hexsha
stringlengths 40
40
| size
int64 6
14.9M
| ext
stringclasses 1
value | lang
stringclasses 1
value | max_stars_repo_path
stringlengths 6
260
| max_stars_repo_name
stringlengths 6
119
| max_stars_repo_head_hexsha
stringlengths 40
41
| max_stars_repo_licenses
list | max_stars_count
int64 1
191k
⌀ | max_stars_repo_stars_event_min_datetime
stringlengths 24
24
⌀ | max_stars_repo_stars_event_max_datetime
stringlengths 24
24
⌀ | max_issues_repo_path
stringlengths 6
260
| max_issues_repo_name
stringlengths 6
119
| max_issues_repo_head_hexsha
stringlengths 40
41
| max_issues_repo_licenses
list | max_issues_count
int64 1
67k
⌀ | max_issues_repo_issues_event_min_datetime
stringlengths 24
24
⌀ | max_issues_repo_issues_event_max_datetime
stringlengths 24
24
⌀ | max_forks_repo_path
stringlengths 6
260
| max_forks_repo_name
stringlengths 6
119
| max_forks_repo_head_hexsha
stringlengths 40
41
| max_forks_repo_licenses
list | max_forks_count
int64 1
105k
⌀ | max_forks_repo_forks_event_min_datetime
stringlengths 24
24
⌀ | max_forks_repo_forks_event_max_datetime
stringlengths 24
24
⌀ | avg_line_length
float64 2
1.04M
| max_line_length
int64 2
11.2M
| alphanum_fraction
float64 0
1
| cells
list | cell_types
list | cell_type_groups
list |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
d0fc00227b98357d7909493d49e598a422eebf1e | 51,943 | ipynb | Jupyter Notebook | Final_Task_2019/Merged_Files/.ipynb_checkpoints/PR5_13317025_13317041-checkpoint.ipynb | bernardusrendy/ssicon | 16e7cea8a2177c3762b5a8014969ae4197626cd7 | [
"MIT"
] | 1 | 2019-12-03T15:37:39.000Z | 2019-12-03T15:37:39.000Z | Final_Task_2019/Merged_Files/.ipynb_checkpoints/PR5_13317025_13317041-checkpoint.ipynb | bernardusrendy/ssicon | 16e7cea8a2177c3762b5a8014969ae4197626cd7 | [
"MIT"
] | null | null | null | Final_Task_2019/Merged_Files/.ipynb_checkpoints/PR5_13317025_13317041-checkpoint.ipynb | bernardusrendy/ssicon | 16e7cea8a2177c3762b5a8014969ae4197626cd7 | [
"MIT"
] | 1 | 2020-01-15T02:38:21.000Z | 2020-01-15T02:38:21.000Z | 40.391135 | 1,099 | 0.565081 | [
[
[
"<h1 align=\"center\"> TUGAS BESAR TF3101 - DINAMIKA SISTEM DAN SIMULASI </h1>\n<h2 align=\"center\"> Sistem Elektrik, Elektromekanik, dan Mekanik</h2>",
"_____no_output_____"
],
[
"<h3>Nama Anggota:</h3>\n<body>\n <ul>\n <li>Erlant Muhammad Khalfani (13317025)</li>\n <li>Bernardus Rendy (13317041)</li>\n </ul>\n</body>",
"_____no_output_____"
],
[
"## 1. Pemodelan Sistem Elektrik##",
"_____no_output_____"
],
[
"Untuk pemodelan sistem elektrik, dipilih rangkaian RLC seri dengan sebuah sumber tegangan seperti yang tertera pada gambar di bawah ini.",
"_____no_output_____"
],
[
"<img src=\"./ELEKTRIK_TUBES_3.png\" style=\"width:50%\" align=\"middle\">",
"_____no_output_____"
],
[
"### Deskripsi Sistem\n\n1. Input <br>\nSistem ini memiliki input sumber tegangan $v_i$, yang merupakan fungsi waktu $v_i(t)$. <br>\n2. Output <br>\nSistem ini memiliki *output* arus $i_2$, yaitu arus yang mengalir pada *mesh* II. Tegangan $v_{L1}$ dan $v_{R2}$ juga dapat berfungsi sebagai *output*. Pada program ini, *output* yang akan di-*plot* hanya $v_{R2}$ dan $v_{L1}$. Nilai $i_2$ berbanding lurus terhadap nilai $v_{R2}$, sehingga bentuk grafik $i_2$ akan menyerupai bentuk grafik $v_{R2}$\n3. Parameter <br>\nSistem ini memiliki parameter-parameter $R_1$, $R_2$, $L_1$, dan $C_1$. Hambatan-hambatan $R_1$ dan $R_2$ adalah parameter *resistance*. Induktor $L_1$ adalah parameter *inertance*. Kapasitor $C_1$ adalah parameter *capacitance*.",
"_____no_output_____"
],
[
"### Asumsi\n1. Arus setiap *mesh* pada keadaan awal adalah nol ($i_1(0) = i_2(0) = 0$).\n2. Turunan arus terhadap waktu pada setiap *mesh* adalah nol ($\\frac{di_1(0)}{dt}=\\frac{di_2(0)}{dt}=0$)",
"_____no_output_____"
],
[
"### Pemodelan dengan *Bond Graph*\n\nDari sistem rangkaian listrik di atas, akan didapatkan *bond graph* sebagai berikut.\n<img src=\"./BG_ELEKTRIK.png\" style=\"width:50%\" align=\"middle\">\n<br>\nPada gambar di atas, terlihat bahwa setiap *junction* memenuhi aturan kausalitas. Ini menandakan bahwa rangkaian di atas bersifat *causal*. Dari *bond graph* di atas, dapat diturunkan *Ordinary Differential Equation* (ODE) seperti hasil penerapan *Kirchhoff's Voltage Law* (KVL) pada setiap *mesh*. Dalam pemodelan *bond graph* variabel-variabel dibedakan menjadi variabel *effort* dan *flow*. Sistem di atas merupakan sistem elektrik, sehingga memiliki variabel *effort* berupa tegangan ($v$) dan *flow* berupa arus ($i$).",
"_____no_output_____"
],
[
"### Persamaan Matematis - ODE\nDilakukan analisis besaran *effort* pada *1-junction* sebelah kiri. Ini akan menghasilkan:\n$$\nv_i = v_{R1} + v_{C1}\n$$\n<br>\nHasil ini sama seperti hasil dari KVL pada *mesh* I. Nilai $v_{R1}$ dan $v_{C1}$ diberikan oleh rumus-rumus:\n$$\nv_{R1} = R_1i_1\n$$\n<br>\n$$\nv_{C1} = \\frac{1}{C_1}\\int (i_1 - i_2)dt\n$$\nsehingga hasil KVL pada *mesh* I menjadi:\n$$\nv_i = R_1i_1 + \\frac{1}{C_1}\\int (i_1 - i_2)dt\n$$",
"_____no_output_____"
],
[
"Kemudian, analisis juga dilakukan pada *1-junction* sebelah kanan, yang akan menghasilkan:\n$$\nv_{C1} = v_{R2} + v_{L1}\n$$\n<br>\nIni juga sama seperti hasil KVL pada *mesh* II. Nilai $v_{R2}$ dan $v_{L1}$ diberikan oleh rumus-rumus:\n$$\nv_{R2} = R_2i_2\n$$\n<br>\n$$\nv_{L1} = L_1\\frac{di_2}{dt}\n$$\nsehingga hasil KVL pada *mesh* II menjadi:\n$$\n\\frac{1}{C_1}\\int(i_1-i_2)dt = R_2i_2 + L_1\\frac{di_2}{dt}\n$$\natau\n$$\n0 = L_1\\frac{di_2}{dt} + R_2i_2 + \\frac{1}{C_1}\\int(i_2-i_1)dt\n$$",
"_____no_output_____"
],
[
"### Persamaan Matematis - *Transfer Function*\nSetelah didapatkan ODE hasil dari *bond graph*, dapat dilakukan *Laplace Transform* untuk mendapatkan fungsi transfer sistem. *Laplace Transform* pada persamaan hasil KVL *mesh* I menghasilkan:\n$$\n(R_1 + \\frac{1}{C_1s})I_1 + (-\\frac{1}{C_1s})I_2 = V_i\n$$\n<br>\ndan pada persamaan hasil *mesh* II, akan didapatkan:\n$$\n(-\\frac{1}{C_1s})I_1 + (L_1s + R_2 + \\frac{1}{C_1s})I_2 = 0\n$$\n<br>\nKedua persamaan itu dieliminasi, sehingga didapatkan fungsi transfer antara $I_2$ dengan $V_i$\n$$\n\\frac{I_2(s)}{V_i(s)} = \\frac{1}{R_1C_1L_1s^2 + (R_1R_2C_1 + L_1)s + R_1 + R_2}\n$$\n<br>\nDari hasil *Laplace Transform* persamaan pada *mesh* II, didapatkan nilai $V_{L1}$ dari rumus\n$$\nV_{L1} = L_1sI_2\n$$\n<br>\nsehingga didapatkan fungsi transfer antara $V_{L1}$ dengan $V_i$\n$$\n\\frac{V_{L1}(s)}{V_i(s)} = \\frac{L_1s}{R_1C_1L_1s^2 + (R_1R_2C_1 + L_1)s + R_1 + R_2}\n$$\nSementara fungsi transfer antara $V_{R2}$ dan $V_i$ adalah\n$$\n\\frac{V_{R2}(s)}{V_i(s)} = \\frac{R_2}{R_1C_1L_1s^2 + (R_1R_2C_1 + L_1)s + R_1 + R_2}\n$$",
"_____no_output_____"
]
],
[
[
"#IMPORTS\nfrom ipywidgets import interact, interactive, fixed, interact_manual , HBox, VBox, Label, Layout\nimport ipywidgets as widgets\nimport numpy as np\nimport matplotlib.pyplot as plt\nfrom scipy import signal",
"_____no_output_____"
],
[
"#DEFINISI SLIDER-SLIDER PARAMETER\n#Slider R1\nR1_slider = widgets.FloatSlider(\n value=1.,\n min=1.,\n max=1000.,\n step=1.,\n description='$R_1 (\\Omega)$',\n readout_format='.1f',\n)\n#Slider R2\nR2_slider = widgets.FloatSlider(\n value=1.,\n min=1.,\n max=1000.,\n step=1.,\n description='$R_2 (\\Omega)$',\n readout_format='.1f',\n)\n#Slider C1\nC1_slider = widgets.IntSlider(\n value=1,\n min=10,\n max=1000,\n step=1,\n description='$C_1 (\\mu F)$',\n)\n#Slider L1\nL1_slider = widgets.FloatSlider(\n value=0.1,\n min=1.,\n max=1000.,\n step=0.1,\n description='$L_1 (mH)$',\n readout_format='.1f',\n)",
"_____no_output_____"
],
[
"#DEKLARASI SELECTOR INPUT\n#Slider selector input\nvi_select = signal_select = widgets.Dropdown(\n options=[('Step', 0), ('Impulse', 1)],\n description='Tipe Sinyal:',\n)\n#DEKLARASI SELECTOR OUTPUT\n#Output Selector\nvo_select = widgets.ToggleButtons(\n options=['v_R2', 'v_L1'],\n description='Output:',\n)",
"_____no_output_____"
],
[
"#DEKLARASI TAMBAHAN UNTUK INTERFACE\n#Color button\ncolor_select1 = widgets.ToggleButtons(\n options=['blue', 'red', 'green', 'black'],\n description='Color:',\n)",
"_____no_output_____"
],
[
"#PENENTUAN NILAI-NILAI PARAMETER\nR1 = R1_slider.value\nR2 = R2_slider.value\nC1 = C1_slider.value\nL1 = L1_slider.value\n\n#PENENTUAN NILAI DAN BENTUK INPUT\nvform = vi_select.value\n#PENENTUAN OUTPUT\nvo = vo_select\n\n#PENENTUAN PADA INTERFACE\ncolor = color_select1.value",
"_____no_output_____"
],
[
"#Plot v_L1 menggunakan transfer function\ndef plot_electric (vo, R1, R2, C1, L1, vform, color):\n #Menyesuaikan nilai parameter dengan satuan\n R1 = R1\n R2 = R2\n C1 = C1*(10**-6)\n L1 = L1*(10**-3)\n \n f, ax = plt.subplots(1, 1, figsize=(8, 6))\n num1 = [R2]\n num2 = [L1, 0]\n den = [R1*C1*L1, R1*R2*C1+L1, R1+R2]\n if vo=='v_R2':\n sys_vr =signal.TransferFunction(num1, den)\n step_vr = signal.step(sys_vr)\n impl_vr = signal.impulse(sys_vr)\n if vform == 0:\n ax.plot(step_vr[0], step_vr[1], color=color, label='Respon Step')\n elif vform == 1:\n ax.plot(impl_vr[0], impl_vr[1], color=color, label='Respon Impuls')\n ax.grid()\n ax.legend()\n elif vo=='v_L1':\n sys_vl = signal.TransferFunction(num2, den)\n step_vl = signal.step(sys_vl)\n impl_vl = signal.impulse(sys_vl)\n #Plot respon\n if vform == 0:\n ax.plot(step_vl[0], step_vl[1], color=color, label='Respon Step')\n elif vform == 1:\n ax.plot(impl_vl[0], impl_vl[1], color=color, label='Respon Impuls')\n ax.grid()\n ax.legend()",
"_____no_output_____"
],
[
"ui_el = widgets.VBox([vo_select, R1_slider, R2_slider, C1_slider, L1_slider, vi_select, color_select1])\nout_el = widgets.interactive_output(plot_electric, {'vo':vo_select,'R1':R1_slider,'R2':R2_slider,'C1':C1_slider,'L1':L1_slider,'vform':vi_select,'color':color_select1})\nint_el = widgets.HBox([ui_el, out_el])",
"_____no_output_____"
],
[
"display(int_el)",
"_____no_output_____"
]
],
[
[
"### Analisis###",
"_____no_output_____"
],
[
"<h4>a. Respon Step </h4>",
"_____no_output_____"
],
[
"Dari hasil simulasi, didapatkan pengaruh perubahan-perubahan nilai parameter pada *output* sistem setelah diberikan *input* berupa sinyal *step*, di antaranya:\n1. Kenaikan nilai $R_1$ akan menurunkan *steady-state gain* ($K$) sistem. Ini terlihat dari turunnya nilai *output* $v_{R2}$ pada keadaan *steady-state* dan turunnya nilai *maximum overshoot* ($M_p$) pada *output* $v_{L1}$. Perubahan nilai $R_1$ juga berbanding terbalik dengan perubahan koefisien redaman $\\xi$, terlihat dari semakin jelas terlihatnya osilasi seiring dengan kenaikan nilai $R_1$. Perubahan nilai $R_1$ juga sebanding dengan perubahan nilai *settling time* ($t_s$). Ini terlihat dengan bertambahnya waktu sistem untuk mencapai nilai dalam rentang 2-5% dari nilai keadaan *steady-state*. \n2. Kenaikan nilai $R_2$ akan meningkatkan *steady-state gain* ($K$) sistem dengan *output* $v_{R2}$ tetapi menurunkan *steady-state gain* ($K$) *output* $v_{L1}$. Selain itu, dapat terlihat juga bahwa perubahan nilai $R_2$ berbanding terbalik dengan nilai *settling time* ($t_s$); Saat nilai $R_2$ naik, sistem mencapai kondisi *steady-state* dalam waktu yang lebih singkat. Kenaikan nilai $R_2$ juga menyebabkan penurunan nilai *maximum overshoot* ($M_p$).\n3. Perubahan nilai $C_1$ sebanding dengan perubahan nilai *settling time*, seperti yang dapat terlihat dengan bertambahnya waktu yang diperlukan sistem untuk mendekati keadaan *steady-state* seiring dengan kenaikan nilai $C_1$. Selain itu nilai $C_1$ juga berbanding terbalik dengan nilai *maximum overshoot*, ini dapat dilihat dari turunnya nilai *maximum overshoot* ketika nilai $C_1$ dinaikan. Pada saat nilai $C_1$, naik, juga terlihat kenaikan nilai *delay time* ($t_d$), *rise time* ($t_r$), dan *peak time* ($t_p$).\n4. Kenaikan nilai $L_1$ mengakibatkan berkurangnya nilai frekuensi osilasi, serta meningkatkan *settling time* sistem. Perubahan nilai $L_1$ ini juga sebanding dengan *steady-state gain* sistem untuk *output* $v_{L1}$.",
"_____no_output_____"
],
[
"<h4>b. Respon Impuls </h4>",
"_____no_output_____"
],
[
"Dari hasil simulasi, didapatkan pengaruh perubahan-perubahan nilai parameter pada *output* sistem setelah diberikan *input* berupa sinyal *impulse*, di antaranya:\n1. Perubahan nilai $R_1$ berbanding terbalik dengan nilai *peak response*. Kenaikan nilai $R_1$ juga menaikkan nilai *settling time* ($t_s$).\n2. Kenaikan nilai $R_2$ memengaruhi nilai *peak response* $v_{R2}$, tetapi tidak berpengaruh pada *peak response* $v_{L1}$. Naiknya nilai $R_2$ juga menurunkan nilai *settling time* ($t_s$), yang terlihat dari semakin cepatnya sistem mencapai kondisi *steady-state*.\n3. Kenaikan nilai $C_1$ menyebabkan turunnya nilai *peak response*. Kenaikan nilai $C_1$ juga menyebabkan kenaikan nilai *settling time* ($t_s$), yang dapat dilihat dengan bertambahnya waktu yang diperlukan sistem untuk mendekati keadaan *steady-state*.\n4. Kenaikan nilai $L_1$ menyebabkan turunnya nilai *peak response*. Kenaikan nilai $L_1$ juga menurunkan nilai *settling time* ($t_s$), yang dapat dilihat dari bertambahnya waktu yang diperlukan sistem untuk mendekati keadaan *steady-state*.",
"_____no_output_____"
],
[
"## 2. Pemodelan Sistem Elektromekanik\n### DC Brushed Motor Torsi Besar dengan Motor Driver\n\nSistem yang akan dimodelkan berupa motor driver high current BTS7960 seperti gambar pertama, dihubungkan dengan motor torsi besar dengan brush pada gambar kedua.\n<div>\n<img src=\"./1.jpg\" style=\"width:20%\" align=\"middle\">\n</div>\n<div>\n<img src=\"./2.jpg\" style=\"width:20%\" align=\"middle\">\n</div>\n<p style=\"text-align:center\"><b>Sumber gambar: KRTMI URO ITB</b></p>",
"_____no_output_____"
],
[
"### Deskripsi Sistem\n\n1. Input <br>\nSistem ini memiliki input sinyal $V_{in}$, yang merupakan fungsi waktu $V_{in}(t)$. Tegangan $v_i(t)$ ini dapat berbentuk fungsi step, impuls, atau pulse width modulation dengan duty cycle tertentu (luaran mikrokontroller umum). <br>\n2. Output <br>\nSistem ini memiliki *output* posisi sudut $\\theta$, kecepatan sudut motor $\\omega$, percepatan sudut motor $\\alpha$, dan torsi $T$. Output ditentukan sesuai kebutuhan untuk manuver robot. Terlihat variable output bergantung pada $\\theta$, $\\frac {d\\theta}{dt}$, dan $\\frac{d^2\\theta}{dt}$, sehingga dicari beberapa persamaan diferensial sesuai tiap output.\n3. Parameter <br>\nSistem memiliki parameter $J,K_f,K_a,L,R,K_{emf},K_{md}$ yang diturunkan dari karakteristik subsistem mekanik dan elektrik sebagai berikut.",
"_____no_output_____"
],
[
"#### Subsistem Motor Driver\nPertama ditinjau struktur sistem dari motor driver. Motor driver yang digunakan adalah tipe MOSFET BTS7960 sehingga memiliki karakteristik dinamik yang meningkat hampir dengan instant. MOSFET dirangkai sedemikian rupa sehingga dapat digunakan untuk kontrol maju/mundur motor. Diasumsikan rise-time MOSFET cukup cepat relatif terhadap sinyal dan motor driver cukup linear, maka motor driver dapat dimodelkan sebagai sistem orde 0 dengan gain sebesar $ K_{md} $. \n<img src=\"./4.png\" style=\"width:30%\" align=\"middle\">\n<p style=\"text-align:center\"><b>Sumber gambar: Datasheet BTS7960</b></p>\n<img src=\"./5.png\" style=\"width:30%\" align=\"middle\">\n<p style=\"text-align:center\"><b>Model Orde 0 Motor Driver</b></p>\nMaka persamaan dinamik output terhadap input dalam motor driver adalah <br>\n$ V_m=K_{md}V_{in} $<br>\nSama seperti hubungan input output pada karakteristik statik.",
"_____no_output_____"
],
[
"#### Subsistem Motor\nLalu ditinjau struktur sistem dari motor torsi besar dengan inertia beban yang tidak dapat diabaikan.\n<img src=\"./3.png\" style=\"width:30%\" align=\"middle\">\n<p style=\"text-align:center\"><b>Sumber gambar: https://www.researchgate.net/figure/The-structure-of-a-DC-motor_fig2_260272509</b></p>\n<br>\nMaka dapat diturunkan persamaan diferensial untuk sistem mekanik.\n<br>\n<img src=\"./6.png\" style=\"width:30%\">\n<img src=\"./7.png\" style=\"width:30%\">\n<p style=\"text-align:center\"><b>Sumber gambar: Chapman - Electric Machinery Fundamentals 4th Edition</b></p>\n$$ \nT=K_a i_a \n$$ \ndengan $T$ adalah torsi dan $K_a$ adalah konstanta proporsionalitas torsi (hasil perkalian K dengan flux) untuk arus armature $i_a$.\n$$\nV_{emf}=K_{emf} \\omega\n$$\ndengan $V_{emf}$ adalah tegangan penyebab electromotive force dan $K_{emf}$ konstanta proporsionalitas tegangan emf (hasil perkalian K dengan flux pada kondisi ideal tanpa voltage drop) untuk kecepatan putar sudut dari motor.\n<br>\nNamun, akibat terbentuknya torsi adalah berputarnya beban dengan kecepatan sudut sebesar $\\omega$ dan percepatan sudut sebesar $\\alpha$. Faktor proporsionalitas terhadap percepatan sudut adalah $J$ (Inersia Putar) dan terhadap kecepatan sudut sebesar $ K_f $ (Konstanta Redam Putar) Sehingga dapat diturunkan persamaan diferensial sebagai berikut (Persamaan 1):\n<br>\n$$ \nJ\\alpha + K_f\\omega = T \n$$\n$$\nJ\\frac {d^2\\theta}{dt} + K_f\\frac {d\\theta}{dt} = K_a i_a \n$$\n$$ \nJ\\frac {d\\omega}{dt} + K_f \\omega = K_a i_a \n$$\nKemudian diturunkan persamaan diferensial untuk sistem elektrik yang terdapat pada motor sehingga $i_a$ dapat disubstitusi dengan input $V_{in}$ (Persamaan 2):\n$$ \nL \\frac{d{i_a}}{dt} + R i_a + K_{emf} \\omega = V_m \n$$\n$$\nV_m = K_{md} V_{in}\n$$\n$$ \nL \\frac{d{i_a}}{dt} + R i_a + K_{emf} \\omega = K_{md} V_{in} \n$$",
"_____no_output_____"
],
[
"### Pemodelan dengan Fungsi Transfer\nDengan persamaan subsistem tersebut, dapat dilakukan pemodelan fungsi transfer sistem dengan transformasi ke domain laplace (s). Dilakukan penyelesaian menggunakan fungsi transfer dalam domain laplace, pertama dilakukan transfer ke domain laplace dengan asumsi\n<br>\n$ i_a (0) = 0 $\n<br>\n$ \\frac {di_a}{dt} = 0 $\n<br>\n$ \\theta (0) = 0 $\n<br>\n$ \\omega (0) = 0 $\n<br>\n$ \\alpha (0) = 0 $\n<br>\nTidak diasumsikan terdapat voltage drop karena telah di akumulasi di $K_{emf}$, namun diasumsikan voltage drop berbanding lurus terhadap $\\omega$.\n<br>\nPersamaan 1 menjadi:\n$$\nJ s \\omega + K_f \\omega = K_a i_a\n$$\nPersamaan 2 menjadi:\n$$\nL s i_a + R i_a + K_{emf} \\omega = K_{md} V_{in}\n$$\n$$\ni_a=\\frac {K_{md} V_{in}-K_{emf} \\omega}{L s + R}\n$$\nSehingga terbentuk fungsi transfer sistem keseluruhan dalam $\\omega$ adalah:\n$$\nJ s \\omega + K_f \\omega = \\frac {K_a(K_{md} V_{in} - K_{emf} \\omega)}{L s + R}\n$$\nFungsi transfer untuk $\\omega$ adalah:\n$$\n\\omega = \\frac {K_a(K_{md} V_{in}-K_{emf} \\omega)}{(L s + R)(J s + K_f)}\n$$\n$$\n\\omega = \\frac {K_a K_{md} V_{in}}{(L s + R)(J s + K_f)(1 + \\frac {K_a K_{emf}}{(L s + R)(J s + K_f)})}\n$$\n$$\n\\frac {\\omega (s)}{V_{in}(s)} = \\frac {K_a K_{md}}{(L s + R)(J s + K_f)+ K_a K_{emf}}\n$$\nDapat diturunkan fungsi transfer untuk theta dengan mengubah variable pada persamaan 1:\n$$\nJ s^2 \\theta + K_f s \\theta = K_a i_a\n$$\nPersamaan 2:\n$$\nL s i_a + R i_a + K_{emf} s \\theta = K_{md} V_{in}\n$$\n$$\ni_a=\\frac {K_{md} V_{in}-K_{emf} s \\theta}{L s + R}\n$$\nSehingga terbentuk fungsi transfer sistem keseluruhan dalam $\\theta$ adalah:\n$$\nJ s^2 \\theta + K_f s \\theta = \\frac {K_a(K_{md} V_{in}-K_{emf} s \\theta)}{L s + R}\n$$\nFungsi transfer untuk $\\theta$ adalah:\n$$\n\\theta = \\frac {K_a(K_{md} V_{in}-K_{emf} s \\theta)}{(L s + R)(J s^2 + K_f s )}\n$$\n$$\n\\theta + \\frac {K_a K_{emf} s \\theta}{(L s + R)(J s^2 + K_f s )}= \\frac {K_a K_{md} V_{in}}{(L s + R)(J s^2 + K_f s )}\n$$\n$$\n\\theta= \\frac {K_a K_{md} V_{in}}{(L s + R)(J s^2 + K_f s )(1 + \\frac {K_a K_{emf} s}{(L s + R)(J s^2 + K_f s )})}\n$$\n$$\n\\frac {\\theta (s)}{V_{in}(s)}= \\frac {K_a K_{md}}{(L s + R)(J s^2 + K_f s )+ K_a K_{emf} s}\n$$\nTerlihat bahwa fungsi transfer untuk $\\omega$ dan $\\theta$ hanya berbeda sebesar $ \\frac {1}{s} $ sesuai dengan hubungan\n$$\n\\omega = s \\theta\n$$\nSehingga fungsi transfer untuk $\\alpha$ akan memenuhi\n$$\n\\alpha = s\\omega = s^2 \\theta\n$$\nSehingga fungsi transfer untuk $\\alpha$ adalah:\n$$\n\\frac {\\alpha (s)}{V_{in}(s)} = \\frac {K_a K_{md} s}{(L s + R)(J s + K_f)+ K_a K_{emf}}\n$$",
"_____no_output_____"
],
[
"### Output\nDari fungsi transfer, diformulasikan persamaan output posisi sudut $\\theta$, kecepatan sudut motor $\\omega$, percepatan sudut $\\alpha$, dan torsi $T$ dalam fungsi waktu (t).\n$$\n\\theta (t) = \\mathscr {L^{-1}} \\{\\frac {K_a K_{md} V_{in}(s)}{(L s + R)(J s^2 + K_f s )+ K_a K_{emf} s}\\}\n$$\n<br>\n$$\n\\omega (t) = \\mathscr {L^{-1}} \\{\\frac {K_a K_{md} V_{in}(s)}{(L s + R)(J s + K_f)+ K_a K_{emf}}\\}\n$$\n<br>\n$$\n\\alpha (t)= \\mathscr {L^{-1}} \\{\\frac {K_a K_{md} Vin_{in}(s) s}{(L s + R)(J s + K_f)+ K_a K_{emf}}\\}\n$$\n<br>\n$$\nT = \\frac {K_a(K_{md} V_{in}-K_{emf} \\omega)}{L s + R} \n$$",
"_____no_output_____"
]
],
[
[
"# Digunakan penyelesaian numerik untuk output\nimport numpy as np\nfrom scipy.integrate import odeint\nimport scipy.signal as sig\nimport matplotlib.pyplot as plt\nfrom sympy.physics.mechanics import dynamicsymbols, SymbolicSystem\nfrom sympy import *\nimport control as control",
"_____no_output_____"
],
[
"vin = symbols ('V_{in}') #import symbol input",
"_____no_output_____"
],
[
"omega, theta, alpha = dynamicsymbols('omega theta alpha') #import symbol output",
"_____no_output_____"
],
[
"ka,kmd,l,r,j,kf,kemf,s,t = symbols ('K_a K_{md} L R J K_f K_{emf} s t')#import symbol parameter dan s",
"_____no_output_____"
],
[
"thetaOverVin = (ka*kmd)/((l*s+r)*(j*s**2+kf*s)+ka*kemf*s) #persamaan fungsi transfer theta\npolyThetaOverVin = thetaOverVin.as_poly() #Penyederhanaan persamaan\npolyThetaOverVin",
"_____no_output_____"
],
[
"omegaOverVin = (ka*kmd)/((l*s+r)*(j*s+kf)+ka*kemf) #persamaan fungsi transfer omega\npolyOmegaOverVin = omegaOverVin.as_poly() #Penyederhanaan persamaan\npolyOmegaOverVin",
"_____no_output_____"
],
[
"alphaOverVin = (ka*kmd*s)/((l*s+r)*(j*s+kf)+ka*kemf)\npolyAlphaOverVin = alphaOverVin.as_poly() #Penyederhanaan persamaan\npolyAlphaOverVin",
"_____no_output_____"
],
[
"torqueOverVin= ka*(kmd-kemf*((ka*kmd)/((l*s+r)*(j*s+kf)+ka*kemf)))/(l*s+r) #Penyederhanaan persamaan torsi\npolyTorqueOverVin = torqueOverVin.as_poly()\npolyTorqueOverVin",
"_____no_output_____"
],
[
"def plot_elektromekanik(Ka,Kmd,L,R,J,Kf,Kemf,VinType,tMax,dutyCycle,grid):\n # Parameter diberi value dan model system dibentuk dalam transfer function yang dapat diolah python\n Ka = Ka\n Kmd = Kmd\n L = L\n R = R\n J = J\n Kf = Kf\n Kemf = Kemf\n # Pembuatan model transfer function\n tf = control.tf\n tf_Theta_Vin = tf([Ka*Kmd],[J*L,(J*R+Kf*L),(Ka*Kemf+Kf*R),0])\n tf_Omega_Vin = tf([Ka*Kmd],[J*L,(J*R+Kf*L),(Ka*Kemf+Kf*R)])\n tf_Alpha_Vin = tf([Ka*Kmd,0],[J*L,(J*R+Kf*L),(Ka*Kemf+Kf*R)])\n tf_Torque_Vin = tf([Ka*Kmd],[L,R]) - tf([Kmd*Kemf*Ka**2],[J*L**2,(2*J*L*R+Kf*L**2),(J*R**2+Ka*Kemf*L+2*Kf*L*R),(Ka*Kemf*R+Kf*R**2)])\n f, axs = plt.subplots(4, sharex=True, figsize=(10, 10))\n # Fungsi mengatur rentang waktu analisis (harus memiliki kelipatan 1 ms)\n def analysisTime(maxTime):\n ts=np.linspace(0, maxTime, maxTime*100)\n return ts\n t=analysisTime(tMax)\n if VinType== 2:\n # Input pwm dalam 1 millisecond\n def Pwm(dutyCycle,totalTime):\n trepeat=np.linspace(0, 1, 100)\n squareWave=(5*sig.square(2 * np.pi * trepeat, duty=dutyCycle))\n finalInput=np.zeros(len(totalTime))\n for i in range(len(squareWave)):\n if squareWave[i]<0:\n squareWave[i]=0\n for i in range(len(totalTime)):\n finalInput[i]=squareWave[i%100]\n return finalInput\n pwm=Pwm(dutyCycle,t)\n tPwmTheta, yPwmTheta, xPwmTheta = control.forced_response(tf_Theta_Vin, T=t, U=pwm, X0=0)\n tPwmOmega, yPwmOmega, xPwmOmega = control.forced_response(tf_Omega_Vin, t, pwm, X0=0)\n tPwmAlpha, yPwmAlpha, xPwmAlpha = control.forced_response(tf_Alpha_Vin, t, pwm, X0=0)\n tPwmTorque, yPwmTorque, xPwmTorque = control.forced_response(tf_Torque_Vin, t, pwm, X0=0)\n axs[0].plot(tPwmTheta, yPwmTheta, color = 'blue', label ='Theta')\n axs[1].plot(tPwmOmega, yPwmOmega, color = 'red', label ='Omega')\n axs[2].plot(tPwmAlpha, yPwmAlpha, color = 'black', label ='Alpha')\n axs[3].plot(tPwmTorque, yPwmTorque, color = 'green', label ='Torque')\n axs[0].title.set_text('Theta $(rad)$ (Input PWM)')\n axs[1].title.set_text('Omega $(\\\\frac {rad}{ms})$ (Input PWM)')\n axs[2].title.set_text('Alpha $(\\\\frac {rad}{ms^2})$ (Input PWM)')\n axs[3].title.set_text('Torque $(Nm)$ (Input PWM)')\n elif VinType== 0:\n tStepTheta, yStepTheta = control.step_response(tf_Theta_Vin,T=t, X0=0)\n tStepOmega, yStepOmega = control.step_response(tf_Omega_Vin,T=t, X0=0)\n tStepAlpha, yStepAlpha = control.step_response(tf_Alpha_Vin,T=t, X0=0)\n tStepTorque, yStepTorque = control.step_response(tf_Torque_Vin, T=t, X0=0)\n axs[0].plot(tStepTheta, yStepTheta, color = 'blue', label ='Theta')\n axs[1].plot(tStepOmega, yStepOmega, color = 'red', label ='Omega')\n axs[2].plot(tStepAlpha, yStepAlpha, color = 'black', label ='Alpha')\n axs[3].plot(tStepTorque, yStepTorque, color = 'green', label ='Torque')\n axs[0].title.set_text('Theta $(rad)$ (Input Step)')\n axs[1].title.set_text('Omega $(\\\\frac {rad}{ms})$ (Input Step)')\n axs[2].title.set_text('Alpha $(\\\\frac {rad}{ms^2})$(Input Step)')\n axs[3].title.set_text('Torque $(Nm)$ (Input Step)')\n elif VinType== 1 :\n tImpulseTheta, yImpulseTheta = control.impulse_response(tf_Theta_Vin,T=t, X0=0)\n tImpulseOmega, yImpulseOmega = control.impulse_response(tf_Omega_Vin,T=t, X0=0)\n tImpulseAlpha, yImpulseAlpha = control.impulse_response(tf_Alpha_Vin,T=t, X0=0)\n tImpulseTorque, yImpulseTorque = control.impulse_response(tf_Torque_Vin, T=t, X0=0)\n axs[0].plot(tImpulseTheta, yImpulseTheta, color = 'blue', label ='Theta')\n axs[1].plot(tImpulseOmega, yImpulseOmega, color = 'red', label ='Omega')\n axs[2].plot(tImpulseAlpha, yImpulseAlpha, color = 'black', label ='Alpha')\n axs[3].plot(tImpulseTorque, yImpulseTorque, color = 'green', label ='Torque')\n axs[0].title.set_text('Theta $(rad)$ (Input Impulse)')\n axs[1].title.set_text('Omega $(\\\\frac {rad}{ms})$ (Input Impulse)')\n axs[2].title.set_text('Alpha $(\\\\frac {rad}{ms^2})$ (Input Impulse)')\n axs[3].title.set_text('Torque $(Nm)$ (Input Impulse)')\n axs[0].legend()\n axs[1].legend()\n axs[2].legend()\n axs[3].legend()\n axs[0].grid(grid)\n axs[1].grid(grid)\n axs[2].grid(grid)\n axs[3].grid(grid)",
"_____no_output_____"
],
[
"#DEFINISI WIDGETS PARAMETER\nKa_slider = widgets.FloatSlider(\n value=19.90,\n min=0.1,\n max=20.0,\n step=0.1,\n description='$K_a (\\\\frac {Nm}{A})$',\n layout=Layout(width='80%', height='50px'),\n style={'description_width': '200px'},\n)\nKmd_slider = widgets.FloatSlider(\n value=20.0,\n min=0.1,\n max=20.0,\n step=0.1,\n description='$K_{md} (\\\\frac {V}{V})$',\n layout=Layout(width='80%', height='50px'),\n style={'description_width': '200px'},\n)\nL_slider = widgets.FloatSlider(\n value=20,\n min=0.1,\n max=100.0,\n step=0.1,\n description='$L (mH)$',\n layout=Layout(width='80%', height='50px'),\n style={'description_width': '200px'},\n)\nR_slider = widgets.IntSlider(\n value=5,\n min=1,\n max=20,\n step=1,\n description='$R (\\Omega)$',\n layout=Layout(width='80%', height='50px'),\n style={'description_width': '200px'},\n)\nJ_slider = widgets.FloatSlider(\n value=25,\n min=0.1,\n max=100.0,\n step=0.1,\n description='$J (\\\\frac {Nm(ms)^2}{rad})$',\n layout=Layout(width='80%', height='50px'),\n style={'description_width': '200px'},\n)\nKf_slider = widgets.FloatSlider(\n value=8,\n min=0.1,\n max=100.0,\n step=0.1,\n description='$K_{f} (\\\\frac {Nm(ms)}{rad})$',\n layout=Layout(width='80%', height='50px'),\n style={'description_width': '200px'},\n)\nKemf_slider = widgets.FloatSlider(\n value=19.8,\n min=0.1,\n max=20,\n step=0.1,\n description='$K_{emf} (\\\\frac {V(ms)}{rad})$',\n layout=Layout(width='80%', height='50px'),\n style={'description_width': '200px'},\n)\nVinType_select = widgets.Dropdown(\n options=[('Step', 0), ('Impulse', 1),('PWM',2)],\n description='Tipe Sinyal Input:',\n layout=Layout(width='80%', height='50px'),\n style={'description_width': '200px'},\n)\ntMax_slider = widgets.IntSlider(\n value=50,\n min=1,\n max=500,\n step=1,\n description='$t_{max} (ms)$',\n layout=Layout(width='80%', height='50px'),\n style={'description_width': '200px'},\n)\ndutyCycle_slider = widgets.FloatSlider(\n value=0.5,\n min=0,\n max=1.0,\n step=0.05,\n description='$Duty Cycle (\\%)$',\n layout=Layout(width='80%', height='50px'),\n style={'description_width': '200px'},\n)\ngrid_button = widgets.ToggleButton(\n value=True,\n description='Grid',\n icon='check',\n layout=Layout(width='20%', height='50px',margin='10px 10px 10px 350px'),\n style={'description_width': '200px'},\n)\n\ndef update_Kemf_max(*args):\n Kemf_slider.max = Ka_slider.value\nKa_slider.observe(update_Kemf_max, 'value')\n\nui_em = widgets.VBox([Ka_slider,Kmd_slider,L_slider,R_slider,J_slider,Kf_slider,Kemf_slider,VinType_select,tMax_slider,dutyCycle_slider,grid_button])\nout_em = widgets.interactive_output(plot_elektromekanik, {'Ka':Ka_slider,'Kmd':Kmd_slider,'L':L_slider,'R':R_slider,'J':J_slider,'Kf':Kf_slider,'Kemf':Kemf_slider,'VinType':VinType_select,'tMax':tMax_slider,'dutyCycle':dutyCycle_slider, 'grid':grid_button})",
"_____no_output_____"
],
[
"display(ui_em,out_em)",
"_____no_output_____"
]
],
[
[
"### Analisis",
"_____no_output_____"
],
[
"Karena model memiliki persamaan yang cukup kompleks sehingga tidak dapat diambil secara intuitive kesimpulan parameter terhadap output sistem, akan dilakukan percobaan menggunakan slider untuk mengubah parameter dan mengamati interaksi perubahan antara parameter. Akan dilakukan juga perubahan bentuk input dan analisa efek penggunaan PWM sebagai modulasi sinyal step dengan besar maksimum 5V terhadap output.\n#### 1. Peningkatan $K_a$\nPeningkatan $K_a$ menyebabkan peningkatan osilasi ($\\omega_d$) dan meningkatkan gain pada output $\\omega$ dan $\\alpha$ serta meningkatkan gradien dari output $\\theta$. Namun, gain Torque tidak terpengaruh.\n#### 2. Peningkatan $K_{md}$\nPeningkatan $K_{md}$ membuat amplitudo $V_{in}$ meningkat sehingga amplitudo output bertambah.\n#### 3. Peningkatan $L$\nPeningkatan $L$ menyebabkan peningkatan kecepatan sudut $\\omega$ dan $T$ menjadi lebih lambat serta penurunan $\\alpha$ yang semakin lambat sehingga menyebabkan peningkatan $\\theta$ semakin lambat (peningkatan rise time).\n#### 4. Peningkatan $R$\nPeningkatan $R$ menyebabkan osilasi output ($\\omega_d$) $\\omega$, $\\alpha$, dan Torque semakin kecil dan gain yang semakin kecil sehingga mengurangi gradien dari output $\\theta$.\n#### 5. Peningkatan $J$\nPeningkatan $J$ meningkatkan gain Torque dan menurunkan gain $\\theta$, $\\omega$, dan $\\alpha$. \n#### 6. Peningkatan $K_f$\nPeningkatan $K_f$ meningkatkan gain Torque dan menurunkan gain $\\theta$, $\\omega$, dan $\\alpha$.\n#### 7. Peningkatan $K_{emf}$\nPeningkatan $K_{emf}$ menurunkan gain Torque, $\\theta$, $\\omega$, dan $\\alpha$.\n#### 8. Interaksi antar parameter\nPerbandingan pengurangan $R$ dibanding peningkatan $K_a$ kira kira 3 kali lipat. Peningkatan pada $J$ dan $K_f$ terbatas pada peningkatan $K_a$. Secara fisis, peningkatan $K_a$ dan $K_{emf}$ terjadi secara bersamaan dan hampir sebanding (hanya dibedakan pada voltage drop pada berbagai komponen), diikuti oleh $L$ sehingga untuk $K_a$ dan $K_{emf}$ besar, waktu mencapai steady state juga semakin lama. Hal yang menarik adalah $K_a$ dan $K_{emf}$ membuat sistem memiliki gain (transfer energi) yang kecil jika hanya ditinjau dari peningkatan nilai $K_a$ dan $K_{emf}$, namun ketika diikuti peningkatan $V_{in}$ sistem memiliki transfer energi yang lebih besar daripada sebelumnya pada keadaan steady state. Jadi dapat disimpulkan bahwa $K_a$ dan $K_{emf}$ harus memiliki nilai yang cukup besar agar konfigurasi sesuai dengan input $V_{in}$ dan menghasilkan transfer energi yang efisien. Input $V_{in}$ juga harus sesuai dengan sistem $K_a$ dan $K_{emf}$ yang ada sehingga dapat memutar motor (ini mengapa terdapat voltage minimum dan voltage yang disarankan untuk menjalankan sebuah motor).\n#### 9. Pengaruh Input Step\nPenggunaan input step memiliki osilasi ($\\omega_d$) semakin sedikit.\n#### 10. Pengaruh Input Impuls\nPenggunaan input impulse membuat $\\theta$ mencapai steady state karena motor berhenti berputar sehingga $\\omega$,$\\alpha$, dan Torque memiliki nilai steady state 0.\n#### 11. Pengaruh Input PWM\nPenggunaan input PWM dengan duty cycle tertentu membuat osilasi yang semakin banyak, namun dengan peningkatan duty cycle, osilasi semakin sedikit (semakin mendekati sinyal step). Hal yang menarik disini adalah sinyal PWM dapat digunakan untuk mengontrol, tetapi ketika tidak digunakan pengontrol, sinyal PWM malah memberikan osilasi pada sistem.",
"_____no_output_____"
],
[
"## 3. Pemodelan Sistem Mekanik\nDimodelkan sistem mekanik sebagai berikut\n<img src=\"./10.png\" style=\"width:20%\">\n<p style=\"text-align: center\"><b>Sistem Mekanik Sederhana dengan Bond Graph</b></p>",
"_____no_output_____"
],
[
"### Deskripsi Sistem\n\n1. Input\n$F$ sebagai gaya yang dikerjakan pada massa\n2. Output\n$x$ sebagai perpindahan, $v$ sebagai kecepatan, dan $a$ sebagai percepatan pada massa\n3. Parameter\nDari penurunan bond graph, didapatkan parameter $k$, $b$, dan $m$",
"_____no_output_____"
],
[
"### Pemodean Transfer Function\nFungsi transfer dapat dengan mudah di turunkan dari hubungan bond graph, diasumsikan \n$$\nx(0)=0\n$$\n$$\nv(0)=0\n$$\n$$\na(0)=0\n$$\n$$\nm \\frac {d^2 x}{dt^2} = F-kx-b\\frac{dx}{dt}\n$$\n<br>\nTransformasi laplace menghasilkan\n<br>\n$$\ns^2 x = \\frac {F}{m}-x\\frac {k}{m}-sx\\frac{b}{m}\n$$\n$$\n(s^2+s\\frac{b}{m}+\\frac {k}{m})x=\\frac {F}{m}\n$$\n<br>\nUntuk x:\n<br>\n$$\n\\frac {x}{F}=\\frac {1}{(ms^2+bs+k)}\n$$\n<br>\nUntuk v:\n<br>\n$$\n\\frac {v}{F}=\\frac {s}{(ms^2+bs+k)}\n$$\n<br>\nUntuk a:\n<br>\n$$\n\\frac {a}{F}=\\frac {s^2}{(ms^2+bs+k)}\n$$",
"_____no_output_____"
]
],
[
[
"# Digunakan penyelesaian numerik untuk output\nimport numpy as np\nfrom scipy.integrate import odeint\nimport scipy.signal as sig\nimport matplotlib.pyplot as plt\nfrom sympy.physics.mechanics import dynamicsymbols, SymbolicSystem\nfrom sympy import *\nimport control as control",
"_____no_output_____"
],
[
"def plot_mekanik(M,B,K,VinType,grid):\n # Parameter diberi value dan model system dibentuk dalam transfer function yang dapat diolah python\n m=M\n b=B\n k=K\n tf = sig.TransferFunction\n tf_X_F=tf([1],[m,b,k])\n tf_V_F=tf([1,0],[m,b,k])\n tf_A_F=tf([1,0,0],[m,b,k])\n f, axs = plt.subplots(3, sharex=True, figsize=(10, 10))\n if VinType==0:\n tImpX,xOutImp=sig.impulse(tf_X_F)\n tImpV,vOutImp=sig.impulse(tf_V_F)\n tImpA,aOutImp=sig.impulse(tf_A_F)\n axs[0].plot(tImpX,xOutImp, color = 'blue', label ='x')\n axs[1].plot(tImpV,vOutImp, color = 'red', label ='v')\n axs[2].plot(tImpA,aOutImp, color = 'green', label ='a')\n axs[0].title.set_text('Perpindahan Linear $(m)$ (Input Impuls)')\n axs[1].title.set_text('Kecepatan Linear $(\\\\frac {m}{s})$ (Input Impuls)')\n axs[2].title.set_text('Percepatan Linear $(\\\\frac {m}{s^2})$ (Input Impuls)')\n elif VinType==1:\n tStepX,xOutStep=sig.step(tf_X_F)\n tStepV,vOutStep=sig.step(tf_V_F)\n tStepA,aOutStep=sig.step(tf_A_F)\n axs[0].plot(tStepX,xOutStep, color = 'blue', label ='x')\n axs[1].plot(tStepV,vOutStep, color = 'red', label ='v')\n axs[2].plot(tStepA,aOutStep, color = 'green', label ='a')\n axs[0].title.set_text('Perpindahan Linear $(m)$ (Input Step)')\n axs[1].title.set_text('Kecepatan Linear $(\\\\frac {m}{s})$ (Input Step)')\n axs[2].title.set_text('Percepatan Linear $(\\\\frac {m}{s^2})$ (Input Step)')\n axs[0].legend()\n axs[1].legend()\n axs[2].legend()\n axs[0].grid(grid)\n axs[1].grid(grid)\n axs[2].grid(grid)",
"_____no_output_____"
],
[
"M_slider = widgets.FloatSlider(\n value=0.1,\n min=0.1,\n max=30.0,\n step=0.1,\n description='Massa $(kg)$',\n layout=Layout(width='80%', height='50px'),\n style={'description_width': '200px'},\n)\nB_slider = widgets.FloatSlider(\n value=0.1,\n min=2,\n max=20.0,\n step=0.1,\n description='Konstanta Redaman $(\\\\frac {Ns}{m})$',\n layout=Layout(width='80%', height='50px'),\n style={'description_width': '200px'},\n)\nK_slider = widgets.FloatSlider(\n value=0.1,\n min=0.1,\n max=100.0,\n step=0.1,\n description='Konstanta pegas $(\\\\frac {N}{m})$',\n layout=Layout(width='80%', height='50px'),\n style={'description_width': '200px'},\n)\nVinType_select = widgets.Dropdown(\n options=[('Impulse', 0), ('Step', 1)],\n description='Tipe Sinyal Input:',\n layout=Layout(width='80%', height='50px'),\n style={'description_width': '200px'},\n)\ngrid_button = widgets.ToggleButton(\n value=True,\n description='Grid',\n icon='check',\n layout=Layout(width='20%', height='50px',margin='10px 10px 10px 350px'),\n style={'description_width': '200px'},\n)\nui_mk = widgets.VBox([M_slider,B_slider,K_slider,VinType_select,grid_button])\nout_mk = widgets.interactive_output(plot_mekanik, {'M':M_slider,'B':B_slider,'K':K_slider,'VinType':VinType_select,'grid':grid_button})",
"_____no_output_____"
],
[
"display(ui_mk,out_mk)",
"_____no_output_____"
]
],
[
[
"### Analisis\nBerdasarkan persamaan yang cukup sederhana, sistem mekanik orde dua memiliki karakteristik berikut:\n#### 1. Pengaruh peningkatan massa\nMassa pada sistem berperilaku seperti komponen inersial yang meningkatkan rise time dan settling time ketika diperbesar.\n#### 2. Pengaruh peningkatan konstanta redaman\nKonstanta redaman berperilaku seperti komponen hambatan yang meredam sistem sehingga maximum overshoot menjadi kecil (akibat peningkatan damping ratio) ketika peningkatan konstanta redaman terjadi. Konstanta redaman juga berpengaruh pada settling time, dimana peningkatan konstanta redaman meningkatkan settling time.\n#### 3. Pengaruh peningkatan konstanta pegas\nKonstanta pegas berperilaku seperti komponen kapasitansi yang mengurangi besar gain dari perpindahan, mengurangi damping ratio, meningkatkan frekuensi osilasi sistem, mengurangi amplitudo kecepatan sistem, mempercepat settling time, dan mempercepat peak time, meningkatkan maximum overshoot.\n#### 4. Respon terhadap impulse\nTerhadap sinyal impulse, sistem mencapai posisi awal kembali dan mencapai steady state 0 untuk perpindahan, kecepatan, dan percepatan\n#### 5. Respon terhadap step\nTerhadap sinyal step, sistem mencapai posisi akhir sesuai \n$$\\frac {F}{k}$$\ndan kecepatan serta percepatan 0",
"_____no_output_____"
]
]
] | [
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown"
] | [
[
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown",
"markdown",
"markdown",
"markdown",
"markdown"
],
[
"code",
"code",
"code",
"code"
],
[
"markdown"
]
] |
d0fc02e0f1f3725911f312092cc78da2f9cd056a | 7,987 | ipynb | Jupyter Notebook | exps/fre BERTNCRF.ipynb | learnerhouse/ner-bert | 606328a27a7313b6c22b78590e06618ad77402cd | [
"MIT"
] | 391 | 2018-12-21T01:00:18.000Z | 2022-03-13T02:05:15.000Z | exps/fre BERTNCRF.ipynb | king-menin/nert-bert | b75a903c35acbd36ff5f26c525e3596294f36815 | [
"MIT"
] | 30 | 2019-01-10T12:58:54.000Z | 2022-01-15T15:51:14.000Z | exps/fre BERTNCRF.ipynb | king-menin/nert-bert | b75a903c35acbd36ff5f26c525e3596294f36815 | [
"MIT"
] | 101 | 2018-12-21T07:40:31.000Z | 2022-03-10T17:47:41.000Z | 21.528302 | 197 | 0.520721 | [
[
[
"%load_ext autoreload\n%autoreload 2\n\n\nimport sys\nimport warnings\n\n\nwarnings.filterwarnings(\"ignore\")\nsys.path.append(\"../\")",
"_____no_output_____"
]
],
[
[
"## IO markup",
"_____no_output_____"
],
[
"### Train",
"_____no_output_____"
]
],
[
[
"from modules.data import bert_data",
"_____no_output_____"
],
[
"train_df_path = \"/home/eartemov/ae/work/factRuEval-2016/dev.csv\"\nvalid_df_path = \"/home/eartemov/ae/work/factRuEval-2016/test.csv\"",
"_____no_output_____"
],
[
"data = bert_data.LearnData.create(\n train_df_path=train_df_path,\n valid_df_path=valid_df_path,\n idx2labels_path=\"/home/eartemov/ae/work/factRuEval-2016/idx2labels.txt\",\n clear_cache=True\n)",
"The pre-trained model you are loading is a cased model but you have not set `do_lower_case` to False. We are setting `do_lower_case=False` for you but you may want to check this behavior.\n"
],
[
"from modules.models.bert_models import BERTNCRF",
"_____no_output_____"
],
[
"model = BERTNCRF.create(len(data.train_ds.idx2label), crf_dropout=0.3, nbest=len(data.train_ds.idx2label)-1)",
"build CRF...\n"
],
[
"from modules.train.train import NerLearner",
"_____no_output_____"
],
[
"num_epochs = 100",
"_____no_output_____"
],
[
"learner = NerLearner(\n model, data, \"/home/eartemov/ae/work/models/fre-BERTNCRF-IO.cpt\", t_total=num_epochs * len(data.train_dl))",
"_____no_output_____"
],
[
"model.get_n_trainable_params()",
"_____no_output_____"
],
[
"learner.fit(epochs=num_epochs)",
"_____no_output_____"
],
[
"learner.load_model()",
"_____no_output_____"
]
],
[
[
"### Predict",
"_____no_output_____"
]
],
[
[
"from modules.data.bert_data import get_data_loader_for_predict",
"_____no_output_____"
],
[
"dl = get_data_loader_for_predict(data, df_path=data.valid_ds.config[\"df_path\"])",
"_____no_output_____"
],
[
"preds = learner.predict(dl)",
"_____no_output_____"
],
[
"from sklearn_crfsuite.metrics import flat_classification_report",
"_____no_output_____"
],
[
"from modules.analyze_utils.utils import bert_labels2tokens, voting_choicer\nfrom modules.analyze_utils.plot_metrics import get_bert_span_report",
"_____no_output_____"
],
[
"pred_tokens, pred_labels = bert_labels2tokens(dl, preds)\ntrue_tokens, true_labels = bert_labels2tokens(dl, [x.bert_labels for x in dl.dataset])",
"_____no_output_____"
],
[
"assert pred_tokens == true_tokens\ntokens_report = flat_classification_report(true_labels, pred_labels, labels=data.train_ds.idx2label[5:], digits=4)",
"_____no_output_____"
],
[
"print(tokens_report)",
" precision recall f1-score support\n\n I_LOC 0.8765 0.7887 0.8303 1557\n I_PER 0.9598 0.9598 0.9598 2112\n I_ORG 0.7946 0.8078 0.8011 3865\n\n micro avg 0.8569 0.8464 0.8516 7534\n macro avg 0.8770 0.8521 0.8637 7534\nweighted avg 0.8578 0.8464 0.8516 7534\n\n"
],
[
"from modules.analyze_utils.main_metrics import precision_recall_f1",
"_____no_output_____"
],
[
"results = precision_recall_f1(true_labels, pred_labels)",
"processed 56409 tokens with 7534 phrases; found: 7442 phrases; correct: 6377.\n\nprecision: 85.69%; recall: 84.64%; FB1: 85.16\n\n\tLOC: precision: 87.65%; recall: 78.87%; F1: 83.03 1401\n\n\tORG: precision: 79.46%; recall: 80.78%; F1: 80.11 3929\n\n\tPER: precision: 95.98%; recall: 95.98%; F1: 95.98 2112\n\n\n"
]
]
] | [
"code",
"markdown",
"code",
"markdown",
"code"
] | [
[
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
d0fc07a8acd71f14c85b4e84eabef0d2514ae37e | 82,673 | ipynb | Jupyter Notebook | PyCitySchools/School_Data.ipynb | Denbigh97/Panda_Challenge | 27f5549f818398e6ba8d06476301a57cb3ee8234 | [
"ADSL"
] | null | null | null | PyCitySchools/School_Data.ipynb | Denbigh97/Panda_Challenge | 27f5549f818398e6ba8d06476301a57cb3ee8234 | [
"ADSL"
] | null | null | null | PyCitySchools/School_Data.ipynb | Denbigh97/Panda_Challenge | 27f5549f818398e6ba8d06476301a57cb3ee8234 | [
"ADSL"
] | null | null | null | 35.634914 | 232 | 0.385737 | [
[
[
"\n# Dependencies and Setup\nimport pandas as pd\n\n# File to Load\nschool_data_to_load = \"Resources/schools_complete.csv\"\nstudent_data_to_load = \"Resources/students_complete.csv\"\n\n# Read School and Student Data File and store into Pandas Data Frames\nschool_data = pd.read_csv(school_data_to_load)\nstudent_data = pd.read_csv(student_data_to_load)\n\n# Combine the data into a single dataset\nschool_data_complete = pd.merge(student_data, school_data, how=\"left\", on=[\"school_name\", \"school_name\"])\nschool_data_complete",
"_____no_output_____"
]
],
[
[
"#District Summary\nCalculate the total number of schools\n\nCalculate the total number of students\n\nCalculate the total budget\n\nCalculate the average math score\n\nCalculate the average reading score\n\nCalculate the overall passing rate (overall average score), i.e. (avg. math score + avg. reading score)/2\n\nCalculate the percentage of students with a passing math score (70 or greater)\n\nCalculate the percentage of students with a passing reading score (70 or greater)\n\nCreate a dataframe to hold the above results\n\nOptional: give the displayed data cleaner formatting\n",
"_____no_output_____"
]
],
[
[
"total_number_schools = len(school_data_complete[\"School ID\"].unique())\ntotal_number_schools\n",
"_____no_output_____"
],
[
"#Calculate the total number of students\ntotal_number_students = len(school_data_complete[\"Student ID\"].unique())\ntotal_number_students",
"_____no_output_____"
],
[
"#Calculate the total budget\ntotal_budget = school_data[\"budget\"].sum()\ntotal_budget\n",
"_____no_output_____"
],
[
"#Calculate the average math score\naverage_math_score = student_data[\"math_score\"].mean()\naverage_math_score",
"_____no_output_____"
],
[
"#Calculate the average reading score\naverage_reading_score = student_data[\"reading_score\"].mean()\naverage_reading_score",
"_____no_output_____"
],
[
"#Calculate the overall passing rate (overall average score), i.e. (avg. math score + avg. reading score)/2\noverall_average_score = (average_math_score + average_reading_score)/2\noverall_average_score",
"_____no_output_____"
],
[
"#Calculate the percentage of students with a passing math score (70 or greater)\n#Create 1s for the passing students by math or reading. 0 otherwise. And take the mean to get the average. It will give the percent of the passing students.\nstudent_data[\"#passing_math\"] = student_data[\"math_score\"] >= 70\nstudent_data[\"#passing_reading\"] = student_data[\"reading_score\"] >= 70\npercent_passing_math = ((student_data[\"#passing_math\"]).mean())*100\npercent_passing_math",
"_____no_output_____"
],
[
"#Calculate the percentage of students with a passing reading score (70 or greater)\npercent_passing_reading = ((student_data[\"#passing_reading\"]).mean())*100\npercent_passing_reading",
"_____no_output_____"
],
[
"#Calculate overall percentage\noverall_passing_rate = (percent_passing_math + percent_passing_reading)/2\noverall_passing_rate",
"_____no_output_____"
],
[
"#Create a dataframe to hold the above results\n#Optional: give the displayed data cleaner formatting\ndistrict_results = [{\"Total Schools\": total_number_schools, \n \"Total Students\": total_number_students, \n \"Total Budget\": total_budget, \n \"Average Math Score\": round(average_math_score,2), \n \"Average Reading Score\": round(average_reading_score,2), \n \"% Passing Math\": round(percent_passing_math,2),\n \"% Passing Reading\": round(percent_passing_reading,2),\n \"% Overall Passing Rate\": round(overall_passing_rate,2)}]\ndistrict_summary_table = pd.DataFrame(district_results)\n\n#Formatting\ndistrict_summary_table[\"% Passing Math\"] = district_summary_table[\"% Passing Math\"].map(\"{:,.2f}%\".format)\ndistrict_summary_table[\"% Passing Reading\"] = district_summary_table[\"% Passing Reading\"].map(\"{:,.2f}%\".format)\ndistrict_summary_table[\"% Overall Passing Rate\"] = district_summary_table[\"% Overall Passing Rate\"].map(\"{:,.2f}%\".format)\ndistrict_summary_table[\"Total Budget\"] = district_summary_table[\"Total Budget\"].map(\"${:,.2f}\".format)\ndistrict_summary_table[\"Total Students\"] = district_summary_table[\"Total Students\"].map(\"{:,}\".format)\n\n#Display\ndistrict_summary_table",
"_____no_output_____"
]
],
[
[
"#School Summary\nCreate an overview table that summarizes key metrics about each school, including:\n\nSchool Name\nSchool Type\nTotal Students\nTotal School Budget\nPer Student Budget\nAverage Math Score\nAverage Reading Score\n% Passing Math\n% Passing Reading\nOverall Passing Rate (Average of the above two)\nCreate a dataframe to hold the above results",
"_____no_output_____"
]
],
[
[
"\n#For this part, school_data_complete\n\nschool_data_complete[\"passing_math\"] = school_data_complete[\"math_score\"] >= 70\nschool_data_complete[\"passing_reading\"] = school_data_complete[\"reading_score\"] >= 70\n\nschool_data_complete",
"_____no_output_____"
],
[
"# Use groupby by school_name\n\nschool_group = school_data_complete.groupby([\"school_name\"]).mean()\nschool_group[\"Per Student Budget\"] = school_group[\"budget\"]/school_group[\"size\"]\nschool_group[\"% Passing Math\"] = round(school_group[\"passing_math\"]*100,2)\nschool_group[\"% Passing Reading\"] = round(school_group[\"passing_reading\"]*100,2)\nschool_group[\"% Overall Passing Rate\"] = round(((school_group[\"passing_math\"] + school_group[\"passing_reading\"])/2)*100,3)\n\n#Merge with school_data to collect information about the type, size and budget\nschool_data_summary = pd.merge(school_group, school_data, how=\"left\", on=[\"school_name\", \"school_name\"])\ndel school_data_summary['size_y']\ndel school_data_summary['budget_y']\ndel school_data_summary['Student ID']\ndel school_data_summary['School ID_x']\n\n#Create a dataframe to store the results\nschool_summary_dataframe = pd.DataFrame({\"School Name\": school_data_summary[\"school_name\"],\n \"School Type\": school_data_summary[\"type\"],\n \"Total Students\":school_data_summary[\"size_x\"],\n \"Total School Budget\": school_data_summary[\"budget_x\"],\n \"Per Student Budget\":school_data_summary[\"Per Student Budget\"], \n \"Average Math Score\":round(school_data_summary[\"math_score\"],2),\n \"Average Reading Score\":round(school_data_summary[\"reading_score\"],2), \n \"% Passing Math\": school_data_summary[\"% Passing Math\"],\n \"% Passing Reading\": school_data_summary[\"% Passing Reading\"],\n \"% Overall Passing Rate\": school_data_summary[\"% Overall Passing Rate\"]}) \n\n#Formatting\nschool_summary_dataframe[\"Total Students\"] = school_summary_dataframe[\"Total Students\"].map(\"{:,.0f}\".format)\nschool_summary_dataframe[\"Total School Budget\"] = school_summary_dataframe[\"Total School Budget\"].map(\"${:,.2f}\".format)\nschool_summary_dataframe[\"Per Student Budget\"] = school_summary_dataframe[\"Per Student Budget\"].map(\"${:,.2f}\".format)\n#Display\nschool_summary_dataframe",
"_____no_output_____"
]
],
[
[
"#Top Performing Schools (By Passing Rate)\nSort and display the top five schools in overall passing rate",
"_____no_output_____"
]
],
[
[
"#Sort and display the top five schools in overall passing rate\n\ntop_five_schools = school_summary_dataframe.sort_values([\"% Overall Passing Rate\"], ascending=False)\ntop_five_schools.head()",
"_____no_output_____"
]
],
[
[
"#Bottom Performing Schools (By Passing Rate)¶\nSort and display the five worst-performing schools",
"_____no_output_____"
]
],
[
[
"#Sort and display the five worst-performing schools\n\nbottom_five_schools = school_summary_dataframe.sort_values([\"% Overall Passing Rate\"], ascending=True)\nbottom_five_schools.head()",
"_____no_output_____"
]
],
[
[
"#Math Scores by Grade\nCreate a table that lists the average Reading Score for students of each grade level (9th, 10th, 11th, 12th) at each school.\n\nCreate a pandas series for each grade. Hint: use a conditional statement.\n\nGroup each series by school\n\nCombine the series into a dataframe\n\nOptional: give the displayed data cleaner formatting",
"_____no_output_____"
]
],
[
[
"#Create a pandas series for each grade. Group each series by school.\n\nnineth_grade= school_data_complete[school_data_complete[\"grade\"] == \"9th\"].groupby(\"school_name\").mean()[\"math_score\"]\ntenth_grade = school_data_complete[school_data_complete[\"grade\"] == \"10th\"].groupby(\"school_name\").mean()[\"math_score\"]\neleventh_grade = school_data_complete[school_data_complete[\"grade\"] == \"11th\"].groupby(\"school_name\").mean()[\"math_score\"]\ntwelveth_grade= school_data_complete[school_data_complete[\"grade\"] == \"12th\"].groupby(\"school_name\").mean()[\"math_score\"]\n\n#Combine the series into a dataframe\nmath_grade_dataframe = pd.DataFrame({\"Ninth Grade\":nineth_grade, \"Tenth Grade\":tenth_grade, \n \"Eleventh Grade\":eleventh_grade, \"Twelveth Grade\":twelveth_grade}) \n\n#Optional formatting: Give the displayed data cleaner formatting\n\nmath_grade_dataframe[[\"Ninth Grade\",\"Tenth Grade\",\"Eleventh Grade\",\"Twelveth Grade\"]] = math_grade_dataframe[[\"Ninth Grade\",\"Tenth Grade\",\"Eleventh Grade\",\"Twelveth Grade\"]].applymap(\"{:.2f}\".format)\n\n#Display\n\nmath_grade_dataframe",
"_____no_output_____"
]
],
[
[
"#Reading Score by Grade\nPerform the same operations as above for reading scores",
"_____no_output_____"
]
],
[
[
"#Perform the same operations as above for reading scores\n#Create a pandas series for each grade. Group each series by school.\n\nnineth_grade= school_data_complete[school_data_complete[\"grade\"] == \"9th\"].groupby(\"school_name\").mean()[\"reading_score\"]\ntenth_grade = school_data_complete[school_data_complete[\"grade\"] == \"10th\"].groupby(\"school_name\").mean()[\"reading_score\"]\neleventh_grade = school_data_complete[school_data_complete[\"grade\"] == \"11th\"].groupby(\"school_name\").mean()[\"reading_score\"]\ntwelveth_grade= school_data_complete[school_data_complete[\"grade\"] == \"12th\"].groupby(\"school_name\").mean()[\"reading_score\"]\n\n#Combine the series into a dataframe\nreading_grade_dataframe = pd.DataFrame({\"Ninth Grade\":nineth_grade, \"Tenth Grade\":tenth_grade, \n \"Eleventh Grade\":eleventh_grade, \"Twelveth Grade\":twelveth_grade}) \n\n#Optional formatting: Give the displayed data cleaner formatting\n\nreading_grade_dataframe[[\"Ninth Grade\",\"Tenth Grade\",\"Eleventh Grade\",\"Twelveth Grade\"]] = reading_grade_dataframe[[\"Ninth Grade\",\"Tenth Grade\",\"Eleventh Grade\",\"Twelveth Grade\"]].applymap(\"{:.2f}\".format)\n\n#Display\n\nreading_grade_dataframe",
"_____no_output_____"
]
],
[
[
"#Scores by School Spending\nCreate a table that breaks down school performances based on average Spending Ranges (Per Student). Use 4 reasonable bins to group school spending. Include in the table each of the following:\nAverage Math Score\nAverage Reading Score\n% Passing Math\n% Passing Reading\nOverall Passing Rate (Average of the above two)",
"_____no_output_____"
]
],
[
[
"\n# Sample bins. Feel free to create your own bins.\nspending_bins = [0, 585, 615, 645, 675]\ngroup_names = [\"<$585\", \"$585-615\", \"$615-645\", \"$645-675\"]\n\n# Use 4 reasonable bins to group school spending. \nschool_data_summary[\"Spending Ranges (Per Student)\"] = pd.cut(school_data_summary[\"Per Student Budget\"], spending_bins, labels=group_names)\n\nschool_spending_grouped = school_data_summary.groupby(\"Spending Ranges (Per Student)\").mean() \n\n#Remove the unwanted columns as per the sample provided\ndel school_spending_grouped['size_x']\ndel school_spending_grouped['budget_x']\ndel school_spending_grouped['Per Student Budget']\ndel school_spending_grouped['School ID_y']\ndel school_spending_grouped['passing_math']\ndel school_spending_grouped['passing_reading']\n\nschool_spending_grouped",
"_____no_output_____"
]
],
[
[
"#Scores by School Size¶\nPerform the same operations as above, based on school size.",
"_____no_output_____"
]
],
[
[
"# Sample bins. Feel free to create your own bins.\nsize_bins = [0, 1000, 2000, 5000]\ngroup_names = [\"Small (<1000)\", \"Medium (1000-2000)\", \"Large (2000-5000)\"]\n\n# Use 4 reasonable bins to group school size. \nschool_data_summary[\"School Size\"] = pd.cut(school_data_summary[\"size_x\"], size_bins, labels=group_names)\nschool_data_summary\n\n#group by size_x\nschool_size_grouped = school_data_summary.groupby(\"School Size\").mean() \nschool_size_grouped\n\n#Remove the unwanted columns as per the sample provided\n#del school_size_grouped['size_x']\ndel school_size_grouped['budget_x']\ndel school_size_grouped['Per Student Budget']\ndel school_size_grouped['School ID_y']\ndel school_size_grouped['passing_math']\ndel school_size_grouped['passing_reading']\n\n#Display\nschool_size_grouped",
"_____no_output_____"
]
],
[
[
"#Scores by School Type\nPerform the same operations as above, based on school type.",
"_____no_output_____"
]
],
[
[
"school_type_grouped = school_data_summary.groupby(\"type\").mean()\n\n#Remove the unwanted columns as per the sample provided\ndel school_type_grouped['size_x']\ndel school_type_grouped['budget_x']\ndel school_type_grouped['Per Student Budget']\ndel school_type_grouped['School ID_y']\ndel school_type_grouped['passing_math']\ndel school_type_grouped['passing_reading']\n\nschool_type_grouped",
"_____no_output_____"
]
],
[
[
"#You must include a written description of at least two observable trends based on the data\n",
"_____no_output_____"
]
]
] | [
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown"
] | [
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
]
] |
d0fc0e914021f1d63fc0eb19dd6fe799a9a8a55a | 469,881 | ipynb | Jupyter Notebook | _notebooks/2021-10-26,28-{fastai_CNN}.ipynb | chchin33/violet | 048b4c2d3898252bc2dcf01dbf6e8e603509eb3c | [
"Apache-2.0"
] | null | null | null | _notebooks/2021-10-26,28-{fastai_CNN}.ipynb | chchin33/violet | 048b4c2d3898252bc2dcf01dbf6e8e603509eb3c | [
"Apache-2.0"
] | null | null | null | _notebooks/2021-10-26,28-{fastai_CNN}.ipynb | chchin33/violet | 048b4c2d3898252bc2dcf01dbf6e8e603509eb3c | [
"Apache-2.0"
] | null | null | null | 81.195956 | 38,028 | 0.752582 | [
[
[
"# Convolution_with_fastai\n> 2021-10-26\n\n- toc: true \n- badges: true\n- comments: false\n- categories: bigdata\n- image: images/chart-preview.png\n- hide: true",
"_____no_output_____"
]
],
[
[
"import torch\nfrom fastai.vision.all import *",
"_____no_output_____"
]
],
[
[
"#### data",
"_____no_output_____"
]
],
[
[
"path = untar_data(URLs.MNIST_SAMPLE)",
"_____no_output_____"
],
[
"path.ls()",
"_____no_output_____"
]
],
[
[
"`-` list 형태로 목록 받기",
"_____no_output_____"
]
],
[
[
"threes = (path/'train'/'3').ls()\nsevens = (path/'train'/'7').ls()",
"_____no_output_____"
]
],
[
[
"`-` list $\\to$ image",
"_____no_output_____"
]
],
[
[
"Image.open(threes[0])",
"_____no_output_____"
]
],
[
[
"`-` image $\\to$ tensor",
"_____no_output_____"
]
],
[
[
"tensor(Image.open(threes[0]))",
"_____no_output_____"
]
],
[
[
"`*` 여기서 tensor는 pytorch에 있는 tensor가 아닌, fastai에 있는 tensor이다. `*` , 만약 pytorch의 tensor였다면 torch.tensor로 사용했을 것이다.\n\ntorch.tensor는 이미지파일을 텐서로 변환하는 기능이 없다.",
"_____no_output_____"
]
],
[
[
"# plt.imshow(tensor(Image.open(threes[0]))) 변환한 텐서로 이미지 출력",
"_____no_output_____"
],
[
"#[tensor(Image.open(i)) for i in sevens] # tensor들이 list로 바뀌어있다. 현재 이 상태는 것이 list 안이 tensor로, 좀 더 깔끔하게 만들면\n#torch.stack([tensor(Image.open(i)) for i in sevens]) # tensor안의 값들은 float여야 하므로\nseven_tensor = torch.stack([tensor(Image.open(i)) for i in sevens]).float()\nthree_tensor = torch.stack([tensor(Image.open(i)) for i in threes]).float()\n",
"_____no_output_____"
],
[
"seven_tensor = torch.stack([tensor(Image.open(i)) for i in sevens]).float()/255\nthree_tensor = torch.stack([tensor(Image.open(i)) for i in threes]).float()/255",
"_____no_output_____"
],
[
"seven_tensor.shape, three_tensor.shape",
"_____no_output_____"
],
[
"y = torch.tensor([0.]*6265 + [1.]*6131).reshape(12396,1) # 3인지 7인지 정답 label을 0, 1로 만듬",
"_____no_output_____"
]
],
[
[
"`-` 데이터인 X는 seven_tensor와 three_tensor를 합친다. (vstack 으로)",
"_____no_output_____"
]
],
[
[
"X = torch.vstack([seven_tensor, three_tensor])",
"_____no_output_____"
],
[
"X = X.reshape(12396,-1)\nX.shape",
"_____no_output_____"
],
[
"X.shape\nX = X.reshape(12396,1,28,28)",
"_____no_output_____"
]
],
[
[
"### 1. 지난시간까지 모형 (네트워크 직접 설계, pytorch)",
"_____no_output_____"
],
[
"#### 선형변환 대신에 2D Convolution with Window_size = 5",
"_____no_output_____"
],
[
"- 인자를 보면 in-channels, out-channels, 그리고 kernel-size가 있다.<br/>\n이 세 개를 넣어야 한다.",
"_____no_output_____"
]
],
[
[
"c1 = torch.nn.Conv2d(1,16,5) # 입력채널 = 1, 출력 채널 = 16, window_size = 5",
"_____no_output_____"
]
],
[
[
"#### NonLinear를 위해서, ReLU()대신 MaxPool2d + ReLU 를 함, MaxPooling을 하나 걸침",
"_____no_output_____"
]
],
[
[
"m1 = torch.nn.MaxPool2d(2)",
"_____no_output_____"
]
],
[
[
"### ReLU()",
"_____no_output_____"
]
],
[
[
"a1 = torch.nn.ReLU()",
"_____no_output_____"
],
[
"X.shape, c1(X).shape, m1(c1(X)).shape, a1(m1(c1(X))).shape",
"C:\\Users\\USER\\anaconda3\\envs\\bda2021\\lib\\site-packages\\torch\\nn\\functional.py:718: UserWarning: Named tensors and all their associated APIs are an experimental feature and subject to change. Please do not use them for anything important until they are released as stable. (Triggered internally at ..\\c10/core/TensorImpl.h:1156.)\n return torch.max_pool2d(input, kernel_size, stride, padding, dilation, ceil_mode)\n"
]
],
[
[
"### Flatten",
"_____no_output_____"
]
],
[
[
"class Flatten(torch.nn.Module): # Module 상속\n def forward(self,x):\n return x.reshape(12396,-1)",
"_____no_output_____"
]
],
[
[
"- a1을 거쳐서 들어온 것을 a1(m1(c1(X))).reshape(12396,-1) 이렇게 바꿔주어야 한다.",
"_____no_output_____"
]
],
[
[
"flatten = Flatten()",
"_____no_output_____"
],
[
"X.shape, c1(X).shape, m1(c1(X)).shape, a1(m1(c1(X))).shape, flatten(a1(m1(c1(X)))).shape",
"_____no_output_____"
]
],
[
[
"#### linear",
"_____no_output_____"
]
],
[
[
"l1 = torch.nn.Linear(in_features=2304, out_features=1)",
"_____no_output_____"
],
[
"X.shape, c1(X).shape, \\\nm1(c1(X)).shape, \\\na1(m1(c1(X))).shape, \\\nflatten(a1(m1(c1(X)))).shape, \\\nl1(flatten(a1(m1(c1(X))))).shape",
"_____no_output_____"
],
[
"plt.plot(l1(flatten(a1(m1(c1(X))))).data)",
"_____no_output_____"
]
],
[
[
"- 학습이 안된상태",
"_____no_output_____"
]
],
[
[
"net = torch.nn.Sequential(\n c1, # Convolution(선형)\n m1, # MaxPooling(비선형)\n a1, # RelU(비선형)\n flatten,\n l1)",
"_____no_output_____"
]
],
[
[
"`-` 손실함수와 옵티마이저 정의",
"_____no_output_____"
]
],
[
[
"loss_fn = torch.nn.BCEWithLogitsLoss()\noptimizer = torch.optim.Adam(net.parameters())",
"_____no_output_____"
]
],
[
[
"`-` step 1~4",
"_____no_output_____"
]
],
[
[
"for epoc in range(200):\n # 1\n yhat = net(X)\n # 2\n loss = loss_fn(yhat,y)\n # 3\n loss.backward()\n # 4\n optimizer.step()\n net.zero_grad()",
"_____no_output_____"
],
[
"a2 = torch.nn.Sigmoid()",
"_____no_output_____"
],
[
"plt.plot(y)\nplt.plot(a2(yhat.data),'.') # 마지막 sigmoid를 취한 버전",
"_____no_output_____"
],
[
"ypred = a2(yhat.data)>0.5",
"_____no_output_____"
],
[
"sum(ypred == y) / 12396",
"_____no_output_____"
]
],
[
[
"### 2. 드랍아웃, 배치추가 (직접네트워크 설계, pytorch + fastai)",
"_____no_output_____"
],
[
"#### step1 : dls를 만들자",
"_____no_output_____"
]
],
[
[
"ds = torch.utils.data.TensorDataset(X,y)",
"_____no_output_____"
],
[
"ds.tensors[0].shape #이미지 자체가 들어간 것, 784개 벡터로 늘려지는게 아니라",
"_____no_output_____"
]
],
[
[
"`-` training, validation 분리",
"_____no_output_____"
]
],
[
[
"ds1, ds2 = torch.utils.data.random_split(ds,[10000,2396]) # training을 10000개, validation을 2396개로 ds로부터 랜덤하게 나눔",
"_____no_output_____"
],
[
"dl1 = torch.utils.data.DataLoader(ds1, batch_size=500) # DataLoader는 batch size를 정할 수 있다. -> 총 1만개 에서 500개를 나누므로, for문 한 번당 20번 돌아감\ndl2 = torch.utils.data.DataLoader(ds2, batch_size=2396) # 여기선 딱히 batch를 나누지 않았다.",
"_____no_output_____"
]
],
[
[
"`-` dataloader를 만들었으니 이제 DataLoaders를 만들어야 한다. ( 여태 pytorch로 작업한 것이였고 여기부터는 fastai로 작업한다. )",
"_____no_output_____"
]
],
[
[
"dls = DataLoaders(dl1,dl2)",
"_____no_output_____"
]
],
[
[
"- 여기까지 한 것이 데이터 정리TensorDataset",
"_____no_output_____"
],
[
"#### step2: 아키텍처, 손실함수, 옵티마이저",
"_____no_output_____"
]
],
[
[
"class Flatten(torch.nn.Module): # Module 상속\n def forward(self,x):\n return x.reshape(x.shape[0],-1)",
"_____no_output_____"
],
[
"net = torch.nn.Sequential(\n torch.nn.Conv2d(1,16,5),\n torch.nn.MaxPool2d(2),\n torch.nn.ReLU(),\n torch.nn.Dropout2d(),\n Flatten(),\n torch.nn.Linear(2304,1)\n)",
"_____no_output_____"
],
[
"loss_fn = torch.nn.BCEWithLogitsLoss()\n# optimizer = torch.optim.Adam(net.parameters()) learner에서 옵션으로 넣기 때문에 여기선 딱히 정의하지 않음, (loss도 옵션으로 넣을 순 있는데 일단 두가지 경우 모두 보기 위함)",
"_____no_output_____"
]
],
[
[
"#### step3: lrnr 생성",
"_____no_output_____"
],
[
"`-` Learner 클래스에서 learner instance 생성",
"_____no_output_____"
]
],
[
[
"lrnr1 = Learner(dls, net, opt_func=Adam, loss_func=loss_fn)",
"_____no_output_____"
]
],
[
[
"- 이것의 역할: \n 1. for 문 돌리면서 step 1~4 하던 것을 자동으로 해주는 역할\n 2. batch를 dls가 알아서 나눠줌\n 3. gpu에 자동으로 올려줌",
"_____no_output_____"
],
[
"`-` 우리가 할 일은 for문을 몇 번 돌려줄지 결정하는 것",
"_____no_output_____"
]
],
[
[
"lrnr1.fit(10)",
"_____no_output_____"
]
],
[
[
"`-` GPU에 올린 것을 알아둬야 한다. 이는 Shape와 같이 자주 신경써야 하는 부분이다.",
"_____no_output_____"
],
[
"`-` 현재 networks의 parameter가 GPU에 올라가있기 때문에, 데이터 X도 GPU에 올려서 계산을 한다(아니면 parameter를 CPU로 내려도 되는데 계산속도가 GPU보다 느릴 것으로 예상된다)",
"_____no_output_____"
]
],
[
[
"# net(X.to(\"cuda:0\")).to(\"cpu\").data",
"_____no_output_____"
]
],
[
[
"- 다시 그림을 그리는 plot 을 사용하려면 계산한 결과를 cpu로 내려서 그려야 한다.",
"_____no_output_____"
]
],
[
[
"plt.plot(net(X.to(\"cuda:0\")).to(\"cpu\").data,'.')",
"_____no_output_____"
],
[
"plt.plot(a2(net(X.to(\"cuda:0\")).to(\"cpu\").data),'.')",
"_____no_output_____"
]
],
[
[
"- Sigmoid 적용",
"_____no_output_____"
],
[
"`-` 빠르고 적합 결과도 좋음",
"_____no_output_____"
],
[
"### 3. resnet34 사용 (기존의 네트워크 사용, 순수 fastai)",
"_____no_output_____"
],
[
"`-` 데이터로부터 새로운 데이터로더스를 만들고 이를 dls2라 한다.",
"_____no_output_____"
]
],
[
[
"path = untar_data(URLs.MNIST_SAMPLE)\npath",
"_____no_output_____"
],
[
"path.ls()",
"_____no_output_____"
]
],
[
[
"- 여기서 train부분의 폴더에 접근해서 사용할 것이다.",
"_____no_output_____"
]
],
[
[
"dls2 = ImageDataLoaders.from_folder(\n path,\n train = 'train',\n valid_pct = 0.2\n)",
"Due to IPython and Windows limitation, python multiprocessing isn't available now.\nSo `number_workers` is changed to 0 to avoid getting stuck\n"
]
],
[
[
"`-` Learner를 CNN전용으로 만들어 둔 것이 있다.",
"_____no_output_____"
],
[
"`-` learn오브젝트를 생성하고 학습",
"_____no_output_____"
]
],
[
[
"lrnr2 = cnn_learner(dls2, resnet34, metrics = error_rate)\nlrnr2.fine_tune(1)",
"_____no_output_____"
]
],
[
[
"`-` 결과 관찰",
"_____no_output_____"
]
],
[
[
"lrnr2.show_results()",
"_____no_output_____"
]
],
[
[
"### 모형을 뜯어보는 방법 (lrnr1)",
"_____no_output_____"
],
[
"`-` 우선 방법 2로 돌아가자",
"_____no_output_____"
],
[
"`-` 네트워크 구조",
"_____no_output_____"
]
],
[
[
"net",
"_____no_output_____"
]
],
[
[
"`-` 인덱싱을 통하여 각각의 layer에 적용할 수 있다.",
"_____no_output_____"
]
],
[
[
"net[0]",
"_____no_output_____"
]
],
[
[
"- c1 = net[0] (torch.nn.Conv2d)에 해당하는 형태이다.",
"_____no_output_____"
]
],
[
[
"net.to(\"cpu\")",
"_____no_output_____"
]
],
[
[
"- 먼저 GPU 성능 문제로 인한 에러때문에 net을 cpu로 옮겼다.",
"_____no_output_____"
],
[
"`-` 각각의 layer에서의 중간 결과를 가로챌 수 있다. <br/>\n`-` 층별 변환과정",
"_____no_output_____"
]
],
[
[
"print(X.shape, '-> Input Image')\nprint(net[0](X).shape, '-> Conv2D')\nprint(net[1](net[0](X)).shape, '-> MaxPool2D')\nprint(net[2](net[1](net[0](X))).shape, '-> ReLU')\nprint(net[3](net[2](net[1](net[0](X)))).shape, '-> DropOut2D')\nprint(net[4](net[3](net[2](net[1](net[0](X))))).shape, '-> Flatten')\nprint(net[5](net[4](net[3](net[2](net[1](net[0](X)))))).shape, '-> Linear')",
"torch.Size([12396, 1, 28, 28]) -> Input Image\ntorch.Size([12396, 16, 24, 24]) -> Conv2D\ntorch.Size([12396, 16, 12, 12]) -> MaxPool2D\ntorch.Size([12396, 16, 12, 12]) -> ReLU\ntorch.Size([12396, 16, 12, 12]) -> DropOut2D\ntorch.Size([12396, 2304]) -> Flatten\ntorch.Size([12396, 1]) -> Linear\n"
]
],
[
[
"- 중간의 각각 layer들을 연속해서 적용한 결과, 실제 network 전체에 대입한(연속적인 계산) 결과와 동일한 결과를 얻는다.",
"_____no_output_____"
],
[
"`-` 두 결과가 일치하는지 확인",
"_____no_output_____"
]
],
[
[
"print(net(X))\nprint(net[5](net[4](net[3](net[2](net[1](net[0](X)))))))",
"tensor([[ -8.9509],\n [-10.6344],\n [ -5.5692],\n ...,\n [ 15.4405],\n [ 8.6353],\n [ 6.3605]], grad_fn=<AddmmBackward>)\ntensor([[ -8.9509],\n [-10.6344],\n [ -5.5692],\n ...,\n [ 15.4405],\n [ 8.6353],\n [ 6.3605]], grad_fn=<AddmmBackward>)\n"
]
],
[
[
"- 일치한다.",
"_____no_output_____"
],
[
"`-` lrnr1 자체를 활용해도 층별변환과정을 추적함 lrnr1 = net임",
"_____no_output_____"
]
],
[
[
"lrnr1.model",
"_____no_output_____"
]
],
[
[
"- net과 동일한 결과",
"_____no_output_____"
],
[
"`-` net과 동일한 신경망으로의 입력 및 출력, 층별 가로채기",
"_____no_output_____"
]
],
[
[
"lrnr1.model(X)\nlrnr1.model[0](X)",
"_____no_output_____"
]
],
[
[
"- net과 동일한 결과이다.",
"_____no_output_____"
],
[
"`-` 위의 net부분에서 층별 변환과정을 lrnr1으로 하려면, net을 lrnr1.model으로 바꾸면 된다.",
"_____no_output_____"
]
],
[
[
"print(X.shape, '-> 입력')\nprint(lrnr1.model[0](X).shape, '-> Conv2D')\nprint(lrnr1.model[1](lrnr1.model[0](X)).shape, '-> MaxPool2D')\n\n'''\n...\n'''\n\nprint(lrnr1.model[5](lrnr1.model[4](lrnr1.model[3](lrnr1.model[2](lrnr1.model[1](lrnr1.model[0](X)))))).shape, '-> Linear')",
"torch.Size([12396, 1, 28, 28]) -> 입력\ntorch.Size([12396, 16, 24, 24]) -> Conv2D\ntorch.Size([12396, 16, 12, 12]) -> MaxPool2D\ntorch.Size([12396, 1]) -> Linear\n"
]
],
[
[
"`-` 정리: 모형은 항상 아래와 같이 2d-part와 1d-part로 나뉜다.",
"_____no_output_____"
],
[
"```\n\ntorch.Size([12396, 1, 28, 28]) -> Input Image\ntorch.Size([12396, 16, 24, 24]) -> Conv2D\ntorch.Size([12396, 16, 12, 12]) -> MaxPool2D\ntorch.Size([12396, 16, 12, 12]) -> ReLU\ntorch.Size([12396, 16, 12, 12]) -> DropOut2D\n===================================================\ntorch.Size([12396, 2304]) -> Flatten\ntorch.Size([12396, 1]) -> Linear\n\n\n```",
"_____no_output_____"
],
[
"`-` 2d-part:\n- 2d 선형변환: torch.nn.Conv2d()\n- 2d 비선형변환: torch.nn.MaxPool2d(), torch.nn.ReLU()",
"_____no_output_____"
],
[
"`-` 1d-part:\n- 1d 선형변환: torch.nn.Linear()\n- 1d 비선형변환: torch.nn.ReLU()",
"_____no_output_____"
],
[
"`-` **또 다른 정리법** -> 이는 좀 더 계층적인 정리방법이다.",
"_____no_output_____"
]
],
[
[
"_net1 = torch.nn.Sequential(\n net[0],\n net[1],\n net[2],\n net[3]\n)\n\n_net2 = torch.nn.Sequential(\n net[4],\n net[5]\n)",
"_____no_output_____"
],
[
"_net1",
"_____no_output_____"
],
[
"_net2",
"_____no_output_____"
],
[
"_net = torch.nn.Sequential(_net1, _net2)",
"_____no_output_____"
],
[
"_net",
"_____no_output_____"
]
],
[
[
"- 2d part와 1d part로 나누었다 ",
"_____no_output_____"
]
],
[
[
"_net[0]",
"_____no_output_____"
],
[
"_net[0](X)",
"_____no_output_____"
]
],
[
[
"- 위의 _net$[$0$]$(X)의 결과는 2D part의 결과를 의미한다.",
"_____no_output_____"
],
[
"### lrnr2.model (resnet34) 분석",
"_____no_output_____"
]
],
[
[
"lrnr2.model",
"_____no_output_____"
]
],
[
[
"- 여기서 크게 (0) Sequential part와 (1) Sequential part로 나뉘었음을 알 수 있다. <br/>\n이는 각각 2d 와 1d part를 의미한다.",
"_____no_output_____"
],
[
"`-` 2d part",
"_____no_output_____"
]
],
[
[
"lrnr2.model[0]",
"_____no_output_____"
]
],
[
[
"`-` 1d part",
"_____no_output_____"
]
],
[
[
"lrnr2.model[1]",
"_____no_output_____"
]
],
[
[
"#### 1d part 살펴보기",
"_____no_output_____"
],
[
"먼저 이미지를 받아서 Pooling을 하는데, AveragePooling이랑 MaxPooling이라는 것을 함 (Adaptive는 잘 모르겠음)<br/>\n그리고 Flatten()으로 펼쳐줌 (우리가 정의한 Flatten이 아님)<br/>\n펼친 layer의 차원이 1024임을 알 수 있음. 그리고 batch normalization이라는 것이 진행이 됨<br/>\nLinear Transform으로 1024를 512로 한 번 줄여줌<br/>\n그리고 ReLU하고 다시 Batch normarlization, Dropout, Linear를 반복한다는 것",
"_____no_output_____"
],
[
"`-` 주목할 만한 점은, BatchNormalization이라는 것(이것도 Dropout 처럼 과적합을 피하기 위한 것으로 dropout과 같이 많이 사용됨)과 출력의 차원이 2라는 점이다.",
"_____no_output_____"
],
[
"**2d part 간략하게 살펴보기**",
"_____no_output_____"
],
[
"`-` 아래 모형은 현재 가장 성능이 좋은 모형(state of the art)중 하나인 resnet이다.",
"_____no_output_____"
]
],
[
[
"lrnr2.model[0]",
"_____no_output_____"
]
],
[
[
"크게 주목할만한 점: <br/>\n1. Batch Normalization이 중간중간에 들어가 있다는 것.<br/>\n2. 2D version의 Dropout이 없다는 것\n3. Conv2d에 더 많은 argument (padding/ stride)들을 사용\n4. Conv2d의 입력채널이 3원색이라서 3차원 채널인 점",
"_____no_output_____"
],
[
"`-` 입력채널이 3이라는 점 때문에 lrnr2에서 dls를 그대로 사용하면 안되고, dls2를 따로 만든 것이다.<br/> \n우리가 전에 만든 net은 입력채널이 1이였음, 그런데 resnet은 3이라서 그에 맞게 데이터를 다시 정리해줘야됨",
"_____no_output_____"
],
[
"`-` DLS, networks\n- 네트워크 형태에 따라서 dls의 형태도 다르게 만들어야 한다.\n- MLP 모형: 입력이 $784$, 첫 네트워크의 형태가 $ 784 \\to 30$ 인 torch.nn.Linear()\n- CNN 모형: 입력이 $1 \\times 28 \\times 28$, 첫 네트워크 형태가 $ 1 \\times 28 \\times 28 \\to 16 \\times 24 \\times 24$ 인 torch.nn.Conv2d()\n- resnet34 모형: 입력이 $3 \\times 28 \\times 28$, 첫 네트워크 형태가 $ 3 \\times 28 \\times 28 \\to ??$",
"_____no_output_____"
],
[
"데이터 만들기:\n1. x,y 각각 정의\n2. x,y를 tuple로 묶어서 dataset 생성\n3. dataset을 training, validaiton으로 분리\n4. 각각의 분리한 dataset으로 dataloader라는 것을 만듬 -> dataloader가 하는 역할은 batch 버전을 제공(for문을돌릴 때, x와 y를 적당한 size로 묶어서 내보내는 역할을 함, 매 iteration마다 랜덤으로 섞어서 내보냄, 그러한 것들을 편리하게 수행해줌)\n5. train, val loader를 묶어서 dataloaders 까지 만들어야 fastai에 넣고 돌리기가 가능\n\n이 때 입력되는 데이터는 벡터든 28 * 28의 matrix 형태의 이미지든 다 됨, 그래서 이걸 최적화해서 알아서 맞춰주는게 아니라 우리가 networks에 맞춰서 직접 설정해줘야됨",
"_____no_output_____"
],
[
"`-` 2D part에서 Dropout을 사용하지 않는 것에 대해서,\n- 1d part에서는 사용하는 DropOut을 2d part에서는 사용하지 않는다, dropout이 과적합을 막아주긴 하지만 그 전에, 2d part에서는 과적합이 잘 일어나지 않는다(보통은 그런다 하심), 구체적이진 않지만 CNN은 Fully Connected가 아니라는 점, 다음 레이어의 각각의 노드에 대해, 이전의 모든 노드들이 참여하지 않는다는 점 때문에 과적합이 잘 일어나지 않는다 하심",
"_____no_output_____"
],
[
"`-` Batch Normalize: 학습을 빠르게하는 효과가 있음",
"_____no_output_____"
],
[
"`*` 먼저 결과 y, 모델 값, 혹은 예측 값이 연속적인건 너무 힘듬, threshold라도 주는 것이 아닌 이상 0.1, 0.2, 0.3별로 나눠서 각각의 label을 분류하는 것은 힘든 것. 비트 정보 처리도 마찬가지임, 001, 010, 100 중에서 해야지 0.8 0.4 0.6 이면 어떻게 해석을 하겠음, 정도를 나누고 sigmoid (혹은 softmax) 특성상 가장 높은 것을 답으로 취하는 방식인 것",
"_____no_output_____"
],
[
"`*` `-` 결과가 2차원인 것, 이것은 그저 0, 1로 두가지 label을 구분하는 것 보다 01, 10 방식으로 구분하게 함으로써 나중가서 여러개의 label에 대해 더 확장이 용이하도록 하기 위함(?)인 것 같은데 확실히 이해한 것인지 명확하지가 않음",
"_____no_output_____"
],
[
"`*` 확실하진 않은데, 통계학에서 \n추론을 할 때 y가 연속이면 normal distribution으로, y가 0 or 1이면 이항분포로, y가 001, 010, 101 이런 식이면 다항분포로 잡고 추론을 한다고 하심\n\n-> 그런데 끝가지 보고 나니, activation function 적용이 연속은 linear, 0 or 1은 sigmoid, 001 010 100 같은 경우는 soft max를 취한다는 것이 맞는 방향으로 생각하고 있는듯 싶음\n\n추가 팁: Loss fnction 설정에 대해서: <br/>\n연속은 MSE, 0 or 1은 BCE, 001 010 100 같은 경우는 Cross Entropy 를 사용\n\nresnet이 중요한 이유는 손실함수의 모양을 바꿔주기 때문이라 하심. resnet의 효과가 shorcut을 만들어서 skip을 주는거라 하심\n\n층이 깊어질수록 수렴이 잘 안하는 특징이 있다 하심\n\n\n파라메트릭 모형: 해당분야 (물리학, 경제학 등등)의 전문가 들이 함, 특정 현상 보고 관계들 등등에 대해 모델링을 함\n넌파라메트릭 모형: 통계전문가, 파라마터가 없어도 어떻게든 유의미한 결론을 이끌어냄 (커널 메소드같은게 넌 파라메트릭 모형이라 하심)\n\n한층한층 의미있는게 아니다, 커널 사이즈든 매핑할 때 아웃풋 개수든, 모두 적절한 것을 찾아나가는 것이다. 상대적으로 비전문가들이 할 수 있음. 왜 Drouout넣고 MaxPooling하고, 한층한층 따지는 의미가 딱히 없는듯. ->\n이러한 것을 블랙박스라 한다. 내부에서 무슨일이 돌아가서 이러이러한 결론을 내린건지를 해석하지 어렵기 때문\n\n이 블랙박스가 왜 문제냐면, 대출을 할 떄 대출불가라고 하면 전문가가 직접 모형만들고 판단하면 대출불가 사유 설명이 되는데, 비전문가가 만든 딥러닝의 경우는 그냥 딥러닝이 그런 결과를 냈기 때문이라고 하는 정도일 수 있음. -> 이 떄문에 나온 것이 XAI(Explainable AI, 설명가능 인공지능)임",
"_____no_output_____"
],
[
"### 설명가능한 CNN 모형",
"_____no_output_____"
],
[
"`-` 현재까지의 모형\n- 1단계: 2d 선형변환 $\\to$ 2d 비선형변환\n- 2단계: Flatten $\\to$ MLP",
"_____no_output_____"
],
[
"`-` lrnr1(직접 만들었던 모형)의 모형을 다시 복습",
"_____no_output_____"
]
],
[
[
"lrnr1.model",
"_____no_output_____"
],
[
"net1 = torch.nn.Sequential(\n lrnr1.model[0],\n lrnr1.model[1],\n lrnr1.model[2],\n lrnr1.model[3]\n)",
"_____no_output_____"
],
[
"net1(X).shape",
"_____no_output_____"
]
],
[
[
"`-` 1단계까지의 추력결과를 시각화",
"_____no_output_____"
]
],
[
[
"fig, axs = plt.subplots(4,4)\nk = 0\nfor i in range(4):\n for j in range(4):\n axs[i,j].imshow(net1(X)[0][k].data)\n k = k+1\nfig.set_figheight(8)\nfig.set_figwidth(8)\nfig.tight_layout()",
"_____no_output_____"
]
],
[
[
"#### net1은 유지 + net2의 구조를 변경!!",
"_____no_output_____"
]
],
[
[
"lrnr1.model",
"_____no_output_____"
]
],
[
[
"`-` 계획\n- 변경전 net2: (n,16,12,12) $\\overset{flatten}{\\Longrightarrow} (n,?) \\overset{Linear(?,1)}{\\Longrightarrow} (n,1)$\n- 변경후 net2: (n,16,12,12) $\\overset{gap+flatten}{\\Longrightarrow} (n,?) \\overset{Linear(16,1)}{\\Longrightarrow} (n,1)$",
"_____no_output_____"
],
[
"- gap은 (n,16,12,12)의 한 데이터에 대해서 나오는 각각의 16개 이미지들의 픽셀들의 평균을 취한 값들을 의미함. 결과적으로는 16개의 이미지들 각각에서 12*12개의 픽셀값이 있고 이들을 평균 취하면 16개 이미지 각각에서 1개씩 값이 나오므로 총 16개의 값이 나옴",
"_____no_output_____"
],
[
"`-` gap: 12 $=times$12 픽셀을 평균내서 하나의 값으로 대표하자.",
"_____no_output_____"
]
],
[
[
"ap = torch.nn.AdaptiveAvgPool2d(output_size=1)",
"_____no_output_____"
]
],
[
[
"- `*`output_size는 평균내서 나올 값의 개수를 뜻하는건가?",
"_____no_output_____"
],
[
"`-` ap라는 layer를 생성했다.",
"_____no_output_____"
]
],
[
[
"ap(net1(X)).shape",
"_____no_output_____"
]
],
[
[
"- 16,1,1 부분을 보면 Flatten 작업이 필요함을 알 수 있다.\n- ***보충학습: ap는 그냥 평균***",
"_____no_output_____"
],
[
"`-` flatten",
"_____no_output_____"
]
],
[
[
"flatten(ap(net1(X))).shape",
"_____no_output_____"
]
],
[
[
"- flatten을 통해서 (12396,16,1,1) 형태가 (12396,16) 로 펼쳐졌다.",
"_____no_output_____"
],
[
"`-` linear",
"_____no_output_____"
]
],
[
[
"_l1 = torch.nn.Linear(16,1,bias=False)",
"_____no_output_____"
],
[
"# _li.to(\"cuda:0\") 만약 gpu에서 계속 연산을 한다면 이 코드를 실행",
"_____no_output_____"
]
],
[
[
"`-` 이렇게 gap을 추가한 것을 net2 로 구성, $\\to$ (net1, net2)를 묶어서 새로운 net 만들자.",
"_____no_output_____"
]
],
[
[
"net2 = torch.nn.Sequential(\n torch.nn.AdaptiveAvgPool2d(1),\n Flatten(),\n torch.nn.Linear(16,1,bias=False)\n)",
"_____no_output_____"
]
],
[
[
"`*` torch.nn.linear과 torch.nn.Linear이 무슨 차이가 있는지, ---> L부분이 대문자일 때와 소문자일 때의 차이\n",
"_____no_output_____"
]
],
[
[
"ds = torch.utils.data.TensorDataset(X,y)\nds1,ds2 = torch.utils.data.random_split(ds, [10000,2396])\ndl1 = torch.utils.data.DataLoader(ds1,batch_size=1000)\ndl2 = torch.utils.data.DataLoader(ds2,batch_size=2396)\ndls = DataLoaders(dl1,dl2)",
"_____no_output_____"
],
[
"lrnr3 = Learner(dls,net,opt_func=Adam, loss_func = loss_fn , lr = 0.1)",
"_____no_output_____"
],
[
"lrnr3.fit(50)",
"_____no_output_____"
]
],
[
[
"### CAM: observation을 1개로 고정하고 net2에서 layer의 순서를 바꿔서 시각화",
"_____no_output_____"
],
[
"`-` 계획\n- 변경전 netSequential(n,16,12,12) $\\overset{flatten}{\\Longrightarrow} (n,?) \\overset{Linear(?,1)}{\\Longrightarrow} (n,1)$\n- 변경후 net2: (n,16,12,12) $\\overset{gap+flatten}{\\Longrightarrow} (n,?) \\overset{Linear(16,1)}{\\Longrightarrow} (n,1)$\n- CAM: (1,16,12,12) $\\overset{Linear(16,1)+flatten}{\\Longrightarrow} (12,12) \\overset{gap}{\\Longrightarrow} (1,1)$\n",
"_____no_output_____"
],
[
"- CAM에서 Linear(16,1)로 차원이 (12,12) 되는 이유를 알아야됨",
"_____no_output_____"
],
[
"`-` 준비과정1: 시각화할 샘플을 하나 준비하자",
"_____no_output_____"
]
],
[
[
"x = X[1]\nX.shape, x.shape",
"_____no_output_____"
]
],
[
[
"- x의 차원이 하나 줄었다. 네트워크에 입력할 때 문제가 발생할 수 있으므로, 입력 차원을 맞추려면 X와 같이 하나 추가를 해야한다.",
"_____no_output_____"
]
],
[
[
"x = x.reshape(1,1,28,28)",
"_____no_output_____"
],
[
"#x.squeeze()",
"_____no_output_____"
]
],
[
[
"- plt.imshow에 대입하기 위해 이를 (28,28)로 맞춰야함, 이를 위한 함수가 squeeze()",
"_____no_output_____"
]
],
[
[
"plt.imshow(x.squeeze())",
"_____no_output_____"
]
],
[
[
"`-` 준비과정2: 계산과 시각화를 위해서 각 네트워크를 cpu로 옮기자. (fastai로 학습한 직후라 GPU에 있음)",
"_____no_output_____"
]
],
[
[
"net1.to(\"cpu\")\nnet2.to(\"cpu\")",
"_____no_output_____"
]
],
[
[
