites
num_train_phrases = round(num_phrases * fraction)
train_size = num_train_phrases * num_sites
train = np.array(numeric[:train_size]).reshape(num_train_phrases, num_sites)
cv =
|
np.array(numeric[train_size:2*train_size])
|
numpy.array
|
# -*- coding: utf-8 -*-
from __future__ import (absolute_import, division,
print_function)
import numpy as np
import os
from enterprise import constants as const
import pickle
import healpy as hp
from scipy.stats import skewnorm, truncnorm
from enterprise import constants as const
from PTMCMCSampler.PTMCMCSampler import PTSampler as ptmcmc
class JumpProposal(object):
def __init__(self, pta, snames=None, empirical_distr=None, f_stat_file=None):
"""Set up some custom jump proposals"""
self.params = pta.params
self.pnames = pta.param_names
self.ndim = sum(p.size or 1 for p in pta.params)
self.plist = [p.name for p in pta.params]
# parameter map
self.pmap = {}
ct = 0
for p in pta.params:
size = p.size or 1
self.pmap[str(p)] = slice(ct, ct+size)
ct += size
# parameter indices map
self.pimap = {}
for ct, p in enumerate(pta.param_names):
self.pimap[p] = ct
# collecting signal parameters across pta
if snames is None:
allsigs = np.hstack([[qq.signal_name for qq in pp._signals]
for pp in pta._signalcollections])
self.snames = dict.fromkeys(np.unique(allsigs))
for key in self.snames: self.snames[key] = []
for sc in pta._signalcollections:
for signal in sc._signals:
self.snames[signal.signal_name].extend(signal.params)
for key in self.snames: self.snames[key] = list(set(self.snames[key]))
else:
self.snames = snames
# empirical distributions
if empirical_distr is not None and os.path.isfile(empirical_distr):
try:
with open(empirical_distr, 'rb') as f:
pickled_distr = pickle.load(f)
except:
try:
with open(empirical_distr, 'rb') as f:
pickled_distr = pickle.load(f)
except:
print('I can\'t open the empirical distribution pickle file!')
pickled_distr = None
self.empirical_distr = pickled_distr
elif isinstance(empirical_distr,list):
pass
else:
self.empirical_distr = None
if self.empirical_distr is not None:
# only save the empirical distributions for parameters that are in the model
mask = []
for idx,d in enumerate(self.empirical_distr):
if d.ndim == 1:
if d.param_name in pta.param_names:
mask.append(idx)
else:
if d.param_names[0] in pta.param_names and d.param_names[1] in pta.param_names:
mask.append(idx)
if len(mask) > 1:
self.empirical_distr = [self.empirical_distr[m] for m in mask]
else:
self.empirical_distr = None
#F-statistic map
if f_stat_file is not None and os.path.isfile(f_stat_file):
npzfile = np.load(f_stat_file)
self.fe_freqs = npzfile['freqs']
self.fe = npzfile['fe']
def draw_from_prior(self, x, iter, beta):
"""Prior draw.
The function signature is specific to PTMCMCSampler.