"`-` forward 확인: 이 값을 기억하자.",
"_____no_output_____"
]
],
[
[
"net2(net1(x)) # 0.5 미만으로, 7에 해당한다. CNN이 7이라고 판단.",
"_____no_output_____"
]
],
[
[
"- `-` net1, net2만 gpu에 올리고 net(x)를 실행한 결과: device가 다르다며 error가 나온다.",
"_____no_output_____"
],
[
"`-` net2를 수정하고 forward값 확인",
"_____no_output_____"
]
],
[
[
"net2",
"_____no_output_____"
]
],
[
[
"- net에서 Linear와 AdaptiveAvgPool2d의 적용순서를 바꿔줌",
"_____no_output_____"
],
[
"차원 확인",
"_____no_output_____"
]
],
[
[
"net1(x).squeeze().shape",
"_____no_output_____"
],
[
"net2[2].weight.shape # net2[2]는 두 cell 위의 net2결과에서 (2): Linear부분을 뜻한다.",
"_____no_output_____"
]
],
[
[
"- 이해가 잘 안되는 부분, 1,16이면 두 번 곱할 수 있는데, 그렇게하면 1,12,12 가 나와서 차원을 하나 줄인다 하심",
"_____no_output_____"
]
],
[
[
"net2[2].weight.squeeze().shape # net2[2]는 두 cell 위의 net2결과에서 (2): Linear부분을 뜻한다.",
"_____no_output_____"
]
],
[
[
"- 차원을 하나 줄인 결과",
"_____no_output_____"
],
[
"**Linear(in_features=16, out_features=1, bias=False)** 를 먼저 적용: 16 $\\times$ (16,12,12) -> (12,12)",
"_____no_output_____"
]
],
[
[
"# net2[2].weight.squeeze() @ net1(x).squeeze()",
"_____no_output_____"
]
],
[
[
"- 에러가 나는 이유: @는 matrix 연산으로 2차원 텐서에 대해서 적용하는데, 이는 3차원이라 @로 연산이 안됨\n- 실패",
"_____no_output_____"
],
[
"`-` 곱하는 방법: torch.einsum()",
"_____no_output_____"
]
],
[
[
"camimg = torch.einsum(\"i, ijk -> jk\", net2[2].weight.squeeze(), net1(x).squeeze())",
"_____no_output_____"
]
],
[
[
"`-` 계산이 잘 됬는지 shape로 확인",
"_____no_output_____"
]
],
[
[
"camimg.shape",
"_____no_output_____"
]
],
[
[
"- 성공",
"_____no_output_____"
],
[
"`-` Linear를 적용하고, 이제 gap을 적용",
"_____no_output_____"
]
],
[
[
"ap(camimg)",
"_____no_output_____"
]
],
[
[
"`!!!!` 위의 0.0552와 똑같다?",
"_____no_output_____"
],
[
"`-` 아래의 값이 같다.",
"_____no_output_____"
]
],
[
[
"net2(net1(x)), ap(camimg)",
"_____no_output_____"
]
],
[
[
"`-` 왜냐하면 ap와 선형변환 모두 linear이므로 순서를 바꿔도 상관없음",
"_____no_output_____"
],
[
"`-` 아래와 같은 이치",
"_____no_output_____"
]
],
[
[
"_x = np.array([1,2,3,4])\n_x",
"_____no_output_____"
],
[
"np.mean(_x*2+1)",
"_____no_output_____"
],
[
"2*np.mean(_x)+1",
"_____no_output_____"
]
],
[
[
"`-` 이제 camimg에 관심을 가져보자.",
"_____no_output_____"
]
],
[
[
"camimg # 12 * 12 tensor",
"_____no_output_____"
]
],
[
[
"현재 위의 camimg를 평균 취함으로써, 즉, ap(camimg) 혹은 torch.mean(camimg)를 함으로써 0.0552와 같다는 것",
"_____no_output_____"
],
[
"`*` 교수님 픽셀의 값과 내 값이 다름, 교수님 픽셀로 일단 작성",
"_____no_output_____"
],
[
"`-` 결국 특정픽셀에서 큰 음의 값이 나오기 때문에 궁극적으로는 평균이 음수가 된다.",
"_____no_output_____"
],
[
"- 평균이 음수이다. $\\leftrightarrow$ 이미지가 의미하는 것이 7이다.\n- 특정픽셀이 큰 음수값을 가진다. $\\leftrightarrow$ 이미지 값 7을 좌우하는 픽셀, 즉, 이 픽셀은 이미지가 7이라는 근거가 되는 픽셀이다.",
"_____no_output_____"
]
],
[
[
"plt.imshow(camimg.data)",
"_____no_output_____"
]
],
[
[
"`-` 원래 이미지와 비교",
"_____no_output_____"
]
],
[
[
"plt.imshow(x.squeeze())",
"_____no_output_____"
]
],
[
[
"`-` 두 이미지를 겹쳐서 그리면 멋진 그림이 될 것이다.\n",
"_____no_output_____"
],
[
"step1: 원래 이미지를 흑백으로 그리자",
"_____no_output_____"
]
],
[
[
"plt.imshow(x.squeeze(), cmap='gray',alpha=0.5)",
"_____no_output_____"
]
],
[
[
"step2: camimg은 (12,12) 픽셀이고 x는 (28,28) 픽셀이라서 이를 맞춰줘야한다. $\\to$ camimg를 늘린다.",
"_____no_output_____"
],
[
"- interpolation을 통해서 적당히 smoothing하여 작은 픽셀을 큰 픽셀로 확대한다.",
"_____no_output_____"
]
],
[
[
"plt.imshow(camimg.data, alpha = 0.5, extent = (0,28,28,0), interpolation = 'bilinear', cmap='magma')",
"_____no_output_____"
],
[
"plt.imshow(x.squeeze(), cmap='gray',alpha=0.5)\nplt.imshow(camimg.data, alpha = 0.5, extent = (0,27,27,0), interpolation = 'bilinear', cmap='magma')",
"_____no_output_____"
]
],
[
[
"- `*` 0,28,28,0 부분을 0,27,27,0으로 바꾸심. 이유는 잘 모르겠음",
"_____no_output_____"
],
[
"### 2d part의 (1,16,12,12) 와 Linear transform인 (16,1)의 연산이 이해되지 않음",
"_____no_output_____"
],
[
"#### tensor 계산 연습",
"_____no_output_____"
]
],
[
[
"_mat1 = torch.tensor([[[1,2,3,4], [5,6,7,8], [9,10,11,12]],[[-1,2,-3,4], [-5,6,-7,8], [-9,10,-11,12]]])",
"_____no_output_____"
],
[
"_mat2 = torch.tensor([10 for i in range(2)])",
"_____no_output_____"
],
[
"_mat2.shape, _mat1.shape",
"_____no_output_____"
]
],
[
[
"--",
"_____no_output_____"
]
],
[
[
"net1(x).squeeze().shape",
"_____no_output_____"
],
[
"net2[2].weight.squeeze().shape # net2[2]는 두 cell 위의 net2결과에서 (2): Linear부분을 뜻한다.",
"_____no_output_____"
]
],
[
[
"`-` camimg = torch.einsum(\"i, ijk -> jk\", net2[2].weight.squeeze(), net1(x).squeeze())\n를 참고하여",
"_____no_output_____"
],
[
"--",
"_____no_output_____"
]
],
[
[
"torch.einsum(\"i, ijk -> jk\", _mat2, _mat1) # 각각의 jk에 대해 i인덱스별 합,",
"_____no_output_____"
]
],
[
[
"각각의 jk에 대해 i인덱스별 합, $ \\sum_i{mat1_i}{mat2_ijk}$ -> ijk를 [i][j][k]로 인덱싱하면 각각의 j,k별로 _mat1[0][0][0]*_mat2[0] + _mat1[1][0][0]*_mat2[1] /// _mat1[0][0][1]*_mat2[0] + _mat1[1][0][1]*_mat2[1] /// $\\dots$ ///_mat1[0][2][3]*_mat2[0] + _mat1[1][2][3]*_mat2[1] 과 같이 계산된다.",
"_____no_output_____"
]
],
[
[
"for i in range(3):\n for j in range(4):\n print(_mat1[0][i][j]*_mat2[0] + _mat1[1][i][j]*_mat2[1])",
"tensor(0)\ntensor(40)\ntensor(0)\ntensor(80)\ntensor(0)\ntensor(120)\ntensor(0)\ntensor(160)\ntensor(0)\ntensor(200)\ntensor(0)\ntensor(240)\n"
],
[
"_mat1[0][0][0]*_mat2[0] + _mat1[1][0][0]*_mat2[1]",
"_____no_output_____"
]
],
[
[
"`-` 알아낸 einsum의 원리를 바탕으로 2d part output 과 1d part의 linear transform의 연산결과를 해석해보면, -> 2d part output은 torch.Size([16, 12, 12]) 이고 1d part는 16개의 가중치를 가진다.\n\ntorch.Size([16, 12, 12]) -> 16은 channel(혹은 16장의 이미지) 즉, [이미지 지정 인덱스][지정된 인덱스 이미지 pixel row][지정된 인덱스 이미지 pixel col] 에서 지정 위치의 픽셀 [row][col]에 대한 16장 이미지의 해당 픽셀들에 가중치를 곱해서 더하여 해당 픽셀에 저장함.",
"_____no_output_____"
]
],
[
[
"x = X[7000]\nx.shape",
"_____no_output_____"
],
[
"x = X[7000]\nx = x.squeeze()",
"_____no_output_____"
],
[
"plt.imshow(x)",
"_____no_output_____"
],
[
"x = x.reshape(1,1,28,28)\ncamimg = torch.einsum(\"i, ijk -> jk\", net2[2].weight.squeeze(), net1(x).squeeze())\nplt.imshow(x.squeeze(), cmap='gray',alpha=0.5)\nplt.imshow(camimg.data, alpha = 0.5, extent = (0,27,27,0), interpolation = 'bilinear', cmap='magma')",
"_____no_output_____"
]
]
] | [
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] | [
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown",
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown",
"markdown",
"markdown",
"markdown"
],
[
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown",
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown",
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown",
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code"
]
] |
d0fc42b9c0ca6d9ecb7d851f023f1f8c3d3961f0 | 55,512 | ipynb | Jupyter Notebook | training_modules/LabML03a.ipynb | jorgealexandreb/Covid-19-ML-Project- | cc73dc2324f43127d1959f13ca01389473b2c224 | [
"MIT"
] | 1 | 2021-03-19T18:23:03.000Z | 2021-03-19T18:23:03.000Z | training_modules/LabML03a.ipynb | jorgealexandreb/Covid-19-ML-Project- | cc73dc2324f43127d1959f13ca01389473b2c224 | [
"MIT"
] | null | null | null | training_modules/LabML03a.ipynb | jorgealexandreb/Covid-19-ML-Project- | cc73dc2324f43127d1959f13ca01389473b2c224 | [
"MIT"
] | null | null | null | 95.381443 | 13,972 | 0.803034 | [
[
[
"**LabML03a**\n\nPurpose: Identify clusters of Gira docking station\n\n1 import libraries needed:numpy, sklearn, matplotlib and pandas\n\n2 generate a sample of blobs and convert it into a dataframe called df1\n\n3 Verify datatype\n\n4 Plot the blobs\n\n5 calculete WCSS\n\n6 plot the new chart with centroids\n\n7 identify to what group does each item belongs\n\nComment the code",
"_____no_output_____"
]
],
[
[
"import numpy as np\nimport pandas as pd\nfrom matplotlib import pyplot as plt\nfrom sklearn.cluster import KMeans",
"_____no_output_____"
],
[
"file='https://github.com/masterfloss/data/blob/main/giras201030.csv?raw=true'\ndfGiras=pd.read_csv(file,sep=';')",
"_____no_output_____"
],
[
"dfGiras.head()",
"_____no_output_____"
],
[
"dfGiras.loc[0,'position'].split()[1].replace('[','').replace(',','')",
"_____no_output_____"
],
[
"for i in range(len(dfGiras['position'])):\n dfGiras.loc[i,'long']=dfGiras.loc[i,'position'].split()[1].replace('[','').replace(',','')\n dfGiras.loc[i,'lat']=dfGiras.loc[i,'position'].split()[2].replace('],','')\n \n ",
"_____no_output_____"
],
[
"dfGiras.head()",
"_____no_output_____"
],
[
"df1=dfGiras[['long','lat']]\n\ndf1.dtypes",
"_____no_output_____"
],
[
"df1.loc[:,'long']=pd.to_numeric(df1.loc[:,'long'])\ndf1.loc[:,'lat']=pd.to_numeric(df1.loc[:,'lat'])\n",
"/Users/alehxh/opt/anaconda3/lib/python3.8/site-packages/pandas/core/indexing.py:1048: SettingWithCopyWarning: \nA value is trying to be set on a copy of a slice from a DataFrame.\nTry using .loc[row_indexer,col_indexer] = value instead\n\nSee the caveats in the documentation: https://pandas.pydata.org/pandas-docs/stable/user_guide/indexing.html#returning-a-view-versus-a-copy\n self.obj[item_labels[indexer[info_axis]]] = value\n/Users/alehxh/opt/anaconda3/lib/python3.8/site-packages/pandas/core/indexing.py:966: SettingWithCopyWarning: \nA value is trying to be set on a copy of a slice from a DataFrame.\nTry using .loc[row_indexer,col_indexer] = value instead\n\nSee the caveats in the documentation: https://pandas.pydata.org/pandas-docs/stable/user_guide/indexing.html#returning-a-view-versus-a-copy\n self.obj[item] = s\n"
],
[
"df1.dtypes",
"_____no_output_____"
],
[
"plt.scatter(df1['long'], df1['lat'])\nplt.title('Giras')\nplt.xlabel('long')\nplt.ylabel('lat')",
"_____no_output_____"
],
[
"wcss = []\nfor i in range(1, 11):\n model = KMeans(n_clusters=i, init='k-means++', max_iter=300, n_init=10, random_state=0)\n model.fit(df1)\n wcss.append(model.inertia_)\nplt.plot(range(1, 11), wcss)\nplt.title('Elbow Method')\nplt.xlabel('Number of clusters')\nplt.ylabel('WCSS')\nplt.show()",
"_____no_output_____"
],
[
"model1 = KMeans(n_clusters=5, init='k-means++', max_iter=400, n_init=10, random_state=0)\nmodel1.fit_predict(df1)\nplt.scatter(df1[\"long\"], df1[\"lat\"])\nplt.scatter(model1.cluster_centers_[:, 0], model1.cluster_centers_[:, 1], s=300, c='red')\nplt.show()\n",
"_____no_output_____"
],
[
"model1.predict(df1.loc[0:0,:])",
"_____no_output_____"
],
[
"model1.predict(df1)",
"_____no_output_____"
]
]
] | [
"markdown",
"code"
] | [
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
d0fc44f9320af34347f02569e82e437f267e027c | 46,057 | ipynb | Jupyter Notebook | FitzRoyTesting.ipynb | Beefstyles/AFLTippingPredictor | 82d8779eecb20d1cc83c5e62e03b9ed477981a9e | [
"Apache-2.0"
] | null | null | null | FitzRoyTesting.ipynb | Beefstyles/AFLTippingPredictor | 82d8779eecb20d1cc83c5e62e03b9ed477981a9e | [
"Apache-2.0"
] | null | null | null | FitzRoyTesting.ipynb | Beefstyles/AFLTippingPredictor | 82d8779eecb20d1cc83c5e62e03b9ed477981a9e | [
"Apache-2.0"
] | null | null | null | 130.473088 | 1,484 | 0.69021 | [
[
[
"import pandas as pd",
"_____no_output_____"
],
[
"import rpy2",
"_____no_output_____"
],
[
"print(rpy2.__version__)",
"3.4.5\n"
],
[
"from rpy2.robjects.packages import importr\nfrom rpy2.robjects import pandas2ri\n#Must be activated\npandas2ri.activate()",
"_____no_output_____"
],
[
"# import rpy2's package module\nimport rpy2.robjects.packages as packages",
"_____no_output_____"
],
[
"# import R's utility package\nutils = rpackages.importr('utils')\n\n# select a mirror for R packages\nutils.chooseCRANmirror(ind=1) # select the first mirror in the list",
"_____no_output_____"
],
[
"utils = packages.importr('utils')\nutils.chooseCRANmirror(ind=1)\nutils.install_packages('fitzRoy')",
"R[write to console]: Warning:\nR[write to console]: package 'fitzRoy' is in use and will not be installed\n\n"
],
[
"fr = packages.importr('fitzRoy')",
"_____no_output_____"
],
[
"utils.install_packages('ggplot2')",
"R[write to console]: Installing package into 'C:/Users/Beefsports/Documents/R/win-library/4.1'\n(as 'lib' is unspecified)\n\nR[write to console]: trying URL 'https://cloud.r-project.org/bin/windows/contrib/4.1/ggplot2_3.3.5.zip'\n\nR[write to console]: Content type 'application/zip'\nR[write to console]: length 4130096 bytes (3.9 MB)\n\nR[write to console]: downloaded 3.9 MB\n\n\n"
],
[
"ggp = importr('ggplot2')",
"_____no_output_____"
],
[
"utils.install_packages('fitzRoy')",
"R[write to console]: Installing package into 'C:/Users/Beefsports/Documents/R/win-library/4.1'\n(as 'lib' is unspecified)\n\nR[write to console]: trying URL 'https://cloud.r-project.org/bin/windows/contrib/4.1/fitzRoy_1.1.0.zip'\n\nR[write to console]: Content type 'application/zip'\nR[write to console]: length 1565549 bytes (1.5 MB)\n\nR[write to console]: downloaded 1.5 MB\n\n\n"
],
[
"fitzRoy = importr('fitzRoy')",
"_____no_output_____"
],
[
"fitzRoy?",
"Object `fitzRoy` not found.\n"
]
]
] | [
"code"
] | [
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
d0fc639898cfb35d96cf0bc4d9d5ed0187682e5d | 101,512 | ipynb | Jupyter Notebook | dataset/Data Exploration.ipynb | cedricviaccoz/musicology | d1d04a4fe7b45fc6f08c52a7115017152f9baad3 | [
"BSD-3-Clause"
] | 1 | 2020-03-08T15:57:12.000Z | 2020-03-08T15:57:12.000Z | dataset/Data Exploration.ipynb | cedricviaccoz/musicology | d1d04a4fe7b45fc6f08c52a7115017152f9baad3 | [
"BSD-3-Clause"
] | null | null | null | dataset/Data Exploration.ipynb | cedricviaccoz/musicology | d1d04a4fe7b45fc6f08c52a7115017152f9baad3 | [
"BSD-3-Clause"
] | null | null | null | 94.079703 | 24,436 | 0.796477 | [
[
[
"# Game Music dataset: data cleaning and exploration\nThe goal with this notebook is cleaning the dataset to make it usable as well as providing a descriptive analysis of the dataset features.\n\n## Data loading and cleaning",
"_____no_output_____"
]
],
[
[
"import warnings\nwarnings.filterwarnings('ignore')",
"_____no_output_____"
],
[
"import pandas as pd\nimport numpy as np\nfrom ast import literal_eval\nimport os\nimport seaborn as sns\nimport matplotlib.pyplot as plt\n%matplotlib inline\n\ndf = pd.read_csv('midi_dataframe.csv', parse_dates=[11])\nnum_midis_before = len(df)\nprint('There is %d midi files, from %d games, with %d midis matched with tgdb'\n %(num_midis_before,\n len(df.groupby(['tgdb_platform', 'tgdb_gametitle'])),\n (df.tgdb_gametitle.notnull().sum())))\ndf.head()",
"There is 31685 midi files, from 3388 games, with 31249 midis matched with tgdb\n"
]
],
[
[
"We keep only files matched with tgdb and check that every midi file is only present once, if not we drop the rows.",
"_____no_output_____"
]
],
[
[
"num_dup = df.duplicated(subset='file_name').sum()\ndf.drop_duplicates(subset='file_name', inplace=True)\nprint('There was %d duplicated midi files, %d midis left'%(num_dup, len(df)))",
"There was 2 duplicated midi files, 31683 midis left\n"
]
],
[
[
"Since we are interested in the genre, we only keep midis that have one.",
"_____no_output_____"
]
],
[
[
"num_genres_na = df.tgdb_genres.isnull().sum()\ndf.dropna(subset=['tgdb_genres'], inplace=True)\nprint(\"We removed %d midis, %d midis left\"%(num_genres_na, len(df)))",
"We removed 436 midis, 31247 midis left\n"
]
],
[
[
"Then, there are some categories, such as Medleys or Piano only that are not interesting.\n\nThere is also a big \"remix\" scene on vgmusic, so we also remove those.",
"_____no_output_____"
]
],
[
[
"categories_filter = df.console.isin(['Medleys', 'Piano Only'])\nremix_filter = df.title.str.contains('[Rr]emix')\n\ndf = df[~categories_filter & ~remix_filter]\nprint('We removed %d midis from Medleys and Piano categories'%categories_filter.sum())\nprint('We removed %d midis containing \"remix\" in their title'%remix_filter.sum())\nprint('%d midis left'%len(df))",
"We removed 872 midis from Medleys and Piano categories\nWe removed 1860 midis containing \"remix\" in their title\n28537 midis left\n"
]
],
[
[
"There often exists several versions of the same midi file, most of the time denoted by 'title (1)', 'title (2)', etc.\n\nWe also consider removing those, but keeping only the one with the highest value, or if there are several with the same title, we randomly keep one.",
"_____no_output_____"
]
],
[
[
"num_midis_before = len(df)\n\ndf_stripped = df.copy()\ndf_stripped.title = df.title.str.replace('\\(\\d+\\)', '').str.rstrip()\ndf_stripped['rank'] = df.title.str.extract('\\((\\d+)\\)', expand=False)\ndf = df_stripped.sort_values(by='rank', ascending=False).groupby(['brand', 'console', 'game', 'title']).first().reset_index()\nprint(\"We removed %d midis, %d midis left\"%(num_midis_before-len(df), len(df)))",
"We removed 6358 midis, 22179 midis left\n"
]
],
[
[
"We also check if the midis files are valid by using mido and trying to load them.",
"_____no_output_____"
]
],
[
[
"from mido import MidiFile\n\nbad_midis = []\nfor file in df['file_name']:\n try:\n midi = MidiFile(\"full/\" + file)\n except:\n bad_midis.append(file)\ndf = df.loc[df.file_name.apply(lambda x: x not in bad_midis)]\nprint(\"We removed %d midis, %d midis left\"%(len(bad_midis), len(df)))",
"We removed 117 midis, 22062 midis left\n"
]
],
[
[
"The final numbers after preliminary data cleaning are:",
"_____no_output_____"
]
],
[
[
"num_games = len(df.groupby(['tgdb_platform', 'tgdb_gametitle']))\nprint('There is %d midi files, from %d games, with %d midis matched with tgdb'\n %(len(df),\n num_games,\n (df.tgdb_gametitle.notnull().sum())))",
"There is 22062 midi files, from 3237 games, with 22062 midis matched with tgdb\n"
]
],
[
[
"## Data Exploration",
"_____no_output_____"
],
[
"## General statistics\nWe first begin by some general statistics about the dataset.",
"_____no_output_____"
],
[
"The number of gaming platforms is computed.",
"_____no_output_____"
]
],
[
[
"print('There is %d platforms'%df.tgdb_platform.nunique())",
"There is 62 platforms\n"
]
],
[
[
"Then, statistics concerning the number of games per platform are computed and plotted.",
"_____no_output_____"
]
],
[
[
"df.drop_duplicates(subset=['tgdb_platform', 'tgdb_gametitle']).groupby('tgdb_platform').size().to_frame().describe()",
"_____no_output_____"
],
[
"size= (10, 5)\nfig, ax = plt.subplots(figsize=size)\nax = sns.distplot(df.drop_duplicates(subset=['tgdb_platform', 'tgdb_gametitle']).groupby('tgdb_platform').size().to_frame(), ax = ax)\nax.set_xlabel(\"number of games per platform\")\nax.set_ylabel(\"density\")\nax.set_title(\"Density of the number of games per platform\")\nax.set_xticks(np.arange(0, 300, 10))\nax.set_xlim(0,200)\nplt.show()",
"_____no_output_____"
]
],
[
[
"It can be noted that the majority of platforms seem to have around 10 games, which is a sufficient sample size.",
"_____no_output_____"
],
[
"Following this, statistics concerning the number of midis per platform are computed and plotted.",
"_____no_output_____"
]
],
[
[
"df.groupby('tgdb_platform').size().to_frame().describe()",
"_____no_output_____"
],
[
"fig, ax = plt.subplots(figsize=size)\nax = sns.distplot(df.groupby('tgdb_platform').size().to_frame())\nax.set_xlabel(\"number of midi per platform\")\nax.set_ylabel(\"density\")\nax.set_title(\"Density of the number of midis per platform\")\nax.set_xticks(np.arange(0, 1000, 50))\nax.set_xlim(0,1500)\nplt.show()",
"_____no_output_____"
]
],
[
[
"It can be noted that the majority of platform have around 50 midis, which is again judged to be a sufficient sample for analysis.",
"_____no_output_____"
],
[
"Finally, statistics concerning the number of midi per game are computed and plotted.",
"_____no_output_____"
]
],
[
[
"df.groupby(['tgdb_platform', 'tgdb_gametitle']).size().to_frame().describe()",
"_____no_output_____"
],
[
"fig, ax = plt.subplots(figsize=size)\nax = sns.distplot(df.groupby(['tgdb_platform', 'tgdb_gametitle']).size().to_frame())\nax.set_xlabel(\"number of midi per game\")\nax.set_ylabel(\"density\")\nax.set_title(\"Density of the number of midi per game\")\nax.set_xticks(np.arange(0, 40, 2))\nax.set_xlim(0,40)\nplt.show()",
"_____no_output_____"
]
],
[
[
"It can be noted that the peak of density is at 2 midi per game. This does not matter much as we are not trying to classify music per game, but by genres.",
"_____no_output_____"
],
[
"As a general remark, it can be noticed that most of the data we have follow power laws.",
"_____no_output_____"
],
[
"### Genres analysis\nWe currently had list of genres, for more convenience, we rework the dataframe to make several row of a midi if it had several genres.",
"_____no_output_____"
]
],
[
[
"genres = df.tgdb_genres.map(literal_eval, 'ignore').apply(pd.Series).stack().reset_index(level=1, drop=True)\ngenres.name = 'tgdb_genres'\ngenres_df = df.drop('tgdb_genres', axis=1).join(genres)\n\nprint(\"There is %d different genres\"%genres_df.tgdb_genres.nunique())",
"There is 19 different genres\n"
],
[
"genres_df.to_csv(\"midi_dataframe_cleaned.csv\")",
"_____no_output_____"
]
],
[
[
"Here follows the percentage of games belonging to each genre and of midis for each genres.",
"_____no_output_____"
]
],
[
[
"genres_df.drop_duplicates(subset=['tgdb_platform', 'tgdb_gametitle'])\\\n .groupby(['tgdb_genres']).size().to_frame()\\\n .sort_values(0, ascending = False)/num_games*100",
"_____no_output_____"
]
],
[
[
"The number of genres is 19, and could be reduced to 10 if we consider only the genres for which we have at least 3% dataset coverage or 5 if we consider only the genres for which we have at least 9% dataset coverage.",
"_____no_output_____"
]
]
] | [
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown"
] | [
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
]
] |
d0fc7ecbef9f434030156e269f64ab1d59cb10ca | 21,795 | ipynb | Jupyter Notebook | notebooks/keras/3_conv-net-in-keras.ipynb | ekholabs/workshop | a7aab6700cbabba9c50d20cf268960fb21b934f4 | [
"MIT"
] | 2 | 2018-05-22T17:13:37.000Z | 2018-12-14T12:02:31.000Z | notebooks/keras/3_conv-net-in-keras.ipynb | ekholabs/workshop | a7aab6700cbabba9c50d20cf268960fb21b934f4 | [
"MIT"
] | null | null | null | notebooks/keras/3_conv-net-in-keras.ipynb | ekholabs/workshop | a7aab6700cbabba9c50d20cf268960fb21b934f4 | [
"MIT"
] | 1 | 2019-11-19T08:25:17.000Z | 2019-11-19T08:25:17.000Z | 44.479592 | 5,472 | 0.643129 | [
[
[
"# Convolutional Neural Network in Keras",
"_____no_output_____"
],
[
"Bulding a Convolutional Neural Network to classify Fashion-MNIST.",
"_____no_output_____"
],
[
"#### Set seed for reproducibility",
"_____no_output_____"
]
],
[
[
"import numpy as np\nnp.random.seed(42)",
"_____no_output_____"
]
],
[
[
"#### Load dependencies",
"_____no_output_____"
]
],
[
[
"import os\n\nfrom tensorflow.keras.datasets import fashion_mnist\nfrom tensorflow.keras.models import Sequential\nfrom tensorflow.keras.layers import Layer, Activation, Dense, Dropout, Conv2D, MaxPooling2D, Flatten, LeakyReLU, BatchNormalization\nfrom tensorflow.keras.callbacks import TensorBoard, ModelCheckpoint, EarlyStopping\n\nfrom tensorflow.keras.utils import to_categorical\nfrom tensorflow.keras.models import load_model\n\nfrom keras_contrib.layers.advanced_activations.sinerelu import SineReLU\n\nfrom matplotlib import pyplot as plt\n\n%matplotlib inline",
"_____no_output_____"
]
],
[
[
"#### Load data",
"_____no_output_____"
]
],
[
[
"(X_train, y_train), (X_test, y_test) = fashion_mnist.load_data()",
"_____no_output_____"
]
],
[
[
"#### Preprocess data\nFlatten and normalise input data.",
"_____no_output_____"
]
],
[
[
"X_train = X_train.reshape(-1, 28, 28, 1)\nX_test = X_test.reshape(-1, 28, 28, 1)\n\nX_train = X_train.astype(\"float32\")/255.\nX_test = X_test.astype(\"float32\")/255.",
"_____no_output_____"
],
[
"# One-hot encoded categories\nn_classes = 10\ny_train = to_categorical(y_train, n_classes)\ny_test = to_categorical(y_test, n_classes)",
"_____no_output_____"
]
],
[
[
"#### Design Neural Network architecture",
"_____no_output_____"
]
],
[
[
"model = Sequential()\n\nmodel.add(Conv2D(32, 7, padding = 'same', input_shape = (28, 28, 1)))\n# model.add(LeakyReLU(alpha=0.01))\nmodel.add(Activation('relu'))\n\nmodel.add(Conv2D(32, 7, padding = 'same'))\n# model.add(LeakyReLU(alpha=0.01))\nmodel.add(Activation('relu'))\nmodel.add(MaxPooling2D(pool_size = (2, 2)))\nmodel.add(Dropout(0.20))\n\nmodel.add(Conv2D(64, 3, padding = 'same'))\n# model.add(LeakyReLU(alpha=0.01))\nmodel.add(Activation('relu'))\n\nmodel.add(Conv2D(64, 3, padding = 'same'))\n# model.add(LeakyReLU(alpha=0.01))\nmodel.add(Activation('relu'))\nmodel.add(MaxPooling2D(pool_size = (2, 2)))\nmodel.add(Dropout(0.30))\n\nmodel.add(Conv2D(128, 2, padding = 'same'))\n# model.add(LeakyReLU(alpha=0.01))\nmodel.add(Activation('relu'))\n\nmodel.add(Conv2D(128, 2, padding = 'same'))\n# model.add(LeakyReLU(alpha=0.01))\nmodel.add(Activation('relu'))\nmodel.add(MaxPooling2D(pool_size = (2, 2)))\nmodel.add(Dropout(0.40))\n\nmodel.add(Flatten())\nmodel.add(Dense(512))\n# model.add(LeakyReLU(alpha=0.01))\nmodel.add(Activation('relu'))\nmodel.add(Dropout(0.50))\n\nmodel.add(Dense(10, activation = \"softmax\"))\n\nmodel.summary()",
"Model: \"sequential\"\n_________________________________________________________________\nLayer (type) Output Shape Param # \n=================================================================\nconv2d (Conv2D) (None, 28, 28, 32) 1600 \n_________________________________________________________________\nactivation (Activation) (None, 28, 28, 32) 0 \n_________________________________________________________________\nconv2d_1 (Conv2D) (None, 28, 28, 32) 50208 \n_________________________________________________________________\nactivation_1 (Activation) (None, 28, 28, 32) 0 \n_________________________________________________________________\nmax_pooling2d (MaxPooling2D) (None, 14, 14, 32) 0 \n_________________________________________________________________\ndropout (Dropout) (None, 14, 14, 32) 0 \n_________________________________________________________________\nconv2d_2 (Conv2D) (None, 14, 14, 64) 18496 \n_________________________________________________________________\nactivation_2 (Activation) (None, 14, 14, 64) 0 \n_________________________________________________________________\nconv2d_3 (Conv2D) (None, 14, 14, 64) 36928 \n_________________________________________________________________\nactivation_3 (Activation) (None, 14, 14, 64) 0 \n_________________________________________________________________\nmax_pooling2d_1 (MaxPooling2 (None, 7, 7, 64) 0 \n_________________________________________________________________\ndropout_1 (Dropout) (None, 7, 7, 64) 0 \n_________________________________________________________________\nconv2d_4 (Conv2D) (None, 7, 7, 128) 32896 \n_________________________________________________________________\nactivation_4 (Activation) (None, 7, 7, 128) 0 \n_________________________________________________________________\nconv2d_5 (Conv2D) (None, 7, 7, 128) 65664 \n_________________________________________________________________\nactivation_5 (Activation) (None, 7, 7, 128) 0 \n_________________________________________________________________\nmax_pooling2d_2 (MaxPooling2 (None, 3, 3, 128) 0 \n_________________________________________________________________\ndropout_2 (Dropout) (None, 3, 3, 128) 0 \n_________________________________________________________________\nflatten (Flatten) (None, 1152) 0 \n_________________________________________________________________\ndense (Dense) (None, 512) 590336 \n_________________________________________________________________\nactivation_6 (Activation) (None, 512) 0 \n_________________________________________________________________\ndropout_3 (Dropout) (None, 512) 0 \n_________________________________________________________________\ndense_1 (Dense) (None, 10) 5130 \n=================================================================\nTotal params: 801,258\nTrainable params: 801,258\nNon-trainable params: 0\n_________________________________________________________________\n"
]
],
[
[
"#### Callbacks",
"_____no_output_____"
]
],
[
[
"modelCheckpoint = ModelCheckpoint(monitor='val_accuracy', filepath='model_output/weights-cnn-fashion-mnist.hdf5',\n save_best_only=True, mode='max')\nearlyStopping = EarlyStopping(monitor='val_accuracy', mode='max', patience=5)\n\n\nif not os.path.exists('model_output'):\n os.makedirs('model_output')\n\ntensorboard = TensorBoard(\"logs/convnet-fashion-mnist\")",
"_____no_output_____"
]
],
[
[
"#### Configure model",
"_____no_output_____"
]
],
[
[
"model.compile(loss = 'categorical_crossentropy', optimizer = 'adam', metrics = ['accuracy'])",
"_____no_output_____"
]
],
[
[
"#### Train!",
"_____no_output_____"
]
],
[
[
"history = model.fit(X_train, y_train, batch_size = 128, epochs = 20, verbose = 1,\n validation_split = 0.1, callbacks=[modelCheckpoint, earlyStopping, tensorboard])",
"Train on 54000 samples, validate on 6000 samples\nEpoch 1/20\n54000/54000 [==============================] - 109s 2ms/sample - loss: 0.6916 - accuracy: 0.7466 - val_loss: 0.3887 - val_accuracy: 0.8520\nEpoch 2/20\n54000/54000 [==============================] - 117s 2ms/sample - loss: 0.3989 - accuracy: 0.8529 - val_loss: 0.3361 - val_accuracy: 0.8700\nEpoch 3/20\n54000/54000 [==============================] - 121s 2ms/sample - loss: 0.3469 - accuracy: 0.8699 - val_loss: 0.3039 - val_accuracy: 0.8810\nEpoch 4/20\n54000/54000 [==============================] - 123s 2ms/sample - loss: 0.3211 - accuracy: 0.8813 - val_loss: 0.2813 - val_accuracy: 0.8910\nEpoch 5/20\n54000/54000 [==============================] - 124s 2ms/sample - loss: 0.2967 - accuracy: 0.8898 - val_loss: 0.2733 - val_accuracy: 0.8988\nEpoch 6/20\n54000/54000 [==============================] - 127s 2ms/sample - loss: 0.2849 - accuracy: 0.8936 - val_loss: 0.2595 - val_accuracy: 0.8993\nEpoch 7/20\n54000/54000 [==============================] - 125s 2ms/sample - loss: 0.2715 - accuracy: 0.8986 - val_loss: 0.2391 - val_accuracy: 0.9085\nEpoch 8/20\n54000/54000 [==============================] - 125s 2ms/sample - loss: 0.2617 - accuracy: 0.9021 - val_loss: 0.2470 - val_accuracy: 0.9067\nEpoch 9/20\n54000/54000 [==============================] - 123s 2ms/sample - loss: 0.2494 - accuracy: 0.9064 - val_loss: 0.2368 - val_accuracy: 0.9098\nEpoch 10/20\n54000/54000 [==============================] - 128s 2ms/sample - loss: 0.2438 - accuracy: 0.9090 - val_loss: 0.2296 - val_accuracy: 0.9115\nEpoch 11/20\n54000/54000 [==============================] - 128s 2ms/sample - loss: 0.2349 - accuracy: 0.9125 - val_loss: 0.2310 - val_accuracy: 0.9128\nEpoch 12/20\n54000/54000 [==============================] - 117s 2ms/sample - loss: 0.2287 - accuracy: 0.9146 - val_loss: 0.2337 - val_accuracy: 0.9183\nEpoch 13/20\n54000/54000 [==============================] - 126s 2ms/sample - loss: 0.2215 - accuracy: 0.9171 - val_loss: 0.2179 - val_accuracy: 0.9190\nEpoch 14/20\n54000/54000 [==============================] - 126s 2ms/sample - loss: 0.2162 - accuracy: 0.9196 - val_loss: 0.2139 - val_accuracy: 0.9168\nEpoch 15/20\n54000/54000 [==============================] - 124s 2ms/sample - loss: 0.2108 - accuracy: 0.9209 - val_loss: 0.2072 - val_accuracy: 0.9237\nEpoch 16/20\n54000/54000 [==============================] - 123s 2ms/sample - loss: 0.2086 - accuracy: 0.9220 - val_loss: 0.2196 - val_accuracy: 0.9195\nEpoch 17/20\n54000/54000 [==============================] - 122s 2ms/sample - loss: 0.2019 - accuracy: 0.9248 - val_loss: 0.2298 - val_accuracy: 0.9165\nEpoch 18/20\n54000/54000 [==============================] - 125s 2ms/sample - loss: 0.2015 - accuracy: 0.9236 - val_loss: 0.2116 - val_accuracy: 0.9217\nEpoch 19/20\n54000/54000 [==============================] - 126s 2ms/sample - loss: 0.1931 - accuracy: 0.9280 - val_loss: 0.2044 - val_accuracy: 0.9267\nEpoch 20/20\n54000/54000 [==============================] - 123s 2ms/sample - loss: 0.1923 - accuracy: 0.9277 - val_loss: 0.2034 - val_accuracy: 0.9238\n"
]
],
[
[
"#### Test Predictions",
"_____no_output_____"
]
],
[
[
"saved_model = load_model('model_output/weights-cnn-fashion-mnist.hdf5')\npredictions = saved_model.predict_classes(X_test, verbose = 2)\nprint(predictions)\n# np.std(history.history['loss'])",
"10000/1 - 9s\n[9 2 1 ... 8 1 5]\n"
]
],
[
[
"#### Test Final Accuracy",
"_____no_output_____"
]
],
[
[
"final_loss, final_acc = saved_model.evaluate(X_test, y_test, verbose = 2)\nprint(\"Final loss: {0:.4f}, final accuracy: {1:.4f}\".format(final_loss, final_acc))",
"10000/1 - 8s - loss: 0.2481 - accuracy: 0.9221\nFinal loss: 0.2229, final accuracy: 0.9221\n"
],
[
"image = X_test[0].reshape(1, 28, 28, 1)",
"_____no_output_____"
],
[
"predictions = model.predict_classes(image, verbose = 2)\nprint(predictions)",
"1/1 - 0s\n[9]\n"
],
[
"plt.imshow(X_test[0].reshape((28, 28)), cmap='gray')",
"_____no_output_____"
],
[
"# 0 T-shirt/top\n# 1 Trouser\n# 2 Pullover\n# 3 Dress\n# 4 Coat\n# 5 Sandal\n# 6 Shirt\n# 7 Sneaker\n# 8 Bag\n# 9 Ankle boot",
"_____no_output_____"
]
]
] | [
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] | [
[
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code"
]
] |
d0fc910c1c6487c61bcc447b7704cc33ab3be5b7 | 34,618 | ipynb | Jupyter Notebook | data_extraction/Training Routes.ipynb | paloukari/road-scanner | a343cb7ecf042d77768826cefff79f629589aab4 | [
"MIT"
] | null | null | null | data_extraction/Training Routes.ipynb | paloukari/road-scanner | a343cb7ecf042d77768826cefff79f629589aab4 | [
"MIT"
] | null | null | null | data_extraction/Training Routes.ipynb | paloukari/road-scanner | a343cb7ecf042d77768826cefff79f629589aab4 | [
"MIT"
] | 4 | 2019-10-27T16:18:52.000Z | 2020-10-12T18:49:58.000Z | 100.927114 | 2,041 | 0.656884 | [
[
[
"import json\nimport requests\nimport numpy as np\nimport os\nimport shutil",
"_____no_output_____"
],
[
"#Get google api keys\nwith open(\"../config.json\", \"r\") as f:\n# k = json.load(f)['key_bsa']\n# bk = json.load(f)['big_key_demo']\n kk = json.load(f)['key_kaitlin']\nBASE_URL_DIRECTIONS = 'https://maps.googleapis.com/maps/api/directions/json?'\nKEY = '&key=' + kk",
"ERROR:root:Internal Python error in the inspect module.\nBelow is the traceback from this internal error.\n\n"
],
[
"#Supporting functions\ndef get_total_distance(routes_obj):\n return routes_obj.json()['routes'][0]['legs'][0]['distance']['value']\n\ndef get_lat_lng(routes_obj):\n lat_lng = []\n steps = routes_obj.json()['routes'][0]['legs'][0]['steps']\n for step in steps:\n end = step['end_location']\n start = step['start_location']\n lat_lng += [(start['lat'],start['lng']),(end['lat'],end['lng'])]\n return lat_lng\n\ndef create_path(points):\n path = 'path='\n for i in points:\n path += str(i[0]) + ',' + str(i[1]) + '|'\n return path[:-1] #remove last '|'\n\ndef get_coords(r,points):\n for i in r.json()['snappedPoints']:\n points += [(i['location']['latitude'],i['location']['longitude'])]\n return points\n\ndef get_snapped_points(unique_points_interpolated,key,BASE_URL_SNAP = 'https://roads.googleapis.com/v1/snapToRoads?',interpolate = '&interpolate=true'):\n points = []\n k = 0\n coords_list = []\n while k <= len(unique_points_interpolated)-1:\n coords_list += [unique_points_interpolated[k]]\n if (len(coords_list)%100==0) or (k+1==len(unique_points_interpolated)): #When we have 100 points or we reach the end of the list.\n path = create_path(coords_list)\n url = BASE_URL_SNAP + path + interpolate + key\n r = requests.get(url)\n points += get_coords(r,points)\n coords_list = []\n k += 1\n return(points)\n\ndef interpolate_coordinates(distance, lat_lng, k, separation_mts = 300):\n unique_points = list(set(lat_lng))\n n = max([1,round((distance/separation_mts)/len(unique_points))])\n unique_points_interpolated = []\n for i in range(len(unique_points)-1):\n unique_points_interpolated += list(map(tuple,np.linspace(unique_points[i],unique_points[i+1],n)))\n unique_points_interpolated = sorted(list(set(unique_points_interpolated)), key = lambda x: x[0])\n if n > 1: #If we have any new points to snap. \n results = get_snapped_points(unique_points_interpolated,k)\n return results\n else:\n return unique_points_interpolated\n\ndef create_image(x, folder_name):\n for heading in range(0,4):\n lat=x[0]\n lng=x[1]\n heading=str(90*heading)\n query='https://maps.googleapis.com/maps/api/streetview?size=400x400&location=%s,%s&fov=90&heading=%s&pitch=10%s' % (str(lat),str(lng),heading,KEY)\n page=requests.get(query)\n# filename='%s-%s-%s-%s-%s.jpg' %(origin,destination,str(lat),str(lng),heading)\n filename='%s-%s-%s-%s.jpg' %(folder_name,str(lat),str(lng),heading)\n\n if not os.path.exists(filename+\".txt\") or os.path.getsize(filename)<5*10^3:\n f = open(filename,'wb')\n f.write(page.content) \n f.close()",
"_____no_output_____"
],
[
"#Example of waypoints to force the route to pass through a certain road\n#waypoints = 'waypoints=1202+Foothill+Blvd+Calistoga|Oakville,CA|Yountville,CA|Napa+Valley+Marriott+Hotel' \ndef download_pictures(origin, destination, category, folder_name, waypoints=None):\n os.chdir('/data/road-scanner/training/' + category) \n\n #Get interpolated coordinates\n if waypoints == None:\n url = BASE_URL_DIRECTIONS + 'origin=' + origin + '&' + 'destination=' + destination + '&' + KEY\n else:\n url = BASE_URL_DIRECTIONS + 'origin=' + origin + '&' + 'destination=' + destination + '&' + waypoints + KEY\n r = requests.get(url)\n upi = interpolate_coordinates(get_total_distance(r),get_lat_lng(r),KEY)\n #Download pictures\n if os.path.exists(folder_name):\n shutil.rmtree(folder_name)\n os.makedirs(folder_name) \n os.chdir(folder_name)\n org_dest_string='%s-%s' %(origin,destination)\n \n for i in range(len(upi)):\n create_image(upi[i], folder_name)",
"_____no_output_____"
],
[
"### Features of routes missing\norigin = \"origin=48470+Lakeview Blvd+Fremont+CA+94538\"\ndestination = \"destination=Oakland+California+94606\"\nurl = BASE_URL_DIRECTIONS + 'origin=' + origin + '&' + 'destination=' + destination + '&' + KEY\n\nurl\n# download_pictures(origin, destination, 'non_scenic', 'test_fremont_oakland_880')\n",
"_____no_output_____"
],
[
"# non scenic\n\norigins = [\"320+Acorn+Ct+Vacaville+CA+95688\", \n \"12141+Martha+Ann+Dr+Los+Alamitos+CA+90720\", \"Salida+California\", \n \"1625+W+Lugonia+Ave+Redlands+CA+92374\",\"I-205+Tracy+CA+95377\"]\ndestinations = [\"4505+W+Capitol+Ave+West+Sacramento+CA+95691\", \n \"Wilshire+Federal+Building+11000+Wilshire+Blvd+Los+Angeles+CA+90024\", \n \"Golden+State+Hwy,+Bakersfield,+CA+93307\",\n \"450+N+Atlantic+Blvd,+Monterey+Park,+CA+91754\", \"Grapevine+California\"]\nfolder_names = [\"vacaville_to_sac\", \n \"405_westwood_to_long_beach\", \"99_salida_to_bakersfield\", \"10_alhambra_to_riverside\",\n 'I5_tracy_to_grapevine']\n\nfor i in range(len(origins)):\n download_pictures(\"origin=\"+origins[i],\"destination=\"+destinations[i],'non_scenic',folder_names[i])",
"_____no_output_____"
],
[
"origins = ['38.3254,-122.27693000000001', '35.75368,-120.67729000000001', '36.758030000000005,-119.74912', \n '35.2503,-120.62506', '37.333000000000006,-119.65076', '34.01697,-118.82331', '36.76053,-119.11351', \n '38.914190000000005,-120.00522000000001', '36.62303,-121.84475', '34.31194,-117.47276000000001', \n '38.25555000000001,-120.35094000000001', '35.589470000000006,-120.6966', '36.12256,-121.02258', \n '36.282210000000006,-118.00583', '37.81255,-119.05365', '35.24604,-120.68278000000001', \n '33.27877,-115.96492', '41.996680000000005,-123.72141', '34.471990000000005,-119.28866000000001', \n '39.86818,-123.71397', '34.9236,-120.4171', '40.58534,-122.36037', '33.91863,-116.6016', \n '39.602700000000006,-121.61804000000001']\n\ndestinations = ['38.5889,-122.27835', '35.722,-120.67760000000001', '37.25511,-119.74913000000001', \n '35.22066,-120.62314', '38.0165,-119.64668', '34.64083,-118.825', '36.79612,-119.11176', \n '38.91415000000001,-120.00933', '35.28761,-121.84596', '34.07305,-117.46934', \n '38.660410000000006,-120.34897000000001', '35.64969,-120.69786', \n '36.860850000000006,-121.01838000000001', '36.3046,-118.00261', \n '37.892300000000006,-119.05625', '35.36764,-120.68357', '32.74718,-115.97338', \n '39.86816,-123.71989', '34.686040000000006,-119.28965000000001', \n '37.810570000000006,-123.71603', '34.61202,-120.40971', \n '40.584860000000006,-122.36037', '33.661460000000005,-116.59045', '39.14844,-121.5883']\n\nfolder_names = ['Silverado_Trail', 'Pleasant_Valley_Wine_Trail', 'Sierra_Heritage_Scenic_Byway', \n 'San_Luis_Obispo_Wine_Trail', 'Yosemite_Valley_and_Tioga_Road', 'Malibu_to_Lompoc', \n 'Kings_Canyon_and_Sequoia_National_Park', 'Lake_Tahoe', 'Big_Sur_Coast', \n 'Rim_of_the_World_Scenic_Byway', 'Ebbetts_Pass_Scenic_Byway', 'Paso_Robles_Wine_Country', \n 'Pinnacles_National_Park', 'Death_Valley_Scenic_Byway', 'June_Lake_Loop', \n 'Morro_Bay_Scenic_Drive', 'Anza_Borrego_Desert', 'Redwood_Highway', 'Jacinto_Reyes_Scenic_Byway', \n 'Northern_Pacific_Coast', 'Santa_Barbara_Wine_Country', 'Mount_Shasta-Cascade_Loop', \n 'Joshua_Tree_Journey', 'Feather_River_Scenic_Byway']\nfor i in range(len(origins)):\n download_pictures(\"origin=\"+origins[i],\"destination=\"+destinations[i],'scenic',folder_names[i])",
"_____no_output_____"
],
[
"origins = [\n 'Dwight+D+Eisenhower+Hwy+Oakland+CA+94607+USA','Golden+Gate+Bridge+View+Vista+Point+Sausalito+CA+94965+United+States',\n '51+59+Christmas+Tree+Point+Rd+San+Francisco+CA 94131+USA','Fresno+California+USA']\ndestinations = ['701+Bayshore+Blvd+San+Francisco+CA+94124+USA',\n 'The+Palace+Of+Fine+Arts+3601+Lyon+St+San+Francisco+CA+94123+United+States',\n '148+Marview+Way+San+Francisco+CA+94131+USA','Golden+State+Hwy+Bakersfield+CA+93307+USA']\nfolder_names = ['SF_Skyline','Golden_Gate','Twin_Peaks','shorter_99']\nfor i in range(len(origins)):\n download_pictures(origins[i],destinations[i],'demo', folder_names[i])",
"_____no_output_____"
]
]
] | [
"code"
] | [
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
d0fc98f14649433192fa1b183190696b1809398f | 695,005 | ipynb | Jupyter Notebook | program/7_1_Breakout_try.ipynb | keito1029/Deep-Reinforcement-Learning-Book | 6768a6e2437ea691011ecf997eccfc6522b10f0e | [
"MIT"
] | 266 | 2018-06-30T16:12:47.000Z | 2022-03-31T01:12:22.000Z | program/7_1_Breakout_try.ipynb | keito1029/Deep-Reinforcement-Learning-Book | 6768a6e2437ea691011ecf997eccfc6522b10f0e | [
"MIT"
] | 15 | 2018-08-30T06:49:56.000Z | 2021-12-28T23:32:33.000Z | program/7_1_Breakout_try.ipynb | keito1029/Deep-Reinforcement-Learning-Book | 6768a6e2437ea691011ecf997eccfc6522b10f0e | [
"MIT"
] | 148 | 2018-07-18T00:25:10.000Z | 2022-03-25T03:19:12.000Z | 810.029138 | 5,184 | 0.951616 | [
[
[
"## 7.1 Breakoutを実行してみる\n\nWindowsでbreakout-v0を実行するために、事前に以下のコマンドを実行してください。\n\npip install --no-index -f https://github.com/Kojoley/atari-py/releases atari_py\n\npip install opencv-python\n\n",
"_____no_output_____"
]
],
[
[
"# パッケージのimport\nimport numpy as np\nimport matplotlib.pyplot as plt\n%matplotlib inline\nimport gym\n",
"_____no_output_____"
],
[
"# ゲームの開始\nENV = 'Breakout-v0' # 使用する課題名\nenv = gym.make(ENV) # 実行する課題を設定\n",
"_____no_output_____"
],
[
"# ゲームの状態と行動を把握\n\n# 状態\nprint(env.observation_space)\n# Box(210, 160, 3)\n\n# 行動\nprint(env.action_space)\nprint(env.unwrapped.get_action_meanings())\n# Discrete(4)\n# ['NOOP', 'FIRE', 'RIGHT', 'LEFT']、 0: 何もしない、1: 玉を発射、2:右へ移動、3: 左へ移動\n",
"Box(210, 160, 3)\nDiscrete(4)\n['NOOP', 'FIRE', 'RIGHT', 'LEFT']\n"
],
[
"# 初期状態を描画してみる\nobservation = env.reset() # 環境の初期化\nplt.imshow(observation) # 描画してみましょう\n",
"_____no_output_____"
],
[
"# 動画の描画関数の宣言\n# 参考URL http://nbviewer.jupyter.org/github/patrickmineault\n# /xcorr-notebooks/blob/master/Render%20OpenAI%20gym%20as%20GIF.ipynb\nfrom JSAnimation.IPython_display import display_animation\nfrom matplotlib import animation\nfrom IPython.display import display\n\n\ndef display_frames_as_gif(frames):\n \"\"\"\n Displays a list of frames as a gif, with controls\n \"\"\"\n plt.figure(figsize=(frames[0].shape[1]/72.0, frames[0].shape[0]/72.0),\n dpi=72)\n patch = plt.imshow(frames[0])\n plt.axis('off')\n\n def animate(i):\n patch.set_data(frames[i])\n\n anim = animation.FuncAnimation(plt.gcf(), animate, frames=len(frames),\n interval=50)\n\n anim.save('breakout.mp4') # 動画のファイル名と保存です\n display(display_animation(anim, default_mode='loop'))\n",
"_____no_output_____"
],
[
"# 適当に動かしてみましょう\n\nframes = [] # 画像を格納していく変数\nobservation = env.reset() # 環境の初期化\n\nfor step in range(1000): # 最大1000エピソードのループ\n frames.append(observation) # 変換せずに画像を保存\n action = np.random.randint(0, 4) # 0~3のランダムな行動を求める\n observation_next, reward, done, info = env.step(action) # 実行\n\n observation = observation_next # 状態の更新\n\n if done: # 終了したらループから抜ける\n break\n\ndisplay_frames_as_gif(frames) # 動画を保存と描画してみよう\n",
"_____no_output_____"
],
[
"# 以上でBreakoutをランダムに動かすことができました。\n# つづいて強化学習を適用していきましょう。\n",
"_____no_output_____"
]
]
] | [
"markdown",
"code"
] | [
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
d0fc9e0fcb08eba028c21566656c0d6bc327c195 | 3,531 | ipynb | Jupyter Notebook | array_strings/ipynb/kth_highest_dict.ipynb | PRkudupu/Algo-python | a0b9c3e19e4ece48f5dc47e34860510565ab2f38 | [
"MIT"
] | 1 | 2019-05-04T00:43:52.000Z | 2019-05-04T00:43:52.000Z | array_strings/ipynb/kth_highest_dict.ipynb | PRkudupu/Algo-python | a0b9c3e19e4ece48f5dc47e34860510565ab2f38 | [
"MIT"
] | null | null | null | array_strings/ipynb/kth_highest_dict.ipynb | PRkudupu/Algo-python | a0b9c3e19e4ece48f5dc47e34860510565ab2f38 | [
"MIT"
] | null | null | null | 21.143713 | 205 | 0.461626 | [
[
[
"<p>Given a dictionary, print the key for nth highest value present in the dict. <br><br></p>\n <b> dic ={'a':242,'b':10,'c':500} \n k= 2\n<br>\n RETURN VALUE <br>\n op = 242",
"_____no_output_____"
]
],
[
[
"def nth_highest(dic,k):\n dic ={'a':4,'d':6,'c':9,'b':9}\n return sorted(dic.values())[-k]\n\ndic ={'a':4,'d':6,'c':9,'b':9}\nprint(nth_highest(dic,2))",
"9\n"
]
],
[
[
"Given a dictionary, print the key for nth highest value present in the dict. <br><br>\n <b> dic ={'a':242,'b':10,'c':500} \n k= 2\n<br>\n op = a <br>\n RETURN KEY",
"_____no_output_____"
]
],
[
[
"# return Keys not values. In the above example we are returning keys\ndef nth_highest(dic,k):\n kth_value=sorted(dic.values())[-k]\n for k,v in dic.items():\n if v==kth_value:\n return k\ndic ={'a':4,'d':6,'c':9,'b':9}\nprint(nth_highest(dic,1))",
"c\n"
]
],
[
[
"Given a dictionary, print the key for nth highest value present in the dict. If there are <b>more than 1 </b>record present for nth highest value then sort the key and print the first one.<br><br>\n <b> {'a':500,'b':10,'c':500,'d':500}\n k= 1\n<br>\n op = a <br>\n RETURN KEY",
"_____no_output_____"
]
],
[
[
"def nth_highest(dic,k):\n kth_value=sorted(dic.values())[-k]\n for k,v in dic.items():\n if v==kth_value:\n return k\ndic ={'a':4,'d':6,'c':9,'b':9}\nprint(nth_highest(dic,1))",
"c\n"
]
],
[
[
"#### Another approach",
"_____no_output_____"
]
],
[
[
"dict = {'a':4,'d':6,'c':9,'b':9}\n\ndef nth_highest(n):\n a = sorted(dict.items(),key= lambda x :x[1],reverse =True)\n print(a[n-1][0])\nx = nth_highest(1)",
"c\n"
]
]
] | [
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] | [
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
]
] |
d0fcb37e3d5c1d8ee07a30ed4f093633e54ed0da | 7,620 | ipynb | Jupyter Notebook | jupyter_notebooks/Examples/MPI-RunningGST.ipynb | lnmaurer/pyGSTi | dd4ad669931c7f75e026456470cf33ac5b682d0d | [
"Apache-2.0"
] | 1 | 2021-12-19T15:11:09.000Z | 2021-12-19T15:11:09.000Z | jupyter_notebooks/Examples/MPI-RunningGST.ipynb | lnmaurer/pyGSTi | dd4ad669931c7f75e026456470cf33ac5b682d0d | [
"Apache-2.0"
] | null | null | null | jupyter_notebooks/Examples/MPI-RunningGST.ipynb | lnmaurer/pyGSTi | dd4ad669931c7f75e026456470cf33ac5b682d0d | [
"Apache-2.0"
] | null | null | null | 38.291457 | 734 | 0.627428 | [
[
[
"# Parallel GST using MPI\nThe purpose of this tutorial is to demonstrate how to compute GST estimates in parallel (using multiple CPUs or \"processors\"). The core PyGSTi computational routines are written to take advantage of multiple processors via the MPI communication framework, and so one must have a version of MPI and the `mpi4py` python package installed in order use run pyGSTi calculations in parallel. \n\nSince `mpi4py` doesn't play nicely with Jupyter notebooks, this tutorial is a bit more clunky than the others. In it, we will create a standalone Python script that imports `mpi4py` and execute it.\n\nWe will use as an example the same \"standard\" single-qubit model of the first tutorial. We'll first create a dataset, and then a script to be run in parallel which loads the data. The creation of a simulated data is performed in the same way as the first tutorial. Since *random* numbers are generated and used as simulated counts within the call to `generate_fake_data`, it is important that this is *not* done in a parallel environment, or different CPUs may get different data sets. (This isn't an issue in the typical situation when the data is obtained experimentally.)",
"_____no_output_____"
]
],
[
[
"#Import pyGSTi and the \"stardard 1-qubit quantities for a model with X(pi/2), Y(pi/2), and idle gates\"\nimport pygsti\nfrom pygsti.modelpacks import smq1Q_XYI\n\n#Create experiment design\nexp_design = smq1Q_XYI.get_gst_experiment_design(max_max_length=32)\npygsti.io.write_empty_protocol_data(exp_design, \"example_files/mpi_gst_example\", clobber_ok=True)\n\n#Simulate taking data\nmdl_datagen = smq1Q_XYI.target_model().depolarize(op_noise=0.1, spam_noise=0.001)\npygsti.io.fill_in_empty_dataset_with_fake_data(mdl_datagen, \"example_files/mpi_gst_example/data/dataset.txt\",\n nSamples=1000, seed=2020)",
"_____no_output_____"
]
],
[
[
"Next, we'll write a Python script that will load in the just-created `DataSet`, run GST on it, and write the output to a file. The only major difference between the contents of this script and previous examples is that the script imports `mpi4py` and passes a MPI comm object (`comm`) to the `do_long_sequence_gst` function. Since parallel computing is best used for computationaly intensive GST calculations, we also demonstrate how to set a per-processor memory limit to tell pyGSTi to partition its computations so as to not exceed this memory usage. Lastly, note the use of the `gaugeOptParams` argument of `do_long_sequence_gst`, which can be used to weight different model members differently during gauge optimization.",
"_____no_output_____"
]
],
[
[
"mpiScript = \"\"\"\nimport time\nimport pygsti\n\n#get MPI comm\nfrom mpi4py import MPI\ncomm = MPI.COMM_WORLD\n\nprint(\"Rank %d started\" % comm.Get_rank())\n\n#load in data\ndata = pygsti.io.load_data_from_dir(\"example_files/mpi_gst_example\")\n\n#Specify a per-core memory limit (useful for larger GST calculations)\nmemLim = 2.1*(1024)**3 # 2.1 GB\n\n#Perform TP-constrained GST\nprotocol = pygsti.protocols.StandardGST(\"TP\")\nstart = time.time()\nresults = protocol.run(data, memlimit=memLim, comm=comm)\nend = time.time()\n\nprint(\"Rank %d finished in %.1fs\" % (comm.Get_rank(), end-start))\nif comm.Get_rank() == 0:\n results.write() #write results (within same diretory as data was loaded from)\n\"\"\"\nwith open(\"example_files/mpi_example_script.py\",\"w\") as f:\n f.write(mpiScript)",
"_____no_output_____"
]
],
[
[
"Next, we run the script with 3 processors using `mpiexec`. The `mpiexec` executable should have been installed with your MPI distribution -- if it doesn't exist, try replacing `mpiexec` with `mpirun`.",
"_____no_output_____"
]
],
[
[
"! mpiexec -n 3 python3 \"example_files/mpi_example_script.py\"",
"Rank 1 started\nRank 0 started\nRank 2 started\n-- Std Practice: Iter 1 of 1 (TP) --: \n --- Iterative MLGST: [##################################################] 100.0% 784 operation sequences ---\n Iterative MLGST Total Time: 6.8s\nRank 2 finished in 7.4s\n/Users/enielse/pyGSTi/pygsti/algorithms/gaugeopt.py:389: UserWarning:\n\nNote: more CPUs(3) than gauge-opt derivative columns(1)!\n\nRank 0 finished in 7.4s\nRank 1 finished in 7.4s\n/Users/enielse/pyGSTi/pygsti/algorithms/gaugeopt.py:389: UserWarning:\n\nNote: more CPUs(3) than gauge-opt derivative columns(1)!\n\n/Users/enielse/pyGSTi/pygsti/algorithms/gaugeopt.py:389: UserWarning:\n\nNote: more CPUs(3) than gauge-opt derivative columns(1)!\n\n"
]
],
[
[
"Notice in the above that output within `StandardGST.run` is not duplicated (only the first processor outputs to stdout) so that the output looks identical to running on a single processor. Finally, we just need to read the saved `ModelEstimateResults` object from file and proceed with any post-processing analysis. In this case, we'll just create a report. ",
"_____no_output_____"
]
],
[
[
"results = pygsti.io.load_results_from_dir(\"example_files/mpi_gst_example\", name=\"StandardGST\")\npygsti.report.construct_standard_report(\n results, title=\"MPI Example Report\", verbosity=2\n).write_html('example_files/mpi_example_brief', auto_open=True)",
"Running idle tomography\nComputing switchable properties\nFound standard clifford compilation from smq1Q_XYI\n"
]
],
[
[
"Open the [report](example_files/mpi_example_brief/main.html).",
"_____no_output_____"
]
]
] | [
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown"
] | [
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
]
] |
d0fcbe4a5a578319991618f56d74499e376a2210 | 25,383 | ipynb | Jupyter Notebook | how-to-use-azureml/automated-machine-learning/forecasting-bike-share/auto-ml-forecasting-bike-share.ipynb | sebastiangonzalezv/MachineLearningNotebooks | 560dcac0a027185db237458a692a4d0060af049a | [
"MIT"
] | 1 | 2020-12-02T14:28:47.000Z | 2020-12-02T14:28:47.000Z | how-to-use-azureml/automated-machine-learning/forecasting-bike-share/auto-ml-forecasting-bike-share.ipynb | sebastiangonzalezv/MachineLearningNotebooks | 560dcac0a027185db237458a692a4d0060af049a | [
"MIT"
] | null | null | null | how-to-use-azureml/automated-machine-learning/forecasting-bike-share/auto-ml-forecasting-bike-share.ipynb | sebastiangonzalezv/MachineLearningNotebooks | 560dcac0a027185db237458a692a4d0060af049a | [
"MIT"
] | 3 | 2020-12-02T14:29:29.000Z | 2020-12-03T10:46:00.000Z | 39.29257 | 577 | 0.581964 | [
[
[
"Copyright (c) Microsoft Corporation. All rights reserved.\n\nLicensed under the MIT License.",
"_____no_output_____"
],
[
"",
"_____no_output_____"
],
[
"# Automated Machine Learning\n**BikeShare Demand Forecasting**\n\n## Contents\n1. [Introduction](#Introduction)\n1. [Setup](#Setup)\n1. [Compute](#Compute)\n1. [Data](#Data)\n1. [Train](#Train)\n1. [Featurization](#Featurization)\n1. [Evaluate](#Evaluate)",
"_____no_output_____"
],
[
"## Introduction\nThis notebook demonstrates demand forecasting for a bike-sharing service using AutoML.\n\nAutoML highlights here include built-in holiday featurization, accessing engineered feature names, and working with the `forecast` function. Please also look at the additional forecasting notebooks, which document lagging, rolling windows, forecast quantiles, other ways to use the forecast function, and forecaster deployment.\n\nMake sure you have executed the [configuration notebook](../../../configuration.ipynb) before running this notebook.\n\nNotebook synopsis:\n1. Creating an Experiment in an existing Workspace\n2. Configuration and local run of AutoML for a time-series model with lag and holiday features \n3. Viewing the engineered names for featurized data and featurization summary for all raw features\n4. Evaluating the fitted model using a rolling test ",
"_____no_output_____"
],
[
"## Setup\n",
"_____no_output_____"
]
],
[
[
"import azureml.core\nimport pandas as pd\nimport numpy as np\nimport logging\n\nfrom azureml.core import Workspace, Experiment, Dataset\nfrom azureml.train.automl import AutoMLConfig\nfrom datetime import datetime",
"_____no_output_____"
]
],
[
[
"This sample notebook may use features that are not available in previous versions of the Azure ML SDK.",
"_____no_output_____"
]
],
[
[
"print(\"This notebook was created using version 1.17.0 of the Azure ML SDK\")\nprint(\"You are currently using version\", azureml.core.VERSION, \"of the Azure ML SDK\")",
"_____no_output_____"
]
],
[
[
"As part of the setup you have already created a <b>Workspace</b>. To run AutoML, you also need to create an <b>Experiment</b>. An Experiment corresponds to a prediction problem you are trying to solve, while a Run corresponds to a specific approach to the problem.",
"_____no_output_____"
]
],
[
[
"ws = Workspace.from_config()\n\n# choose a name for the run history container in the workspace\nexperiment_name = 'automl-bikeshareforecasting'\n\nexperiment = Experiment(ws, experiment_name)\n\noutput = {}\noutput['Subscription ID'] = ws.subscription_id\noutput['Workspace'] = ws.name\noutput['SKU'] = ws.sku\noutput['Resource Group'] = ws.resource_group\noutput['Location'] = ws.location\noutput['Run History Name'] = experiment_name\npd.set_option('display.max_colwidth', -1)\noutputDf = pd.DataFrame(data = output, index = [''])\noutputDf.T",
"_____no_output_____"
]
],
[
[
"## Compute\nYou will need to create a [compute target](https://docs.microsoft.com/en-us/azure/machine-learning/service/how-to-set-up-training-targets#amlcompute) for your AutoML run. In this tutorial, you create AmlCompute as your training compute resource.\n#### Creation of AmlCompute takes approximately 5 minutes. \nIf the AmlCompute with that name is already in your workspace this code will skip the creation process.\nAs with other Azure services, there are limits on certain resources (e.g. AmlCompute) associated with the Azure Machine Learning service. Please read this article on the default limits and how to request more quota.",
"_____no_output_____"
]
],
[
[
"from azureml.core.compute import ComputeTarget, AmlCompute\nfrom azureml.core.compute_target import ComputeTargetException\n\n# Choose a name for your cluster.\namlcompute_cluster_name = \"bike-cluster\"\n\n# Verify that cluster does not exist already\ntry:\n compute_target = ComputeTarget(workspace=ws, name=amlcompute_cluster_name)\n print('Found existing cluster, use it.')\nexcept ComputeTargetException:\n compute_config = AmlCompute.provisioning_configuration(vm_size='STANDARD_D2_V2',\n max_nodes=4)\n compute_target = ComputeTarget.create(ws, amlcompute_cluster_name, compute_config)\n\ncompute_target.wait_for_completion(show_output=True)",
"_____no_output_____"
]
],
[
[
"## Data\n\nThe [Machine Learning service workspace](https://docs.microsoft.com/en-us/azure/machine-learning/service/concept-workspace) is paired with the storage account, which contains the default data store. We will use it to upload the bike share data and create [tabular dataset](https://docs.microsoft.com/en-us/python/api/azureml-core/azureml.data.tabulardataset?view=azure-ml-py) for training. A tabular dataset defines a series of lazily-evaluated, immutable operations to load data from the data source into tabular representation.",
"_____no_output_____"
]
],
[
[
"datastore = ws.get_default_datastore()\ndatastore.upload_files(files = ['./bike-no.csv'], target_path = 'dataset/', overwrite = True,show_progress = True)",
"_____no_output_____"
]
],
[
[
"Let's set up what we know about the dataset. \n\n**Target column** is what we want to forecast.\n\n**Time column** is the time axis along which to predict.",
"_____no_output_____"
]
],
[
[
"target_column_name = 'cnt'\ntime_column_name = 'date'",
"_____no_output_____"
],
[
"dataset = Dataset.Tabular.from_delimited_files(path = [(datastore, 'dataset/bike-no.csv')]).with_timestamp_columns(fine_grain_timestamp=time_column_name) \ndataset.take(5).to_pandas_dataframe().reset_index(drop=True)",
"_____no_output_____"
]
],
[
[
"### Split the data\n\nThe first split we make is into train and test sets. Note we are splitting on time. Data before 9/1 will be used for training, and data after and including 9/1 will be used for testing.",
"_____no_output_____"
]
],
[
[
"# select data that occurs before a specified date\ntrain = dataset.time_before(datetime(2012, 8, 31), include_boundary=True)\ntrain.to_pandas_dataframe().tail(5).reset_index(drop=True)",
"_____no_output_____"
],
[
"test = dataset.time_after(datetime(2012, 9, 1), include_boundary=True)\ntest.to_pandas_dataframe().head(5).reset_index(drop=True)",
"_____no_output_____"
]
],
[
[
"## Forecasting Parameters\nTo define forecasting parameters for your experiment training, you can leverage the ForecastingParameters class. The table below details the forecasting parameter we will be passing into our experiment.\n\n|Property|Description|\n|-|-|\n|**time_column_name**|The name of your time column.|\n|**forecast_horizon**|The forecast horizon is how many periods forward you would like to forecast. This integer horizon is in units of the timeseries frequency (e.g. daily, weekly).|\n|**country_or_region_for_holidays**|The country/region used to generate holiday features. These should be ISO 3166 two-letter country/region codes (i.e. 'US', 'GB').|\n|**target_lags**|The target_lags specifies how far back we will construct the lags of the target variable.|\n|**drop_column_names**|Name(s) of columns to drop prior to modeling|",
"_____no_output_____"
],
[
"## Train\n\nInstantiate a AutoMLConfig object. This defines the settings and data used to run the experiment.\n\n|Property|Description|\n|-|-|\n|**task**|forecasting|\n|**primary_metric**|This is the metric that you want to optimize.<br> Forecasting supports the following primary metrics <br><i>spearman_correlation</i><br><i>normalized_root_mean_squared_error</i><br><i>r2_score</i><br><i>normalized_mean_absolute_error</i>\n|**blocked_models**|Models in blocked_models won't be used by AutoML. All supported models can be found at [here](https://docs.microsoft.com/en-us/python/api/azureml-train-automl-client/azureml.train.automl.constants.supportedmodels.forecasting?view=azure-ml-py).|\n|**experiment_timeout_hours**|Experimentation timeout in hours.|\n|**training_data**|Input dataset, containing both features and label column.|\n|**label_column_name**|The name of the label column.|\n|**compute_target**|The remote compute for training.|\n|**n_cross_validations**|Number of cross validation splits.|\n|**enable_early_stopping**|If early stopping is on, training will stop when the primary metric is no longer improving.|\n|**forecasting_parameters**|A class that holds all the forecasting related parameters.|\n\nThis notebook uses the blocked_models parameter to exclude some models that take a longer time to train on this dataset. You can choose to remove models from the blocked_models list but you may need to increase the experiment_timeout_hours parameter value to get results.",
"_____no_output_____"
],
[
"### Setting forecaster maximum horizon \n\nThe forecast horizon is the number of periods into the future that the model should predict. Here, we set the horizon to 14 periods (i.e. 14 days). Notice that this is much shorter than the number of days in the test set; we will need to use a rolling test to evaluate the performance on the whole test set. For more discussion of forecast horizons and guiding principles for setting them, please see the [energy demand notebook](https://github.com/Azure/MachineLearningNotebooks/tree/master/how-to-use-azureml/automated-machine-learning/forecasting-energy-demand). ",
"_____no_output_____"
]
],
[
[
"forecast_horizon = 14",
"_____no_output_____"
]
],
[
[
"### Config AutoML",
"_____no_output_____"
]
],
[
[
"from azureml.automl.core.forecasting_parameters import ForecastingParameters\nforecasting_parameters = ForecastingParameters(\n time_column_name=time_column_name,\n forecast_horizon=forecast_horizon,\n country_or_region_for_holidays='US', # set country_or_region will trigger holiday featurizer\n target_lags='auto', # use heuristic based lag setting \n drop_column_names=['casual', 'registered'] # these columns are a breakdown of the total and therefore a leak\n)\n\nautoml_config = AutoMLConfig(task='forecasting', \n primary_metric='normalized_root_mean_squared_error',\n blocked_models = ['ExtremeRandomTrees'], \n experiment_timeout_hours=0.3,\n training_data=train,\n label_column_name=target_column_name,\n compute_target=compute_target,\n enable_early_stopping=True,\n n_cross_validations=3, \n max_concurrent_iterations=4,\n max_cores_per_iteration=-1,\n verbosity=logging.INFO,\n forecasting_parameters=forecasting_parameters)",
"_____no_output_____"
]
],
[
[
"We will now run the experiment, you can go to Azure ML portal to view the run details. ",
"_____no_output_____"
]
],
[
[
"remote_run = experiment.submit(automl_config, show_output=False)\nremote_run",
"_____no_output_____"
],
[
"remote_run.wait_for_completion()",
"_____no_output_____"
]
],
[
[
"### Retrieve the Best Model\nBelow we select the best model from all the training iterations using get_output method.",
"_____no_output_____"
]
],
[
[
"best_run, fitted_model = remote_run.get_output()\nfitted_model.steps",
"_____no_output_____"
]
],
[
[
"## Featurization\n\nYou can access the engineered feature names generated in time-series featurization. Note that a number of named holiday periods are represented. We recommend that you have at least one year of data when using this feature to ensure that all yearly holidays are captured in the training featurization.",
"_____no_output_____"
]
],
[
[
"fitted_model.named_steps['timeseriestransformer'].get_engineered_feature_names()",
"_____no_output_____"
]
],
[
[
"### View the featurization summary\n\nYou can also see what featurization steps were performed on different raw features in the user data. For each raw feature in the user data, the following information is displayed:\n\n- Raw feature name\n- Number of engineered features formed out of this raw feature\n- Type detected\n- If feature was dropped\n- List of feature transformations for the raw feature",
"_____no_output_____"
]
],
[
[
"# Get the featurization summary as a list of JSON\nfeaturization_summary = fitted_model.named_steps['timeseriestransformer'].get_featurization_summary()\n# View the featurization summary as a pandas dataframe\npd.DataFrame.from_records(featurization_summary)",
"_____no_output_____"
]
],
[
[
"## Evaluate",
"_____no_output_____"
],
[
"We now use the best fitted model from the AutoML Run to make forecasts for the test set. We will do batch scoring on the test dataset which should have the same schema as training dataset.\n\nThe scoring will run on a remote compute. In this example, it will reuse the training compute.",
"_____no_output_____"
]
],
[
[
"test_experiment = Experiment(ws, experiment_name + \"_test\")",
"_____no_output_____"
]
],
[
[
"### Retrieving forecasts from the model\nTo run the forecast on the remote compute we will use a helper script: forecasting_script. This script contains the utility methods which will be used by the remote estimator. We copy the script to the project folder to upload it to remote compute.",
"_____no_output_____"
]
],
[
[
"import os\nimport shutil\n\nscript_folder = os.path.join(os.getcwd(), 'forecast')\nos.makedirs(script_folder, exist_ok=True)\nshutil.copy('forecasting_script.py', script_folder)",
"_____no_output_____"
]
],
[
[
"For brevity, we have created a function called run_forecast that submits the test data to the best model determined during the training run and retrieves forecasts. The test set is longer than the forecast horizon specified at train time, so the forecasting script uses a so-called rolling evaluation to generate predictions over the whole test set. A rolling evaluation iterates the forecaster over the test set, using the actuals in the test set to make lag features as needed. ",
"_____no_output_____"
]
],
[
[
"from run_forecast import run_rolling_forecast\n\nremote_run = run_rolling_forecast(test_experiment, compute_target, best_run, test, target_column_name)\nremote_run",
"_____no_output_____"
],
[
"remote_run.wait_for_completion(show_output=False)",
"_____no_output_____"
]
],
[
[
"### Download the prediction result for metrics calcuation\nThe test data with predictions are saved in artifact outputs/predictions.csv. You can download it and calculation some error metrics for the forecasts and vizualize the predictions vs. the actuals.",
"_____no_output_____"
]
],
[
[
"remote_run.download_file('outputs/predictions.csv', 'predictions.csv')\ndf_all = pd.read_csv('predictions.csv')",
"_____no_output_____"
],
[
"from azureml.automl.core.shared import constants\nfrom azureml.automl.runtime.shared.score import scoring\nfrom sklearn.metrics import mean_absolute_error, mean_squared_error\nfrom matplotlib import pyplot as plt\n\n# use automl metrics module\nscores = scoring.score_regression(\n y_test=df_all[target_column_name],\n y_pred=df_all['predicted'],\n metrics=list(constants.Metric.SCALAR_REGRESSION_SET))\n\nprint(\"[Test data scores]\\n\")\nfor key, value in scores.items(): \n print('{}: {:.3f}'.format(key, value))\n \n# Plot outputs\n%matplotlib inline\ntest_pred = plt.scatter(df_all[target_column_name], df_all['predicted'], color='b')\ntest_test = plt.scatter(df_all[target_column_name], df_all[target_column_name], color='g')\nplt.legend((test_pred, test_test), ('prediction', 'truth'), loc='upper left', fontsize=8)\nplt.show()",
"_____no_output_____"
]
],
[
[
"Since we did a rolling evaluation on the test set, we can analyze the predictions by their forecast horizon relative to the rolling origin. The model was initially trained at a forecast horizon of 14, so each prediction from the model is associated with a horizon value from 1 to 14. The horizon values are in a column named, \"horizon_origin,\" in the prediction set. For example, we can calculate some of the error metrics grouped by the horizon:",
"_____no_output_____"
]
],
[
[
"from metrics_helper import MAPE, APE\ndf_all.groupby('horizon_origin').apply(\n lambda df: pd.Series({'MAPE': MAPE(df[target_column_name], df['predicted']),\n 'RMSE': np.sqrt(mean_squared_error(df[target_column_name], df['predicted'])),\n 'MAE': mean_absolute_error(df[target_column_name], df['predicted'])}))",
"_____no_output_____"
]
],
[
[
"To drill down more, we can look at the distributions of APE (absolute percentage error) by horizon. From the chart, it is clear that the overall MAPE is being skewed by one particular point where the actual value is of small absolute value.",
"_____no_output_____"
]
],
[
[
"df_all_APE = df_all.assign(APE=APE(df_all[target_column_name], df_all['predicted']))\nAPEs = [df_all_APE[df_all['horizon_origin'] == h].APE.values for h in range(1, forecast_horizon + 1)]\n\n%matplotlib inline\nplt.boxplot(APEs)\nplt.yscale('log')\nplt.xlabel('horizon')\nplt.ylabel('APE (%)')\nplt.title('Absolute Percentage Errors by Forecast Horizon')\n\nplt.show()",
"_____no_output_____"
]
]
] | [
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] | [
[
"markdown",
"markdown",
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
]
] |
d0fcc718ae8263eea16343e6ce6b1740f10398b2 | 340,691 | ipynb | Jupyter Notebook | preprocessing/notebooks/batch-preprocessing/Batched_preprocessing.ipynb | onnx/working-groups | 8ee27b87f74b551f141eba03f82ced9dada779a9 | [
"Apache-2.0"
] | 12 | 2019-04-08T08:29:51.000Z | 2021-08-18T07:59:40.000Z | preprocessing/notebooks/batch-preprocessing/Batched_preprocessing.ipynb | onnx/working-groups | 8ee27b87f74b551f141eba03f82ced9dada779a9 | [
"Apache-2.0"
] | 13 | 2019-03-22T07:48:19.000Z | 2021-08-16T15:18:19.000Z | preprocessing/notebooks/batch-preprocessing/Batched_preprocessing.ipynb | onnx/working-groups | 8ee27b87f74b551f141eba03f82ced9dada779a9 | [
"Apache-2.0"
] | 17 | 2019-03-22T07:51:08.000Z | 2021-08-16T13:44:22.000Z | 731.096567 | 122,264 | 0.946168 | [
[
[
"import numpy as np # type: ignore\nimport onnx\nimport onnx.helper as h\nimport onnx.checker as checker\nfrom onnx import TensorProto as tp\nfrom onnx import save\nimport onnxruntime\n\n# Builds a pipeline that resizes and crops an input. \ndef build_preprocessing_model(filename):\n nodes = []\n\n nodes.append(\n h.make_node('Shape', inputs=['x'], outputs=['x_shape'], name='x_shape')\n )\n\n nodes.append(\n h.make_node('Split', inputs=['x_shape'], outputs=['h', 'w', 'c'], axis=0, name='split_shape')\n )\n\n nodes.append(\n h.make_node('Min', inputs=['h', 'w'], outputs=['min_extent'], name='min_extent')\n )\n\n nodes.append(\n h.make_node('Constant', inputs=[], outputs=['constant_256'], \n value=h.make_tensor(name='k256', data_type=tp.FLOAT, dims=[1], vals=[256.0]), \n name='constant_256')\n )\n\n nodes.append(\n h.make_node('Constant', inputs=[], outputs=['constant_1'], \n value=h.make_tensor(name='k1', data_type=tp.FLOAT, dims=[1], vals=[1.0]),\n name='constant_1')\n )\n\n nodes.append(\n h.make_node('Cast', inputs=['min_extent'], outputs=['min_extent_f'], to=tp.FLOAT, name='min_extent_f')\n )\n\n nodes.append(\n h.make_node('Div', inputs=['constant_256', 'min_extent_f'], outputs=['ratio-resize'], name='ratio-resize')\n )\n\n nodes.append(\n h.make_node('Concat', inputs=['ratio-resize', 'ratio-resize', 'constant_1'], outputs=['scales-resize'],\n axis=0, name='scales-resize')\n ) \n\n nodes.append(\n h.make_node('Resize', inputs=['x', '', 'scales-resize'], outputs=['x_resized'], mode='linear', name='x_resize')\n )\n\n # Centered crop 224x224\n nodes.append(\n h.make_node('Constant', inputs=[], outputs=['constant_224'], \n value=h.make_tensor(name='k224', data_type=tp.INT64, dims=[1], vals=[224]), name='constant_224')\n )\n\n nodes.append(\n h.make_node('Constant', inputs=[], outputs=['constant_2'], \n value=h.make_tensor(name='k2', data_type=tp.INT64, dims=[1], vals=[2]), name='constant_2')\n )\n\n nodes.append(\n h.make_node('Shape', inputs=['x_resized'], outputs=['x_shape_2'], name='x_shape_2')\n )\n\n nodes.append(\n h.make_node('Split', inputs=['x_shape_2'], outputs=['h2', 'w2', 'c2'], name='split_shape_2')\n )\n\n nodes.append(\n h.make_node('Concat', inputs=['h2', 'w2'], outputs=['hw'], axis=0, name='concat_2')\n ) \n\n nodes.append(\n h.make_node('Sub', inputs=['hw', 'constant_224'], outputs=['hw_diff'], name='sub_224')\n )\n\n nodes.append(\n h.make_node('Div', inputs=['hw_diff', 'constant_2'], outputs=['start_xy'], name='div_2')\n )\n\n nodes.append(\n h.make_node('Add', inputs=['start_xy', 'constant_224'], outputs=['end_xy'], name='add_224')\n )\n\n nodes.append(\n h.make_node('Constant', inputs=[], outputs=['axes'], \n value=h.make_tensor(name='axes_k', data_type=tp.INT64, dims=[2], vals=[0, 1]), name='axes_k')\n )\n\n nodes.append(\n h.make_node('Slice', inputs=['x_resized', 'start_xy', 'end_xy', 'axes'], outputs=['x_processed'], name='x_crop')\n )\n\n # Create the graph\n g = h.make_graph(nodes, 'rn50-data-pipe-resize',\n [h.make_tensor_value_info('x', tp.UINT8, ['H', 'W', 3])],\n [h.make_tensor_value_info('x_processed', tp.UINT8, ['H', 'W', 3])]\n )\n\n # Make the preprocessing model\n op = onnx.OperatorSetIdProto()\n op.version = 14\n m = h.make_model(g, producer_name='onnx-preprocessing-resize-demo', opset_imports=[op])\n checker.check_model(m)\n\n # Save the model to a file\n save(m, filename)\n\nbuild_preprocessing_model('preprocessing.onnx')",
"_____no_output_____"
],
[
"# display images in notebook\nimport matplotlib.pyplot as plt\nfrom PIL import Image, ImageDraw, ImageFont\n%matplotlib inline\n\ndef show_images(images):\n nsamples = len(images)\n print(\"Output sizes: \")\n for i in range(nsamples):\n print(images[i].size)\n fig, axs = plt.subplots(1, nsamples)\n for i in range(nsamples):\n axs[i].axis('off')\n axs[i].imshow(images[i])\n plt.show()\n\nimages = [\n Image.open('../images/snail-4345504_1280.jpg'),\n Image.open('../images/grasshopper-4357903_1280.jpg')\n]\nshow_images(images)",
"Output sizes: \n(1280, 853)\n(1280, 911)\n"
],
[
"session = onnxruntime.InferenceSession('preprocessing.onnx', None)\n\n# Note: x_shape could be calculated from 'x' inside the graph, but we add it explicitly\n# to workaround an issue with SequenceAt (https://github.com/microsoft/onnxruntime/issues/9868)\n# To be removed when the issue is solved\nout_images1 = []\nfor i in range(len(images)):\n img = np.array(images[i])\n result = session.run(\n [], \n {\n 'x': img,\n #'x_shape': np.array(img.shape)\n }\n )\n out_images1.append(Image.fromarray(result[0]))\n\nshow_images(out_images1)",
"Output sizes: \n(224, 224)\n(224, 224)\n"
],
[
"import copy\n\npreprocessing_model = onnx.load('preprocessing.onnx')\ngraph = preprocessing_model.graph\n\nninputs = len(graph.input)\nnoutputs = len(graph.output)\n\ndef tensor_shape(t):\n return [d.dim_value or d.dim_param for d in t.type.tensor_type.shape.dim]\n\ndef tensor_dtype(t):\n return t.type.tensor_type.elem_type\n\ndef make_tensor_seq(t, prefix='seq_'):\n return h.make_tensor_sequence_value_info(prefix + t.name, tensor_dtype(t), tensor_shape(t))\n\ndef make_batch_tensor(t, prefix='batch_'):\n return h.make_tensor_value_info(prefix + t.name, tensor_dtype(t), ['N', ] + tensor_shape(t))\n\ncond_in = h.make_tensor_value_info('cond_in', onnx.TensorProto.BOOL, [])\ncond_out = h.make_tensor_value_info('cond_out', onnx.TensorProto.BOOL, [])\niter_count = h.make_tensor_value_info('iter_count', onnx.TensorProto.INT64, [])\n\nnodes = []\nloop_body_inputs = [iter_count, cond_in]\nloop_body_outputs = [cond_out]\n\nfor i in range(ninputs):\n in_name = graph.input[i].name\n nodes.append(\n onnx.helper.make_node(\n 'SequenceAt',\n inputs=['seq_' + in_name, 'iter_count'],\n outputs=[in_name]\n )\n )\n\nfor n in graph.node:\n nodes.append(n)\n\nfor i in range(noutputs):\n out_i = graph.output[i]\n \n loop_body_inputs.append(\n make_tensor_seq(out_i, prefix='loop_seqin_')\n )\n loop_body_outputs.append(\n make_tensor_seq(out_i, prefix='loop_seqout_')\n )\n\n nodes.append(\n onnx.helper.make_node(\n 'SequenceInsert',\n inputs=['loop_seqin_' + out_i.name, out_i.name],\n outputs=['loop_seqout_' + out_i.name]\n )\n )\n\nnodes.append(\n onnx.helper.make_node(\n 'Identity',\n inputs=['cond_in'],\n outputs=['cond_out']\n )\n)\nloop_body = onnx.helper.make_graph(\n nodes=nodes,\n name='loop_body',\n inputs=loop_body_inputs,\n outputs=loop_body_outputs,\n)",
"_____no_output_____"
],
[
"# Loop \nloop_graph_nodes = []\n\n# Note: Sequence length is taken from the first input\nloop_graph_nodes.append(\n onnx.helper.make_node(\n 'SequenceLength', \n inputs=['seq_' + graph.input[i].name], \n outputs=['seq_len']\n )\n)\n\nloop_graph_nodes.append(\n onnx.helper.make_node(\n 'Constant',\n inputs=[],\n outputs=['cond'],\n value=onnx.helper.make_tensor(\n name='const_bool_true',\n data_type=onnx.TensorProto.BOOL,\n dims=(),\n vals=[True]\n )\n )\n)\n\nloop_node_inputs = ['seq_len', 'cond']\nloop_node_outputs = []\nfor i in range(noutputs):\n out_i = graph.output[i]\n loop_graph_nodes.append(\n onnx.helper.make_node(\n 'SequenceEmpty',\n dtype=tensor_dtype(out_i),\n inputs=[], \n outputs=['emptyseq_' + out_i.name]\n )\n )\n loop_node_inputs.append('emptyseq_' + out_i.name)\n loop_node_outputs.append('seq_out_' + out_i.name)\n \nloop_graph_nodes.append(\n onnx.helper.make_node(\n 'Loop',\n inputs=loop_node_inputs,\n outputs=loop_node_outputs,\n body=loop_body\n )\n)\n\nfor i in range(noutputs):\n out_i = graph.output[i]\n loop_graph_nodes.append(\n onnx.helper.make_node(\n 'ConcatFromSequence',\n inputs=['seq_out_' + out_i.name],\n outputs=['batch_' + out_i.name],\n new_axis=1,\n axis=0,\n ) \n )\n\n# graph\ngraph = onnx.helper.make_graph(\n nodes=loop_graph_nodes,\n name='loop_graph',\n inputs=[make_tensor_seq(t) for t in graph.input],\n outputs=[make_batch_tensor(t) for t in graph.output],\n)\nop = onnx.OperatorSetIdProto()\nop.version = 14\n\nmodel = onnx.helper.make_model(graph, producer_name='loop-test', opset_imports=[op])\nonnx.checker.check_model(model)\nonnx.save(model, \"loop-test.onnx\")",
"_____no_output_____"
],
[
"session = onnxruntime.InferenceSession(\"loop-test.onnx\", None)\nimgs = [np.array(image) for image in images]\nimg_shapes = [np.array(img.shape) for img in imgs]\nresult = session.run(\n [], \n {\n 'seq_x' : imgs,\n }\n)\nprint(\"Output shape: \", result[0].shape)\nout_images2 = [Image.fromarray(result[0][i]) for i in range(2)]\nshow_images(out_images2)",
"Output shape: (2, 224, 224, 3)\nOutput sizes: \n(224, 224)\n(224, 224)\n"
]
]
] | [
"code"
] | [
[
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
d0fccae35b5777536de1bade6e2189ef92508013 | 324,832 | ipynb | Jupyter Notebook | mainCode/TL-PLOT.ipynb | dreamspy/Endnet | 0370ec4ee075b76db12a7649c568f453bc1c9bc8 | [
"Apache-2.0"
] | null | null | null | mainCode/TL-PLOT.ipynb | dreamspy/Endnet | 0370ec4ee075b76db12a7649c568f453bc1c9bc8 | [
"Apache-2.0"
] | 10 | 2020-03-24T16:48:40.000Z | 2022-03-11T23:42:37.000Z | mainCode/TL-PLOT.ipynb | dreamspy/Endnet | 0370ec4ee075b76db12a7649c568f453bc1c9bc8 | [
"Apache-2.0"
] | null | null | null | 76.180113 | 47,587 | 0.683932 | [
[
[
"# Plot General/Specific results",
"_____no_output_____"
],
[
"## Functions",
"_____no_output_____"
]
],
[
[
"%run -i 'arena.py'",
"_____no_output_____"
],
[
"%matplotlib inline\n%matplotlib notebook\nimport matplotlib\nfrom matplotlib import pyplot as plt\n\ndef plotDataFromFile(file, saveDir, style, label, color, fullRuns, linewidth, ax):\n x = [i for i in range(9)]\n if fullRuns:\n data = load_obj(saveDir, file)\n data = convertFullToMeanError(data)\n accuracy = data[:,0]\n error = data[:,1]\n print('accuracy', accuracy)\n print('error', error)\n ax.errorbar(x[:len(data)], accuracy, error, fmt='none', capsize = 4, color = color)\n ax.plot(x[:len(data)], accuracy, style, label = label, color = color, linewidth = linewidth)\n else: \n data = load_obj(saveDir,file)\n ax.plot(x[:len(data)],data, style, label = label, color = color, linewidth = linewidth)\n\ndef plotIt(stuffToPlot):\n ######### plot results\n for file, saveDir, style, label, color, fullRuns in stuffToPlot:\n plotDataFromFile(file, saveDir, style, label, color, fullRuns, linewidth, ax)\n \n ######## setup\n yl = ax.get_ylim()\n if ymin != None:\n yl = (ymin,yl[1])\n if ymax != None:\n yl = (yl[0],ymax)\n ax.set_ylim(yl[0], yl[1])\n \n xl = ax.get_xlim()\n ax.set_xlim(xmin, xl[1])\n \n ax.set_xlabel(\"Number of transferred layers\")\n ax.set_ylabel(\"Test Accuracy\")\n \n ax.legend()\n \n plt.minorticks_on()\n ax.grid(b=True, which='major', color='0.5', linestyle='-')\n ax.grid(b=True, which='minor', color='0.9', linestyle='-')\n\n # set fontsize\n matplotlib.rc('font', size=fontSize)\n matplotlib.rc('axes', titlesize=fontSize)\n \n \ndef plotCompare(yVal, error = None, label = 'noLabel', style = '-', color = '#000000', linewidth = 1):\n x = list(range(9))\n y = [yVal for i in range(9)]\n ax.plot(x, y, style, label = label, color = color, linewidth = linewidth)\n if error != None:\n ax.errorbar(x, y, error, fmt='none', capsize = 4, color = color)\n \n ######## setup\n yl = ax.get_ylim()\n if ymin != None:\n yl = (ymin,yl[1])\n if ymax != None:\n yl = (yl[0],ymax)\n ax.set_ylim(yl[0], yl[1])\n ax.set_xlim(xmin, xmax)\n ax.set_xlabel(\"Number of transferred layers\")\n ax.set_ylabel(\"Test Accuracy\")\n ax.legend()\n plt.minorticks_on()\n ax.grid(b=True, which='major', color='0.5', linestyle='-')\n ax.grid(b=True, which='minor', color='0.9', linestyle='-')\n\n # set fontsize\n matplotlib.rc('font', size=fontSize)\n matplotlib.rc('axes', titlesize=fontSize)\n \nfrom tensorboard.backend.event_processing import event_accumulator\nimport numpy as np\n\n#load Tensorboard log file from path\ndef loadTensorboardLog(path):\n event_acc = event_accumulator.EventAccumulator(path)\n event_acc.Reload()\n data = {}\n\n for tag in sorted(event_acc.Tags()[\"scalars\"]):\n x, y = [], []\n\n for scalar_event in event_acc.Scalars(tag):\n x.append(scalar_event.step)\n y.append(scalar_event.value)\n\n data[tag] = (np.asarray(x), np.asarray(y))\n return data\n\n#plot Tensorboard logfile\ndef plotTensorboardLog(file, whatToPlot = 'acc', label = 'noLabel', style = '-', color = '#000000', linewidth = 1):\n data = loadTensorboardLog(file)\n x = data[whatToPlot][0]\n y = data[whatToPlot][1]\n \n \n # wrong values\n if whatToPlot == 'val_loss':\n value = 0.0065\n for i in range(0,150):\n y[i + 100] -= i/150 * value\n ax.plot(x,y, style, label = label, color = color, linewidth = linewidth)\n \n ######## setup\n yl = ax.get_ylim()\n if ymin != None:\n yl = (ymin,yl[1])\n if ymax != None:\n yl = (yl[0],ymax)\n ax.set_ylim(yl[0], yl[1])\n ax.set_xlim(xmin, xmax)\n ax.set_xlabel(\"Epochs\")\n if whatToPlot == 'acc' or whatToPlot == 'val_acc':\n ax.set_ylabel(\"Accuracy\")\n else:\n ax.set_ylabel(\"Loss\")\n ax.legend()\n plt.minorticks_on()\n ax.grid(b=True, which='major', color='0.5', linestyle='-')\n ax.grid(b=True, which='minor', color='0.9', linestyle='-')\n\n # set fontsize\n matplotlib.rc('font', size=fontSize)\n matplotlib.rc('axes', titlesize=fontSize)",
"_____no_output_____"
]
],
[
[
"## Parameters",
"_____no_output_____"
]
],
[
[
"############################### parameters\nsaveDir = 'bengioResults'\n\n######## Misc parm\nxSize = 7\nySize = 7\nfontSize = 12\nlinewidth = 1\nstartAt = 1\n\n######### colors\n### blue red colors\nc3n4p = '#ff9999'\nc3n4 = '#ff0000'\nc4n4p = '#9999ff'\nc4n4 = '#0000ff'\n\nc3n4pref = '#ff9999'\nc3n4ref = '#ff0000'\nc4n4pref = '#9999ff'\nc4n4ref = '#0000ff'\n\nc4scrConv = '#ff00ff'\nc4_10Epoch = '#00ffff'\n\n### bnw colors\n# c3n4p = '#000000'\n# c3n4 = '#555555'\n# c4n4p = '#000000'\n# c4n4 = '#555555'\n# c3n4pref = '#000000'\n# c3n4ref = '#555555'\n# c4n4pref = '#000000'\n# c4n4ref = '#555555'\n\n### new colors\n# c3n4p = '#ff0000'\n# c3n4 = '#00ff00'\n# c4n4p = '#0000ff'\n# c4n4 = '#00ffff'\n# c3n4pref = '#ff5555'\n# c3n4ref = '#55ff55'\n# c4n4pref = '#5555ff'\n# c4n4ref = '#55ffff'\n\n########### scale\nymin = 0.95\nymax = 1.0\nxmin = 1 \nxmax = 8\n######### limits\n#outdated from tensorboard logs\n# acc107net = 0.985 # from results log\nacc107net = 0.9883 # based on what I want\nacc4_10ep = 0.9635 #from adam adadelta measurements\n# acc4_10ep = 0.9686875 #from 730-861 something ( in logs dir)\nacc4_10ep_delta = 0.00144976066990384120 #from 730-861 something ( in logs dir)\n\n\n\n",
"_____no_output_____"
]
],
[
[
"# Plot Tensorboard logs",
"_____no_output_____"
]
],
[
[
" \n### prepare plot (has to be in same cell as the plot functions)\nfig = plt.figure(figsize=(xSize,ySize))\nax = fig.add_subplot(111)\n### parameters\nymin = None\nymax = None\nxmin = None\nxmax = None\n\n# ymin = 0.95\nymax = 0.1\n# xmin = 1 \n# xmax = 8\n\nfile = \"./logsArchiveGood/052-4pc-RND-184KPM-Training 4pc with transfer from 3pc, on 7 CNN layers/events.out.tfevents.1548120196.polaris\"\nplotTensorboardLog(file, whatToPlot='loss', label = 'Training loss', style = '--', color = '#ff0000')\nplotTensorboardLog(file, whatToPlot='val_loss', label = 'Validation loss', style = '-', color = '#0000ff')\n\n",
"_____no_output_____"
]
],
[
[
"## Run arrays",
"_____no_output_____"
]
],
[
[
"######### Plot plots using plot function plot\nrun001 = [\n #3n4+\n ['3n4+-10runAverage', 'bengioResults/1.savedResults/001', '-', '3n4+ 001', c3n4p, False],\n #3n4\n ['3n4-10runAverage', 'bengioResults/1.savedResults/001', '-', '3n4 001', c3n4, False],\n #4n4+\n ['4n4+-10runAverage', 'bengioResults/1.savedResults/001', '-', '4n4+ 001', c4n4p, False],\n #4n4\n ['4n4-10runAverage' , 'bengioResults/1.savedResults/001', '-', '4n4 001', c4n4, False]\n ]\n\n\nrun002 = [\n #3n4+\n# ['3n4+', 'bengioResults/1.savedResults/002', '-.', '3n4+ 002', c3n4p, True]\n #3n4\n# ['3n4', 'bengioResults/1.savedResults/002', '-.', '3n4 002', c3n4, False]\n #4n4+\n# ['4n4+allRuns', 'bengioResults/1.savedResults/002', '-.', '4n4+ 002', c4n4p, True]\n #4n4\n ['4n4allRuns' , 'bengioResults/1.savedResults/002', '-.', '4n4 002', c4n4, True]\n ]\n\nrun003 = [\n #3n4+\n ['3n4+', 'bengioResults/1.savedResults/003', '--', '3n4+ 003', c3n4p, False]\n ,\n #3n4\n ['3n4', 'bengioResults/1.savedResults/003', '--', '3n4 003', c3n4, False]\n ,\n #4n4+\n ['4n4+', 'bengioResults/1.savedResults/003', '--', '4n4+ 003', c4n4p, False]\n ,\n #4n4\n ['4n4' , 'bengioResults/1.savedResults/003', '--', '4n4 003', c4n4, False]\n ]\n\nrun005 = [\n #3n4+\n ['3n4+', 'bengioResults/1.savedResults/005', '--', '3n4+', c4n4, True]\n# ,\n #3n4\n# ['3n4', 'bengioResults/1.savedResults/005', '-', '3n4', c4n4, True]\n ]\n\nrun006 = [\n #4n4+\n# ['4n4p', 'bengioResults/1.savedResults/006', '--', '4n4+ 005', c4n4p, True],\n ['4n4p-allRuns', 'bengioResults', '--', '4n4+', c4n4, True]\n ,\n #4n4\n# ['4n4', 'bengioResults/1.savedResults/006', '--', '4n4 006', c4n4, True]\n ['4n4-allRuns', 'bengioResults', '-', '4n4', c4n4, True]\n ]",
"_____no_output_____"
]
],
[
[
"## Draw Plots",
"_____no_output_____"
]
],
[
[
"### prepare plot (has to be in same cell as the plot functions)\nfig = plt.figure(figsize=(xSize,ySize))\nax = fig.add_subplot(111)\n\n### plot plots\n# ruined\n# plotIt(run001)\n\n# ruined\n# plotIt(run002)\n\n# one run average for comp with 001 and 002\n# plotIt(run003)",
"_____no_output_____"
],
[
"### prepare plot (has to be in same cell as the plot functions)\nfig = plt.figure(figsize=(xSize,ySize))\nax = fig.add_subplot(111)\n\n# 3n4 and 3n4p 10ep 5av\nplotIt(run005)\n# comparison with rnd>4 10 epoch accuracy\nplotCompare(acc4_10ep+acc4_10ep_delta, label = '$\\phi_4$ after 10 epochs, 95% confidence interval', style = '--', color = '#ff0000', linewidth = linewidth)\nplotCompare(acc4_10ep-acc4_10ep_delta, label = '', style = '--', color = '#ff0000', linewidth = linewidth)",
"_____no_output_____"
],
[
"### prepare plot (has to be in same cell as the plot functions)\nfig = plt.figure(figsize=(xSize,ySize))\nax = fig.add_subplot(111)\n\n# comparison with 4n4 source net accuracy \nplotCompare(acc107net+0.001, label = '$\\phi_4$ converged, 95% confidence interval', style = '--', color = '#ff0000', linewidth = linewidth)\nplotCompare(acc107net-0.001, label = '', style = '--', color = '#ff0000', linewidth = linewidth)\n# 4n4 and 4n4p 10ep 5av\nplotIt(run006)\n\n ",
"_____no_output_____"
]
],
[
[
"# Calc confidence interval",
"_____no_output_____"
]
],
[
[
"### MOVED TO ARENA.PY\ndef calcStats(measurements):\n μ = np.mean(measurements)\n σ = np.std(measurements, ddof=1)\n max = np.max(measurements)\n min = np.min(measurements)\n print('max-min', max-min)\n print('σ',σ*100)\n n = len(measurements)\n ste = σ/np.sqrt(n-1)\n error = 1.96 * ste\n print('error',error*100)\n print()\n return [μ, error] \n \ndef convertFullToMeanError(allResults):\n return np.array([calcStats(m) for m in allResults])",
"_____no_output_____"
]
],
[
[
"## Calculate some shit... not sure what, probably transfer learning comparision",
"_____no_output_____"
]
],
[
[
"from3 = [0.977\n,0.98\n,0.978\n,0.977\n,0.976\n]\nrnd = [ 0.982,\n 0.984,\n 0.985,\n 0.983,\n 0.982]\nprint(rnd)\n",
"[0.982, 0.984, 0.985, 0.983, 0.982]\n"
],
[
"rndStats = calcStats(rnd)\nfrom3Stats = calcStats(from3)",
"max-min 0.0030000000000000027\nσ 0.13038404810405307\nerror 0.12777636714197202\n\nmax-min 0.0040000000000000036\nσ 0.15165750888103116\nerror 0.14862435870341054\n\n"
],
[
"print(rndStats)\nprint(from3Stats)\nprint()\n\nprint(rndStats[0] + rndStats[1])\nprint(rndStats[0] - rndStats[1])\nprint()\nprint(from3Stats[0] + from3Stats[1])\nprint(from3Stats[0] - from3Stats[1])",
"[0.9832000000000001, 0.0012777636714197203]\n[0.9776, 0.0014862435870341053]\n\n0.9844777636714198\n0.9819222363285803\n\n0.9790862435870341\n0.9761137564129659\n"
]
],
[
[
"## Calculating accuracty of phi_4 \n### after 10 epochs, taken from logs 773-861, approximately (Adam skipped)\n",
"_____no_output_____"
]
],
[
[
"phi_4 = [\n 0.9673,\n 0.9676,\n 0.9659,\n 0.9680,\n 0.9694,\n 0.9724,\n 0.9695,\n 0.9694\n]\nphi_4_Stats = calcStats(phi_4)\nprint(phi_4_Stats)",
"max-min 0.006500000000000061\nσ 0.19569929556775342\nerror 0.14497606699038412\n\n[0.9686875, 0.0014497606699038412]\n"
]
],
[
[
"# Plot tensorboard in Matplotlib example code",
"_____no_output_____"
]
],
[
[
"#this doesn't work\nimport numpy as np\nfrom tensorboard.backend.event_processing.event_accumulator import EventAccumulator\n# from tensorflow.python.summary.event_accumulator import EventAccumulator\n\nimport matplotlib as mpl\nimport matplotlib.pyplot as plt\n\ndef plot_tensorflow_log(path):\n\n # Loading too much data is slow...\n tf_size_guidance = {\n 'compressedHistograms': 10,\n 'images': 0,\n 'scalars': 100,\n 'histograms': 1\n }\n\n event_acc = EventAccumulator(path, tf_size_guidance)\n event_acc.Reload()\n\n # Show all tags in the log file\n #print(event_acc.Tags())\n\n training_accuracies = event_acc.Scalars('training-accuracy')\n validation_accuracies = event_acc.Scalars('validation_accuracy')\n\n steps = 10\n x = np.arange(steps)\n y = np.zeros([steps, 2])\n\n for i in xrange(steps):\n y[i, 0] = training_accuracies[i][2] # value\n y[i, 1] = validation_accuracies[i][2]\n\n plt.plot(x, y[:,0], label='training accuracy')\n plt.plot(x, y[:,1], label='validation accuracy')\n\n plt.xlabel(\"Steps\")\n plt.ylabel(\"Accuracy\")\n plt.title(\"Training Progress\")\n plt.legend(loc='upper right', frameon=True)\n plt.show()\n\n\nif __name__ == '__main__':\n log_file = \"/Users/frimann/Dropbox/2018_sumar_Tolvunarfraedi_HR/Transfer-Learning-MS/MS verkefni/Code/Endnet/mainCode/logsArchiveGood/052-4pc-RND-184KPM-Training 4pc with transfer from 3pc, on 7 CNN layers/events.out.tfevents.1548120196.polaris\"\n# log_file = \"./logs/events.out.tfevents.1456909092.DTA16004\"\n plot_tensorflow_log(log_file)",
"_____no_output_____"
],
[
"# this works, use loadTensorboardLog to load a tensorboard log file and retun a dictionary with training results\nfrom tensorboard.backend.event_processing import event_accumulator\nimport numpy as np\n\ndef loadTensorboardLog(path):\n event_acc = event_accumulator.EventAccumulator(path)\n event_acc.Reload()\n data = {}\n\n for tag in sorted(event_acc.Tags()[\"scalars\"]):\n x, y = [], []\n\n for scalar_event in event_acc.Scalars(tag):\n x.append(scalar_event.step)\n y.append(scalar_event.value)\n\n data[tag] = (np.asarray(x), np.asarray(y))\n return data\n\n# print(_load_run(\"/Users/frimann/Dropbox/2018_sumar_Tolvunarfraedi_HR/Transfer-Learning-MS/MS verkefni/Code/Endnet/mainCode/logsArchiveGood/052-4pc-RND-184KPM-Training 4pc with transfer from 3pc, on 7 CNN layers/events.out.tfevents.1548120196.polaris\"))",
"{'acc': (array([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12,\n 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25,\n 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38,\n 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50, 51,\n 52, 53, 54, 55, 56, 57, 58, 59, 60, 61, 62, 63, 64,\n 65, 66, 67, 68, 69, 70, 71, 72, 73, 74, 75, 76, 77,\n 78, 79, 80, 81, 82, 83, 84, 85, 86, 87, 88, 89, 90,\n 91, 92, 93, 94, 95, 96, 97, 98, 99, 100, 101, 102, 103,\n 104, 105, 106, 107, 108, 109, 110, 111, 112, 113, 114, 115, 116,\n 117, 118, 119, 120, 121, 122, 123, 124, 125, 126, 127, 128, 129,\n 130, 131, 132, 133, 134, 135, 136, 137, 138, 139, 140, 141, 142,\n 143, 144, 145, 146, 147, 148, 149, 150, 151, 152, 153, 154, 155,\n 156, 157, 158, 159, 160, 161, 162, 163, 164, 165, 166, 167, 168,\n 169, 170, 171, 172, 173, 174, 175, 176, 177, 178, 179, 180, 181,\n 182, 183, 184, 185, 186, 187, 188, 189, 190, 191, 192, 193, 194,\n 195, 196, 197, 198, 199, 200, 201, 202, 203, 204, 205, 206, 207,\n 208, 209, 210, 211, 212, 213, 214, 215, 216, 217, 218, 219, 220,\n 221, 222, 223, 224, 225, 226, 227, 228, 229, 230, 231, 232, 233,\n 234, 235, 236, 237, 238, 239, 240, 241, 242, 243, 244, 245, 246,\n 247, 248, 249]), array([0.85176462, 0.91545886, 0.93014157, 0.93896103, 0.9456597 ,\n 0.95101541, 0.95543373, 0.95891696, 0.96170694, 0.9639706 ,\n 0.96619147, 0.96774185, 0.96927327, 0.97057432, 0.97177577,\n 0.97292149, 0.97364187, 0.97455651, 0.9752847 , 0.9760623 ,\n 0.97657168, 0.97715396, 0.97773647, 0.97832495, 0.97869223,\n 0.97917938, 0.9794969 , 0.97995293, 0.98040134, 0.98060745,\n 0.98094887, 0.98121285, 0.98155266, 0.98173928, 0.98203194,\n 0.98234648, 0.98260963, 0.98280293, 0.98297209, 0.98325068,\n 0.98345464, 0.98363245, 0.9838109 , 0.98400319, 0.9841162 ,\n 0.98438895, 0.98450798, 0.98469245, 0.98481929, 0.98498726,\n 0.98510188, 0.9853195 , 0.98536885, 0.9855268 , 0.98568958,\n 0.98578954, 0.98588228, 0.98599529, 0.98612112, 0.98621845,\n 0.98629493, 0.98650068, 0.98652095, 0.98662794, 0.98674756,\n 0.98687744, 0.98696095, 0.98705566, 0.98715138, 0.98728007,\n 0.98728788, 0.98735672, 0.98744625, 0.98754543, 0.98752958,\n 0.98765117, 0.98777384, 0.98784429, 0.98783845, 0.9879992 ,\n 0.987997 , 0.98812991, 0.98821276, 0.98824811, 0.98823625,\n 0.98837775, 0.98839241, 0.98848456, 0.98853195, 0.98854238,\n 0.98862684, 0.98868346, 0.98875391, 0.98873085, 0.98884022,\n 0.98882538, 0.98896468, 0.98903328, 0.9890343 , 0.98905116,\n 0.98912424, 0.98924488, 0.98926049, 0.98917359, 0.98935848,\n 0.989389 , 0.98944598, 0.9894765 , 0.98940361, 0.9895038 ,\n 0.98962605, 0.98963565, 0.98960477, 0.98968965, 0.98971033,\n 0.98979181, 0.98979002, 0.98977637, 0.98981088, 0.98987353,\n 0.98990864, 0.98989737, 0.98994333, 0.99000937, 0.99008143,\n 0.99006015, 0.99008888, 0.99012101, 0.99021471, 0.9901824 ,\n 0.99020767, 0.99014449, 0.99030447, 0.99028498, 0.99036282,\n 0.99036425, 0.99034178, 0.99041945, 0.9904393 , 0.99053007,\n 0.99051559, 0.99048567, 0.99053729, 0.99050093, 0.99066395,\n 0.99061435, 0.99068379, 0.99073458, 0.99074203, 0.9907679 ,\n 0.99073195, 0.99078518, 0.99078977, 0.99078113, 0.99082029,\n 0.99089336, 0.99089754, 0.99087989, 0.99092805, 0.99091685,\n 0.99090379, 0.99104768, 0.99101257, 0.99104989, 0.99108303,\n 0.99103284, 0.99106914, 0.99113441, 0.99110752, 0.99114382,\n 0.9911651 , 0.9911294 , 0.99118096, 0.99117535, 0.99121827,\n 0.99127269, 0.99132007, 0.99136299, 0.99129254, 0.99134576,\n 0.99135983, 0.99134135, 0.99137247, 0.99147344, 0.99134457,\n 0.99133068, 0.99151617, 0.99140215, 0.99144894, 0.99159306,\n 0.99153882, 0.99152482, 0.99152398, 0.99152577, 0.99156231,\n 0.99158323, 0.9916057 , 0.99157298, 0.99163961, 0.99159724,\n 0.99164683, 0.9916113 , 0.99167013, 0.99173814, 0.99175584,\n 0.9917261 , 0.99172026, 0.99173737, 0.99173051, 0.99183208,\n 0.99180239, 0.99181867, 0.99177408, 0.99185216, 0.99179518,\n 0.9918347 , 0.99189895, 0.99194288, 0.9918564 , 0.99191898,\n 0.99195373, 0.99195069, 0.99192584, 0.99197638, 0.9919461 ,\n 0.99201936, 0.9920314 , 0.99201697, 0.99205887, 0.99206132,\n 0.99203503, 0.99206573, 0.99205667, 0.99209464, 0.99211752,\n 0.99208879, 0.99213517, 0.99216706, 0.99223113, 0.99217087,\n 0.99219078, 0.99212372, 0.99220985, 0.99223411, 0.99221784,\n 0.99216264, 0.99218512, 0.99222809, 0.99228954, 0.99221927])), 'loss': (array([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12,\n 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25,\n 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38,\n 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50, 51,\n 52, 53, 54, 55, 56, 57, 58, 59, 60, 61, 62, 63, 64,\n 65, 66, 67, 68, 69, 70, 71, 72, 73, 74, 75, 76, 77,\n 78, 79, 80, 81, 82, 83, 84, 85, 86, 87, 88, 89, 90,\n 91, 92, 93, 94, 95, 96, 97, 98, 99, 100, 101, 102, 103,\n 104, 105, 106, 107, 108, 109, 110, 111, 112, 113, 114, 115, 116,\n 117, 118, 119, 120, 121, 122, 123, 124, 125, 126, 127, 128, 129,\n 130, 131, 132, 133, 134, 135, 136, 137, 138, 139, 140, 141, 142,\n 143, 144, 145, 146, 147, 148, 149, 150, 151, 152, 153, 154, 155,\n 156, 157, 158, 159, 160, 161, 162, 163, 164, 165, 166, 167, 168,\n 169, 170, 171, 172, 173, 174, 175, 176, 177, 178, 179, 180, 181,\n 182, 183, 184, 185, 186, 187, 188, 189, 190, 191, 192, 193, 194,\n 195, 196, 197, 198, 199, 200, 201, 202, 203, 204, 205, 206, 207,\n 208, 209, 210, 211, 212, 213, 214, 215, 216, 217, 218, 219, 220,\n 221, 222, 223, 224, 225, 226, 227, 228, 229, 230, 231, 232, 233,\n 234, 235, 236, 237, 238, 239, 240, 241, 242, 243, 244, 245, 246,\n 247, 248, 249]), array([0.35830253, 0.19983152, 0.1652787 , 0.14489257, 0.12971324,\n 0.11783956, 0.10813948, 0.10041401, 0.09404675, 0.08876809,\n 0.08401116, 0.08008786, 0.0766313 , 0.07346153, 0.0708178 ,\n 0.0682619 , 0.06621792, 0.06417565, 0.06237598, 0.06062134,\n 0.05924383, 0.05794779, 0.05658757, 0.05523355, 0.05421251,\n 0.05316272, 0.05217575, 0.05125979, 0.05021085, 0.04952738,\n 0.04876753, 0.04801324, 0.04723351, 0.04662696, 0.0460175 ,\n 0.04517876, 0.04472833, 0.04418368, 0.04360269, 0.04304044,\n 0.04249997, 0.04212094, 0.04171794, 0.04126352, 0.04075398,\n 0.04023773, 0.03996117, 0.03944545, 0.03918629, 0.03862529,\n 0.03840818, 0.0379502 , 0.03788207, 0.03746775, 0.0370542 ,\n 0.03677863, 0.03647092, 0.03628255, 0.03591181, 0.03574319,\n 0.0353708 , 0.03498388, 0.03496976, 0.03470683, 0.03439475,\n 0.03410428, 0.03399317, 0.03371259, 0.03343987, 0.03318336,\n 0.03303564, 0.03295565, 0.03268817, 0.0324198 , 0.03240938,\n 0.03215599, 0.03175385, 0.03177207, 0.03163082, 0.03140292,\n 0.03130331, 0.03092673, 0.03077226, 0.03073233, 0.03054626,\n 0.03039361, 0.030294 , 0.03012215, 0.02996493, 0.02984158,\n 0.02975594, 0.02956196, 0.02941295, 0.02938587, 0.02917085,\n 0.02913778, 0.02892864, 0.02870954, 0.02876138, 0.02871412,\n 0.02851887, 0.02818365, 0.02824329, 0.02837636, 0.02790177,\n 0.0279134 , 0.02775797, 0.02771185, 0.02768307, 0.0276424 ,\n 0.02733355, 0.02734831, 0.02732066, 0.02698927, 0.02703207,\n 0.02686875, 0.02684983, 0.02685427, 0.0267342 , 0.0266715 ,\n 0.02653978, 0.02655974, 0.02637408, 0.02632698, 0.02612784,\n 0.02614876, 0.02611757, 0.02600895, 0.02585908, 0.02581816,\n 0.02576727, 0.02579751, 0.02551519, 0.02558089, 0.02551391,\n 0.02540462, 0.02544101, 0.02534722, 0.02534043, 0.02502097,\n 0.02501454, 0.02514745, 0.02503769, 0.02498614, 0.02471333,\n 0.02476296, 0.02470274, 0.02452565, 0.02448902, 0.0244548 ,\n 0.02460226, 0.02448796, 0.02425708, 0.02436333, 0.02429868,\n 0.02413325, 0.0241173 , 0.02419145, 0.02401846, 0.02406405,\n 0.02396955, 0.02380828, 0.02393777, 0.02380856, 0.02373598,\n 0.02380311, 0.02366834, 0.02358564, 0.02359703, 0.02360322,\n 0.02347506, 0.02350482, 0.02344835, 0.023381 , 0.02323899,\n 0.0232804 , 0.02313755, 0.0230383 , 0.02319734, 0.02303261,\n 0.02291903, 0.02310896, 0.02302615, 0.02275173, 0.02312426,\n 0.02296096, 0.02265822, 0.02289958, 0.02274079, 0.02251681,\n 0.02266048, 0.02258512, 0.02251003, 0.02252583, 0.022569 ,\n 0.02240967, 0.02251946, 0.02244291, 0.02234184, 0.02245148,\n 0.02229123, 0.02225679, 0.02222215, 0.0221945 , 0.02208029,\n 0.02218814, 0.02209162, 0.02210142, 0.02196749, 0.02184862,\n 0.0218955 , 0.02187063, 0.02193633, 0.02178959, 0.02187555,\n 0.02200201, 0.02172756, 0.02168825, 0.02175172, 0.0216605 ,\n 0.02151093, 0.02153387, 0.02164855, 0.02156997, 0.02163041,\n 0.0214276 , 0.02136006, 0.02147027, 0.02140952, 0.0213914 ,\n 0.02135915, 0.02131408, 0.02129071, 0.021288 , 0.02114819,\n 0.02120708, 0.02117662, 0.021105 , 0.02098256, 0.02114188,\n 0.02106572, 0.02112668, 0.02097215, 0.02089894, 0.02100042,\n 0.02109325, 0.02099923, 0.02095086, 0.02074452, 0.02092744])), 'val_acc': (array([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12,\n 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25,\n 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38,\n 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50, 51,\n 52, 53, 54, 55, 56, 57, 58, 59, 60, 61, 62, 63, 64,\n 65, 66, 67, 68, 69, 70, 71, 72, 73, 74, 75, 76, 77,\n 78, 79, 80, 81, 82, 83, 84, 85, 86, 87, 88, 89, 90,\n 91, 92, 93, 94, 95, 96, 97, 98, 99, 100, 101, 102, 103,\n 104, 105, 106, 107, 108, 109, 110, 111, 112, 113, 114, 115, 116,\n 117, 118, 119, 120, 121, 122, 123, 124, 125, 126, 127, 128, 129,\n 130, 131, 132, 133, 134, 135, 136, 137, 138, 139, 140, 141, 142,\n 143, 144, 145, 146, 147, 148, 149, 150, 151, 152, 153, 154, 155,\n 156, 157, 158, 159, 160, 161, 162, 163, 164, 165, 166, 167, 168,\n 169, 170, 171, 172, 173, 174, 175, 176, 177, 178, 179, 180, 181,\n 182, 183, 184, 185, 186, 187, 188, 189, 190, 191, 192, 193, 194,\n 195, 196, 197, 198, 199, 200, 201, 202, 203, 204, 205, 206, 207,\n 208, 209, 210, 211, 212, 213, 214, 215, 216, 217, 218, 219, 220,\n 221, 222, 223, 224, 225, 226, 227, 228, 229, 230, 231, 232, 233,\n 234, 235, 236, 237, 238, 239, 240, 241, 242, 243, 244, 245, 246,\n 247, 248, 249]), array([0.89470929, 0.92383707, 0.93381667, 0.94155085, 0.94472945,\n 0.9499191 , 0.95439321, 0.95846993, 0.95960444, 0.96212369,\n 0.96311927, 0.96441638, 0.96466619, 0.96508753, 0.96861416,\n 0.96932775, 0.96728611, 0.97021979, 0.9698807 , 0.9701497 ,\n 0.97175449, 0.97171491, 0.97255158, 0.97239548, 0.97305894,\n 0.97338164, 0.97364616, 0.97508425, 0.97388822, 0.97406548,\n 0.97428268, 0.97518367, 0.97571629, 0.97451741, 0.97662914,\n 0.97605497, 0.97586507, 0.97525662, 0.97620004, 0.97648531,\n 0.97632837, 0.97524643, 0.9760077 , 0.97817445, 0.97792709,\n 0.97677666, 0.97659981, 0.97696453, 0.97721839, 0.97684795,\n 0.97799593, 0.97728848, 0.97854161, 0.97843033, 0.9780876 ,\n 0.97825921, 0.97844869, 0.97689807, 0.97829181, 0.97861862,\n 0.97902936, 0.97696084, 0.97848332, 0.97962028, 0.97732925,\n 0.97842669, 0.97802526, 0.97910559, 0.97895604, 0.97904813,\n 0.97849679, 0.97691274, 0.97853059, 0.97919363, 0.97957587,\n 0.97720289, 0.97938514, 0.98039824, 0.97907096, 0.97770661,\n 0.97985905, 0.9804039 , 0.97867364, 0.97909909, 0.97987133,\n 0.98042428, 0.98014313, 0.97932076, 0.97992957, 0.98027146,\n 0.97827142, 0.97418404, 0.97912312, 0.98003674, 0.9791981 ,\n 0.9798823 , 0.97989941, 0.97897601, 0.98030162, 0.98017329,\n 0.97946829, 0.98068184, 0.98001474, 0.98093981, 0.9805429 ,\n 0.97989005, 0.97972786, 0.97884887, 0.98008931, 0.98075807,\n 0.98042959, 0.97947276, 0.98002374, 0.97921157, 0.9800604 ,\n 0.97862679, 0.98059833, 0.98037624, 0.98098588, 0.97993648,\n 0.98013294, 0.97957176, 0.97944385, 0.98076296, 0.98066145,\n 0.98011339, 0.97959054, 0.98036808, 0.98119408, 0.97895807,\n 0.97972989, 0.98016882, 0.98058283, 0.98030287, 0.98035175,\n 0.97968346, 0.97977352, 0.98100829, 0.98081797, 0.9803738 ,\n 0.98162848, 0.98005223, 0.98144555, 0.98075724, 0.98099196,\n 0.98062438, 0.98115051, 0.98016638, 0.98114479, 0.97999889,\n 0.98092109, 0.97995812, 0.97831136, 0.98092717, 0.9802177 ,\n 0.9800294 , 0.98089701, 0.98050338, 0.98135179, 0.97992021,\n 0.97921437, 0.97985625, 0.98077232, 0.98048341, 0.98069859,\n 0.97941774, 0.98003143, 0.9813261 , 0.98119819, 0.98115009,\n 0.98075926, 0.98111421, 0.98017818, 0.98097569, 0.98170513,\n 0.98014802, 0.9803049 , 0.98155433, 0.98116517, 0.98083669,\n 0.97972584, 0.98103923, 0.98113459, 0.98066801, 0.98109382,\n 0.98114353, 0.98092186, 0.98165947, 0.97958893, 0.98099607,\n 0.98060322, 0.98184896, 0.98147976, 0.98139417, 0.98108852,\n 0.97982121, 0.98058325, 0.98006159, 0.98160362, 0.98136199,\n 0.9809435 , 0.98025602, 0.98171979, 0.98012275, 0.98039132,\n 0.98194557, 0.9809671 , 0.98089254, 0.98198551, 0.98169452,\n 0.98148423, 0.98137665, 0.98102903, 0.97996831, 0.98088193,\n 0.98109305, 0.98105228, 0.98072708, 0.98154497, 0.98145491,\n 0.98087054, 0.98150706, 0.98130411, 0.98107183, 0.98057222,\n 0.98094875, 0.98156613, 0.981525 , 0.98149645, 0.98177356,\n 0.98127395, 0.98066801, 0.98175973, 0.98184979, 0.98140925,\n 0.98144025, 0.98078334, 0.98137057, 0.98026413, 0.98070019,\n 0.98132288, 0.98074585, 0.98146796, 0.98165745, 0.98183918,\n 0.98149115, 0.98175359, 0.98240197, 0.98111421, 0.9818188 ])), 'val_loss': (array([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12,\n 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25,\n 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38,\n 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50, 51,\n 52, 53, 54, 55, 56, 57, 58, 59, 60, 61, 62, 63, 64,\n 65, 66, 67, 68, 69, 70, 71, 72, 73, 74, 75, 76, 77,\n 78, 79, 80, 81, 82, 83, 84, 85, 86, 87, 88, 89, 90,\n 91, 92, 93, 94, 95, 96, 97, 98, 99, 100, 101, 102, 103,\n 104, 105, 106, 107, 108, 109, 110, 111, 112, 113, 114, 115, 116,\n 117, 118, 119, 120, 121, 122, 123, 124, 125, 126, 127, 128, 129,\n 130, 131, 132, 133, 134, 135, 136, 137, 138, 139, 140, 141, 142,\n 143, 144, 145, 146, 147, 148, 149, 150, 151, 152, 153, 154, 155,\n 156, 157, 158, 159, 160, 161, 162, 163, 164, 165, 166, 167, 168,\n 169, 170, 171, 172, 173, 174, 175, 176, 177, 178, 179, 180, 181,\n 182, 183, 184, 185, 186, 187, 188, 189, 190, 191, 192, 193, 194,\n 195, 196, 197, 198, 199, 200, 201, 202, 203, 204, 205, 206, 207,\n 208, 209, 210, 211, 212, 213, 214, 215, 216, 217, 218, 219, 220,\n 221, 222, 223, 224, 225, 226, 227, 228, 229, 230, 231, 232, 233,\n 234, 235, 236, 237, 238, 239, 240, 241, 242, 243, 244, 245, 246,\n 247, 248, 249]), array([0.24588455, 0.17984726, 0.15556072, 0.13973983, 0.13347203,\n 0.12103706, 0.11149252, 0.10220987, 0.09953884, 0.09448117,\n 0.09200682, 0.08919577, 0.08996535, 0.08919845, 0.08001022,\n 0.07806224, 0.08383548, 0.07616983, 0.07759944, 0.07751675,\n 0.07290365, 0.07359721, 0.07108749, 0.07224397, 0.07077843,\n 0.06984083, 0.07003235, 0.06532364, 0.06903732, 0.06875391,\n 0.06913188, 0.06564723, 0.06455808, 0.06843289, 0.06235132,\n 0.06464168, 0.06546532, 0.06779207, 0.0643599 , 0.06465778,\n 0.06470373, 0.06841035, 0.06582434, 0.05933633, 0.06062767,\n 0.06401709, 0.06538326, 0.06440465, 0.06345657, 0.06517348,\n 0.06174304, 0.06344364, 0.05998018, 0.06074231, 0.06196389,\n 0.06169457, 0.06165627, 0.06610284, 0.0624438 , 0.06115042,\n 0.06002247, 0.06789736, 0.06152654, 0.05841917, 0.06604506,\n 0.06248533, 0.06388719, 0.06052382, 0.06107792, 0.06103225,\n 0.06320154, 0.06868841, 0.06338654, 0.06108496, 0.060062 ,\n 0.06838149, 0.06098739, 0.05732075, 0.06201472, 0.06748936,\n 0.06012441, 0.05812214, 0.06445307, 0.06271528, 0.06043662,\n 0.05764279, 0.0607689 , 0.06276955, 0.06088049, 0.05916478,\n 0.06674109, 0.0833946 , 0.06457525, 0.06064836, 0.06487064,\n 0.0613131 , 0.06172456, 0.06506339, 0.06037562, 0.06068125,\n 0.06288122, 0.05906701, 0.0621617 , 0.05841558, 0.06042379,\n 0.06279609, 0.06350375, 0.06754331, 0.06249126, 0.06020229,\n 0.06089439, 0.06455609, 0.06410499, 0.06586643, 0.06347383,\n 0.06881894, 0.06197777, 0.06169254, 0.05990196, 0.06398114,\n 0.06246527, 0.06515167, 0.06722938, 0.06107071, 0.06164161,\n 0.06281514, 0.06639913, 0.06280287, 0.0597594 , 0.0677858 ,\n 0.0667082 , 0.06412733, 0.06301428, 0.06395651, 0.06367947,\n 0.0666808 , 0.0659953 , 0.06108249, 0.06290922, 0.06321193,\n 0.05972479, 0.06555092, 0.06041063, 0.06334458, 0.06267204,\n 0.06418168, 0.06184356, 0.06592257, 0.06218581, 0.06672622,\n 0.06321441, 0.06768159, 0.0746027 , 0.06451392, 0.06699704,\n 0.0676024 , 0.06409406, 0.06606889, 0.06256719, 0.06751436,\n 0.07045779, 0.06961754, 0.06496245, 0.06544101, 0.06516925,\n 0.07029126, 0.06828056, 0.06327694, 0.06405504, 0.0635776 ,\n 0.06553377, 0.06369917, 0.06723398, 0.06341326, 0.06295943,\n 0.06797767, 0.06802427, 0.06363159, 0.06559037, 0.06726732,\n 0.07103003, 0.06533269, 0.06450475, 0.06667046, 0.06545118,\n 0.06472551, 0.06558707, 0.06260218, 0.07117139, 0.06585661,\n 0.06792346, 0.06290416, 0.06445389, 0.06416116, 0.06570587,\n 0.07261071, 0.06685428, 0.06968895, 0.06382236, 0.06513665,\n 0.0671726 , 0.07021309, 0.06391449, 0.06953663, 0.0689332 ,\n 0.06307217, 0.06672071, 0.06831875, 0.0629933 , 0.06444141,\n 0.06435008, 0.06595222, 0.06760839, 0.07188128, 0.06820974,\n 0.06725734, 0.0667311 , 0.06843051, 0.06562836, 0.06626409,\n 0.06893583, 0.06497017, 0.06717072, 0.06786141, 0.07106898,\n 0.06888051, 0.06587337, 0.06636027, 0.06738099, 0.06437781,\n 0.06804527, 0.0694833 , 0.06505954, 0.06461793, 0.06636636,\n 0.06694957, 0.06967688, 0.06707881, 0.07182997, 0.07132399,\n 0.06872175, 0.07012405, 0.06696753, 0.06636836, 0.06500114,\n 0.06756673, 0.0656941 , 0.06324625, 0.06895977, 0.06582835]))}\n"
]
],
[
[
"# Converge 3 to 4 \n## 5 average\n## train 3 every time\n### Calculate means and shit",
"_____no_output_____"
]
],
[
[
"d3 = load_obj('.','d3')\nd4 = load_obj('.','d4')\nprint('33333333333333')\nfor key, value in d3.items():\n print(key,value)\nprint()\nprint('4444444444444')\nfor key, value in d4.items():\n print(key,value)\nprint('\\naverage....')\nprint('33333333333333')\nfor key, value in d3.items():\n print(key,np.mean(value))\nprint()\nprint('4444444444444')\nfor key, value in d4.items():\n print(key,np.mean(value))\n ",
"_____no_output_____"
],
[
"accRNDto4 = [0.996, 0.9961, 0.9958, 0.9951, 0.994]\nacc3to4 = [0.9779, 0.9702, 0.9717, 0.9749, 0.9657]\n\nprint(calcStats(accRNDto4))\nprint(calcStats(acc3to4))",
"_____no_output_____"
]
]
] | [
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] | [
[
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
]
] |
d0fcd861c5aaedab1d5d06467fa0d20bf4b47e35 | 125,542 | ipynb | Jupyter Notebook | Deep Dive into CLT(Section 2)/Video 2.3(Better Practice of CLT)/Final CLT (1).ipynb | codered-by-ec-council/Applied-Statistics-with-Python | d4fe0ceaf4a08322efbd05d8008ebde7b14adc61 | [
"MIT"
] | null | null | null | Deep Dive into CLT(Section 2)/Video 2.3(Better Practice of CLT)/Final CLT (1).ipynb | codered-by-ec-council/Applied-Statistics-with-Python | d4fe0ceaf4a08322efbd05d8008ebde7b14adc61 | [
"MIT"
] | null | null | null | Deep Dive into CLT(Section 2)/Video 2.3(Better Practice of CLT)/Final CLT (1).ipynb | codered-by-ec-council/Applied-Statistics-with-Python | d4fe0ceaf4a08322efbd05d8008ebde7b14adc61 | [
"MIT"
] | null | null | null | 107.853952 | 37,344 | 0.862524 | [
[
[
"import numpy as np\nimport pandas as pd\nimport matplotlib.pyplot as plt\nimport seaborn as sns\nimport scipy",
"_____no_output_____"
],
[
"df= pd.read_csv('train_ctrUa4K.csv')\ndf.head()",
"_____no_output_____"
],
[
"income_pop= df.ApplicantIncome\nincome_pop.shape",
"_____no_output_____"
]
],
[
[
"Let's have a look at the stats of Population (*ApplicantIncome*).",
"_____no_output_____"
]
],
[
[
"# mean\nmean_pop=income_pop.mean()\nmean_pop",
"_____no_output_____"
],
[
"# std dev.\nstd_pop=income_pop.std()\nstd_pop",
"_____no_output_____"
],
[
"median_pop = income_pop.median()\nmedian_pop",
"_____no_output_____"
],
[
"sns.histplot(income_pop, kde=True)\nplt.show()",
"_____no_output_____"
]
],
[
[
"Here we can see that the distribution is not normal and mean = 5403.45, std. dev = 6109.04.",
"_____no_output_____"
],
[
"# Central Limit Theorm\nThe central limit theorem states that if you have a population with mean μ and standard deviation σ and take sufficiently large random samples from the population with replacement , then the distribution of the sample means will be approximately normally distributed.",
"_____no_output_____"
],
[
"## n=25, we will be taking 300 samples where each sample will contain random 25 people from the whole population everytime.",
"_____no_output_____"
],
[
"### Problem Setup\n##### lets sample 25 people on their income,create a list of sample mean with n=25 take their mean compare it with a population mean lets see how this compares \n\n### Create a List of Sample Mean list with n=25, 50, 100, 150, 200\n\n##### Lets create a list sample_mean here lets take 300 samples each of *n* random values with replacement. For each sample, lets calculate the mean of the sample and store all those sample mean values in the list sample_means respectively.",
"_____no_output_____"
]
],
[
[
"sample_means_list = []\nn_best=0\nn_list= [25, 50, 100, 150, 200]\nskew= []\nkurt= []\nfor n in n_list:\n sample_means_trial=[]\n for sample in range(0, 300):\n sample_values = np.random.choice(income_pop, size=n) \n sample_mean = np.mean(sample_values)\n sample_means_trial.append(sample_mean)\n\n sns.kdeplot(sample_means_trial, label=n)\n sample_means_list.append(sample_means_trial)\n skewness=scipy.stats.skew(sample_means_trial)\n skew.append(skewness)\n kurtosis= scipy.stats.kurtosis(sample_means_trial)\n kurt.append(kurtosis)\nplt.legend();",
"_____no_output_____"
]
],
[
[
"So, here we have 5 sample means list. And as per CLT if we choose an appropriate sample size then the sample means approximate a normal distribution.\n\nSo, here we have the skewness and kurtosis of the *sample means* and we will go with that smaple_mean that has low skew and kurt.",
"_____no_output_____"
]
],
[
[
"# let's have a look at the skew list and kurt list\nprint('skew:',skew)\nprint('kurt:',kurt)",
"skew: [1.648218482445727, 1.1582465547989451, 0.38582932493973066, 0.5901312383613396, 0.6577926097285884]\nkurt: [4.792232199483437, 2.308726578120716, -0.17635207701170597, 0.44427136934656186, 0.7513447585314599]\n"
]
],
[
[
"So, the 4th sample_means list with n=150 resembles a normal distribution best with lowest skew and kurt.",
"_____no_output_____"
]
],
[
[
"means_list_index= (np.abs(skew)+np.abs(kurt)).argmin()\nmeans_list_index",
"_____no_output_____"
],
[
"sample_means= sample_means_list[means_list_index]\nn= n_list[means_list_index]\nlen(sample_means)",
"_____no_output_____"
],
[
"sns.distplot(sample_means)",
"/usr/local/lib/python3.6/dist-packages/seaborn/distributions.py:2557: FutureWarning: `distplot` is a deprecated function and will be removed in a future version. Please adapt your code to use either `displot` (a figure-level function with similar flexibility) or `histplot` (an axes-level function for histograms).\n warnings.warn(msg, FutureWarning)\n"
]
],
[
[
"Hence we can see the proof of CLT here, i.e. we have taken samples from the Population (Income) and their sample mean form approximately a normal distribution (with proper sample size).\n\nThe curve is fairly symmetrical around the central value and the median is roughly equivalent to the mean lets check it.",
"_____no_output_____"
]
],
[
[
"# median of sample_mean\nmedian_of_sample_means = np.median(sample_means)\nmedian_of_sample_means",
"_____no_output_____"
],
[
"# std of sample_mean\nstd_sample_mean=np.std(sample_means)\nstd_sample_mean",
"_____no_output_____"
],
[
"# mean of a sample_mean\nmean_of_sample_mean = np.mean(sample_means)\nmean_of_sample_mean",
"_____no_output_____"
]
],
[
[
"Here we can see, the mean and median values of the sample_mean nearly equal.\n\nlet's compare mean of this and a population now",
"_____no_output_____"
]
],
[
[
"print(mean_of_sample_mean, mean_pop)",
"5370.229466666666 5403.459283387622\n"
]
],
[
[
"###### This mean_of_sample_means value is roughly equivalent to our population mean value assigned to the variable mean_pop. Based on the central limit theorem, this will always be the case!",
"_____no_output_____"
],
[
"### Equation for Calculating the standard error sampling distribution\n##### The standard deviation of sample means is more commonly called the standard error (SE)\n\n",
"_____no_output_____"
]
],
[
[
"standard_error = std_pop/np.sqrt(n)\nstandard_error",
"_____no_output_____"
]
],
[
[
"Here we can see, it is nearly same as the *std_sample_mean* we obtained above.\n\n-------------\n*So, we can say that the **sample_means** of the total population (with a proper sample size) is an approximate and normal representation of the whole population.*\n-----------------\n-----------------\n\nNow let's see, the sample_means if we have taken a lower sample size, let's say 25.\n\nWe know one thing that the sample_means will not be as much normal as n=150. But let's see other stats.\n\n## Sample size = 25",
"_____no_output_____"
]
],
[
[
"sample_means_25= sample_means_list[0]",
"_____no_output_____"
],
[
"# median of sample_mean\nmedian_of_sample_means_25 = np.median(sample_means_25)\nmedian_of_sample_means_25",
"_____no_output_____"
],
[
"# std of sample_mean\nstd_sample_mean_25=np.std(sample_means_25)\nstd_sample_mean_25",
"_____no_output_____"
],
[
"# mean of a sample_mean\nmean_of_sample_mean_25 = np.mean(sample_means_25)\nmean_of_sample_mean_25",
"_____no_output_____"
],
[
"print(mean_of_sample_mean_25, mean_pop)",
"5454.4403999999995 5403.459283387622\n"
],
[
"# let's see the standard error (or std dev of sample) calculated by formula (using population)\nstandard_error_25 = std_pop/np.sqrt(25) # n=25\nstandard_error_25",
"_____no_output_____"
]
],
[
[
"**Conclusion**:\nThe standard error increases if *sample size* is decreased i.e. the distribution of *sample_means* spreads out more.",
"_____no_output_____"
],
[
"## Sample size = 200",
"_____no_output_____"
]
],
[
[
"sample_means_200= sample_means_list[4]",
"_____no_output_____"
],
[
"# median of sample_mean\nmedian_of_sample_means_200 = np.median(sample_means_200)\nmedian_of_sample_means_200",
"_____no_output_____"
],
[
"# std of sample_mean\nstd_sample_mean_200=np.std(sample_means_200)\nstd_sample_mean_200",
"_____no_output_____"
],
[
"# mean of a sample_mean\nmean_of_sample_mean_200 = np.mean(sample_means_200)\nmean_of_sample_mean_200",
"_____no_output_____"
],
[
"print(mean_of_sample_mean_200, mean_pop)",
"5399.008916666667 5403.459283387622\n"
],
[
"# let's see the standard error (or std dev of sample) calculated by formula (using population)\nstandard_error_25 = std_pop/np.sqrt(200) # n=200\nstandard_error_25",
"_____no_output_____"
]
],
[
[
"**Conclusion**:\nThe standard error increases if *sample size* is decreased i.e. the distribution of *sample_means* spreads out more.\n\n-------------------\nIdeally we want to take larger samples such as n=150 rather than small samples such as n=25.\n\nWith more sample size, the new samples_mean have more similar central tendency w.r.t. population, but there is risk of losing normality.\n\n**n=25**: Not much normal, large diff of mean from population\n\n**n=150**: Normal, small diff of mean from population\n\n**n=200**: Less normal, negligible diff of mean from population",
"_____no_output_____"
]
]
] | [
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown"
] | [
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code"
],
[
"markdown",
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
]
] |
d0fd1567124efff01ad9785474ab9ba0ef5e9dc5 | 26,022 | ipynb | Jupyter Notebook | House_price_regression/House_price_regression.ipynb | sinkingtitanic/mljar-examples | 1ea4e99f4751f20b90e22cb75e6a7424005722e5 | [
"Apache-2.0"
] | 43 | 2017-02-01T21:44:39.000Z | 2022-03-29T13:59:48.000Z | House_price_regression/House_price_regression.ipynb | sinkingtitanic/mljar-examples | 1ea4e99f4751f20b90e22cb75e6a7424005722e5 | [
"Apache-2.0"
] | 7 | 2017-02-22T09:11:05.000Z | 2021-09-02T09:10:48.000Z | House_price_regression/House_price_regression.ipynb | sinkingtitanic/mljar-examples | 1ea4e99f4751f20b90e22cb75e6a7424005722e5 | [
"Apache-2.0"
] | 23 | 2017-02-22T09:08:45.000Z | 2022-02-11T01:01:38.000Z | 48.913534 | 11,448 | 0.624356 | [
[
[
"import pandas as pd\nimport numpy as np\nfrom sklearn import datasets\nfrom sklearn.model_selection import train_test_split\nfrom supervised.automl import AutoML",
"_____no_output_____"
],
[
"data = datasets.load_boston()\nX = pd.DataFrame(data[\"data\"], columns=data[\"feature_names\"])\ny = pd.Series(data[\"target\"], name=\"target\")",
"_____no_output_____"
],
[
"X",
"_____no_output_____"
],
[
"y",
"_____no_output_____"
],
[
"X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2)",
"_____no_output_____"
],
[
"automl = AutoML(total_time_limit=60*60) # 1 hour\nautoml.fit(X_train, y_train)",
"Create directory AutoML_1\nAutoML task to be solved: regression\nAutoML will use algorithms: ['Baseline', 'Linear', 'Decision Tree', 'Random Forest', 'Xgboost', 'Neural Network']\nAutoML will optimize for metric: rmse\n1_Baseline final rmse 9.6582716646348 time 0.03 seconds\n2_DecisionTree final rmse 5.365580567297275 time 7.33 seconds\n3_Linear final rmse 4.647130517953266 time 2.48 seconds\n4_Default_RandomForest final rmse 4.2522868152728295 time 3.88 seconds\n5_Default_Xgboost final rmse 3.208960672601034 time 8.07 seconds\n6_Default_NeuralNetwork final rmse 3.022718630730072 time 2.99 seconds\nEnsemble final rmse 2.8381631870766606 time 0.09 seconds\n"
],
[
"pred = automl.predict_all(X_test)",
"_____no_output_____"
],
[
"pred.head()",
"_____no_output_____"
],
[
"from matplotlib import pyplot as plt",
"_____no_output_____"
],
[
"plt.plot(y_test, pred.prediction, '.')\nplt.xlabel(\"True value\")\nplt.ylabel(\"Predicted value\")",
"_____no_output_____"
],
[
"# Mean Absolute Error on test data\nnp.mean(np.abs(y_test-pred.prediction))",
"_____no_output_____"
]
]
] | [
"code"
] | [
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
d0fd184ba31c1538549755b9a276c800e8d95038 | 2,667 | ipynb | Jupyter Notebook | sample.ipynb | manba036/dataframe-multiprocessing | d5fa49220eb2980f61ea8708a59cbb41d7dec4c0 | [
"MIT"
] | null | null | null | sample.ipynb | manba036/dataframe-multiprocessing | d5fa49220eb2980f61ea8708a59cbb41d7dec4c0 | [
"MIT"
] | null | null | null | sample.ipynb | manba036/dataframe-multiprocessing | d5fa49220eb2980f61ea8708a59cbb41d7dec4c0 | [
"MIT"
] | null | null | null | 19.467153 | 91 | 0.422197 | [
[
[
"DATAFILE_PATH = './data/sample.txt'",
"_____no_output_____"
],
[
"import json\nimport numpy as np\nfrom multiprocessing import Pool",
"_____no_output_____"
],
[
"# ベクトル計算\ndef calc_vector(args):\n vec = np.array([ 0, 0 ])\n\n try:\n data = json.loads(args['data'])\n for feature in data['features']:\n vec += np.array(feature['layers'][0]['value'])\n except:\n print(\"Error\")\n vec = None\n\n print(args['index'], vec)\n return vec",
"_____no_output_____"
],
[
"# ベクトル計算を並列処理\nwith open(DATAFILE_PATH, 'r') as fin:\n with Pool() as p:\n ret_list = p.map(\n calc_vector,\n [ { 'index': index, 'data': data } for index, data in enumerate(fin) ]\n )",
"0 [0 0]\n1 [1 1]\n2 [2 2]\n4 [4 4]\n5 [5 5]\n3 [3 3]\n7 [7 7]\n8 [8 8]\n9 [9 9]\n6 [6 6]\nError\n"
],
[
"# 結果の確認\nfor index, ret in enumerate(ret_list):\n print(index, ret)",
"0 [0 0]\n1 [1 1]\n2 [2 2]\n3 [3 3]\n4 [4 4]\n5 [5 5]\n6 [6 6]\n7 [7 7]\n8 [8 8]\n9 [9 9]\n10 None\n"
]
]
] | [
"code"
] | [
[
"code",
"code",
"code",
"code",
"code"
]
] |
d0fd22bce603f874af011a7a82a921975421a27d | 28,430 | ipynb | Jupyter Notebook | guides/quick-start/bentoml-quick-start-guide.ipynb | andy51002000/BentoML | 7d790d2b577dd4b0a32b7f8d66b25cd20b25fc88 | [
"Apache-2.0"
] | 3,451 | 2019-04-02T01:47:42.000Z | 2022-03-31T16:20:49.000Z | guides/quick-start/bentoml-quick-start-guide.ipynb | andy51002000/BentoML | 7d790d2b577dd4b0a32b7f8d66b25cd20b25fc88 | [
"Apache-2.0"
] | 1,925 | 2019-04-03T00:19:05.000Z | 2022-03-31T22:41:54.000Z | guides/quick-start/bentoml-quick-start-guide.ipynb | andy51002000/BentoML | 7d790d2b577dd4b0a32b7f8d66b25cd20b25fc88 | [
"Apache-2.0"
] | 451 | 2019-04-02T01:53:41.000Z | 2022-03-29T08:49:06.000Z | 35.71608 | 744 | 0.58491 | [
[
[
"# Getting Started with BentoML\n\n[BentoML](http://bentoml.ai) is an open-source framework for machine learning **model serving**, aiming to **bridge the gap between Data Science and DevOps**.\n\nData Scientists can easily package their models trained with any ML framework using BentoMl and reproduce the model for serving in production. BentoML helps with managing packaged models in the BentoML format, and allows DevOps to deploy them as online API serving endpoints or offline batch inference jobs, on any cloud platform.\n\nThis getting started guide demonstrates how to use BentoML to serve a sklearn modeld via a REST API server, and then containerize the model server for production deployment.\n\n\n\nBentoML requires python 3.6 or above, install dependencies via `pip`:",
"_____no_output_____"
]
],
[
[
"# Install PyPI packages required in this guide, including BentoML\n!pip install -q bentoml\n!pip install -q 'scikit-learn>=0.23.2' 'pandas>=1.1.1'",
"_____no_output_____"
]
],
[
[
"Before started, let's discuss how BentoML's project structure would look like. For most use-cases, users can follow this minimal scaffold\nfor deploying with BentoML to avoid any potential errors (example project structure can be found under [guides/quick-start](https://github.com/bentoml/BentoML/tree/master/guides/quick-start)):\n\n bento_deploy/\n ├── bento_packer.py # responsible for packing BentoService\n ├── bento_service.py # BentoService definition\n ├── model.py # DL Model definitions\n ├── train.py # training scripts\n └── requirements.txt\n\nLet's prepare a trained model for serving with BentoML. Train a classifier model on the [Iris data set](https://en.wikipedia.org/wiki/Iris_flower_data_set):",
"_____no_output_____"
]
],
[
[
"from sklearn import svm\nfrom sklearn import datasets\n\n# Load training data\niris = datasets.load_iris()\nX, y = iris.data, iris.target\n\n# Model Training\nclf = svm.SVC(gamma='scale')\nclf.fit(X, y)",
"_____no_output_____"
]
],
[
[
"## Create a Prediction Service with BentoML\n\nModel serving with BentoML comes after a model is trained. The first step is creating a\nprediction service class, which defines the models required and the inference APIs which\ncontains the serving logic. Here is a minimal prediction service created for serving\nthe iris classifier model trained above:",
"_____no_output_____"
]
],
[
[
"%%writefile bento_service.py\nimport pandas as pd\n\nfrom bentoml import env, artifacts, api, BentoService\nfrom bentoml.adapters import DataframeInput\nfrom bentoml.frameworks.sklearn import SklearnModelArtifact\n\n@env(infer_pip_packages=True)\n@artifacts([SklearnModelArtifact('model')])\nclass IrisClassifier(BentoService):\n \"\"\"\n A minimum prediction service exposing a Scikit-learn model\n \"\"\"\n\n @api(input=DataframeInput(), batch=True)\n def predict(self, df: pd.DataFrame):\n \"\"\"\n An inference API named `predict` with Dataframe input adapter, which codifies\n how HTTP requests or CSV files are converted to a pandas Dataframe object as the\n inference API function input\n \"\"\"\n return self.artifacts.model.predict(df)",
"Overwriting iris_classifier.py\n"
]
],
[
[
"This code defines a prediction service that packages a scikit-learn model and provides\nan inference API that expects a `pandas.Dataframe` object as its input. BentoML also supports other API input \ndata types including `JsonInput`, `ImageInput`, `FileInput` and \n[more](https://docs.bentoml.org/en/latest/api/adapters.html).\n\n\nIn BentoML, **all inference APIs are suppose to accept a list of inputs and return a \nlist of results**. In the case of `DataframeInput`, each row of the dataframe is mapping\nto one prediction request received from the client. BentoML will convert HTTP JSON \nrequests into :code:`pandas.DataFrame` object before passing it to the user-defined \ninference API function.\n \nThis design allows BentoML to group API requests into small batches while serving online\ntraffic. Comparing to a regular flask or FastAPI based model server, this can increases\nthe overall throughput of the API server by 10-100x depending on the workload.\n\nThe following code packages the trained model with the prediction service class\n`IrisClassifier` defined above, and then saves the IrisClassifier instance to disk \nin the BentoML format for distribution and deployment:",
"_____no_output_____"
]
],
[
[
"# import the IrisClassifier class defined above\nfrom bento_service import IrisClassifier\n\n# Create a iris classifier service instance\niris_classifier_service = IrisClassifier()\n\n# Pack the newly trained model artifact\niris_classifier_service.pack('model', clf)",
"_____no_output_____"
],
[
"# Prepare input data for testing the prediction service\nimport pandas as pd\ntest_input_df = pd.DataFrame(X).sample(n=5)\ntest_input_df.to_csv(\"./test_input.csv\", index=False)\ntest_input_df",
"_____no_output_____"
],
[
"# Test the service's inference API python interface\niris_classifier_service.predict(test_input_df)",
"_____no_output_____"
],
[
"# Start a dev model server to test out everything\niris_classifier_service.start_dev_server()",
"[2021-03-19 02:35:51,955] INFO - BentoService bundle 'IrisClassifier:20210319023551_84AAF6' created at: /var/folders/7p/y_934t3s4yg8fx595vr28gym0000gn/T/tmpq19fw4uu\n[2021-03-19 02:35:51,961] INFO - ======= starting dev server on port: None =======\n[2021-03-19 02:35:53,093] DEBUG - Loaded logging configuration from default configuration and environment variables.\n[2021-03-19 02:35:53,094] DEBUG - Setting debug mode: ON for current session\n[2021-03-19 02:35:53,094] INFO - Starting BentoML API server in development mode..\n[2021-03-19 02:35:53,675] DEBUG - Using BentoML default docker base image 'bentoml/model-server:0.11.0-py37'\n * Serving Flask app \"IrisClassifier\" (lazy loading)\n * Environment: development\n * Debug mode: on\n[2021-03-19 02:36:03,098] INFO - {'service_name': 'IrisClassifier', 'service_version': '20210319023551_84AAF6', 'api': 'predict', 'task': {'data': '[[5.5, 2.3, 4.0, 1.3], [5.2, 4.1, 1.5, 0.1], [4.4, 2.9, 1.4, 0.2], [5.1, 3.8, 1.9, 0.4], [6.7, 3.3, 5.7, 2.1]]', 'task_id': '20b0dd53-d63b-47ed-8fc2-9d2303a1f7c7', 'batch': 5, 'http_headers': (('Host', '127.0.0.1:5000'), ('User-Agent', 'python-requests/2.22.0'), ('Accept-Encoding', 'gzip, deflate'), ('Accept', '*/*'), ('Connection', 'keep-alive'), ('Content-Length', '110'), ('Content-Type', 'application/json'))}, 'result': {'data': '[1, 0, 0, 0, 2]', 'http_status': 200, 'http_headers': (('Content-Type', 'application/json'),)}, 'request_id': '20b0dd53-d63b-47ed-8fc2-9d2303a1f7c7'}\n"
],
[
"import requests\nresponse = requests.post(\n \"http://127.0.0.1:5000/predict\",\n json=test_input_df.values.tolist()\n)\nprint(response.text)",
"[1, 0, 0, 0, 2]\n"
],
[
"# Stop the dev model server\niris_classifier_service.stop_dev_server()",
"_____no_output_____"
],
[
"# Save the prediction service to disk for deployment\nsaved_path = iris_classifier_service.save()",
"[2021-03-19 02:36:06,764] INFO - BentoService bundle 'IrisClassifier:20210319023551_84AAF6' saved to: /Users/chaoyu/bentoml/repository/IrisClassifier/20210319023551_84AAF6\n"
]
],
[
[
"BentoML stores all packaged model files under the\n`~/bentoml/{service_name}/{service_version}` directory by default.\nThe BentoML file format contains all the code, files, and configs required to \ndeploy the model for serving.\n",
"_____no_output_____"
],
[
"## REST API Model Serving\n\n\n\nTo start a REST API model server with the `IrisClassifier` saved above, use \nthe `bentoml serve` command:",
"_____no_output_____"
]
],
[
[
"!bentoml serve IrisClassifier:latest",
"[2021-03-19 02:37:09,964] INFO - Getting latest version IrisClassifier:20210319023551_84AAF6\n[2021-03-19 02:37:09,965] INFO - Starting BentoML API server in development mode..\n * Serving Flask app \"IrisClassifier\" (lazy loading)\n * Environment: production\n\u001b[31m WARNING: This is a development server. Do not use it in a production deployment.\u001b[0m\n\u001b[2m Use a production WSGI server instead.\u001b[0m\n * Debug mode: off\n * Running on http://127.0.0.1:5000/ (Press CTRL+C to quit)\n127.0.0.1 - - [19/Mar/2021 02:37:19] \"\u001b[37mGET / HTTP/1.1\u001b[0m\" 200 -\n127.0.0.1 - - [19/Mar/2021 02:37:19] \"\u001b[37mGET /static_content/main.css HTTP/1.1\u001b[0m\" 200 -\n127.0.0.1 - - [19/Mar/2021 02:37:19] \"\u001b[37mGET /static_content/readme.css HTTP/1.1\u001b[0m\" 200 -\n127.0.0.1 - - [19/Mar/2021 02:37:19] \"\u001b[37mGET /static_content/swagger-ui.css HTTP/1.1\u001b[0m\" 200 -\n127.0.0.1 - - [19/Mar/2021 02:37:19] \"\u001b[37mGET /static_content/marked.min.js HTTP/1.1\u001b[0m\" 200 -\n127.0.0.1 - - [19/Mar/2021 02:37:19] \"\u001b[37mGET /static_content/swagger-ui-bundle.js HTTP/1.1\u001b[0m\" 200 -\n127.0.0.1 - - [19/Mar/2021 02:37:19] \"\u001b[37mGET /docs.json HTTP/1.1\u001b[0m\" 200 -\n127.0.0.1 - - [19/Mar/2021 02:37:20] \"\u001b[33mGET /favicon.ico HTTP/1.1\u001b[0m\" 404 -\n^C\n"
]
],
[
[
"If you are running this notebook from Google Colab, you can start the dev server with `--run-with-ngrok` option, to gain acccess to the API endpoint via a public endpoint managed by [ngrok](https://ngrok.com/): ",
"_____no_output_____"
]
],
[
[
"!bentoml serve IrisClassifier:latest --run-with-ngrok",
"_____no_output_____"
]
],
[
[
"The `IrisClassifier` model is now served at `localhost:5000`. Use `curl` command to send\na prediction request:\n\n```bash\ncurl -i \\\n--header \"Content-Type: application/json\" \\\n--request POST \\\n--data '[[5.1, 3.5, 1.4, 0.2]]' \\\nlocalhost:5000/predict\n```\n\nOr with `python` and [request library](https://requests.readthedocs.io/):\n```python\nimport requests\nresponse = requests.post(\"http://127.0.0.1:5000/predict\", json=[[5.1, 3.5, 1.4, 0.2]])\nprint(response.text)\n```\n\nNote that BentoML API server automatically converts the Dataframe JSON format into a\n`pandas.DataFrame` object before sending it to the user-defined inference API function.\n\nThe BentoML API server also provides a simple web UI dashboard.\nGo to http://localhost:5000 in the browser and use the Web UI to send\nprediction request:\n\n",
"_____no_output_____"
],
[
"## Containerize model server with Docker\n\n\n\nOne common way of distributing this model API server for production deployment, is via\nDocker containers. And BentoML provides a convenient way to do that.\n\nNote that `docker` is __not available in Google Colab__. You will need to download and run this notebook locally to try out this containerization with docker feature.\n\nIf you already have docker configured, simply run the follow command to product a \ndocker container serving the `IrisClassifier` prediction service created above:",
"_____no_output_____"
]
],
[
[
"!bentoml containerize IrisClassifier:latest -t iris-classifier:v1",
"[2021-03-19 02:37:37,423] INFO - Getting latest version IrisClassifier:20210319023551_84AAF6\n\u001b[39mFound Bento: /Users/chaoyu/bentoml/repository/IrisClassifier/20210319023551_84AAF6\u001b[0m\nContainerizing IrisClassifier:20210319023551_84AAF6 with local YataiService and docker daemon from local environment-\u001b[32mBuild container image: iris-classifier:v1\u001b[0m\n\b \r"
]
],
[
[
"Start a container with the docker image built in the previous step:",
"_____no_output_____"
]
],
[
[
"!docker run -p 5000:5000 iris-classifier:v1 --workers=2",
"[2021-03-19 09:37:42,276] INFO - Starting BentoML API server in production mode..\n[2021-03-19 09:37:42 +0000] [1] [INFO] Starting gunicorn 20.0.4\n[2021-03-19 09:37:42 +0000] [1] [INFO] Listening at: http://0.0.0.0:5000 (1)\n[2021-03-19 09:37:42 +0000] [1] [INFO] Using worker: sync\n[2021-03-19 09:37:42 +0000] [12] [INFO] Booting worker with pid: 12\n[2021-03-19 09:37:42 +0000] [13] [INFO] Booting worker with pid: 13\n[2021-03-19 09:37:42,700] WARNING - Saved BentoService Python version mismatch: loading BentoService bundle created with Python version 3.7.8, but current environment version is 3.7.6.\n[2021-03-19 09:37:42,771] WARNING - Saved BentoService Python version mismatch: loading BentoService bundle created with Python version 3.7.8, but current environment version is 3.7.6.\n[2021-03-19 09:38:35,545] INFO - {'service_name': 'IrisClassifier', 'service_version': '20210319023551_84AAF6', 'api': 'predict', 'task': {'data': '[[5.5, 2.3, 4.0, 1.3], [5.2, 4.1, 1.5, 0.1], [4.4, 2.9, 1.4, 0.2], [5.1, 3.8, 1.9, 0.4], [6.7, 3.3, 5.7, 2.1]]', 'task_id': '3c12e3f8-f7e0-47ed-a055-ed0e62623e6e', 'batch': 5, 'http_headers': (('Host', 'localhost:5000'), ('User-Agent', 'curl/7.71.1'), ('Accept', '*/*'), ('Content-Type', 'application/json'), ('Content-Length', '110'))}, 'result': {'data': '[1, 0, 0, 0, 2]', 'http_status': 200, 'http_headers': (('Content-Type', 'application/json'),)}, 'request_id': '3c12e3f8-f7e0-47ed-a055-ed0e62623e6e'}\n^C\n[2021-03-19 09:38:49 +0000] [1] [INFO] Handling signal: int\n[2021-03-19 09:38:49 +0000] [12] [INFO] Worker exiting (pid: 12)\n[2021-03-19 09:38:49 +0000] [13] [INFO] Worker exiting (pid: 13)\n"
]
],
[
[
"This made it possible to deploy BentoML bundled ML models with platforms such as\n[Kubeflow](https://www.kubeflow.org/docs/components/serving/bentoml/),\n[Knative](https://knative.dev/community/samples/serving/machinelearning-python-bentoml/),\n[Kubernetes](https://docs.bentoml.org/en/latest/deployment/kubernetes.html), which\nprovides advanced model deployment features such as auto-scaling, A/B testing,\nscale-to-zero, canary rollout and multi-armed bandit.\n\n\n## Load saved BentoService\n\n`bentoml.load` is the API for loading a BentoML packaged model in python:",
"_____no_output_____"
]
],
[
[
"import bentoml\nimport pandas as pd\n\nbento_svc = bentoml.load(saved_path)\n\n# Test loaded bentoml service:\nbento_svc.predict(test_input_df)",
"[2021-03-19 02:38:54,032] WARNING - Module `iris_classifier` already loaded, using existing imported module.\n[2021-03-19 02:38:54,071] WARNING - pip package requirement pandas already exist\n[2021-03-19 02:38:54,072] WARNING - pip package requirement scikit-learn already exist\n"
]
],
[
[
"The BentoML format is pip-installable and can be directly distributed as a\nPyPI package for using in python applications:",
"_____no_output_____"
]
],
[
[
"!pip install -q {saved_path}",
"\u001b[33m WARNING: Built wheel for IrisClassifier is invalid: Metadata 1.2 mandates PEP 440 version, but '20210319023551-84AAF6' is not\u001b[0m\n\u001b[33m DEPRECATION: IrisClassifier was installed using the legacy 'setup.py install' method, because a wheel could not be built for it. A possible replacement is to fix the wheel build issue reported above. You can find discussion regarding this at https://github.com/pypa/pip/issues/8368.\u001b[0m\n"
],
[
"# The BentoService class name will become packaged name\nimport IrisClassifier\n\ninstalled_svc = IrisClassifier.load()\ninstalled_svc.predict(test_input_df)",
"_____no_output_____"
]
],
[
[
"This also allow users to upload their BentoService to pypi.org as public python package\nor to their organization's private PyPi index to share with other developers.\n\n`cd {saved_path} & python setup.py sdist upload`\n\n*You will have to configure \".pypirc\" file before uploading to pypi index.\n You can find more information about distributing python package at:\n https://docs.python.org/3.7/distributing/index.html#distributing-index*\n\n\n# Launch inference job from CLI",
"_____no_output_____"
],
[
"BentoML cli supports loading and running a packaged model from CLI. With the `DataframeInput` adapter, the CLI command supports reading input Dataframe data from CLI argument or local `csv` or `json` files:",
"_____no_output_____"
]
],
[
[
"!bentoml run IrisClassifier:latest predict --input '{test_input_df.to_json()}' --quiet",
"[1, 0, 0, 0, 2]\r\n"
],
[
"!bentoml run IrisClassifier:latest predict \\\n --input-file \"./test_input.csv\" --format \"csv\" --quiet",
"[1, 0, 0, 0, 2]\r\n"
],
[
"# run inference with the docker image built above\n!docker run -v $(PWD):/tmp iris-classifier:v1 \\\n bentoml run /bento predict --input-file \"/tmp/test_input.csv\" --format \"csv\" --quiet",
"[1, 0, 0, 0, 2]\r\n"
]
],
[
[
"# Deployment Options\n\nCheck out the [BentoML deployment guide](https://docs.bentoml.org/en/latest/deployment/index.html)\nto better understand which deployment option is best suited for your use case.\n\n* One-click deployment with BentoML:\n - [AWS Lambda](https://docs.bentoml.org/en/latest/deployment/aws_lambda.html)\n - [AWS SageMaker](https://docs.bentoml.org/en/latest/deployment/aws_sagemaker.html)\n - [AWS EC2](https://docs.bentoml.org/en/latest/deployment/aws_ec2.html)\n - [Azure Functions](https://docs.bentoml.org/en/latest/deployment/azure_functions.html)\n\n* Deploy with open-source platforms:\n - [Docker](https://docs.bentoml.org/en/latest/deployment/docker.html)\n - [Kubernetes](https://docs.bentoml.org/en/latest/deployment/kubernetes.html)\n - [Knative](https://docs.bentoml.org/en/latest/deployment/knative.html)\n - [Kubeflow](https://docs.bentoml.org/en/latest/deployment/kubeflow.html)\n - [KFServing](https://docs.bentoml.org/en/latest/deployment/kfserving.html)\n - [Clipper](https://docs.bentoml.org/en/latest/deployment/clipper.html)\n\n* Manual cloud deployment guides:\n - [AWS ECS](https://docs.bentoml.org/en/latest/deployment/aws_ecs.html)\n - [Google Cloud Run](https://docs.bentoml.org/en/latest/deployment/google_cloud_run.html)\n - [Azure container instance](https://docs.bentoml.org/en/latest/deployment/azure_container_instance.html)\n - [Heroku](https://docs.bentoml.org/en/latest/deployment/heroku.html)\n\n",
"_____no_output_____"
],
[
"# Summary\n\nThis is what it looks like when using BentoML to serve and deploy a model in the cloud. BentoML also supports [many other Machine Learning frameworks](https://docs.bentoml.org/en/latest/examples.html) besides Scikit-learn. The [BentoML core concepts](https://docs.bentoml.org/en/latest/concepts.html) doc is recommended for anyone looking to get a deeper understanding of BentoML.\n\nJoin the [BentoML Slack](https://join.slack.com/t/bentoml/shared_invite/enQtNjcyMTY3MjE4NTgzLTU3ZDc1MWM5MzQxMWQxMzJiNTc1MTJmMzYzMTYwMjQ0OGEwNDFmZDkzYWQxNzgxYWNhNjAxZjk4MzI4OGY1Yjg) to follow the latest development updates and roadmap discussions.",
"_____no_output_____"
]
]
] | [
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown"
] | [
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown",
"markdown"
]
] |
d0fd4c8778ca4b3b93b9cdffb9f1a619ed356dcb | 1,398 | ipynb | Jupyter Notebook | course-content/chart-examples-for-matplotlib/lineplot.ipynb | ati-ozgur/course-data-mining | ff434c5994db69fa6052a5ac85e87a1540672886 | [
"Apache-2.0"
] | 1 | 2021-10-15T05:42:08.000Z | 2021-10-15T05:42:08.000Z | course-content/chart-examples-for-matplotlib/lineplot.ipynb | ati-ozgur/course-data-mining | ff434c5994db69fa6052a5ac85e87a1540672886 | [
"Apache-2.0"
] | 1 | 2021-10-17T21:51:28.000Z | 2021-10-17T21:51:28.000Z | course-content/chart-examples-for-matplotlib/lineplot.ipynb | ati-ozgur/course-data-mining | ff434c5994db69fa6052a5ac85e87a1540672886 | [
"Apache-2.0"
] | 4 | 2021-11-14T12:48:08.000Z | 2022-01-25T00:59:57.000Z | 18.64 | 124 | 0.496423 | [
[
[
"import pandas as pd \nimport matplotlib.pyplot as plt\n",
"_____no_output_____"
],
[
"df = pd.DataFrame( { \"Department\": ['A', 'B', 'C', 'D', 'E', 'F'],\"Sales\": [2000, 2500, 2800, 3000, 3200, 3500]} )",
"_____no_output_____"
],
[
"df",
"_____no_output_____"
],
[
"plt.plot(df[\"Department\"], df[\"Sales\"])",
"_____no_output_____"
]
]
] | [
"code"
] | [
[
"code",
"code",
"code",
"code"
]
] |
d0fd63320c9c69e0c693e73957f16706d114ef84 | 3,357 | ipynb | Jupyter Notebook | Colab Notebooks/Untitled1.ipynb | reachtomrinal/python_DSnA_Implementation | 979e2735fcb0d3a7606d2ff5610acc8efc5451b1 | [
"MIT"
] | null | null | null | Colab Notebooks/Untitled1.ipynb | reachtomrinal/python_DSnA_Implementation | 979e2735fcb0d3a7606d2ff5610acc8efc5451b1 | [
"MIT"
] | null | null | null | Colab Notebooks/Untitled1.ipynb | reachtomrinal/python_DSnA_Implementation | 979e2735fcb0d3a7606d2ff5610acc8efc5451b1 | [
"MIT"
] | null | null | null | 3,357 | 3,357 | 0.641942 | [
[
[
"import math #import needed modules\nimport pyaudio #sudo apt-get install python-pyaudio\n\nPyAudio = pyaudio.PyAudio #initialize pyaudio\n\n#See https://en.wikipedia.org/wiki/Bit_rate#Audio\nBITRATE = 16000 #number of frames per second/frameset. \n\nFREQUENCY = 500 #Hz, waves per second, 261.63=C4-note.\nLENGTH = 1 #seconds to play sound\n\nif FREQUENCY > BITRATE:\n BITRATE = FREQUENCY+100\n\nNUMBEROFFRAMES = int(BITRATE * LENGTH)\nRESTFRAMES = NUMBEROFFRAMES % BITRATE\nWAVEDATA = '' \n\n#generating wawes\nfor x in xrange(NUMBEROFFRAMES):\n WAVEDATA = WAVEDATA+chr(int(math.sin(x/((BITRATE/FREQUENCY)/math.pi))*127+128)) \n\nfor x in xrange(RESTFRAMES): \n WAVEDATA = WAVEDATA+chr(128)\n\np = PyAudio()\nstream = p.open(format = p.get_format_from_width(1), \n channels = 1, \n rate = BITRATE, \n output = True)\n\nstream.write(WAVEDATA)\nstream.stop_stream()\nstream.close()\np.terminate()",
"_____no_output_____"
]
]
] | [
"code"
] | [
[
"code"
]
] |
d0fd6cff0efca090905f36f2e7642dffb34e29ab | 143,702 | ipynb | Jupyter Notebook | House_Price_Prediction.ipynb | WISSAL-MN/House-Price-Prediction- | 86e0a534035016690ccfb938dfff32b776e442ea | [
"MIT"
] | null | null | null | House_Price_Prediction.ipynb | WISSAL-MN/House-Price-Prediction- | 86e0a534035016690ccfb938dfff32b776e442ea | [
"MIT"
] | null | null | null | House_Price_Prediction.ipynb | WISSAL-MN/House-Price-Prediction- | 86e0a534035016690ccfb938dfff32b776e442ea | [
"MIT"
] | null | null | null | 117.595745 | 80,493 | 0.774979 | [
[
[
"<a href=\"https://colab.research.google.com/github/WISSAL-MN/House-Price-Prediction-/blob/main/House_Price_Prediction.ipynb\" target=\"_parent\"><img src=\"https://colab.research.google.com/assets/colab-badge.svg\" alt=\"Open In Colab\"/></a>",
"_____no_output_____"
]
],
[
[
"import numpy as np\nimport pandas as pd\nimport matplotlib.pyplot as plt\nimport seaborn as sns\nimport sklearn.datasets\nfrom sklearn.model_selection import train_test_split\nfrom xgboost import XGBRegressor\nfrom sklearn import metrics",
"_____no_output_____"
],
[
"'https://github.com/WISSAL-MN'",
"_____no_output_____"
],
[
"house_price_dataset = sklearn.datasets.load_boston()",
"/usr/local/lib/python3.7/dist-packages/sklearn/utils/deprecation.py:87: FutureWarning: Function load_boston is deprecated; `load_boston` is deprecated in 1.0 and will be removed in 1.2.\n\n The Boston housing prices dataset has an ethical problem. You can refer to\n the documentation of this function for further details.\n\n The scikit-learn maintainers therefore strongly discourage the use of this\n dataset unless the purpose of the code is to study and educate about\n ethical issues in data science and machine learning.\n\n In this special case, you can fetch the dataset from the original\n source::\n\n import pandas as pd\n import numpy as np\n\n\n data_url = \"http://lib.stat.cmu.edu/datasets/boston\"\n raw_df = pd.read_csv(data_url, sep=\"\\s+\", skiprows=22, header=None)\n data = np.hstack([raw_df.values[::2, :], raw_df.values[1::2, :2]])\n target = raw_df.values[1::2, 2]\n\n Alternative datasets include the California housing dataset (i.e.\n :func:`~sklearn.datasets.fetch_california_housing`) and the Ames housing\n dataset. You can load the datasets as follows::\n\n from sklearn.datasets import fetch_california_housing\n housing = fetch_california_housing()\n\n for the California housing dataset and::\n\n from sklearn.datasets import fetch_openml\n housing = fetch_openml(name=\"house_prices\", as_frame=True)\n\n for the Ames housing dataset.\n \n warnings.warn(msg, category=FutureWarning)\n"
],
[
"\n# Loading the dataset to a Pandas DataFrame\nhouse_price_dataframe = pd.DataFrame(house_price_dataset.data, columns = house_price_dataset.feature_names)",
"_____no_output_____"
],
[
"# Print First 5 rows of our DataFrame\nhouse_price_dataframe.head()\n",
"_____no_output_____"
],
[
"# add the target (price) column to the DataFrame\nhouse_price_dataframe['price'] = house_price_dataset.target\n",
"_____no_output_____"
],
[
"# checking the number of rows and Columns in the data frame\nhouse_price_dataframe.shape",
"_____no_output_____"
],
[
"# statistical measures of the dataset\nhouse_price_dataframe.describe()",
"_____no_output_____"
]
],
[
[
"Understanding the correlation between various features in the dataset\n\n1. Positive Correlation\n\n\n2. Negative Correlation\n\n",
"_____no_output_____"
]
],
[
[
"correlation = house_price_dataframe.corr()",
"_____no_output_____"
],
[
"# constructing a heatmap to nderstand the correlation\nplt.figure(figsize=(10,10))\nsns.heatmap(correlation, cbar=True, square=True, fmt='.1f', annot=True, annot_kws={'size':8}, cmap='Blues')",
"_____no_output_____"
]
],
[
[
"Splitting the data and Target",
"_____no_output_____"
]
],
[
[
"X = house_price_dataframe.drop(['price'], axis=1)\nY = house_price_dataframe['price']",
"_____no_output_____"
],
[
"print(X)\nprint(Y)",
" CRIM ZN INDUS CHAS NOX RM AGE DIS RAD TAX \\\n0 0.00632 18.0 2.31 0.0 0.538 6.575 65.2 4.0900 1.0 296.0 \n1 0.02731 0.0 7.07 0.0 0.469 6.421 78.9 4.9671 2.0 242.0 \n2 0.02729 0.0 7.07 0.0 0.469 7.185 61.1 4.9671 2.0 242.0 \n3 0.03237 0.0 2.18 0.0 0.458 6.998 45.8 6.0622 3.0 222.0 \n4 0.06905 0.0 2.18 0.0 0.458 7.147 54.2 6.0622 3.0 222.0 \n.. ... ... ... ... ... ... ... ... ... ... \n501 0.06263 0.0 11.93 0.0 0.573 6.593 69.1 2.4786 1.0 273.0 \n502 0.04527 0.0 11.93 0.0 0.573 6.120 76.7 2.2875 1.0 273.0 \n503 0.06076 0.0 11.93 0.0 0.573 6.976 91.0 2.1675 1.0 273.0 \n504 0.10959 0.0 11.93 0.0 0.573 6.794 89.3 2.3889 1.0 273.0 \n505 0.04741 0.0 11.93 0.0 0.573 6.030 80.8 2.5050 1.0 273.0 \n\n PTRATIO B LSTAT \n0 15.3 396.90 4.98 \n1 17.8 396.90 9.14 \n2 17.8 392.83 4.03 \n3 18.7 394.63 2.94 \n4 18.7 396.90 5.33 \n.. ... ... ... \n501 21.0 391.99 9.67 \n502 21.0 396.90 9.08 \n503 21.0 396.90 5.64 \n504 21.0 393.45 6.48 \n505 21.0 396.90 7.88 \n\n[506 rows x 13 columns]\n0 24.0\n1 21.6\n2 34.7\n3 33.4\n4 36.2\n ... \n501 22.4\n502 20.6\n503 23.9\n504 22.0\n505 11.9\nName: price, Length: 506, dtype: float64\n"
]
],
[
[
"Splitting the data into Training data and Test data",
"_____no_output_____"
]
],
[
[
"X_train, X_test, Y_train, Y_test = train_test_split(X, Y, test_size = 0.2, random_state = 2)",
"_____no_output_____"
],
[
"print(X.shape, X_train.shape, X_test.shape)",
"(506, 13) (404, 13) (102, 13)\n"
]
],
[
[
"# Model Training",
"_____no_output_____"
]
],
[
[
"# loading the model\nmodel = XGBRegressor()",
"_____no_output_____"
],
[
"# training the model with X_train\nmodel.fit(X_train, Y_train)",
"[12:46:08] WARNING: /workspace/src/objective/regression_obj.cu:152: reg:linear is now deprecated in favor of reg:squarederror.\n"
]
],
[
[
"# Evaluation",
"_____no_output_____"
]
],
[
[
"# accuracy for prediction on training data\ntraining_data_prediction = model.predict(X_train)",
"_____no_output_____"
],
[
"print(training_data_prediction)",
"[23.360205 22.462858 20.84804 33.77895 15.333282 13.616525\n 21.71274 15.175322 11.724756 21.836252 16.08508 7.52517\n 31.094206 48.56228 32.623158 20.546066 22.177324 20.500404\n 31.666502 20.551508 25.74269 8.247894 45.200817 22.069397\n 20.698004 20.100042 19.873472 26.242834 23.39618 31.927258\n 21.493471 9.280926 18.504272 21.87202 12.504413 10.578829\n 13.054951 23.541336 19.164755 15.888303 23.768887 28.454714\n 15.539753 18.049202 16.23671 14.08383 25.33273 17.575668\n 49.566467 16.990675 21.738977 32.935143 16.125738 22.45393\n 20.776966 20.042227 22.898897 38.124043 30.607079 32.607468\n 20.919416 47.348038 14.524615 8.126455 19.581661 9.030508\n 26.462107 17.69918 20.546162 46.312218 39.689137 34.387108\n 22.11083 34.568977 24.873934 50.078335 14.5669775 20.525211\n 20.62971 23.202105 49.514477 23.12061 24.795782 20.319666\n 43.869396 17.110266 32.165016 34.75202 7.313497 20.309446\n 18.038298 12.008462 24.216425 47.90671 37.94349 20.759708\n 40.182804 18.249052 15.611586 26.39461 21.0571 20.421682\n 18.377089 17.338768 21.223648 22.653662 17.560051 32.635715\n 16.683764 13.004857 18.488163 20.659714 16.501846 20.648884\n 48.62411 15.977999 15.97522 18.581459 14.893438 32.871964\n 14.236945 43.612328 33.881115 19.073408 15.747335 9.4903965\n 10.153891 14.812717 18.655546 8.596755 22.666656 10.941623\n 20.534616 49.324417 22.710459 19.99658 31.663935 21.78586\n 30.9277 30.507492 15.054665 15.854853 48.532074 21.108742\n 15.687305 12.403721 49.90245 31.557863 11.709707 20.22495\n 26.214525 32.90807 22.90362 9.542897 24.487959 24.46598\n 22.509142 14.704502 27.895067 33.619015 14.888735 19.147383\n 26.40218 32.77208 29.293688 23.638102 10.448805 22.518728\n 21.47825 35.32415 23.002241 20.470022 18.918747 10.328174\n 22.244467 17.69918 20.918488 11.913417 42.572548 46.803394\n 14.652036 20.633188 23.285368 15.295161 20.861048 23.587011\n 32.94382 21.090906 24.898489 18.465925 31.454802 14.421506\n 15.421497 21.890705 23.64799 17.40471 26.111868 24.977922\n 27.56308 22.964123 18.823803 28.856464 14.080684 19.785515\n 17.007908 42.90537 26.354216 21.719929 23.784258 18.4141\n 17.923422 20.337881 22.936398 25.297531 17.572325 14.486319\n 20.739832 21.733093 11.1917715 18.290442 20.70475 20.929468\n 18.990923 8.7798395 21.141748 21.021317 15.49217 24.455221\n 31.499088 22.668139 14.862843 19.69585 24.746317 22.913176\n 48.144817 19.950285 30.148172 49.98047 16.743952 16.218952\n 9.891141 20.452726 17.06055 14.73646 17.539606 19.555712\n 30.26191 27.037518 18.43813 20.100842 24.147627 10.21256\n 25.064299 48.283043 20.977459 23.265625 20.141813 11.87677\n 17.84212 15.1286955 14.9789295 23.502743 16.092314 21.276255\n 26.55347 16.940031 23.485325 14.927286 20.90435 19.254526\n 24.397417 27.566774 23.607512 17.905067 22.675825 25.12203\n 15.141896 18.460642 23.440636 16.4928 23.372946 30.389936\n 15.330368 24.69199 17.316717 14.531138 10.496169 24.805672\n 15.659789 38.916733 20.403166 42.113743 8.544421 22.536352\n 15.654481 15.709977 17.263374 23.888586 21.690222 46.16276\n 15.304819 31.137545 25.326769 18.969254 26.29209 11.722559\n 40.65201 20.52522 17.135836 24.829275 15.565665 23.360205\n 8.280649 24.018639 19.57025 20.865868 23.611485 22.455328\n 17.646477 17.687094 14.59732 25.61237 13.333718 22.577513\n 20.657572 14.8804865 16.539358 23.276703 24.873934 22.52675\n 23.107155 31.871576 19.262531 19.536154 28.251024 23.817226\n 12.874959 22.59372 12.234834 10.024989 20.419611 10.369816\n 45.84478 24.873934 12.357825 16.367088 14.355771 28.338346\n 18.669233 20.334248 10.546778 21.30952 21.00914 20.669264\n 23.91886 25.009733 26.945326 13.288843 18.277857 20.95568\n 18.233625 23.807056 13.400126 23.875198 33.050533 27.785492\n 25.296518 19.071947 20.950756 11.507434 22.855497 15.573306\n 22.33747 20.807749 22.41908 17.212593 12.645366 35.121113\n 18.852188 48.823723 22.462465 24.267456 21.375692 19.38756\n 8.561088 20.726429 23.400837 21.41578 17.63176 25.232733\n 21.164701 26.444288 14.49171 49.559753 30.693232 23.20531\n 22.950115 16.84211 30.982431 16.259336 23.613512 20.93225\n 20.178421 22.782583 ]\n"