"""
q = x.copy()
lqxy = 0
# randomly choose parameter
param = np.random.choice(self.params)
# if vector parameter jump in random component
if param.size:
idx2 = np.random.randint(0, param.size)
q[self.pmap[str(param)]][idx2] = param.sample()[idx2]
# scalar parameter
else:
q[self.pmap[str(param)]] = param.sample()
# forward-backward jump probability
lqxy = (param.get_logpdf(x[self.pmap[str(param)]]) -
param.get_logpdf(q[self.pmap[str(param)]]))
return q, float(lqxy)
def draw_from_red_prior(self, x, iter, beta):
q = x.copy()
lqxy = 0
signal_name = 'red noise'
# draw parameter from signal model
param = np.random.choice(self.snames[signal_name])
if param.size:
idx2 = np.random.randint(0, param.size)
q[self.pmap[str(param)]][idx2] = param.sample()[idx2]
# scalar parameter
else:
q[self.pmap[str(param)]] = param.sample()
# forward-backward jump probability
lqxy = (param.get_logpdf(x[self.pmap[str(param)]]) -
param.get_logpdf(q[self.pmap[str(param)]]))
return q, float(lqxy)
def draw_from_empirical_distr(self, x, iter, beta):
q = x.copy()
lqxy = 0
if self.empirical_distr is not None:
# randomly choose one of the empirical distributions
distr_idx = np.random.randint(0, len(self.empirical_distr))
if self.empirical_distr[distr_idx].ndim == 1:
idx = self.pnames.index(self.empirical_distr[distr_idx].param_name)
q[idx] = self.empirical_distr[distr_idx].draw()
lqxy = (self.empirical_distr[distr_idx].logprob(x[idx]) -
self.empirical_distr[distr_idx].logprob(q[idx]))
else:
oldsample = [x[self.pnames.index(p)]
for p in self.empirical_distr[distr_idx].param_names]
newsample = self.empirical_distr[distr_idx].draw()
for p,n in zip(self.empirical_distr[distr_idx].param_names, newsample):
q[self.pnames.index(p)] = n
lqxy = (self.empirical_distr[distr_idx].logprob(oldsample) -
self.empirical_distr[distr_idx].logprob(newsample))
return q, float(lqxy)
def draw_from_dm_gp_prior(self, x, iter, beta):
q = x.copy()
lqxy = 0
signal_name = 'dm_gp'
# draw parameter from signal model
param = np.random.choice(self.snames[signal_name])
if param.size:
idx2 = np.random.randint(0, param.size)
q[self.pmap[str(param)]][idx2] = param.sample()[idx2]
# scalar parameter
else:
q[self.pmap[str(param)]] = param.sample()
# forward-backward jump probability
lqxy = (param.get_logpdf(x[self.pmap[str(param)]]) -
param.get_logpdf(q[self.pmap[str(param)]]))
return q, float(lqxy)
def draw_from_dm1yr_prior(self, x, iter, beta):
q = x.copy()
lqxy = 0
dm1yr_names = [dmname for dmname in self.pnames if 'dm_s1yr' in dmname]
dmname = np.random.choice(dm1yr_names)
idx = self.pnames.index(dmname)
if 'log10_Amp' in dmname:
q[idx] = np.random.uniform(-10, -2)
elif 'phase' in dmname:
q[idx] = np.random.uniform(0, 2*np.pi)
return q, 0
def draw_from_dmexpdip_prior(self, x, iter, beta):
q = x.copy()
lqxy = 0
dmexp_names = [dmname for dmname in self.pnames if 'dmexp' in dmname]
dmname = np.random.choice(dmexp_names)
idx = self.pnames.index(dmname)
if 'log10_Amp' in dmname:
q[idx] = np.random.uniform(-10, -2)
elif 'log10_tau' in dmname:
q[idx] = np.random.uniform(0, 2.5)
elif 'sign_param' in dmname:
q[idx] = np.random.uniform(-1.0, 1.0)
return q, 0
def draw_from_dmexpcusp_prior(self, x, iter, beta):
q = x.copy()
lqxy = 0
dmexp_names = [dmname for dmname in self.pnames if 'dm_cusp' in dmname]
dmname = np.random.choice(dmexp_names)
idx = self.pnames.index(dmname)
if 'log10_Amp' in dmname:
q[idx] = np.random.uniform(-10, -2)
elif 'log10_tau' in dmname:
q[idx] = np.random.uniform(0, 2.5)
#elif 't0' in dmname:
# q[idx] = np.random.uniform(53393.0, 57388.0)
elif 'sign_param' in dmname:
q[idx] = np.random.uniform(-1.0, 1.0)
return q, 0
def draw_from_dmx_prior(self, x, iter, beta):
q = x.copy()
lqxy = 0
signal_name = 'dmx_signal'
# draw parameter from signal model
param = np.random.choice(self.snames[signal_name])
if param.size:
idx2 = np.random.randint(0, param.size)
q[self.pmap[str(param)]][idx2] = param.sample()[idx2]
# scalar parameter
else:
q[self.pmap[str(param)]] = param.sample()
# forward-backward jump probability
lqxy = (param.get_logpdf(x[self.pmap[str(param)]]) -
param.get_logpdf(q[self.pmap[str(param)]]))
return q, float(lqxy)
def draw_from_gwb_log_uniform_distribution(self, x, iter, beta):
q = x.copy()
lqxy = 0
# draw parameter from signal model
idx = self.pnames.index('gw_log10_A')
q[idx] = np.random.uniform(-18, -11)
return q, 0
def draw_from_dipole_log_uniform_distribution(self, x, iter, beta):
q = x.copy()
lqxy = 0
# draw parameter from signal model
idx = self.pnames.index('dipole_log10_A')
q[idx] = np.random.uniform(-18, -11)
return q, 0
def draw_from_monopole_log_uniform_distribution(self, x, iter, beta):
q = x.copy()
lqxy = 0
# draw parameter from signal model
idx = self.pnames.index('monopole_log10_A')
q[idx] = np.random.uniform(-18, -11)
return q, 0
def draw_from_altpol_log_uniform_distribution(self, x, iter, beta):
q = x.copy()
lqxy = 0
# draw parameter from signal model
polnames = [pol for pol in self.pnames if 'log10Apol' in pol]
if 'kappa' in self.pnames:
polnames.append('kappa')
pol = np.random.choice(polnames)
idx = self.pnames.index(pol)
if pol == 'log10Apol_tt':
q[idx] = np.random.uniform(-18, -12)
elif pol == 'log10Apol_st':
q[idx] = np.random.uniform(-18, -12)
elif pol == 'log10Apol_vl':
q[idx] = np.random.uniform(-18, -15)
elif pol == 'log10Apol_sl':
q[idx] = np.random.uniform(-18, -16)
elif pol == 'kappa':
q[idx] = np.random.uniform(0, 10)
return q, 0
def draw_from_ephem_prior(self, x, iter, beta):
q = x.copy()
lqxy = 0
signal_name = 'phys_ephem'
# draw parameter from signal model
param = np.random.choice(self.snames[signal_name])
if param.size:
idx2 = np.random.randint(0, param.size)
q[self.pmap[str(param)]][idx2] = param.sample()[idx2]
# scalar parameter
else:
q[self.pmap[str(param)]] = param.sample()
# forward-backward jump probability
lqxy = (param.get_logpdf(x[self.pmap[str(param)]]) -
param.get_logpdf(q[self.pmap[str(param)]]))
return q, float(lqxy)
def draw_from_bwm_prior(self, x, iter, beta):
q = x.copy()
lqxy = 0
signal_name = 'bwm'
# draw parameter from signal model
param = np.random.choice(self.snames[signal_name])
if param.size:
idx2 = np.random.randint(0, param.size)
q[self.pmap[str(param)]][idx2] = param.sample()[idx2]
# scalar parameter
else:
q[self.pmap[str(param)]] = param.sample()
# forward-backward jump probability
lqxy = (param.get_logpdf(x[self.pmap[str(param)]]) -
param.get_logpdf(q[self.pmap[str(param)]]))
return q, float(lqxy)
def draw_from_cw_prior(self, x, iter, beta):
q = x.copy()
lqxy = 0
signal_name = 'cw'
# draw parameter from signal model
param = np.random.choice(self.snames[signal_name])
if param.size:
idx2 = np.random.randint(0, param.size)
q[self.pmap[str(param)]][idx2] = param.sample()[idx2]
# scalar parameter
else:
q[self.pmap[str(param)]] = param.sample()
# forward-backward jump probability
lqxy = (param.get_logpdf(x[self.pmap[str(param)]]) -
param.get_logpdf(q[self.pmap[str(param)]]))
return q, float(lqxy)
def draw_from_cw_log_uniform_distribution(self, x, iter, beta):
q = x.copy()
lqxy = 0
# draw parameter from signal model
idx = self.pnames.index('log10_h')
q[idx] = np.random.uniform(-18, -11)
return q, 0
def draw_from_dm_sw_prior(self, x, iter, beta):
q = x.copy()
lqxy = 0
signal_name = 'gp_sw'
# draw parameter from signal model
param = np.random.choice(self.snames[signal_name])
if param.size:
idx2 = np.random.randint(0, param.size)