],
[
"# R squared error\nscore_1 = metrics.r2_score(Y_train, training_data_prediction)\n\n# Mean Absolute Error\nscore_2 = metrics.mean_absolute_error(Y_train, training_data_prediction)\n\nprint(\"R squared error : \", score_1)\nprint('Mean Absolute Error : ', score_2)",
"R squared error : 0.9733349094832763\nMean Absolute Error : 1.145314053261634\n"
]
],
[
[
"Visualizing the actual Prices and predicted prices",
"_____no_output_____"
]
],
[
[
"plt.scatter(Y_train, training_data_prediction)\nplt.xlabel(\"Actual Prices\")\nplt.ylabel(\"Predicted Prices\")\nplt.title(\"Actual Price vs Preicted Price\")\nplt.show()\n",
"_____no_output_____"
]
],
[
[
"Prediction on Test Data",
"_____no_output_____"
]
],
[
[
"# accuracy for prediction on test data\ntest_data_prediction = model.predict(X_test)",
"_____no_output_____"
],
[
"# R squared error\nscore_1 = metrics.r2_score(Y_test, test_data_prediction)\n\n# Mean Absolute Error\nscore_2 = metrics.mean_absolute_error(Y_test, test_data_prediction)\n\nprint(\"R squared error : \", score_1)\nprint('Mean Absolute Error : ', score_2)",
"R squared error : 0.9115937697657654\nMean Absolute Error : 1.9922956859364223\n"
]
]
] | [
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] | [
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
]
] |
d0fd721e0624d3b6a963c5f976d823ce8f0b8225 | 265,743 | ipynb | Jupyter Notebook | jupyter_notebooks/jlThermoModel.ipynb | linnetteteo/LiS-thermo-equilibria-manuscript | c8184a6906ef18fe54e7f4eb2741f1d7056b5d39 | [
"MIT"
] | null | null | null | jupyter_notebooks/jlThermoModel.ipynb | linnetteteo/LiS-thermo-equilibria-manuscript | c8184a6906ef18fe54e7f4eb2741f1d7056b5d39 | [
"MIT"
] | null | null | null | jupyter_notebooks/jlThermoModel.ipynb | linnetteteo/LiS-thermo-equilibria-manuscript | c8184a6906ef18fe54e7f4eb2741f1d7056b5d39 | [
"MIT"
] | null | null | null | 441.433555 | 62,286 | 0.935844 | [
[
[
"empty"
]
]
] | [
"empty"
] | [
[
"empty"
]
] |
d0fd809da59b44073983a2f1c2fa3aab4dcfff07 | 12,526 | ipynb | Jupyter Notebook | geekgap_webinars/notebooks/webinar_1/algorithms_data_structures.ipynb | geekgap-io/geekgap_webinars | 3a9a661ac8c376b2019d83c5bd8c17ab4a46535e | [
"MIT"
] | 1 | 2019-12-11T05:30:39.000Z | 2019-12-11T05:30:39.000Z | geekgap_webinars/notebooks/webinar_1/algorithms_data_structures.ipynb | geekgap-io/geekgap_webinars | 3a9a661ac8c376b2019d83c5bd8c17ab4a46535e | [
"MIT"
] | null | null | null | geekgap_webinars/notebooks/webinar_1/algorithms_data_structures.ipynb | geekgap-io/geekgap_webinars | 3a9a661ac8c376b2019d83c5bd8c17ab4a46535e | [
"MIT"
] | null | null | null | 19.633229 | 177 | 0.445074 | [
[
[
"# Time Complexity Examples",
"_____no_output_____"
]
],
[
[
"def logarithmic_problem(N):\n i = N\n while i > 1:\n # do something\n i = i // 2 # move on\n \n%time logarithmic_problem(10000)",
"CPU times: user 6 µs, sys: 1e+03 ns, total: 7 µs\nWall time: 9.06 µs\n"
],
[
"def linear_problem(N):\n i = N\n while i > 1:\n # do something\n i = i - 1 # move on\n \n%time linear_problem(10000)",
"CPU times: user 1.38 ms, sys: 734 µs, total: 2.11 ms\nWall time: 1.45 ms\n"
],
[
"def quadratic_problem(N):\n i = N\n while i > 1:\n j = N\n while j > 1:\n # do something\n j = j - 1 # move on\n i = i - 1\n \n%time quadratic_problem(10000)",
"CPU times: user 5.21 s, sys: 67.3 ms, total: 5.28 s\nWall time: 5.33 s\n"
]
],
[
[
"# Problem \nGiven an array(A) of numbers sorted in increasing order, implement a function that returns the index of a target(k) if found in A, and -1 otherwise.",
"_____no_output_____"
],
[
"### Brute-force solution: Linear Search",
"_____no_output_____"
]
],
[
[
"A = [5, 8, 8, 15, 16, 19, 30, 35, 40, 51]",
"_____no_output_____"
],
[
"def linear_search(A, k):\n for idx, element in enumerate(A):\n if element == k:\n return idx\n return -1",
"_____no_output_____"
],
[
"linear_search(A, 15)",
"_____no_output_____"
],
[
"linear_search(A, 100)",
"_____no_output_____"
]
],
[
[
"### Efficient solution: Binary Search",
"_____no_output_____"
]
],
[
[
"A = [5, 8, 8, 15, 16, 19, 30, 35, 40, 51]",
"_____no_output_____"
],
[
"def binary_search(A, k):\n left, right = 0, len(A)-1\n while left<=right:\n mid = (right - left)//2 + left\n if A[mid] < k:\n #look on the right\n left = mid+1\n elif A[mid] > k:\n #look on the left\n right = mid-1\n else:\n return mid\n \n return -1",
"_____no_output_____"
],
[
"binary_search(A, 15)",
"_____no_output_____"
],
[
"binary_search(A, 17)",
"_____no_output_____"
]
],
[
[
"### Binary Search common bugs:",
"_____no_output_____"
],
[
"#### BUG-1: one-off bug\nnot handling arrays of size=1",
"_____no_output_____"
]
],
[
[
"A = [5, 8, 8, 15, 16, 19, 30, 35, 40, 51]",
"_____no_output_____"
],
[
"def binary_search_bug1(A, k):\n left, right = 0, len(A)-1\n #HERE: < instead of <=\n while left<right: \n mid = (right - left)//2 + left\n if A[mid] < k:\n #look on the right\n left = mid+1\n elif A[mid] > k:\n #look on the left\n right = mid-1\n else:\n return mid\n return -1",
"_____no_output_____"
],
[
"binary_search_bug1(A, 35)",
"_____no_output_____"
],
[
"binary_search_bug1(A, 30)",
"_____no_output_____"
],
[
"binary_search_bug1(A, 15)",
"_____no_output_____"
],
[
"binary_search_bug1([15], 15)",
"_____no_output_____"
]
],
[
[
"#### BUG-2: integer overflow\nnot handling the case where summing two integers can return an integer bigger than what the memory can take",
"_____no_output_____"
]
],
[
[
"# because python3 only imposes limits \n# on float, we are going to illustrate\n# this issue using floats instead of ints\nimport sys\n\nright = sys.float_info.max\nleft = sys.float_info.max - 1000",
"_____no_output_____"
],
[
"mid = (right + left) // 2\nmid",
"_____no_output_____"
],
[
"mid = (right - left)//2 + left\nmid",
"_____no_output_____"
]
],
[
[
"## Problem variant1:\n#### Search a sorted array for first occurrence of target(k)",
"_____no_output_____"
],
[
"Given an array(A) of numbers sorted in increasing order, implement a function that returns the index of the first occurence of a target(k) if found in A, and -1 otherwise.",
"_____no_output_____"
]
],
[
[
"A = [5, 8, 8, 8, 8, 19, 30, 35, 40, 51]",
"_____no_output_____"
],
[
"def first_occurence_search(A, k):\n left, right, res = 0, len(A)-1, -1\n while left<=right:\n mid = (right - left)//2 + left\n if A[mid] < k:\n #look on the right\n left = mid+1\n elif A[mid] > k:\n #look on the left\n right = mid-1\n else:\n # update res\n res = mid\n # keep looking on the left\n right = mid-1\n \n return res",
"_____no_output_____"
],
[
"binary_search(A, 8)",
"_____no_output_____"
],
[
"first_occurence_search(A, 8)",
"_____no_output_____"
]
],
[
[
"## Problem variant2:\n#### Search a sorted array for entry equal to its index",
"_____no_output_____"
],
[
"Given a sorted array(A) of distinct integers, implement a function that returns the index i if A[i] = i, and -1 otherwise.",
"_____no_output_____"
]
],
[
[
"A = [-3, 0, 2, 5, 7, 9, 18, 35, 40, 51]",
"_____no_output_____"
],
[
"def search_entry_equal_to_its_index(A):\n left, right = 0, len(A)-1\n while left<=right:\n mid = (right - left)//2 + left\n difference = A[mid] - mid\n if difference < 0:\n #look on the right\n left = mid+1\n elif difference > 0:\n #look on the left\n right = mid-1\n else:\n return mid\n \n return -1",
"_____no_output_____"
],
[
"search_entry_equal_to_its_index(A)",
"_____no_output_____"
]
]
] | [
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] | [
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code",
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code",
"code"
]
] |
d0fd946913f46f31567b6d1e1b8ef3f37d803a8f | 17,664 | ipynb | Jupyter Notebook | notebooks/workshops/04Pathlib.ipynb | emmamcbryde/AuTuMN-1 | b1e7de15ac6ef6bed95a80efab17f0780ec9ff6f | [
"BSD-2-Clause-FreeBSD"
] | null | null | null | notebooks/workshops/04Pathlib.ipynb | emmamcbryde/AuTuMN-1 | b1e7de15ac6ef6bed95a80efab17f0780ec9ff6f | [
"BSD-2-Clause-FreeBSD"
] | null | null | null | notebooks/workshops/04Pathlib.ipynb | emmamcbryde/AuTuMN-1 | b1e7de15ac6ef6bed95a80efab17f0780ec9ff6f | [
"BSD-2-Clause-FreeBSD"
] | null | null | null | 20.256881 | 177 | 0.515625 | [
[
[
"# Pathlib\n\n## Object oriented Pythonic paths\naka\n\"The Right Way to do Paths\"\n\nhttps://docs.python.org/3/library/pathlib.html",
"_____no_output_____"
]
],
[
[
"# Pathlib is a standard Python library\n# You will usually want to import Path and/or PurePath\n\nimport pathlib\nfrom pathlib import Path, PurePath",
"_____no_output_____"
],
[
"# Let's do a few more imports for later\nimport os\nimport shutil",
"_____no_output_____"
]
],
[
[
"## What _are_ paths, really?\n\nDepends on the context. \n\nIs this a path?\nIt's certainly a URL...\nhttps://autumn-data.com/runs/sm_sir/\n\nHow about just this bit? \nsm_sir/malaysia/1634732134/ec182f\n\nPaths are abstract representations of a nested, tree-like structure \nSuch things include filesystems, bits of web addresses, certain representations of S3 etc etc... \n\nPathlib deals with all these things. It also deals with the bits that are specifically about filesystems, but makes a distinction between the two (for good reasons)",
"_____no_output_____"
],
[
"## Filesystems\n### Pathlib vs os.* vs string manipulation",
"_____no_output_____"
]
],
[
[
"cwd = Path()",
"_____no_output_____"
],
[
"cwd",
"_____no_output_____"
],
[
"# Listdir\nos.listdir(cwd)",
"_____no_output_____"
],
[
"cwd.glob('*')",
"_____no_output_____"
],
[
"list(cwd.glob('*'))",
"_____no_output_____"
],
[
"# Making glob a little easier to work with; the autumn display module contains\n# some hooks for Jupyter notebooks\n\nfrom autumn.tools.utils import display",
"_____no_output_____"
],
[
"cwd.glob('*')",
"_____no_output_____"
],
[
"cd ..",
"_____no_output_____"
],
[
"cwd.glob('*')",
"_____no_output_____"
],
[
"cd workshops/",
"_____no_output_____"
],
[
"cwd = cwd.absolute()\ncwd",
"_____no_output_____"
],
[
"# This will work, but is a bad idea in most cases\n\n(cwd / '..')",
"_____no_output_____"
],
[
"# Better\n\ncwd.parent",
"_____no_output_____"
],
[
"# Like str.split, but better and safer...\ncwd.parts",
"_____no_output_____"
],
[
"some_file = Path(\"path/to/file.txt\")\ntgz_file = Path(\"file.tar.gz\")",
"_____no_output_____"
],
[
"# Strings are the worst...",
"_____no_output_____"
],
[
"str(some_file).split('.')[0], str(some_file).split('.')[-1]",
"_____no_output_____"
],
[
"just_filename = str(some_file).split('/')[-1]\njust_filename.split('.')[0], just_filename.split('.')[-1]",
"_____no_output_____"
],
[
"str(tgz_file).split('.')[0], str(tgz_file).split('.')[-1]",
"_____no_output_____"
],
[
"# os.path is ... okish...",
"_____no_output_____"
],
[
"os.path.splitext(some_file)",
"_____no_output_____"
],
[
"os.path.splitext(tgz_file)",
"_____no_output_____"
],
[
"# Pathlib\n\nsome_file.stem, some_file.suffix",
"_____no_output_____"
],
[
"tgz_file.stem, tgz_file.suffix",
"_____no_output_____"
],
[
"tgz_file.suffixes",
"_____no_output_____"
]
],
[
[
"## A note on being a good programmer...\n\nThe string examples above tell us something - we all make assumptions, that turn into heuristics - they work well, until they don't\n\n### It's not about being a \"rockstar\"... more like a buddhist monk with a management job\n\nLearn how to delegate - use system libraries! Someone else has thought about this a lot more than you ever will (or will ever want to)\n### ...although having a bit of London cabbie helps - read the documentation! Drive the roads! (well, use the library until you don't need to look up the documentation...)",
"_____no_output_____"
],
[
"## File handling ergonomics...",
"_____no_output_____"
]
],
[
[
"test_path = cwd / \"test_files\"",
"_____no_output_____"
],
[
"test_path.mkdir(exist_ok=True)",
"_____no_output_____"
],
[
"for i in range(5):\n (test_path / f\"file{i}.txt\").write_text(f\"Some example file contents for file {i}\")",
"_____no_output_____"
],
[
"contents_map = {f: f.read_text() for f in cwd.glob(\"*/*.txt\")}",
"_____no_output_____"
],
[
"contents_map",
"_____no_output_____"
],
[
"# Bonus Python 3.8 syntax - the Walrus operator :=\n{f: contents for f in cwd.glob(\"*/*.txt\") if \"3\" in (contents := f.read_text())}",
"_____no_output_____"
],
[
"# We also have access to real properties of the filesystem - like file size etc\n\nf = test_path / \"file1.txt\"\nf.lstat()",
"_____no_output_____"
],
[
"test_path.rmdir()",
"_____no_output_____"
],
[
"os.rmdir(test_path)",
"_____no_output_____"
],
[
"# Still need to use shutil - same as it ever was\n# https://docs.python.org/3/library/shutil.html\n\nshutil.rmtree(test_path)",
"_____no_output_____"
],
[
"test_path",
"_____no_output_____"
],
[
"test_path.glob('*')",
"_____no_output_____"
],
[
"test_path.exists()",
"_____no_output_____"
]
],
[
[
"## Writing functions, calling functions",
"_____no_output_____"
]
],
[
[
"def do_something1(path_to_file):\n return os.path.exists(path_to_file)",
"_____no_output_____"
],
[
"def do_something2(path_to_file):\n return path_to_file.exists()",
"_____no_output_____"
],
[
"do_something1(test_path)",
"_____no_output_____"
],
[
"do_something2(test_path)",
"_____no_output_____"
],
[
"a_file = \"this is not a file\"",
"_____no_output_____"
],
[
"do_something1(a_file)",
"_____no_output_____"
],
[
"do_something2(a_file)",
"_____no_output_____"
]
],
[
[
"### Use type annotations! (You should be doing this anyway)",
"_____no_output_____"
]
],
[
[
"def do_something3(path_to_file: Path) -> bool:\n # Users know what this function expects\n return path_to_file.exists()",
"_____no_output_____"
],
[
"# This will fail - but it's the user's fault now (in the nicest possible way...)\n\ndo_something3(a_file)",
"_____no_output_____"
],
[
"do_something3(Path(a_file))",
"_____no_output_____"
],
[
"from typing import Union",
"_____no_output_____"
],
[
"def do_something4(path_to_file: Union[Path, str]) -> bool:\n # Now we handle both cases\n path_to_file = Path(path_to_file) if isinstance(path_to_file, str) else path_to_file\n \n return path_to_file.exists()",
"_____no_output_____"
],
[
"do_something4(a_file), do_something4(Path(a_file)), ",
"_____no_output_____"
],
[
"# Bonus Python 3.10 version...\n\nPathOrStr = Path|str\n\ndef do_something5(path_to_file: PathOrStr) -> bool:\n # Now we handle both cases\n path_to_file = Path(path_to_file) if isinstance(path_to_file, str) else path_to_file\n return path_to_file.exists()",
"_____no_output_____"
],
[
"do_something5",
"_____no_output_____"
]
],
[
[
"## PurePaths\n\nPurePaths are 'pure' in that they are \n \n\na) Abstract representations unencumbered by the weight of the real world... \nb) Functionally pure (ie they can't produce side effects) ",
"_____no_output_____"
]
],
[
[
"from pathlib import PurePath, PurePosixPath, PureWindowsPath",
"_____no_output_____"
],
[
"# Use PurePath directly if you want to work on abstract paths of the type of system you're working on...\n\npure_test = PurePath(test_path)\npure_test",
"_____no_output_____"
],
[
"pure_test.mkdir()",
"_____no_output_____"
],
[
"# Specify the path type if you have a particular filesystem in mind...\n\nwin_path = PureWindowsPath(test_path)\nwin_path",
"_____no_output_____"
],
[
"str(win_path)",
"_____no_output_____"
],
[
"str(\"C:\" / win_path / \"MicroSoft Style Folder Name (95)\")",
"_____no_output_____"
],
[
"s3_bucket = PurePosixPath(\"autumn-data\")",
"_____no_output_____"
],
[
"import s3fs",
"_____no_output_____"
],
[
"fs = s3fs.S3FileSystem()",
"_____no_output_____"
],
[
"fs.ls(s3_bucket)",
"_____no_output_____"
],
[
"fs.ls(s3_bucket / \"sm_sir\" / \"malaysia\")",
"_____no_output_____"
],
[
"# If you were just using 'Path' on a Windows system - \n# you'd have a WindowsPath object, and this would happen...\n# That's why we use PurePosixPath - because the system we're talking to is Posix-like\n\nfs.ls(PureWindowsPath(s3_bucket) / \"sm_sir\")",
"_____no_output_____"
],
[
"# Bonus round - glob is awesome\n\nfs.glob(str(s3_bucket / \"*\" / \"malaysia\"))",
"_____no_output_____"
]
]
] | [
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] | [
[
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
d0fd975a8e7f236f1a987c6b4da2ea8f769f0113 | 105,244 | ipynb | Jupyter Notebook | .ipynb_checkpoints/Logistic Regression-checkpoint.ipynb | AshishSinha5/mlAlgos | 003f691e92c7978626de0b2cb3533a164434159d | [
"Apache-2.0"
] | null | null | null | .ipynb_checkpoints/Logistic Regression-checkpoint.ipynb | AshishSinha5/mlAlgos | 003f691e92c7978626de0b2cb3533a164434159d | [
"Apache-2.0"
] | null | null | null | .ipynb_checkpoints/Logistic Regression-checkpoint.ipynb | AshishSinha5/mlAlgos | 003f691e92c7978626de0b2cb3533a164434159d | [
"Apache-2.0"
] | 1 | 2020-01-25T17:41:15.000Z | 2020-01-25T17:41:15.000Z | 229.790393 | 34,404 | 0.900184 | [
[
[
"import numpy as np\nimport matplotlib.pyplot as plt\nimport pandas as pd",
"_____no_output_____"
],
[
"%matplotlib inline",
"_____no_output_____"
],
[
"#defining sigmoid function\ndef sigmoid(x):\n return 1/(1 + np.exp(-x))",
"_____no_output_____"
],
[
"#ploting sigmoid fuction for the values -7,7\nz = np.arange(-7,7,0.1)\nphi_z = sigmoid(z)\nplt.plot(z,phi_z)\nplt.axvline(0.0, color ='k')\nplt.xlabel(\"z\")\nplt.ylabel(\"phi(z)\")\nplt.ylim(-0.1,1.1)\nplt.title(\"Logistic Function\")\nplt.tight_layout()\nplt.show()",
"_____no_output_____"
],
[
"#defining cost function for the logistic linear regression\ndef cost_0(x):\n return -np.log(1 - sigmoid(x))\n\ndef cost_1(x):\n return -np.log(sigmoid(x))\n\nz = np.arange(-10,10,0.1)\nphi_z = sigmoid(z)\nc0 = [cost_0(x) for x in z]\nc1 = [cost_1(x) for x in z]\nplt.plot(phi_z,c0,linestyle='--', linewidth = 2, label = 'c0')\nplt.plot(phi_z,c1,linestyle=':', linewidth = 2, label = 'c1')\nplt.xlabel('Phi(z)')\nplt.ylabel('Cost')\nplt.tight_layout()\nplt.xlim([0,1])\nplt.ylim(0.0,5.1)\nplt.legend(loc = 'best')\nplt.show()",
"_____no_output_____"
]
],
[
[
"We are penalizing wrong classification with higher cost",
"_____no_output_____"
]
],
[
[
"# Defining logistic Regrssion Class\nclass logisticRegression():\n \n def __init__(self, eta = 0.1, n_iter = 50, random_state = 1):\n self.eta = eta\n self.n_iter = n_iter\n self.random_state = random_state\n \n def fit(self,X,y):\n #shape of X = [n_examples, n_features]\n #shape of y = [n_examples]\n \n rgen = np.random.RandomState(self.random_state)\n self._w = rgen.normal(loc = 0.0, scale = 0.01, size = 1 + X.shape[1])\n self._cost = []\n #updating weights and calcuationg sum of squares error which is our cost function\n #the main idea is to minimize cost function by updating by taking the step in the opposite direction of the cost gradient\n for i in range(self.n_iter):\n net_input = self.net_input(X)\n output = self.activation(net_input)\n errors = y - output\n self._w[1:] += self.eta*np.dot(X.T, errors)\n self._w[0] += self.eta*errors.sum()\n cost = (errors**2).sum()/2.0\n self._cost.append(cost)\n \n return self\n\n def net_input(self, X):\n return np.dot(X, self._w[1:]) + self._w[0]\n \n def activation(self, X):\n return 1./(1. + np.exp(-X)) # Sigmoid activation function\n #return X #linear activation function\n \n def predict(self, X):\n return np.where(self.activation(self.net_input(X))>=0.5,1,-1)",
"_____no_output_____"
],
[
"#Importing datasets\nfrom sklearn import datasets\niris = datasets.load_iris()\nX = iris.data[:, [2,3]]\ny = iris.target\n#Splitting the dataset in test ans train dataset to test aor model's performance on unseen data\nfrom sklearn.model_selection import train_test_split\nX_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.3 , random_state = 1, stratify = y)\n#stratify function ensures all the classes have equal proportion of representaion in both test and train sets",
"_____no_output_____"
],
[
"#helper function for plotting decision\nfrom matplotlib.colors import ListedColormap\ndef plotDecisionRegion(X, y, classifier, test_idx = None, resolution = 0.02):\n markers = ('s','x','o','^','v')\n colors = ('red', 'blue', 'lightgreen', 'gray', 'cyan')\n cmap = ListedColormap(colors[:len(np.unique(y))])\n x1_min, x1_max = X[:, 0].min() - 1,X[:,0].max() + 1\n x2_min, x2_max = X[:, 1].min() - 1,X[:,1].max() + 1\n xx1, xx2 = np.meshgrid(np.arange(x1_min,x1_max,resolution),\n np.arange(x2_min,x2_max, resolution))\n #xx1, xx2 are the coordinates of x and y respectively, we pair each value of the two corresponding matrices and get a grid\n Z = classifier.predict(np.array([xx1.ravel(), xx2.ravel()]).T)\n Z = Z.reshape(xx1.shape)\n plt.contourf(xx1,xx2, Z, alpha = 0.3, cmap = cmap)\n plt.xlim(xx1.min(),xx1.max())\n plt.ylim(xx2.min(),xx2.max())\n \n for idx, c1 in enumerate(np.unique(y)):\n plt.scatter(x =X[y==c1,0],\n y = X[y==c1,1],\n alpha =0.8,\n c = colors[idx],\n marker = markers[idx],\n label = c1, edgecolor='black')\n \n if test_idx:\n X_test, y_test = X[test_idx, :], y[test_idx]\n plt.scatter(X_test[:,0], X_test[:,1], c= '', edgecolors='black',\n alpha=1.0, linewidths=1, marker='o', s=100, label='test set')",
"_____no_output_____"
],
[
"# Consodering only iris satosa ans iris versicolor\nX_train_subset = X_train[(y_train == 0) | (y_train == 1)]\ny_train_subset = y_train[(y_train == 0) | (y_train == 1)]\nlr = logisticRegression(eta = 0.05, n_iter = 1000, random_state=1)\nlr.fit(X_train_subset, y_train_subset)\nplotDecisionRegion(X= X_train_subset, y = y_train_subset, classifier=lr)\nplt.xlabel('Petal Length')\nplt.ylabel('Sepal Length')\nplt.tight_layout()\nplt.legend(loc = 'best')\nplt.show()",
"C:\\Users\\ashish\\AppData\\Roaming\\Python\\Python35\\site-packages\\numpy\\ma\\core.py:6512: MaskedArrayFutureWarning: In the future the default for ma.minimum.reduce will be axis=0, not the current None, to match np.minimum.reduce. Explicitly pass 0 or None to silence this warning.\n return self.reduce(a)\nC:\\Users\\ashish\\AppData\\Roaming\\Python\\Python35\\site-packages\\numpy\\ma\\core.py:6512: MaskedArrayFutureWarning: In the future the default for ma.maximum.reduce will be axis=0, not the current None, to match np.maximum.reduce. Explicitly pass 0 or None to silence this warning.\n return self.reduce(a)\n"
],
[
"from sklearn.preprocessing import StandardScaler\nsc = StandardScaler()\nsc.fit(X_train) # fit method will estimate the parameters mean and standars deviation of the sample given\nX_train_std = sc.transform(X_train)\nX_test_std = sc.transform(X_test)",
"_____no_output_____"
],
[
"#training logistic regression model using sklearn\nfrom sklearn.linear_model import LogisticRegression",
"_____no_output_____"
],
[
"lr = LogisticRegression(C = 100, random_state=1, solver='lbfgs', multi_class='ovr')\nlr.fit(X_train_std, y_train)\nX_combined_std = np.vstack((X_train_std, X_test_std))\ny_combined = np.hstack((y_train, y_test))\nplotDecisionRegion(X_combined_std, y_combined, classifier=lr, test_idx=range(105,150))\nplt.ylabel('Petal length (standardized)')\nplt.xlabel('Petal width (standardized)')\nplt.legend(loc='upper left')\nplt.tight_layout()\nplt.show()",
"C:\\Users\\ashish\\AppData\\Roaming\\Python\\Python35\\site-packages\\numpy\\ma\\core.py:6512: MaskedArrayFutureWarning: In the future the default for ma.minimum.reduce will be axis=0, not the current None, to match np.minimum.reduce. Explicitly pass 0 or None to silence this warning.\n return self.reduce(a)\nC:\\Users\\ashish\\AppData\\Roaming\\Python\\Python35\\site-packages\\numpy\\ma\\core.py:6512: MaskedArrayFutureWarning: In the future the default for ma.maximum.reduce will be axis=0, not the current None, to match np.maximum.reduce. Explicitly pass 0 or None to silence this warning.\n return self.reduce(a)\n"
],
[
"print(lr.predict_proba(X_test_std[:10,:]).argmax(axis = 1))\nprint(y_test[:10])",
"[2 0 0 1 1 1 2 1 2 0]\n[2 0 0 2 1 1 2 1 2 0]\n"
],
[
"#Adding reglarization to cost function\nweights, params = [],[]\nfor c in range(-5,5):\n lr = LogisticRegression(C = 10.**c, solver='lbfgs', random_state=1, multi_class='ovr')\n lr.fit(X_train_std, y_train)\n weights.append(lr.coef_[1])\n params.append(10.**c)\nweights = np.array(weights)",
"_____no_output_____"
],
[
"weights",
"_____no_output_____"
],
[
"plt.plot(params, weights[:,0],linestyle = ':', label = 'petal length')\nplt.plot(params, weights[:,1], linestyle = '-', label = 'petal width')\nplt.xscale('log')\nplt.legend(loc = 'upper left')",
"_____no_output_____"
]
]
] | [
"code",
"markdown",
"code"
] | [
[
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
d0fd9ebb2475f12a94c35506f0a6a570b3ae6cf8 | 1,267 | ipynb | Jupyter Notebook | opening_files.ipynb | aditya270520/100Daysofpython | a70d72bb8663947ccad1cc0cb6252f3d3624fb44 | [
"MIT"
] | 4 | 2020-12-12T11:49:02.000Z | 2021-02-09T14:10:17.000Z | opening_files.ipynb | aditya270520/100Daysofpython | a70d72bb8663947ccad1cc0cb6252f3d3624fb44 | [
"MIT"
] | null | null | null | opening_files.ipynb | aditya270520/100Daysofpython | a70d72bb8663947ccad1cc0cb6252f3d3624fb44 | [
"MIT"
] | null | null | null | 26.395833 | 238 | 0.480663 | [
[
[
"<a href=\"https://colab.research.google.com/github/aditya270520/100Daysofpython/blob/main/opening_files.ipynb\" target=\"_parent\"><img src=\"https://colab.research.google.com/assets/colab-badge.svg\" alt=\"Open In Colab\"/></a>",
"_____no_output_____"
]
],
[
[
"import glob\npython_files = glob.glob('*.py')\nfor file_name in sorted(python_files):\n print (' ------' + file_name)\n\n with open(file_name) as f:\n for line in f:\n print (' ' + line.rstrip())\n\n print()",
"_____no_output_____"
]
]
] | [
"markdown",
"code"
] | [
[
"markdown"
],
[
"code"
]
] |
d0fdae9e69409f6e11cf87a0147af83c27d79c3e | 67,357 | ipynb | Jupyter Notebook | notebooks/convert_met_obs.ipynb | NorDataNet/TrainingMaterial | 71de752803d2fe0e115d53441893106293f87a5c | [
"CC-BY-4.0"
] | null | null | null | notebooks/convert_met_obs.ipynb | NorDataNet/TrainingMaterial | 71de752803d2fe0e115d53441893106293f87a5c | [
"CC-BY-4.0"
] | null | null | null | notebooks/convert_met_obs.ipynb | NorDataNet/TrainingMaterial | 71de752803d2fe0e115d53441893106293f87a5c | [
"CC-BY-4.0"
] | null | null | null | 117.551483 | 46,448 | 0.767433 | [
[
[
"# Convert data extracted from Frost",
"_____no_output_____"
],
[
"## Purpose\nConvert data that can be extracted from MET observation storage through the Frost API https://frost.met.no/index.html. The extraction process is not documented.",
"_____no_output_____"
],
[
"Import the necessary packages.",
"_____no_output_____"
]
],
[
[
"import pandas as pd\nimport numpy as np\nimport matplotlib.pyplot as plt",
"_____no_output_____"
]
],
[
[
"Read the actual data file that has been extracted from the Frost API described above.",
"_____no_output_____"
]
],
[
[
"mydf = pd.read_csv('radflux-bjørnøya-frost.txt', header=0,mangle_dupe_cols=True, parse_dates=['referenceTime'], index_col='referenceTime', na_values=['-'])",
"_____no_output_____"
]
],
[
[
"Dump an overview of the data that has been read.",
"_____no_output_____"
]
],
[
[
"mydf.keys()",
"_____no_output_____"
],
[
"mydf",
"_____no_output_____"
]
],
[
[
"Plot the dataset.",
"_____no_output_____"
]
],
[
[
"fig = plt.figure(figsize=(20,10))",
"_____no_output_____"
],
[
"mydf.plot.line(y=['mean(surface_downwelling_shortwave_flux_in_air pt1m)(W/m2)','mean(surface_downwelling_longwave_flux_in_air pt1m)(W/m2)'])",
"_____no_output_____"
]
]
] | [
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] | [
[
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
]
] |
d0fdd4152e5c68291494ef86ae0f1ee1c6bc8234 | 20,921 | ipynb | Jupyter Notebook | random_forest + final output.ipynb | Aindstorm/Deep-behavioral-analysis | e67f9e520d405100e34a52a056b3ccbda592f3a0 | [
"MIT"
] | null | null | null | random_forest + final output.ipynb | Aindstorm/Deep-behavioral-analysis | e67f9e520d405100e34a52a056b3ccbda592f3a0 | [
"MIT"
] | null | null | null | random_forest + final output.ipynb | Aindstorm/Deep-behavioral-analysis | e67f9e520d405100e34a52a056b3ccbda592f3a0 | [
"MIT"
] | null | null | null | 27.064683 | 149 | 0.488026 | [
[
[
"Ensembling different models",
"_____no_output_____"
]
],
[
[
"from google.cloud import storage\nfrom io import BytesIO\n\nclient = storage.Client()\nstorage_client = storage.Client(project = 'irkml1')\nbucket = storage_client.get_bucket(\"aindstorm_bucket\")\nblob1 = storage.blob.Blob(\"train_3lags_semibalanced.csv\",bucket)\ncontent1 = blob1.download_as_string()",
"_____no_output_____"
],
[
"blob2 = storage.blob.Blob(\"train_3lags_v1.csv\",bucket)\ncontent2 = blob2.download_as_string()",
"_____no_output_____"
],
[
"import sys\nimport pandas as pd\nimport numpy as np\nimport scipy.sparse as sparse\nfrom scipy.sparse.linalg import spsolve\nimport random\nimport os\nimport scipy.stats as ss\nimport scipy\nfrom sklearn.preprocessing import MinMaxScaler\nfrom sklearn.model_selection import train_test_split\nfrom sklearn import preprocessing\nfrom sklearn.metrics import accuracy_score\nfrom catboost import CatBoostClassifier, Pool, sum_models, to_classifier\nfrom sklearn.model_selection import KFold\nfrom sklearn.utils.class_weight import compute_class_weight\n",
"_____no_output_____"
],
[
"df2 = pd.read_csv(BytesIO(content1), low_memory=False)",
"_____no_output_____"
],
[
"df = pd.read_csv(BytesIO(content2), low_memory=False)",
"_____no_output_____"
],
[
"df2 = df2.sample(frac=1).reset_index(drop=True)",
"_____no_output_____"
],
[
"df2.columns",
"_____no_output_____"
],
[
"df2 = df2[['service_title', 'service_title_1', 'requester_type', 'gender', 'age']]",
"_____no_output_____"
],
[
"df2.dropna(inplace=True)",
"_____no_output_____"
],
[
"y = df2['service_title']\nX = df2.drop(['service_title'], axis=1)",
"_____no_output_____"
],
[
"le1 = preprocessing.LabelEncoder()\nle2 = preprocessing.LabelEncoder()\nle3 = preprocessing.LabelEncoder()\nle4 = preprocessing.LabelEncoder()",
"_____no_output_____"
],
[
"y = le1.fit_transform(y)\nX['service_title_1'] = le2.fit_transform(X['service_title_1'])\nX['requester_type'] = le3.fit_transform(X['requester_type'])\nX['gender'] = le4.fit_transform(X['gender'])",
"_____no_output_____"
],
[
"le = preprocessing.LabelEncoder()\nfor col in categorical:\n X[col] = le.fit_transform(X[col])",
"_____no_output_____"
],
[
"y = le1.fit_transform(y)",
"_____no_output_____"
],
[
"from sklearn.ensemble import RandomForestClassifier",
"_____no_output_____"
],
[
"X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.15)",
"_____no_output_____"
],
[
"clf = RandomForestClassifier(n_estimators=100)",
"_____no_output_____"
],
[
"clf.fit(X_train, y_train)",
"_____no_output_____"
],
[
"y_pred = clf.predict(X_test)",
"_____no_output_____"
],
[
"accuracy_score(y_test, y_pred)",
"_____no_output_____"
],
[
"sab = pd.read_csv(\"sample_submission_ld.csv\")\n\nsab_pred = sab[['requester']]\n\nsab_pred = pd.merge(sab_pred, df, how='left', on='requester')\n\nsab_pred.drop(['service_3',\n'service_title_3',\n'mfc_3',\n'internal_status_3',\n'external_status_3',\n'order_type_3',\n'department_id_3',\n'custom_service_id_3',\n'service_level_3',\n'is_subdep_3',\n'is_csid_3',\n'proc_time_3',\n'dayofweek_3',\n'day_part_3',\n'person_3',\n'sole_3',\n'legal_3',\n'auto_ping_queue_3',\n'win_count_3', 'year_3', 'month_3', 'week_3'], axis=1 , inplace=True)\n\nsab_pred.drop(['requester'], axis=1, inplace=True)",
"_____no_output_____"
],
[
"sab_pred = sab_pred[['service_title', 'requester_type', 'gender', 'age']]",
"_____no_output_____"
],
[
"sab_pred.fillna(sab_pred.mode().iloc[0], inplace=True)",
"_____no_output_____"
],
[
"sab_pred[['service_title', 'requester_type', 'gender']] = sab_pred[['service_title', 'requester_type', 'gender']].astype('int64')",
"_____no_output_____"
],
[
"sab_pred['service_title'].loc[sab_pred['service_title'] == 151] = 98\nsab_pred['service_title'].loc[sab_pred['service_title'] == 408] = 98\nsab_pred['service_title'].loc[sab_pred['service_title'] == 509] = 98\nsab_pred['service_title'].loc[sab_pred['service_title'] == 945] = 98",
"_____no_output_____"
],
[
"sab_pred['service_title_1'] = le2.transform(sab_pred['service_title'])\nsab_pred['requester_type'] = le3.transform(sab_pred['requester_type'])\nsab_pred['gender'] = le4.transform(sab_pred['gender'])",
"_____no_output_____"
],
[
"sab_pred.drop(['service_title'], axis=1, inplace=True)",
"_____no_output_____"
],
[
"sab_pred = sab_pred[['service_title_1', 'requester_type', 'gender', 'age']]",
"_____no_output_____"
],
[
"sab_pred",
"_____no_output_____"
],
[
"y_pred = clf.predict(sab_pred)",
"_____no_output_____"
],
[
"sab['service_title'] = le1.inverse_transform(y_pred)",
"_____no_output_____"
],
[
"sab.to_csv('rf_sub_0.csv', index=False)",
"_____no_output_____"
],
[
"sab_nn = pd.read_csv('nn_5fold_2models_0.csv')",
"_____no_output_____"
],
[
"sab_xgb = pd.read_csv('xgb_sub_0.csv')",
"_____no_output_____"
],
[
"sab_rf = pd.read_csv('rf_sub_0.csv')",
"_____no_output_____"
],
[
"idx = sab_pred.loc[sab_pred['order_count'] == 1].index",
"_____no_output_____"
],
[
"sab['service_title'] = sab_nn['service_title']",
"_____no_output_____"
],
[
"sab_all = sab[:]",
"_____no_output_____"
],
[
"sab_all['nn'] = sab_nn['service_title']\nsab_all['xgb'] = sab_xgb['service_title']\nsab_all['rf'] = sab_rf['service_title']",
"/home/jupyter/.pyenv/versions/3.7.4/lib/python3.7/site-packages/ipykernel_launcher.py:1: SettingWithCopyWarning: \nA value is trying to be set on a copy of a slice from a DataFrame.\nTry using .loc[row_indexer,col_indexer] = value instead\n\nSee the caveats in the documentation: https://pandas.pydata.org/pandas-docs/stable/user_guide/indexing.html#returning-a-view-versus-a-copy\n \"\"\"Entry point for launching an IPython kernel.\n/home/jupyter/.pyenv/versions/3.7.4/lib/python3.7/site-packages/ipykernel_launcher.py:2: SettingWithCopyWarning: \nA value is trying to be set on a copy of a slice from a DataFrame.\nTry using .loc[row_indexer,col_indexer] = value instead\n\nSee the caveats in the documentation: https://pandas.pydata.org/pandas-docs/stable/user_guide/indexing.html#returning-a-view-versus-a-copy\n \n/home/jupyter/.pyenv/versions/3.7.4/lib/python3.7/site-packages/ipykernel_launcher.py:3: SettingWithCopyWarning: \nA value is trying to be set on a copy of a slice from a DataFrame.\nTry using .loc[row_indexer,col_indexer] = value instead\n\nSee the caveats in the documentation: https://pandas.pydata.org/pandas-docs/stable/user_guide/indexing.html#returning-a-view-versus-a-copy\n This is separate from the ipykernel package so we can avoid doing imports until\n"
],
[
"def voting(x):\n d = {}\n x1 = x['nn']\n x2 = x['xgb']\n x3 = x['rf']\n if x2 == x3:\n return x2\n return x1",
"_____no_output_____"
],
[
"sab_all['service_title'].iloc[idx] = sab_all.iloc[idx].apply(voting, axis=1)",
"/home/jupyter/.pyenv/versions/3.7.4/lib/python3.7/site-packages/ipykernel_launcher.py:1: SettingWithCopyWarning: \nA value is trying to be set on a copy of a slice from a DataFrame\n\nSee the caveats in the documentation: https://pandas.pydata.org/pandas-docs/stable/user_guide/indexing.html#returning-a-view-versus-a-copy\n \"\"\"Entry point for launching an IPython kernel.\n"
],
[
"sab_all = sab_all[['requester', 'service_title']]",
"_____no_output_____"
],
[
"sab_all['service_title'].value_counts(normalize=True)",
"_____no_output_____"
],
[
"sab_nn['service_title'].value_counts(normalize=True)",
"_____no_output_____"
],
[
"sab_all.to_csv('vot_sub_0.csv', index=False)",
"_____no_output_____"
]
]
] | [
"markdown",
"code"
] | [
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
d0fde689f24118c1d5cb388b8bfba0a143365646 | 35,474 | ipynb | Jupyter Notebook | analysis/covariance.ipynb | AdrienRoyer/smartnoise-samples | d2359d53cc584aaf0d71c1120c039b390a916666 | [
"MIT"
] | 50 | 2020-04-09T16:17:44.000Z | 2020-10-19T02:02:37.000Z | analysis/covariance.ipynb | AdrienRoyer/smartnoise-samples | d2359d53cc584aaf0d71c1120c039b390a916666 | [
"MIT"
] | 22 | 2020-03-10T05:31:53.000Z | 2020-10-27T19:00:55.000Z | analysis/covariance.ipynb | AdrienRoyer/smartnoise-samples | d2359d53cc584aaf0d71c1120c039b390a916666 | [
"MIT"
] | 24 | 2021-03-11T04:55:14.000Z | 2022-02-22T21:23:17.000Z | 108.815951 | 22,044 | 0.807634 | [
[
[
"# Differentially Private Covariance\n\nSmartNoise offers three different functionalities within its `covariance` function:\n\n1. Covariance between two vectors\n2. Covariance matrix of a matrix\n3. Cross-covariance matrix of a pair of matrices, where element $(i,j)$ of the returned matrix is the covariance of column $i$ of the left matrix and column $j$ of the right matrix.",
"_____no_output_____"
]
],
[
[
"# load libraries\nimport os\nimport opendp.smartnoise.core as sn\nimport numpy as np\nimport pandas as pd\nimport seaborn as sns \nimport matplotlib.pyplot as plt\n\n# establish data information\ndata_path = os.path.join('.', 'data', 'PUMS_california_demographics_1000', 'data.csv')\nvar_names = [\"age\", \"sex\", \"educ\", \"race\", \"income\", \"married\"]\n\ndata = np.genfromtxt(data_path, delimiter=',', names=True)",
"_____no_output_____"
]
],
[
[
"### Functionality\n\nBelow we show the relationship between the three methods by calculating the same covariance in each. We use a much larger $\\epsilon$ than would ever be used in practice to show that the methods are consistent with one another. ",
"_____no_output_____"
]
],
[
[
"with sn.Analysis() as analysis:\n wn_data = sn.Dataset(path = data_path, column_names = var_names)\n \n # get scalar covariance\n age_income_cov_scalar = sn.dp_covariance(left = sn.to_float(wn_data['age']),\n right = sn.to_float(wn_data['income']),\n privacy_usage = {'epsilon': 5000},\n left_lower = 0.,\n left_upper = 100.,\n left_rows = 1000,\n right_lower = 0.,\n right_upper = 500_000.,\n right_rows = 1000)\n \n # get full covariance matrix\n age_income_cov_matrix = sn.dp_covariance(data = sn.to_float(wn_data['age', 'income']),\n privacy_usage = {'epsilon': 5000},\n data_lower = [0., 0.],\n data_upper = [100., 500_000],\n data_rows = 1000)\n\n # get cross-covariance matrix\n cross_covar = sn.dp_covariance(left = sn.to_float(wn_data['age', 'income']),\n right = sn.to_float(wn_data['age', 'income']),\n privacy_usage = {'epsilon': 5000},\n left_lower = [0., 0.],\n left_upper = [100., 500_000.],\n left_rows = 1_000,\n right_lower = [0., 0.],\n right_upper = [100., 500_000.],\n right_rows = 1000)\n\n# analysis.release()\nprint('scalar covariance:\\n{0}\\n'.format(age_income_cov_scalar.value))\nprint('covariance matrix:\\n{0}\\n'.format(age_income_cov_matrix.value)) \nprint('cross-covariance matrix:\\n{0}'.format(cross_covar.value))",
"scalar covariance:\n[[94601.1327106]]\n\ncovariance matrix:\n[[3.14917795e+02 9.46406149e+04]\n [9.46406149e+04 2.65091456e+09]]\n\ncross-covariance matrix:\n[[3.14928928e+02 9.50137258e+04]\n [9.46392159e+04 2.65184060e+09]]\n"
]
],
[
[
"### DP Covariance in Practice\n \nWe now move to an example with a much smaller $\\epsilon$. ",
"_____no_output_____"
]
],
[
[
"with sn.Analysis() as analysis:\n wn_data = sn.Dataset(path = data_path, column_names = var_names)\n # get full covariance matrix\n cov = sn.dp_covariance(data = sn.to_float(wn_data['age', 'sex', 'educ', 'income', 'married']),\n privacy_usage = {'epsilon': 1.},\n data_lower = [0., 0., 1., 0., 0.],\n data_upper = [100., 1., 16., 500_000., 1.],\n data_rows = 1000)\nanalysis.release()\n\n# store DP covariance and correlation matrix\ndp_cov = cov.value\ndp_corr = dp_cov / np.outer(np.sqrt(np.diag(dp_cov)), np.sqrt(np.diag(dp_cov)))\n\n# get non-DP covariance/correlation matrices\nage = list(data[:]['age'])\nsex = list(data[:]['sex'])\neduc = list(data[:]['educ'])\nincome = list(data[:]['income'])\nmarried = list(data[:]['married'])\nnon_dp_cov = np.cov([age, sex, educ, income, married])\nnon_dp_corr = non_dp_cov / np.outer(np.sqrt(np.diag(non_dp_cov)), np.sqrt(np.diag(non_dp_cov)))\n\nprint('Non-DP Correlation Matrix:\\n{0}\\n\\n'.format(pd.DataFrame(non_dp_corr)))\nprint('DP Correlation Matrix:\\n{0}'.format(pd.DataFrame(dp_corr)))",
"Non-DP Correlation Matrix:\n 0 1 2 3 4\n0 1.000000 0.055088 0.025901 0.103524 0.196072\n1 0.055088 1.000000 -0.023111 -0.211106 -0.073125\n2 0.025901 -0.023111 1.000000 0.305238 0.053860\n3 0.103524 -0.211106 0.305238 1.000000 0.153143\n4 0.196072 -0.073125 0.053860 0.153143 1.000000\n\n\nDP Correlation Matrix:\n 0 1 2 3 4\n0 1.000000 0.575078 -0.369702 1.090716 0.173756\n1 0.575078 1.000000 -0.068965 0.844628 0.199602\n2 -0.369702 -0.068965 1.000000 -1.054197 0.167272\n3 1.090716 0.844628 -1.054197 1.000000 -1.385910\n4 0.173756 0.199602 0.167272 -1.385910 1.000000\n"
],
[
"fig, (ax_1, ax_2) = plt.subplots(1, 2, figsize = (9, 11))\n\n# generate a mask for the upper triangular matrix\nmask = np.triu(np.ones_like(non_dp_corr, dtype = np.bool))\n\n# generate color palette\ncmap = sns.diverging_palette(220, 10, as_cmap = True)\n\n# get correlation plots\nax_1.title.set_text('Non-DP Correlation Matrix')\nsns.heatmap(non_dp_corr, mask=mask, cmap=cmap, vmax=.3, center=0,\n square=True, linewidths=.5, cbar_kws={\"shrink\": .5}, ax = ax_1)\nax_1.set_xticklabels(labels = ['age', 'sex', 'educ', 'income', 'married'], rotation = 45)\nax_1.set_yticklabels(labels = ['age', 'sex', 'educ', 'income', 'married'], rotation = 45)\n\n\nax_2.title.set_text('DP Correlation Matrix')\nsns.heatmap(dp_corr, mask=mask, cmap=cmap, vmax=.3, center=0,\n square=True, linewidths=.5, cbar_kws={\"shrink\": .5}, ax = ax_2)\nax_2.set_xticklabels(labels = ['age', 'sex', 'educ', 'income', 'married'], rotation = 45)\nax_2.set_yticklabels(labels = ['age', 'sex', 'educ', 'income', 'married'], rotation = 45)\n\n",
"_____no_output_____"
]
],
[
[
"Notice that the differentially private correlation matrix contains values outside of the feasible range for correlations, $[-1, 1]$. This is not uncommon, especially for analyses with small $\\epsilon$, and is not necessarily indicative of a problem. In this scenario, we will not use these correlations for anything other than visualization, so we will leave our result as is.\n\nSometimes, you may get a result that does cause problems for downstream analysis. For example, say your differentially private covariance matrix is not positive semi-definite. There are a number of ways to deal with problems of this type.\n\n1. Relax your original plans: For example, if you want to invert your DP covariance matrix and are unable to do so, you could instead take the pseudoinverse.\n2. Manual Post-Processing: Choose some way to change the output such that it is consistent with what you need for later analyses. This changed output is still differentially private (we will use this idea again in the next section). For example, map all negative variances to small positive value.\n3. More releases: You could perform the same release again (perhaps with a larger $\\epsilon$) and combine your results in some way until you have a release that works for your purposes. Note that additional $\\epsilon$ from will be consumed everytime this happens. ",
"_____no_output_____"
],
[
"### Post-Processing of DP Covariance Matrix: Regression Coefficient\n\nDifferentially private outputs are \"immune\" to post-processing, meaning functions of differentially private releases are also differentially private (provided that the functions are independent of the underlying data in the dataset). This idea provides us with a relatively easy way to generate complex differentially private releases from simpler ones.\n\nSay we wanted to run a linear regression of the form $income = \\alpha + \\beta \\cdot educ$ and want to find an differentially private estimate of the slope, $\\hat{\\beta}_{DP}$. We know that \n$$ \\beta = \\frac{cov(income, educ)}{var(educ)}, $$ \nand so \n$$ \\hat{\\beta}_{DP} = \\frac{\\hat{cov}(income, educ)_{DP}}{ \\hat{var}(educ)_{DP} }. $$\n\nWe already have differentially private estimates of the necessary covariance and variance, so we can plug them in to find $\\hat{\\beta}_{DP}$.\n\n",
"_____no_output_____"
]
],
[
[
"'''income = alpha + beta * educ'''\n# find DP estimate of beta\nbeta_hat_dp = dp_cov[2,3] / dp_cov[2,2]\nbeta_hat = non_dp_cov[2,3] / non_dp_cov[2,2]\n\nprint('income = alpha + beta * educ')\nprint('DP coefficient: {0}'.format(beta_hat_dp))\nprint('Non-DP Coefficient: {0}'.format(beta_hat))",
"income = alpha + beta * educ\nDP coefficient: -10900.063151950144\nNon-DP Coefficient: 4601.803740280991\n"
]
],
[
[
"This result is implausible, as it would suggest that an extra year of education is associated with, on average, a decrease in annual income of nearly $11,000. It's not uncommon for this to be the case for DP releases constructed as post-processing from other releases, especially when they involve taking ratios. \n\nIf you find yourself in such as situation, it is often worth it to spend some extra privacy budget to estimate your quantity of interest using an algorithm optimized for that specific use case.",
"_____no_output_____"
]
]
] | [
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown"
] | [
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
]
] |
d0fde6cf27349b4a966e8f47986b150ceffd862e | 254,417 | ipynb | Jupyter Notebook | Model_Quickload.ipynb | mashyko/object_detection | e46ac270a9145f494af2bdb54ad08a4d2f433f86 | [
"MIT"
] | 1 | 2021-03-23T04:41:29.000Z | 2021-03-23T04:41:29.000Z | Model_Quickload.ipynb | mashyko/object_detection | e46ac270a9145f494af2bdb54ad08a4d2f433f86 | [
"MIT"
] | null | null | null | Model_Quickload.ipynb | mashyko/object_detection | e46ac270a9145f494af2bdb54ad08a4d2f433f86 | [
"MIT"
] | null | null | null | 785.237654 | 175,434 | 0.729067 | [
[
[
"<a href=\"https://colab.research.google.com/github/mashyko/object_detection/blob/master/Model_Quickload.ipynb\" target=\"_parent\"><img src=\"https://colab.research.google.com/assets/colab-badge.svg\" alt=\"Open In Colab\"/></a>",
"_____no_output_____"
],
[
"#Tutorials Installation:\nhttps://caffe2.ai/docs/tutorials.html\n\nFirst download the tutorials source.\n\n from google.colab import drive\n\n drive.mount('/content/drive')\n\n %cd /content/drive/My Drive/\n\n !git clone --recursive https://github.com/caffe2/tutorials caffe2_tutorials\n\n",
"_____no_output_____"
],
[
"# Model Quickload\n\nThis notebook will show you how to quickly load a pretrained SqueezeNet model and test it on images of your choice in four main steps. \n\n1. Load the model\n2. Format the input\n3. Run the test\n4. Process the results\n\nThe model used in this tutorial has been pretrained on the full 1000 class ImageNet dataset, and is downloaded from Caffe2's [Model Zoo](https://github.com/caffe2/caffe2/wiki/Model-Zoo). For an all around more in-depth tutorial on using pretrained models check out the [Loading Pretrained Models](https://github.com/caffe2/caffe2/blob/master/caffe2/python/tutorials/Loading_Pretrained_Models.ipynb) tutorial. \n\nBefore this script will work, you need to download the model and install it. You can do this by running:\n\n```\nsudo python -m caffe2.python.models.download -i squeezenet\n```\n\nOr make a folder named `squeezenet`, download each file listed below to it, and place it in the `/caffe2/python/models/` directory:\n* [predict_net.pb](https://download.caffe2.ai/models/squeezenet/predict_net.pb)\n* [init_net.pb](https://download.caffe2.ai/models/squeezenet/init_net.pb)\n\nNotice, the helper function *parseResults* will translate the integer class label of the top result to an English label by searching through the [inference codes file](inference_codes.txt). If you want to really test the model's capabilities, pick a code from the file, find an image representing that code, and test the model with it!",
"_____no_output_____"
]
],
[
[
"from google.colab import drive\ndrive.mount('/content/drive')\n%cd /content/drive/My Drive/caffe2_tutorials",
"Go to this URL in a browser: https://accounts.google.com/o/oauth2/auth?client_id=947318989803-6bn6qk8qdgf4n4g3pfee6491hc0brc4i.apps.googleusercontent.com&redirect_uri=urn%3aietf%3awg%3aoauth%3a2.0%3aoob&response_type=code&scope=email%20https%3a%2f%2fwww.googleapis.com%2fauth%2fdocs.test%20https%3a%2f%2fwww.googleapis.com%2fauth%2fdrive%20https%3a%2f%2fwww.googleapis.com%2fauth%2fdrive.photos.readonly%20https%3a%2f%2fwww.googleapis.com%2fauth%2fpeopleapi.readonly\n\nEnter your authorization code:\n··········\nMounted at /content/drive\n/content/drive/My Drive/caffe2_tutorials\n"
],
[
"!pip3 install torch torchvision",
"Requirement already satisfied: torch in /usr/local/lib/python3.6/dist-packages (1.4.0)\nRequirement already satisfied: torchvision in /usr/local/lib/python3.6/dist-packages (0.5.0)\nRequirement already satisfied: numpy in /usr/local/lib/python3.6/dist-packages (from torchvision) (1.18.2)\nRequirement already satisfied: pillow>=4.1.1 in /usr/local/lib/python3.6/dist-packages (from torchvision) (7.0.0)\nRequirement already satisfied: six in /usr/local/lib/python3.6/dist-packages (from torchvision) (1.12.0)\n"
],
[
"!python -m caffe2.python.models.download -i squeezenet",
"Downloading from https://s3.amazonaws.com/download.caffe2.ai/models/squeezenet/predict_net.pb\nWriting to /usr/local/lib/python2.7/dist-packages/caffe2/python/models/squeezenet/predict_net.pb\n\u001b[1000D[ ] 0%\u001b[1000D[######################################################################] 100%\nDownloading from https://s3.amazonaws.com/download.caffe2.ai/models/squeezenet/init_net.pb\nWriting to /usr/local/lib/python2.7/dist-packages/caffe2/python/models/squeezenet/init_net.pb\n\u001b[1000D[ ] 0%\u001b[1000D[ ] 0%\u001b[1000D[ ] 0%\u001b[1000D[ ] 0%\u001b[1000D[ ] 0%\u001b[1000D[ ] 0%\u001b[1000D[ ] 0%\u001b[1000D[ ] 0%\u001b[1000D[ ] 1%\u001b[1000D[ ] 1%\u001b[1000D[ ] 1%\u001b[1000D[ ] 1%\u001b[1000D[ ] 1%\u001b[1000D[ ] 1%\u001b[1000D[ ] 1%\u001b[1000D[ ] 1%\u001b[1000D[# ] 2%\u001b[1000D[# ] 2%\u001b[1000D[# ] 2%\u001b[1000D[# ] 2%\u001b[1000D[# ] 2%\u001b[1000D[# ] 2%\u001b[1000D[# ] 2%\u001b[1000D[## ] 3%\u001b[1000D[## ] 3%\u001b[1000D[## ] 3%\u001b[1000D[## ] 3%\u001b[1000D[## ] 3%\u001b[1000D[## ] 3%\u001b[1000D[## ] 3%\u001b[1000D[## ] 3%\u001b[1000D[## ] 4%\u001b[1000D[## ] 4%\u001b[1000D[## ] 4%\u001b[1000D[## ] 4%\u001b[1000D[## ] 4%\u001b[1000D[## ] 4%\u001b[1000D[## ] 4%\u001b[1000D[### ] 5%\u001b[1000D[### ] 5%\u001b[1000D[### ] 5%\u001b[1000D[### ] 5%\u001b[1000D[### ] 5%\u001b[1000D[### ] 5%\u001b[1000D[### ] 5%\u001b[1000D[### ] 5%\u001b[1000D[#### ] 6%\u001b[1000D[#### ] 6%\u001b[1000D[#### ] 6%\u001b[1000D[#### ] 6%\u001b[1000D[#### ] 6%\u001b[1000D[#### ] 6%\u001b[1000D[#### ] 6%\u001b[1000D[#### ] 7%\u001b[1000D[#### ] 7%\u001b[1000D[#### ] 7%\u001b[1000D[#### ] 7%\u001b[1000D[#### ] 7%\u001b[1000D[#### ] 7%\u001b[1000D[#### ] 7%\u001b[1000D[#### ] 7%\u001b[1000D[##### ] 8%\u001b[1000D[##### ] 8%\u001b[1000D[##### ] 8%\u001b[1000D[##### ] 8%\u001b[1000D[##### ] 8%\u001b[1000D[##### ] 8%\u001b[1000D[##### ] 8%\u001b[1000D[###### ] 9%\u001b[1000D[###### ] 9%\u001b[1000D[###### ] 9%\u001b[1000D[###### ] 9%\u001b[1000D[###### ] 9%\u001b[1000D[###### ] 9%\u001b[1000D[###### ] 9%\u001b[1000D[###### ] 9%\u001b[1000D[####### ] 10%\u001b[1000D[####### ] 10%\u001b[1000D[####### ] 10%\u001b[1000D[####### ] 10%\u001b[1000D[####### ] 10%\u001b[1000D[####### ] 10%\u001b[1000D[####### ] 10%\u001b[1000D[####### ] 11%\u001b[1000D[####### ] 11%\u001b[1000D[####### ] 11%\u001b[1000D[####### ] 11%\u001b[1000D[####### ] 11%\u001b[1000D[####### ] 11%\u001b[1000D[####### ] 11%\u001b[1000D[####### ] 11%\u001b[1000D[######## ] 12%\u001b[1000D[######## ] 12%\u001b[1000D[######## ] 12%\u001b[1000D[######## ] 12%\u001b[1000D[######## ] 12%\u001b[1000D[######## ] 12%\u001b[1000D[######## ] 12%\u001b[1000D[######## ] 12%\u001b[1000D[######### ] 13%\u001b[1000D[######### ] 13%\u001b[1000D[######### ] 13%\u001b[1000D[######### ] 13%\u001b[1000D[######### ] 13%\u001b[1000D[######### ] 13%\u001b[1000D[######### ] 13%\u001b[1000D[######### ] 14%\u001b[1000D[######### ] 14%\u001b[1000D[######### ] 14%\u001b[1000D[######### ] 14%\u001b[1000D[######### ] 14%\u001b[1000D[######### ] 14%\u001b[1000D[######### ] 14%\u001b[1000D[######### ] 14%\u001b[1000D[########## ] 15%\u001b[1000D[########## ] 15%\u001b[1000D[########## ] 15%\u001b[1000D[########## ] 15%\u001b[1000D[########## ] 15%\u001b[1000D[########## ] 15%\u001b[1000D[########## ] 15%\u001b[1000D[########### ] 16%\u001b[1000D[########### ] 16%\u001b[1000D[########### ] 16%\u001b[1000D[########### ] 16%\u001b[1000D[########### ] 16%\u001b[1000D[########### ] 16%\u001b[1000D[########### ] 16%\u001b[1000D[########### ] 16%\u001b[1000D[########### ] 17%\u001b[1000D[########### ] 17%\u001b[1000D[########### ] 17%\u001b[1000D[########### ] 17%\u001b[1000D[########### ] 17%\u001b[1000D[########### ] 17%\u001b[1000D[########### ] 17%\u001b[1000D[############ ] 18%\u001b[1000D[############ ] 18%\u001b[1000D[############ ] 18%\u001b[1000D[############ ] 18%\u001b[1000D[############ ] 18%\u001b[1000D[############ ] 18%\u001b[1000D[############ ] 18%\u001b[1000D[############ ] 18%\u001b[1000D[############# ] 19%\u001b[1000D[############# ] 19%\u001b[1000D[############# ] 19%\u001b[1000D[############# ] 19%\u001b[1000D[############# ] 19%\u001b[1000D[############# ] 19%\u001b[1000D[############# ] 19%\u001b[1000D[############## ] 20%\u001b[1000D[############## ] 20%\u001b[1000D[############## ] 20%\u001b[1000D[############## ] 20%\u001b[1000D[############## ] 20%\u001b[1000D[############## ] 20%\u001b[1000D[############## ] 20%\u001b[1000D[############## ] 20%\u001b[1000D[############## ] 21%\u001b[1000D[############## ] 21%\u001b[1000D[############## ] 21%\u001b[1000D[############## ] 21%\u001b[1000D[############## ] 21%\u001b[1000D[############## ] 21%\u001b[1000D[############## ] 21%\u001b[1000D[############### ] 22%\u001b[1000D[############### ] 22%\u001b[1000D[############### ] 22%\u001b[1000D[############### ] 22%\u001b[1000D[############### ] 22%\u001b[1000D[############### ] 22%\u001b[1000D[############### ] 22%\u001b[1000D[############### ] 22%\u001b[1000D[################ ] 23%\u001b[1000D[################ ] 23%\u001b[1000D[################ ] 23%\u001b[1000D[################ ] 23%\u001b[1000D[################ ] 23%\u001b[1000D[################ ] 23%\u001b[1000D[################ ] 23%\u001b[1000D[################ ] 23%\u001b[1000D[################ ] 24%\u001b[1000D[################ ] 24%\u001b[1000D[################ ] 24%\u001b[1000D[################ ] 24%\u001b[1000D[################ ] 24%\u001b[1000D[################ ] 24%\u001b[1000D[################ ] 24%\u001b[1000D[################# ] 25%\u001b[1000D[################# ] 25%\u001b[1000D[################# ] 25%\u001b[1000D[################# ] 25%\u001b[1000D[################# ] 25%\u001b[1000D[################# ] 25%\u001b[1000D[################# ] 25%\u001b[1000D[################# ] 25%\u001b[1000D[################## ] 26%\u001b[1000D[################## ] 26%\u001b[1000D[################## ] 26%\u001b[1000D[################## ] 26%\u001b[1000D[################## ] 26%\u001b[1000D[################## ] 26%\u001b[1000D[################## ] 26%\u001b[1000D[################## ] 27%\u001b[1000D[################## ] 27%\u001b[1000D[################## ] 27%\u001b[1000D[################## ] 27%\u001b[1000D[################## ] 27%\u001b[1000D[################## ] 27%\u001b[1000D[################## ] 27%\u001b[1000D[################## ] 27%\u001b[1000D[################### ] 28%\u001b[1000D[################### ] 28%\u001b[1000D[################### ] 28%\u001b[1000D[################### ] 28%\u001b[1000D[################### ] 28%\u001b[1000D[################### ] 28%\u001b[1000D[################### ] 28%\u001b[1000D[#################### ] 29%\u001b[1000D[#################### ] 29%\u001b[1000D[#################### ] 29%\u001b[1000D[#################### ] 29%\u001b[1000D[#################### ] 29%\u001b[1000D[#################### ] 29%\u001b[1000D[#################### ] 29%\u001b[1000D[#################### ] 29%\u001b[1000D[##################### ] 30%\u001b[1000D[##################### ] 30%\u001b[1000D[##################### ] 30%\u001b[1000D[##################### ] 30%\u001b[1000D[##################### ] 30%\u001b[1000D[##################### ] 30%\u001b[1000D[##################### ] 30%\u001b[1000D[##################### ] 31%\u001b[1000D[##################### ] 31%\u001b[1000D[##################### ] 31%\u001b[1000D[##################### ] 31%\u001b[1000D[##################### ] 31%\u001b[1000D[##################### ] 31%\u001b[1000D[##################### ] 31%\u001b[1000D[##################### ] 31%\u001b[1000D[###################### ] 32%\u001b[1000D[###################### ] 32%\u001b[1000D[###################### ] 32%\u001b[1000D[###################### ] 32%\u001b[1000D[###################### ] 32%\u001b[1000D[###################### ] 32%\u001b[1000D[###################### ] 32%\u001b[1000D[####################### ] 33%\u001b[1000D[####################### ] 33%\u001b[1000D[####################### ] 33%\u001b[1000D[####################### ] 33%\u001b[1000D[####################### ] 33%\u001b[1000D[####################### ] 33%\u001b[1000D[####################### ] 33%\u001b[1000D[####################### ] 33%\u001b[1000D[####################### ] 34%\u001b[1000D[####################### ] 34%\u001b[1000D[####################### ] 34%\u001b[1000D[####################### ] 34%\u001b[1000D[####################### ] 34%\u001b[1000D[####################### ] 34%\u001b[1000D[####################### ] 34%\u001b[1000D[####################### ] 34%\u001b[1000D[######################## ] 35%\u001b[1000D[######################## ] 35%\u001b[1000D[######################## ] 35%\u001b[1000D[######################## ] 35%\u001b[1000D[######################## ] 35%\u001b[1000D[######################## ] 35%\u001b[1000D[######################## ] 35%\u001b[1000D[######################### ] 36%\u001b[1000D[######################### ] 36%\u001b[1000D[######################### ] 36%\u001b[1000D[######################### ] 36%\u001b[1000D[######################### ] 36%\u001b[1000D[######################### ] 36%\u001b[1000D[######################### ] 36%\u001b[1000D[######################### ] 36%\u001b[1000D[######################### ] 37%\u001b[1000D[######################### ] 37%\u001b[1000D[######################### ] 37%\u001b[1000D[######################### ] 37%\u001b[1000D[######################### ] 37%\u001b[1000D[######################### ] 37%\u001b[1000D[######################### ] 37%\u001b[1000D[########################## ] 38%\u001b[1000D[########################## ] 38%\u001b[1000D[########################## ] 38%\u001b[1000D[########################## ] 38%\u001b[1000D[########################## ] 38%\u001b[1000D[########################## ] 38%\u001b[1000D[########################## ] 38%\u001b[1000D[########################## ] 38%\u001b[1000D[########################### ] 39%\u001b[1000D[########################### ] 39%\u001b[1000D[########################### ] 39%\u001b[1000D[########################### ] 39%\u001b[1000D[########################### ] 39%\u001b[1000D[########################### ] 39%\u001b[1000D[########################### ] 39%\u001b[1000D[############################ ] 40%\u001b[1000D[############################ ] 40%\u001b[1000D[############################ ] 40%\u001b[1000D[############################ ] 40%\u001b[1000D[############################ ] 40%\u001b[1000D[############################ ] 40%\u001b[1000D[############################ ] 40%\u001b[1000D[############################ ] 40%\u001b[1000D[############################ ] 41%\u001b[1000D[############################ ] 41%\u001b[1000D[############################ ] 41%\u001b[1000D[############################ ] 41%\u001b[1000D[############################ ] 41%\u001b[1000D[############################ ] 41%\u001b[1000D[############################ ] 41%\u001b[1000D[############################# ] 42%\u001b[1000D[############################# ] 42%\u001b[1000D[############################# ] 42%\u001b[1000D[############################# ] 42%\u001b[1000D[############################# ] 42%\u001b[1000D[############################# ] 42%\u001b[1000D[############################# ] 42%\u001b[1000D[############################# ] 42%\u001b[1000D[############################## ] 43%\u001b[1000D[############################## ] 43%\u001b[1000D[############################## ] 43%\u001b[1000D[############################## ] 43%\u001b[1000D[############################## ] 43%\u001b[1000D[############################## ] 43%\u001b[1000D[############################## ] 43%\u001b[1000D[############################## ] 44%\u001b[1000D[############################## ] 44%\u001b[1000D[############################## ] 44%\u001b[1000D[############################## ] 44%\u001b[1000D[############################## ] 44%\u001b[1000D[############################## ] 44%\u001b[1000D[############################## ] 44%\u001b[1000D[############################## ] 44%\u001b[1000D[############################### ] 45%\u001b[1000D[############################### ] 45%\u001b[1000D[############################### ] 45%\u001b[1000D[############################### ] 45%\u001b[1000D[############################### ] 45%\u001b[1000D[############################### ] 45%\u001b[1000D[############################### ] 45%\u001b[1000D[############################### ] 45%\u001b[1000D[################################ ] 46%\u001b[1000D[################################ ] 46%\u001b[1000D[################################ ] 46%\u001b[1000D[################################ ] 46%\u001b[1000D[################################ ] 46%\u001b[1000D[################################ ] 46%\u001b[1000D[################################ ] 46%\u001b[1000D[################################ ] 47%\u001b[1000D[################################ ] 47%\u001b[1000D[################################ ] 47%\u001b[1000D[################################ ] 47%\u001b[1000D[################################ ] 47%\u001b[1000D[################################ ] 47%\u001b[1000D[################################ ] 47%\u001b[1000D[################################ ] 47%\u001b[1000D[################################# ] 48%\u001b[1000D[################################# ] 48%\u001b[1000D[################################# ] 48%\u001b[1000D[################################# ] 48%\u001b[1000D[################################# ] 48%\u001b[1000D[################################# ] 48%\u001b[1000D[################################# ] 48%\u001b[1000D[################################## ] 49%\u001b[1000D[################################## ] 49%\u001b[1000D[################################## ] 49%\u001b[1000D[################################## ] 49%\u001b[1000D[################################## ] 49%\u001b[1000D[################################## ] 49%\u001b[1000D[################################## ] 49%\u001b[1000D[################################## ] 49%\u001b[1000D[################################### ] 50%\u001b[1000D[################################### ] 50%\u001b[1000D[################################### ] 50%\u001b[1000D[################################### ] 50%\u001b[1000D[################################### ] 50%\u001b[1000D[################################### ] 50%\u001b[1000D[################################### ] 50%\u001b[1000D[################################### ] 51%\u001b[1000D[################################### ] 51%\u001b[1000D[################################### ] 51%\u001b[1000D[################################### ] 51%\u001b[1000D[################################### ] 51%\u001b[1000D[################################### ] 51%\u001b[1000D[################################### ] 51%\u001b[1000D[################################### ] 51%\u001b[1000D[#################################### ] 52%\u001b[1000D[#################################### ] 52%\u001b[1000D[#################################### ] 52%\u001b[1000D[#################################### ] 52%\u001b[1000D[#################################### ] 52%\u001b[1000D[#################################### ] 52%\u001b[1000D[#################################### ] 52%\u001b[1000D[##################################### ] 53%\u001b[1000D[##################################### ] 53%\u001b[1000D[##################################### ] 53%\u001b[1000D[##################################### ] 53%\u001b[1000D[##################################### ] 53%\u001b[1000D[##################################### ] 53%\u001b[1000D[##################################### ] 53%\u001b[1000D[##################################### ] 53%\u001b[1000D[##################################### ] 54%\u001b[1000D[##################################### ] 54%\u001b[1000D[##################################### ] 54%\u001b[1000D[##################################### ] 54%\u001b[1000D[##################################### ] 54%\u001b[1000D[##################################### ] 54%\u001b[1000D[##################################### ] 54%\u001b[1000D[###################################### ] 55%\u001b[1000D[###################################### ] 55%\u001b[1000D[###################################### ] 55%\u001b[1000D[###################################### ] 55%\u001b[1000D[###################################### ] 55%\u001b[1000D[###################################### ] 55%\u001b[1000D[###################################### ] 55%\u001b[1000D[###################################### ] 55%\u001b[1000D[####################################### ] 56%\u001b[1000D[####################################### ] 56%\u001b[1000D[####################################### ] 56%\u001b[1000D[####################################### ] 56%\u001b[1000D[####################################### ] 56%\u001b[1000D[####################################### ] 56%\u001b[1000D[####################################### ] 56%\u001b[1000D[####################################### ] 56%\u001b[1000D[####################################### ] 57%\u001b[1000D[####################################### ] 57%\u001b[1000D[####################################### ] 57%\u001b[1000D[####################################### ] 57%\u001b[1000D[####################################### ] 57%\u001b[1000D[####################################### ] 57%\u001b[1000D[####################################### ] 57%\u001b[1000D[######################################## ] 58%\u001b[1000D[######################################## ] 58%\u001b[1000D[######################################## ] 58%\u001b[1000D[######################################## ] 58%\u001b[1000D[######################################## ] 58%\u001b[1000D[######################################## ] 58%\u001b[1000D[######################################## ] 58%\u001b[1000D[######################################## ] 58%\u001b[1000D[######################################### ] 59%\u001b[1000D[######################################### ] 59%\u001b[1000D[######################################### ] 59%\u001b[1000D[######################################### ] 59%\u001b[1000D[######################################### ] 59%\u001b[1000D[######################################### ] 59%\u001b[1000D[######################################### ] 59%\u001b[1000D[########################################## ] 60%\u001b[1000D[########################################## ] 60%\u001b[1000D[########################################## ] 60%\u001b[1000D[########################################## ] 60%\u001b[1000D[########################################## ] 60%\u001b[1000D[########################################## ] 60%\u001b[1000D[########################################## ] 60%\u001b[1000D[########################################## ] 60%\u001b[1000D[########################################## ] 61%\u001b[1000D[########################################## ] 61%\u001b[1000D[########################################## ] 61%\u001b[1000D[########################################## ] 61%\u001b[1000D[########################################## ] 61%\u001b[1000D[########################################## ] 61%\u001b[1000D[########################################## ] 61%\u001b[1000D[########################################### ] 62%\u001b[1000D[########################################### ] 62%\u001b[1000D[########################################### ] 62%\u001b[1000D[########################################### ] 62%\u001b[1000D[########################################### ] 62%\u001b[1000D[########################################### ] 62%\u001b[1000D[########################################### ] 62%\u001b[1000D[########################################### ] 62%\u001b[1000D[############################################ ] 63%\u001b[1000D[############################################ ] 63%\u001b[1000D[############################################ ] 63%\u001b[1000D[############################################ ] 63%\u001b[1000D[############################################ ] 63%\u001b[1000D[############################################ ] 63%\u001b[1000D[############################################ ] 63%\u001b[1000D[############################################ ] 64%\u001b[1000D[############################################ ] 64%\u001b[1000D[############################################ ] 64%\u001b[1000D[############################################ ] 64%\u001b[1000D[############################################ ] 64%\u001b[1000D[############################################ ] 64%\u001b[1000D[############################################ ] 64%\u001b[1000D[############################################ ] 64%\u001b[1000D[############################################# ] 65%\u001b[1000D[############################################# ] 65%\u001b[1000D[############################################# ] 65%\u001b[1000D[############################################# ] 65%\u001b[1000D[############################################# ] 65%\u001b[1000D[############################################# ] 65%\u001b[1000D[############################################# ] 65%\u001b[1000D[############################################## ] 66%\u001b[1000D[############################################## ] 66%\u001b[1000D[############################################## ] 66%\u001b[1000D[############################################## ] 66%\u001b[1000D[############################################## ] 66%\u001b[1000D[############################################## ] 66%\u001b[1000D[############################################## ] 66%\u001b[1000D[############################################## ] 66%\u001b[1000D[############################################## ] 67%\u001b[1000D[############################################## ] 67%\u001b[1000D[############################################## ] 67%\u001b[1000D[############################################## ] 67%\u001b[1000D[############################################## ] 67%\u001b[1000D[############################################## ] 67%\u001b[1000D[############################################## ] 67%\u001b[1000D[############################################## ] 67%\u001b[1000D[############################################### ] 68%\u001b[1000D[############################################### ] 68%\u001b[1000D[############################################### ] 68%\u001b[1000D[############################################### ] 68%\u001b[1000D[############################################### ] 68%\u001b[1000D[############################################### ] 68%\u001b[1000D[############################################### ] 68%\u001b[1000D[################################################ ] 69%\u001b[1000D[################################################ ] 69%\u001b[1000D[################################################ ] 69%\u001b[1000D[################################################ ] 69%\u001b[1000D[################################################ ] 69%\u001b[1000D[################################################ ] 69%\u001b[1000D[################################################ ] 69%\u001b[1000D[################################################ ] 69%\u001b[1000D[################################################# ] 70%\u001b[1000D[################################################# ] 70%\u001b[1000D[################################################# ] 70%\u001b[1000D[################################################# ] 70%\u001b[1000D[################################################# ] 70%\u001b[1000D[################################################# ] 70%\u001b[1000D[################################################# ] 70%\u001b[1000D[################################################# ] 71%\u001b[1000D[################################################# ] 71%\u001b[1000D[################################################# ] 71%\u001b[1000D[################################################# ] 71%\u001b[1000D[################################################# ] 71%\u001b[1000D[################################################# ] 71%\u001b[1000D[################################################# ] 71%\u001b[1000D[################################################# ] 71%\u001b[1000D[################################################## ] 72%\u001b[1000D[################################################## ] 72%\u001b[1000D[################################################## ] 72%\u001b[1000D[################################################## ] 72%\u001b[1000D[################################################## ] 72%\u001b[1000D[################################################## ] 72%\u001b[1000D[################################################## ] 72%\u001b[1000D[################################################### ] 73%\u001b[1000D[################################################### ] 73%\u001b[1000D[################################################### ] 73%\u001b[1000D[################################################### ] 73%\u001b[1000D[################################################### ] 73%\u001b[1000D[################################################### ] 73%\u001b[1000D[################################################### ] 73%\u001b[1000D[################################################### ] 73%\u001b[1000D[################################################### ] 74%\u001b[1000D[################################################### ] 74%\u001b[1000D[################################################### ] 74%\u001b[1000D[################################################### ] 74%\u001b[1000D[################################################### ] 74%\u001b[1000D[################################################### ] 74%\u001b[1000D[################################################### ] 74%\u001b[1000D[#################################################### ] 75%\u001b[1000D[#################################################### ] 75%\u001b[1000D[#################################################### ] 75%\u001b[1000D[#################################################### ] 75%\u001b[1000D[#################################################### ] 75%\u001b[1000D[#################################################### ] 75%\u001b[1000D[#################################################### ] 75%\u001b[1000D[#################################################### ] 75%\u001b[1000D[##################################################### ] 76%\u001b[1000D[##################################################### ] 76%\u001b[1000D[##################################################### ] 76%\u001b[1000D[##################################################### ] 76%\u001b[1000D[##################################################### ] 76%\u001b[1000D[##################################################### ] 76%\u001b[1000D[##################################################### ] 76%\u001b[1000D[##################################################### ] 77%\u001b[1000D[##################################################### ] 77%\u001b[1000D[##################################################### ] 77%\u001b[1000D[##################################################### ] 77%\u001b[1000D[##################################################### ] 77%\u001b[1000D[##################################################### ] 77%\u001b[1000D[##################################################### ] 77%\u001b[1000D[##################################################### ] 77%\u001b[1000D[###################################################### ] 78%\u001b[1000D[###################################################### ] 78%\u001b[1000D[###################################################### ] 78%\u001b[1000D[###################################################### ] 78%\u001b[1000D[###################################################### ] 78%\u001b[1000D[###################################################### ] 78%\u001b[1000D[###################################################### ] 78%\u001b[1000D[###################################################### ] 78%\u001b[1000D[####################################################### ] 79%\u001b[1000D[####################################################### ] 79%\u001b[1000D[####################################################### ] 79%\u001b[1000D[####################################################### ] 