q[self.pmap[str(param)]][idx2] = param.sample()[idx2]
# scalar parameter
else:
q[self.pmap[str(param)]] = param.sample()
# forward-backward jump probability
lqxy = (param.get_logpdf(x[self.pmap[str(param)]]) -
param.get_logpdf(q[self.pmap[str(param)]]))
return q, float(lqxy)
def draw_from_signal_prior(self, x, iter, beta):
q = x.copy()
lqxy = 0
std = ['linear timing model',
'red noise',
'phys_ephem',
'gw',
'cw',
'bwm',
'gp_sw',
'ecorr_sherman-morrison',
'ecorr',
'efac',
'equad',
]
non_std = [nm for nm in self.snames.keys() if nm not in std]
# draw parameter from signal model
signal_name = np.random.choice(non_std)
param = np.random.choice(self.snames[signal_name])
if param.size:
idx2 = np.random.randint(0, param.size)
q[self.pmap[str(param)]][idx2] = param.sample()[idx2]
# scalar parameter
else:
q[self.pmap[str(param)]] = param.sample()
# forward-backward jump probability
lqxy = (param.get_logpdf(x[self.pmap[str(param)]]) -
param.get_logpdf(q[self.pmap[str(param)]]))
return q, float(lqxy)
def draw_from_par_prior(self, par_names):
# Preparing and comparing par_names with PTA parameters
par_names = np.atleast_1d(par_names)
par_list = []
name_list = []
for par_name in par_names:
pn_list = [n for n in self.plist if par_name in n]
if pn_list:
par_list.append(pn_list)
name_list.append(par_name)
if not par_list:
raise UserWarning("No parameter prior match found between {} and PTA.object."
.format(par_names))
par_list = np.concatenate(par_list,axis=None)
def draw(x, iter, beta):
"""Prior draw function generator for custom par_names.
par_names: list of strings
The function signature is specific to PTMCMCSampler.
"""
q = x.copy()
lqxy = 0
# randomly choose parameter
idx_name = np.random.choice(par_list)
idx = self.plist.index(idx_name)
# if vector parameter jump in random component
param = self.params[idx]
if param.size:
idx2 = np.random.randint(0, param.size)
q[self.pmap[str(param)]][idx2] = param.sample()[idx2]
# scalar parameter
else:
q[self.pmap[str(param)]] = param.sample()
# forward-backward jump probability
lqxy = (param.get_logpdf(x[self.pmap[str(param)]]) -
param.get_logpdf(q[self.pmap[str(param)]]))
return q, float(lqxy)
name_string = '_'.join(name_list)
draw.__name__ = 'draw_from_{}_prior'.format(name_string)
return draw
def draw_from_par_log_uniform(self, par_dict):
# Preparing and comparing par_dict.keys() with PTA parameters
par_list = []
name_list = []
for par_name in par_dict.keys():
pn_list = [n for n in self.plist if par_name in n and 'log' in n]
if pn_list:
par_list.append(pn_list)
name_list.append(par_name)
if not par_list:
raise UserWarning("No parameter dictionary match found between {} and PTA.object."
.format(par_dict.keys()))
par_list = np.concatenate(par_list,axis=None)
def draw(x, iter, beta):
"""log uniform prior draw function generator for custom par_names.
par_dict: dictionary with {"par_names":(lower bound,upper bound)}
{ "string":(float,float)}
The function signature is specific to PTMCMCSampler.
"""
q = x.copy()
lqxy = 0
# draw parameter from signal model
idx_name = np.random.choice(par_list)
idx = self.plist.index(idx_name)
q[idx] = np.random.uniform(par_dict[par_name][0],par_dict[par_name][1])
return q, 0
name_string = '_'.join(name_list)
draw.__name__ = 'draw_from_{}_log_uniform'.format(name_string)
return draw
def draw_from_signal(self, signal_names):
# Preparing and comparing signal_names with PTA signals
signal_names = np.atleast_1d(signal_names)
signal_list = []
name_list = []
for signal_name in signal_names:
try:
param_list = self.snames[signal_name]
signal_list.append(param_list)
name_list.append(signal_name)
except:
pass
if not signal_list:
raise UserWarning("No signal match found between {} and PTA.object!"