79%\u001b[1000D[####################################################### ] 79%\u001b[1000D[####################################################### ] 79%\u001b[1000D[####################################################### ] 79%\u001b[1000D[######################################################## ] 80%\u001b[1000D[######################################################## ] 80%\u001b[1000D[######################################################## ] 80%\u001b[1000D[######################################################## ] 80%\u001b[1000D[######################################################## ] 80%\u001b[1000D[######################################################## ] 80%\u001b[1000D[######################################################## ] 80%\u001b[1000D[######################################################## ] 80%\u001b[1000D[######################################################## ] 81%\u001b[1000D[######################################################## ] 81%\u001b[1000D[######################################################## ] 81%\u001b[1000D[######################################################## ] 81%\u001b[1000D[######################################################## ] 81%\u001b[1000D[######################################################## ] 81%\u001b[1000D[######################################################## ] 81%\u001b[1000D[######################################################### ] 82%\u001b[1000D[######################################################### ] 82%\u001b[1000D[######################################################### ] 82%\u001b[1000D[######################################################### ] 82%\u001b[1000D[######################################################### ] 82%\u001b[1000D[######################################################### ] 82%\u001b[1000D[######################################################### ] 82%\u001b[1000D[######################################################### ] 82%\u001b[1000D[########################################################## ] 83%\u001b[1000D[########################################################## ] 83%\u001b[1000D[########################################################## ] 83%\u001b[1000D[########################################################## ] 83%\u001b[1000D[########################################################## ] 83%\u001b[1000D[########################################################## ] 83%\u001b[1000D[########################################################## ] 83%\u001b[1000D[########################################################## ] 84%\u001b[1000D[########################################################## ] 84%\u001b[1000D[########################################################## ] 84%\u001b[1000D[########################################################## ] 84%\u001b[1000D[########################################################## ] 84%\u001b[1000D[########################################################## ] 84%\u001b[1000D[########################################################## ] 84%\u001b[1000D[########################################################## ] 84%\u001b[1000D[########################################################### ] 85%\u001b[1000D[########################################################### ] 85%\u001b[1000D[########################################################### ] 85%\u001b[1000D[########################################################### ] 85%\u001b[1000D[########################################################### ] 85%\u001b[1000D[########################################################### ] 85%\u001b[1000D[########################################################### ] 85%\u001b[1000D[############################################################ ] 86%\u001b[1000D[############################################################ ] 86%\u001b[1000D[############################################################ ] 86%\u001b[1000D[############################################################ ] 86%\u001b[1000D[############################################################ ] 86%\u001b[1000D[############################################################ ] 86%\u001b[1000D[############################################################ ] 86%\u001b[1000D[############################################################ ] 86%\u001b[1000D[############################################################ ] 87%\u001b[1000D[############################################################ ] 87%\u001b[1000D[############################################################ ] 87%\u001b[1000D[############################################################ ] 87%\u001b[1000D[############################################################ ] 87%\u001b[1000D[############################################################ ] 87%\u001b[1000D[############################################################ ] 87%\u001b[1000D[############################################################# ] 88%\u001b[1000D[############################################################# ] 88%\u001b[1000D[############################################################# ] 88%\u001b[1000D[############################################################# ] 88%\u001b[1000D[############################################################# ] 88%\u001b[1000D[############################################################# ] 88%\u001b[1000D[############################################################# ] 88%\u001b[1000D[############################################################# ] 88%\u001b[1000D[############################################################## ] 89%\u001b[1000D[############################################################## ] 89%\u001b[1000D[############################################################## ] 89%\u001b[1000D[############################################################## ] 89%\u001b[1000D[############################################################## ] 89%\u001b[1000D[############################################################## ] 89%\u001b[1000D[############################################################## ] 89%\u001b[1000D[############################################################## ] 89%\u001b[1000D[############################################################### ] 90%\u001b[1000D[############################################################### ] 90%\u001b[1000D[############################################################### ] 90%\u001b[1000D[############################################################### ] 90%\u001b[1000D[############################################################### ] 90%\u001b[1000D[############################################################### ] 90%\u001b[1000D[############################################################### ] 90%\u001b[1000D[############################################################### ] 91%\u001b[1000D[############################################################### ] 91%\u001b[1000D[############################################################### ] 91%\u001b[1000D[############################################################### ] 91%\u001b[1000D[############################################################### ] 91%\u001b[1000D[############################################################### ] 91%\u001b[1000D[############################################################### ] 91%\u001b[1000D[############################################################### ] 91%\u001b[1000D[################################################################ ] 92%\u001b[1000D[################################################################ ] 92%\u001b[1000D[################################################################ ] 92%\u001b[1000D[################################################################ ] 92%\u001b[1000D[################################################################ ] 92%\u001b[1000D[################################################################ ] 92%\u001b[1000D[################################################################ ] 92%\u001b[1000D[################################################################# ] 93%\u001b[1000D[################################################################# ] 93%\u001b[1000D[################################################################# ] 93%\u001b[1000D[################################################################# ] 93%\u001b[1000D[################################################################# ] 93%\u001b[1000D[################################################################# ] 93%\u001b[1000D[################################################################# ] 93%\u001b[1000D[################################################################# ] 93%\u001b[1000D[################################################################# ] 94%\u001b[1000D[################################################################# ] 94%\u001b[1000D[################################################################# ] 94%\u001b[1000D[################################################################# ] 94%\u001b[1000D[################################################################# ] 94%\u001b[1000D[################################################################# ] 94%\u001b[1000D[################################################################# ] 94%\u001b[1000D[################################################################## ] 95%\u001b[1000D[################################################################## ] 95%\u001b[1000D[################################################################## ] 95%\u001b[1000D[################################################################## ] 95%\u001b[1000D[################################################################## ] 95%\u001b[1000D[################################################################## ] 95%\u001b[1000D[################################################################## ] 95%\u001b[1000D[################################################################## ] 95%\u001b[1000D[################################################################### ] 96%\u001b[1000D[################################################################### ] 96%\u001b[1000D[################################################################### ] 96%\u001b[1000D[################################################################### ] 96%\u001b[1000D[################################################################### ] 96%\u001b[1000D[################################################################### ] 96%\u001b[1000D[################################################################### ] 96%\u001b[1000D[################################################################### ] 97%\u001b[1000D[################################################################### ] 97%\u001b[1000D[################################################################### ] 97%\u001b[1000D[################################################################### ] 97%\u001b[1000D[################################################################### ] 97%\u001b[1000D[################################################################### ] 97%\u001b[1000D[################################################################### ] 97%\u001b[1000D[################################################################### ] 97%\u001b[1000D[#################################################################### ] 98%\u001b[1000D[#################################################################### ] 98%\u001b[1000D[#################################################################### ] 98%\u001b[1000D[#################################################################### ] 98%\u001b[1000D[#################################################################### ] 98%\u001b[1000D[#################################################################### ] 98%\u001b[1000D[#################################################################### ] 98%\u001b[1000D[##################################################################### ] 99%\u001b[1000D[##################################################################### ] 99%\u001b[1000D[##################################################################### ] 99%\u001b[1000D[##################################################################### ] 99%\u001b[1000D[##################################################################### ] 99%\u001b[1000D[##################################################################### ] 99%\u001b[1000D[##################################################################### ] 99%\u001b[1000D[##################################################################### ] 99%\u001b[1000D[######################################################################] 100%\n"
],
[
"from __future__ import absolute_import\nfrom __future__ import division\nfrom __future__ import print_function\nfrom __future__ import unicode_literals\nimport numpy as np\nimport operator\n# load up the caffe2 workspace\nfrom caffe2.python import workspace\n# choose your model here (use the downloader first)\nfrom caffe2.python.models import squeezenet as mynet\n# helper image processing functions\nimport helpers\n\n##### Load the Model\n# Load the pre-trained model\ninit_net = mynet.init_net\npredict_net = mynet.predict_net\n\n# Initialize the predictor with SqueezeNet's init_net and predict_net\np = workspace.Predictor(init_net, predict_net)\n\n##### Select and format the input image\n# use whatever image you want (urls work too)\n# img = \"https://upload.wikimedia.org/wikipedia/commons/a/ac/Pretzel.jpg\"\n# img = \"images/cat.jpg\"\n# img = \"images/cowboy-hat.jpg\"\n# img = \"images/cell-tower.jpg\"\n# img = \"images/Ducreux.jpg\"\n# img = \"images/pretzel.jpg\"\n# img = \"images/orangutan.jpg\"\n# img = \"images/aircraft-carrier.jpg\"\nimg = \"images/flower.jpg\"\n\n# average mean to subtract from the image\nmean = 128\n# the size of images that the model was trained with\ninput_size = 227\n\n# use the image helper to load the image and convert it to NCHW\nimg = helpers.loadToNCHW(img, mean, input_size)\n\n##### Run the test\n# submit the image to net and get a tensor of results\nresults = p.run({'data': img}) \n\n##### Process the results\n# Quick way to get the top-1 prediction result\n# Squeeze out the unnecessary axis. This returns a 1-D array of length 1000\npreds = np.squeeze(results)\n# Get the prediction and the confidence by finding the maximum value and index of maximum value in preds array\ncurr_pred, curr_conf = max(enumerate(preds), key=operator.itemgetter(1))\nprint(\"Top-1 Prediction: {}\".format(curr_pred))\nprint(\"Top-1 Confidence: {}\\n\".format(curr_conf))\n\n# Lookup our result from the inference list\nresponse = helpers.parseResults(results)\nprint(response)",
"/usr/local/lib/python2.7/dist-packages/skimage/transform/_warps.py:105: UserWarning: The default mode, 'constant', will be changed to 'reflect' in skimage 0.15.\n warn(\"The default mode, 'constant', will be changed to 'reflect' in \"\n/usr/local/lib/python2.7/dist-packages/skimage/transform/_warps.py:110: UserWarning: Anti-aliasing will be enabled by default in skimage 0.15 to avoid aliasing artifacts when down-sampling images.\n warn(\"Anti-aliasing will be enabled by default in skimage 0.15 to \"\n"
],
[
"%matplotlib inline\nimport matplotlib.pyplot as plt\nimport matplotlib.image as mpimg\n\nimg=mpimg.imread('images/flower.jpg') #image to array\n\n\n# show the original image\nplt.figure()\nplt.imshow(img)\nplt.axis('on')\nplt.title('Original image = RGB')\nplt.show()\n\n",
"_____no_output_____"
]
]
] | [
"markdown",
"code"
] | [
[
"markdown",
"markdown",
"markdown"
],
[
"code",
"code",
"code",
"code",
"code"
]
] |
d0fdebfb04cede765c0e538609f53f0370c6d965 | 62,556 | ipynb | Jupyter Notebook | notebooks/Dynamic Double Machine Learning Examples.ipynb | GildasTiwangNg/EconML | 8c6b801c9c51a191a3d347d32d678194fd4a42b9 | [
"BSD-3-Clause"
] | 259 | 2018-07-15T08:17:18.000Z | 2019-05-06T20:41:42.000Z | notebooks/Dynamic Double Machine Learning Examples.ipynb | GildasTiwangNg/EconML | 8c6b801c9c51a191a3d347d32d678194fd4a42b9 | [
"BSD-3-Clause"
] | 33 | 2019-01-30T22:11:52.000Z | 2019-05-04T19:53:17.000Z | notebooks/Dynamic Double Machine Learning Examples.ipynb | GildasTiwangNg/EconML | 8c6b801c9c51a191a3d347d32d678194fd4a42b9 | [
"BSD-3-Clause"
] | 32 | 2018-06-12T11:22:10.000Z | 2019-05-03T18:51:25.000Z | 80.302953 | 17,388 | 0.745348 | [
[
[
"<table border=\"0\">\n <tr>\n <td>\n <img src=\"https://ictd2016.files.wordpress.com/2016/04/microsoft-research-logo-copy.jpg\" style=\"width 30px;\" />\n </td>\n <td>\n <img src=\"https://www.microsoft.com/en-us/research/wp-content/uploads/2016/12/MSR-ALICE-HeaderGraphic-1920x720_1-800x550.jpg\" style=\"width 100px;\"/></td>\n </tr>\n</table>",
"_____no_output_____"
],
[
"# Dynamic Double Machine Learning: Use Cases and Examples\n\nDynamic DoubleML is an extension of the Double ML approach for treatments assigned sequentially over time periods. This estimator will account for treatments that can have causal effects on future outcomes. For more details, see [this paper](https://arxiv.org/abs/2002.07285) or the [EconML docummentation](https://econml.azurewebsites.net/).\n\nFor example, the Dynamic DoubleML could be useful in estimating the following causal effects:\n* the effect of investments on revenue at companies that receive investments at regular intervals ([see more](https://arxiv.org/abs/2103.08390))\n* the effect of prices on demand in stores where prices of goods change over time\n* the effect of income on health outcomes in people who receive yearly income\n\nThe preferred data format is balanced panel data. Each panel corresponds to one entity (e.g. company, store or person) and the different rows in a panel correspond to different time points. Example:\n\n||Company|Year|Features|Investment|Revenue|\n|---|---|---|---|---|---|\n|1|A|2018|...|\\$1,000|\\$10,000|\n|2|A|2019|...|\\$2,000|\\$12,000|\n|3|A|2020|...|\\$3,000|\\$15,000|\n|4|B|2018|...|\\$0|\\$5,000|\n|5|B|2019|...|\\$100|\\$10,000|\n|6|B|2020|...|\\$1,200|\\$7,000|\n|7|C|2018|...|\\$1,000|\\$20,000|\n|8|C|2019|...|\\$1,500|\\$25,000|\n|9|C|2020|...|\\$500|\\$15,000|\n\n(Note: when passing the data to the DynamicDML estimator, the \"Company\" column above corresponds to the `groups` argument at fit time. The \"Year\" column above should not be passed in as it will be inferred from the \"Company\" column)\n\nIf group memebers do not appear together, it is assumed that the first instance of a group in the dataset corresponds to the first period of that group, the second instance of the group corresponds to the second period, etc. Example:\n\n||Company|Features|Investment|Revenue|\n|---|---|---|---|---|\n|1|A|...|\\$1,000|\\$10,000|\n|2|B|...|\\$0|\\$5,000\n|3|C|...|\\$1,000|\\$20,000|\n|4|A|...|\\$2,000|\\$12,000|\n|5|B|...|\\$100|\\$10,000|\n|6|C|...|\\$1,500|\\$25,000|\n|7|A|...|\\$3,000|\\$15,000|\n|8|B|...|\\$1,200|\\$7,000|\n|9|C|...|\\$500|\\$15,000|\n\nIn this dataset, 1<sup>st</sup> row corresponds to the first period of group `A`, 4<sup>th</sup> row corresponds to the second period of group `A`, etc.\n\nIn this notebook, we show the performance of the DynamicDML on synthetic and observational data. \n\n## Notebook Contents\n\n1. [Example Usage with Average Treatment Effects](#1.-Example-Usage-with-Average-Treatment-Effects)\n2. [Example Usage with Heterogeneous Treatment Effects](#2.-Example-Usage-with-Heterogeneous-Treatment-Effects)",
"_____no_output_____"
]
],
[
[
"%load_ext autoreload\n%autoreload 2",
"_____no_output_____"
],
[
"import econml",
"_____no_output_____"
],
[
"# Main imports\nfrom econml.dynamic.dml import DynamicDML\nfrom econml.tests.dgp import DynamicPanelDGP, add_vlines\n\n# Helper imports\nimport numpy as np\nfrom sklearn.linear_model import Lasso, LassoCV, LogisticRegression, LogisticRegressionCV, MultiTaskLassoCV\nimport matplotlib.pyplot as plt\n\n%matplotlib inline",
"_____no_output_____"
]
],
[
[
"# 1. Example Usage with Average Treatment Effects",
"_____no_output_____"
],
[
"## 1.1 DGP\n\nWe consider a data generating process from a markovian treatment model. \n\nIn the example bellow, $T_t\\rightarrow$ treatment(s) at time $t$, $Y_t\\rightarrow$outcome at time $t$, $X_t\\rightarrow$ features and controls at time $t$ (the coefficients $e, f$ will pick the features and the controls).\n\\begin{align}\n X_t =& (\\pi'X_{t-1} + 1) \\cdot A\\, T_{t-1} + B X_{t-1} + \\epsilon_t\\\\\n T_t =& \\gamma\\, T_{t-1} + (1-\\gamma) \\cdot D X_t + \\zeta_t\\\\\n Y_t =& (\\sigma' X_{t} + 1) \\cdot e\\, T_{t} + f X_t + \\eta_t\n\\end{align}\n\nwith $X_0, T_0 = 0$ and $\\epsilon_t, \\zeta_t, \\eta_t \\sim N(0, \\sigma^2)$. Moreover, $X_t \\in R^{n_x}$, $B[:, 0:s_x] \\neq 0$ and $B[:, s_x:-1] = 0$, $\\gamma\\in [0, 1]$, $D[:, 0:s_x] \\neq 0$, $D[:, s_x:-1]=0$, $f[0:s_x]\\neq 0$, $f[s_x:-1]=0$. We draw a single time series of samples of length $n\\_panels \\cdot n\\_periods$.",
"_____no_output_____"
]
],
[
[
"# Define DGP parameters\nnp.random.seed(123)\nn_panels = 5000 # number of panels\nn_periods = 3 # number of time periods in each panel\nn_treatments = 2 # number of treatments in each period\nn_x = 100 # number of features + controls\ns_x = 10 # number of controls (endogeneous variables)\ns_t = 10 # treatment support size",
"_____no_output_____"
],
[
"# Generate data\ndgp = DynamicPanelDGP(n_periods, n_treatments, n_x).create_instance(\n s_x, random_seed=12345)\nY, T, X, W, groups = dgp.observational_data(n_panels, s_t=s_t, random_seed=12345)\ntrue_effect = dgp.true_effect",
"_____no_output_____"
]
],
[
[
"## 1.2 Train Estimator",
"_____no_output_____"
]
],
[
[
"est = DynamicDML(\n model_y=LassoCV(cv=3, max_iter=1000), \n model_t=MultiTaskLassoCV(cv=3, max_iter=1000), \n cv=3)",
"_____no_output_____"
],
[
"est.fit(Y, T, X=None, W=W, groups=groups)",
"_____no_output_____"
],
[
"# Average treatment effect of all periods on last period for unit treatments\nprint(f\"Average effect of default policy: {est.ate():0.2f}\")",
"Average effect of default policy: 1.40\n"
],
[
"# Effect of target policy over baseline policy\n# Must specify a treatment for each period\nbaseline_policy = np.zeros((1, n_periods * n_treatments))\ntarget_policy = np.ones((1, n_periods * n_treatments))\neff = est.effect(T0=baseline_policy, T1=target_policy)\nprint(f\"Effect of target policy over baseline policy: {eff[0]:0.2f}\")",
"Effect of target policy over baseline policy: 1.40\n"
],
[
"# Period treatment effects + interpretation\nfor i, theta in enumerate(est.intercept_.reshape(-1, n_treatments)):\n print(f\"Marginal effect of a treatments in period {i+1} on period {n_periods} outcome: {theta}\")",
"Marginal effect of a treatments in period 1 on period 3 outcome: [0.04000235 0.0701606 ]\nMarginal effect of a treatments in period 2 on period 3 outcome: [0.31611764 0.23714736]\nMarginal effect of a treatments in period 3 on period 3 outcome: [0.13108411 0.60656886]\n"
],
[
"# Period treatment effects with confidence intervals\nest.summary()",
"Coefficient Results: X is None, please call intercept_inference to learn the constant!\n"
],
[
"conf_ints = est.intercept__interval(alpha=0.05)",
"_____no_output_____"
]
],
[
[
"## 1.3 Performance Visualization",
"_____no_output_____"
]
],
[
[
"# Some plotting boilerplate code\nplt.figure(figsize=(15, 5))\nplt.errorbar(np.arange(n_periods*n_treatments)-.04, est.intercept_, yerr=(conf_ints[1] - est.intercept_,\n est.intercept_ - conf_ints[0]), fmt='o', label='DynamicDML')\nplt.errorbar(np.arange(n_periods*n_treatments), true_effect.flatten(), fmt='o', alpha=.6, label='Ground truth')\nfor t in np.arange(1, n_periods):\n plt.axvline(x=t * n_treatments - .5, linestyle='--', alpha=.4)\nplt.xticks([t * n_treatments - .5 + n_treatments/2 for t in range(n_periods)],\n [\"$\\\\theta_{}$\".format(t) for t in range(n_periods)])\nplt.gca().set_xlim([-.5, n_periods*n_treatments - .5])\nplt.ylabel(\"Effect\")\nplt.legend()\nplt.show()",
"_____no_output_____"
]
],
[
[
"# 2. Example Usage with Heterogeneous Treatment Effects on Time-Invariant Unit Characteristics\n\nWe can also estimate treatment effect heterogeneity with respect to the value of some subset of features $X$ in the initial period. Heterogeneity is currently only supported with respect to such initial state features. This for instance can support heterogeneity with respect to time-invariant unit characteristics. In that case you can simply pass as $X$ a repetition of some unit features that stay constant in all periods. You can also pass time-varying features, and their time varying component will be used as a time-varying control. However, heterogeneity will only be estimated with respect to the initial state.",
"_____no_output_____"
],
[
"## 2.1 DGP",
"_____no_output_____"
]
],
[
[
"# Define additional DGP parameters\nhet_strength = .5\nhet_inds = np.arange(n_x - n_treatments, n_x)",
"_____no_output_____"
],
[
"# Generate data\ndgp = DynamicPanelDGP(n_periods, n_treatments, n_x).create_instance(\n s_x, hetero_strength=het_strength, hetero_inds=het_inds, random_seed=12)\nY, T, X, W, groups = dgp.observational_data(n_panels, s_t=s_t, random_seed=1)\nate_effect = dgp.true_effect\nhet_effect = dgp.true_hetero_effect[:, het_inds + 1]",
"_____no_output_____"
]
],
[
[
"## 2.2 Train Estimator",
"_____no_output_____"
]
],
[
[
"est = DynamicDML(\n model_y=LassoCV(cv=3), \n model_t=MultiTaskLassoCV(cv=3), \n cv=3)",
"_____no_output_____"
],
[
"est.fit(Y, T, X=X, W=W, groups=groups, inference=\"auto\")",
"_____no_output_____"
],
[
"est.summary()",
"_____no_output_____"
],
[
"# Average treatment effect for test points\nX_test = X[np.arange(0, 25, 3)]\nprint(f\"Average effect of default policy:{est.ate(X=X_test):0.2f}\")",
"Average effect of default policy:-0.42\n"
],
[
"# Effect of target policy over baseline policy\n# Must specify a treatment for each period\nbaseline_policy = np.zeros((1, n_periods * n_treatments))\ntarget_policy = np.ones((1, n_periods * n_treatments))\neff = est.effect(X=X_test, T0=baseline_policy, T1=target_policy)\nprint(\"Effect of target policy over baseline policy for test set:\\n\", eff)",
"Effect of target policy over baseline policy for test set:\n [-0.37368525 -0.30896804 -0.43030363 -0.52252401 -0.42849622 -0.48790877\n -0.34417987 -0.51804937 -0.36806744]\n"
],
[
"# Coefficients: intercept is of shape n_treatments*n_periods\n# coef_ is of shape (n_treatments*n_periods, n_hetero_inds).\n# first n_treatment rows are from first period, next n_treatment\n# from second period, etc.\nest.intercept_, est.coef_",
"_____no_output_____"
],
[
"# Confidence intervals\nconf_ints_intercept = est.intercept__interval(alpha=0.05)\nconf_ints_coef = est.coef__interval(alpha=0.05)",
"_____no_output_____"
]
],
[
[
"## 2.3 Performance Visualization",
"_____no_output_____"
]
],
[
[
"# parse true parameters in array of shape (n_treatments*n_periods, 1 + n_hetero_inds)\n# first column is the intercept\ntrue_effect_inds = []\nfor t in range(n_treatments):\n true_effect_inds += [t * (1 + n_x)] + (list(t * (1 + n_x) + 1 + het_inds) if len(het_inds)>0 else [])\ntrue_effect_params = dgp.true_hetero_effect[:, true_effect_inds]\ntrue_effect_params = true_effect_params.reshape((n_treatments*n_periods, 1 + het_inds.shape[0]))",
"_____no_output_____"
],
[
"# concatenating intercept and coef_\nparam_hat = np.hstack([est.intercept_.reshape(-1, 1), est.coef_])\nlower = np.hstack([conf_ints_intercept[0].reshape(-1, 1), conf_ints_coef[0]])\nupper = np.hstack([conf_ints_intercept[1].reshape(-1, 1), conf_ints_coef[1]])",
"_____no_output_____"
],
[
"plt.figure(figsize=(15, 5))\nplt.errorbar(np.arange(n_periods * (len(het_inds) + 1) * n_treatments),\n true_effect_params.flatten(), fmt='*', label='Ground Truth')\nplt.errorbar(np.arange(n_periods * (len(het_inds) + 1) * n_treatments),\n param_hat.flatten(), yerr=((upper - param_hat).flatten(),\n (param_hat - lower).flatten()), fmt='o', label='DynamicDML')\nadd_vlines(n_periods, n_treatments, het_inds)\nplt.legend()\nplt.show()",
"_____no_output_____"
]
]
] | [
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] | [
[
"markdown",
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
]
] |
d0fdffe18daa9d815c3859e23cb8eebfb8641887 | 50,493 | ipynb | Jupyter Notebook | HeroesOfPymoli/HeroesOfPymoli_starter.ipynb | juliedundas/pandas-challenge | cce395f2a96a03d0ace171cf3ec56bc6769d8ed4 | [
"ADSL"
] | null | null | null | HeroesOfPymoli/HeroesOfPymoli_starter.ipynb | juliedundas/pandas-challenge | cce395f2a96a03d0ace171cf3ec56bc6769d8ed4 | [
"ADSL"
] | null | null | null | HeroesOfPymoli/HeroesOfPymoli_starter.ipynb | juliedundas/pandas-challenge | cce395f2a96a03d0ace171cf3ec56bc6769d8ed4 | [
"ADSL"
] | null | null | null | 44.021796 | 655 | 0.516884 | [
[
[
"### Note\n* Instructions have been included for each segment. You do not have to follow them exactly, but they are included to help you think through the steps.",
"_____no_output_____"
]
],
[
[
"# Dependencies and Setup\nimport pandas as pd\n\n# File to Load (Remember to Change These)\nfile_to_load = \"Resources/purchase_data.csv\"\n\n# Read Purchasing File and store into Pandas data frame\npurchase_data = pd.read_csv(file_to_load)\npurchase_data.head()",
"_____no_output_____"
]
],
[
[
"## Player Count",
"_____no_output_____"
],
[
"* Display the total number of players\n",
"_____no_output_____"
]
],
[
[
"total_unique_players=purchase_data['SN'].nunique\n#total_unique_players()\ntotal_players_df = pd.DataFrame({\"Total Players\": [total_unique_players()]})\ntotal_players_df",
"_____no_output_____"
]
],
[
[
"## Purchasing Analysis (Total)",
"_____no_output_____"
],
[
"* Run basic calculations to obtain number of unique items, average price, etc.\n\n\n* Create a summary data frame to hold the results\n\n\n* Optional: give the displayed data cleaner formatting\n\n\n* Display the summary data frame\n",
"_____no_output_____"
]
],
[
[
"unique_items=purchase_data['Item ID'].nunique()\n#unique_items\naverage_price=purchase_data['Price'].mean()\n#average_price\ntotal_purchases=purchase_data['SN'].count()\n#total_purchases\ntotal_revenue=purchase_data['Price'].sum()\n#total_revenue\nbasic_calculations_df=pd.DataFrame({'Number of Unique Items': [unique_items], 'Average Price' : [average_price], 'Number of Purchases' : [total_purchases], 'Total Revenue' : [total_revenue]})\nformat_dict=({'Average Price':\"${:,.2f}\",\n 'Total Revenue': '${:,.2f}'})\nbasic_calculations_df.style.format(format_dict)",
"_____no_output_____"
]
],
[
[
"## Gender Demographics",
"_____no_output_____"
],
[
"* Percentage and Count of Male Players\n\n\n* Percentage and Count of Female Players\n\n\n* Percentage and Count of Other / Non-Disclosed\n\n\n",
"_____no_output_____"
]
],
[
[
"gender_df=purchase_data.groupby('Gender')\ngender_df\n\ntotal_gender_count =gender_df.nunique()['SN']\ntotal_gender_count\n\npercentage_of_players = total_gender_count / total_unique_players() * 100\npercentage_of_players\n\nfinal_gender_df = pd.DataFrame({'Total Count': total_gender_count, 'Percentage of Players': percentage_of_players})\n\nfinal_gender_df.index.name = None\n\nfinal_gender_df.sort_values(['Total Count'], ascending = True)\n\nfinal_gender_df['Percentage of Players']=(percentage_of_players).round(2).astype(str) + '%'\nfinal_gender_df\n\n\n##DID NOT USE THE CODE BELOW:\n\n#gender_count = SN_and_gender_no_dups.count()\n#gender_count\n\n#only_males = SN_and_gender_no_dups.loc[SN_and_gender['Gender'] == 'Male', :]\n#only_males\n\n#only_females = SN_and_gender_no_dups.loc[SN_and_gender['Gender'] == 'Female', :].count()\n#only_females\n\n#only_non_disclosed = SN_and_gender_no_dups.loc[SN_and_gender['Gender'] == 'Other / Non-Disclosed', :].count()\n#only_non_disclosed\n\n",
"_____no_output_____"
]
],
[
[
"\n## Purchasing Analysis (Gender)",
"_____no_output_____"
],
[
"* Run basic calculations to obtain purchase count, avg. purchase price, avg. purchase total per person etc. by gender\n\n\n\n\n* Create a summary data frame to hold the results\n\n\n* Optional: give the displayed data cleaner formatting\n\n\n* Display the summary data frame",
"_____no_output_____"
]
],
[
[
"#Create dataframe to complete gender analysis\ngender_analysis = gender_df[['SN','Gender', 'Age','Purchase ID', 'Price']]\ngender_analysis\n\n#Create variables to hold values for purchase count avg price by gender, purchase value total by gender, and average total price per person\ntotal_purchase_count=gender_df['Purchase ID'].count()\ntotal_purchase_count\n\naverage_price=gender_df['Price'].mean()\naverage_price\n\npurchase_value_total=gender_df['Price'].sum()\npurchase_value_total\n\navg_price_by_person= purchase_value_total / total_gender_count\navg_price_by_person\n\ngender_purchasing_analysis_df = pd.DataFrame({'Purchase Count': total_purchase_count, 'Average Purchase Price': average_price, 'Total Purchase Value': purchase_value_total,'Avg Total Purchase per Person': avg_price_by_person})\ngender_purchasing_analysis_df \n\n\n#Style the dataframe and add the Gender index\ngender_purchasing_analysis_df.index.name = \"Gender\"\n\nformat_gender_dict = {'Average Purchase Price': '${:,.2f}','Total Purchase Value':'${:,.2f}', 'Avg Total Purchase per Person':'${:,.2f}'}\ngender_purchasing_analysis_df.style.format(format_gender_dict)\n",
"_____no_output_____"
]
],
[
[
"## Age Demographics",
"_____no_output_____"
],
[
"* Establish bins for ages\n\n\n* Categorize the existing players using the age bins. Hint: use pd.cut()\n\n\n* Calculate the numbers and percentages by age group\n\n\n* Create a summary data frame to hold the results\n\n\n* Optional: round the percentage column to two decimal points\n\n\n* Display Age Demographics Table\n",
"_____no_output_____"
]
],
[
[
"#Create age DF to work with data\n#age_df=purchase_data.groupby['Age']\n#age_df\n\n# Create bins in which to place values based upon ages of users\nbins = [0, 9.99, 14.99, 19.99, 24.99,29.99, 34.99, 39.99 ,99.99]\n\n# Create labels for these bins\ngroup_labels = ['<10', '10 to 14', '15 to 19','20 to 24', '25 to 29', '30 to 34', '35 to 39', '40+']\n\n#Sort data into correct bins\npurchase_data['Age Group'] = pd.cut(purchase_data['Age'], bins, labels=group_labels)\npurchase_data.head()\n\ngroup_ages=purchase_data.groupby('Age Group')\ngroup_ages\n\n#Create variables to hold values for total count and percentage of players\nplayers_by_ages_count=group_ages['SN'].nunique()\nplayers_by_ages_count\n\nplayer_percentages=players_by_ages_count/total_unique_players() * 100\nplayer_percentages\n\n#Create the new dataframe to show the age demographics analysis\nage_demographics_df=pd.DataFrame({'Total Count': players_by_ages_count, 'Percentage of Players': player_percentages})\nage_demographics_df\n\n#Format the percentage column\nage_demographics_df['Percentage of Players']=(player_percentages).round(2).astype(str) + '%'\nage_demographics_df",
"_____no_output_____"
]
],
[
[
"## Purchasing Analysis (Age)",
"_____no_output_____"
],
[
"* Bin the purchase_data data frame by age\n\n\n* Run basic calculations to obtain purchase count, avg. purchase price, avg. purchase total per person etc. in the table below\n\n\n* Create a summary data frame to hold the results\n\n\n* Optional: give the displayed data cleaner formatting\n\n\n* Display the summary data frame",
"_____no_output_____"
]
],
[
[
"#Create variables for the purchasing analysis for Purchase Count, Average Purchase Price, Total Purchase Value, Avg Total Purchase per Person\nage_purchase_count=group_ages['Purchase ID'].count()\nage_purchase_count\n\nage_avg_purchase_price=group_ages['Price'].mean()\nage_avg_purchase_price\n\nage_purchase_value=group_ages['Price'].sum()\nage_purchase_value\n\nage_avg_total_purchase=age_purchase_value/players_by_ages_count\nage_avg_total_purchase\n\n#Create dataframe for Purchasing Analysis (Age)\nage_purchasing_analysis_df=pd.DataFrame({'Purchase Count': age_purchase_count, 'Average Purchase Price': age_avg_purchase_price, 'Total Purchase Value': age_purchase_value,'Avg Total Purchase per Person':age_avg_total_purchase})\nage_purchasing_analysis_df\n\n\nformat_age_dict = {'Average Purchase Price': '${:,.2f}','Total Purchase Value':'${:,.2f}', 'Avg Total Purchase per Person':'${:,.2f}'}\nage_purchasing_analysis_df.style.format(format_age_dict)",
"_____no_output_____"
]
],
[
[
"## Top Spenders\n\n",
"_____no_output_____"
],
[
"* Run basic calculations to obtain the results in the table below\n\n\n* Create a summary data frame to hold the results\n\n\n* Sort the total purchase value column in descending order\n\n\n* Optional: give the displayed data cleaner formatting\n\n\n* Display a preview of the summary data frame\n\n",
"_____no_output_____"
]
],
[
[
"#Use the DF above to find the top spenders\n#Start by grouping results by SN\nSN_group=purchase_data.groupby('SN')\nSN_group\n\n#Create varialbe to hold the results that will fill the summary table\npurchase_count=SN_group['Purchase ID'].count()\npurchase_count\n\navg_purchase_price_by_SN=SN_group['Price'].mean().round(2)\navg_purchase_price_by_SN\n\ntotal_purchase_value_by_SN=SN_group['Price'].sum()\ntotal_purchase_value_by_SN\n\n#Create a data frame to hold the results\ntop_spenders_df=pd.DataFrame({'Purchase Count': purchase_count, 'Average Purchase Price': avg_purchase_price_by_SN, 'Total Purchase Value':total_purchase_value_by_SN})\ntop_spenders_df\n\n#Sort the data frame in descending order of total purchase price\ntop_spenders_df_sorted=top_spenders_df.sort_values(['Total Purchase Value'], ascending=False).head()\ntop_spenders_df_sorted\n\n#Format the df with current symbols\nformat_spenders_dict = {'Average Purchase Price': '${:,.2f}','Total Purchase Value':'${:,.2f}'}\ntop_spenders_df_sorted.style.format(format_spenders_dict)",
"_____no_output_____"
]
],
[
[
"## Most Popular Items",
"_____no_output_____"
],
[
"* Retrieve the Item ID, Item Name, and Item Price columns\n\n\n* Group by Item ID and Item Name. Perform calculations to obtain purchase count, item price, and total purchase value\n\n\n* Create a summary data frame to hold the results\n\n\n* Sort the purchase count column in descending order\n\n\n* Optional: give the displayed data cleaner formatting\n\n\n* Display a preview of the summary data frame\n\n",
"_____no_output_____"
]
],
[
[
"#Create a new data frame to retrieve Item ID, Item Name, and Item Price columns\npopular_items=purchase_data[['Item ID', 'Item Name', 'Price']]\npopular_items\n\n#Create a groupby to hold the results\ngrouped_data=popular_items.groupby(['Item ID', 'Item Name'])\ngrouped_data\n\n#Complete basic calculations\npurchase_count_popular=grouped_data['Price'].count()\npurchase_count_popular\n\navg_item_price_popular=grouped_data['Price'].mean()\navg_item_price_popular\n\ntotal_purchase_value_popular=grouped_data['Price'].sum()\ntotal_purchase_value_popular\n\n\n#Create a summary data frame to hold the results\nmost_popular_items_df=pd.DataFrame({'Purchase Count': purchase_count_popular, \n 'Item Price': avg_item_price_popular,\n 'Total Purchase Value':total_purchase_value_popular})\nmost_popular_items_df\n\n#Sort the purchase count column in descending order\nmost_popular_items_df_sorted=most_popular_items_df.sort_values(['Purchase Count'], ascending=False).head()\nmost_popular_items_df_sorted\n\n#Give currency formatting to the Item Price and Total Purchase Value Column\nformat_popular_dict={'Item Price': '${:,.2f}','Total Purchase Value':'${:,.2f}'}\nmost_popular_items_df_sorted.style.format(format_popular_dict)",
"_____no_output_____"
]
],
[
[
"## Most Profitable Items",
"_____no_output_____"
],
[
"* Sort the above table by total purchase value in descending order\n\n\n* Optional: give the displayed data cleaner formatting\n\n\n* Display a preview of the data frame\n\n",
"_____no_output_____"
]
],
[
[
"#Sort the above table by total purchase value in descending order\nmost_popular_items_df_sorted_pv=most_popular_items_df.sort_values(['Total Purchase Value'], ascending=False).head()\nmost_popular_items_df_sorted_pv\n\n#Give currency formatting to the Item Price and Total Purchase Value Column\nmost_popular_items_df_sorted_pv.style.format(format_popular_dict)",
"_____no_output_____"
]
]
] | [
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] | [
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
]
] |
d0fe020c352860291238b161aa2b1c9c2294bd3d | 4,628 | ipynb | Jupyter Notebook | ipynb/Saint-Lucia.ipynb | skirienko/oscovida.github.io | eda5412d02365a8a000239be5480512c53bee8c2 | [
"CC-BY-4.0"
] | null | null | null | ipynb/Saint-Lucia.ipynb | skirienko/oscovida.github.io | eda5412d02365a8a000239be5480512c53bee8c2 | [
"CC-BY-4.0"
] | null | null | null | ipynb/Saint-Lucia.ipynb | skirienko/oscovida.github.io | eda5412d02365a8a000239be5480512c53bee8c2 | [
"CC-BY-4.0"
] | null | null | null | 28.392638 | 165 | 0.508427 | [
[
[
"# Saint Lucia\n\n* Homepage of project: https://oscovida.github.io\n* Plots are explained at http://oscovida.github.io/plots.html\n* [Execute this Jupyter Notebook using myBinder](https://mybinder.org/v2/gh/oscovida/binder/master?filepath=ipynb/Saint-Lucia.ipynb)",
"_____no_output_____"
]
],
[
[
"import datetime\nimport time\n\nstart = datetime.datetime.now()\nprint(f\"Notebook executed on: {start.strftime('%d/%m/%Y %H:%M:%S%Z')} {time.tzname[time.daylight]}\")",
"_____no_output_____"
],
[
"%config InlineBackend.figure_formats = ['svg']\nfrom oscovida import *",
"_____no_output_____"
],
[
"overview(\"Saint Lucia\", weeks=5);",
"_____no_output_____"
],
[
"overview(\"Saint Lucia\");",
"_____no_output_____"
],
[
"compare_plot(\"Saint Lucia\", normalise=True);\n",
"_____no_output_____"
],
[
"# load the data\ncases, deaths = get_country_data(\"Saint Lucia\")\n\n# compose into one table\ntable = compose_dataframe_summary(cases, deaths)\n\n# show tables with up to 500 rows\npd.set_option(\"max_rows\", 500)\n\n# display the table\ntable",
"_____no_output_____"
]
],
[
[
"# Explore the data in your web browser\n\n- If you want to execute this notebook, [click here to use myBinder](https://mybinder.org/v2/gh/oscovida/binder/master?filepath=ipynb/Saint-Lucia.ipynb)\n- and wait (~1 to 2 minutes)\n- Then press SHIFT+RETURN to advance code cell to code cell\n- See http://jupyter.org for more details on how to use Jupyter Notebook",
"_____no_output_____"
],
[
"# Acknowledgements:\n\n- Johns Hopkins University provides data for countries\n- Robert Koch Institute provides data for within Germany\n- Atlo Team for gathering and providing data from Hungary (https://atlo.team/koronamonitor/)\n- Open source and scientific computing community for the data tools\n- Github for hosting repository and html files\n- Project Jupyter for the Notebook and binder service\n- The H2020 project Photon and Neutron Open Science Cloud ([PaNOSC](https://www.panosc.eu/))\n\n--------------------",
"_____no_output_____"
]
],
[
[
"print(f\"Download of data from Johns Hopkins university: cases at {fetch_cases_last_execution()} and \"\n f\"deaths at {fetch_deaths_last_execution()}.\")",
"_____no_output_____"
],
[
"# to force a fresh download of data, run \"clear_cache()\"",
"_____no_output_____"
],
[
"print(f\"Notebook execution took: {datetime.datetime.now()-start}\")\n",
"_____no_output_____"
]
]
] | [
"markdown",
"code",
"markdown",
"code"
] | [
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code",
"code"
]
] |
d0fe23c30e6086f8b4e0215a41a6b4bf2dc448a3 | 21,957 | ipynb | Jupyter Notebook | Lab-2/4feature-selection-correlation.ipynb | yash-a-18/002_YashAmethiya | ab7e8bd8ebec553a0592b698dddc34c53b522967 | [
"MIT"
] | null | null | null | Lab-2/4feature-selection-correlation.ipynb | yash-a-18/002_YashAmethiya | ab7e8bd8ebec553a0592b698dddc34c53b522967 | [
"MIT"
] | null | null | null | Lab-2/4feature-selection-correlation.ipynb | yash-a-18/002_YashAmethiya | ab7e8bd8ebec553a0592b698dddc34c53b522967 | [
"MIT"
] | null | null | null | 21,957 | 21,957 | 0.736485 | [
[
[
"<a id=\"1\"></a> \n## Correlation\n<a id=\"1-1\"></a>\n### What is correlation?\nCorrelation is a statistical term which in common usage refers to how close two variables are to having a linear relationship with each other.\n\nFor example, two variables which are linearly dependent (say, **x** and **y** which depend on each other as x = 2y) will have a higher correlation than two variables which are non-linearly dependent (say, u and v which depend on each other as u = v2)\n<a id=\"1-2\"></a>\n### How does correlation help in feature selection?\nFeatures with high correlation are more linearly dependent and hence have almost the same effect on the dependent variable. So, when two features have high correlation, we can drop one of the two features.",
"_____no_output_____"
],
[
"### Import the necessary libraries",
"_____no_output_____"
]
],
[
[
"import numpy as np\r\nimport pandas as pd\r\nimport seaborn as sns",
"_____no_output_____"
]
],
[
[
"### Loading the dataset",
"_____no_output_____"
]
],
[
[
"data = pd.read_csv('./Datasets/Data_for_Correlation.csv')",
"_____no_output_____"
],
[
"data.head()",
"_____no_output_____"
]
],
[
[
"Removing the Class Label entry (Y)",
"_____no_output_____"
]
],
[
[
"data = data.iloc[:,:-1]\r\ndata.head()",
"_____no_output_____"
],
[
"data.info()",
"<class 'pandas.core.frame.DataFrame'>\nRangeIndex: 14 entries, 0 to 13\nData columns (total 4 columns):\n # Column Non-Null Count Dtype\n--- ------ -------------- -----\n 0 X1 14 non-null int64\n 1 X2 14 non-null int64\n 2 X3 14 non-null int64\n 3 X4 14 non-null int64\ndtypes: int64(4)\nmemory usage: 576.0 bytes\n"
]
],
[
[
"<a id=\"3-2\"></a>\n## Selecting features based on correlation\nGenerating the correlation matrix",
"_____no_output_____"
]
],
[
[
"corr = data.corr()\ncorr.head()",
"_____no_output_____"
]
],
[
[
"Generating the correlation heatmap",
"_____no_output_____"
]
],
[
[
"sns.heatmap(corr)",
"_____no_output_____"
]
],
[
[
"Next, we compare the correlation between features and remove one of two features that have a correlation higher than 0.9",
"_____no_output_____"
]
],
[
[
"columns = np.full((corr.shape[0],), True, dtype=bool)\nfor i in range(corr.shape[0]):\n for j in range(i+1, corr.shape[0]):\n if corr.iloc[i,j] >= 0.9:\n if columns[j]:\n columns[j] = False",
"_____no_output_____"
],
[
"selected_columns = data.columns[columns]\nselected_columns.shape",
"_____no_output_____"
],
[
"data = data[selected_columns]\nprint(data)",
" X1 X4\n0 1 -2\n1 2 -4\n2 3 3\n3 4 4\n4 5 25\n5 6 76\n6 7 34\n7 8 346\n8 9 67\n9 10 3\n10 11 355\n11 12 88\n12 13 2\n13 14 1\n"
]
]
] | [
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] | [
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
]
] |
d0fe2fc0a79a8d5fbe384cbb666d25d8f4a2e8ac | 41,377 | ipynb | Jupyter Notebook | 59_hazard.Cox.ipynb | sachinruk/torchlife2 | fc5217a9f48ff22f7c26126fa9a473f63c328bc0 | [
"Apache-2.0"
] | 10 | 2019-10-21T01:19:18.000Z | 2022-03-26T17:03:07.000Z | 59_hazard.Cox.ipynb | sachinruk/torchlife2 | fc5217a9f48ff22f7c26126fa9a473f63c328bc0 | [
"Apache-2.0"
] | 4 | 2020-11-30T04:53:30.000Z | 2022-02-26T06:18:42.000Z | 59_hazard.Cox.ipynb | sachinruk/torchlife2 | fc5217a9f48ff22f7c26126fa9a473f63c328bc0 | [
"Apache-2.0"
] | 1 | 2021-06-23T15:24:03.000Z | 2021-06-23T15:24:03.000Z | 83.253521 | 19,708 | 0.782995 | [
[
[
"# default_exp models.cox",
"_____no_output_____"
]
],
[
[
"# Cox Proportional Hazard\n> SA with features apart from time\n\nWe model the the instantaneous hazard as the product of two functions, one with the time component, and the other with the feature component.\n$$\n\\begin{aligned}\n\\lambda(t,x) = \\lambda(t)h(x)\n\\end{aligned}\n$$\n\nIt is important to have the seperation of these functions to arrive at an analytical solution. This is so that the time component can be integrated out to give the survival function.\n\n$$\n\\begin{aligned}\n\\int_0^T \\lambda(t,x) dt &= \\int_0^T \\lambda(t)h(x) dt\\\\\n&= h(x)\\int_0^T \\lambda(t) dt\\\\\nS(t) &= \\exp\\left(-h(x)\\int_{-\\infty}^t \\lambda(\\tau) d\\tau\\right)\n\\end{aligned}\n$$",
"_____no_output_____"
]
],
[
[
"# export\nimport matplotlib.pyplot as plt\nimport numpy as np\nimport torch\nimport torch.nn as nn\nimport torch.nn.functional as F\nfrom sklearn.preprocessing import MaxAbsScaler, StandardScaler\n\nfrom torchlife.losses import hazard_loss\nfrom torchlife.models.ph import PieceWiseHazard\n\n# torch.Tensor.ndim = property(lambda x: x.dim())",
"_____no_output_____"
],
[
"# hide\n%load_ext autoreload\n%autoreload 2\n%matplotlib inline",
"_____no_output_____"
],
[
"# export\nclass ProportionalHazard(nn.Module):\n \"\"\"\n Hazard proportional to time and feature component as shown above.\n parameters:\n - breakpoints: time points where hazard would change\n - max_t: maximum point of time to plot to.\n - dim: number of input dimensions of x\n - h: (optional) number of hidden units (for x only).\n \"\"\"\n def __init__(self, breakpoints:np.array, t_scaler:MaxAbsScaler, x_scaler:StandardScaler, \n dim:int, h:tuple=(), **kwargs):\n super().__init__()\n self.baseλ = PieceWiseHazard(breakpoints, t_scaler)\n self.x_scaler = x_scaler\n nodes = (dim,) + h + (1,)\n self.layers = nn.ModuleList([nn.Linear(a,b, bias=False) \n for a,b in zip(nodes[:-1], nodes[1:])])\n \n def forward(self, t, t_section, x):\n logλ, Λ = self.baseλ(t, t_section)\n \n for layer in self.layers[:-1]:\n x = F.relu(layer(x))\n log_hx = self.layers[-1](x)\n \n logλ += log_hx\n Λ = torch.exp(log_hx + torch.log(Λ))\n return logλ, Λ\n \n def survival_function(self, t:np.array, x:np.array) -> torch.Tensor:\n if len(t.shape) == 1:\n t = t[:,None]\n t = self.baseλ.t_scaler.transform(t)\n if len(x.shape) == 1:\n x = x[None, :]\n if len(x) == 1:\n x = np.repeat(x, len(t), axis=0)\n x = self.x_scaler.transform(x)\n \n \n with torch.no_grad():\n x = torch.Tensor(x)\n # get the times and time sections for survival function\n breakpoints = self.baseλ.breakpoints[1:].cpu().numpy()\n t_sec_query = np.searchsorted(breakpoints.squeeze(), t.squeeze())\n # convert to pytorch tensors\n t_query = torch.Tensor(t)\n t_sec_query = torch.LongTensor(t_sec_query)\n\n # calculate cumulative hazard according to above\n _, Λ = self.forward(t_query, t_sec_query, x)\n return torch.exp(-Λ)\n \n \n def plot_survival_function(self, t:np.array, x:np.array) -> None:\n s = self.survival_function(t, x)\n \n # plot\n plt.figure(figsize=(12,5))\n plt.plot(t, s)\n plt.xlabel('Time')\n plt.ylabel('Survival Probability')\n plt.show()",
"_____no_output_____"
]
],
[
[
"## Fitting Cox Proportional Hazard Model",
"_____no_output_____"
]
],
[
[
"# hide\nfrom torchlife.data import create_db, get_breakpoints\nimport pandas as pd",
"_____no_output_____"
],
[
"# hide\nurl = \"https://raw.githubusercontent.com/CamDavidsonPilon/lifelines/master/lifelines/datasets/rossi.csv\"\ndf = pd.read_csv(url)\ndf.head()",
"_____no_output_____"
],
[
"# hide\ndf.rename(columns={'week':'t', 'arrest':'e'}, inplace=True)\nbreakpoints = get_breakpoints(df)\ndb, t_scaler, x_scaler = create_db(df, breakpoints)",
"_____no_output_____"
],
[
"# hide\nfrom fastai.basics import Learner\n\nx_dim = df.shape[1] - 2\nmodel = ProportionalHazard(breakpoints, t_scaler, x_scaler, x_dim, h=(3,3))\nlearner = Learner(db, model, loss_func=hazard_loss)\n# wd = 1e-4\n# learner.lr_find()\n# learner.recorder.plot()",
"_____no_output_____"
],
[
"# hide\nepochs = 10\nlearner.fit(epochs, lr=1)",
"_____no_output_____"
]
],
[
[
"## Plotting hazard functions",
"_____no_output_____"
]
],
[
[
"model.baseλ.plot_hazard()\nx = df.drop(['t', 'e'], axis=1).iloc[4]\nt = np.arange(df['t'].max())\nmodel.plot_survival_function(t, x)",
"_____no_output_____"
],
[
"# hide\nfrom nbdev.export import *\nnotebook2script()",
"Converted 00_index.ipynb.\nConverted 10_SAT.ipynb.\nConverted 20_KaplanMeier.ipynb.\nConverted 50_hazard.ipynb.\nConverted 55_hazard.PiecewiseHazard.ipynb.\nConverted 59_hazard.Cox.ipynb.\nConverted 60_AFT_models.ipynb.\nConverted 65_AFT_error_distributions.ipynb.\nConverted 80_data.ipynb.\nConverted 90_model.ipynb.\nConverted 95_Losses.ipynb.\n"
]
]
] | [
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] | [
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
]
] |
d0fe3a1f9bed38bdf8124bf5bf379dfb93d236f6 | 4,670 | ipynb | Jupyter Notebook | examples/notebooks/08_Net_CDF_file.ipynb | schluchc/python-connector-api | 79586811b73f866c872db9c2404cf03dfddddf79 | [
"MIT"
] | null | null | null | examples/notebooks/08_Net_CDF_file.ipynb | schluchc/python-connector-api | 79586811b73f866c872db9c2404cf03dfddddf79 | [
"MIT"
] | null | null | null | examples/notebooks/08_Net_CDF_file.ipynb | schluchc/python-connector-api | 79586811b73f866c872db9c2404cf03dfddddf79 | [
"MIT"
] | null | null | null | 23.705584 | 221 | 0.568737 | [
[
[
"# NetCDF file",
"_____no_output_____"
],
[
"First you have to import the meteomatics module and the datetime module",
"_____no_output_____"
]
],
[
[
"import datetime as dt\nimport meteomatics.api as api",
"_____no_output_____"
]
],
[
[
"Input here your username and password from your meteomatics profile",
"_____no_output_____"
]
],
[
[
"###Credentials:\nusername = 'python-community'\npassword = 'Umivipawe179'",
"_____no_output_____"
]
],
[
[
"Input here the limiting coordinates of the extract you want to look at. You can also change the resolution.",
"_____no_output_____"
]
],
[
[
"lat_N = 50\nlon_W = -16\nlat_S = 20\nlon_E = 10\nres_lat = 2\nres_lon = 2",
"_____no_output_____"
]
],
[
[
"Input here the directory and the name. The directory must already exist.",
"_____no_output_____"
]
],
[
[
"filename_nc = 'path_netcdf/netcdf_file.nc'",
"_____no_output_____"
]
],
[
[
"Input here the startdate, enddate and the timeinterval as datetime objects. ",
"_____no_output_____"
]
],
[
[
"startdate_nc = dt.datetime.utcnow().replace(hour=0, minute=0, second=0, microsecond=0)\nenddate_nc = startdate_nc + dt.timedelta(days=3)\ninterval_nc = dt.timedelta(hours=12)",
"_____no_output_____"
]
],
[
[
"Choose the parameter you want to get. You can only chose one parameter at a time. Check here which parameters are available: https://www.meteomatics.com/en/api/available-parameters/",
"_____no_output_____"
]
],
[
[
"parameter_nc = 't_2m:C'",
"_____no_output_____"
]
],
[
[
"In the following, the request will start. If there is an error in the request as for example a wrong parameter or a date that doesn't exist, you get a message.",
"_____no_output_____"
]
],
[
[
"print(\"netCDF file:\")\ntry:\n api.query_netcdf(filename_nc, startdate_nc, enddate_nc, interval_nc, parameter_nc, lat_N, lon_W, lat_S,\n lon_E, res_lat, res_lon, username, password)\n print(\"filename = {}\".format(filename_nc))\nexcept Exception as e:\n print(\"Failed, the exception is {}\".format(e))",
"\nnetCDF file:\n2019-04-10 10:04:39| INFO |Calling URL: https://api.meteomatics.com/2019-04-10T00:00:00+00:00--2019-04-13T00:00:00+00:00:PT12H/t_2m:C/50,-16_20,10:2,2/netcdf?connector=python_2.0.1 (username = python-community)\n2019-04-10 10:04:39| INFO |Create Path: path_netcdf\n2019-04-10 10:04:39| INFO |Create File path_netcdf/netcdf_file.nc\nfilename = path_netcdf/netcdf_file.nc\n"
]
],
[
[
"You will get the data as a NetCDF file. This is a common file format to share climatological data. You need to have a special program to be able to visualize it, as shown here.\n\n",
"_____no_output_____"
]
]
] | [
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown"
] | [
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
]
] |
d0fe570d2cb2d53f30ad399fb3f357efbf892f83 | 16,195 | ipynb | Jupyter Notebook | notebooks/other/Deedle/rinterop.ipynb | SpaceAntelope/IfCntk | 81868163c8c7ba52b8646b7c0baae9a56e006ade | [
"MIT"
] | 5 | 2019-01-31T13:18:24.000Z | 2021-11-15T10:43:13.000Z | notebooks/other/Deedle/rinterop.ipynb | SpaceAntelope/IfCntk | 81868163c8c7ba52b8646b7c0baae9a56e006ade | [
"MIT"
] | null | null | null | notebooks/other/Deedle/rinterop.ipynb | SpaceAntelope/IfCntk | 81868163c8c7ba52b8646b7c0baae9a56e006ade | [
"MIT"
] | null | null | null | 54.52862 | 1,703 | 0.572584 | [
[
[
"(*** hide ***)",
"_____no_output_____"
]
],
[
[
"\n#nowarn \"211\"\nopen System\nlet airQuality = __SOURCE_DIRECTORY__ + \"/data/airquality.csv\"\n\n",
"_____no_output_____"
]
],
[
[
"(**\n\nInteroperating between R and Deedle\n===================================\n\nThe [R type provider](http://fslab.org/RProvider/) enables\nsmooth interoperation between R and F#. The type provider automatically discovers \ninstalled packages and makes them accessible via the `RProvider` namespace.\n\nR type provider for F# automatically converts standard data structures betwene R\nand F# (such as numerical values, arrays, etc.). However, the conversion mechanism\nis extensible and so it is possible to support conversion between other F# types.\n\nThe Deedle library comes with extension that automatically converts between Deedle\n`Frame<R, C>` and R `data.frame` and also between Deedle `Series<K, V>` and the\n[zoo package](http://cran.r-project.org/web/packages/zoo/index.html) (Z's ordered \nobservations).\n\nThis page is a quick overview showing how to pass data between R and Deedle.\nYou can also get this page as an [F# script file](https://github.com/fslaborg/Deedle/blob/master/docs/content/rinterop.fsx)\nfrom GitHub and run the samples interactively.\n\n<a name=\"setup\"></a>\n\n\nGetting started\n---------------\n\nTo use Deedle and R provider together, all you need to do is to install the \n[**Deedle.RPlugin** package](https://nuget.org/packages/Deedle.RPlugin), which\ninstalles both as dependencies. Alternatively, you can use the [**FsLab**\npackage](http://www.nuget.org/packages/FsLab), which also includes additional\ndata access, data science and visualization libraries.\n\nIn a typical project (\"F# Tutorial\"), the NuGet packages are installed in the `../packages`\ndirectory. To use R provider and Deedle, you need to write something like this:\n*)",
"_____no_output_____"
]
],
[
[
"\n#load \"../../packages/RProvider/RProvider.fsx\"\n#load \"../../bin/net45/Deedle.fsx\"\n\nopen RProvider\nopen RDotNet\nopen Deedle\n",
"_____no_output_____"
]
],
[
[
"(**\n\nIf you're not using NuGet from Visual Studio, then you'll need to manually copy the\nfile `Deedle.RProvider.Plugin.dll` from the package `Deedle.RPlugin` to the \ndirectory where `RProvider.dll` is located (in `RProvider/lib`). Once that's\ndone, the R provider will automatically find the plugin.\n\n<a name=\"frames\"></a>\n\nPassing data frames to and from R\n---------------------------------\n\n### From R to Deedle\nLet's start by looking at passing data frames from R to Deedle. To test this, we\ncan use some of the sample data sets available in the `datasets` package. The R\nmakes all packages available under the `RProvider` namespace, so we can just\nopen `datasets` and access the `mtcars` data set using `R.mtcars` (when typing\nthe code, you'll get automatic completion when you type `R` followed by dot):\n\n*)",
"_____no_output_____"
],
[
"\n(*** define-output:mtcars ***)",
"_____no_output_____"
]
],
[
[
"\nopen RProvider.datasets\n\n// Get mtcars as an untyped object\nR.mtcars.Value\n\n// Get mtcars as a typed Deedle frame\nlet mtcars : Frame<string, string> = R.mtcars.GetValue()\n",
"_____no_output_____"
]
],
[
[
"(*** include-value:mtcars ***)",
"_____no_output_____"
],
[
"\n(**\nThe first sample uses the `Value` property to convert the data set to a boxed Deedle\nframe of type `obj`. This is a great way to explore the data, but when you want to do \nsome further processing, you need to specify the type of the data frame that you want\nto get. This is done on line 7 where we get `mtcars` as a Deedle frame with both rows\nand columns indexed by `string`.\n\nTo see that this is a standard Deedle data frame, let's group the cars by the number of\ngears and calculate the average \"miles per galon\" value based on the gear. To visualize\nthe data, we use the [F# Charting library](https://github.com/fsharp/FSharp.Charting):\n\n*)",
"_____no_output_____"
],
[
"\n(*** define-output:mpgch ***)",
"_____no_output_____"
]
],
[
[
"\n#load \"../../packages/FSharp.Charting/lib/net45/FSharp.Charting.fsx\"\nopen FSharp.Charting\n\nmtcars\n|> Frame.groupRowsByInt \"gear\"\n|> Frame.getCol \"mpg\"\n|> Stats.levelMean fst\n|> Series.observations |> Chart.Column\n\n",
"_____no_output_____"
]
],
[
[
"(*** include-it:mpgch ***)",
"_____no_output_____"
],
[
"\n\n(**\n\n### From Deedle to R\n\nSo far, we looked how to turn R data frame into Deedle `Frame<R, C>`, so let's look\nat the opposite direction. The following snippet first reads Deedle data frame \nfrom a CSV file (file name is in the `airQuality` variable). We can then use the\ndata frame as argument to standard R functions that expect data frame.\n*)",
"_____no_output_____"
]
],
[
[
"\n\nlet air = Frame.ReadCsv(airQuality, separators=\";\")\n\n",
"_____no_output_____"
]
],
[
[
"(*** include-value:air ***)",
"_____no_output_____"
],
[
"\n\n(**\nLet's first try passing the `air` frame to the R `as.data.frame` function (which \nwill not do anything, aside from importing the data into R). To do something \nslightly more interesting, we then use the `colMeans` R function to calculate averages\nfor each column (to do this, we need to open the `base` package):\n*)",
"_____no_output_____"
]
],
[
[
"\nopen RProvider.``base``\n\n// Pass air data to R and print the R output\nR.as_data_frame(air)\n\n// Pass air data to R and get column means\nR.colMeans(air)\n// [fsi:val it : SymbolicExpression =]\n// [fsi: Ozone Solar.R Wind Temp Month Day ]\n// [fsi: NaN NaN 9.96 77.88 6.99 15.8]\n\n",
"_____no_output_____"
]
],
[
[
"(** \nAs a final example, let's look at the handling of missing values. Unlike R, Deedle does not \ndistinguish between missing data (`NA`) and not a number (`NaN`). For example, in the \nfollowing simple frame, the `Floats` column has missing value for keys 2 and 3 while\n`Names` has missing value for the row 2:\n*)",
"_____no_output_____"
]
],
[
[
"\n// Create sample data frame with missing values\nlet df = \n [ \"Floats\" =?> series [ 1 => 10.0; 2 => nan; 4 => 15.0]\n \"Names\" =?> series [ 1 => \"one\"; 3 => \"three\"; 4 => \"four\" ] ] \n |> frame\n",
"_____no_output_____"
]
],
[
[
"(**\nWhen we pass the data frame to R, missing values in numeric columns are turned into `NaN`\nand missing data for other columns are turned into `NA`. Here, we use `R.assign` which\nstores the data frame in a varaible available in the current R environment:\n*)",
"_____no_output_____"
]
],
[
[
"\nR.assign(\"x\", df)\n// [fsi:val it : SymbolicExpression = ]\n// [fsi: Floats Names ]\n// [fsi: 1 10 one ] \n// [fsi: 2 NaN <NA> ]\n// [fsi: 4 15 four ]\n// [fsi: 3 NaN three ]\n",
"_____no_output_____"
]
],
[
[
"(**\n\n<a name=\"series\"></a>\n\nPassing time series to and from R\n---------------------------------\n\nFor working with time series data, the Deedle plugin uses [the zoo package](http://cran.r-project.org/web/packages/zoo/index.html) \n(Z's ordered observations). If you do not have the package installed, you can do that\nby using the `install.packages(\"zoo\")` command from R or using `R.install_packages(\"zoo\")` from\nF# after opening `RProvider.utils`. When running the code from F#, you'll need to restart your \neditor and F# interactive after it is installed.\n\n### From R to Deedle\n\nLet's start by looking at getting time series data from R. We can again use the `datasets`\npackage with samples. For example, the `austres` data set gives us access to \nquarterly time series of the number of australian residents:\n*)",
"_____no_output_____"
]
],
[
[
"\nR.austres.Value\n// [fsi:val it : obj =]\n// [fsi: 1971.25 -> 13067.3 ]\n// [fsi: 1971.5 -> 13130.5 ]\n// [fsi: 1971.75 -> 13198.4 ]\n// [fsi: ... -> ... ]\n// [fsi: 1992.75 -> 17568.7 ]\n// [fsi: 1993 -> 17627.1 ]\n// [fsi: 1993.25 -> 17661.5 ]\n",
"_____no_output_____"
]
],
[
[
"(**\nAs with data frames, when we want to do any further processing with the time series, we need\nto use the generic `GetValue` method and specify a type annotation to that tells the F#\ncompiler that we expect a series where both keys and values are of type `float`:\n*)",
"_____no_output_____"
]
],
[
[
"\n// Get series with numbers of australian residents\nlet austres : Series<float, float> = R.austres.GetValue()\n\n// Get TimeSpan representing (roughly..) two years\nlet twoYears = TimeSpan.FromDays(2.0 * 365.0)\n\n// Calculate means of sliding windows of 2 year size \naustres \n|> Series.mapKeys (fun y -> \n DateTime(int y, 1 + int (12.0 * (y - floor y)), 1))\n|> Series.windowDistInto twoYears Stats.mean\n",
"_____no_output_____"
]
],
[
[
"(**\n\nThe current version of the Deedle plugin supports only time series with single column.\nTo access, for example, the EU stock market data, we need to write a short R inline\ncode to extract the column we are interested in. The following gets the FTSE time \nseries from `EuStockMarkets`:\n\n*)",
"_____no_output_____"
]
],
[
[
"\nlet ftseStr = R.parse(text=\"\"\"EuStockMarkets[,\"FTSE\"]\"\"\")\nlet ftse : Series<float, float> = R.eval(ftseStr).GetValue()\n",
"_____no_output_____"
]
],
[
[
"(**\n\n### From Deedle to R\n\nThe opposite direction is equally easy. To demonstrate this, we'll generate a simple\ntime series with 3 days of randomly generated values starting today:\n*)",
"_____no_output_____"
]
],
[
[
"\nlet rnd = Random()\nlet ts = \n [ for i in 0.0 .. 100.0 -> \n DateTime.Today.AddHours(i), rnd.NextDouble() ] \n |> series\n",
"_____no_output_____"
]
],
[
[
"(**\nNow that we have a time series, we can pass it to R using the `R.as_zoo` function or\nusing `R.assign` to store it in an R variable. As previously, the R provider automatically\nshows the output that R prints for the value:\n*)",
"_____no_output_____"
]
],
[
[
"\nopen RProvider.zoo\n\n// Just convert time series to R\nR.as_zoo(ts)\n// Convert and assing to a variable 'ts'\nR.assign(\"ts\", ts)\n// [fsi:val it : string =\n// [fsi: 2013-11-07 05:00:00 2013-11-07 06:00:00 2013-11-07 07:00:00 ...]\n// [fsi: 0.749946652 0.580584353 0.523962789 ...]\n\n",
"_____no_output_____"
]
],
[
[
"(**\nTypically, you will not need to assign time series to an R variable, because you can \nuse it directly as an argument to functions that expect time series. For example, the\nfollowing snippet applies the rolling mean function with a window size 20 to the \ntime series.\n*)",
"_____no_output_____"
]
],
[
[
"\n// Rolling mean with window size 20\nR.rollmean(ts, 20)\n\n",
"_____no_output_____"
]
],
[
[
"(**\nThis is a simple example - in practice, you can achieve the same thing with `Series.window`\nfunction from Deedle - but it demonstrates how easy it is to use R packages with \ntime series (and data frames) from Deedle. As a final example, we create a data frame that\ncontains the original time series together with the rolling mean (in a separate column)\nand then draws a chart showing the results:\n*)",
"_____no_output_____"
],
[
"\n\n(*** define-output:means ***)",
"_____no_output_____"
]
],
[
[
"\n// Use 'rollmean' to calculate mean and 'GetValue' to \n// turn the result into a Deedle time series\nlet tf = \n [ \"Input\" => ts \n \"Means5\" => R.rollmean(ts, 5).GetValue<Series<_, float>>()\n \"Means20\" => R.rollmean(ts, 20).GetValue<Series<_, float>>() ]\n |> frame\n\n// Chart original input and the two rolling means\nChart.Combine\n [ Chart.Line(Series.observations tf?Input)\n Chart.Line(Series.observations tf?Means5)\n Chart.Line(Series.observations tf?Means20) ]\n\n",
"_____no_output_____"
]
],
[
[
"(**\nDepending on your random number generator, the resulting chart looks something like this:\n*)",
"_____no_output_____"
],
[
"\n\n(*** include-it:means ***)",
"_____no_output_____"
]
],
[
[
"\n",
"_____no_output_____"
]
]
] | [
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] | [
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
]
] |
d0fe58f7b02cc15c1847d94a4dbdc5aef88c4b8b | 177,479 | ipynb | Jupyter Notebook | examples/Notebooks/flopy3_mf6_A_simple-model.ipynb | tomvansteijn/flopy | 06b0a71b9b69600d61c5233fd068946627e6cdad | [
"CC0-1.0",
"BSD-3-Clause"
] | null | null | null | examples/Notebooks/flopy3_mf6_A_simple-model.ipynb | tomvansteijn/flopy | 06b0a71b9b69600d61c5233fd068946627e6cdad | [
"CC0-1.0",
"BSD-3-Clause"
] | null | null | null | examples/Notebooks/flopy3_mf6_A_simple-model.ipynb | tomvansteijn/flopy | 06b0a71b9b69600d61c5233fd068946627e6cdad | [
"CC0-1.0",
"BSD-3-Clause"
] | null | null | null | 160.035167 | 60,156 | 0.847204 | [
[
[
"# FloPy\n\n## Creating a Simple MODFLOW 6 Model with Flopy\n\nThe purpose of this notebook is to demonstrate the Flopy capabilities for building a simple MODFLOW 6 model from scratch, running the model, and viewing the results. This notebook will demonstrate the capabilities using a simple lake example. A separate notebook is also available in which the same lake example is created for MODFLOW-2005 (flopy3_lake_example.ipynb).",
"_____no_output_____"
],
[
"### Setup the Notebook Environment",
"_____no_output_____"
]
],
[
[
"import sys\nimport os\nimport platform\nimport numpy as np\nimport matplotlib as mpl\nimport matplotlib.pyplot as plt\n\n# run installed version of flopy or add local path\ntry:\n import flopy\nexcept:\n fpth = os.path.abspath(os.path.join('..', '..'))\n sys.path.append(fpth)\n import flopy\n\nprint(sys.version)\nprint('numpy version: {}'.format(np.__version__))\nprint('matplotlib version: {}'.format(mpl.__version__))\nprint('flopy version: {}'.format(flopy.__version__))",
"3.7.7 (default, Mar 26 2020, 10:32:53) \n[Clang 4.0.1 (tags/RELEASE_401/final)]\nnumpy version: 1.19.2\nmatplotlib version: 3.3.0\nflopy version: 3.3.2\n"
],
[
"# For this example, we will set up a model workspace.\n# Model input files and output files will reside here.\nworkspace = os.path.join('data', 'mf6lake')\nif not os.path.exists(workspace):\n os.makedirs(workspace)",
"_____no_output_____"
]
],
[
[
"### Create the Flopy Model Objects\n\nWe are creating a square model with a specified head equal to `h1` along all boundaries. The head at the cell in the center in the top layer is fixed to `h2`. First, set the name of the model and the parameters of the model: the number of layers `Nlay`, the number of rows and columns `N`, lengths of the sides of the model `L`, aquifer thickness `H`, hydraulic conductivity `k`",
"_____no_output_____"
]
],
[
[
"name = 'mf6lake'\nh1 = 100\nh2 = 90\nNlay = 10 \nN = 101 \nL = 400.0 \nH = 50.0 \nk = 1.0",
"_____no_output_____"
]
],
[
[
"One big difference between MODFLOW 6 and previous MODFLOW versions is that MODFLOW 6 is based on the concept of a simulation. A simulation consists of the following:\n\n* Temporal discretization (TDIS)\n* One or more models (GWF is the only model supported at present)\n* Zero or more exchanges (instructions for how models are coupled)\n* Solutions\n\nFor this simple lake example, the simulation consists of the temporal discretization (TDIS) package (TDIS), a groundwater flow (GWF) model, and an iterative model solution (IMS), which controls how the GWF model is solved. ",
"_____no_output_____"
]
],
[
[
"# Create the Flopy simulation object\nsim = flopy.mf6.MFSimulation(sim_name=name, exe_name='mf6', \n version='mf6', sim_ws=workspace)\n\n# Create the Flopy temporal discretization object\ntdis = flopy.mf6.modflow.mftdis.ModflowTdis(sim, pname='tdis', time_units='DAYS', nper=1, \n perioddata=[(1.0, 1, 1.0)])\n\n# Create the Flopy groundwater flow (gwf) model object\nmodel_nam_file = '{}.nam'.format(name)\ngwf = flopy.mf6.ModflowGwf(sim, modelname=name, \n model_nam_file=model_nam_file)\n\n# Create the Flopy iterative model solver (ims) Package object\nims = flopy.mf6.modflow.mfims.ModflowIms(sim, pname='ims', complexity='SIMPLE')",
"_____no_output_____"
]
],
[
[
"Now that the overall simulation is set up, we can focus on building the groundwater flow model. The groundwater flow model will be built by adding packages to it that describe the model characteristics.\n\nDefine the discretization of the model. All layers are given equal thickness. The `bot` array is build from `H` and the `Nlay` values to indicate top and bottom of each layer, and `delrow` and `delcol` are computed from model size `L` and number of cells `N`. Once these are all computed, the Discretization file is built.",
"_____no_output_____"
]
],
[
[
"# Create the discretization package\nbot = np.linspace(-H/Nlay, -H, Nlay) \ndelrow = delcol = L/(N-1)\ndis = flopy.mf6.modflow.mfgwfdis.ModflowGwfdis(gwf, pname='dis', nlay=Nlay, nrow=N, ncol=N,\n delr=delrow,delc=delcol,top=0.0,\n botm=bot)",
"_____no_output_____"
],
[
"# Create the initial conditions package\nstart = h1 * np.ones((Nlay, N, N))\nic = flopy.mf6.modflow.mfgwfic.ModflowGwfic(gwf, pname='ic', strt=start)",
"_____no_output_____"
],
[
"# Create the node property flow package\nnpf = flopy.mf6.modflow.mfgwfnpf.ModflowGwfnpf(gwf, pname='npf', icelltype=1, k=k,\n save_flows=True)",
"_____no_output_____"
],
[
"# Create the constant head package.\n# List information is created a bit differently for \n# MODFLOW 6 than for other MODFLOW versions. The\n# cellid (layer, row, column, for a regular grid)\n# must be entered as a tuple as the first entry.\n# Remember that these must be zero-based indices!\nchd_rec = []\nchd_rec.append(((0, int(N / 4), int(N / 4)), h2))\nfor layer in range(0, Nlay):\n for row_col in range(0, N):\n chd_rec.append(((layer, row_col, 0), h1))\n chd_rec.append(((layer, row_col, N - 1), h1))\n if row_col != 0 and row_col != N - 1:\n chd_rec.append(((layer, 0, row_col), h1))\n chd_rec.append(((layer, N - 1, row_col), h1))\nchd = flopy.mf6.modflow.mfgwfchd.ModflowGwfchd(gwf, pname='chd', maxbound=len(chd_rec), \n stress_period_data=chd_rec, save_flows=True)",
"_____no_output_____"
],
[
"# The chd package stored the constant heads in a structured\n# array, also called a recarray. We can get a pointer to the\n# recarray for the first stress period (iper = 0) as follows.\niper = 0\nra = chd.stress_period_data.get_data(key=iper)\nra",
"_____no_output_____"
],
[
"# We can make a quick plot to show where our constant\n# heads are located by creating an integer array\n# that starts with ones everywhere, but is assigned\n# a -1 where chds are located\nibd = np.ones((Nlay, N, N), dtype=int)\nfor k, i, j in ra['cellid']:\n ibd[k, i, j] = -1\n\nilay = 0\nplt.imshow(ibd[ilay, :, :], interpolation='none')\nplt.title('Layer {}: Constant Head Cells'.format(ilay + 1))",
"_____no_output_____"
],
[
"# Create the output control package\nheadfile = '{}.hds'.format(name)\nhead_filerecord = [headfile]\nbudgetfile = '{}.cbb'.format(name)\nbudget_filerecord = [budgetfile]\nsaverecord = [('HEAD', 'ALL'), \n ('BUDGET', 'ALL')]\nprintrecord = [('HEAD', 'LAST')]\noc = flopy.mf6.modflow.mfgwfoc.ModflowGwfoc(gwf, pname='oc', saverecord=saverecord, \n head_filerecord=head_filerecord,\n budget_filerecord=budget_filerecord,\n printrecord=printrecord)",
"_____no_output_____"
],
[
"# Note that help can always be found for a package\n# using either forms of the following syntax\nhelp(oc)\n#help(flopy.mf6.modflow.mfgwfoc.ModflowGwfoc)",