.format(signal_names))
signal_list = np.concatenate(signal_list,axis=None)
def draw(x, iter, beta):
"""Signal draw function generator for custom signal_names.
signal_names: list of strings
The function signature is specific to PTMCMCSampler.
"""
q = x.copy()
lqxy = 0
# draw parameter from signal model
param = np.random.choice(signal_list)
if param.size:
idx2 = np.random.randint(0, param.size)
q[self.pmap[str(param)]][idx2] = param.sample()[idx2]
# scalar parameter
else:
q[self.pmap[str(param)]] = param.sample()
# forward-backward jump probability
lqxy = (param.get_logpdf(x[self.pmap[str(param)]]) -
param.get_logpdf(q[self.pmap[str(param)]]))
return q, float(lqxy)
name_string = '_'.join(name_list)
draw.__name__ = 'draw_from_{}_signal'.format(name_string)
return draw
def fe_jump(self, x, iter, beta):
q = x.copy()
lqxy = 0
fe_limit = np.max(self.fe)
#draw skylocation and frequency from f-stat map
accepted = False
while accepted==False:
log_f_new = self.params[self.pimap['log10_fgw']].sample()
f_idx = (np.abs(np.log10(self.fe_freqs) - log_f_new)).argmin()
gw_theta = np.arccos(self.params[self.pimap['cos_gwtheta']].sample())
gw_phi = self.params[self.pimap['gwphi']].sample()
hp_idx = hp.ang2pix(hp.get_nside(self.fe), gw_theta, gw_phi)
fe_new_point = self.fe[f_idx, hp_idx]
if np.random.uniform()<(fe_new_point/fe_limit):
accepted = True
#draw other parameters from prior
cos_inc = self.params[self.pimap['cos_inc']].sample()
psi = self.params[self.pimap['psi']].sample()
phase0 = self.params[self.pimap['phase0']].sample()
log10_h = self.params[self.pimap['log10_h']].sample()
#put new parameters into q
signal_name = 'cw'
for param_name, new_param in zip(['log10_fgw','gwphi','cos_gwtheta','cos_inc','psi','phase0','log10_h'],
[log_f_new, gw_phi, np.cos(gw_theta), cos_inc, psi, phase0, log10_h]):
q[self.pimap[param_name]] = new_param
#calculate Hastings ratio
log_f_old = x[self.pimap['log10_fgw']]
f_idx_old = (np.abs(np.log10(self.fe_freqs) - log_f_old)).argmin()
gw_theta_old = np.arccos(x[self.pimap['cos_gwtheta']])
gw_phi_old = x[self.pimap['gwphi']]
hp_idx_old = hp.ang2pix(hp.get_nside(self.fe), gw_theta_old, gw_phi_old)
fe_old_point = self.fe[f_idx_old, hp_idx_old]
if fe_old_point>fe_limit:
fe_old_point = fe_limit
log10_h_old = x[self.pimap['log10_h']]
phase0_old = x[self.pimap['phase0']]
psi_old = x[self.pimap['psi']]
cos_inc_old = x[self.pimap['cos_inc']]
hastings_extra_factor = self.params[self.pimap['log10_h']].get_pdf(log10_h_old)
hastings_extra_factor *= 1/self.params[self.pimap['log10_h']].get_pdf(log10_h)
hastings_extra_factor = self.params[self.pimap['phase0']].get_pdf(phase0_old)
hastings_extra_factor *= 1/self.params[self.pimap['phase0']].get_pdf(phase0)
hastings_extra_factor = self.params[self.pimap['psi']].get_pdf(psi_old)
hastings_extra_factor *= 1/self.params[self.pimap['psi']].get_pdf(psi)
hastings_extra_factor = self.params[self.pimap['cos_inc']].get_pdf(cos_inc_old)
hastings_extra_factor *= 1/self.params[self.pimap['cos_inc']].get_pdf(cos_inc)
lqxy = np.log(fe_old_point/fe_new_point * hastings_extra_factor)
return q, float(lqxy)
class JumpProposalCW(object):
def __init__(self, pta, fgw=8e-9,psr_dist = None, snames=None, empirical_distr=None, f_stat_file=None):
"""Set up some custom jump proposals"""
self.params = pta.params
self.pnames = pta.param_names
self.ndim = sum(p.size or 1 for p in pta.params)
self.plist = [p.name for p in pta.params]
# parameter map
self.pmap = {}
ct = 0
for p in pta.params:
size = p.size or 1
self.pmap[str(p)] = slice(ct, ct+size)
ct += size
# parameter indices map
self.pimap = {}
for ct, p in enumerate(pta.param_names):
self.pimap[p] = ct
# collecting signal parameters across pta
if snames is None:
allsigs = np.hstack([[qq.signal_name for qq in pp._signals]
for pp in pta._signalcollections])
self.snames = dict.fromkeys(np.unique(allsigs))
for key in self.snames: self.snames[key] = []
for sc in pta._signalcollections:
for signal in sc._signals:
self.snames[signal.signal_name].extend(signal.params)
for key in self.snames: self.snames[key] = list(set(self.snames[key]))
else:
self.snames = snames
self.fgw = fgw
self.psr_dist = psr_dist
# empirical distributions
if empirical_distr is not None and os.path.isfile(empirical_distr):
try:
with open(empirical_distr, 'rb') as f:
pickled_distr = pickle.load(f)
except:
try:
with open(empirical_distr, 'rb') as f:
pickled_distr = pickle.load(f)
except:
print('I can\'t open the empirical distribution pickle file!')
pickled_distr = None
self.empirical_distr = pickled_distr
elif isinstance(empirical_distr,list):
pass
else:
self.empirical_distr = None
if self.empirical_distr is not None:
# only save the empirical distributions for parameters that are in the model
mask = []
for idx,d in enumerate(self.empirical_distr):
if d.ndim == 1:
if d.param_name in pta.param_names:
mask.append(idx)
else:
if d.param_names[0] in pta.param_names and d.param_names[1] in pta.param_names:
mask.append(idx)
if len(mask) > 1:
self.empirical_distr = [self.empirical_distr[m] for m in mask]
else:
self.empirical_distr = None
#F-statistic map
if f_stat_file is not None and os.path.isfile(f_stat_file):
npzfile = np.load(f_stat_file)
self.fe_freqs = npzfile['freqs']
self.fe = npzfile['fe']
def draw_from_many_par_prior(self, par_names, string_name):
# Preparing and comparing par_names with PTA parameters
par_names = np.atleast_1d(par_names)
par_list = []
name_list = []
for par_name in par_names:
pn_list = [n for n in self.plist if par_name in n]
if pn_list:
par_list.append(pn_list)
name_list.append(par_name)
if not par_list:
raise UserWarning("No parameter prior match found between {} and PTA.object."
.format(par_names))
par_list =
|
np.concatenate(par_list,axis=None)
|
numpy.concatenate
|
from numba import jit
import time
from project_code.misc_functions import sub_matrix, combine_sets
from project_code.classes import Result_IF, Result_IF_generators
import numpy as np
import logging
def compute_IFs(branches, setI, setT, setR, LODF, PATL, PTDF):
t0 = time.clock()
results = []
sizeI = len(setI)
sizeT = len(setT)
current_ring = 1
setR_this_ring = [branch for branch in setR if branch.ring == current_ring]
while len(setR_this_ring) > 0:
sizeR = len(setR_this_ring)
logging.info(f"Assessing IF for ring # {current_ring} with {sizeR} elements.")