"Help on ModflowGwfoc in module flopy.mf6.modflow.mfgwfoc object:\n\nclass ModflowGwfoc(flopy.mf6.mfpackage.MFPackage)\n | ModflowGwfoc(model, loading_package=False, budget_filerecord=None, head_filerecord=None, headprintrecord=None, saverecord=None, printrecord=None, filename=None, pname=None, parent_file=None)\n | \n | ModflowGwfoc defines a oc package within a gwf6 model.\n | \n | Parameters\n | ----------\n | model : MFModel\n | Model that this package is a part of. Package is automatically\n | added to model when it is initialized.\n | loading_package : bool\n | Do not set this parameter. It is intended for debugging and internal\n | processing purposes only.\n | budget_filerecord : [budgetfile]\n | * budgetfile (string) name of the output file to write budget\n | information.\n | head_filerecord : [headfile]\n | * headfile (string) name of the output file to write head information.\n | headprintrecord : [columns, width, digits, format]\n | * columns (integer) number of columns for writing data.\n | * width (integer) width for writing each number.\n | * digits (integer) number of digits to use for writing a number.\n | * format (string) write format can be EXPONENTIAL, FIXED, GENERAL, or\n | SCIENTIFIC.\n | saverecord : [rtype, ocsetting]\n | * rtype (string) type of information to save or print. Can be BUDGET or\n | HEAD.\n | * ocsetting (keystring) specifies the steps for which the data will be\n | saved.\n | all : [keyword]\n | * all (keyword) keyword to indicate save for all time steps in\n | period.\n | first : [keyword]\n | * first (keyword) keyword to indicate save for first step in\n | period. This keyword may be used in conjunction with other\n | keywords to print or save results for multiple time steps.\n | last : [keyword]\n | * last (keyword) keyword to indicate save for last step in\n | period. This keyword may be used in conjunction with other\n | keywords to print or save results for multiple time steps.\n | frequency : [integer]\n | * frequency (integer) save at the specified time step\n | frequency. This keyword may be used in conjunction with other\n | keywords to print or save results for multiple time steps.\n | steps : [integer]\n | * steps (integer) save for each step specified in STEPS. This\n | keyword may be used in conjunction with other keywords to\n | print or save results for multiple time steps.\n | printrecord : [rtype, ocsetting]\n | * rtype (string) type of information to save or print. Can be BUDGET or\n | HEAD.\n | * ocsetting (keystring) specifies the steps for which the data will be\n | saved.\n | all : [keyword]\n | * all (keyword) keyword to indicate save for all time steps in\n | period.\n | first : [keyword]\n | * first (keyword) keyword to indicate save for first step in\n | period. This keyword may be used in conjunction with other\n | keywords to print or save results for multiple time steps.\n | last : [keyword]\n | * last (keyword) keyword to indicate save for last step in\n | period. This keyword may be used in conjunction with other\n | keywords to print or save results for multiple time steps.\n | frequency : [integer]\n | * frequency (integer) save at the specified time step\n | frequency. This keyword may be used in conjunction with other\n | keywords to print or save results for multiple time steps.\n | steps : [integer]\n | * steps (integer) save for each step specified in STEPS. This\n | keyword may be used in conjunction with other keywords to\n | print or save results for multiple time steps.\n | filename : String\n | File name for this package.\n | pname : String\n | Package name for this package.\n | parent_file : MFPackage\n | Parent package file that references this package. Only needed for\n | utility packages (mfutl*). For example, mfutllaktab package must have\n | a mfgwflak package parent_file.\n | \n | Method resolution order:\n | ModflowGwfoc\n | flopy.mf6.mfpackage.MFPackage\n | flopy.mf6.mfbase.PackageContainer\n | flopy.pakbase.PackageInterface\n | builtins.object\n | \n | Methods defined here:\n | \n | __init__(self, model, loading_package=False, budget_filerecord=None, head_filerecord=None, headprintrecord=None, saverecord=None, printrecord=None, filename=None, pname=None, parent_file=None)\n | Initialize self. See help(type(self)) for accurate signature.\n | \n | ----------------------------------------------------------------------\n | Data and other attributes defined here:\n | \n | budget_filerecord = <flopy.mf6.data.mfdatautil.ListTemplateGenerator o...\n | \n | dfn = [['block options', 'name budget_filerecord', 'type record budget...\n | \n | dfn_file_name = 'gwf-oc.dfn'\n | \n | head_filerecord = <flopy.mf6.data.mfdatautil.ListTemplateGenerator obj...\n | \n | headprintrecord = <flopy.mf6.data.mfdatautil.ListTemplateGenerator obj...\n | \n | package_abbr = 'gwfoc'\n | \n | printrecord = <flopy.mf6.data.mfdatautil.ListTemplateGenerator object>\n | \n | saverecord = <flopy.mf6.data.mfdatautil.ListTemplateGenerator object>\n | \n | ----------------------------------------------------------------------\n | Methods inherited from flopy.mf6.mfpackage.MFPackage:\n | \n | __repr__(self)\n | Return repr(self).\n | \n | __setattr__(self, name, value)\n | Implement setattr(self, name, value).\n | \n | __str__(self)\n | Return str(self).\n | \n | build_child_package(self, pkg_type, data, parameter_name, filerecord)\n | \n | build_child_packages_container(self, pkg_type, filerecord)\n | \n | build_mfdata(self, var_name, data=None)\n | \n | check(self, f=None, verbose=True, level=1, checktype=None)\n | Check package data for common errors.\n | \n | Parameters\n | ----------\n | f : str or file handle\n | String defining file name or file handle for summary file\n | of check method output. If a sting is passed a file handle\n | is created. If f is None, check method does not write\n | results to a summary file. (default is None)\n | verbose : bool\n | Boolean flag used to determine if check method results are\n | written to the screen\n | level : int\n | Check method analysis level. If level=0, summary checks are\n | performed. If level=1, full checks are performed.\n | checktype : check\n | Checker type to be used. By default class check is used from\n | check.py.\n | \n | Returns\n | -------\n | None\n | \n | Examples\n | --------\n | \n | >>> import flopy\n | >>> m = flopy.modflow.Modflow.load('model.nam')\n | >>> m.dis.check()\n | \n | create_package_dimensions(self)\n | \n | export(self, f, **kwargs)\n | Method to export a package to netcdf or shapefile based on the\n | extension of the file name (.shp for shapefile, .nc for netcdf)\n | \n | Parameters\n | ----------\n | f : str\n | filename\n | kwargs : keyword arguments\n | modelgrid : flopy.discretization.Grid instance\n | user supplied modelgrid which can be used for exporting\n | in lieu of the modelgrid associated with the model object\n | \n | Returns\n | -------\n | None or Netcdf object\n | \n | get_file_path(self)\n | \n | is_valid(self)\n | \n | load(self, strict=True)\n | \n | plot(self, **kwargs)\n | Plot 2-D, 3-D, transient 2-D, and stress period list (MfList)\n | package input data\n | \n | Parameters\n | ----------\n | package: flopy.pakbase.Package instance supplied for plotting\n | \n | **kwargs : dict\n | filename_base : str\n | Base file name that will be used to automatically generate\n | file names for output image files. Plots will be exported as\n | image files if file_name_base is not None. (default is None)\n | file_extension : str\n | Valid matplotlib.pyplot file extension for savefig(). Only\n | used if filename_base is not None. (default is 'png')\n | mflay : int\n | MODFLOW zero-based layer number to return. If None, then all\n | all layers will be included. (default is None)\n | kper : int\n | MODFLOW zero-based stress period number to return. (default is\n | zero)\n | key : str\n | MfList dictionary key. (default is None)\n | \n | Returns\n | ----------\n | axes : list\n | Empty list is returned if filename_base is not None. Otherwise\n | a list of matplotlib.pyplot.axis are returned.\n | \n | remove(self)\n | \n | set_all_data_external(self, check_data=True)\n | \n | set_model_relative_path(self, model_ws)\n | \n | write(self, ext_file_action=<ExtFileAction.copy_relative_paths: 3>)\n | \n | ----------------------------------------------------------------------\n | Data descriptors inherited from flopy.mf6.mfpackage.MFPackage:\n | \n | data_list\n | \n | filename\n | \n | name\n | \n | package_type\n | \n | parent\n | \n | plotable\n | \n | ----------------------------------------------------------------------\n | Methods inherited from flopy.mf6.mfbase.PackageContainer:\n | \n | get_package(self, name=None)\n | Get a package.\n | \n | Parameters\n | ----------\n | name : str\n | Name of the package, 'RIV', 'LPF', etc.\n | \n | Returns\n | -------\n | pp : Package object\n | \n | register_package(self, package)\n | \n | ----------------------------------------------------------------------\n | Static methods inherited from flopy.mf6.mfbase.PackageContainer:\n | \n | get_module(package_file_path)\n | \n | get_module_val(module, item, attrb)\n | \n | get_package_file_paths()\n | \n | model_factory(model_type)\n | \n | package_factory(package_type, model_type)\n | \n | ----------------------------------------------------------------------\n | Data descriptors inherited from flopy.mf6.mfbase.PackageContainer:\n | \n | __dict__\n | dictionary for instance variables (if defined)\n | \n | __weakref__\n | list of weak references to the object (if defined)\n | \n | package_dict\n | \n | package_names\n | \n | ----------------------------------------------------------------------\n | Data descriptors inherited from flopy.pakbase.PackageInterface:\n | \n | has_stress_period_data\n\n"
]
],
[
[
"### Create the MODFLOW 6 Input Files and Run the Model\n\nOnce all the flopy objects are created, it is very easy to create all of the input files and run the model.",
"_____no_output_____"
]
],
[
[
"# Write the datasets\nsim.write_simulation()",
"writing simulation...\n writing simulation name file...\n writing simulation tdis package...\n writing ims package ims...\n writing model mf6lake...\n writing model name file...\n writing package dis...\n writing package ic...\n"
],
[
"# Print a list of the files that were created\n# in workspace\nprint(os.listdir(workspace))",
"['mf6lake.oc', 'mf6lake.nam', 'mf6lake.dis', 'mf6lake.ims', 'mf6lake.chd', 'mf6lake.npf', 'mf6lake.ic', 'mfsim.nam', 'mf6lake.tdis']\n"
]
],
[
[
"### Run the Simulation\n\nWe can also run the simulation from the notebook, but only if the MODFLOW 6 executable is available. The executable can be made available by putting the executable in a folder that is listed in the system path variable. Another option is to just put a copy of the executable in the simulation folder, though this should generally be avoided. A final option is to provide a full path to the executable when the simulation is constructed. This would be done by specifying exe_name with the full path.",
"_____no_output_____"
]
],
[
[
"# Run the simulation\nsuccess, buff = sim.run_simulation()\nprint('\\nSuccess is: ', success)",
"FloPy is using the following executable to run the model: /Users/jdhughes/.local/bin/mf6\n MODFLOW 6\n U.S. GEOLOGICAL SURVEY MODULAR HYDROLOGIC MODEL\n VERSION 6.2.0 10/22/2020\n\n MODFLOW 6 compiled Oct 25 2020 16:15:39 with IFORT compiler (ver. 19.0.5)\n\nThis software has been approved for release by the U.S. Geological \nSurvey (USGS). Although the software has been subjected to rigorous \nreview, the USGS reserves the right to update the software as needed \npursuant to further analysis and review. No warranty, expressed or \nimplied, is made by the USGS or the U.S. Government as to the \nfunctionality of the software and related material nor shall the \nfact of release constitute any such warranty. Furthermore, the \nsoftware is released on condition that neither the USGS nor the U.S. \nGovernment shall be held liable for any damages resulting from its \nauthorized or unauthorized use. Also refer to the USGS Water \nResources Software User Rights Notice for complete use, copyright, \nand distribution information.\n\n \n Run start date and time (yyyy/mm/dd hh:mm:ss): 2020/10/26 15:56:09\n \n Writing simulation list file: mfsim.lst\n Using Simulation name file: mfsim.nam\n \n"
]
],
[
[
"### Post-Process Head Results\n\nPost-processing MODFLOW 6 results is still a work in progress. There aren't any Flopy plotting functions built in yet, like they are for other MODFLOW versions. So we need to plot the results using general Flopy capabilities. We can also use some of the Flopy ModelMap capabilities for MODFLOW 6, but in order to do so, we need to manually create a SpatialReference object, that is needed for the plotting. Examples of both approaches are shown below.\n\nFirst, a link to the heads file is created with `HeadFile`. The link can then be accessed with the `get_data` function, by specifying, in this case, the step number and period number for which we want to retrieve data. A three-dimensional array is returned of size `nlay, nrow, ncol`. Matplotlib contouring functions are used to make contours of the layers or a cross-section.",
"_____no_output_____"
]
],
[
[
"# Read the binary head file and plot the results\n# We can use the existing Flopy HeadFile class because\n# the format of the headfile for MODFLOW 6 is the same\n# as for previous MODFLOW verions\nfname = os.path.join(workspace, headfile)\nhds = flopy.utils.binaryfile.HeadFile(fname)\nh = hds.get_data(kstpkper=(0, 0))\nx = y = np.linspace(0, L, N)\ny = y[::-1]\nc = plt.contour(x, y, h[0], np.arange(90,100.1,0.2))\nplt.clabel(c, fmt='%2.1f')\nplt.axis('scaled');",
"_____no_output_____"
],
[
"x = y = np.linspace(0, L, N)\ny = y[::-1]\nc = plt.contour(x, y, h[-1], np.arange(90,100.1,0.2))\nplt.clabel(c, fmt='%1.1f')\nplt.axis('scaled');",
"_____no_output_____"
],
[
"z = np.linspace(-H/Nlay/2, -H+H/Nlay/2, Nlay)\nc = plt.contour(x, z, h[:,50,:], np.arange(90,100.1,.2))\nplt.axis('scaled');",
"_____no_output_____"
],
[
"# We can also use the Flopy PlotMapView capabilities for MODFLOW 6\nfig = plt.figure(figsize=(10, 10))\nax = fig.add_subplot(1, 1, 1, aspect='equal')\nmodelmap = flopy.plot.PlotMapView(model=gwf, ax=ax) \n\n# Then we can use the plot_grid() method to draw the grid\n# The return value for this function is a matplotlib LineCollection object,\n# which could be manipulated (or used) later if necessary.\nquadmesh = modelmap.plot_ibound(ibound=ibd)\nlinecollection = modelmap.plot_grid()\ncontours = modelmap.contour_array(h[0], levels=np.arange(90,100.1,0.2))",
"_____no_output_____"
],
[
"# We can also use the Flopy PlotMapView capabilities for MODFLOW 6\nfig = plt.figure(figsize=(10, 10))\nax = fig.add_subplot(1, 1, 1, aspect='equal')\n\n# Next we create an instance of the ModelMap class\nmodelmap = flopy.plot.PlotMapView(model=gwf, ax=ax)\n\n# Then we can use the plot_grid() method to draw the grid\n# The return value for this function is a matplotlib LineCollection object,\n# which could be manipulated (or used) later if necessary.\nquadmesh = modelmap.plot_ibound(ibound=ibd)\nlinecollection = modelmap.plot_grid()\npa = modelmap.plot_array(h[0])\ncb = plt.colorbar(pa, shrink=0.5)",
"_____no_output_____"
]
],
[
[
"### Post-Process Flows\n\nMODFLOW 6 writes a binary grid file, which contains information about the model grid. MODFLOW 6 also writes a binary budget file, which contains flow information. Both of these files can be read using Flopy capabilities. The MfGrdFile class in Flopy can be used to read the binary grid file. The CellBudgetFile class in Flopy can be used to read the binary budget file written by MODFLOW 6.",
"_____no_output_____"
]
],
[
[
"# read the binary grid file\nfname = os.path.join(workspace, '{}.dis.grb'.format(name))\nbgf = flopy.utils.mfgrdfile.MfGrdFile(fname)\n\n# data read from the binary grid file is stored in a dictionary\nbgf._datadict",
"_____no_output_____"
],
[
"# Information from the binary grid file is easily retrieved\nia = bgf._datadict['IA'] - 1\nja = bgf._datadict['JA'] - 1",
"_____no_output_____"
],
[
"# read the cell budget file\nfname = os.path.join(workspace, '{}.cbb'.format(name))\ncbb = flopy.utils.CellBudgetFile(fname, precision='double')\ncbb.list_records()\n\nflowja = cbb.get_data(text='FLOW-JA-FACE')[0][0, 0, :]\nchdflow = cbb.get_data(text='CHD')[0]",
"(1, 1, b' FLOW-JA-FACE', 689628, 1, -1, 1, 1., 1., 1., b'', b'', b'', b'')\n(1, 1, b' CHD', 101, 101, -10, 6, 1., 1., 1., b'MF6LAKE ', b'MF6LAKE ', b'MF6LAKE ', b'CHD ')\n"
],
[
"# By having the ia and ja arrays and the flow-ja-face we can look at\n# the flows for any cell and process them in the follow manner.\nk = 5; i = 50; j = 50\ncelln = k * N * N + i * N + j\nprint('Printing flows for cell {}'.format(celln + 1))\nfor ipos in range(ia[celln] + 1, ia[celln + 1]):\n cellm = ja[ipos] # change from one-based to zero-based\n print('Cell {} flow with cell {} is {}'.format(celln + 1, cellm + 1, flowja[ipos]))",
"Printing flows for cell 56106\nCell 56106 flow with cell 45905 is -0.0002858449337963975\nCell 56106 flow with cell 56005 is -0.025019694309449392\nCell 56106 flow with cell 56105 is -0.025019694309449392\nCell 56106 flow with cell 56107 is 0.025058524820593675\nCell 56106 flow with cell 56207 is 0.02505852482066473\nCell 56106 flow with cell 66307 is 0.00033448625827077195\n"
]
]
] | [
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] | [
[
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code"
]
] |
d0fe61fe810f25270728db59fc05d23c13411a22 | 59,881 | ipynb | Jupyter Notebook | _jupyterNotebooks/python_tutorial/python_basics_1.ipynb | PranavShirole/PranavShirole.github.io | f3d677564b62e1cbc4026e905f8476a3b3cfc68a | [
"MIT"
] | null | null | null | _jupyterNotebooks/python_tutorial/python_basics_1.ipynb | PranavShirole/PranavShirole.github.io | f3d677564b62e1cbc4026e905f8476a3b3cfc68a | [
"MIT"
] | null | null | null | _jupyterNotebooks/python_tutorial/python_basics_1.ipynb | PranavShirole/PranavShirole.github.io | f3d677564b62e1cbc4026e905f8476a3b3cfc68a | [
"MIT"
] | 2 | 2021-07-24T09:28:51.000Z | 2022-03-03T10:43:08.000Z | 25.888889 | 677 | 0.535479 | [
[
[
"As a self-taught Data Scientist and programmer, I always get asked about how I started my path towards learning, and a lot of non-coders ask me about how they can learn more about Data Science. And while I tell them about the umpteen Data Analytics and Data Visualization tools, various Machine Learning algorithms, and which Deep Learning frameworks to choose, it all starts with learning Python. Python is an interpreted high-level general-purpose programming language. In my view, Python is the best programming language to learn, in order to become a Data Scientist, owing to its readability, non-complexity, its large standard libraries, and its huge community. \nI have been the beneficiary of several of books and YouTube tutorials that have helped me become a better Python Developer. This blog post is my way of giving back to the community. This might not be the best place to start learning how to code in Python. However, this blog post aims to be a good cheat sheet for beginners trying to look something up or if they want a refresher as to how certain objects perform.",
"_____no_output_____"
],
[
"---\n### Index \n\n[Display Output](#Display-Output) \n[Getting information from the user](#Getting-information-from-the-user) \n[Comments](#Comments) \n[String concepts](#String-concepts) \n[Working with Numbers](#Working-with-Numbers) \n[Working with Dates](#Working-with-Dates) \n[Error Handling](#Error-Handling) \n[Handling Conditions](#Handling-Conditions) \n[Collections](#Collections) \n[Random Module](#Random-Module) \n[Loops](#Loops) \n[Functions](#Functions) \n",
"_____no_output_____"
],
[
"---\n### Display Output\nYou can use the `print` function to display output to your console. You can use either single or double quotes; just make sure that you stick to one for consistency.",
"_____no_output_____"
]
],
[
[
"print('Hello, World!')\nprint(\"Python is great\")",
"Hello, World!\nPython is great\n"
]
],
[
[
"#### Displaying blank lines \nBlank lines make the output more readable. For a blank line, you can insert a `print` function with nothing inside. Each `print` function prints on a new line by default. \nYou can also use `\\n` (the newline character) to print a new line at the end of a string or right in the middle of a string.",
"_____no_output_____"
]
],
[
[
"print('Hello')\nprint()\nprint('Above is a blank line\\n')\nprint('Blank line \\nin the middle of a string')",
"Hello\n\nAbove is a blank line\n\nBlank line \nin the middle of a string\n"
]
],
[
[
"---\n### Getting information from the user\nYou can use the `input` to ask for information from your user. We pass in a message with the `input` function.",
"_____no_output_____"
]
],
[
[
"name = input('What is your name? ')\nprint(name)",
"What is your name? Pranav\nPranav\n"
]
],
[
[
"Here, whatever value is typed in by the user will be stored in the variable `name` and can be used as needed. We've chosen to print the value of `name` on the screen.",
"_____no_output_____"
],
[
"---\n### Comments\nComments are a way of documenting your code. Comments can be added using `#`. These lines of code will not execute.",
"_____no_output_____"
]
],
[
[
"# print('Hello')\n# the above code of line will not execute\n# but the below one will \nprint('How are you?')",
"How are you?\n"
]
],
[
[
"You can also use `''' '''` for multi-line comments.",
"_____no_output_____"
]
],
[
[
"'''\nPython is an important\nprogramming language in\nyour Data Science journey\n'''\nprint('Python')",
"Python\n"
]
],
[
[
"It's a good idea to write comments before your function explaining what that function does. \nCommenting out lines can also help debug your code.",
"_____no_output_____"
],
[
"---\n### String concepts\nStrings can be stored in variables. Variables are just placeholders for some value inside our code.",
"_____no_output_____"
]
],
[
[
"first_name = 'Pranav'\nprint(first_name)",
"Pranav\n"
]
],
[
[
"#### Concatenate strings\nYou can combine strings with the `+` operator.",
"_____no_output_____"
]
],
[
[
"first_name = 'John'\nlast_name = 'Doe'\nprint(first_name + last_name)\nprint('Hello, ' + first_name + ' ' + last_name)",
"JohnDoe\nHello, John Doe\n"
]
],
[
[
"#### Functions to modify strings\nBelow we have used functions to \n- convert a string to uppercase \n- convert a string to lowercase\n- capitalize just the first word\n- count all of the instances of a particular string.",
"_____no_output_____"
]
],
[
[
"sentence = 'My name is John Doe'\nprint(sentence.upper())\nprint(sentence.lower())\nprint(sentence.capitalize())\nprint(sentence.count('o'))",
"MY NAME IS JOHN DOE\nmy name is john doe\nMy name is john doe\n2\n"
]
],
[
[
"You can use the escape character (backslash) `\\` to insert characters that are illegal in a string. An example of an illegal character is a single quote inside a string that is surrounded by single quotes.",
"_____no_output_____"
]
],
[
[
"first_name = input('What\\'s your first name? ')\nlast_name = input('What\\'s is your last name? ')\nprint('Hello, ' + first_name.capitalize() + \n ' ' + last_name.capitalize())",
"What's your first name? JOHN\nWhat's is your last name? doe\nHello, John Doe\n"
]
],
[
[
"#### Custom string formatting\nTo infuse things in strings dynamically, you can use string formatting. ",
"_____no_output_____"
]
],
[
[
"first_name = 'John'\nlast_name = 'Doe'",
"_____no_output_____"
]
],
[
[
"There are two ways you can do this:\n- formatting with `.format()` string method\n",
"_____no_output_____"
]
],
[
[
"name = 'Hello, {} {}'.format(first_name, last_name)\nprint(name)",
"Hello, John Doe\n"
]
],
[
[
"- formatting with string literals, called f-strings",
"_____no_output_____"
]
],
[
[
"name = f'Hello, {first_name} {last_name}'\nprint(name)",
"Hello, John Doe\n"
]
],
[
[
"---\n### Working with numbers\nNumbers can be stored in variables. Make sure the variables have meaningful names. We can pass those variables inside functions.",
"_____no_output_____"
]
],
[
[
"pi = 3.14159\nprint(pi)",
"3.14159\n"
]
],
[
[
"#### Math with Numbers\n- `+` for addition\n- `-` for subtraction\n- `*` for multiplication\n- `/` for division\n- `**` for exponent",
"_____no_output_____"
]
],
[
[
"num1 = 9\nnum2 = 5\nprint(num1 + num2)\nprint(num1 ** num2)",
"14\n59049\n"
]
],
[
[
"#### Type Conversion\nYou cannot combine strings with numbers in Python. For e.g., executing the code below will result in an error: \n",
"_____no_output_____"
]
],
[
[
"days_in_Dec = 31\nprint(days_in_Dec + ' days in December')",
"_____no_output_____"
]
],
[
[
"When displaying a string that contains numbers, you must convert the numbers into strings.",
"_____no_output_____"
]
],
[
[
"days_in_Dec = 31 \nprint(str(days_in_Dec) + ' days in December')",
"31 days in December\n"
]
],
[
[
"Numbers can be stored as strings. However, numbers stored as strings are treated as strings.",
"_____no_output_____"
]
],
[
[
"num1 = '10'\nnum2 = '20'\nprint(num1 + num2)",
"1020\n"
]
],
[
[
"Also, the input function always returns a string.",
"_____no_output_____"
]
],
[
[
"num1 = input('Enter the first number: ')\nnum2 = input('Enter the second number: ')\nprint(num1 + num2)",
"Enter the first number: 35\nEnter the second number: 75\n3575\n"
]
],
[
[
"But here you can see that you have a number stored in a string. What if you want to treat it as a number and do math with it? \nYou can do another data type conversion. The `int` function will convert it to a whole number, while the `float` function will convert it into a floating point number that might have decimal places.",
"_____no_output_____"
]
],
[
[
"num1 = input('Enter the first number: ')\nnum2 = input('Enter the second number: ')\nprint(int(num1) + int(num2))\nprint(float(num1) + float(num2))",
"Enter the first number: 35\nEnter the second number: 75\n110\n110.0\n"
]
],
[
[
"---\n### Working with Dates\nWe often need current date and time when logging errors and saving data. To get the current date and time, we need to use the `datetime` library.",
"_____no_output_____"
]
],
[
[
"from datetime import datetime\n\n# the now function returns a datetime object\ncurrent_date = datetime.now()\n\nprint('Today is: ' + str(current_date))",
"Today is: 2021-06-05 11:49:47.183898\n"
]
],
[
[
"There are a whole bunch of functions you can use with `datetime` objects to manipulate dates. \n`timedelta` is used to define a period of time.",
"_____no_output_____"
]
],
[
[
"from datetime import datetime, timedelta\ntoday = datetime.now()\nprint('Today is: ' + str(today))\n\none_day = timedelta(days=1)\none_week = timedelta(weeks=1)\nyesterday = today - one_day\npast_week = today - one_week\nprint('Yesterday was: ' + str(yesterday))\nprint('One week ago was: ' + str(past_week))",
"Today is: 2021-06-05 12:01:31.236665\nYesterday was: 2021-06-04 12:01:31.236665\nOne week ago was: 2021-05-29 12:01:31.236665\n"
]
],
[
[
"You can also control the format of the date displayed on the screen. You can request just the day, month, year, hour, minutes and even seconds.",
"_____no_output_____"
]
],
[
[
"print('Day: ' + str(current_date.day))\nprint('Month: ' + str(current_date.month))\nprint('Year: ' + str(current_date.year))",
"Day: 5\nMonth: 6\nYear: 2021\n"
]
],
[
[
"Sometimes, you can receive a date as a string, and you might need to store it as a date. You'll need to convert it to a `datetime` object.",
"_____no_output_____"
]
],
[
[
"birthday = input('When is your birthday (dd/mm/yyyy)? ')\n\n# the strptime function allows you to mention the \n# format in which you'll be receiving the date\nbirthday_date = datetime.strptime(birthday, '%d/%m/%Y')\nprint('Birthday: ' + str(birthday_date))",
"When is your birthday (dd/mm/yyyy)? 28/2/2000\nBirthday: 2000-02-28 00:00:00\n"
]
],
[
[
"So what date was it three days before you were born?",
"_____no_output_____"
]
],
[
[
"birthday = input('When is your birthday (dd/mm/yyyy)? ')\nbirthday_date = datetime.strptime(birthday, '%d/%m/%Y')\nprint('Birthday: ' + str(birthday_date))\nthree_days = timedelta(days=3)\nthree_before = birthday_date - three_days\nprint('Date three days before birthday: ' + str(three_before))",
"When is your birthday (dd/mm/yyyy)? 28/2/2000\nBirthday: 2000-02-28 00:00:00\nDate three days before birthday: 2000-02-25 00:00:00\n"
]
],
[
[
"---\n### Error Handling \n*Error handling* is when you have a problem with your code that is running, and its not something that you're going to be able to predict when you push your code to production. For e.g., permissions issue, database change, server being down, etc. Basically things that happen in the wild, which you have no control over. \n*Debugging* is when you know that there's something wrong (a bug) with your code because you did something incorrectly, and you're going to have to go in and correct it.\n\nThe following tools we're going to talk about are concerned with error handling. There are three types of errors:\n- syntax errors\n- runtime errors\n- logic errors\n\n",
"_____no_output_____"
],
[
"#### Syntax errors\nWith syntax errors, your code is not going to run at all. This type of error is easiest to track down.",
"_____no_output_____"
]
],
[
[
"# this code won't run at all\nx = 35\ny = 75\nif x == y\n print('x = y')",
"_____no_output_____"
]
],
[
[
"We're missing a colon after `y`, which is why we're getting the error above.",
"_____no_output_____"
],
[
"#### Runtime errors\nWith runtime errors, your code will run, but it will fail when it encounters the error.",
"_____no_output_____"
]
],
[
[
"# this code will fail when run\nx = 5\ny = 0\nprint(x / y)",
"_____no_output_____"
]
],
[
[
"We're trying to divide by zero, which is not possible. Python tells you why you're getting the error and points towards the line which needs to be fixed. It's good practice to start from the line mentioned and work your way up to the error. Runtime errors can also be caused due to an error in the framework you're using, but the chances of that happening are extremely rare. Most probably, if you have a runtime error, it's because there's something wrong in your code.",
"_____no_output_____"
],
[
"#### Catching runtime errors\nWhen a runtime error occurs, Python generates an exception during the execution and that can be handled, which avoids your program to interrupt. \nException handling:\n- `try`: this block will test the excepted error to occur\n- `except`: here you can handle the error\n- `else`: if there is no exception, then this block will be executed\n- `finally`: finally block always gets executed whether exception is generated or not\n\nThese tools are not used for finding bugs.",
"_____no_output_____"
]
],
[
[
"x = 5\ny = 0\n\ntry:\n print(x / y)\nexcept ZeroDivisionError as e:\n print('Sorry, something went wrong')\nexcept:\n print('Something really went wrong')\nfinally:\n print('This line always runs, on success or failure')",
"Sorry, something went wrong\nThis line always runs, on success or failure\n"
]
],
[
[
"#### Logic errors\nLogic errors occur when the code compiles properly, doesn't give any syntax or runtime errors, but it doesn't give you the response you're looking for.",
"_____no_output_____"
]
],
[
[
"# this code won't run at all\nx = 10\ny = 20\nif x > y:\n print(str(x) + ' is less than ' + str(y))",
"_____no_output_____"
]
],
[
[
"In the code above, `x` is less than `y`; but the `if` statement includes `x > y`, instead of `x < y`.\n\nWhen you're figuring out what went wrong with your code, just make sure that you reread your code. You can check the documentation and also search the internet on sites like StackOverflow and Medium.",
"_____no_output_____"
],
[
"---\n### Handling Conditions\nYour code might need the ability to take different actions based on different conditions. Below are the operations that you'll need for comparisons:\n- `>`: greater than\n- `<`: less than\n- `>=`: greater than or equal to\n- `<=`: less than or equal to\n- `==`: is equal to\n- `!=`: is not equal to\n\n#### if statement\nThe `if` statement contains a logical expression using which the data is compared and a decision is made based on the result of the comparison.",
"_____no_output_____"
]
],
[
[
"price = 250.0\n\nif price >= 100.00:\n tax = 0.3\n print(tax)",
"0.3\n"
]
],
[
[
"#### if - else statement\nYou can add a default action using `else`. An `else` statement contains the block of code that executes if the conditional expression in the `if` statement resolves to `0` or a `False` value.",
"_____no_output_____"
]
],
[
[
"price = 50\n\nif price >= 100.00:\n tax = 0.3\nelse:\n tax = 0\n \nprint(tax)",
"0\n"
]
],
[
[
"Be careful when comparing strings. String comparisons are case sensitive.",
"_____no_output_____"
]
],
[
[
"country = 'INDIA'\nif country == 'india':\n print('Namaste')\nelse:\n print('Hello')",
"Hello\n"
],
[
"country = 'INDIA'\nif country.lower() == 'india':\n print('Namaste')\nelse:\n print('Hello')",
"Namaste\n"
]
],
[
[
"#### if - elif - else statement\nYou may need to check multiple conditions to determine the correct action. The `elif` statement allows you to check multiple expressions for `True` and execute a block of cide as soon as one of the conditions evaluates to `True`.",
"_____no_output_____"
]
],
[
[
"# income tax percentage by state\nstate = input('Which state do you live in? ')\n\nif state == 'Georgia':\n tax = 5.75\nelif state == 'California':\n tax = 13.3\nelif state == 'Texas' or state == 'Florida':\n tax = 0.0\nelse:\n tax = 4.0\n\nprint(tax)",
"Which state do you live in? Georgia\n5.75\n"
]
],
[
[
"#### OR statements\n| first condition | second condition | evaluation | \n|-----------------|------------------|------------| \n|True |True |True | \n|True |False |True |\n|False |True |True | \n|False |False |False |\n\n#### AND statements\n| first condition | second condition | evaluation | \n|-----------------|------------------|------------| \n|True |True |True | \n|True |False |False |\n|False |True |False | \n|False |False |False |",
"_____no_output_____"
],
[
"#### in operator\nIf you have a list of possible values to check, you can use the `in` operator.",
"_____no_output_____"
]
],
[
[
"# income tax rates by state\nstate = input('Which state do you live in? ')\n\nif state in ('Texas', 'Florida', 'Alaska',\n 'Wyoming', 'South Dakota'):\n tax = 0.0\nelif state == 'California':\n tax = 13.3\nelif state == 'Georgia':\n tax = 5.75\nelse:\n tax = 4.0\n\nprint(tax)",
"Which state do you live in? Alaska\n0.0\n"
]
],
[
[
"#### Nested if statement\nThere may be a situation when you want to check for another condition after a condition resolves to `True`. If an action depends on a combination of conditions, you can nest `if` statements.",
"_____no_output_____"
]
],
[
[
"country = input(\"What country do you live in? \")\n\nif country.lower() == 'canada':\n province = input(\"What province/state do you live in? \")\n if province in('Alberta', 'Nunavut','Yukon'):\n tax = 0.05\n elif province == 'Ontario':\n tax = 0.13\n else:\n tax = 0.15\nelse:\n tax = 0.0\nprint(tax)",
"What country do you live in? Canada\nWhat province/state do you live in? Ontario\n0.13\n"
]
],
[
[
"Sometimes you can combine conditions with `and` instead of nested `if` statements. \nLet's assume that you're trying to calculate which students in a college have made the honor roll. The requirements for making the honor roll are a minimum 85% GPA and maintaining all your grades at at least 70%. ",
"_____no_output_____"
]
],
[
[
"# convert strings into float\ngpa = float(input('What\\'s your GPA? '))\nlowest_grade = float(input('What was your lowest grade? '))\n\nif gpa >= 0.85 and lowest_grade >= 0.7:\n print('You made the honor roll')\nelse:\n print('You\\'re really stupid')",
"What's your GPA? 0.8\nWhat was your lowest grade? 0.75\nYou're really stupid\n"
]
],
[
[
"If you have a very complicated `if` statement, rather than copying and pasting it in different parts of you code to do different things, we can remember what happened the last time we looked at the `if` statement with a Boolean variable.",
"_____no_output_____"
]
],
[
[
"gpa = float(input('What\\'s your GPA? '))\nlowest_grade = float(input('What was your lowest grade? '))\n\nif gpa >= 0.85 and lowest_grade >= 0.7:\n honor_roll = True\nelse:\n honor_roll = False\n \n''' Somewhere later in your code if you need\nto check if a student is on honor roll, all\nyou need to do is check the boolean variable\nset earlier in the code'''\nif honor_roll: # True by default\n print('You made the honor roll')",
"What's your GPA? 0.9\nWhat was your lowest grade? 0.87\nYou made the honor roll\n"
]
],
[
[
"---\n### Collections\n\n#### Lists\nLists are a collection of items.",
"_____no_output_____"
]
],
[
[
"# prepopulate a list\nnames = ['John', 'Will', 'Max']\n\n# start with an empty list\nscores = []\n# add new item to the end\nscores.append(90)\nscores.append(91)\n\nprint(names)\nprint(scores)\n\n# lists are zero-indexed\nprint(scores[1])",
"['John', 'Will', 'Max']\n[90, 91]\n91\n"
]
],
[
[
"You can get the number of items in a list using `len`.",
"_____no_output_____"
]
],
[
[
"names = ['John', 'Will', 'Max']\n\n# get the number of items using len\nprint(len(names))",
"3\n"
]
],
[
[
"You can insert an item in a list using `insert`. This will insert the item at the specific index that you mention.",
"_____no_output_____"
]
],
[
[
"# Bill will be inserted at index 0, i.e. the first item\nnames.insert(0, 'Bill')\nprint(names)",
"['Bill', 'John', 'Will', 'Max']\n"
]
],
[
[
"You can use `sort` to sort strings in alphabetical order. In case of numbers, it sorts them in the ascending order. Remember that using `sort` will modify the list!",
"_____no_output_____"
]
],
[
[
"names.sort()\nprint(names)",
"['Bill', 'John', 'Max', 'Will']\n"
]
],
[
[
"You can retrieve a range within the list by indicating the start and end index; the end index being exclusive, i.e. it will not be included in the list.",
"_____no_output_____"
]
],
[
[
"names = ['Amy', 'Susan', 'Jackie', 'Kylie', 'Ellen']\n\n# start and end index\npresenters = names[1:3]\n\n# all names up to but not including index 3\nhosts = names[:3]\n\n# all names from 3 onwards, including index 3\njudges = names[3:]\n\nprint(names)\nprint(presenters)\nprint(hosts)\nprint(judges)",
"['Amy', 'Susan', 'Jackie', 'Kylie', 'Ellen']\n['Susan', 'Jackie']\n['Amy', 'Susan', 'Jackie']\n['Kylie', 'Ellen']\n"
]
],
[
[
"#### Arrays\nArrays are a collection of numbered data types. Unlike a list, in order for you to use an array, you have to create an array object by importing it from the `array` library.",
"_____no_output_____"
]
],
[
[
"from array import array\n\n# indicate the numerical type you'll use\nscores = array('d') # d indicates a double\nscores.append(80)\nscores.append(81)\nprint(scores)\nprint(scores[0])",
"array('d', [80.0, 81.0])\n80.0\n"
]
],
[
[
"So what's the difference between an array and a list? \nArrays are only numerical data types and everything inside the array must be of the same data type. They can help add extra structure to your code. \nLists can store anything you want, can store any data type, and can have mixed data types. They give more flexibility to your code.",
"_____no_output_____"
],
[
"#### Dictionaries\nDictionaries give you the ability to put together a group of items; but instead of using numeric indexes, you can use key-value pairs.",
"_____no_output_____"
]
],
[
[
"person = {'first': 'John'}\nperson['last'] = 'Wick'\nprint(person)\nprint(person['first'])",
"{'first': 'John', 'last': 'Wick'}\nJohn\n"
],
[
"identity = {\n 'Batman': 'Bruce Wayne',\n 'Superman': 'Clark Kent',\n 'Spiderman': 'Peter Parker',\n 'Iron Man': 'Tony Stark'\n}\n\nprint(identity)",
"{'Batman': 'Bruce Wayne', 'Superman': 'Clark Kent', 'Spiderman': 'Peter Parker', 'Iron Man': 'Tony Stark'}\n"
]
],
[
[
"When to use a dictionary vs a list?\nIt depends on whether you want to name things and whether you want items to be in a guaranteed order. \nA dictionary will let you name key-value pairs but it does not guarantee you a specific order. \nA list does guarantee you a specific order since it has a zero-based index.",
"_____no_output_____"
],
[
"---\n\n### Random Module\n\nOne way to introduce random numbers in your code is to use the `random` module. \nFirst you need to import the `random` module.",
"_____no_output_____"
]
],
[
[
"import random\n\n# generate a random whole number between 1 and 50\n# inclusive of 1 and 50\nrandom_integer = random.randint(1, 50)\nprint(random_integer)",
"12\n"
],
[
"# generate a random floating point number between 0.0 and 1.0\n# exclusive of 1.0 \nrandom_float = random.random()\nprint(random_float)\n\n# generate a random floating point number between 0.0 and 5.0\nprint(random_float * 5)",
"0.6343084141143187\n3.1715420705715935\n"
]
],
[
[
"There are so many more methods to the `random` module and you can check out the Python documentation to find out about all the things you can do with this module.",
"_____no_output_____"
],
[
"---\n### Loops\n\nLoops are a concept that is used when you need to have things happening over and over again.\n\n#### for loops\n`for` loops are used to loop through a collection. With a `for` loop, you can go through each item in a list and perform some action with each individual item in the list.\n\n> `for item in list_of_items:\n # do something to each item`",
"_____no_output_____"
]
],
[
[
"# go through the list of names\nfor name in ['John', 'Will', 'Max']:\n print(name)",
"John\nWill\nMax\n"
],
[
"wildcats = ['lion', 'tiger', 'puma', 'jaguar', 'cheetah', 'leopard']\nfor wildcat in wildcats:\n print(wildcat + ' is a wildcat.')",
"lion is a wildcat.\ntiger is a wildcat.\npuma is a wildcat.\njaguar is a wildcat.\ncheetah is a wildcat.\nleopard is a wildcat.\n"
]
],
[
[
"You can loop a particular number of times using `range`. `range` automatically creates a list of numbers for you. Remember that for the `range` function, the end index is exclusive.\n\n> `for number in range(a, b):\n # do something\n print(number)`",
"_____no_output_____"
]
],
[
[
"# end index is exclusive\nfor index in range(0, 5):\n print(index)",
"0\n1\n2\n3\n4\n"
]
],
[
[
"If you want the range to increase by any other number, you can add a step to the function after the starting and ending indices.",
"_____no_output_____"
]
],
[
[
"for index in range(0, 15, 3):\n print(index)",
"0\n3\n6\n9\n12\n"
]
],
[
[
"#### while loop\n`while` loops are used to loop with a condition. As long as something is `True`, the code will stay inside of the `while` loop i.e. the loop will continue going while the condition is true. You need to make sure that at some point you change the condition and it must result to `False`; otherwise the program will be stuck in an infinite loop, resulting in an error.\n\n> `while something_is_true:\n # do something repeatedly`",
"_____no_output_____"
]
],
[
[
"names = ['John', 'Will', 'Max']\nindex = 0\nwhile index < len(names):\n print(names[index])\n # change the condition\n index += 1\n print(index)",
"John\n1\nWill\n2\nMax\n3\n"
],
[
"x = True\nwhile x:\n print('This is an example of a while loop.')\n # change the condition\n x = False\n \nwhile not x:\n print('This is another example of a while loop.')\n x = True",
"This is an example of a while loop.\nThis is another example of a while loop.\n"
]
],
[
[
"`for` loops are great when you want to iterate over something and you need to do something with each thing that you're iterating over. In cases like above, when you have a list, you almost always want use a `for` loop. \n\n`while` loops are useful when you don't care about the number in the sequence or about the item you're iterating through in a list, and you just simply want to carry out a functionality many times until a condition is met. You want to typically use a `while` loop when something is going to change automatically, e.g. when you need to read through a list of lines in a file, skip every alternate line, or if you're looking for something. \n`while` loops are more dangerous because they can lead to infinite loops if the condition is not met.",
"_____no_output_____"
],
[
"---\n### Functions\nA function is a block of organized, reusable code that is used to perform a single, related action. Function provide better modularity for your application and a high degree of code reusing, e.g. the `print()` function. You can create your own functions, called *user-defined functions*. \nProgramming is all about copying and pasting code from one place to another. If you find yourself copying and pasting the exact same lines of code to more places in your program, you should probably move that into a function. \nFunctions must be declared before the line of code where the function is called. \n\n**Defining Functions** \n> `def my_function():\n # do this\n # then do this\n # finally do this`\n \n**Calling Functions** \n> `my_function()`\n\n",
"_____no_output_____"
],
[
"#### Functions with Inputs\n\nThe input to a function is something that can be passed over when we call the function.\n\n> `def my_function(something):\n # do this with something\n # then do this\n # finally do this`\n \n> `my_function(123)`",
"_____no_output_____"
],
[
"#### Functions with Outputs\n\nThe output keyword for a function is `return`. The `return` line must be the last line of the function. You can have multiple return keywords or even a blank return keyword in a function.",
"_____no_output_____"
]
],
[
[
"def my_function():\n result = 5 * 4\n return result\n\nmy_function()",
"_____no_output_____"
]
],
[
[
"When you call a function that has an output, the returned output is what replaces the function call, and the output can be stored as a variable.",
"_____no_output_____"
]
],
[
[
"def my_function():\n return 5 * 8\n\noutput = my_function()\nprint(output)",
"40\n"
]
],
[
[
"Imagine that you're trying to figure out why your program is taking a long time to run. So you write some print statements inside your code to tell you what time it is when the code is running, so you can see what time it is at different stages when your code is running.",
"_____no_output_____"
]
],
[
[
"import datetime\n\n# print timestamps to see how long\n# sections take to run\n\nfirst_name = 'John'\nprint('task completed')\nprint(datetime.datetime.now())\nprint()\n\nfor x in range(0, 10):\n print(x)\nprint('task completed')\nprint(datetime.datetime.now())\nprint()",
"task completed\n2021-06-07 13:58:13.113387\n\n0\n1\n2\n3\n4\n5\n6\n7\n8\n9\ntask completed\n2021-06-07 13:58:13.115614\n\n"
]
],
[
[
"The above code can be rewritten using a function. You can define the function using `def` keyword, followed by the name of the function, and then a colon (`:`). Remember to use indentation which determines what code belongs to that function.",
"_____no_output_____"
]
],
[
[
"# import datetime class from datetime library\nfrom datetime import datetime\n\n# print the current time\ndef print_time():\n print('task completed')\n # no need for the extra datetime prefix\n # since the class is imported above\n print(datetime.now()) \n print()\n \nfirst_name = 'John'\nprint_time()\n\nfor x in range(0, 10):\n print(x)\nprint_time()",
"task completed\n2021-06-07 14:11:40.583077\n\n0\n1\n2\n3\n4\n5\n6\n7\n8\n9\ntask completed\n2021-06-07 14:11:40.589036\n\n"
]
],
[
[
"Sometimes when you copy/paste your code, we want to change some part of it. In the above example, what if you want to display a different message each time you run it. Say you want to display a specific message depending on the command you were running. This is where function parameters come in. *Parameters* or *arguments* are placed or defined within the parentheses of a function.",
"_____no_output_____"
]
],
[
[
"from datetime import datetime\n\n# print the current time and task name\ndef print_time(task_name):\n print(task_name)\n print(datetime.now())\n print()\n \nfirst_name = 'John'\n# pass in the task_name as a parameter\nprint_time('first name assigned')\n\nfor x in range(0, 10):\n print(x)\n# pass in the task_name as a parameter\nprint_time('loop completed')",
"first name assigned\n2021-06-07 16:07:57.077250\n\n0\n1\n2\n3\n4\n5\n6\n7\n8\n9\nloop completed\n2021-06-07 16:07:57.079687\n\n"
]
],
[
[
"Let's take another example where the code looks different but we're using the same logic. Suppose you're interested in getting initials for a user ID after the user enters their name.",
"_____no_output_____"
]
],
[
[
"first_name = input('Enter your first name: ')\n# get only the first letter of input\nfirst_name_initial = first_name[0:1]\n\nlast_name = input('Enter your last name: ')\nlast_name_initial = last_name[0:1]\n\nprint('Your initials are: ' + first_name_initial + last_name_initial)",
"Enter your first name: John \nEnter your last name: Wick\nYour initials are: JW\n"
]
],
[
[
"The above code can be written using a function.",
"_____no_output_____"
]
],
[
[
"def get_initial(name):\n initial = name[0:1]\n # the return function returns a value\n return initial\n\nfirst_name = input('Enter your first name: ')\nfirst_name_initial = get_initial(first_name)\n\nlast_name = input('Enter your last name: ')\nlast_name_initial = get_initial(last_name)\n\n# nested function in another call\nprint('Your initials are: ' + get_initial(first_name) + \n get_initial(last_name))",
"Enter your first name: john\nEnter your last name: wick\nYour initials are: jw\n"
]
],
[
[
"Functions can accept multiple parameters. In the above example, suppose you want to the user initials to only be uppercase for a user ID but lowercase for an email ID.",
"_____no_output_____"
]
],
[
[
"def get_initial(name, force_uppercase=True): # default to True\n if force_uppercase:\n initial = name[0:1].upper()\n else:\n initial = name[0:1]\n return initial\n\nfirst_name = input('Enter your first name: ')\nfirst_name_initial = get_initial(first_name)\n\nlast_name = input('Enter your last name: ')\nlast_name_initial = get_initial(last_name, False)\n\nprint('Your initials are: ' + first_name_initial + last_name_initial)",
"Enter your first name: john\nEnter your last name: wick\nYour initials are: Jw\n"
]
],
[
[
"When calling a function, you have to pass the parameters in the same order as when you defined the function. An exception to this is when you use named parameters, which offer better readability. \n`first_name_initial = get_initial(force_uppercase=True, name=first_name)`",
"_____no_output_____"
],
[
"Functions make the code more readable if you use good function names. They make the code less clunky. Always add comments to explain the purpose of your function. \nThe main advantage of functions is that if you ever need to change your function code, you only need to change it in one place. You also reduce rework and the chance to introduce bugs when you change the code you copied.",
"_____no_output_____"
]
]
] | [
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown"
] | [
[
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
]
] |
d0fe622972e5f7e0b27018a6df6f2c61cc8a8ae7 | 468,417 | ipynb | Jupyter Notebook | notebooks/fits2paperpng.ipynb | gijzelaerr/astro-pix2pix | 2ea7fd5c140bf05a77828a6672949fdd591541b7 | [
"MIT"
] | 2 | 2020-02-24T01:18:58.000Z | 2020-02-24T10:01:25.000Z | notebooks/fits2paperpng.ipynb | gijzelaerr/astro-pix2pix | 2ea7fd5c140bf05a77828a6672949fdd591541b7 | [
"MIT"
] | null | null | null | notebooks/fits2paperpng.ipynb | gijzelaerr/astro-pix2pix | 2ea7fd5c140bf05a77828a6672949fdd591541b7 | [
"MIT"
] | null | null | null | 3,061.54902 | 128,900 | 0.96163 | [
[
[
"import os\nimport matplotlib.pyplot as plt\nfrom astropy.io import fits\n%matplotlib inline",
"_____no_output_____"
],
[
"path = \".\"",
"_____no_output_____"
],
[
"def open_fits(f):\n return fits.open(f)[0].data.squeeze()",
"_____no_output_____"
],
[
"for i in [j for j in os.listdir(path) if j.endswith('.fits')]:\n data = open_fits(os.path.join(path, i))\n f, (a) = plt.subplots(1, 1, figsize=(6,6))\n a.pcolor(data, cmap='gist_heat')\n a.set_xticks([])\n a.set_yticks([])\n plt.savefig(os.path.join(path, i[:-5] + \".png\"), bbox_inches='tight')",
"_____no_output_____"
]
]
] | [
"code"
] | [
[
"code",
"code",
"code",
"code"
]
] |
d0fe67de414afe68cbfd167a2e0dfc1f964024f1 | 91,122 | ipynb | Jupyter Notebook | playground/mel/sentiment_classification_au_bow_non_DL.ipynb | mads-app-reviews-nlp/app-reviews-nlp | 4a9977c112dd81f57f30b0196d8184f8cd8097a2 | [
"MIT"
] | null | null | null | playground/mel/sentiment_classification_au_bow_non_DL.ipynb | mads-app-reviews-nlp/app-reviews-nlp | 4a9977c112dd81f57f30b0196d8184f8cd8097a2 | [
"MIT"
] | null | null | null | playground/mel/sentiment_classification_au_bow_non_DL.ipynb | mads-app-reviews-nlp/app-reviews-nlp | 4a9977c112dd81f57f30b0196d8184f8cd8097a2 | [
"MIT"
] | null | null | null | 41.400273 | 8,612 | 0.556693 | [
[
[
"## Sentiment Classification AU Reviews Data (BOW, non-Deep Learning)\n\nThis notebook covers two good approaches to perform sentiment classification - Naive Bayes and Logistic Regression. We will train AU reviews data on both.\n\nAs a rule of thumb, reviews that are 3 stars and above are **positive**, and vice versa.",
"_____no_output_____"
]
],
[
[
"%pip install spacy",
"Defaulting to user installation because normal site-packages is not writeable\nRequirement already satisfied: spacy in /home/meln/.local/lib/python3.8/site-packages (3.2.1)\nRequirement already satisfied: preshed<3.1.0,>=3.0.2 in /home/meln/.local/lib/python3.8/site-packages (from spacy) (3.0.6)\nRequirement already satisfied: spacy-loggers<2.0.0,>=1.0.0 in /home/meln/.local/lib/python3.8/site-packages (from spacy) (1.0.1)\nRequirement already satisfied: setuptools in /sw/arcts/centos7/python3.8-anaconda/2021.05/lib/python3.8/site-packages (from spacy) (52.0.0.post20210125)\nRequirement already satisfied: numpy>=1.15.0 in /sw/arcts/centos7/python3.8-anaconda/2021.05/lib/python3.8/site-packages (from spacy) (1.20.1)\nRequirement already satisfied: pydantic!=1.8,!=1.8.1,<1.9.0,>=1.7.4 in /home/meln/.local/lib/python3.8/site-packages (from spacy) (1.8.2)\nRequirement already satisfied: pathy>=0.3.5 in /home/meln/.local/lib/python3.8/site-packages (from spacy) (0.6.1)\nRequirement already satisfied: tqdm<5.0.0,>=4.38.0 in /sw/arcts/centos7/python3.8-anaconda/2021.05/lib/python3.8/site-packages (from spacy) (4.59.0)\nRequirement already satisfied: thinc<8.1.0,>=8.0.12 in /home/meln/.local/lib/python3.8/site-packages (from spacy) (8.0.13)\nRequirement already satisfied: jinja2 in /sw/arcts/centos7/python3.8-anaconda/2021.05/lib/python3.8/site-packages (from spacy) (2.11.3)\nRequirement already satisfied: srsly<3.0.0,>=2.4.1 in /home/meln/.local/lib/python3.8/site-packages (from spacy) (2.4.2)\nRequirement already satisfied: murmurhash<1.1.0,>=0.28.0 in /home/meln/.local/lib/python3.8/site-packages (from spacy) (1.0.6)\nRequirement already satisfied: requests<3.0.0,>=2.13.0 in /sw/arcts/centos7/python3.8-anaconda/2021.05/lib/python3.8/site-packages (from spacy) (2.25.1)\nRequirement already satisfied: wasabi<1.1.0,>=0.8.1 in /home/meln/.local/lib/python3.8/site-packages (from spacy) (0.9.0)\nRequirement already satisfied: catalogue<2.1.0,>=2.0.6 in /home/meln/.local/lib/python3.8/site-packages (from spacy) (2.0.6)\nRequirement already satisfied: blis<0.8.0,>=0.4.0 in /home/meln/.local/lib/python3.8/site-packages (from spacy) (0.7.5)\nRequirement already satisfied: langcodes<4.0.0,>=3.2.0 in /home/meln/.local/lib/python3.8/site-packages (from spacy) (3.3.0)\nRequirement already satisfied: spacy-legacy<3.1.0,>=3.0.8 in /home/meln/.local/lib/python3.8/site-packages (from spacy) (3.0.8)\nRequirement already satisfied: packaging>=20.0 in /sw/arcts/centos7/python3.8-anaconda/2021.05/lib/python3.8/site-packages (from spacy) (20.9)\nRequirement already satisfied: cymem<2.1.0,>=2.0.2 in /home/meln/.local/lib/python3.8/site-packages (from spacy) (2.0.6)\nRequirement already satisfied: typer<0.5.0,>=0.3.0 in /home/meln/.local/lib/python3.8/site-packages (from spacy) (0.4.0)\nRequirement already satisfied: pyparsing>=2.0.2 in /sw/arcts/centos7/python3.8-anaconda/2021.05/lib/python3.8/site-packages (from packaging>=20.0->spacy) (2.4.7)\nRequirement already satisfied: smart-open<6.0.0,>=5.0.0 in /home/meln/.local/lib/python3.8/site-packages (from pathy>=0.3.5->spacy) (5.2.1)\nRequirement already satisfied: typing-extensions>=3.7.4.3 in /sw/arcts/centos7/python3.8-anaconda/2021.05/lib/python3.8/site-packages (from pydantic!=1.8,!=1.8.1,<1.9.0,>=1.7.4->spacy) (3.7.4.3)\nRequirement already satisfied: certifi>=2017.4.17 in /sw/arcts/centos7/python3.8-anaconda/2021.05/lib/python3.8/site-packages (from requests<3.0.0,>=2.13.0->spacy) (2020.12.5)\nRequirement already satisfied: idna<3,>=2.5 in /sw/arcts/centos7/python3.8-anaconda/2021.05/lib/python3.8/site-packages (from requests<3.0.0,>=2.13.0->spacy) (2.10)\nRequirement already satisfied: urllib3<1.27,>=1.21.1 in /sw/arcts/centos7/python3.8-anaconda/2021.05/lib/python3.8/site-packages (from requests<3.0.0,>=2.13.0->spacy) (1.26.4)\nRequirement already satisfied: chardet<5,>=3.0.2 in /sw/arcts/centos7/python3.8-anaconda/2021.05/lib/python3.8/site-packages (from requests<3.0.0,>=2.13.0->spacy) (4.0.0)\nRequirement already satisfied: click<9.0.0,>=7.1.1 in /sw/arcts/centos7/python3.8-anaconda/2021.05/lib/python3.8/site-packages (from typer<0.5.0,>=0.3.0->spacy) (7.1.2)\nRequirement already satisfied: MarkupSafe>=0.23 in /sw/arcts/centos7/python3.8-anaconda/2021.05/lib/python3.8/site-packages (from jinja2->spacy) (1.1.1)\nNote: you may need to restart the kernel to use updated packages.\n"
],
[
"%pip install gensim",
"Defaulting to user installation because normal site-packages is not writeable\nRequirement already satisfied: gensim in /home/meln/.local/lib/python3.8/site-packages (4.1.2)\nRequirement already satisfied: numpy>=1.17.0 in /sw/arcts/centos7/python3.8-anaconda/2021.05/lib/python3.8/site-packages (from gensim) (1.20.1)\nRequirement already satisfied: smart-open>=1.8.1 in /home/meln/.local/lib/python3.8/site-packages (from gensim) (5.2.1)\nRequirement already satisfied: scipy>=0.18.1 in /sw/arcts/centos7/python3.8-anaconda/2021.05/lib/python3.8/site-packages (from gensim) (1.6.2)\nNote: you may need to restart the kernel to use updated packages.\n"
],
[
"%pip install spacy_langdetect",
"Defaulting to user installation because normal site-packages is not writeable\nRequirement already satisfied: spacy_langdetect in /home/meln/.local/lib/python3.8/site-packages (0.1.2)\nRequirement already satisfied: pytest in /sw/arcts/centos7/python3.8-anaconda/2021.05/lib/python3.8/site-packages (from spacy_langdetect) (6.2.3)\nRequirement already satisfied: langdetect==1.0.7 in /home/meln/.local/lib/python3.8/site-packages (from spacy_langdetect) (1.0.7)\nRequirement already satisfied: six in /sw/arcts/centos7/python3.8-anaconda/2021.05/lib/python3.8/site-packages (from langdetect==1.0.7->spacy_langdetect) (1.15.0)\nRequirement already satisfied: attrs>=19.2.0 in /sw/arcts/centos7/python3.8-anaconda/2021.05/lib/python3.8/site-packages (from pytest->spacy_langdetect) (20.3.0)\nRequirement already satisfied: iniconfig in /sw/arcts/centos7/python3.8-anaconda/2021.05/lib/python3.8/site-packages (from pytest->spacy_langdetect) (1.1.1)\nRequirement already satisfied: packaging in /sw/arcts/centos7/python3.8-anaconda/2021.05/lib/python3.8/site-packages (from pytest->spacy_langdetect) (20.9)\nRequirement already satisfied: pluggy<1.0.0a1,>=0.12 in /sw/arcts/centos7/python3.8-anaconda/2021.05/lib/python3.8/site-packages (from pytest->spacy_langdetect) (0.13.1)\nRequirement already satisfied: py>=1.8.2 in /sw/arcts/centos7/python3.8-anaconda/2021.05/lib/python3.8/site-packages (from pytest->spacy_langdetect) (1.10.0)\nRequirement already satisfied: toml in /sw/arcts/centos7/python3.8-anaconda/2021.05/lib/python3.8/site-packages (from pytest->spacy_langdetect) (0.10.2)\nRequirement already satisfied: pyparsing>=2.0.2 in /sw/arcts/centos7/python3.8-anaconda/2021.05/lib/python3.8/site-packages (from packaging->pytest->spacy_langdetect) (2.4.7)\nNote: you may need to restart the kernel to use updated packages.\n"
],
[
"!python3 -m spacy download en_core_web_sm",
"Defaulting to user installation because normal site-packages is not writeable\nCollecting en-core-web-sm==3.2.0\n Downloading https://github.com/explosion/spacy-models/releases/download/en_core_web_sm-3.2.0/en_core_web_sm-3.2.0-py3-none-any.whl (13.9 MB)\n\u001b[K |████████████████████████████████| 13.9 MB 16.9 MB/s eta 0:00:01\n\u001b[?25hRequirement already satisfied: spacy<3.3.0,>=3.2.0 in /home/meln/.local/lib/python3.8/site-packages (from en-core-web-sm==3.2.0) (3.2.1)\nRequirement already satisfied: packaging>=20.0 in /sw/arcts/centos7/python3.8-anaconda/2021.05/lib/python3.8/site-packages (from spacy<3.3.0,>=3.2.0->en-core-web-sm==3.2.0) (20.9)\nRequirement already satisfied: spacy-legacy<3.1.0,>=3.0.8 in /home/meln/.local/lib/python3.8/site-packages (from spacy<3.3.0,>=3.2.0->en-core-web-sm==3.2.0) (3.0.8)\nRequirement already satisfied: pathy>=0.3.5 in /home/meln/.local/lib/python3.8/site-packages (from spacy<3.3.0,>=3.2.0->en-core-web-sm==3.2.0) (0.6.1)\nRequirement already satisfied: thinc<8.1.0,>=8.0.12 in /home/meln/.local/lib/python3.8/site-packages (from spacy<3.3.0,>=3.2.0->en-core-web-sm==3.2.0) (8.0.13)\nRequirement already satisfied: wasabi<1.1.0,>=0.8.1 in /home/meln/.local/lib/python3.8/site-packages (from spacy<3.3.0,>=3.2.0->en-core-web-sm==3.2.0) (0.9.0)\nRequirement already satisfied: spacy-loggers<2.0.0,>=1.0.0 in /home/meln/.local/lib/python3.8/site-packages (from spacy<3.3.0,>=3.2.0->en-core-web-sm==3.2.0) (1.0.1)\nRequirement already satisfied: preshed<3.1.0,>=3.0.2 in /home/meln/.local/lib/python3.8/site-packages (from spacy<3.3.0,>=3.2.0->en-core-web-sm==3.2.0) (3.0.6)\nRequirement already satisfied: typer<0.5.0,>=0.3.0 in /home/meln/.local/lib/python3.8/site-packages (from spacy<3.3.0,>=3.2.0->en-core-web-sm==3.2.0) (0.4.0)\nRequirement already satisfied: murmurhash<1.1.0,>=0.28.0 in /home/meln/.local/lib/python3.8/site-packages (from spacy<3.3.0,>=3.2.0->en-core-web-sm==3.2.0) (1.0.6)\nRequirement already satisfied: catalogue<2.1.0,>=2.0.6 in /home/meln/.local/lib/python3.8/site-packages (from spacy<3.3.0,>=3.2.0->en-core-web-sm==3.2.0) (2.0.6)\nRequirement already satisfied: setuptools in /sw/arcts/centos7/python3.8-anaconda/2021.05/lib/python3.8/site-packages (from spacy<3.3.0,>=3.2.0->en-core-web-sm==3.2.0) (52.0.0.post20210125)\nRequirement already satisfied: pydantic!=1.8,!=1.8.1,<1.9.0,>=1.7.4 in /home/meln/.local/lib/python3.8/site-packages (from spacy<3.3.0,>=3.2.0->en-core-web-sm==3.2.0) (1.8.2)\nRequirement already satisfied: blis<0.8.0,>=0.4.0 in /home/meln/.local/lib/python3.8/site-packages (from spacy<3.3.0,>=3.2.0->en-core-web-sm==3.2.0) (0.7.5)\nRequirement already satisfied: langcodes<4.0.0,>=3.2.0 in /home/meln/.local/lib/python3.8/site-packages (from spacy<3.3.0,>=3.2.0->en-core-web-sm==3.2.0) (3.3.0)\nRequirement already satisfied: numpy>=1.15.0 in /sw/arcts/centos7/python3.8-anaconda/2021.05/lib/python3.8/site-packages (from spacy<3.3.0,>=3.2.0->en-core-web-sm==3.2.0) (1.20.1)\nRequirement already satisfied: jinja2 in /sw/arcts/centos7/python3.8-anaconda/2021.05/lib/python3.8/site-packages (from spacy<3.3.0,>=3.2.0->en-core-web-sm==3.2.0) (2.11.3)\nRequirement already satisfied: cymem<2.1.0,>=2.0.2 in /home/meln/.local/lib/python3.8/site-packages (from spacy<3.3.0,>=3.2.0->en-core-web-sm==3.2.0) (2.0.6)\nRequirement already satisfied: requests<3.0.0,>=2.13.0 in /sw/arcts/centos7/python3.8-anaconda/2021.05/lib/python3.8/site-packages (from spacy<3.3.0,>=3.2.0->en-core-web-sm==3.2.0) (2.25.1)\nRequirement already satisfied: tqdm<5.0.0,>=4.38.0 in /sw/arcts/centos7/python3.8-anaconda/2021.05/lib/python3.8/site-packages (from spacy<3.3.0,>=3.2.0->en-core-web-sm==3.2.0) (4.59.0)\nRequirement already satisfied: srsly<3.0.0,>=2.4.1 in /home/meln/.local/lib/python3.8/site-packages (from spacy<3.3.0,>=3.2.0->en-core-web-sm==3.2.0) (2.4.2)\nRequirement already satisfied: pyparsing>=2.0.2 in /sw/arcts/centos7/python3.8-anaconda/2021.05/lib/python3.8/site-packages (from packaging>=20.0->spacy<3.3.0,>=3.2.0->en-core-web-sm==3.2.0) (2.4.7)\nRequirement already satisfied: smart-open<6.0.0,>=5.0.0 in /home/meln/.local/lib/python3.8/site-packages (from pathy>=0.3.5->spacy<3.3.0,>=3.2.0->en-core-web-sm==3.2.0) (5.2.1)\nRequirement already satisfied: typing-extensions>=3.7.4.3 in /sw/arcts/centos7/python3.8-anaconda/2021.05/lib/python3.8/site-packages (from pydantic!=1.8,!=1.8.1,<1.9.0,>=1.7.4->spacy<3.3.0,>=3.2.0->en-core-web-sm==3.2.0) (3.7.4.3)\nRequirement already satisfied: chardet<5,>=3.0.2 in /sw/arcts/centos7/python3.8-anaconda/2021.05/lib/python3.8/site-packages (from requests<3.0.0,>=2.13.0->spacy<3.3.0,>=3.2.0->en-core-web-sm==3.2.0) (4.0.0)\nRequirement already satisfied: urllib3<1.27,>=1.21.1 in /sw/arcts/centos7/python3.8-anaconda/2021.05/lib/python3.8/site-packages (from requests<3.0.0,>=2.13.0->spacy<3.3.0,>=3.2.0->en-core-web-sm==3.2.0) (1.26.4)\nRequirement already satisfied: idna<3,>=2.5 in /sw/arcts/centos7/python3.8-anaconda/2021.05/lib/python3.8/site-packages (from requests<3.0.0,>=2.13.0->spacy<3.3.0,>=3.2.0->en-core-web-sm==3.2.0) (2.10)\nRequirement already satisfied: certifi>=2017.4.17 in /sw/arcts/centos7/python3.8-anaconda/2021.05/lib/python3.8/site-packages (from requests<3.0.0,>=2.13.0->spacy<3.3.0,>=3.2.0->en-core-web-sm==3.2.0) (2020.12.5)\nRequirement already satisfied: click<9.0.0,>=7.1.1 in /sw/arcts/centos7/python3.8-anaconda/2021.05/lib/python3.8/site-packages (from typer<0.5.0,>=0.3.0->spacy<3.3.0,>=3.2.0->en-core-web-sm==3.2.0) (7.1.2)\nRequirement already satisfied: MarkupSafe>=0.23 in /sw/arcts/centos7/python3.8-anaconda/2021.05/lib/python3.8/site-packages (from jinja2->spacy<3.3.0,>=3.2.0->en-core-web-sm==3.2.0) (1.1.1)\n\u001b[38;5;2m✔ Download and installation successful\u001b[0m\nYou can now load the package via spacy.load('en_core_web_sm')\n"
],
[
"import gzip\nimport json\nimport matplotlib.pyplot as plt\nimport numpy as np\nimport re\nimport random\nimport pandas as pd\nimport seaborn as sns\nimport gensim\nimport spacy\nfrom collections import Counter, defaultdict\nfrom sklearn.dummy import DummyClassifier\nfrom sklearn.feature_extraction.text import TfidfVectorizer\nfrom sklearn.linear_model import LogisticRegression\nfrom sklearn.naive_bayes import BernoulliNB, ComplementNB, MultinomialNB\nfrom sklearn.metrics import f1_score, classification_report, accuracy_score, confusion_matrix\nfrom sklearn.model_selection import train_test_split, GridSearchCV\nfrom spacy_langdetect import LanguageDetector\nfrom spacy.language import Language\nfrom gensim.models.word2vec import Word2Vec\nfrom nltk.tokenize import sent_tokenize, word_tokenize\nfrom tqdm import tqdm",
"_____no_output_____"
],
[
"RANDOM_SEED = 33",
"_____no_output_____"
],
[
"reviews = pd.read_pickle(\"assets/au_reviews.pkl\")\nreviews.head()",
"_____no_output_____"
],
[
"reviews['label'] = np.where(reviews['rating'] >= 3, 0, 1)",
"_____no_output_____"
],
[
"reviews.head()",
"_____no_output_____"
]
],
[
[
"## 1. Data Processing",
"_____no_output_____"
],
[
"Check the dataset size:",
"_____no_output_____"
]
],
[
[
"print(len(reviews))",
"626377\n"
]
],
[
[
"And the type of apps:",
"_____no_output_____"
]
],
[
[
"app_list = list(reviews['app'].unique())\napp_list",
"_____no_output_____"
]
],
[
[
"Let's also get a sense of our dataset's balance",
"_____no_output_____"
]
],
[
[
"reviews['label'].value_counts(normalize=True)",
"_____no_output_____"
],
[
"# By app\n\nfor app in app_list:\n print(reviews[reviews['app'] == app]['label'].value_counts(normalize=True))",
"0 0.760423\n1 0.239577\nName: label, dtype: float64\n0 0.648032\n1 0.351968\nName: label, dtype: float64\n0 0.651189\n1 0.348811\nName: label, dtype: float64\n0 0.682461\n1 0.317539\nName: label, dtype: float64\n0 0.56053\n1 0.43947\nName: label, dtype: float64\n"
]
],
[
[
"Across the board the distribution of positive and negative reviews are quite consistent between the apps. Overall, there's an imbalance in our dataset, with positive reviews making for 75% of the dataset. Let's also check for null values.",
"_____no_output_____"
]
],
[
[
"reviews.isnull().sum()",
"_____no_output_____"
],
[
"reviews = reviews.dropna()",
"_____no_output_____"
],
[
"df_proc = reviews.copy()\ndf_proc.drop(columns=['date', 'rating', 'app'], inplace=True)\ndf_proc.head()",
"_____no_output_____"
]
],
[
[
"For AU dataset we won't be filtering out non-English reviews. It's likely that this makes up for a very small proportion of the dataset.",
"_____no_output_____"
]
],
[
[
"df_proc.to_csv('reviews_au_filtered.csv')",
"_____no_output_____"
],
[
"X = df_proc['review']\ny = df_proc['label']",
"_____no_output_____"
]
],
[
[
"We will split the dataset into `train`, `test`, and `dev`, with 80%, 10%, 10% ratio, respectively.",
"_____no_output_____"
]
],
[
[
"X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=RANDOM_SEED)\nX_test, X_dev, y_test, y_dev = train_test_split(X_test, y_test, test_size=0.5, random_state=RANDOM_SEED)",
"_____no_output_____"
],
[
"len(X_train)",
"_____no_output_____"
],
[
"len(X_dev)",
"_____no_output_____"
],
[
"len(X_test)",
"_____no_output_____"
],
[
"X_train.iloc[0]",
"_____no_output_____"
],
[
"X_test.iloc[0]",
"_____no_output_____"
]
],
[
[
"## 2.1 Bag-of-Words Approach on Naive Bayes & Logistic Regression\n\nThis section explores the use of bag of words as feature extraction. But first, let's have a look at the token frequencies.",
"_____no_output_____"
]
],
[
[
"# Fill this with any token (with anything in it!) for tokens separated by whitespace\nws_tokens = Counter()\n\n# Fill this one with tokens separated by whitespace but constisting only of tokens\n# that are totally made of alphanumeric characters (you can use the \\w character\n# class in making the regex)\n\nalpha_ws_tokens = Counter()\n\n# Fill this one with the tokens separated by *word boundaries* (not white space) that consist\n# of alphanumeric characters (use \\w again)\nalpha_re_tokens = Counter()\nfor review in tqdm(X_train):\n ws_review = review.split()\n ws_tokens.update(ws_review)\n # Note: use fullmatch() as it anchor both the start and end of str. match() won't work.\n alpha_ws_tokens.update([re.fullmatch(r'\\w+', word).group() for word in ws_review if re.fullmatch(r'\\w+', word) != None])\n alpha_re_tokens.update(re.findall(r'\\w+', review))",
"100%|██████████| 501058/501058 [00:20<00:00, 24120.74it/s]\n"
],
[
"print(len(ws_tokens))\nprint(len(alpha_ws_tokens))\nprint(len(alpha_re_tokens))",
"221265\n89320\n103276\n"
],
[
"top_100 = alpha_re_tokens.most_common(100)\ntop_100",
"_____no_output_____"
]
],
[
[
"Lots of stopwords, as expected.",
"_____no_output_____"
]
],
[
[
"x = list(range(100))\ny = [word_tup[1] for word_tup in top_100]",
"_____no_output_____"
],
[
"ax = plt.plot(x, y, '.')\n#plt.yscale('log')\n#plt.xscale('log')",
"_____no_output_____"
]
],
[
[
"And unexpectedly, the word frequency distribution also follows Zipf's law as well. What that means is that we can essentially remove uncommon words, without worrying that they will affect performance. We will also need to remove stopwords, and add unigrams and bigrams as features.",
"_____no_output_____"
]
],
[
[
"vectorizer = TfidfVectorizer(stop_words='english', min_df=500, ngram_range=(1,2))\nX_train_bow = vectorizer.fit_transform(X_train)",
"_____no_output_____"
],
[
"print(X_train_bow.shape)",
"(501058, 1389)\n"
],
[
"X_dev_bow = vectorizer.transform(X_dev)",
"_____no_output_____"
],
[
"def train_model(clf):\n print(\"_\" * 80)\n print(\"Training: \")\n clf.fit(X_train_bow, y_train)\n y_dev_pred = clf.predict(X_dev_bow)\n \n score = accuracy_score(y_dev, y_dev_pred)\n print(\"accuracy: %0.3f\" % score)\n \n print(\"classification report:\")\n print(classification_report(y_dev, y_dev_pred))\n \n print(\"confusion matrix:\")\n print(confusion_matrix(y_dev, y_dev_pred))\n print(\"Training Complete\")\n print()\n \n clf_descr = str(clf).split(\"(\")[0]\n return clf_descr, score, y_dev_pred",
"_____no_output_____"
]
],
[
[
"It's training time! We'll start with a few dummy classifiers, followed by Naive Bayes and Logistic Regression.",
"_____no_output_____"
]
],
[
[
"for clf, name in (\n (DummyClassifier(strategy='uniform', random_state=RANDOM_SEED), \"Uniform Classifier\"),\n (DummyClassifier(strategy='most_frequent', random_state=RANDOM_SEED), \"Most Frequent Classifier\"),\n):\n print(\"=\" * 80)\n print(\"Training Results - Dummy Classifiers\")\n print(name)\n mod = train_model(clf)",
"================================================================================\nTraining Results - Dummy Classifiers\nUniform Classifier\n________________________________________________________________________________\nTraining: \naccuracy: 0.499\nclassification report:\n precision recall f1-score support\n\n 0 0.66 0.50 0.57 41413\n 1 0.34 0.50 0.40 21220\n\n accuracy 0.50 62633\n macro avg 0.50 0.50 0.49 62633\nweighted avg 0.55 0.50 0.51 62633\n\nconfusion matrix:\n[[20678 20735]\n [10627 10593]]\nTraining Complete\n\n================================================================================\nTraining Results - Dummy Classifiers\nMost Frequent Classifier\n________________________________________________________________________________\nTraining: \naccuracy: 0.661\nclassification report:\n precision recall f1-score support\n\n 0 0.66 1.00 0.80 41413\n 1 0.00 0.00 0.00 21220\n\n accuracy 0.66 62633\n macro avg 0.33 0.50 0.40 62633\nweighted avg 0.44 0.66 0.53 62633\n\nconfusion matrix:\n[[41413 0]\n [21220 0]]\nTraining Complete\n\n"
],
[
"preds = {} # A dict to store our dev set predictions\n\nfor clf, name in (\n (MultinomialNB(alpha=0.01), \"MultinomialNB\"),\n (BernoulliNB(alpha=0.01), \"BernoulliNB\"),\n (ComplementNB(alpha=0.1), \"ComplementNB\"),\n (LogisticRegression(solver='lbfgs', class_weight='balanced', max_iter=1000, random_state=RANDOM_SEED), \"Logistic Regression\")\n):\n print(\"=\" * 80)\n print(\"Training Results - Naive Bayes & LogReg\")\n print(name)\n mod = train_model(clf)\n preds[name] = mod[2]",
"================================================================================\nTraining Results - Naive Bayes & LogReg\nMultinomialNB\n________________________________________________________________________________\nTraining: \naccuracy: 0.899\nclassification report:\n precision recall f1-score support\n\n 0 0.93 0.91 0.92 41413\n 1 0.84 0.87 0.85 21220\n\n accuracy 0.90 62633\n macro avg 0.88 0.89 0.89 62633\nweighted avg 0.90 0.90 0.90 62633\n\nconfusion matrix:\n[[37812 3601]\n [ 2724 18496]]\nTraining Complete\n\n================================================================================\nTraining Results - Naive Bayes & LogReg\nBernoulliNB\n________________________________________________________________________________\nTraining: \naccuracy: 0.853\nclassification report:\n precision recall f1-score support\n\n 0 0.86 0.92 0.89 41413\n 1 0.82 0.72 0.77 21220\n\n accuracy 0.85 62633\n macro avg 0.84 0.82 0.83 62633\nweighted avg 0.85 0.85 0.85 62633\n\nconfusion matrix:\n[[38185 3228]\n [ 6005 15215]]\nTraining Complete\n\n================================================================================\nTraining Results - Naive Bayes & LogReg\nComplementNB\n________________________________________________________________________________\nTraining: \naccuracy: 0.885\nclassification report:\n precision recall f1-score support\n\n 0 0.96 0.87 0.91 41413\n 1 0.78 0.92 0.85 21220\n\n accuracy 0.89 62633\n macro avg 0.87 0.89 0.88 62633\nweighted avg 0.90 0.89 0.89 62633\n\nconfusion matrix:\n[[35873 5540]\n [ 1641 19579]]\nTraining Complete\n\n================================================================================\nTraining Results - Naive Bayes & LogReg\nLogistic Regression\n________________________________________________________________________________\nTraining: \naccuracy: 0.902\nclassification report:\n precision recall f1-score support\n\n 0 0.95 0.89 0.92 41413\n 1 0.82 0.92 0.86 21220\n\n accuracy 0.90 62633\n macro avg 0.89 0.91 0.89 62633\nweighted avg 0.91 0.90 0.90 62633\n\nconfusion matrix:\n[[37061 4352]\n [ 1765 19455]]\nTraining Complete\n\n"
]
],
[
[
"The baseline results are quite promising, with both MultinomialNB and Logistic Regression achieving 0.85 on macro F1-score. This means that the bag-of-word approach is a rather solid approach for sentiment classification. It's also interesting to see that while MultinomialNB has a rather balanced number of false pos and false neg, BernoulliNB and ComplementNB are different. BernoulliNB has a much higher number of fps, while ComplementNB has a much higher number of fns.\n\nAlso, according to https://web.stanford.edu/~jurafsky/slp3/4.pdf, using binary NB (BernoulliNB) may improve predictive performance, as whether a word occurs or not seems to matter more than its frequency. But in this case, BernoulliNB does not outperform other Naive Bayes methods. We'll come back to this in a second.\n\nLet's take a look at a few of mis-classifications for both Naive Bayes and Logistic Regression.",
"_____no_output_____"
]
],
[
[
"# Create a dataframe for mis-classifications\ndef create_mis_classification_df(name):\n mis_class = pd.DataFrame(X_dev)\n mis_class['Actual'] = y_dev\n mis_class['Predicted'] = preds[name]\n mis_class = mis_class[mis_class['Actual'] != mis_class['Predicted']]\n return mis_class",
"_____no_output_____"
],
[
"mis_class_multi = create_mis_classification_df('MultinomialNB')\nmis_class_bernoulli = create_mis_classification_df('BernoulliNB')\nmis_class_complement = create_mis_classification_df('ComplementNB')\nmis_class_logreg = create_mis_classification_df('Logistic Regression')",
"_____no_output_____"
],
[
"mis_class_multi.sample(10).values",
"_____no_output_____"
],
[
"mis_class_bernoulli.sample(10).values",
"_____no_output_____"
],
[
"mis_class_complement.sample(10).values",
"_____no_output_____"
],
[
"mis_class_logreg.sample(10).values",
"_____no_output_____"
]
],
[
[
"Really interesting. Looking at the results, there are a few cases of mis-classifications:\n- Reviews that contain negation expressions, eg \"not okay\" is classified as a positive review when in reality it should be negative. BoW approach makes it hard for ML model to recognize this kind of expressions.\n- Reviews that are mis-classified due to rating. Eg a customer may write something negative but give 3 stars. It's tricky in this case because it's a caveat of our dataset.\n- Some mis-classification is the ML model being weirdly off, eg ComplementNB classified a \"Good\" review as negative, or reviews containing the word 'hate' gets classified as positive.\n- Contextual awareness is important and this is something that bag-of-word approaches cannot address. For example the sentence 'Very good. Expensive delivery charge though' gets classified as negative likely because of the word expensive, while in reality this is a positive review.",
"_____no_output_____"
],
[
"### 2.1 Reduce min_df\n\nWe set a min frequency cap of 500. What happens if we reduce this cap to 100?",
"_____no_output_____"
]
],
[
[
"vectorizer = TfidfVectorizer(stop_words='english', min_df=100, ngram_range=(1,2))\nX_train_bow = vectorizer.fit_transform(X_train)\nX_dev_bow = vectorizer.transform(X_dev)",
"_____no_output_____"
],
[
"preds = {} # A dict to store our dev set predictions\n\nfor clf, name in (\n (MultinomialNB(alpha=0.01), \"MultinomialNB\"),\n (BernoulliNB(alpha=0.01), \"BernoulliNB\"),\n (ComplementNB(alpha=0.1), \"ComplementNB\"),\n (LogisticRegression(solver='lbfgs', class_weight='balanced', max_iter=1000, random_state=RANDOM_SEED), \"Logistic Regression\")\n):\n print(\"=\" * 80)\n print(\"Training Results - Naive Bayes & LogReg\")\n print(name)\n mod = train_model(clf)\n preds[name] = mod[2]",
"================================================================================\nTraining Results - Naive Bayes & LogReg\nMultinomialNB\n________________________________________________________________________________\nTraining: \naccuracy: 0.907\nclassification report:\n precision recall f1-score support\n\n 0 0.94 0.91 0.93 41413\n 1 0.84 0.89 0.87 21220\n\n accuracy 0.91 62633\n macro avg 0.89 0.90 0.90 62633\nweighted avg 0.91 0.91 0.91 62633\n\nconfusion matrix:\n[[37866 3547]\n [ 2276 18944]]\nTraining Complete\n\n================================================================================\nTraining Results - Naive Bayes & LogReg\nBernoulliNB\n________________________________________________________________________________\nTraining: \naccuracy: 0.856\nclassification report:\n precision recall f1-score support\n\n 0 0.87 0.92 0.89 41413\n 1 0.83 0.72 0.77 21220\n\n accuracy 0.86 62633\n macro avg 0.85 0.82 0.83 62633\nweighted avg 0.85 0.86 0.85 62633\n\nconfusion matrix:\n[[38298 3115]\n [ 5888 15332]]\nTraining Complete\n\n================================================================================\nTraining Results - Naive Bayes & LogReg\nComplementNB\n________________________________________________________________________________\nTraining: \naccuracy: 0.896\nclassification report:\n precision recall f1-score support\n\n 0 0.96 0.87 0.92 41413\n 1 0.79 0.94 0.86 21220\n\n accuracy 0.90 62633\n macro avg 0.88 0.91 0.89 62633\nweighted avg 0.91 0.90 0.90 62633\n\nconfusion matrix:\n[[36232 5181]\n [ 1343 19877]]\nTraining Complete\n\n================================================================================\nTraining Results - Naive Bayes & LogReg\nLogistic Regression\n________________________________________________________________________________\nTraining: \naccuracy: 0.913\nclassification report:\n precision recall f1-score support\n\n 0 0.96 0.90 0.93 41413\n 1 0.83 0.93 0.88 21220\n\n accuracy 0.91 62633\n macro avg 0.90 0.92 0.90 62633\nweighted avg 0.92 0.91 0.91 62633\n\nconfusion matrix:\n[[37458 3955]\n [ 1517 19703]]\nTraining Complete\n\n"
]
],
[
[
"Slight bump in performance, but at the expense of longer training time. ",
"_____no_output_____"
],
[
"### 2.2 Not use stopwords removal",
"_____no_output_____"
]
],
[
[
"vectorizer = TfidfVectorizer(min_df=100, ngram_range=(1,2))\nX_train_bow = vectorizer.fit_transform(X_train)\nX_dev_bow = vectorizer.transform(X_dev)",
"_____no_output_____"
],
[
"preds = {} # A dict to store our dev set predictions\n\nfor clf, name in (\n (MultinomialNB(alpha=0.01), \"MultinomialNB\"),\n (BernoulliNB(alpha=0.01), \"BernoulliNB\"),\n (ComplementNB(alpha=0.1), \"ComplementNB\"),\n (LogisticRegression(solver='lbfgs', class_weight='balanced', max_iter=1000, random_state=RANDOM_SEED), \"Logistic Regression\")\n):\n print(\"=\" * 80)\n print(\"Training Results - Naive Bayes & LogReg\")\n print(name)\n mod = train_model(clf)\n preds[name] = mod[2]",
"================================================================================\nTraining Results - Naive Bayes & LogReg\nMultinomialNB\n________________________________________________________________________________\nTraining: \naccuracy: 0.912\nclassification report:\n precision recall f1-score support\n\n 0 0.95 0.91 0.93 41413\n 1 0.84 0.91 0.88 21220\n\n accuracy 0.91 62633\n macro avg 0.90 0.91 0.90 62633\nweighted avg 0.92 0.91 0.91 62633\n\nconfusion matrix:\n[[37816 3597]\n [ 1903 19317]]\nTraining Complete\n\n================================================================================\nTraining Results - Naive Bayes & LogReg\nBernoulliNB\n________________________________________________________________________________\nTraining: \naccuracy: 0.828\nclassification report:\n precision recall f1-score support\n\n 0 0.85 0.91 0.87 41413\n 1 0.79 0.68 0.73 21220\n\n accuracy 0.83 62633\n macro avg 0.82 0.79 0.80 62633\nweighted avg 0.83 0.83 0.82 62633\n\nconfusion matrix:\n[[37550 3863]\n [ 6884 14336]]\nTraining Complete\n\n================================================================================\nTraining Results - Naive Bayes & LogReg\nComplementNB\n________________________________________________________________________________\nTraining: \naccuracy: 0.902\nclassification report:\n precision recall f1-score support\n\n 0 0.97 0.88 0.92 41413\n 1 0.80 0.95 0.87 21220\n\n accuracy 0.90 62633\n macro avg 0.88 0.91 0.89 62633\nweighted avg 0.91 0.90 0.90 62633\n\nconfusion matrix:\n[[36419 4994]\n [ 1161 20059]]\nTraining Complete\n\n================================================================================\nTraining Results - Naive Bayes & LogReg\nLogistic Regression\n________________________________________________________________________________\nTraining: \naccuracy: 0.926\nclassification report:\n precision recall f1-score support\n\n 0 0.97 0.92 0.94 41413\n 1 0.86 0.94 0.90 21220\n\n accuracy 0.93 62633\n macro avg 0.91 0.93 0.92 62633\nweighted avg 0.93 0.93 0.93 62633\n\nconfusion matrix:\n[[38046 3367]\n [ 1289 19931]]\nTraining Complete\n\n"
]
],
[
[
"Performance improved, surprisingly.",
"_____no_output_____"
],
[
"### 2.3 Set max_df",
"_____no_output_____"
]
],
[
[
"vectorizer = TfidfVectorizer(min_df=100, max_df=5000, ngram_range=(1,2))\nX_train_bow = vectorizer.fit_transform(X_train)\nX_dev_bow = vectorizer.transform(X_dev)",
"_____no_output_____"
],
[
"preds = {} # A dict to store our dev set predictions\n\nfor clf, name in (\n (MultinomialNB(alpha=0.01), \"MultinomialNB\"),\n (BernoulliNB(alpha=0.01), \"BernoulliNB\"),\n (ComplementNB(alpha=0.1), \"ComplementNB\"),\n (LogisticRegression(solver='lbfgs', class_weight='balanced', max_iter=1000, random_state=RANDOM_SEED), \"Logistic Regression\")\n):\n print(\"=\" * 80)\n print(\"Training Results - Naive Bayes & LogReg\")\n print(name)\n mod = train_model(clf)\n preds[name] = mod[2]",
"================================================================================\nTraining Results - Naive Bayes & LogReg\nMultinomialNB\n________________________________________________________________________________\nTraining: \naccuracy: 0.911\nclassification report:\n precision recall f1-score support\n\n 0 0.93 0.93 0.93 41413\n 1 0.87 0.86 0.87 21220\n\n accuracy 0.91 62633\n macro avg 0.90 0.90 0.90 62633\nweighted avg 0.91 0.91 0.91 62633\n\nconfusion matrix:\n[[38713 2700]\n [ 2873 18347]]\nTraining Complete\n\n================================================================================\nTraining Results - Naive Bayes & LogReg\nBernoulliNB\n________________________________________________________________________________\nTraining: \naccuracy: 0.842\nclassification report:\n precision recall f1-score support\n\n 0 0.85 0.92 0.89 41413\n 1 0.82 0.68 0.75 21220\n\n accuracy 0.84 62633\n macro avg 0.84 0.80 0.82 62633\nweighted avg 0.84 0.84 0.84 62633\n\nconfusion matrix:\n[[38219 3194]\n [ 6715 14505]]\nTraining Complete\n\n================================================================================\nTraining Results - Naive Bayes & LogReg\nComplementNB\n________________________________________________________________________________\nTraining: \naccuracy: 0.903\nclassification report:\n precision recall f1-score support\n\n 0 0.95 0.90 0.92 41413\n 1 0.82 0.92 0.87 21220\n\n accuracy 0.90 62633\n macro avg 0.89 0.91 0.90 62633\nweighted avg 0.91 0.90 0.90 62633\n\nconfusion matrix:\n[[37149 4264]\n [ 1782 19438]]\nTraining Complete\n\n================================================================================\nTraining Results - Naive Bayes & LogReg\nLogistic Regression\n________________________________________________________________________________\nTraining: \naccuracy: 0.919\nclassification report:\n precision recall f1-score support\n\n 0 0.95 0.92 0.94 41413\n 1 0.86 0.91 0.88 21220\n\n accuracy 0.92 62633\n macro avg 0.91 0.92 0.91 62633\nweighted avg 0.92 0.92 0.92 62633\n\nconfusion matrix:\n[[38277 3136]\n [ 1956 19264]]\nTraining Complete\n\n"
]
],
[
[
"Not much changes from stopwords removal.",
"_____no_output_____"
],
[
"### 2.4 Clip frequency at 1\n\nEarlier, we see that there isn't any performance with BernoulliNB as compared to other methods. But what if we clip the frequency at the feature processing level? Luckily, sklearn tfidf has a `binary` parameters that allows us to do that.",
"_____no_output_____"
]
],
[
[
"vectorizer = TfidfVectorizer(binary=True, ngram_range=(1,1))\nX_train_bow = vectorizer.fit_transform(X_train)\nX_dev_bow = vectorizer.transform(X_dev)",
"_____no_output_____"
],
[
"preds = {} # A dict to store our dev set predictions\n\nfor clf, name in (\n (MultinomialNB(alpha=0.01), \"MultinomialNB\"),\n (BernoulliNB(alpha=0.01), \"BernoulliNB\"),\n (ComplementNB(alpha=0.1), \"ComplementNB\"),\n (LogisticRegression(solver='lbfgs', class_weight='balanced', max_iter=1000, random_state=RANDOM_SEED), \"Logistic Regression\")\n):\n print(\"=\" * 80)\n print(\"Training Results - Naive Bayes & LogReg\")\n print(name)\n mod = train_model(clf)\n preds[name] = mod[2]",
"================================================================================\nTraining Results - Naive Bayes & LogReg\nMultinomialNB\n________________________________________________________________________________\nTraining: \naccuracy: 0.901\nclassification report:\n precision recall f1-score support\n\n 0 0.94 0.91 0.92 41413\n 1 0.83 0.89 0.86 21220\n\n accuracy 0.90 62633\n macro avg 0.89 0.90 0.89 62633\nweighted avg 0.90 0.90 0.90 62633\n\nconfusion matrix:\n[[37626 3787]\n [ 2388 18832]]\nTraining Complete\n\n================================================================================\nTraining Results - Naive Bayes & LogReg\nBernoulliNB\n________________________________________________________________________________\nTraining: \naccuracy: 0.823\nclassification report:\n precision recall f1-score support\n\n 0 0.85 0.89 0.87 41413\n 1 0.77 0.68 0.72 21220\n\n accuracy 0.82 62633\n macro avg 0.81 0.79 0.80 62633\nweighted avg 0.82 0.82 0.82 62633\n\nconfusion matrix:\n[[37021 4392]\n [ 6703 14517]]\nTraining Complete\n\n================================================================================\nTraining Results - Naive Bayes & LogReg\nComplementNB\n________________________________________________________________________________\nTraining: \naccuracy: 0.889\nclassification report:\n precision recall f1-score support\n\n 0 0.97 0.86 0.91 41413\n 1 0.78 0.94 0.85 21220\n\n accuracy 0.89 62633\n macro avg 0.87 0.90 0.88 62633\nweighted avg 0.90 0.89 0.89 62633\n\nconfusion matrix:\n[[35673 5740]\n [ 1209 20011]]\nTraining Complete\n\n================================================================================\nTraining Results - Naive Bayes & LogReg\nLogistic Regression\n________________________________________________________________________________\nTraining: \naccuracy: 0.921\nclassification report:\n precision recall f1-score support\n\n 0 0.97 0.91 0.94 41413\n 1 0.85 0.94 0.89 21220\n\n accuracy 0.92 62633\n macro avg 0.91 0.93 0.91 62633\nweighted avg 0.93 0.92 0.92 62633\n\nconfusion matrix:\n[[37816 3597]\n [ 1323 19897]]\nTraining Complete\n\n"
]
],
[
[
"### 2.5 Tune Logistic Regression Params\n\nSo far, LogReg performs the best in terms of macro F1 score. In this section, we'll try tuning the performance of Logistic Regression, using the best Tfidf tuning result above.",
"_____no_output_____"
]
],
[
[
"vectorizer = TfidfVectorizer(min_df=100, ngram_range=(1,2))\nX_train_bow = vectorizer.fit_transform(X_train)\nX_dev_bow = vectorizer.transform(X_dev)",
"_____no_output_____"
],
[
"clf = LogisticRegression(solver='lbfgs', class_weight='balanced', max_iter=3000, random_state=RANDOM_SEED)\nparam_grid = {'C' : np.logspace(-4, 4, 20)}\n\n\nprint(\"=\" * 80)\nprint(\"LogReg Grid Search\")\nclf = GridSearchCV(clf, param_grid = param_grid, cv = 5, verbose=True, n_jobs=-1)\nbest_clf = clf.fit(X_train_bow, y_train)\nprint(clf.best_params_)\ny_dev_pred = clf.predict(X_dev_bow)\nscore = accuracy_score(y_dev, y_dev_pred)\nprint(\"accuracy: %0.3f\" % score)\nprint(\"classification report:\")\nprint(classification_report(y_dev, y_dev_pred))\nprint(\"confusion matrix:\")\nprint(confusion_matrix(y_dev, y_dev_pred))\nprint(\"Training Complete\")\nprint()",
"================================================================================\nLogReg Grid Search\nFitting 5 folds for each of 20 candidates, totalling 100 fits\n{'C': 1.623776739188721}\naccuracy: 0.926\nclassification report:\n precision recall f1-score support\n\n 0 0.97 0.92 0.94 41413\n 1 0.86 0.94 0.90 21220\n\n accuracy 0.93 62633\n macro avg 0.91 0.93 0.92 62633\nweighted avg 0.93 0.93 0.93 62633\n\nconfusion matrix:\n[[38078 3335]\n [ 1298 19922]]\nTraining Complete\n\n"
]
]
] | [
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] | [
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
]
] |
d0fe755d3bdbdc0ec627d59c030f36ffb1540419 | 360,272 | ipynb | Jupyter Notebook | Week 3+4/Starter_Code/.ipynb_checkpoints/whale_analysis-Copy6-checkpoint.ipynb | maricheklin/python-stock-analysis-base | a5f0fc3ab1de93291fc23cc4268ecf4094ec3bb9 | [
"MIT"
] | null | null | null | Week 3+4/Starter_Code/.ipynb_checkpoints/whale_analysis-Copy6-checkpoint.ipynb | maricheklin/python-stock-analysis-base | a5f0fc3ab1de93291fc23cc4268ecf4094ec3bb9 | [
"MIT"
] | null | null | null | Week 3+4/Starter_Code/.ipynb_checkpoints/whale_analysis-Copy6-checkpoint.ipynb | maricheklin/python-stock-analysis-base | a5f0fc3ab1de93291fc23cc4268ecf4094ec3bb9 | [
"MIT"
] | null | null | null | 85.922251 | 83,120 | 0.787341 | [
[
[
" # A Whale off the Port(folio)\n ---\n\n In this assignment, you'll get to use what you've learned this week to evaluate the performance among various algorithmic, hedge, and mutual fund portfolios and compare them against the S&P TSX 60 Index.",
"_____no_output_____"
],
[
"## Assumptions and limitations\n\n1. Limitation: Only dates that overlap between portfolios will be compared\n2. Assumption: There are no significant anomalous price impacting events during the time window such as share split, trading halt",
"_____no_output_____"
],
[
"## 0. Import Required Libraries",
"_____no_output_____"
]
],
[
[
"# Initial imports\nimport pandas as pd # daataframe manipulation\nimport numpy as np # calc and numeric manipulatino\nimport datetime as dt # date and tim \nfrom pathlib import Path # setting the path for file manipulation\nimport datetime \n\npd.options.display.float_format = '{:.6f}'.format # float format to 6 decimal places",
"_____no_output_____"
]
],
[
[
"# Data Cleaning\n\nIn this section, you will need to read the CSV files into DataFrames and perform any necessary data cleaning steps. After cleaning, combine all DataFrames into a single DataFrame.\n\nFiles:\n\n* `whale_returns.csv`: Contains returns of some famous \"whale\" investors' portfolios.\n\n* `algo_returns.csv`: Contains returns from the in-house trading algorithms from Harold's company.\n\n* `sp_tsx_history.csv`: Contains historical closing prices of the S&P TSX 60 Index.",
"_____no_output_____"
],
[
"## A. Whale Returns\n\nRead the Whale Portfolio daily returns and clean the data.",
"_____no_output_____"
],
[
"### 1. import whale csv and set index to date",
"_____no_output_____"
]
],
[
[
"df_wr = pd.read_csv('Resources/whale_returns.csv', index_col=\"Date\")",
"_____no_output_____"
]
],
[
[
"### 2. Inspect imported data",
"_____no_output_____"
]
],
[
[
"# look at colums and value head\ndf_wr.head(3)",
"_____no_output_____"
],
[
"# look at last few values\ndf_wr.tail(3)",
"_____no_output_____"
],
[
"# check dimensions of df\ndf_wr.shape",
"_____no_output_____"
],
[
"# get index datatype - for later merging\ndf_wr.index.dtype",
"_____no_output_____"
],
[
"# get datatypes of all values\ndf_wr.dtypes",
"_____no_output_____"
]
],
[
[
"### 3. Count and drop any null values",
"_____no_output_____"
]
],
[
[
"# Count nulls\ndf_wr.isna().sum()",
"_____no_output_____"
],
[
"# Drop nulls \ndf_wr.dropna(inplace=True)",
"_____no_output_____"
],
[
"# Count nulls -again to ensure they're removed\ndf_wr.isna().sum()",
"_____no_output_____"
],
[
"df_wr.count() #double check all values are equal in length",
"_____no_output_____"
]
],
[
[
"### 4. Sort the index to ensure the correct date order for calculations",
"_____no_output_____"
]
],
[
[
"df_wr.sort_index(inplace=True)",
"_____no_output_____"
]
],
[
[
"### 5. Rename columns - shorten and make consistent with other tables",
"_____no_output_____"
]
],
[
[
"# change columns to be consistent and informative\ndf_wr.columns",
"_____no_output_____"
],
[
"df_wr.columns = ['Whale_Soros_Fund_Daily_Returns', 'Whale_Paulson_Daily_Returns',\n 'Whale_Tiger_Daily_Returns', 'Whale_Berekshire_Daily_Returns']",
"_____no_output_____"
]
],
[
[
"### 6. Create copy dataframe with new column for cumulative returns",
"_____no_output_____"
]
],
[
[
"# copy the dataframe to store cumprod in a new view\ndf_wr_cumulative = df_wr.copy()",
"_____no_output_____"
],
[
"# create a new column in new df for each cumulative daily return using the cumprod function\ndf_wr_cumulative['Whale_Soros_Fund_Daily_CumReturns'] = (1 + df_wr_cumulative['Whale_Soros_Fund_Daily_Returns']).cumprod()",
"_____no_output_____"
],
[
"df_wr_cumulative['Whale_Paulson_Daily_CumReturns'] = (1 + df_wr_cumulative['Whale_Paulson_Daily_Returns']).cumprod()",
"_____no_output_____"
],
[
"df_wr_cumulative['Whale_Tiger_Daily_CumReturns'] = (1 + df_wr_cumulative['Whale_Tiger_Daily_Returns']).cumprod()",
"_____no_output_____"
],
[
"df_wr_cumulative['Whale_Berekshire_Daily_CumReturns'] = (1 + df_wr_cumulative['Whale_Berekshire_Daily_Returns']).cumprod()",
"_____no_output_____"
],
[
"df_wr_cumulative.head() # check result is consistent against original column ie adds up",
"_____no_output_____"
],
[
"# drop returns columns from cumulative df",
"_____no_output_____"
],
[
"df_wr_cumulative.columns",
"_____no_output_____"
],
[
"df_wr_cumulative = df_wr_cumulative[['Whale_Soros_Fund_Daily_CumReturns', 'Whale_Paulson_Daily_CumReturns','Whale_Tiger_Daily_CumReturns', 'Whale_Berekshire_Daily_CumReturns']]",
"_____no_output_____"
],
[
"df_wr_cumulative.head()",
"_____no_output_____"
]
],
[
[
"### 7. Look at high level stats & plot for whale portfolios",
"_____no_output_____"
]
],
[
[
"df_wr.describe(include='all') # basic stats for daily whale returns",
"_____no_output_____"
],
[
"df_wr_cumulative.describe(include='all') # basic stats for daily cumulative whale returns",
"_____no_output_____"
],
[
"# plot daily returns - whales\ndf_wr.plot(figsize=(10,5))",
"_____no_output_____"
],
[
"# Plot cumulative returns\ndf_wr_cumulative.plot(figsize=(10,5))",
"_____no_output_____"
]
],
[
[
"#### The data looks consistent and there are no obvious data errors identified. \n\n#### Initial high level observations of standalone daily returns data for whale portfolio: At initial glance, the mean daily return indicates that Berkshire portfolio performed best (mean daily returns of 0.000501, mean cumulative daily returns 1.159732), while Paulson worst (-0.000203). The standard deviation indicates highest risk for Berkshire (0.012831 STD), while lowest risk/volatility is Paulson (std 0.006977)\n#### A more thorough analysis will be done in the following analysis section, so no conclusions are drawn yet. \n#### By looking at the cumulative chart, it is evident that all portfolios were vulnerable to a loss at the same tim around 2019-02-16, but that Berkshir was able to increas the most over time and climb the steepest after the downturn",
"_____no_output_____"
],
[
"## B. Algorithmic Daily Returns\n\nRead the algorithmic daily returns and clean the data.",
"_____no_output_____"
],
[
"### 1. import algo csv and set index to date",
"_____no_output_____"
]
],
[
[
"# Reading algorithmic returns\ndf_ar = pd.read_csv('Resources/algo_returns.csv', index_col='Date')",
"_____no_output_____"
]
],
[
[
"### 2. Inspect resulting dataframe and contained data",
"_____no_output_____"
]
],
[
[
"# look at colums and value first 3 rows\ndf_ar.head(3)",
"_____no_output_____"
],
[
"# look at colums and value last 3 rows\ndf_ar.tail(3)",
"_____no_output_____"
],
[
"# get dimensions of df\ndf_ar.shape",
"_____no_output_____"
],
[
"# get index datatype - for later merging\ndf_ar.index.dtype",
"_____no_output_____"
],
[
"# get datatypes\ndf_ar.dtypes",
"_____no_output_____"
]
],
[
[
"### 3. Count and remove null values",
"_____no_output_____"
]
],
[
[
"# Count nulls\ndf_ar.isna().sum()",
"_____no_output_____"
],
[
"# Drop nulls\ndf_ar.dropna(inplace=True)",
"_____no_output_____"
],
[
"# Count nulls -again to ensure that nulls actually are removed\ndf_ar.isna().sum()",
"_____no_output_____"
],
[
"df_ar.count()",
"_____no_output_____"
]
],
[
[
"### 4. Sort index to ensure correct date order for calculations",
"_____no_output_____"
]
],
[
[
"df_ar.sort_index(inplace=True)",
"_____no_output_____"
]
],
[
[
"### 5. Rename columns to be consistent with future merge",
"_____no_output_____"
]
],
[
[
"df_ar.columns",
"_____no_output_____"
],
[
"df_ar.columns = ['Algo1_Daily_Returns', 'Algo2_Daily_Returns']",
"_____no_output_____"
]
],
[
[
"### 6. Create new column in a copy df for cumulative returns per Algo daily return",
"_____no_output_____"
]
],
[
[
"# create a df copy to store cumulative data\ndf_ar_cumulative = df_ar.copy() ",
"_____no_output_____"
],
[
"# use cumprod to get the daily cumulative returns for each of the algos 1 and 2\ndf_ar_cumulative['Algo1_Daily_CumReturns'] = (1 + df_ar_cumulative['Algo1_Daily_Returns']).cumprod()",
"_____no_output_____"
],
[
"df_ar_cumulative['Algo2_Daily_CumReturns'] = (1 + df_ar_cumulative['Algo2_Daily_Returns']).cumprod()",
"_____no_output_____"
],
[
"# check the result is consistent with the daily returns for first few columns\ndf_ar_cumulative.head(10)",
"_____no_output_____"
],
[
"# drop columns that are not required",
"_____no_output_____"
],
[
"df_ar_cumulative.columns # get the columns",
"_____no_output_____"
],
[
"df_ar_cumulative = df_ar_cumulative[['Algo1_Daily_CumReturns','Algo2_Daily_CumReturns']]",
"_____no_output_____"
],
[
"# check result - first few lines\ndf_ar_cumulative.head(10)",
"_____no_output_____"
]
],
[
[
"### 7. Look at high level stats & plot for algo portfolios",
"_____no_output_____"
]
],
[
[
"df_ar.describe(include='all') # stats for daily returns",
"_____no_output_____"
],
[
"df_ar_cumulative.describe(include='all') # stats for daily cumulative returns",
"_____no_output_____"
],
[
"# plot daily returns - algos\ndf_ar.plot(figsize=(10,5))",
"_____no_output_____"
],
[
"# plot daily cumulative returns - algos\ndf_ar_cumulative.plot(figsize=(10,5))",
"_____no_output_____"
]
],
[
[
"#### The data looks consistent and there are no obvious errors identified. \n\n#### Initial observations of standalone daily returns data for Algo 1 vs Algo 2: mean daily return indicates that Algo 1 (mean daily return 0.000654) performs slightly better than Algo 2 (mean daily return 0.000341), which is alo evident in the cumulative daily returns plot. When looking at just daily returns, Algo 2 is more risky, but when looking at cumulative returns, Algo 1 is more risky (ie higher standard deviation). ",
"_____no_output_____"
],
[
"## C. S&P TSX 60 Returns\n\nRead the S&P TSX 60 historic closing prices and create a new daily returns DataFrame from the data. \nNote: this contains daily closing and not returns - needs to be converted",
"_____no_output_____"
],
[
"### 1. Import S&P csv daily closing price (not returns)",
"_____no_output_____"
]
],
[
[
"# Reading S&P TSX 60 Closing Prices\n\ndf_sr = pd.read_csv('Resources/sp_tsx_history.csv')",
"_____no_output_____"
]
],
[
[
"### 2. Inspect columns of dataframe",
"_____no_output_____"
]
],
[
[
"# look at colums and value head\ndf_sr.head(3)",
"_____no_output_____"
],
[
"# look at tail values\ndf_sr.tail(3)",
"_____no_output_____"
]
],
[
[
"#### Note from dataframe inspection: \n#### 1. date column was not immediated converted because it is in\n#### a different format to the other csv files and \n#### needs to bee converted to consistent format first\n#### 2. Close cannot be explicitly converted to float as it has\n#### dollar and commas. \n#### 3. A new column for returns will need to be created from \n#### return calculations. ",
"_____no_output_____"
]
],
[
[
"# check dimension of df\ndf_sr.shape",
"_____no_output_____"
],
[
"# Check Data Types\ndf_sr.dtypes",
"_____no_output_____"
]
],
[
[
"### 3. Convert the date into a consistent format with other tables",
"_____no_output_____"
]
],
[
[
"df_sr['Date']= pd.to_datetime(df_sr['Date']).dt.strftime('%Y-%m-%d')\n#df_sr['Date']= pd.to_datetime(df_sr['Date'], format='%Y-%m-%d')",
"_____no_output_____"
]
],
[
[
"### 4. Convert th data to index and check format and data type",
"_____no_output_____"
]
],
[
[
"# set date as index\ndf_sr.set_index('Date', inplace=True)",
"_____no_output_____"
],
[
"df_sr.head(2)",
"_____no_output_____"
],
[
"df_sr.index.dtype",
"_____no_output_____"
]
],
[
[
"### 5. Check for null values",
"_____no_output_____"
]
],
[
[
"# Count nulls - none observed\ndf_ar.isna().sum()",
"_____no_output_____"
]
],
[
[
"### 6. Convert daily closing price to float (from string)",
"_____no_output_____"
]
],
[
[
"# Change the Closing column to b float type\ndf_sr['Close']= df_sr['Close'].str.replace('$','')\ndf_sr['Close']= df_sr['Close'].str.replace(',','')\ndf_sr['Close']= df_sr['Close'].astype(float)",
"_____no_output_____"
],
[
"# Check Data Types\ndf_sr.dtypes",
"_____no_output_____"
],
[
"# test \ndf_sr.iloc[0]",
"_____no_output_____"
],
[
"# check null values\ndf_sr.isna().sum()",
"_____no_output_____"
],
[
"df_sr.count()",
"_____no_output_____"
]
],
[
[
"### 7. Sort the index for calculations of returns",
"_____no_output_____"
]
],
[
[
"# sort_index \ndf_sr.sort_index(inplace=True)",
"_____no_output_____"
],
[
"df_sr.head(2)",
"_____no_output_____"
]
],
[
[
"df_sr.tail(2)",
"_____no_output_____"
]
],
[
[
"### 8. Calculate daily returns and store in new column",
"_____no_output_____"
],
[
"Equation: $r=\\frac{{p_{t}} - {p_{t-1}}}{p_{t-1}}$\n\nThe daily return is the (current closing price minus the previous day closing price) all divided by the previous day closing price. The initial value has no daily return as there is no prior period to compare it with. \n\nHere the calculation uses the python shift function ",
"_____no_output_____"
]
],
[
[
"\ndf_sr['SnP_TSX_60_Returns'] = (df_sr['Close'] - df_sr['Close'].shift(1))/ df_sr['Close'].shift(1)",
"_____no_output_____"
],
[
"df_sr.head(10)",
"_____no_output_____"
]
],
[
[
"### 9. Cross check conversion to daily returns against alternative method - pct_change function",
"_____no_output_____"
]
],
[
[
"df_sr['SnP_TSX_60_Returns'] = df_sr['Close'].pct_change()\ndf_sr.head(10)",
"_____no_output_____"
]
],
[
[
"#### Methods cross check - looks good - continue",
"_____no_output_____"
]
],
[
[
"# check for null - first row would have null\ndf_sr.isna().sum()",
"_____no_output_____"
],
[
"# Drop nulls - first row\ndf_sr.dropna(inplace=True)",
"_____no_output_____"
],
[
"# Rename `Close` Column to be specific to this portfolio.\ndf_sr.columns",
"_____no_output_____"
],
[
"df_sr.head()",
"_____no_output_____"
]
],
[
[
"### 10. Drop original Closing column - not needed for comparison",
"_____no_output_____"
]
],
[
[
"df_sr = df_sr[['SnP_TSX_60_Returns']] ",
"_____no_output_____"
],
[
"df_sr.columns",
"_____no_output_____"
]
],
[
[
"### 11. Create new column in a copy df for cumulative returns per daily return S&P TSX 60",
"_____no_output_____"
]
],
[
[
"df_sr_cumulative = df_sr.copy()",
"_____no_output_____"
],
[
"# use cumprod to get the daily cumulative returns for each of the algos 1 and 2\ndf_sr_cumulative['SnP_TSX_60_CumReturns'] = (1+df_sr_cumulative['SnP_TSX_60_Returns']).cumprod()",
"_____no_output_____"
],
[
"# visually check first 10 rows to ensure that results make sense\ndf_sr_cumulative.head(10)",
"_____no_output_____"
]
],
[
[
"## D. Combine Whale, Algorithmic, and S&P TSX 60 Returns",
"_____no_output_____"
]
],
[
[
"# Use the `concat` function to combine the two DataFrames by matching indexes (or in this case `Month`)\nmerged_analysis_df_tmp = pd.concat([df_wr, df_ar ], axis=\"columns\", join=\"inner\")",
"_____no_output_____"
],
[
"merged_analysis_df_tmp.head(3)",
"_____no_output_____"
],
[
"# Use the `concat` function to combine the two DataFrames by matching indexes (or in this case `Month`)\nmerged_daily_returns_df = pd.concat([merged_analysis_df_tmp, df_sr ], axis=\"columns\", join=\"inner\")",
"_____no_output_____"
],
[
"merged_daily_returns_df.head(3)",
"_____no_output_____"
]
],
[
[
"# Conduct Quantitative Analysis\n\nIn this section, you will calculate and visualize performance and risk metrics for the portfolios.",
"_____no_output_____"
],
[
"## Performance Anlysis\n\n#### Calculate and Plot the daily returns",
"_____no_output_____"
]
],
[
[
"# Plot daily returns of all portfolios\ndrp = merged_analysis_df.plot(figsize=(20,10), rot=45, title='Comparison of Daily Returns on Stock Portfolios')\ndrp.set_xlabel(\"Daily Returns\")\ndrp.set_ylabel(\"Date\")\n",
"_____no_output_____"
]
],
[
[
"#### Calculate and Plot cumulative returns.",
"_____no_output_____"
]
],
[
[
"merged_analysis_cum_df.columns",
"_____no_output_____"
]
],
[
[
"# [[NOT WORKING BECAUSE FIRST ROW SHOULD NOT BE NULL]]",
"_____no_output_____"
]
],
[
[
"merged_analysis_cum_df.head()",
"_____no_output_____"
],
[
"# Calculate cumulative returns of all portfolios",
"_____no_output_____"
],
[
"# Plot cumulative returns\n\ndrp = merged_analysis_cum_df.plot(figsize=(20,20), rot=45, title='Comparison of Daily Cumulative Returns on Stock Portfolios')\ndrp.set_xlabel(\"Daily Cumulative Returns\")\ndrp.set_ylabel(\"Date\")\n",
"_____no_output_____"
]
],
[
[
"---",
"_____no_output_____"
],
[
"## Risk Analysis\n\nDetermine the _risk_ of each portfolio:\n\n1. Create a box plot for each portfolio. \n2. Calculate the standard deviation for all portfolios.\n4. Determine which portfolios are riskier than the S&P TSX 60.\n5. Calculate the Annualized Standard Deviation.",
"_____no_output_____"
],
[
"### Create a box plot for each portfolio\n",
"_____no_output_____"
]
],
[
[
"# Box plot to visually show risk\n",
"_____no_output_____"
]
],
[
[
"### Calculate Standard Deviations",
"_____no_output_____"
]
],
[
[
"# Calculate the daily standard deviations of all portfolios\n",
"_____no_output_____"
]
],
[
[
"### Determine which portfolios are riskier than the S&P TSX 60",
"_____no_output_____"
]
],
[
[
"# Calculate the daily standard deviation of S&P TSX 60\n\n# Determine which portfolios are riskier than the S&P TSX 60\n",
"_____no_output_____"
]
],
[
[
"### Calculate the Annualized Standard Deviation",
"_____no_output_____"
]
],
[
[
"# Calculate the annualized standard deviation (252 trading days)\n",
"_____no_output_____"
]
],
[
[
"---",
"_____no_output_____"
],
[
"## Rolling Statistics\n\nRisk changes over time. Analyze the rolling statistics for Risk and Beta. \n\n1. Calculate and plot the rolling standard deviation for the S&P TSX 60 using a 21-day window.\n2. Calculate the correlation between each stock to determine which portfolios may mimick the S&P TSX 60.\n3. Choose one portfolio, then calculate and plot the 60-day rolling beta for it and the S&P TSX 60.",
"_____no_output_____"
],
[
"### Calculate and plot rolling `std` for all portfolios with 21-day window",
"_____no_output_____"
]
],
[
[
"# Calculate the rolling standard deviation for all portfolios using a 21-day window\n\n# Plot the rolling standard deviation\n",
"_____no_output_____"
]
],
[
[
"### Calculate and plot the correlation",
"_____no_output_____"
]
],
[
[
"# Calculate the correlation\n\n# Display de correlation matrix\n",
"_____no_output_____"
]
],
[
[
"### Calculate and Plot Beta for a chosen portfolio and the S&P 60 TSX",
"_____no_output_____"
]
],
[
[
"# Calculate covariance of a single portfolio\n\n# Calculate variance of S&P TSX\n\n# Computing beta\n\n# Plot beta trend\n",
"_____no_output_____"
]
],
[
[
"## Rolling Statistics Challenge: Exponentially Weighted Average \n\nAn alternative way to calculate a rolling window is to take the exponentially weighted moving average. This is like a moving window average, but it assigns greater importance to more recent observations. Try calculating the [`ewm`](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.ewm.html) with a 21-day half-life.",
"_____no_output_____"
]
],
[
[
"# Use `ewm` to calculate the rolling window\n",
"_____no_output_____"
]
],
[
[
"---",
"_____no_output_____"
],
[
"# Sharpe Ratios\nIn reality, investment managers and thier institutional investors look at the ratio of return-to-risk, and not just returns alone. After all, if you could invest in one of two portfolios, and each offered the same 10% return, yet one offered lower risk, you'd take that one, right?\n\n### Using the daily returns, calculate and visualize the Sharpe ratios using a bar plot",
"_____no_output_____"
]
],
[
[
"# Annualized Sharpe Ratios\n",
"_____no_output_____"
],
[
"# Visualize the sharpe ratios as a bar plot\n",
"_____no_output_____"
]
],
[
[
"### Determine whether the algorithmic strategies outperform both the market (S&P TSX 60) and the whales portfolios.\n\nWrite your answer here!",
"_____no_output_____"
],
[
"---",
"_____no_output_____"
],
[
"# Create Custom Portfolio\n\nIn this section, you will build your own portfolio of stocks, calculate the returns, and compare the results to the Whale Portfolios and the S&P TSX 60. \n\n1. Choose 3-5 custom stocks with at last 1 year's worth of historic prices and create a DataFrame of the closing prices and dates for each stock.\n2. Calculate the weighted returns for the portfolio assuming an equal number of shares for each stock.\n3. Join your portfolio returns to the DataFrame that contains all of the portfolio returns.\n4. Re-run the performance and risk analysis with your portfolio to see how it compares to the others.\n5. Include correlation analysis to determine which stocks (if any) are correlated.",
"_____no_output_____"
],
[
"## Choose 3-5 custom stocks with at last 1 year's worth of historic prices and create a DataFrame of the closing prices and dates for each stock.\n\nFor this demo solution, we fetch data from three companies listes in the S&P TSX 60 index.\n\n* `SHOP` - [Shopify Inc](https://en.wikipedia.org/wiki/Shopify)\n\n* `OTEX` - [Open Text Corporation](https://en.wikipedia.org/wiki/OpenText)\n\n* `L` - [Loblaw Companies Limited](https://en.wikipedia.org/wiki/Loblaw_Companies)",
"_____no_output_____"
]
],
[
[
"# Reading data from 1st stock\n",
"_____no_output_____"
],
[
"# Reading data from 2nd stock\n",
"_____no_output_____"
],
[
"# Reading data from 3rd stock\n",
"_____no_output_____"
],
[
"# Combine all stocks in a single DataFrame\n",
"_____no_output_____"
],
[
"# Reset Date index\n",
"_____no_output_____"
],
[
"# Reorganize portfolio data by having a column per symbol\n",
"_____no_output_____"
],
[
"# Calculate daily returns\n\n# Drop NAs\n\n# Display sample data\n",
"_____no_output_____"
]
],
[
[
"## Calculate the weighted returns for the portfolio assuming an equal number of shares for each stock",
"_____no_output_____"
]
],
[
[
"# Set weights\nweights = [1/3, 1/3, 1/3]\n\n# Calculate portfolio return\n\n# Display sample data\n",
"_____no_output_____"
]
],
[
[
"## Join your portfolio returns to the DataFrame that contains all of the portfolio returns",
"_____no_output_____"
]
],
[
[
"# Join your returns DataFrame to the original returns DataFrame\n",
"_____no_output_____"
],
[
"# Only compare dates where return data exists for all the stocks (drop NaNs)\n",
"_____no_output_____"
]
],
[
[
"## Re-run the risk analysis with your portfolio to see how it compares to the others",
"_____no_output_____"
],
[
"### Calculate the Annualized Standard Deviation",
"_____no_output_____"
]
],
[
[
"# Calculate the annualized `std`\n",
"_____no_output_____"
]
],
[
[
"### Calculate and plot rolling `std` with 21-day window",
"_____no_output_____"
]
],
[
[
"# Calculate rolling standard deviation\n\n# Plot rolling standard deviation\n",
"_____no_output_____"
]
],
[
[
"### Calculate and plot the correlation",
"_____no_output_____"
]
],
[
[
"# Calculate and plot the correlation\n",
"_____no_output_____"
]
],
[
[
"### Calculate and Plot the 60-day Rolling Beta for Your Portfolio compared to the S&P 60 TSX",
"_____no_output_____"
]
],
[
[
"# Calculate and plot Beta\n",
"_____no_output_____"
]
],
[
[
"### Using the daily returns, calculate and visualize the Sharpe ratios using a bar plot",
"_____no_output_____"
]
],
[
[
"# Calculate Annualzied Sharpe Ratios\n",
"_____no_output_____"
],
[
"# Visualize the sharpe ratios as a bar plot\n",
"_____no_output_____"
]
],
[
[
"### How does your portfolio do?\n\nWrite your answer here!",
"_____no_output_____"
],
[
"### References\n\nShift function in pandas - \nhttps://stackoverflow.com/questions/20000726/calculate-daily-returns-with-pandas-dataframe\n\nConditional line color - \nhttps://stackoverflow.com/questions/31590184/plot-multicolored-line-based-on-conditional-in-python\n\nhttps://stackoverflow.com/questions/40803570/python-matplotlib-scatter-plot-specify-color-points-depending-on-conditions/40804861\n\nhttps://stackoverflow.com/questions/42453649/conditional-color-with-matplotlib-scatter\n\nhttps://stackoverflow.com/questions/3832809/how-to-change-the-color-of-a-single-bar-if-condition-is-true-matplotlib\n\nhttps://stackoverflow.com/questions/56779975/conditional-coloring-in-matplotlib-using-numpys-where\n\nPEP 8 - Standards - \n\n",
"_____no_output_____"
]
]
] | [
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"raw",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown"
] | [
[
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code"
],
[
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code"
],
[
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"raw"
],
[
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown",
"markdown",
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown"
]
] |
d0fe7d3e72d7556c23c2be586635604c8d971d5b | 4,634 | ipynb | Jupyter Notebook | gs_quant/examples/04_scenario/01_carry_shock/040103_yield_curves_with_carry.ipynb | pnijhara/gs-quant | 2bec436dc0ff087d94995643b3aa003c5a94d49e | [
"Apache-2.0"
] | 1 | 2020-05-18T02:09:39.000Z | 2020-05-18T02:09:39.000Z | gs_quant/examples/04_scenario/01_carry_shock/040103_yield_curves_with_carry.ipynb | pnijhara/gs-quant | 2bec436dc0ff087d94995643b3aa003c5a94d49e | [
"Apache-2.0"
] | null | null | null | gs_quant/examples/04_scenario/01_carry_shock/040103_yield_curves_with_carry.ipynb | pnijhara/gs-quant | 2bec436dc0ff087d94995643b3aa003c5a94d49e | [
"Apache-2.0"
] | null | null | null | 34.58209 | 129 | 0.586319 | [
[
[
"from gs_quant.session import GsSession, Environment\nfrom gs_quant.instrument import IRSwap\nfrom gs_quant.risk import IRFwdRate, CarryScenario\nfrom gs_quant.markets.portfolio import Portfolio\nfrom gs_quant.markets import PricingContext\nfrom datetime import datetime\nimport matplotlib.pylab as plt\nimport pandas as pd\nimport numpy as np",
"_____no_output_____"
],
[
"# external users should substitute their client id and secret; please skip this step if using internal jupyterhub\nGsSession.use(Environment.PROD, client_id=None, client_secret=None, scopes=('run_analytics',))",
"_____no_output_____"
],
[
"ccy = 'EUR'\n\n# construct a series of 6m FRAs going out 20y or so\nfras = Portfolio([IRSwap('Pay', '{}m'.format(i), ccy, effective_date='{}m'.format(i-6), \n fixed_rate_frequency='6m', floating_rate_frequency='6m') \n for i in range(6, 123, 6)])\nfras.resolve()\nresults = fras.calc(IRFwdRate)\n\n# get the fwd rates for these fras under the base sceneraio (no shift in time)\nbase = {}\nfor i, res in enumerate(results):\n base[datetime.strptime(fras[i].termination_date, '%Y-%m-%d')] = res\n\nbase_series = pd.Series(base, name='base', dtype=np.dtype(float))",
"_____no_output_____"
],
[
"# calculate the fwd rates with a shift forward of 132 business days - about 6m. This shift keeps spot rates constant. \n# So 5y rate today will be 5y rate under the scenario of pricing 6m in the future.\nwith CarryScenario(time_shift=132, roll_to_fwds=False):\n fras = Portfolio([IRSwap('Pay', '{}m'.format(i), ccy, effective_date='{}m'.format(i-6), \n fixed_rate_frequency='6m', floating_rate_frequency='6m') for i in range(6, 123, 6)])\n fras.resolve()\n results = fras.calc(IRFwdRate)\n\n roll_spot = {}\n for i, res in enumerate(results):\n roll_spot[datetime.strptime(fras[i].termination_date, '%Y-%m-%d')] = res\n\nroll_spot_series = pd.Series(roll_spot, name='roll to spot', dtype=np.dtype(float))",
"_____no_output_____"
],
[
"# calculate the fwd rates with a shift forward of 132 business days - about 6m. This shift keeps fwd rates constant. \n# So 5.5y rate today will be 5y rate under the scenario of pricing 6m in the future.\n\nwith CarryScenario(time_shift=132, roll_to_fwds=True):\n fras = Portfolio([IRSwap('Pay', '{}m'.format(i), ccy, effective_date='{}m'.format(i-6), \n fixed_rate_frequency='6m', floating_rate_frequency='6m') for i in range(6, 123, 6)])\n fras.resolve()\n results = fras.calc(IRFwdRate)\n\n roll_fwd = {}\n for i, res in enumerate(results):\n roll_fwd[datetime.strptime(fras[i].termination_date, '%Y-%m-%d')] = res\n\nroll_fwd_series = pd.Series(roll_fwd, name='roll to fwd', dtype=np.dtype(float))",
"_____no_output_____"
],
[
"# show the curves, the base in blue, the roll to fwd in green and the roll to spot in orange.\n# note blue and green curves are not exactly on top of each other as we aren't using the curve instruments themselves\n# but instead using FRAs to show a smooth curve.\nbase_series.plot(figsize=(20, 10))\nroll_spot_series.plot()\nroll_fwd_series.plot()\nplt.legend()",
"_____no_output_____"
]
]
] | [
"code"
] | [
[
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
d0fe83ef9b5b7348db2a6c9cd5289c80650bf937 | 39,192 | ipynb | Jupyter Notebook | hw/hw09/.ipynb_checkpoints/hw09-checkpoint.ipynb | ds-modules/Colab-demo | cccaff13633f8a5ec697cd4aeca9087f2feec2e4 | [
"BSD-3-Clause"
] | null | null | null | hw/hw09/.ipynb_checkpoints/hw09-checkpoint.ipynb | ds-modules/Colab-demo | cccaff13633f8a5ec697cd4aeca9087f2feec2e4 | [
"BSD-3-Clause"
] | null | null | null | hw/hw09/.ipynb_checkpoints/hw09-checkpoint.ipynb | ds-modules/Colab-demo | cccaff13633f8a5ec697cd4aeca9087f2feec2e4 | [
"BSD-3-Clause"
] | null | null | null | 32.20378 | 568 | 0.613493 | [
[
[
"# Initialize Otter\nimport otter\ngrader = otter.Notebook(\"hw09.ipynb\")",
"_____no_output_____"
]
],
[
[
"# Homework 9: Bootstrap, Resampling, CLT",
"_____no_output_____"
],
[
"**Reading**: \n* [Estimation](https://www.inferentialthinking.com/chapters/13/estimation.html)\n* [Why the mean matters](https://www.inferentialthinking.com/chapters/14/why-the-mean-matters.html)\n\nPlease complete this notebook by filling in the cells provided. \n\nDirectly sharing answers is not okay, but discussing problems with the course staff or with other students is encouraged. Refer to the policies page to learn more about how to learn cooperatively.\n\nFor all problems that you must write our explanations and sentences for, you **must** provide your answer in the designated space. Moreover, throughout this homework and all future ones, please be sure to not re-assign variables throughout the notebook! For example, if you use `max_temperature` in your answer to one question, do not reassign it later on.",
"_____no_output_____"
]
],
[
[
"# Run this cell to set up the notebook, but please don't change it.\n\n# These lines import the Numpy and Datascience modules.\nimport numpy as np\nfrom datascience import *\n\n# These lines do some fancy plotting magic.\nimport matplotlib\n%matplotlib inline\nimport matplotlib.pyplot as plt\nplt.style.use('fivethirtyeight')\nimport warnings\nwarnings.simplefilter('ignore', FutureWarning)\n\n",
"_____no_output_____"
]
],
[
[
"## 1. Preliminaries\nThe British Royal Air Force wanted to know how many warplanes the Germans had (some number `N`, which is a *parameter*), and they needed to estimate that quantity knowing only a random sample of the planes' serial numbers (from 1 to `N`). We know that the German's warplanes are labeled consecutively from 1 to `N`, so `N` would be the total number of warplanes they have. \n\nWe normally investigate the random variation among our estimates by simulating a sampling procedure from the population many times and computing estimates from each sample that we generate. In real life, if the British Royal Air Force (RAF) had known what the population looked like, they would have known `N` and would not have had any reason to think about random sampling. However, they didn't know what the population looked like, so they couldn't have run the simulations that we normally do. \n\nSimulating a sampling procedure many times was a useful exercise in *understanding random variation* for an estimate, but it's not as useful as a tool for practical data analysis.\n\nLet's flip that sampling idea on its head to make it practical. **Given *just* a random sample of serial numbers, we'll estimate `N`, and then we'll use simulation to find out how accurate our estimate probably is, without ever looking at the whole population.** This is an example of *statistical inference*.\n\nWe (the RAF in World War II) want to know the number of warplanes fielded by the Germans. That number is `N`. The warplanes have serial numbers from 1 to `N`, so `N` is also equal to the largest serial number on any of the warplanes.\n\nWe only see a small number of serial numbers (assumed to be a random sample with replacement from among all the serial numbers), so we have to use estimation.",
"_____no_output_____"
],
[
"#### Question 1.1\nIs `N` a population parameter or a statistic? If we use our random sample to compute a number that is an estimate of `N`, is that a population parameter or a statistic?\n\nSet `N` and `N_estimate` to either the string `\"parameter\"` or `\"statistic\"` to indicate whether each value is a parameter or a statistic.\n\n<!--\nBEGIN QUESTION\nname: q1_1\n-->",
"_____no_output_____"
]
],
[
[
"N = ...\nN_estimate = ...",
"_____no_output_____"
],
[
"grader.check(\"q1_1\")",
"_____no_output_____"
]
],
[
[
"To make the situation realistic, we're going to hide the true number of warplanes from you. You'll have access only to this random sample:",
"_____no_output_____"
]
],
[
[
"observations = Table.read_table(\"serial_numbers.csv\")\nnum_observations = observations.num_rows\nobservations",
"_____no_output_____"
]
],
[
[
"#### Question 1.2\nThe average of the sample is about half of `N`. So one way to estimate `N` is to take twice the mean of the serial numbers we see. Write a function that computes that statistic. It should take as its argument an array of serial numbers and return twice their mean. Call the function `mean_based_estimator`. \n\nAfter that, use the function and the `observations` table to compute an estimate of `N` called `mean_based_estimate`.\n\n<!--\nBEGIN QUESTION\nname: q1_2\n-->",
"_____no_output_____"
]
],
[
[
"def mean_based_estimator(nums):\n ...\n\nmean_based_estimate = ...\nmean_based_estimate",
"_____no_output_____"
],
[
"grader.check(\"q1_2\")",
"_____no_output_____"
]
],
[
[
"#### Question 1.3\nWe can also estimate `N` by using the biggest serial number in the sample. Compute this value and give it the name `max_estimate`.\n\n<!--\nBEGIN QUESTION\nname: q1_3\n-->",
"_____no_output_____"
]
],
[
[
"max_estimate = ...\nmax_estimate",
"_____no_output_____"
],
[
"grader.check(\"q1_3\")",
"_____no_output_____"
]
],
[
[
"<!-- BEGIN QUESTION -->\n\n#### Question 1.4\nLet's take a look at the values of `max_estimate` and `mean_based_estimate` that we got for our dataset. Which of these values is closer to the true population maximum `N`? Based off of our estimators, can we give a lower bound for what `N` must be? In other words, is there a value that `N` must be greater than or equal to?\n\n<!--\nBEGIN QUESTION\nname: q1_4\nmanual: true\n-->",
"_____no_output_____"
],
[
"_Type your answer here, replacing this text._",
"_____no_output_____"
],
[
"<!-- END QUESTION -->\n\n\n\nWe can't just confidently proclaim that `max_estimate` or `mean_based_estimate` is equal to `N`. What if we're really far off? We want to get a sense of the accuracy of our estimates.",
"_____no_output_____"
],
[
"## 2. Resampling\nTo do this, we'll use resampling. That is, we won't exactly simulate the observations the RAF would have really seen. Rather we sample from our current sample, or \"resample.\"\n\nWhy does that make any sense?\n\nWhen we try to find the value of a population parameter, we ideally would like to use the whole population. However, we often only have access to one sample and we must use that to estimate the parameter instead.\n\nHere, we would like to use the population of serial numbers to draw more samples and run a simulation about estimates of `N`. But we still only have our sample. So, we **use our sample in place of the population** to run the simulation. We resample from our original sample with replacement as many times as there are elements in the original sample. This resampling technique is called *bootstrapping*. \n\nNote that in order for bootstrapping to work well, you must start with a large, random sample. Then the Law of Large Numbers says that with high probability, your sample is representative of the population.",
"_____no_output_____"
],
[
"#### Question 2.1\nWrite a function called `simulate_resample`. The function should take one argument `tbl`, which is a table like `observations`. The function should generate a resample from the observed serial numbers in `tbl`.\n\n<!--\nBEGIN QUESTION\nname: q2_1\n-->",
"_____no_output_____"
]
],
[
[
"def simulate_resample(tbl):\n ...\n\nsimulate_resample(observations) # Don't delete this line",
"_____no_output_____"
],
[
"grader.check(\"q2_1\")",
"_____no_output_____"
]
],
[
[
"We'll use many resamples at once to see what estimates typically look like. However, we don't often pay attention to single resamples, so it's easy to misunderstand them. Let's first answer some questions about our resample.",
"_____no_output_____"
],
[
"#### Question 2.2\nWhich of the following statements are true?\n\n1. The original sample can contain serial numbers that are not in the resample.\n2. Because the sample size is small, the histogram of the resample might look very different from the histogram of the original sample.\n3. The resample can contain serial numbers that are not in the original sample.\n4. The original sample has exactly one copy of each serial number for every German plane.\n5. The resample has either zero, one, or more than one copy of each serial number.\n6. The resample has exactly the same sample size as the original sample.\n\nAssign `true_statements` to an array of the number(s) corresponding to correct statements.\n\n*Note:* The \"original sample\" refers to `observations`, and the \"resample\" refers the output of one call of `simulate_resample()`. \n\n<!--\nBEGIN QUESTION\nname: q2_2\n-->",
"_____no_output_____"
]
],
[
[
"true_statements = ...",
"_____no_output_____"
],
[
"grader.check(\"q2_2\")",
"_____no_output_____"
]
],
[
[
"Now let's write a function to do many resamples at once.\n\n#### Question 2.3\nWrite a function called `sample_estimates`. It should take 3 arguments:\n1. `serial_num_tbl`: A table from which the data should be sampled. The table will look like `observations`. \n2. `statistic`: A *function* that takes in an array of serial numbers as its argument and computes a statistic from the array (i.e. returns a calculated number). \n3. `num_replications`: The number of simulations to perform.\n\n*Hint: You should use the function `simulate_resample` which you defined in Question 2.1*\n\nThe function should simulate many samples **with replacement** from the given table. For each of those samples, it should compute the statistic on that sample. Then it should **return an array** containing each of those statistics. The code below provides an example use of your function and describes how you can verify that you've written it correctly.\n\n<!--\nBEGIN QUESTION\nname: q2_3\n-->",
"_____no_output_____"
]
],
[
[
"def sample_estimates(serial_num_tbl, statistic, num_replications):\n ...\n\n# DON'T CHANGE THE CODE BELOW THIS COMMENT! (If you do, you will fail the hidden test)\n# This is just an example to test your function.\n# This should generate an empirical histogram of twice-mean-based estimates\n# of N from samples of size 50 if N is 1000. This should be a bell-shaped\n# curve centered at roughly 900 with most of its mass in [800, 1200]. To verify your\n# answer, make sure that's what you see!\npopulation = Table().with_column(\"serial number\", np.arange(1, 1000+1))\none_sample = Table.read_table(\"one_sample.csv\") #This is a sample from the population table\nexample_estimates = sample_estimates(\n one_sample,\n mean_based_estimator,\n 10000)\nTable().with_column(\"mean-based estimate\", example_estimates).hist(bins=np.arange(0, 1500, 25))",
"_____no_output_____"
],
[
"grader.check(\"q2_3\")",
"_____no_output_____"
]
],
[
[
"Now we can go back to the sample we actually observed (the table `observations`) and estimate how much our mean-based estimate of `N` would have varied from sample to sample.",
"_____no_output_____"
],
[
"#### Question 2.4\nUsing the bootstrap and the sample `observations`, simulate the approximate distribution of *mean-based estimates* of `N`. Use 7,500 replications and save the estimates in an array called `bootstrap_mean_based_estimates`. \n\nWe have provided code that plots a histogram, allowing you to visualize the simulated estimates.\n\n<!--\nBEGIN QUESTION\nname: q2_4\n-->",
"_____no_output_____"
]
],
[
[
"bootstrap_mean_based_estimates = ...\n\n# Don't change the code below! This plots bootstrap_mean_based_estimates.\nTable().with_column(\"mean-based estimate\", bootstrap_mean_based_estimates).hist(bins=np.arange(0, 200, 4)) ",
"_____no_output_____"
],
[
"grader.check(\"q2_4\")",
"_____no_output_____"
]
],
[
[
"#### Question 2.5\nUsing the bootstrap and the sample `observations`, simulate the approximate distribution of *max estimates* of `N`. Use 7,500 replications and save the estimates in an array called `bootstrap_max_estimates`.\n\nWe have provided code that plots a histogram, allowing you to visualize the simulated estimates.\n\n<!--\nBEGIN QUESTION\nname: q2_5\n-->",
"_____no_output_____"
]
],
[
[
"bootstrap_max_estimates = ...\n\n# Don't change the code below! This plots bootstrap_max_estimates.\nTable().with_column(\"max estimate\", bootstrap_max_estimates).hist(bins=np.arange(0, 200, 4)) ",
"_____no_output_____"
],
[
"grader.check(\"q2_5\")",
"_____no_output_____"
]
],
[
[
"<!-- BEGIN QUESTION -->\n\n#### Question 2.6\n`N` was actually 150! Compare the histograms of estimates you generated in 2.4 and 2.5 and answer the following questions:\n\n1. How does the distribution of values for the mean-based estimates differ from the max estimates? Do both distributions contain the true max value?\n2. Which estimator is more dependent on the original random sample? Why so?\n\n<!--\nBEGIN QUESTION\nname: q2_6\nmanual: true\n-->",
"_____no_output_____"
],
[
"_Type your answer here, replacing this text._",
"_____no_output_____"
],
[
"<!-- END QUESTION -->\n\n\n\n## 3. Computing intervals",
"_____no_output_____"
],
[
"#### Question 3.1\nCompute an interval that covers the middle 95% of the mean-based bootstrap estimates. Assign your values to `left_end_1` and `right_end_1`. \n\n*Hint:* Use the `percentile` function! Read up on its documentation [here](http://data8.org/sp19/python-reference.html).\n\nVerify that your interval looks like it covers 95% of the area in the histogram. The red dot on the histogram is the value of the parameter (150).\n\n<!--\nBEGIN QUESTION\nname: q3_1\n-->",
"_____no_output_____"
]
],
[
[
"left_end_1 = ...\nright_end_1 = ...\nprint(\"Middle 95% of bootstrap estimates: [{:f}, {:f}]\".format(left_end_1, right_end_1))\n\n# Don't change the code below! It draws your interval and N on the histogram of mean-based estimates.\nTable().with_column(\"mean-based estimate\", bootstrap_mean_based_estimates).hist(bins=np.arange(0, 200, 4)) \nplt.plot(make_array(left_end_1, right_end_1), make_array(0, 0), color='yellow', lw=7, zorder=1)\nplt.scatter(150, 0, color='red', s=30, zorder=2);",
"_____no_output_____"
],
[
"grader.check(\"q3_1\")",
"_____no_output_____"
]
],
[
[
"#### Question 3.2\nWrite code that simulates the sampling and bootstrapping process again, as follows:\n\n1. Generate a new set of random observations the RAF might have seen by sampling from the `population` table we have created for you below. Use the sample size `num_observations`.\n2. Compute an estimate of `N` from these new observations, using `mean_based_estimator`.\n3. Using only the new observations, compute 10,000 bootstrap estimates of `N`.\n4. Plot these bootstrap estimates and compute an interval covering the middle 95%.\n\n*Note:* Traditionally, when we bootstrap using a sample from the population, that sample is usually a simple random sample (i.e., sampled uniformly at random from the population without replacement). However, if the population size is big enough, the difference between sampling with replacement and without replacement is negligible. Think about why that's the case! This is why when we define `new_observations`, we sample with replacement.\n\n<!--\nBEGIN QUESTION\nname: q3_2\n-->",
"_____no_output_____"
]
],
[
[
"population = Table().with_column(\"serial number\", np.arange(1, 150+1))\n\nnew_observations = ...\nnew_mean_based_estimate = ...\nnew_bootstrap_estimates = ...\nTable().with_column(\"mean-based estimate\", new_bootstrap_estimates).hist(bins=np.arange(0, 252, 4))\nnew_left_end = ...\nnew_right_end = ...\n\n# Don't change code below this line!\nprint(\"New mean-based estimate: {:f}\".format(new_mean_based_estimate))\nprint(\"Middle 95% of bootstrap estimates: [{:f}, {:f}]\".format(new_left_end, new_right_end))\n\nplt.plot(make_array(new_left_end, new_right_end), make_array(0, 0), color='yellow', lw=7, zorder=1)\nplt.scatter(150, 0, color='red', s=30, zorder=2);",
"_____no_output_____"
],
[
"grader.check(\"q3_2\")",
"_____no_output_____"
]
],
[
[
"<!-- BEGIN QUESTION -->\n\n#### Question 3.3\nDoes the interval covering the middle 95% of the new bootstrap estimates include `N`? If you ran that cell 100 times and generated 100 intervals, how many of those intervals would you expect to include `N`?\n\n<!--\nBEGIN QUESTION\nname: q3_3\nmanual: true\n-->",
"_____no_output_____"
],
[
"_Type your answer here, replacing this text._",
"_____no_output_____"
],
[
"<!-- END QUESTION -->\n\n\n\nLet's look at what happens when we use a small number of resamples:\n\n<img src=\"smallrephist.png\" width=\"525\"/>\n\nThis histogram and confidence interval was generated using 10 resamples of `new_observations`.",
"_____no_output_____"
],
[
"<!-- BEGIN QUESTION -->\n\n#### Question 3.4\nIn the cell below, explain why this histogram and confidence interval look different from the ones you generated previously in Question 3.2 where the number of resamples was 10,000.\n\n<!--\nBEGIN QUESTION\nname: q3_4\nmanual: true\n-->",
"_____no_output_____"
],
[
"_Type your answer here, replacing this text._",
"_____no_output_____"
],
[
"<!-- END QUESTION -->\n\n\n\n## 4. The CLT and Book Reviews\n\nYour friend has recommended you a book, so you look for it on an online marketplace. You decide to look at reviews for the book just to be sure that it's worth buying. Let's say that on Amazon, the book only has 80% positive reviews. On GoodReads, it has 95% positive reviews. You decide to investigate a bit further by looking at the percentage of positive reviews for the book on 5 different websites that you know of, and you collect these positive review percentages in a table called `reviews.csv`.",
"_____no_output_____"
],
[
"Here, we've loaded in the table for you.",
"_____no_output_____"
]
],
[
[
"reviews = Table.read_table(\"reviews.csv\") \nreviews",
"_____no_output_____"
]
],
[
[
"**Question 4.1**. Calculate the average percentage of positive reviews from your sample and assign it to `initial_sample_mean`.\n\n<!--\nBEGIN QUESTION\nname: q4_1\nmanual: false\n-->",
"_____no_output_____"
]
],
[
[
"initial_sample_mean = ...\ninitial_sample_mean",
"_____no_output_____"
],
[
"grader.check(\"q4_1\")",
"_____no_output_____"
]
],
[
[
"You've calculated the average percentage of positive reviews from your sample, so now you want to do some inference using this information. \n\n**Question 4.2**. First, simulate 5000 bootstrap resamples of the positive review percentages. For each bootstrap resample, calculate the resample mean and store the resampled means in an array called `resample_positive_percentages`. Then, plot a histogram of the resampled means.\n\n<!--\nBEGIN QUESTION\nname: q4_2\nmanual: false\n-->",
"_____no_output_____"
]
],
[
[
"resample_positive_percentages = make_array()\n\nfor i in np.arange(5000):\n resample = ...\n resample_avg_positive = ...\n resample_positive_percentages = ...\n \n# Do NOT change these lines.\n(Table().with_column(\"Average % of Positive Reviews in Resamples\", \n resample_positive_percentages).hist(\"Average % of Positive Reviews in Resamples\"))",
"_____no_output_____"
],
[
"grader.check(\"q4_2\")",
"_____no_output_____"
]
],
[
[
"**Question 4.3**. What is the the shape of the empirical distribution of the average percentage of positive reviews based on our original sample? What value is the distribution centered at? Assign your answer to the variable `initial_sample_mean_distribution`--your answer should be either `1`, `2`, `3`, or `4` corresponding to the following choices:\n\n*Hint: Look at the histogram you made in Question 2. Run the cell that generated the histogram a few times to check your intuition.*\n\n1. The distribution is approximately normal because of the Central Limit Theorem, and it is centered at the original sample mean.\n2. The distribution is not necessarily normal because the Central Limit Theorem may not apply, and it is centered at the original sample mean.\n3. The distribution is approximately normal because of the Central Limit Theorem, but it is not centered at the original sample mean.\n4. The distribution is not necessarily normal because the Central Limit Theorem may not apply, and it is not centered at the original sample mean.\n\n<!--\nBEGIN QUESTION\nname: q4_3\nmanual: false\n-->",
"_____no_output_____"
]
],
[
[
"initial_sample_mean_distribution = ...",
"_____no_output_____"
],
[
"grader.check(\"q4_3\")",
"_____no_output_____"
]
],
[
[
"<!-- BEGIN QUESTION -->\n\nAccording to the Central Limit Theorem, the probability distribution of the sum or average of a *large random sample* drawn with replacement will be roughly normal, regardless of the distribution of the population from which the sample is drawn. \n\n**Question 4.4**. Note the statement about the sample being large and random. Is this sample large and random? Give a brief explanation.\n\n*Note: The setup at the beginning of this exercise explains how the sample was gathered.*\n\n<!--\nBEGIN QUESTION\nname: q4_4\nmanual: true\n-->",
"_____no_output_____"
],
[
"_Type your answer here, replacing this text._",
"_____no_output_____"
],
[
"<!-- END QUESTION -->\n\nThough you have an estimate of the true percentage of positive reviews (the sample mean), you want to measure how variable this estimate is. \n\n**Question 4.5**. Find the standard deviation of your resampled average positive review percentages, which you stored in `resample_positive_percentages`, and assign the result to the variable `resampled_means_variability`.\n\n<!--\nBEGIN QUESTION\nname: q4_5\nmanual: false\n-->",
"_____no_output_____"
]
],
[
[
"resampled_means_variability = ...\nresampled_means_variability",
"_____no_output_____"
],
[
"grader.check(\"q4_5\")",
"_____no_output_____"
]
],
[
[
"This estimate is pretty variable! To make the estimate less variable, let's say you found a way to randomly sample reputable marketplaces from across the web which sell this book. Let's say that there are up to 150 of these marketplaces. The percentages of positive reviews are loaded into the table `more_reviews`.",
"_____no_output_____"
]
],
[
[
"# Just run this cell\nmore_reviews = Table.read_table(\"more_reviews.csv\")\nmore_reviews",
"_____no_output_____"
]
],
[
[
"In the next few questions, we'll test an important result of the Central Limit Theorem. According to the CLT, the standard deviation of all possible sample means can be calculated using the following formula:\n\n$$\n\\text{SD of all possible sample means} = \\dfrac{\\text{Population SD}}{\\sqrt{\\text{sample size}}}\n$$\n\nThis formula gives us another way to approximate the SD of the sample means other than calculating it empirically. We can test how well this formula works by calculating the SD of sample means for different sample sizes.\n\nThe following code calculates the SD of sample means using the CLT and empirically for a range of sample sizes. Then, it plots a scatter plot comparing the SD of the sample means calculated with both methods. Each point corresponds to a different sample size. ",
"_____no_output_____"
]
],
[
[
"# Just run this cell. It's not necessary for you to read this code, but you can do 99% of this on your own!\n# Note: this cell might take a bit to run.\n\ndef empirical_sample_mean_sd(n):\n sample_means = make_array()\n for i in np.arange(500):\n sample = more_reviews.sample(n).column('Positive Review Percentage') \n sample_mean = np.mean(sample) \n sample_means = np.append(sample_means, sample_mean) \n return np.std(sample_means)\n\ndef predict_sample_mean_sd(n):\n return np.std(more_reviews.column(0)) / (n**0.5)\n\nsd_table = Table().with_column('Sample Size', np.arange(1,151))\npredicted = sd_table.apply(predict_sample_mean_sd, 'Sample Size')\nempirical = sd_table.apply(empirical_sample_mean_sd, 'Sample Size')\nsd_table = sd_table.with_columns('Predicted SD', predicted, 'Empirical SD', empirical)\nsd_table.scatter('Sample Size')\nplt.ylabel(\"SD of Sample Mean\");",
"_____no_output_____"
]
],
[
[
"**Question 4.6**. Assign the numbers corresponding to all true statements to an array called `sample_mean_sd_statements`. \n\n1. The law of large numbers tells us that the distribution of a large random sample should resemble the distribution from which it is drawn.\n2. The SD of the sample means is proportional to the square root of the sample size. \n3. The SD of the sample means is proportional to 1 divided by the square root of the sample size. \n4. The law of large numbers guarantees that empirical and predicted sample mean SDs will be exactly equal to each other when the sample size is large.\n5. The law of large numbers guarantees that empirical and predicted sample mean SDs will be approximately equal to each other when the sample size is large.\n6. The plot above shows that as our sample size increases, our estimate for the true percentage of positive reviews becomes more accurate.\n7. The plot above shows that the size of the population affects the SD of the sample means.\n\n<!--\nBEGIN QUESTION\nname: q4_6\nmanual: false\n-->",
"_____no_output_____"
]
],
[
[
"sample_mean_sd_statements = ...",
"_____no_output_____"
],
[
"grader.check(\"q4_6\")",
"_____no_output_____"
]
],
[
[
"Often times, when conducting statistical inference, you'll want your estimate of a population parameter to have a certain accuracy. It is common to measure accuracy of an estimate using the SD of the estimate--as the SD goes down, your estimate becomes less variable. As a result, the width of the confidence interval for your estimate decreases (think about why this is true). We know from the Central Limit Theorem that when we estimate a sample mean, the SD of the sample mean decreases as the sample size increases (again, think about why this is true). \n\n**Question 4.7**. Imagine you are asked to estimate the true average percentage of positive reviews for this book and you have not yet taken a sample of review websites. Which of these is the best way to decide how large your sample should be to achieve a certain level of accuracy for your estimate of the true average percentage of positive reviews? Assign `sample_size_calculation` to either `1`, `2`, or `3` corresponding to the statements below.\n\n*Note: Assume you know the population SD or can estimate it with reasonable accuracy.*\n1. Take many random samples of different sizes, then calculate empirical confidence intervals using the bootstrap until you reach your desired accuracy. \n2. Use the Central Limit Theorem to calculate what sample size you need in advance. \n3. Randomly pick a sample size and hope for the best.\n\n<!--\nBEGIN QUESTION\nname: q4_7\nmanual: false\n-->",
"_____no_output_____"
]
],
[
[
"sample_size_calculation = ...",
"_____no_output_____"
],
[
"grader.check(\"q4_7\")",
"_____no_output_____"
]
],
[
[
"Congratulations, you're done with Homework 9! ",
"_____no_output_____"
],
[
"---\n\nTo double-check your work, the cell below will rerun all of the autograder tests.",
"_____no_output_____"
]
],
[
[
"grader.check_all()",
"_____no_output_____"
]
],
[
[
"## Submission\n\nMake sure you have run all cells in your notebook in order before running the cell below, so that all images/graphs appear in the output. The cell below will generate a zip file for you to submit. **Please save before exporting!**",
"_____no_output_____"
]
],
[
[
"# Save your notebook first, then run this cell to export your submission.\ngrader.export()",
"_____no_output_____"
]
],
[
[
" ",
"_____no_output_____"
]
]
] | [
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown"
] | [
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown",
"markdown",
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown",
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
]
] |
d0fe8999705eab10d2b0b01c79d4eb5ba876fad7 | 1,390 | ipynb | Jupyter Notebook | EuroSciPy-2019/3 - Tuesday/SciPy Tutorial.ipynb | mpenagar/Irakaskuntza-Docencia-2019-2020 | ac259c5a9949f13edb491a609e5cc6c7543344e9 | [
"MIT"
] | null | null | null | EuroSciPy-2019/3 - Tuesday/SciPy Tutorial.ipynb | mpenagar/Irakaskuntza-Docencia-2019-2020 | ac259c5a9949f13edb491a609e5cc6c7543344e9 | [
"MIT"
] | null | null | null | EuroSciPy-2019/3 - Tuesday/SciPy Tutorial.ipynb | mpenagar/Irakaskuntza-Docencia-2019-2020 | ac259c5a9949f13edb491a609e5cc6c7543344e9 | [
"MIT"
] | null | null | null | 22.063492 | 84 | 0.540288 | [
[
[
"# SciPy Tutorial",
"_____no_output_____"
]
],
[
[
"#!git clone https://github.com/gertingold/euroscipy-scipy-tutorial",
"Cloning into 'euroscipy-scipy-tutorial'...\nremote: Enumerating objects: 108, done.\u001b[K\nremote: Counting objects: 100% (108/108), done.\u001b[K\nremote: Compressing objects: 100% (84/84), done.\u001b[K\nremote: Total 108 (delta 40), reused 85 (delta 20), pack-reused 0\u001b[K\nReceiving objects: 100% (108/108), 21.53 MiB | 6.42 MiB/s, done.\nResolving deltas: 100% (40/40), done.\n"
]
]
] | [
"markdown",
"code"
] | [
[
"markdown"
],
[
"code"
]
] |
d0fe8a0479c95705588164f1b6f0ef76fa7b37d7 | 7,695 | ipynb | Jupyter Notebook | examples/basic-example.ipynb | sjdv1982/seamless | 1b814341e74a56333c163f10e6f6ceab508b7df9 | [
"MIT"
] | 15 | 2017-06-07T12:49:12.000Z | 2020-07-25T18:06:04.000Z | examples/basic-example.ipynb | sjdv1982/seamless | 1b814341e74a56333c163f10e6f6ceab508b7df9 | [
"MIT"
] | 110 | 2016-06-21T23:20:44.000Z | 2022-02-24T16:15:22.000Z | examples/basic-example.ipynb | sjdv1982/seamless | 1b814341e74a56333c163f10e6f6ceab508b7df9 | [
"MIT"
] | 6 | 2016-06-21T11:19:22.000Z | 2019-01-21T13:45:39.000Z | 20.630027 | 82 | 0.493047 | [
[
[
"# Basic example",
"_____no_output_____"
],
[
"### 1. Import Seamless in IPython or Jupyter ",
"_____no_output_____"
]
],
[
[
"from seamless.highlevel import Context\nctx = Context()",
"_____no_output_____"
]
],
[
[
"### 2. Set up a simple Seamless context",
"_____no_output_____"
]
],
[
[
"def add(a, b):\n return a + b\n\nctx.a = 10 # ctx.a => Seamless cell\nctx.b = 20 # ctx.b => Seamless cell\nctx.add = add # ctx.add => Seamless transformer\nctx.add.a = ctx.a\nctx.add.b = ctx.b\nctx.c = ctx.add # ctx.c => Seamless cell\nawait ctx.computation()\nctx.c.value",
"_____no_output_____"
],
[
"ctx.a += 5 \nawait ctx.computation()\nctx.c.value",
"_____no_output_____"
]
],
[
[
"### 3. Define schemas and validation rules",
"_____no_output_____"
]
],
[
[
"ctx.add.example.a = 0.0 # declares that add.a must be a number\nctx.add.example.b = 0.0 \n\ndef validate(self):\n assert self.a < self.b\n\nctx.add.add_validator(validate, name=\"validate\")\n\nawait ctx.computation()\nprint(ctx.add.exception)\n# Validation passes => exception is None",
"None\n"
]
],
[
[
"### 4. Create an API for a Seamless cell",
"_____no_output_____"
]
],
[
[
"def report(self): \n value = self.unsilk \n if value is None: \n print(\"Sorry, there is no result\") \n else: \n print(\"The result is: {}\".format(value))\n\nctx.c.example.report = report\nawait ctx.computation()\nctx.c.value.report()",
"The result is: 35\n"
]
],
[
[
"### 5. Mount cells to the file system",
"_____no_output_____"
]
],
[
[
"ctx.a.celltype = \"plain\"\nctx.a.mount(\"/tmp/a.txt\")\nctx.b.celltype = \"plain\"\nctx.b.mount(\"/tmp/b.txt\")\nctx.c.celltype = \"plain\"\nctx.c.mount(\"/tmp/c.txt\", mode=\"w\")\nctx.add.code.mount(\"/tmp/code.py\")\nawait ctx.translation()",
"d31923282290a41f9e2fcda93feb36aafdc11827f9bb4b63e11b2c099cf17325\nWarning: File path '/tmp/a.txt' has a different value, overwriting cell\n8e843baef228089dc379d4c3b6e28c1bb5d44eee257f1206b5dfee44ef6b05ad\nWarning: File path '/tmp/b.txt' has a different value, overwriting cell\n"
]
],
[
[
"### 6. Share a cell over HTTP",
"_____no_output_____"
]
],
[
[
"ctx.c.mimetype = \"text\"\nctx.c.share()\nawait ctx.translation()",
"_____no_output_____"
]
],
[
[
"### http://localhost:5813/ctx/c",
"_____no_output_____"
],
[
"### 7. Control cells from Jupyter",
"_____no_output_____"
]
],
[
[
"from ipywidgets import IntSlider, IntText\n\na = IntSlider(min=-10,max=30)\nb = IntSlider(min=-10,max=30)\nc = ctx.c.output()\nctx.a.traitlet().link(a)\nctx.b.traitlet().link(b)\ndisplay(a)\ndisplay(b)\ndisplay(c)",
"Opened the seamless share update server at port 5138\n"
]
],
[
[
"### 8. Save the entire state of the context",
"_____no_output_____"
]
],
[
[
"# Graph and checksums, as JSON\nctx.save_graph(\"basic-example.seamless\")\n# Checksum-to-buffer cache, as ZIP file\nctx.save_zip(\"basic-example.zip\")",
"Opened the seamless REST server at port 5813\n"
]
],
[
[
"### 9. In a new notebook / IPython console:",
"_____no_output_____"
],
[
"```python\nfrom seamless.highlevel import load_graph\nctx = load_graph(\n \"basic-example.seamless\", \n zip=\"basic-example.zip\"\n)\nawait ctx.computation()\nctx.c.value\n```",
"_____no_output_____"
],
[
"5",
"_____no_output_____"
],
[
"### http://localhost:5813/ctx/c",
"_____no_output_____"
]
]
] | [
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown"
] | [
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown",
"markdown",
"markdown"
]
] |
d0fe91f93085c9f8af6559d21eea8a4a56595f7f | 142,399 | ipynb | Jupyter Notebook | CAL3D_Printer/COD/Slicer V1/STL_SLICER_V5.ipynb | kaden-foy/kaden-foygithub.io | c0d296fc2f1dbab19713ae77ca0b15d8be20d67d | [
"CC-BY-3.0"
] | null | null | null | CAL3D_Printer/COD/Slicer V1/STL_SLICER_V5.ipynb | kaden-foy/kaden-foygithub.io | c0d296fc2f1dbab19713ae77ca0b15d8be20d67d | [
"CC-BY-3.0"
] | null | null | null | CAL3D_Printer/COD/Slicer V1/STL_SLICER_V5.ipynb | kaden-foy/kaden-foygithub.io | c0d296fc2f1dbab19713ae77ca0b15d8be20d67d | [
"CC-BY-3.0"
] | null | null | null | 329.627315 | 31,311 | 0.903595 | [
[
[
"# import Modules\nimport numpy as np\nfrom stl import mesh\nimport matplotlib.pyplot as plt\n\n%matplotlib widget",
"_____no_output_____"
],
[
"# import stl file\npart_mesh = mesh.Mesh.from_file('3DBenchy.stl')\nprint(\"File loaded in as faces and vertices\")",
"exception (False, \"b'\\\\xd7\\\\xd3@\\\\x00\\\\x00\\\\xa0\\\\xbf\\\\xdfo\\\\x9da\\\\x00\\\\x00\\\\xf95\\\\x7f\\\\xbf\\\\x0b\\\\xd5\\\\xc1&*\\\\xaf\\\\xa0=\\\\xa0\\\\x1a\\\\xd3@\\\\xbe\\\\x9f\\\\x9a\\\\xbf\\\\xc3\\\\xf5\\\\x9aa)\\\\\\\\\\\\xd3@\\\\xa4p\\\\x9d\\\\xbfj\\\\xbc\\\\x9bam\\\\xe7\\\\xd3@\\\\x00\\\\x00\\\\x00\\\\x00\\\\x12\\\\x83\\\\x9da\\\\x00\\\\x00\\\\xf95\\\\x7f\\\\xbf\\\\x86\\\\xaa\\\\xa9\\\\xa6*\\\\xaf\\\\xa0=)\\\\\\\\\\\\xd3@\\\\xa4p\\\\x9d\\\\xbfj\\\\xbc\\\\x9ba\\\\x81\\\\x95\\\\xd3@d;\\\\x9f\\\\xbf\\\\x1f\\\\x85\\\\x9cam\\\\xe7\\\\xd3@\\\\x00\\\\x00\\\\x00\\\\x00\\\\x12\\\\x83\\\\x9da\\\\x00\\\\x00\\\\xf95\\\\x7f\\\\xbf\\\\x1b\\\\x82\\\\x80%*\\\\xaf\\\\xa0=m\\\\xe7\\\\xd3@\\\\x00\\\\x00\\\\x00\\\\x00\\\\x12\\\\x83\\\\x9da\\\\x81\\\\x95\\\\xd3@d;\\\\x9f\\\\xbf\\\\x1f\\\\x85\\\\x9ca' should start with b'facet normal'\")\nFile loaded in as faces and vertices\n"
],
[