set_size_RIT = np.array([sizeR, sizeI, sizeT], dtype=np.int32)
vPTDF_I = [i.PTDF for i in setI]
vPTDF_R = [r.PTDF for r in setR_this_ring]
mxPTDF_IR = sub_matrix(setI, setR_this_ring, PTDF)
mxPTDF_IT = sub_matrix(setI, setT, PTDF)
mxPTDF_RI = sub_matrix(setR_this_ring, setI, PTDF)
mxPTDF_RT = sub_matrix(setR_this_ring, setT, PTDF)
res_T = np.zeros((sizeI, sizeR), dtype=np.int32) # Most influenced t element in N-i-r
res_IF = np.zeros((sizeI, sizeR)) # IF of the most influenced t element in N-i-r situation
set_IR = combine_sets(setI, setR_this_ring) # elms i in R set to avoid i = r situation
set_RT = combine_sets(setR_this_ring, setT) # elms r in T set to avoid r = t situation
set_TI = combine_sets(setT, setI)
mxPATL_RT = sub_matrix(setR_this_ring, setT, PATL)
res_norm_T = np.zeros((sizeI, sizeR), dtype=np.int32) # same but normalized
res_norm_IF = np.zeros((sizeI, sizeR)) # same but normalized
res_norm_IF_non_norm = np.zeros((sizeI, sizeR))
res_T_max = np.zeros(sizeR, dtype=np.int32) # most influenced t element
res_norm_T_max = np.zeros(sizeR, dtype=np.int32) # same but normalized
res_I_max = np.zeros(sizeR, dtype=np.int32)
res_norm_I_max = np.zeros(sizeR, dtype=np.int32)
res_IF_max = np.zeros(sizeR)
res_norm_IF_max = np.zeros(sizeR)
res_norm_IF_non_norm_max = np.zeros(sizeR)
LODF_RT = sub_matrix(setR_this_ring, setT, LODF)
LODFn_RT = LODF_RT * sub_matrix(setR_this_ring, setT, PATL)
compute_IF_CPU(set_size_RIT,
vPTDF_I, vPTDF_R, mxPTDF_IR, mxPTDF_IT, mxPTDF_RI, mxPTDF_RT,
res_T, res_IF, set_IR, set_RT, set_TI,
mxPATL_RT, res_norm_IF, res_norm_T, res_norm_IF_non_norm)
get_max_results(res_T, res_IF, res_norm_T, res_norm_IF, res_norm_IF_non_norm,
res_T_max, res_norm_T_max, res_I_max, res_norm_I_max, res_IF_max,
res_norm_IF_max, res_norm_IF_non_norm_max)
for idx in range(len(setR_this_ring)):
# Template : "name,N-1 IF, N-1 nIF,IF,i,t,nIF,i,t,NNnIF"
r = setR_this_ring[idx]
IF_1 = max(np.absolute(LODF_RT[:, idx]))
norm_IF_1 = max(np.absolute(LODFn_RT[:, idx]))
IF_2 = res_IF_max[idx]
norm_IF_2 = res_norm_IF_max[idx]
i = setI[res_I_max[idx]]
t = setT[res_T_max[idx]]
i_norm = setI[res_norm_I_max[idx]]
t_norm = setT[res_norm_T_max[idx]]
LODF_it = LODF[t_norm.index, i_norm.index]
LODF_ir = LODF[r.index, i_norm.index]
results.append(Result_IF(r, IF_1, norm_IF_1, IF_2, norm_IF_2,
i, t, i_norm, t_norm, LODF_it, LODF_ir))
current_ring += 1
setR_this_ring = [elt for elt in branches if elt.ring == current_ring]
logging.info("IF computed in " + str(round(time.clock() - t0, 1)) + " seconds.")
return results
# Function defined to compute N-2 IF on CPU
@jit('void(int32[:], float64[:], float64[:], float64[:,:], float64[:,:], float64[:,:], float64[:,'
':], int32[:,:], float64[:,:], int32[:], int32[:], int32[:], float64[:], float64[:], '
'int32[:], float64[:,:])')
def compute_IF_CPU(set_size_RIT, vPTDF_I, vPTDF_R, mxPTDF_IR, mxPTDF_IT, mxPTDF_RI, mxPTDF_RT,
res_T, res_IF, set_IR, set_RT, set_TI, mxPATL_RT, res_norm_IF, res_norm_T,
res_norm_IF_non_norm):
epsilon = 0.00001
for (r, i) in np.ndindex((set_size_RIT[0], set_size_RIT[1])):
PTDF_ir = mxPTDF_IR[r, i]
PTDF_ri = mxPTDF_RI[i, r]
PTDF_i = vPTDF_I[i]
PTDF_r = vPTDF_R[r]
denominator = (1 - PTDF_i) * (1 - PTDF_r) - PTDF_ir * PTDF_ri
if abs(denominator) > epsilon:
for t in range(set_size_RIT[2]):
if set_IR[i] != r and set_RT[r] != t and set_TI[t] != i:
PTDF_it = mxPTDF_IT[t, i]
PTDF_rt = mxPTDF_RT[t, r]
PATL_rt = mxPATL_RT[t, r]
numerator = PTDF_it * PTDF_ri + (1 - PTDF_i) * PTDF_rt
IF = numerator / denominator
if abs(IF) > res_IF[i, r]:
res_IF[i, r] = abs(IF)
res_T[i, r] = t
norm_IF = PATL_rt * abs(IF)
if norm_IF > res_norm_IF[i, r]:
res_norm_IF[i, r] = norm_IF
res_norm_IF_non_norm[i, r] = abs(IF)
res_norm_T[i, r] = t
# Function defined to get IF, t and i from 2-D matrices previously computed (CPU compiled)
@jit(
'void(int32[:,:], float64[:,:], int32[:,:], float64[:,:], float64[:,:], int32[:], int32[:], '
'int32[:], int32[:], float64[:], float64[:], float64[:])')
def get_max_results(res_T, res_IF, res_norm_T, res_norm_IF, res_norm_IF_non_norm,
res_T_max, res_norm_T_max, res_I_max, res_norm_I_max, res_IF_max,
res_norm_IF_max, res_norm_IF_non_norm_max):
for (i, r) in np.ndindex(res_T.shape):
if res_IF[i, r] > res_IF_max[r]:
res_IF_max[r] = res_IF[i, r]
res_T_max[r] = res_T[i, r]
res_I_max[r] = i
if res_norm_IF[i, r] > res_norm_IF_max[r]:
res_norm_IF_max[r] = res_norm_IF[i, r]
res_norm_T_max[r] = res_norm_T[i, r]
res_norm_I_max[r] = i
res_norm_IF_non_norm_max[r] = res_norm_IF_non_norm[i, r]
def compute_IFs_generators(branches, setT, setI, setR_gens, LODF, LODF_gens, PATL):
t0 = time.clock()
logging.info("computing IF for generators")
LODF_gens_norm = LODF_gens * normalize_generators(branches, setR_gens)
list_LODF_gens = []
list_LODFnorm_gens = []
for t in setT:
list_LODF_gens.append(LODF_gens[t.index, :])
list_LODFnorm_gens.append(LODF_gens_norm[t.index, :])
mxLODF_gens_TR =
|
np.array(list_LODF_gens)
|
numpy.array
|
import numpy as np
import cv2
import img_to_depth as itd
import time
img = cv2.imread("cam2/tape_1.jpg")
itd_cvter = itd.ImageToDepth(0)
pixmm = itd_cvter.pix_to_mm
_, hm = itd_cvter.convert(img)
def hm2pos(hm):
depth_max = np.max(hm)
hm_map = hm/depth_max * 255
width, height = np.shape(hm_map)[:]
hm_map = hm_map.astype('uint8')
img = hm_map
f = np.fft.fft2(img)
fshift = np.fft.fftshift(f)
magnitude_spectrum = 100 * np.log(np.abs(fshift))
rows, cols = img.shape
crow, ccol = int(rows / 2), int(cols / 2)
p = 100
fshift[crow - p:crow + p, ccol - p:ccol + p] = 0
f_ishift = np.fft.ifftshift(fshift)
img_back = np.fft.ifft2(f_ishift)
img_back = np.abs(img_back)
img_back = (img_back /
|
np.amax(img_back)
|
numpy.amax
|
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.