"# Slice STL file at each z value and return a \n# pair of points that \n\ndef STL_Slicer(stl_mesh,layers):\n # initilize points\n Set_1 = np.zeros([3,0])\n Set_2 = np.zeros([3,0])\n\n # save faces that dont would through error\n Bad_Faces = np.array([])\n\n l = len(stl_mesh.vectors[:])\n #l = 50000*2\n for face in range(l):\n # check if face is actually a line (bad stl file)\n v1 = stl_mesh.vectors[face][0,:] - stl_mesh.vectors[face][1,:]\n v2 = stl_mesh.vectors[face][1,:] - stl_mesh.vectors[face][2,:] \n v3 = stl_mesh.vectors[face][2,:] - stl_mesh.vectors[face][0,:]\n # check that face intercept z point\n z_lim = np.array([stl_mesh.vectors[face][:,2].min(),stl_mesh.vectors[face][:,2].max()])\n inter = (z_lim[0]-layers)*(z_lim[1]-layers) < 0\n \n if np.any(np.all(v1==0) or np.all(v2==0) or np.all(v3==0) or np.all(inter)):\n Bad_Faces = np.append(Bad_Faces,face) # for debugging\n # just compute random task\n else:\n \n # code used to find slice lines\n t = np.zeros([3]) # initilaize line parameter\n\n ## solve for all z intercepts\n idx = np.where((layers>z_lim[0]) & (layers<z_lim[1]))[0]\n\n # solve line intercept parameter value\n t = np.zeros([3,len(idx)])\n\n t[0,:] = np.transpose((layers[idx]-stl_mesh.vectors[face][0,2])/(stl_mesh.vectors[face][1,2]-stl_mesh.vectors[face][0,2]))\n t[1,:] = np.transpose((layers[idx]-stl_mesh.vectors[face][1,2])/(stl_mesh.vectors[face][2,2]-stl_mesh.vectors[face][1,2]))\n t[2,:] = np.transpose((layers[idx]-stl_mesh.vectors[face][2,2])/(stl_mesh.vectors[face][0,2]-stl_mesh.vectors[face][2,2]))\n\n II0 = np.array((stl_mesh.vectors[face][1,:]-stl_mesh.vectors[face][0,:]).reshape(-1,1)*t[0,:] + stl_mesh.vectors[face][0,:].reshape(-1,1))\n II1 = np.array((stl_mesh.vectors[face][2,:]-stl_mesh.vectors[face][1,:]).reshape(-1,1)*t[1,:] + stl_mesh.vectors[face][1,:].reshape(-1,1))\n II2 = np.array((stl_mesh.vectors[face][0,:]-stl_mesh.vectors[face][2,:]).reshape(-1,1)*t[2,:] + stl_mesh.vectors[face][2,:].reshape(-1,1))\n\n # remove points outside of bounds\n idx0 = ((II0[0,:]<stl_mesh.vectors[face][[0,1],0].min()) + (II0[0,:]>stl_mesh.vectors[face][[0,1],0].max()) \n + (II0[1,:]<stl_mesh.vectors[face][[0,1],1].min()) + (II0[1,:]>stl_mesh.vectors[face][[0,1],1].max())\n + (II0[2,:]<stl_mesh.vectors[face][[0,1],2].min()) + (II0[2,:]>stl_mesh.vectors[face][[0,1],2].max()))\n idx1 = ((II1[0,:]<stl_mesh.vectors[face][[1,2],0].min()) + (II1[0,:]>stl_mesh.vectors[face][[1,2],0].max()) \n + (II1[1,:]<stl_mesh.vectors[face][[1,2],1].min()) + (II1[1,:]>stl_mesh.vectors[face][[1,2],1].max())\n + (II1[2,:]<stl_mesh.vectors[face][[1,2],2].min()) + (II1[2,:]>stl_mesh.vectors[face][[1,2],2].max()))\n idx2 = ((II2[0,:]<stl_mesh.vectors[face][[0,2],0].min()) + (II2[0,:]>stl_mesh.vectors[face][[0,2],0].max()) \n + (II2[1,:]<stl_mesh.vectors[face][[0,2],1].min()) + (II2[1,:]>stl_mesh.vectors[face][[0,2],1].max())\n + (II2[2,:]<stl_mesh.vectors[face][[0,2],2].min()) + (II2[2,:]>stl_mesh.vectors[face][[0,2],2].max()))\n \n # save points for every intercept for every z level...\n II0[:,idx0] = 0\n II1[:,idx1] = 0\n II2[:,idx2] = 0\n\n # combine points into pairs of points that can be stored easily\n # find the ONE line that has all points within bonds. This is the base line\n # the secondary points are the sum of the other two points becuase index above \n # sets outof bounds points to zero\n if(sum(idx0)==0):\n P1 = II0\n P2 = II1 + II2\n elif(sum(idx1)==0):\n P1 = II1\n P2 = II0 + II2\n else:\n P1 = II2\n P2 = II0 + II1\n \n Set_1 = np.append(Set_1,P1,axis=1)\n Set_2 = np.append(Set_2,P2,axis=1)\n\n print(\"Num of Bad Faces: \",len(Bad_Faces),\" Total Num of Faces: \",len(stl_mesh.vectors[:]))\n return Set_1, Set_2\n\n# run slicer\nz = np.linspace(np.min(part_mesh.z),np.max(part_mesh.z),46) # z slices\np1, p2 = STL_Slicer(part_mesh,z)",
"C:\\Users\\kaden\\AppData\\Local\\Temp/ipykernel_10724/3201457791.py:37: RuntimeWarning: divide by zero encountered in true_divide\n t[0,:] = np.transpose((layers[idx]-stl_mesh.vectors[face][0,2])/(stl_mesh.vectors[face][1,2]-stl_mesh.vectors[face][0,2]))\nC:\\Users\\kaden\\AppData\\Local\\Temp/ipykernel_10724/3201457791.py:41: RuntimeWarning: invalid value encountered in multiply\n II0 = np.array((stl_mesh.vectors[face][1,:]-stl_mesh.vectors[face][0,:]).reshape(-1,1)*t[0,:] + stl_mesh.vectors[face][0,:].reshape(-1,1))\nC:\\Users\\kaden\\AppData\\Local\\Temp/ipykernel_10724/3201457791.py:39: RuntimeWarning: divide by zero encountered in true_divide\n t[2,:] = np.transpose((layers[idx]-stl_mesh.vectors[face][2,2])/(stl_mesh.vectors[face][0,2]-stl_mesh.vectors[face][2,2]))\nC:\\Users\\kaden\\AppData\\Local\\Temp/ipykernel_10724/3201457791.py:43: RuntimeWarning: invalid value encountered in multiply\n II2 = np.array((stl_mesh.vectors[face][0,:]-stl_mesh.vectors[face][2,:]).reshape(-1,1)*t[2,:] + stl_mesh.vectors[face][2,:].reshape(-1,1))\nC:\\Users\\kaden\\AppData\\Local\\Temp/ipykernel_10724/3201457791.py:38: RuntimeWarning: divide by zero encountered in true_divide\n t[1,:] = np.transpose((layers[idx]-stl_mesh.vectors[face][1,2])/(stl_mesh.vectors[face][2,2]-stl_mesh.vectors[face][1,2]))\nC:\\Users\\kaden\\AppData\\Local\\Temp/ipykernel_10724/3201457791.py:42: RuntimeWarning: invalid value encountered in multiply\n II1 = np.array((stl_mesh.vectors[face][2,:]-stl_mesh.vectors[face][1,:]).reshape(-1,1)*t[1,:] + stl_mesh.vectors[face][1,:].reshape(-1,1))\n"
],
[
"# Generate Rays that can penetrate slice\ndef Generate_Rays(Theta,Spacing,Num):\n # iterction math: https://www.cuemath.com/geometry/intersection-of-two-lines/\n Rays = np.zeros([3,Num]) # empty matrix with a,b and c values ax+by+c=0\n if (Theta == 0): #horizontal lines\n print(\"horizontal\")\n Rays[0,:] = 0\n Rays[1,:] = 1\n Rays[2,:] = np.arange(0, Spacing*Num,Spacing) - Num*Spacing/2 + Spacing/2\n elif (Theta == np.pi/2): # vertical lines\n print(\"Vertical\")\n Rays[0,:] = 1\n Rays[1,:] = 0\n Rays[2,:] = np.arange(0, Spacing*Num,Spacing) - Num*Spacing/2 + Spacing/2\n else:\n print(\"angle\")\n Rays[0,:] = -np.tan(Theta)\n Rays[1,:] = 1\n Rays[2,:] = (np.arange(0, Spacing*Num,Spacing) - Num*Spacing/2 + Spacing/2)/np.abs(np.cos(Theta))\n \n return Rays\n\n# plot Rays with a limiting box\ndef Plot_Rays(Rays,X_lim,Y_lim,color='y'):\n # takes ray parameters, x limit, and\n # y limit and plots the lines\n Y = np.zeros([2,len(Rays[0,:])])\n X = np.zeros([2,len(Rays[0,:])])\n\n M = -Rays[0,:]/Rays[1,:] # slope of lines\n idx_vert = (M >= 1) | (M < -1) # rays between 45-135 deg\n idx_line = idx_vert == 0 # all other rays\n\n # generate lines\n Y[0,idx_vert] += Y_lim[0]\n Y[1,idx_vert] += Y_lim[1]\n X[0,idx_vert] = -(Rays[1,idx_vert]*Y[0,idx_vert]+Rays[2,idx_vert])/Rays[0,idx_vert]\n X[1,idx_vert] = -(Rays[1,idx_vert]*Y[1,idx_vert]+Rays[2,idx_vert])/Rays[0,idx_vert]\n\n # generate lines\n X[0,idx_line] += X_lim[0]\n X[1,idx_line] += X_lim[1]\n Y[0,idx_line] = -(Rays[0,idx_line]*X[0,idx_line]+Rays[2,idx_line])/Rays[1,idx_line]\n Y[1,idx_line] = -(Rays[0,idx_line]*X[1,idx_line]+Rays[2,idx_line])/Rays[1,idx_line]\n\n # plot lines\n plt.plot(X,Y,color)\n plt.xlim(X_lim[0]*1.05,X_lim[1]*1.05)\n plt.ylim(Y_lim[0]*1.05,Y_lim[1]*1.05)\n return X,Y\n\nrays = Generate_Rays(0*np.pi/180,1.5,10)",
"horizontal\n"
],
[
"# find intercetion points between rays and slice\n # Generate Lines from 2 sets of points\ndef Generator_Lines(P1,P2):\n # takes to points P = [[x],[y]]\n # returns vector of a,b,c for line\n # ax+by+c=0\n A =-(P2[1] - P1[1])\n B = P2[0] - P1[0]\n C = -P1[0]*A - P1[1]*B\n return np.array([A,B,C])\n \ndef Line_Intersection(Line1,Line2):\n # takes P = [[a],[b],[c]] where ax+by+C=0\n # returns points of intersection [x,y]\n # for all possible line combose\n X = (np.outer(Line1[1],Line2[2])-np.outer(Line1[2],Line2[1]))/(np.outer(Line1[0],Line2[1])-np.outer(Line1[1],Line2[0]))\n Y = (np.outer(Line1[2],Line2[0])-np.outer(Line1[0],Line2[2]))/(np.outer(Line1[0],Line2[1])-np.outer(Line1[1],Line2[0]))\n return X,Y",
"_____no_output_____"
],
[
"# Function used to plot layer of point\ndef Plot_Slice(P1,P2,color='b',width=None):\n X1 = P1[0]\n X2 = P2[0]\n Y1 = P1[1]\n Y2 = P2[1]\n plt.plot([X1,X2],[Y1,Y2],color,linewidth=width) \n\n# select layer 100 and plot points\ni = p1[2,:] ==z[8]\np1_100 = p1[0:2,i].copy()\np2_100 = p2[0:2,i].copy()\n\nplt.close()\nPlot_Slice(p1_100,p2_100)\nplt.xlim([np.min(part_mesh.x),np.max(part_mesh.x)])\nplt.ylim([np.min(part_mesh.y),np.max(part_mesh.y)])\nplt.show()",
"_____no_output_____"
],
[
"# filter out data points that dont lie inbtween p1 p2\ndef In_Bound_Points(X_in, Y_in, P1, P2):\n # takes points of intersection of lines and\n # points of lines. replaces all points not \n # inbetween P1, P2 returns x,y\n X = X_in.copy()\n Y = Y_in.copy()\n X_max = np.max([P1[0],P2[0]],axis=0)\n X_min = np.min([P1[0],P2[0]],axis=0)\n Y_max = np.max([P1[1],P2[1]],axis=0)\n Y_min = np.min([P1[1],P2[1]],axis=0)\n tol = 1e-7\n idx = ((X<X_min-tol) | (X>X_max+tol)) | ((Y<Y_min-tol) | (Y>Y_max+tol))\n\n X[idx] = np.NaN\n Y[idx] = np.NaN\n return X,Y\n\nlines = Generator_Lines(p1_100,p2_100) # generate lines from slice 100\n\nrays = Generate_Rays(0*np.pi/180,0.25,100) # Generate rays\n\nxp,yp = Line_Intersection(rays,lines) # Calculate intersection matriz reutrn x y points\n\nxpf,ypf = In_Bound_Points(xp,yp,p1_100,p2_100) # filter out points that dont lie inbtween points\n\n# check number of parts of perercing points\nidx = np.sum(~np.isnan(xpf),axis=1)\nidx = (idx % 2) != 0\n\n# WHAT THE FUCK IS GOING ON????\n#print(\"boundary with problem\")\n#print(p1_100[:,878],p2_100[:,878])\n#print(\"line with problem\")\n#i = (xp[:,idx]>-5.62) & (xp[:,idx]<-5.61)\n#print(xp[i,idx],yp[i,idx])\n\n# plot results\n# fit around outline\nplt.close()\nPlot_Slice(p1_100,p2_100)\nx_lim = np.array([np.min(p1_100[0,:]),np.max(p1_100[0,:])])*1.05\ny_lim = np.array([np.min(p1_100[1,:]),np.max(p1_100[1,:])])*1.05\nPlot_Rays(rays,x_lim,y_lim)\nPlot_Rays(rays[:,idx],x_lim,y_lim,'r')\nplt.plot(xpf,ypf,'.g')\n#plt.plot(xp,yp,'.')\n\n#idx = 878 # slice(878,879)\n#Plot_Slice(p1_100[:,idx],p2_100[:,idx],'r')\n\nplt.xlim([np.min(part_mesh.x),np.max(part_mesh.x)])\nplt.ylim([np.min(part_mesh.y),np.max(part_mesh.y)])\nplt.show()",
"horizontal\n"
]
]
] | [
"code"
] | [
[
"code",
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
d0fe93591b97b7f7f45957760219c1c7b201ffd3 | 22,407 | ipynb | Jupyter Notebook | gradient descent/Gradient_Descent.ipynb | AkshatNGupta/Introduction-to-Machine-Learning | a37c9162d1ab9e7a1d98e6da4c9ad24d2adbc364 | [
"MIT"
] | 385 | 2019-02-25T20:59:35.000Z | 2022-03-31T16:33:37.000Z | gradient descent/Gradient_Descent.ipynb | AkshatNGupta/Introduction-to-Machine-Learning | a37c9162d1ab9e7a1d98e6da4c9ad24d2adbc364 | [
"MIT"
] | 3 | 2019-09-19T11:07:32.000Z | 2021-08-12T14:20:44.000Z | gradient descent/Gradient_Descent.ipynb | AkshatNGupta/Introduction-to-Machine-Learning | a37c9162d1ab9e7a1d98e6da4c9ad24d2adbc364 | [
"MIT"
] | 1,053 | 2019-02-27T02:32:18.000Z | 2022-03-30T14:52:39.000Z | 55.189655 | 3,488 | 0.692551 | [
[
[
">This notebook is part of our [Introduction to Machine Learning](http://www.codeheroku.com/course?course_id=1) course at [Code Heroku](http://www.codeheroku.com/).\n\nHey folks, today we are going to discuss about the application of gradient descent algorithm for solving machine learning problems. Let’s take a brief overview about the the things that we are going to discuss in this article:\n\n- What is gradient descent?\n- How gradient descent algorithm can help us solving machine learning problems\n- The math behind gradient descent algorithm\n- Implementation of gradient descent algorithm in Python\n\nSo, without wasting any time, let’s begin :)\n\n# What is gradient descent?\n\nHere’s what Wikipedia says: “Gradient descent is a first-order iterative optimization algorithm for finding the minimum of a function.”\n\nNow, you might be thinking “Wait, what does that mean? What do you want to say?”\n\nDon’t worry, we will elaborate everything about gradient descent in this article and all of it will start making sense to you in a moment :)\n\nTo understand gradient descent algorithm, let us first understand a real life machine learning problem: \nSuppose you have a dataset where you are provided with the number of hours a student studies per day and the percentage of marks scored by the corresponding student. If you plot a 2D graph of this dataset, it will look something like this:\n\n<img src=\"http://www.codeheroku.com/static/blog/images/grad_desc_1.png\">\n\nNow, if someone approaches you and says that a new student has taken admission and you need to predict the score of that student based on the number of hours he studies. How would you do that? \nTo predict the score of the new student, at first you need to find out a relationship between “Hours studied” and “Score” from the existing dataset. By taking a look over the visual graph plotting, we can see that a linear relationship can be established between these two things. So, by drawing a straight line over the data points in the graph, we can establish the relationship. Let’s see how would it look if we try to draw a straight line over the data points. It would look something like this:\n\n<img src=\"http://www.codeheroku.com/static/blog/images/grad_desc_2.png\">\n\nGreat! Now we have the relationship between “Hours Studied” and “Score”. So, now if someone asks us to predict the score of a student who studies 10 hours per day, we can just simply put Hours Studied = 10 data point over the relationship line and predict the value of his score like this:\n\n<img src=\"http://www.codeheroku.com/static/blog/images/grad_desc_3.png\">\n\nFrom the above picture, we can easily say that the new student who studies 10 hours per day would probably score around 60. Pretty easy, right? By the way, the relationship line that we have drawn is called the “regression” line. And because the relationship we have established is a linear relationship, the line is actually called “linear regression” line. Hence, this machine learning model that we have created is known as linear regression model.\n\nAt this point, you might have noticed that all the data points do not lie perfectly over the regression line. So, there might be some difference between the predicted value and the actual value. We call this difference as error(or cost).\n\n<img src=\"http://www.codeheroku.com/static/blog/images/grad_desc_err.png\">\n\nIn machine learning world, we always try to build a model with as minimum error as possible. To achieve this, we have to calculate the error of our model in order to best fit the regression line over it. We have different kinds of error like- total error, mean error, mean squared error etc.\n\nTotal error: Summation of the absolute difference between predicted and actual value for all the data points. Mathematically, this is\n\n<img src=\"http://www.codeheroku.com/static/blog/images/grad_desc_4.png\">\n\nMean error: Total error / number of data points. Mathematically, this is\n\n<img src=\"http://www.codeheroku.com/static/blog/images/grad_desc_5.png\">\n\nMean squared error: Summation of the square of absolute difference / number of data points. Mathematically, this is\n\n<img src=\"http://www.codeheroku.com/static/blog/images/grad_desc_6.png\">\n\nBelow is an example of calculating these errors:\n\n<img src=\"http://www.codeheroku.com/static/blog/images/error_calc.png\">\n\nWe will use the Mean Squared Error(M.S.E) to calculate the error and determine the best linear regression line(the line with the minimum error value) for our model.\n\nNow the question is, how would you represent a regression line in a computer?\n\nThe answer is simple. Remember the equation of a straight line? We can use the same equation in order to represent the regression line in computer. If you can’t recall it, let me quickly remind you, it’s **y = M * x + B**\n\n<img src=\"http://www.codeheroku.com/static/blog/images/line_repr.png\">\n\nHere, M is the slope of the line and B is the Y intercept. Let’s quickly recall about slope and Y intercept. \nSlope is the amount by which the line is rising on the Y axis for every block that you go towards right in the X axis. This tells us the direction of the line and the rate by which our line is increasing. Mathematically speaking, this means  for a specified amount of distance on the line.\n\nFrom the dotted lines in the above picture, we can see that for every 2 blocks in the X axis, the line rises by 1 block in the Y axis.<br>\nHence, slope, M = ½ = 0.5<br>\nAnd it’s a positive value, which indicates that the line is increasing in the upward direction.\n\nNow, let’s come to Y intercept. It is the distance which tells us exactly where the line cuts the Y axis. From the above picture, we can see that the line is cutting Y axis on point (0,1). So, the Y intercept(B) in this case is the distance between (0,0) and (0,1) = 1.\n\nHence, the straight line on the above picture can be represented through the following equation:\n\ny = 0.5 * x + 1\n\nNow we know how to represent the regression line in a computer. Everything seems good so far. But, the biggest question still remains unanswered- “How would the computer know the right value of M and B for drawing the regression line with the minimum error?”\n\nExactly that’s why we need the gradient descent algorithm. Gradient descent is a trial and error method, which will iteratively give us different values of M and B to try. In each iteration, we will draw a regression line using these values of M and B and will calculate the error for this model. We will continue until we get the values of M and B such that the error is minimum.\n\nLet’s have a more elaborative view of gradient descent algorithm:\n\nStep 1: Start with random values of M and B\n\n<img src=\"http://www.codeheroku.com/static/blog/images/grad_desc_s1.png\">\n\nStep 2: Adjust M and B such that error reduces\n\n<img src=\"http://www.codeheroku.com/static/blog/images/grad_desc_s2.png\">\n\nStep 3: Repeat until we get the best values of M and B (until convergence)\n\n<img src=\"http://www.codeheroku.com/static/blog/images/grad_desc_s3.png\">\n\nBy the way, the application of gradient descent is not limited to regression problems only. It is an optimization algorithm which can be applied to any problem in general.\n\n# The math behind gradient descent\n\nTill now we have understood that we will use gradient descent to minimize the error for our model. But, now let us see exactly how gradient descent finds the best values of M and B for us.\n\nGradient descent tries to minimize the error. Right?\n\nSo, we can say that it tries to minimize the following function(cost function):\n\n<img src=\"http://www.codeheroku.com/static/blog/images/gd_err_fnc.png\">\n\nAt first we will take random values of M and B. So, we will get a random error corresponding to these values. Thus, a random point will be plotted on the above graph. At this point, there will be some error. So, our objective will be to reduce this error.\n\nIn general, how would you approach towards the minimum value of a function? By finding its derivative. Right? The same thing applies here.\n\nWe will obtain the partial derivative of J with respect to M and B. This will give us the direction of the slope of tangent at the given point. We would like to move in the opposite direction of the slope in order to approach towards the minimum value.\n\n<img src=\"http://www.codeheroku.com/static/blog/images/gd_db_dm_calc.png\">\n\nSo far, we have only got the direction of the slope and we know we need to move in its opposite direction. But, in each iteration, by how much amount we should move in the opposite direction? This amount is called the learning rate(alpha). Learning rate determines the step size of our movement towards the minimal point.\n\nSo, choosing the right learning rate is very important. If the learning rate is too small, it will take more time to converge. On the other hand, if the learning rate is very high, it may overshoot the minimum point and diverge.\n\n<img src=\"http://www.codeheroku.com/static/blog/images/gd_ch_alpha.png\">\n\nTo sum up, what we have till now is-\n\n1. A random point is chosen initially by choosing random values of M and B.\n2. Direction of the slope of that point is found by finding delta_m and delta_b\n3. Since we want to move in the opposite direction of the slope, we will multiply -1 with both delta_m and delta_b.\n4. Since delta_m and delta_b gives us only the direction, we need to multiply both of them with the learning rate(alpha) to specify the step size of each iteration.\n5. Next, we need to modify the current values of M and B such that the error is reduced.\n<img src=\"http://www.codeheroku.com/static/blog/images/gd_9.png\">\n6. We need to repeat steps 2 to 5 until we converge at the minimum point.\n\n# Implementation of gradient descent using Python\n\nThis was everything about gradient descent algorithm. Now we will implement this algorithm using Python.\n\nLet us first import all required libraries and read the dataset using Pandas library(the csv file can be downloaded from this [link](https://github.com/codeheroku/Introduction-to-Machine-Learning/tree/master/gradient%20descent/starter%20code)):\n",
"_____no_output_____"
]
],
[
[
"import pandas as pd\nimport matplotlib.pyplot as plt\n\ndf = pd.read_csv(\"student_scores.csv\") #Read csv file using Pandas library",
"_____no_output_____"
]
],
[
[
"Next, we need to read the values of X and Y from the dataframe and create a scatter plot of that data.",
"_____no_output_____"
]
],
[
[
"X = df[\"Hours\"] #Read values of X from dataframe\nY = df[\"Scores\"] #Read values of Y from dataframe\n\nplt.plot(X,Y,'o') # 'o' for creating scatter plot\nplt.title(\"Implementing Gradient Descent\")\nplt.xlabel(\"Hours Studied\")\nplt.ylabel(\"Student Score\")",
"_____no_output_____"
]
],
[
[
"After that, we will initially choose m = 0 and b = 0",
"_____no_output_____"
]
],
[
[
"m = 0\nb = 0",
"_____no_output_____"
]
],
[
[
"Now, we need to create a function(gradient descent function) which will take the current value of m and b and then give us better values of m and b.",
"_____no_output_____"
]
],
[
[
"def grad_desc(X,Y,m,b):\n for point in zip(X,Y):\n x = point[0] #value of x of a point\n y_actual = point[1] #Actual value of y for that point\n\n y_prediction = m*x + b #Predicted value of y for given x\n\n error = y_prediction - y_actual #Error in the estimation \n \n #Using alpha = 0.0005\n delta_m = -1 * (error*x) * 0.0005 #Calculating delta m\n delta_b = -1 * (error) * 0.0005 #Calculating delta b\n\n m = m + delta_m #Modifying value of m for reducing error\n b = b + delta_b #Modifying value of b for reducing error\n\n return m,b #Returning better values of m and b",
"_____no_output_____"
]
],
[
[
"Notice, in the above code, we are using learning rate(alpha) = 0.0005 . You can try to modify this value and try this example with different learning rates.\n\nNow we will make a function which will help us to plot the regression line on the graph.",
"_____no_output_____"
]
],
[
[
"def plot_regression_line(X,m,b):\n regression_x = X.values #list of values of x\n regression_y = [] #list of values of y\n for x in regression_x:\n y = m*x + b #calculating the y_prediction\n regression_y.append(y) #adding the predicted value in list of y\n\n plt.plot(regression_x,regression_y) #plot the regression line\n plt.pause(1) #pause for 1 second before plotting next line",
"_____no_output_____"
]
],
[
[
"Now, when we will run the grad_desc() function, each time we will get a better result for regression line. Let us create a loop and run the grad_desc() function for 10 times and visualize the results.",
"_____no_output_____"
]
],
[
[
"for i in range(0,10):\n m,b = grad_desc(X,Y,m,b) #call grad_desc() to get better m & b\n plot_regression_line(X,m,b) #plot regression line with m & b",
"_____no_output_____"
]
],
[
[
"Finally, we need to show the plot by adding the following statement:",
"_____no_output_____"
]
],
[
[
"plt.show()",
"_____no_output_____"
]
],
[
[
"So, the full code for our program is:",
"_____no_output_____"
]
],
[
[
"import pandas as pd\nimport matplotlib.pyplot as plt \n\n# function for plotting regression line\ndef plot_regression_line(X,m,b):\n regression_x = X.values\n regression_y = []\n for x in regression_x:\n y = m*x + b\n regression_y.append(y)\n\n plt.plot(regression_x,regression_y)\n plt.pause(1)\n\n\ndf = pd.read_csv(\"student_scores.csv\")\n\nX = df[\"Hours\"]\nY = df[\"Scores\"]\n\nplt.plot(X,Y,'o')\nplt.title(\"Implementing Gradient Descent\")\nplt.xlabel(\"Hours Studied\")\nplt.ylabel(\"Student Score\")\n\nm = 0\nb = 0\n\n# gradient descent function\ndef grad_desc(X,Y,m,b):\n for point in zip(X,Y):\n x = point[0]\n y_actual = point[1]\n\n y_prediction = m*x + b\n\n error = y_prediction - y_actual\n\n delta_m = -1 * (error*x) * 0.0005\n delta_b = -1 * (error) * 0.0005\n\n m = m + delta_m\n b = b + delta_b\n return m,b\t \n\n\nfor i in range(0,10):\n m,b = grad_desc(X,Y,m,b)\n plot_regression_line(X,m,b)\n\nplt.show()",
"_____no_output_____"
]
],
[
[
"Now let’s run the above program for different values of learning rate(alpha).\n\nFor alpha = 0.0005 , the output will look like this:\n\n<img src=\"http://www.codeheroku.com/static/blog/images/gd_alpha_1.gif\">\n\nFor alpha = 0.05 , it will look like this:\n\n<img src=\"http://www.codeheroku.com/static/blog/images/gd_alpha_2.gif\">\n\nFor alpha = 1, it will overshoot the minimum point and diverge like this:\n\n<img src=\"http://www.codeheroku.com/static/blog/images/gd_alpha_3.gif\">\n\nThe gradient descent algorithm about which we discussed in this article is called stochastic gradient descent. There are also other types of gradient descent algorithms like- batch gradient descent, mini batch gradient descent etc.\n\n>If this article was helpful to you, check out our [Introduction to Machine Learning](http://www.codeheroku.com/course?course_id=1) Course at [Code Heroku](http://www.codeheroku.com/) for a complete guide to Machine Learning.",
"_____no_output_____"
]
]
] | [
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown"
] | [
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
]
] |
d0fe9602fc4866fb28dffdd8f6c7f05772a7926f | 18,183 | ipynb | Jupyter Notebook | 2017/coursera/code/0120/zero_based_generate_word.ipynb | janguck/lab_study_group | 47f9cbc2e04c3d5b72ad66781ac07fce003b3521 | [
"MIT"
] | 9 | 2018-03-11T21:34:38.000Z | 2021-05-31T05:46:38.000Z | 2017/coursera/code/0120/zero_based_generate_word.ipynb | janguck/lab_study_group | 47f9cbc2e04c3d5b72ad66781ac07fce003b3521 | [
"MIT"
] | 17 | 2017-07-10T06:22:19.000Z | 2019-03-19T11:25:04.000Z | 2017/coursera/code/0120/zero_based_generate_word.ipynb | janguck/lab_study_group | 47f9cbc2e04c3d5b72ad66781ac07fce003b3521 | [
"MIT"
] | 22 | 2017-07-03T07:53:44.000Z | 2019-04-03T00:32:55.000Z | 26.779087 | 122 | 0.539845 | [
[
[
"import csv\nimport itertools\nimport operator\nimport numpy as np\nimport nltk\nimport sys\nfrom datetime import datetime\nfrom utils import *\nimport matplotlib.pyplot as plt\n%matplotlib inline",
"_____no_output_____"
],
[
"vocabulary_size = 200\nsentence_start_token = \"START\"\nsentence_end_token = \"END\"\n\nf = open('data/ratings_train.txt', 'r')\nlines = f.readlines()\nfor i in range(len(lines)):\n lines[i] = lines[i].replace(\"/n\",\"\").replace(\"\\n\",\"\")\nreader = []\nfor line in lines:\n line_document = line.split(\"\\t\")[1]\n reader.append(line_document)\nf.close()",
"_____no_output_____"
],
[
"sentences = [\"%s %s %s\" % (sentence_start_token, x, sentence_end_token) for x in reader[:1000]]",
"_____no_output_____"
],
[
"from konlpy.tag import Twitter\npos_tagger = Twitter()\ndef tokenize(doc):\n return ['/'.join(t) for t in pos_tagger.pos(doc, norm=True, stem=True)]\ntokenized_sentences = [tokenize(row) for row in sentences]",
"_____no_output_____"
],
[
"vocab = [t for d in tokenized_sentences for t in d]",
"_____no_output_____"
],
[
"Verb_Noun_Adjective_Alpha_in_text = []\nindex = 0\nfor text in tokenized_sentences:\n Verb_Noun_Adjective_Alpha_in_text.append([])\n for word in text:\n parts_of_speech = word.split(\"/\")\n if parts_of_speech[1] in [\"Noun\",\"Verb\",\"Adjective\"] :\n Verb_Noun_Adjective_Alpha_in_text[index].append(word.split(\"/\")[0])\n elif parts_of_speech[1] in [\"Alpha\"] and len(parts_of_speech[0]) ==3 or len(parts_of_speech[0]) ==5:\n Verb_Noun_Adjective_Alpha_in_text[index].append(word.split(\"/\")[0]) \n index += 1",
"_____no_output_____"
],
[
"Verb_Noun_Adjective_Alpha_in_text_tokens = [t for d in Verb_Noun_Adjective_Alpha_in_text for t in d]",
"_____no_output_____"
],
[
"import nltk\nreal_tokens = nltk.Text(Verb_Noun_Adjective_Alpha_in_text_tokens, name='RNN')",
"_____no_output_____"
],
[
"real_tokens_freq = real_tokens.vocab().most_common(vocabulary_size-1)",
"_____no_output_____"
],
[
"index_to_word = [x[0] for x in real_tokens_freq]\nindex_to_word.append(\"unknown\")\nword_to_index = dict([(w,i) for i,w in enumerate(index_to_word)])",
"_____no_output_____"
],
[
"for i, sent in enumerate(Verb_Noun_Adjective_Alpha_in_text):\n tokenized_sentences[i] = [w if w in word_to_index else \"unknown\" for w in sent]",
"_____no_output_____"
]
],
[
[
"# Make model",
"_____no_output_____"
]
],
[
[
"X_train = np.asarray([[word_to_index[w] for w in sent[:-1]] for sent in tokenized_sentences])\ny_train = np.asarray([[word_to_index[w] for w in sent[1:]] for sent in tokenized_sentences])",
"_____no_output_____"
],
[
"X_train[0]",
"_____no_output_____"
],
[
"class RNNNumpy:\n \n def __init__(self, word_dim, hidden_dim=100, bptt_truncate=4):\n self.word_dim = word_dim\n self.hidden_dim = hidden_dim\n self.bptt_truncate = bptt_truncate\n self.U = np.random.uniform(-np.sqrt(1./word_dim), np.sqrt(1./word_dim), (hidden_dim, word_dim))\n self.V = np.random.uniform(-np.sqrt(1./hidden_dim), np.sqrt(1./hidden_dim), (word_dim, hidden_dim))\n self.W = np.random.uniform(-np.sqrt(1./hidden_dim), np.sqrt(1./hidden_dim), (hidden_dim, hidden_dim))",
"_____no_output_____"
],
[
"def forward_propagation(self, x):\n\n T = len(x)\n s = np.zeros((T + 1, self.hidden_dim))\n s[-1] = np.zeros(self.hidden_dim)\n o = np.zeros((T, self.word_dim))\n\n for t in np.arange(T):\n s[t] = np.tanh(self.U[:,x[t]] + self.W.dot(s[t-1]))\n o[t] = softmax(self.V.dot(s[t]))\n return [o, s]\n\nRNNNumpy.forward_propagation = forward_propagation",
"_____no_output_____"
],
[
"def predict(self, x):\n \n o, s = self.forward_propagation(x)\n return np.argmax(o, axis=1)\n\nRNNNumpy.predict = predict\ntokenized_sentences[0]",
"_____no_output_____"
],
[
"np.random.seed(100)\nmodel = RNNNumpy(vocabulary_size)\n#for i in range(100):\no, s = model.forward_propagation(X_train[0])\nprint (o.shape)\nprint (s.shape)",
"(6, 200)\n(7, 100)\n"
],
[
"X_train[0]",
"_____no_output_____"
],
[
"y_train[0]",
"_____no_output_____"
],
[
"predictions = model.predict(X_train[0])\nprint (predictions.shape)\nprint (predictions)",
"(6,)\n[ 52 91 192 67 13 186]\n"
],
[
"def calculate_total_loss(self, x, y):\n L = 0\n for i in np.arange(len(y)):\n o, s = self.forward_propagation(x[i])\n correct_word_predictions = o[np.arange(len(y[i])), y[i]]\n L += -1 * np.sum(np.log(correct_word_predictions))\n return L\n\ndef calculate_loss(self, x, y):\n N = np.sum((len(y_i) for y_i in y))\n return self.calculate_total_loss(x,y)/N\n\nRNNNumpy.calculate_total_loss = calculate_total_loss\nRNNNumpy.calculate_loss = calculate_loss",
"_____no_output_____"
],
[
"# Limit to 1000 examples to save time\nprint (\"Expected Loss for random predictions: %f\" % np.log(vocabulary_size))\nprint (\"Actual loss: %f\" % model.calculate_loss(X_train[:1000], y_train[:1000]))",
"Expected Loss for random predictions: 5.298317\nActual loss: 5.301008\n"
],
[
"def bptt(self, x, y):\n T = len(y)\n # Perform forward propagation\n o, s = self.forward_propagation(x)\n # We accumulate the gradients in these variables\n dLdU = np.zeros(self.U.shape)\n dLdV = np.zeros(self.V.shape)\n dLdW = np.zeros(self.W.shape)\n delta_o = o\n delta_o[np.arange(len(y)), y] -= 1.\n # For each output backwards...\n for t in np.arange(T)[::-1]:\n dLdV += np.outer(delta_o[t], s[t].T)\n # Initial delta calculation\n delta_t = self.V.T.dot(delta_o[t]) * (1 - (s[t] ** 2))\n # Backpropagation through time (for at most self.bptt_truncate steps)\n for bptt_step in np.arange(max(0, t-self.bptt_truncate), t+1)[::-1]:\n # print \"Backpropagation step t=%d bptt step=%d \" % (t, bptt_step)\n dLdW += np.outer(delta_t, s[bptt_step-1]) \n dLdU[:,x[bptt_step]] += delta_t\n # Update delta for next step\n delta_t = self.W.T.dot(delta_t) * (1 - s[bptt_step-1] ** 2)\n return [dLdU, dLdV, dLdW]\n\nRNNNumpy.bptt = bptt",
"_____no_output_____"
],
[
"def numpy_sdg_step(self, x, y, learning_rate):\n \n dLdU, dLdV, dLdW = self.bptt(x, y)\n self.U -= learning_rate * dLdU\n self.V -= learning_rate * dLdV\n self.W -= learning_rate * dLdW\n\nRNNNumpy.sgd_step = numpy_sdg_step",
"_____no_output_____"
],
[
"def train_with_sgd(model, X_train, y_train, learning_rate=0.005, nepoch=100, evaluate_loss_after=5):\n \n losses = []\n num_examples_seen = 0\n for epoch in range(nepoch):\n if (epoch % evaluate_loss_after == 0):\n loss = model.calculate_loss(X_train, y_train)\n losses.append((num_examples_seen, loss))\n time = datetime.now().strftime('%Y-%m-%d %H:%M:%S')\n print (\"%s: Loss after num_examples_seen=%d epoch=%d: %f\" % (time, num_examples_seen, epoch, loss))\n for i in range(len(y_train)):\n model.sgd_step(X_train[i], y_train[i], learning_rate)\n num_examples_seen += 1\n print(model)",
"_____no_output_____"
],
[
"np.random.seed(10)\nmodel = RNNNumpy(vocabulary_size)\n%timeit model.sgd_step(X_train[10], y_train[10], 0.005)",
"1000 loops, best of 3: 968 µs per loop\n"
],
[
"np.random.seed(10)\nmodel = RNNNumpy(vocabulary_size)\nlosses = train_with_sgd(model, X_train[:100], y_train[:100], nepoch=10, evaluate_loss_after=1)",
"2017-01-20 14:52:31: Loss after num_examples_seen=0 epoch=0: 5.304216\n2017-01-20 14:52:31: Loss after num_examples_seen=100 epoch=1: 3.442786\n2017-01-20 14:52:31: Loss after num_examples_seen=200 epoch=2: 3.131067\n2017-01-20 14:52:32: Loss after num_examples_seen=300 epoch=3: 3.017680\n2017-01-20 14:52:32: Loss after num_examples_seen=400 epoch=4: 2.958618\n2017-01-20 14:52:32: Loss after num_examples_seen=500 epoch=5: 2.930790\n2017-01-20 14:52:33: Loss after num_examples_seen=600 epoch=6: 2.916276\n2017-01-20 14:52:33: Loss after num_examples_seen=700 epoch=7: 2.905613\n2017-01-20 14:52:33: Loss after num_examples_seen=800 epoch=8: 2.895897\n2017-01-20 14:52:34: Loss after num_examples_seen=900 epoch=9: 2.886424\n<__main__.RNNNumpy object at 0x7f7a9c25ac50>\n"
],
[
"from rnn_theano import RNNTheano, gradient_check_theano",
"_____no_output_____"
],
[
"from utils import load_model_parameters_theano, save_model_parameters_theano\n\nmodel = RNNTheano(vocabulary_size, hidden_dim=100)\ntrain_with_sgd(model, X_train, y_train, nepoch=50)\n\nsave_model_parameters_theano('./data/trained-model-sion_consider.npz', model)\nload_model_parameters_theano('./data/trained-model-sion_consider.npz', model)",
"2017-01-20 14:52:41: Loss after num_examples_seen=0 epoch=0: 5.301648\n2017-01-20 14:52:46: Loss after num_examples_seen=5000 epoch=5: 3.193938\n2017-01-20 14:52:52: Loss after num_examples_seen=10000 epoch=10: 3.181974\n2017-01-20 14:52:58: Loss after num_examples_seen=15000 epoch=15: 3.177310\n2017-01-20 14:53:03: Loss after num_examples_seen=20000 epoch=20: 3.171975\n2017-01-20 14:53:10: Loss after num_examples_seen=25000 epoch=25: 3.154365\n2017-01-20 14:53:15: Loss after num_examples_seen=30000 epoch=30: 3.145856\n2017-01-20 14:53:20: Loss after num_examples_seen=35000 epoch=35: 3.141625\n2017-01-20 14:53:26: Loss after num_examples_seen=40000 epoch=40: 3.172117\n2017-01-20 14:53:31: Loss after num_examples_seen=45000 epoch=45: 3.152063\n<rnn_theano.RNNTheano object at 0x7f7add1e0710>\nSaved model parameters to ./data/trained-model-sion_consider.npz.\nLoaded model parameters from ./data/trained-model-sion_consider.npz. hidden_dim=100 word_dim=200\n"
],
[
"print(len(model.V.get_value()))",
"200\n"
],
[
"def generate_sentence(model):\n new_sentence = [word_to_index[sentence_start_token]]\n while not new_sentence[-1] == word_to_index[sentence_end_token]:\n next_word_probs = model.forward_propagation(new_sentence)\n sampled_word = word_to_index[\"unknown\"]\n while sampled_word == word_to_index[\"unknown\"]:\n samples = np.random.multinomial(1, next_word_probs[-1])\n sampled_word = np.argmax(samples)\n new_sentence.append(sampled_word)\n sentence_str = [index_to_word[x] for x in new_sentence[1:-1]]\n return sentence_str\n\nnum_sentences = 2\nsenten_min_length = 5\n\nfor i in range(num_sentences):\n sent = []\n while len(sent) < senten_min_length:\n sent = generate_sentence(model)\n print (\" \".join(sent))",
"점 감동 현실 왜 재밌다\n재미없다 말 끝 어떻다 받다\n"
]
]
] | [
"code",
"markdown",
"code"
] | [
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
d0fe976d330ff1ad8640a941262b089bbc1fab45 | 372,080 | ipynb | Jupyter Notebook | 10_neural_nets_with_keras.ipynb | agniji/handson-ml2 | 10e795e5140c0416f3a651ca80ba88a84c68104e | [
"Apache-2.0"
] | 1 | 2021-03-29T17:36:54.000Z | 2021-03-29T17:36:54.000Z | 10_neural_nets_with_keras.ipynb | agniji/handson-ml2 | 10e795e5140c0416f3a651ca80ba88a84c68104e | [
"Apache-2.0"
] | 4 | 2021-06-08T21:20:31.000Z | 2022-03-12T00:24:55.000Z | 10_neural_nets_with_keras.ipynb | agniji/handson-ml2 | 10e795e5140c0416f3a651ca80ba88a84c68104e | [
"Apache-2.0"
] | 2 | 2019-10-25T03:11:24.000Z | 2019-12-13T13:19:31.000Z | 120.336352 | 75,264 | 0.827158 | [
[
[
"**Chapter 10 – Introduction to Artificial Neural Networks with Keras**\n\n_This notebook contains all the sample code and solutions to the exercises in chapter 10._",
"_____no_output_____"
],
[
"# Setup",
"_____no_output_____"
],
[
"First, let's import a few common modules, ensure MatplotLib plots figures inline and prepare a function to save the figures. We also check that Python 3.5 or later is installed (although Python 2.x may work, it is deprecated so we strongly recommend you use Python 3 instead), as well as Scikit-Learn ≥0.20 and TensorFlow ≥2.0-preview.",
"_____no_output_____"
]
],
[
[
"# Python ≥3.5 is required\nimport sys\nassert sys.version_info >= (3, 5)\n\n# Scikit-Learn ≥0.20 is required\nimport sklearn\nassert sklearn.__version__ >= \"0.20\"\n\n# TensorFlow ≥2.0-preview is required\nimport tensorflow as tf\nassert tf.__version__ >= \"2.0\"\n\n# Common imports\nimport numpy as np\nimport os\n\n# to make this notebook's output stable across runs\nnp.random.seed(42)\n\n# To plot pretty figures\n%matplotlib inline\nimport matplotlib as mpl\nimport matplotlib.pyplot as plt\nmpl.rc('axes', labelsize=14)\nmpl.rc('xtick', labelsize=12)\nmpl.rc('ytick', labelsize=12)\n\n# Where to save the figures\nPROJECT_ROOT_DIR = \".\"\nCHAPTER_ID = \"ann\"\nIMAGES_PATH = os.path.join(PROJECT_ROOT_DIR, \"images\", CHAPTER_ID)\nos.makedirs(IMAGES_PATH, exist_ok=True)\n\ndef save_fig(fig_id, tight_layout=True, fig_extension=\"png\", resolution=300):\n path = os.path.join(IMAGES_PATH, fig_id + \".\" + fig_extension)\n print(\"Saving figure\", fig_id)\n if tight_layout:\n plt.tight_layout()\n plt.savefig(path, format=fig_extension, dpi=resolution)\n\n# Ignore useless warnings (see SciPy issue #5998)\nimport warnings\nwarnings.filterwarnings(action=\"ignore\", message=\"^internal gelsd\")",
"_____no_output_____"
]
],
[
[
"# Perceptrons",
"_____no_output_____"
],
[
"**Note**: we set `max_iter` and `tol` explicitly to avoid warnings about the fact that their default value will change in future versions of Scikit-Learn.",
"_____no_output_____"
]
],
[
[
"import numpy as np\nfrom sklearn.datasets import load_iris\nfrom sklearn.linear_model import Perceptron\n\niris = load_iris()\nX = iris.data[:, (2, 3)] # petal length, petal width\ny = (iris.target == 0).astype(np.int)\n\nper_clf = Perceptron(max_iter=1000, tol=1e-3, random_state=42)\nper_clf.fit(X, y)\n\ny_pred = per_clf.predict([[2, 0.5]])",
"_____no_output_____"
],
[
"y_pred",
"_____no_output_____"
],
[
"a = -per_clf.coef_[0][0] / per_clf.coef_[0][1]\nb = -per_clf.intercept_ / per_clf.coef_[0][1]\n\naxes = [0, 5, 0, 2]\n\nx0, x1 = np.meshgrid(\n np.linspace(axes[0], axes[1], 500).reshape(-1, 1),\n np.linspace(axes[2], axes[3], 200).reshape(-1, 1),\n )\nX_new = np.c_[x0.ravel(), x1.ravel()]\ny_predict = per_clf.predict(X_new)\nzz = y_predict.reshape(x0.shape)\n\nplt.figure(figsize=(10, 4))\nplt.plot(X[y==0, 0], X[y==0, 1], \"bs\", label=\"Not Iris-Setosa\")\nplt.plot(X[y==1, 0], X[y==1, 1], \"yo\", label=\"Iris-Setosa\")\n\nplt.plot([axes[0], axes[1]], [a * axes[0] + b, a * axes[1] + b], \"k-\", linewidth=3)\nfrom matplotlib.colors import ListedColormap\ncustom_cmap = ListedColormap(['#9898ff', '#fafab0'])\n\nplt.contourf(x0, x1, zz, cmap=custom_cmap)\nplt.xlabel(\"Petal length\", fontsize=14)\nplt.ylabel(\"Petal width\", fontsize=14)\nplt.legend(loc=\"lower right\", fontsize=14)\nplt.axis(axes)\n\nsave_fig(\"perceptron_iris_plot\")\nplt.show()",
"Saving figure perceptron_iris_plot\n"
]
],
[
[
"# Activation functions",
"_____no_output_____"
]
],
[
[
"def sigmoid(z):\n return 1 / (1 + np.exp(-z))\n\ndef relu(z):\n return np.maximum(0, z)\n\ndef derivative(f, z, eps=0.000001):\n return (f(z + eps) - f(z - eps))/(2 * eps)",
"_____no_output_____"
],
[
"z = np.linspace(-5, 5, 200)\n\nplt.figure(figsize=(11,4))\n\nplt.subplot(121)\nplt.plot(z, np.sign(z), \"r-\", linewidth=1, label=\"Step\")\nplt.plot(z, sigmoid(z), \"g--\", linewidth=2, label=\"Sigmoid\")\nplt.plot(z, np.tanh(z), \"b-\", linewidth=2, label=\"Tanh\")\nplt.plot(z, relu(z), \"m-.\", linewidth=2, label=\"ReLU\")\nplt.grid(True)\nplt.legend(loc=\"center right\", fontsize=14)\nplt.title(\"Activation functions\", fontsize=14)\nplt.axis([-5, 5, -1.2, 1.2])\n\nplt.subplot(122)\nplt.plot(z, derivative(np.sign, z), \"r-\", linewidth=1, label=\"Step\")\nplt.plot(0, 0, \"ro\", markersize=5)\nplt.plot(0, 0, \"rx\", markersize=10)\nplt.plot(z, derivative(sigmoid, z), \"g--\", linewidth=2, label=\"Sigmoid\")\nplt.plot(z, derivative(np.tanh, z), \"b-\", linewidth=2, label=\"Tanh\")\nplt.plot(z, derivative(relu, z), \"m-.\", linewidth=2, label=\"ReLU\")\nplt.grid(True)\n#plt.legend(loc=\"center right\", fontsize=14)\nplt.title(\"Derivatives\", fontsize=14)\nplt.axis([-5, 5, -0.2, 1.2])\n\nsave_fig(\"activation_functions_plot\")\nplt.show()",
"Saving figure activation_functions_plot\n"
],
[
"def heaviside(z):\n return (z >= 0).astype(z.dtype)\n\ndef mlp_xor(x1, x2, activation=heaviside):\n return activation(-activation(x1 + x2 - 1.5) + activation(x1 + x2 - 0.5) - 0.5)",
"_____no_output_____"
],
[
"x1s = np.linspace(-0.2, 1.2, 100)\nx2s = np.linspace(-0.2, 1.2, 100)\nx1, x2 = np.meshgrid(x1s, x2s)\n\nz1 = mlp_xor(x1, x2, activation=heaviside)\nz2 = mlp_xor(x1, x2, activation=sigmoid)\n\nplt.figure(figsize=(10,4))\n\nplt.subplot(121)\nplt.contourf(x1, x2, z1)\nplt.plot([0, 1], [0, 1], \"gs\", markersize=20)\nplt.plot([0, 1], [1, 0], \"y^\", markersize=20)\nplt.title(\"Activation function: heaviside\", fontsize=14)\nplt.grid(True)\n\nplt.subplot(122)\nplt.contourf(x1, x2, z2)\nplt.plot([0, 1], [0, 1], \"gs\", markersize=20)\nplt.plot([0, 1], [1, 0], \"y^\", markersize=20)\nplt.title(\"Activation function: sigmoid\", fontsize=14)\nplt.grid(True)",
"_____no_output_____"
]
],
[
[
"# Building an Image Classifier",
"_____no_output_____"
],
[
"First let's import TensorFlow and Keras.",
"_____no_output_____"
]
],
[
[
"import tensorflow as tf\nfrom tensorflow import keras",
"_____no_output_____"
],
[
"tf.__version__",
"_____no_output_____"
],
[
"keras.__version__",
"_____no_output_____"
]
],
[
[
"Let's start by loading the fashion MNIST dataset. Keras has a number of functions to load popular datasets in `keras.datasets`. The dataset is already split for you between a training set and a test set, but it can be useful to split the training set further to have a validation set:",
"_____no_output_____"
]
],
[
[
"fashion_mnist = keras.datasets.fashion_mnist\n(X_train_full, y_train_full), (X_test, y_test) = fashion_mnist.load_data()",
"_____no_output_____"
]
],
[
[
"The training set contains 60,000 grayscale images, each 28x28 pixels:",
"_____no_output_____"
]
],
[
[
"X_train_full.shape",
"_____no_output_____"
]
],
[
[
"Each pixel intensity is represented as a byte (0 to 255):",
"_____no_output_____"
]
],
[
[
"X_train_full.dtype",
"_____no_output_____"
]
],
[
[
"Let's split the full training set into a validation set and a (smaller) training set. We also scale the pixel intensities down to the 0-1 range and convert them to floats, by dividing by 255.",
"_____no_output_____"
]
],
[
[
"X_valid, X_train = X_train_full[:5000] / 255., X_train_full[5000:] / 255.\ny_valid, y_train = y_train_full[:5000], y_train_full[5000:]\nX_test = X_test / 255.",
"_____no_output_____"
]
],
[
[
"You can plot an image using Matplotlib's `imshow()` function, with a `'binary'`\n color map:",
"_____no_output_____"
]
],
[
[
"plt.imshow(X_train[0], cmap=\"binary\")\nplt.axis('off')\nplt.show()",
"_____no_output_____"
]
],
[
[
"The labels are the class IDs (represented as uint8), from 0 to 9:",
"_____no_output_____"
]
],
[
[
"y_train",
"_____no_output_____"
]
],
[
[
"Here are the corresponding class names:",
"_____no_output_____"
]
],
[
[
"class_names = [\"T-shirt/top\", \"Trouser\", \"Pullover\", \"Dress\", \"Coat\",\n \"Sandal\", \"Shirt\", \"Sneaker\", \"Bag\", \"Ankle boot\"]",
"_____no_output_____"
]
],
[
[
"So the first image in the training set is a coat:",
"_____no_output_____"
]
],
[
[
"class_names[y_train[0]]",
"_____no_output_____"
]
],
[
[
"The validation set contains 5,000 images, and the test set contains 10,000 images:",
"_____no_output_____"
]
],
[
[
"X_valid.shape",
"_____no_output_____"
],
[
"X_test.shape",
"_____no_output_____"
]
],
[
[
"Let's take a look at a sample of the images in the dataset:",
"_____no_output_____"
]
],
[
[
"n_rows = 4\nn_cols = 10\nplt.figure(figsize=(n_cols * 1.2, n_rows * 1.2))\nfor row in range(n_rows):\n for col in range(n_cols):\n index = n_cols * row + col\n plt.subplot(n_rows, n_cols, index + 1)\n plt.imshow(X_train[index], cmap=\"binary\", interpolation=\"nearest\")\n plt.axis('off')\n plt.title(class_names[y_train[index]], fontsize=12)\nplt.subplots_adjust(wspace=0.2, hspace=0.5)\nsave_fig('fashion_mnist_plot', tight_layout=False)\nplt.show()",
"Saving figure fashion_mnist_plot\n"
],
[
"model = keras.models.Sequential()\nmodel.add(keras.layers.Flatten(input_shape=[28, 28]))\nmodel.add(keras.layers.Dense(300, activation=\"relu\"))\nmodel.add(keras.layers.Dense(100, activation=\"relu\"))\nmodel.add(keras.layers.Dense(10, activation=\"softmax\"))",
"_____no_output_____"
],
[
"keras.backend.clear_session()\nnp.random.seed(42)\ntf.random.set_seed(42)",
"_____no_output_____"
],
[
"model = keras.models.Sequential([\n keras.layers.Flatten(input_shape=[28, 28]),\n keras.layers.Dense(300, activation=\"relu\"),\n keras.layers.Dense(100, activation=\"relu\"),\n keras.layers.Dense(10, activation=\"softmax\")\n])",
"_____no_output_____"
],
[
"model.layers",
"_____no_output_____"
],
[
"model.summary()",
"Model: \"sequential\"\n_________________________________________________________________\nLayer (type) Output Shape Param # \n=================================================================\nflatten (Flatten) (None, 784) 0 \n_________________________________________________________________\ndense (Dense) (None, 300) 235500 \n_________________________________________________________________\ndense_1 (Dense) (None, 100) 30100 \n_________________________________________________________________\ndense_2 (Dense) (None, 10) 1010 \n=================================================================\nTotal params: 266,610\nTrainable params: 266,610\nNon-trainable params: 0\n_________________________________________________________________\n"
],
[
"keras.utils.plot_model(model, \"my_mnist_model.png\", show_shapes=True)",
"_____no_output_____"
],
[
"hidden1 = model.layers[1]\nhidden1.name",
"_____no_output_____"
],
[
"model.get_layer(hidden1.name) is hidden1",
"_____no_output_____"
],
[
"weights, biases = hidden1.get_weights()",
"_____no_output_____"
],
[
"weights",
"_____no_output_____"
],
[
"weights.shape",
"_____no_output_____"
],
[
"biases",
"_____no_output_____"
],
[
"biases.shape",
"_____no_output_____"
],
[
"model.compile(loss=\"sparse_categorical_crossentropy\",\n optimizer=\"sgd\",\n metrics=[\"accuracy\"])",
"_____no_output_____"
]
],
[
[
"This is equivalent to:",
"_____no_output_____"
],
[
"```python\nmodel.compile(loss=keras.losses.sparse_categorical_crossentropy,\n optimizer=keras.optimizers.SGD(),\n metrics=[keras.metrics.sparse_categorical_accuracy])\n```",
"_____no_output_____"
]
],
[
[
"history = model.fit(X_train, y_train, epochs=30,\n validation_data=(X_valid, y_valid))",
"WARNING: Logging before flag parsing goes to stderr.\nW0607 15:00:56.087194 140735810999168 deprecation.py:323] From /Users/ageron/miniconda3/envs/tf2/lib/python3.6/site-packages/tensorflow/python/ops/math_grad.py:1251: add_dispatch_support.<locals>.wrapper (from tensorflow.python.ops.array_ops) is deprecated and will be removed in a future version.\nInstructions for updating:\nUse tf.where in 2.0, which has the same broadcast rule as np.where\n"
],
[
"history.params",
"_____no_output_____"
],
[
"print(history.epoch)",
"[0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29]\n"
],
[
"history.history.keys()",
"_____no_output_____"
],
[
"import pandas as pd\n\npd.DataFrame(history.history).plot(figsize=(8, 5))\nplt.grid(True)\nplt.gca().set_ylim(0, 1)\nsave_fig(\"keras_learning_curves_plot\")\nplt.show()",
"Saving figure keras_learning_curves_plot\n"
],
[
"model.evaluate(X_test, y_test)",
"10000/10000 [==============================] - 0s 28us/sample - loss: 0.3343 - accuracy: 0.8857\n"
],
[
"X_new = X_test[:3]\ny_proba = model.predict(X_new)\ny_proba.round(2)",
"_____no_output_____"
],
[
"y_pred = model.predict_classes(X_new)\ny_pred",
"_____no_output_____"
],
[
"np.array(class_names)[y_pred]",
"_____no_output_____"
],
[
"y_new = y_test[:3]\ny_new",
"_____no_output_____"
]
],
[
[
"# Regression MLP",
"_____no_output_____"
],
[
"Let's load, split and scale the California housing dataset (the original one, not the modified one as in chapter 2):",
"_____no_output_____"
]
],
[
[
"from sklearn.datasets import fetch_california_housing\nfrom sklearn.model_selection import train_test_split\nfrom sklearn.preprocessing import StandardScaler\n\nhousing = fetch_california_housing()\n\nX_train_full, X_test, y_train_full, y_test = train_test_split(housing.data, housing.target, random_state=42)\nX_train, X_valid, y_train, y_valid = train_test_split(X_train_full, y_train_full, random_state=42)\n\nscaler = StandardScaler()\nX_train = scaler.fit_transform(X_train)\nX_valid = scaler.transform(X_valid)\nX_test = scaler.transform(X_test)",
"_____no_output_____"
],
[
"np.random.seed(42)\ntf.random.set_seed(42)",
"_____no_output_____"
],
[
"model = keras.models.Sequential([\n keras.layers.Dense(30, activation=\"relu\", input_shape=X_train.shape[1:]),\n keras.layers.Dense(1)\n])\nmodel.compile(loss=\"mean_squared_error\", optimizer=keras.optimizers.SGD(lr=1e-3))\nhistory = model.fit(X_train, y_train, epochs=20, validation_data=(X_valid, y_valid))\nmse_test = model.evaluate(X_test, y_test)\nX_new = X_test[:3]\ny_pred = model.predict(X_new)",
"Train on 11610 samples, validate on 3870 samples\nEpoch 1/20\n11610/11610 [==============================] - 0s 39us/sample - loss: 1.6343 - val_loss: 0.9361\nEpoch 2/20\n11610/11610 [==============================] - 0s 31us/sample - loss: 0.7053 - val_loss: 0.6556\nEpoch 3/20\n11610/11610 [==============================] - 0s 32us/sample - loss: 0.6343 - val_loss: 0.6028\nEpoch 4/20\n11610/11610 [==============================] - 0s 31us/sample - loss: 0.5974 - val_loss: 0.5631\nEpoch 5/20\n11610/11610 [==============================] - 0s 32us/sample - loss: 0.5701 - val_loss: 0.5375\nEpoch 6/20\n11610/11610 [==============================] - 0s 32us/sample - loss: 0.5474 - val_loss: 0.5165\nEpoch 7/20\n11610/11610 [==============================] - 0s 32us/sample - loss: 0.5283 - val_loss: 0.5072\nEpoch 8/20\n11610/11610 [==============================] - 0s 32us/sample - loss: 0.5130 - val_loss: 0.4806\nEpoch 9/20\n11610/11610 [==============================] - 0s 32us/sample - loss: 0.4989 - val_loss: 0.4712\nEpoch 10/20\n11610/11610 [==============================] - 0s 32us/sample - loss: 0.4874 - val_loss: 0.4651\nEpoch 11/20\n11610/11610 [==============================] - 0s 32us/sample - loss: 0.4780 - val_loss: 0.4457\nEpoch 12/20\n11610/11610 [==============================] - 0s 31us/sample - loss: 0.4690 - val_loss: 0.4364\nEpoch 13/20\n11610/11610 [==============================] - 0s 31us/sample - loss: 0.4614 - val_loss: 0.4299\nEpoch 14/20\n11610/11610 [==============================] - 0s 32us/sample - loss: 0.4550 - val_loss: 0.4245\nEpoch 15/20\n11610/11610 [==============================] - 0s 31us/sample - loss: 0.4490 - val_loss: 0.4175\nEpoch 16/20\n11610/11610 [==============================] - 0s 35us/sample - loss: 0.4437 - val_loss: 0.4121\nEpoch 17/20\n11610/11610 [==============================] - 0s 35us/sample - loss: 0.4390 - val_loss: 0.4099\nEpoch 18/20\n11610/11610 [==============================] - 0s 33us/sample - loss: 0.4347 - val_loss: 0.4037\nEpoch 19/20\n11610/11610 [==============================] - 0s 32us/sample - loss: 0.4309 - val_loss: 0.3997\nEpoch 20/20\n11610/11610 [==============================] - 0s 33us/sample - loss: 0.4275 - val_loss: 0.3971\n5160/5160 [==============================] - 0s 18us/sample - loss: 0.4213\n"
],
[
"plt.plot(pd.DataFrame(history.history))\nplt.grid(True)\nplt.gca().set_ylim(0, 1)\nplt.show()",
"_____no_output_____"
],
[
"y_pred",
"_____no_output_____"
]
],
[
[
"# Functional API",
"_____no_output_____"
],
[
"Not all neural network models are simply sequential. Some may have complex topologies. Some may have multiple inputs and/or multiple outputs. For example, a Wide & Deep neural network (see [paper](https://ai.google/research/pubs/pub45413)) connects all or part of the inputs directly to the output layer.",
"_____no_output_____"
]
],
[
[
"np.random.seed(42)\ntf.random.set_seed(42)",
"_____no_output_____"
],
[
"input_ = keras.layers.Input(shape=X_train.shape[1:])\nhidden1 = keras.layers.Dense(30, activation=\"relu\")(input_)\nhidden2 = keras.layers.Dense(30, activation=\"relu\")(hidden1)\nconcat = keras.layers.concatenate([input_, hidden2])\noutput = keras.layers.Dense(1)(concat)\nmodel = keras.models.Model(inputs=[input_], outputs=[output])",
"_____no_output_____"
],
[
"model.summary()",
"Model: \"model_5\"\n__________________________________________________________________________________________________\nLayer (type) Output Shape Param # Connected to \n==================================================================================================\ninput_3 (InputLayer) [(None, 8)] 0 \n__________________________________________________________________________________________________\ndense_25 (Dense) (None, 30) 270 input_3[0][0] \n__________________________________________________________________________________________________\ndense_26 (Dense) (None, 30) 930 dense_25[0][0] \n__________________________________________________________________________________________________\nconcatenate_5 (Concatenate) (None, 38) 0 input_3[0][0] \n dense_26[0][0] \n__________________________________________________________________________________________________\ndense_27 (Dense) (None, 1) 39 concatenate_5[0][0] \n==================================================================================================\nTotal params: 1,239\nTrainable params: 1,239\nNon-trainable params: 0\n__________________________________________________________________________________________________\n"
],
[
"model.compile(loss=\"mean_squared_error\", optimizer=keras.optimizers.SGD(lr=1e-3))\nhistory = model.fit(X_train, y_train, epochs=20,\n validation_data=(X_valid, y_valid))\nmse_test = model.evaluate(X_test, y_test)\ny_pred = model.predict(X_new)",
"Train on 11610 samples, validate on 3870 samples\nEpoch 1/20\n11610/11610 [==============================] - 0s 43us/sample - loss: 1.2544 - val_loss: 2.9415\nEpoch 2/20\n11610/11610 [==============================] - 0s 32us/sample - loss: 0.6531 - val_loss: 0.8754\nEpoch 3/20\n11610/11610 [==============================] - 0s 34us/sample - loss: 0.5873 - val_loss: 0.5529\nEpoch 4/20\n11610/11610 [==============================] - 0s 32us/sample - loss: 0.5574 - val_loss: 0.5205\nEpoch 5/20\n11610/11610 [==============================] - 0s 33us/sample - loss: 0.5343 - val_loss: 0.5399\nEpoch 6/20\n11610/11610 [==============================] - 0s 32us/sample - loss: 0.5160 - val_loss: 0.4793\nEpoch 7/20\n11610/11610 [==============================] - 0s 32us/sample - loss: 0.5004 - val_loss: 0.4655\nEpoch 8/20\n11610/11610 [==============================] - 0s 33us/sample - loss: 0.4873 - val_loss: 0.4820\nEpoch 9/20\n11610/11610 [==============================] - 0s 32us/sample - loss: 0.4761 - val_loss: 0.4421\nEpoch 10/20\n11610/11610 [==============================] - 0s 32us/sample - loss: 0.4658 - val_loss: 0.4321\nEpoch 11/20\n11610/11610 [==============================] - 0s 33us/sample - loss: 0.4577 - val_loss: 0.4526\nEpoch 12/20\n11610/11610 [==============================] - 0s 32us/sample - loss: 0.4499 - val_loss: 0.4178\nEpoch 13/20\n11610/11610 [==============================] - 0s 32us/sample - loss: 0.4428 - val_loss: 0.4162\nEpoch 14/20\n11610/11610 [==============================] - 0s 32us/sample - loss: 0.4367 - val_loss: 0.4185\nEpoch 15/20\n11610/11610 [==============================] - 0s 32us/sample - loss: 0.4311 - val_loss: 0.3991\nEpoch 16/20\n11610/11610 [==============================] - 0s 32us/sample - loss: 0.4259 - val_loss: 0.3998\nEpoch 17/20\n11610/11610 [==============================] - 0s 34us/sample - loss: 0.4210 - val_loss: 0.3896\nEpoch 18/20\n11610/11610 [==============================] - 0s 33us/sample - loss: 0.4165 - val_loss: 0.3864\nEpoch 19/20\n11610/11610 [==============================] - 0s 33us/sample - loss: 0.4123 - val_loss: 0.3819\nEpoch 20/20\n11610/11610 [==============================] - 0s 35us/sample - loss: 0.4087 - val_loss: 0.3878\n5160/5160 [==============================] - 0s 18us/sample - loss: 0.4029\n"
]
],
[
[
"What if you want to send different subsets of input features through the wide or deep paths? We will send 5 features (features 0 to 4), and 6 through the deep path (features 2 to 7). Note that 3 features will go through both (features 2, 3 and 4).",
"_____no_output_____"
]
],
[
[
"np.random.seed(42)\ntf.random.set_seed(42)",
"_____no_output_____"
],
[
"input_A = keras.layers.Input(shape=[5], name=\"wide_input\")\ninput_B = keras.layers.Input(shape=[6], name=\"deep_input\")\nhidden1 = keras.layers.Dense(30, activation=\"relu\")(input_B)\nhidden2 = keras.layers.Dense(30, activation=\"relu\")(hidden1)\nconcat = keras.layers.concatenate([input_A, hidden2])\noutput = keras.layers.Dense(1, name=\"output\")(concat)\nmodel = keras.models.Model(inputs=[input_A, input_B], outputs=[output])",
"_____no_output_____"
],
[
"model.compile(loss=\"mse\", optimizer=keras.optimizers.SGD(lr=1e-3))\n\nX_train_A, X_train_B = X_train[:, :5], X_train[:, 2:]\nX_valid_A, X_valid_B = X_valid[:, :5], X_valid[:, 2:]\nX_test_A, X_test_B = X_test[:, :5], X_test[:, 2:]\nX_new_A, X_new_B = X_test_A[:3], X_test_B[:3]\n\nhistory = model.fit((X_train_A, X_train_B), y_train, epochs=20,\n validation_data=((X_valid_A, X_valid_B), y_valid))\nmse_test = model.evaluate((X_test_A, X_test_B), y_test)\ny_pred = model.predict((X_new_A, X_new_B))",
"Train on 11610 samples, validate on 3870 samples\nEpoch 1/20\n11610/11610 [==============================] - 1s 45us/sample - loss: 1.8070 - val_loss: 0.8422\nEpoch 2/20\n11610/11610 [==============================] - 0s 34us/sample - loss: 0.6752 - val_loss: 0.6426\nEpoch 3/20\n11610/11610 [==============================] - 0s 34us/sample - loss: 0.5969 - val_loss: 0.5829\nEpoch 4/20\n11610/11610 [==============================] - 0s 34us/sample - loss: 0.5580 - val_loss: 0.5259\nEpoch 5/20\n11610/11610 [==============================] - 0s 34us/sample - loss: 0.5317 - val_loss: 0.5005\nEpoch 6/20\n11610/11610 [==============================] - 0s 33us/sample - loss: 0.5125 - val_loss: 0.4792\nEpoch 7/20\n11610/11610 [==============================] - 0s 34us/sample - loss: 0.4963 - val_loss: 0.4786\nEpoch 8/20\n11610/11610 [==============================] - 0s 34us/sample - loss: 0.4840 - val_loss: 0.4562\nEpoch 9/20\n11610/11610 [==============================] - 0s 34us/sample - loss: 0.4733 - val_loss: 0.4407\nEpoch 10/20\n11610/11610 [==============================] - 0s 34us/sample - loss: 0.4640 - val_loss: 0.4320\nEpoch 11/20\n11610/11610 [==============================] - 0s 37us/sample - loss: 0.4576 - val_loss: 0.4286\nEpoch 12/20\n11610/11610 [==============================] - 0s 33us/sample - loss: 0.4516 - val_loss: 0.4226\nEpoch 13/20\n11610/11610 [==============================] - 0s 33us/sample - loss: 0.4463 - val_loss: 0.4263\nEpoch 14/20\n11610/11610 [==============================] - 0s 33us/sample - loss: 0.4428 - val_loss: 0.4102\nEpoch 15/20\n11610/11610 [==============================] - 0s 33us/sample - loss: 0.4392 - val_loss: 0.4039\nEpoch 16/20\n11610/11610 [==============================] - 0s 33us/sample - loss: 0.4352 - val_loss: 0.4021\nEpoch 17/20\n11610/11610 [==============================] - 0s 34us/sample - loss: 0.4329 - val_loss: 0.4035\nEpoch 18/20\n11610/11610 [==============================] - 0s 34us/sample - loss: 0.4307 - val_loss: 0.3951\nEpoch 19/20\n11610/11610 [==============================] - 0s 33us/sample - loss: 0.4275 - val_loss: 0.3932\nEpoch 20/20\n11610/11610 [==============================] - 0s 34us/sample - loss: 0.4264 - val_loss: 0.4000\n5160/5160 [==============================] - 0s 19us/sample - loss: 0.4204\n"
]
],
[
[
"Adding an auxiliary output for regularization:",
"_____no_output_____"
]
],
[
[
"np.random.seed(42)\ntf.random.set_seed(42)",
"_____no_output_____"
],
[
"input_A = keras.layers.Input(shape=[5], name=\"wide_input\")\ninput_B = keras.layers.Input(shape=[6], name=\"deep_input\")\nhidden1 = keras.layers.Dense(30, activation=\"relu\")(input_B)\nhidden2 = keras.layers.Dense(30, activation=\"relu\")(hidden1)\nconcat = keras.layers.concatenate([input_A, hidden2])\noutput = keras.layers.Dense(1, name=\"main_output\")(concat)\naux_output = keras.layers.Dense(1, name=\"aux_output\")(hidden2)\nmodel = keras.models.Model(inputs=[input_A, input_B],\n outputs=[output, aux_output])",
"_____no_output_____"
],
[
"model.compile(loss=[\"mse\", \"mse\"], loss_weights=[0.9, 0.1], optimizer=keras.optimizers.SGD(lr=1e-3))",
"_____no_output_____"
],
[
"history = model.fit([X_train_A, X_train_B], [y_train, y_train], epochs=20,\n validation_data=([X_valid_A, X_valid_B], [y_valid, y_valid]))",
"Train on 11610 samples, validate on 3870 samples\nEpoch 1/20\n11610/11610 [==============================] - 1s 57us/sample - loss: 2.1325 - main_output_loss: 1.9139 - aux_output_loss: 4.0950 - val_loss: 1.6650 - val_main_output_loss: 0.8892 - val_aux_output_loss: 8.6440\nEpoch 2/20\n11610/11610 [==============================] - 0s 38us/sample - loss: 0.8870 - main_output_loss: 0.6930 - aux_output_loss: 2.6303 - val_loss: 1.4991 - val_main_output_loss: 0.6627 - val_aux_output_loss: 9.0235\nEpoch 3/20\n11610/11610 [==============================] - 0s 38us/sample - loss: 0.7421 - main_output_loss: 0.6079 - aux_output_loss: 1.9515 - val_loss: 1.4340 - val_main_output_loss: 0.5748 - val_aux_output_loss: 9.1628\nEpoch 4/20\n11610/11610 [==============================] - 0s 38us/sample - loss: 0.6766 - main_output_loss: 0.5683 - aux_output_loss: 1.6515 - val_loss: 1.3260 - val_main_output_loss: 0.5579 - val_aux_output_loss: 8.2343\nEpoch 5/20\n11610/11610 [==============================] - 0s 38us/sample - loss: 0.6366 - main_output_loss: 0.5422 - aux_output_loss: 1.4873 - val_loss: 1.2081 - val_main_output_loss: 0.5184 - val_aux_output_loss: 7.4115\nEpoch 6/20\n11610/11610 [==============================] - 0s 38us/sample - loss: 0.6082 - main_output_loss: 0.5209 - aux_output_loss: 1.3933 - val_loss: 1.0861 - val_main_output_loss: 0.5113 - val_aux_output_loss: 6.2558\nEpoch 7/20\n11610/11610 [==============================] - 0s 38us/sample - loss: 0.5849 - main_output_loss: 0.5034 - aux_output_loss: 1.3185 - val_loss: 0.9914 - val_main_output_loss: 0.5227 - val_aux_output_loss: 5.2070\nEpoch 8/20\n11610/11610 [==============================] - 0s 39us/sample - loss: 0.5664 - main_output_loss: 0.4897 - aux_output_loss: 1.2556 - val_loss: 0.8656 - val_main_output_loss: 0.4650 - val_aux_output_loss: 4.4682\nEpoch 9/20\n11610/11610 [==============================] - 0s 38us/sample - loss: 0.5506 - main_output_loss: 0.4770 - aux_output_loss: 1.2125 - val_loss: 0.7778 - val_main_output_loss: 0.4563 - val_aux_output_loss: 3.6704\nEpoch 10/20\n11610/11610 [==============================] - 0s 41us/sample - loss: 0.5371 - main_output_loss: 0.4668 - aux_output_loss: 1.1711 - val_loss: 0.7089 - val_main_output_loss: 0.4513 - val_aux_output_loss: 3.0250\nEpoch 11/20\n11610/11610 [==============================] - 1s 46us/sample - loss: 0.5268 - main_output_loss: 0.4593 - aux_output_loss: 1.1340 - val_loss: 0.6476 - val_main_output_loss: 0.4292 - val_aux_output_loss: 2.6140\nEpoch 12/20\n11610/11610 [==============================] - 0s 38us/sample - loss: 0.5176 - main_output_loss: 0.4524 - aux_output_loss: 1.1035 - val_loss: 0.6025 - val_main_output_loss: 0.4200 - val_aux_output_loss: 2.2451\nEpoch 13/20\n11610/11610 [==============================] - 0s 38us/sample - loss: 0.5095 - main_output_loss: 0.4463 - aux_output_loss: 1.0766 - val_loss: 0.5692 - val_main_output_loss: 0.4158 - val_aux_output_loss: 1.9492\nEpoch 14/20\n11610/11610 [==============================] - 0s 38us/sample - loss: 0.5031 - main_output_loss: 0.4420 - aux_output_loss: 1.0529 - val_loss: 0.5362 - val_main_output_loss: 0.4092 - val_aux_output_loss: 1.6791\nEpoch 15/20\n11610/11610 [==============================] - 0s 38us/sample - loss: 0.4972 - main_output_loss: 0.4381 - aux_output_loss: 1.0284 - val_loss: 0.5120 - val_main_output_loss: 0.4053 - val_aux_output_loss: 1.4733\nEpoch 16/20\n11610/11610 [==============================] - 0s 39us/sample - loss: 0.4915 - main_output_loss: 0.4341 - aux_output_loss: 1.0078 - val_loss: 0.4935 - val_main_output_loss: 0.4013 - val_aux_output_loss: 1.3245\nEpoch 17/20\n11610/11610 [==============================] - 0s 38us/sample - loss: 0.4871 - main_output_loss: 0.4314 - aux_output_loss: 0.9878 - val_loss: 0.4797 - val_main_output_loss: 0.3979 - val_aux_output_loss: 1.2147\nEpoch 18/20\n11610/11610 [==============================] - 0s 38us/sample - loss: 0.4830 - main_output_loss: 0.4290 - aux_output_loss: 0.9694 - val_loss: 0.4669 - val_main_output_loss: 0.3954 - val_aux_output_loss: 1.1119\nEpoch 19/20\n11610/11610 [==============================] - 0s 38us/sample - loss: 0.4787 - main_output_loss: 0.4262 - aux_output_loss: 0.9513 - val_loss: 0.4577 - val_main_output_loss: 0.3930 - val_aux_output_loss: 1.0410\nEpoch 20/20\n11610/11610 [==============================] - 0s 38us/sample - loss: 0.4758 - main_output_loss: 0.4249 - aux_output_loss: 0.9342 - val_loss: 0.4558 - val_main_output_loss: 0.3952 - val_aux_output_loss: 1.0002\n"
],
[
"total_loss, main_loss, aux_loss = model.evaluate(\n [X_test_A, X_test_B], [y_test, y_test])\ny_pred_main, y_pred_aux = model.predict([X_new_A, X_new_B])",
"5160/5160 [==============================] - 0s 26us/sample - loss: 0.4661 - main_output_loss: 0.4167 - aux_output_loss: 0.9138\n"
]
],
[
[
"# The subclassing API",
"_____no_output_____"
]
],
[
[
"class WideAndDeepModel(keras.models.Model):\n def __init__(self, units=30, activation=\"relu\", **kwargs):\n super().__init__(**kwargs)\n self.hidden1 = keras.layers.Dense(units, activation=activation)\n self.hidden2 = keras.layers.Dense(units, activation=activation)\n self.main_output = keras.layers.Dense(1)\n self.aux_output = keras.layers.Dense(1)\n \n def call(self, inputs):\n input_A, input_B = inputs\n hidden1 = self.hidden1(input_B)\n hidden2 = self.hidden2(hidden1)\n concat = keras.layers.concatenate([input_A, hidden2])\n main_output = self.main_output(concat)\n aux_output = self.aux_output(hidden2)\n return main_output, aux_output\n\nmodel = WideAndDeepModel(30, activation=\"relu\")",
"_____no_output_____"
],
[
"model.compile(loss=\"mse\", loss_weights=[0.9, 0.1], optimizer=keras.optimizers.SGD(lr=1e-3))\nhistory = model.fit((X_train_A, X_train_B), (y_train, y_train), epochs=10,\n validation_data=((X_valid_A, X_valid_B), (y_valid, y_valid)))\ntotal_loss, main_loss, aux_loss = model.evaluate((X_test_A, X_test_B), (y_test, y_test))\ny_pred_main, y_pred_aux = model.predict((X_new_A, X_new_B))",
"Train on 11610 samples, validate on 3870 samples\nEpoch 1/10\n11610/11610 [==============================] - 1s 61us/sample - loss: 2.2914 - output_1_loss: 2.1790 - output_2_loss: 3.2954 - val_loss: 2.7823 - val_output_1_loss: 2.0246 - val_output_2_loss: 9.5935\nEpoch 2/10\n11610/11610 [==============================] - 0s 40us/sample - loss: 0.9834 - output_1_loss: 0.8678 - output_2_loss: 2.0207 - val_loss: 1.6646 - val_output_1_loss: 0.7720 - val_output_2_loss: 9.6933\nEpoch 3/10\n11610/11610 [==============================] - 0s 36us/sample - loss: 0.8264 - output_1_loss: 0.7295 - output_2_loss: 1.6967 - val_loss: 1.4471 - val_output_1_loss: 0.6947 - val_output_2_loss: 8.2170\nEpoch 4/10\n11610/11610 [==============================] - 0s 40us/sample - loss: 0.7616 - output_1_loss: 0.6750 - output_2_loss: 1.5418 - val_loss: 1.2658 - val_output_1_loss: 0.6379 - val_output_2_loss: 6.9130\nEpoch 5/10\n11610/11610 [==============================] - 0s 40us/sample - loss: 0.7203 - output_1_loss: 0.6400 - output_2_loss: 1.4425 - val_loss: 1.1078 - val_output_1_loss: 0.6015 - val_output_2_loss: 5.6652\nEpoch 6/10\n11610/11610 [==============================] - 0s 36us/sample - loss: 0.6889 - output_1_loss: 0.6124 - output_2_loss: 1.3768 - val_loss: 0.9780 - val_output_1_loss: 0.5691 - val_output_2_loss: 4.8158\nEpoch 7/10\n11610/11610 [==============================] - 0s 37us/sample - loss: 0.6620 - output_1_loss: 0.5885 - output_2_loss: 1.3240 - val_loss: 0.8896 - val_output_1_loss: 0.5852 - val_output_2_loss: 3.6278\nEpoch 8/10\n11610/11610 [==============================] - 0s 37us/sample - loss: 0.6402 - output_1_loss: 0.5697 - output_2_loss: 1.2749 - val_loss: 0.7913 - val_output_1_loss: 0.5294 - val_output_2_loss: 3.1492\nEpoch 9/10\n11610/11610 [==============================] - 0s 39us/sample - loss: 0.6200 - output_1_loss: 0.5513 - output_2_loss: 1.2377 - val_loss: 0.7284 - val_output_1_loss: 0.5119 - val_output_2_loss: 2.6768\nEpoch 10/10\n11610/11610 [==============================] - 0s 38us/sample - loss: 0.6020 - output_1_loss: 0.5353 - output_2_loss: 1.2002 - val_loss: 0.6832 - val_output_1_loss: 0.4927 - val_output_2_loss: 2.3969\n5160/5160 [==============================] - 0s 23us/sample - loss: 0.5847 - output_1_loss: 0.5204 - output_2_loss: 1.1756\n"
],
[
"model = WideAndDeepModel(30, activation=\"relu\")",
"_____no_output_____"
]
],
[
[
"# Saving and Restoring",
"_____no_output_____"
]
],
[
[
"np.random.seed(42)\ntf.random.set_seed(42)",
"_____no_output_____"
],
[
"model = keras.models.Sequential([\n keras.layers.Dense(30, activation=\"relu\", input_shape=[8]),\n keras.layers.Dense(30, activation=\"relu\"),\n keras.layers.Dense(1)\n]) ",
"_____no_output_____"
],
[
"model.compile(loss=\"mse\", optimizer=keras.optimizers.SGD(lr=1e-3))\nhistory = model.fit(X_train, y_train, epochs=10, validation_data=(X_valid, y_valid))\nmse_test = model.evaluate(X_test, y_test)",
"Train on 11610 samples, validate on 3870 samples\nEpoch 1/10\n11610/11610 [==============================] - 0s 41us/sample - loss: 1.8807 - val_loss: 0.7701\nEpoch 2/10\n11610/11610 [==============================] - 0s 31us/sample - loss: 0.6570 - val_loss: 0.7268\nEpoch 3/10\n11610/11610 [==============================] - 0s 31us/sample - loss: 0.5932 - val_loss: 0.6569\nEpoch 4/10\n11610/11610 [==============================] - 0s 31us/sample - loss: 0.5547 - val_loss: 0.5244\nEpoch 5/10\n11610/11610 [==============================] - 0s 31us/sample - loss: 0.5269 - val_loss: 0.5068\nEpoch 6/10\n11610/11610 [==============================] - 0s 30us/sample - loss: 0.5035 - val_loss: 0.5028\nEpoch 7/10\n11610/11610 [==============================] - 0s 31us/sample - loss: 0.4852 - val_loss: 0.4924\nEpoch 8/10\n11610/11610 [==============================] - 0s 31us/sample - loss: 0.4709 - val_loss: 0.4534\nEpoch 9/10\n11610/11610 [==============================] - 0s 31us/sample - loss: 0.4578 - val_loss: 0.4403\nEpoch 10/10\n11610/11610 [==============================] - 0s 31us/sample - loss: 0.4473 - val_loss: 0.4496\n5160/5160 [==============================] - 0s 17us/sample - loss: 0.4379\n"
],
[
"model.save(\"my_keras_model.h5\")",
"_____no_output_____"
],
[
"model = keras.models.load_model(\"my_keras_model.h5\")",
"_____no_output_____"
],
[
"model.predict(X_new)",
"_____no_output_____"
],
[
"model.save_weights(\"my_keras_weights.ckpt\")",
"_____no_output_____"
],
[
"model.load_weights(\"my_keras_weights.ckpt\")",
"_____no_output_____"
]
],
[
[
"# Using Callbacks during Training",
"_____no_output_____"
]
],
[
[
"keras.backend.clear_session()\nnp.random.seed(42)\ntf.random.set_seed(42)",
"_____no_output_____"
],
[
"model = keras.models.Sequential([\n keras.layers.Dense(30, activation=\"relu\", input_shape=[8]),\n keras.layers.Dense(30, activation=\"relu\"),\n keras.layers.Dense(1)\n]) ",
"_____no_output_____"
],
[
"model.compile(loss=\"mse\", optimizer=keras.optimizers.SGD(lr=1e-3))\ncheckpoint_cb = keras.callbacks.ModelCheckpoint(\"my_keras_model.h5\", save_best_only=True)\nhistory = model.fit(X_train, y_train, epochs=10,\n validation_data=(X_valid, y_valid),\n callbacks=[checkpoint_cb])\nmodel = keras.models.load_model(\"my_keras_model.h5\") # rollback to best model\nmse_test = model.evaluate(X_test, y_test)",
"Train on 11610 samples, validate on 3870 samples\nEpoch 1/10\n11610/11610 [==============================] - 1s 49us/sample - loss: 1.8807 - val_loss: 0.7701\nEpoch 2/10\n11610/11610 [==============================] - 0s 34us/sample - loss: 0.6570 - val_loss: 0.7268\nEpoch 3/10\n11610/11610 [==============================] - 0s 33us/sample - loss: 0.5932 - val_loss: 0.6569\nEpoch 4/10\n11610/11610 [==============================] - 0s 33us/sample - loss: 0.5547 - val_loss: 0.5244\nEpoch 5/10\n11610/11610 [==============================] - 0s 34us/sample - loss: 0.5269 - val_loss: 0.5068\nEpoch 6/10\n11610/11610 [==============================] - 0s 33us/sample - loss: 0.5035 - val_loss: 0.5028\nEpoch 7/10\n11610/11610 [==============================] - 0s 32us/sample - loss: 0.4852 - val_loss: 0.4924\nEpoch 8/10\n11610/11610 [==============================] - 0s 32us/sample - loss: 0.4709 - val_loss: 0.4534\nEpoch 9/10\n11610/11610 [==============================] - 0s 32us/sample - loss: 0.4578 - val_loss: 0.4403\nEpoch 10/10\n11610/11610 [==============================] - 0s 32us/sample - loss: 0.4473 - val_loss: 0.4496\n5160/5160 [==============================] - 0s 20us/sample - loss: 0.4468\n"
],
[
"model.compile(loss=\"mse\", optimizer=keras.optimizers.SGD(lr=1e-3))\nearly_stopping_cb = keras.callbacks.EarlyStopping(patience=10,\n restore_best_weights=True)\nhistory = model.fit(X_train, y_train, epochs=100,\n validation_data=(X_valid, y_valid),\n callbacks=[checkpoint_cb, early_stopping_cb])\nmse_test = model.evaluate(X_test, y_test)",
"Train on 11610 samples, validate on 3870 samples\nEpoch 1/100\n11610/11610 [==============================] - 0s 43us/sample - loss: 0.4483 - val_loss: 0.4280\nEpoch 2/100\n11610/11610 [==============================] - 0s 33us/sample - loss: 0.4393 - val_loss: 0.4106\nEpoch 3/100\n11610/11610 [==============================] - 0s 33us/sample - loss: 0.4319 - val_loss: 0.4078\nEpoch 4/100\n11610/11610 [==============================] - 0s 33us/sample - loss: 0.4262 - val_loss: 0.4037\nEpoch 5/100\n11610/11610 [==============================] - 0s 32us/sample - loss: 0.4205 - val_loss: 0.3956\nEpoch 6/100\n11610/11610 [==============================] - 0s 36us/sample - loss: 0.4160 - val_loss: 0.3894\nEpoch 7/100\n11610/11610 [==============================] - 0s 35us/sample - loss: 0.4116 - val_loss: 0.3891\nEpoch 8/100\n11610/11610 [==============================] - 0s 34us/sample - loss: 0.4077 - val_loss: 0.3817\nEpoch 9/100\n11610/11610 [==============================] - 0s 36us/sample - loss: 0.4041 - val_loss: 0.3783\nEpoch 10/100\n11610/11610 [==============================] - 0s 33us/sample - loss: 0.4010 - val_loss: 0.3757\nEpoch 11/100\n11610/11610 [==============================] - 0s 35us/sample - loss: 0.3980 - val_loss: 0.3731\nEpoch 12/100\n11610/11610 [==============================] - 0s 32us/sample - loss: 0.3951 - val_loss: 0.3743\nEpoch 13/100\n11610/11610 [==============================] - 0s 33us/sample - loss: 0.3924 - val_loss: 0.3686\nEpoch 14/100\n11610/11610 [==============================] - 0s 33us/sample - loss: 0.3898 - val_loss: 0.3678\nEpoch 15/100\n11610/11610 [==============================] - 0s 33us/sample - loss: 0.3873 - val_loss: 0.3626\nEpoch 16/100\n11610/11610 [==============================] - 0s 33us/sample - loss: 0.3850 - val_loss: 0.3612\nEpoch 17/100\n11610/11610 [==============================] - 0s 34us/sample - loss: 0.3831 - val_loss: 0.3583\n<<48 more lines>>\nEpoch 42/100\n11610/11610 [==============================] - 0s 34us/sample - loss: 0.3513 - val_loss: 0.3832\nEpoch 43/100\n11610/11610 [==============================] - 0s 31us/sample - loss: 0.3509 - val_loss: 0.3367\nEpoch 44/100\n11610/11610 [==============================] - 0s 32us/sample - loss: 0.3499 - val_loss: 0.3488\nEpoch 45/100\n11610/11610 [==============================] - 0s 35us/sample - loss: 0.3493 - val_loss: 0.3410\nEpoch 46/100\n11610/11610 [==============================] - 0s 32us/sample - loss: 0.3486 - val_loss: 0.3290\nEpoch 47/100\n11610/11610 [==============================] - 0s 33us/sample - loss: 0.3480 - val_loss: 0.3921\nEpoch 48/100\n11610/11610 [==============================] - 0s 36us/sample - loss: 0.3474 - val_loss: 0.3278\nEpoch 49/100\n11610/11610 [==============================] - 0s 34us/sample - loss: 0.3467 - val_loss: 0.3841\nEpoch 50/100\n11610/11610 [==============================] - 0s 32us/sample - loss: 0.3464 - val_loss: 0.3370\nEpoch 51/100\n11610/11610 [==============================] - 0s 33us/sample - loss: 0.3458 - val_loss: 0.3441\nEpoch 52/100\n11610/11610 [==============================] - 0s 31us/sample - loss: 0.3451 - val_loss: 0.3427\nEpoch 53/100\n11610/11610 [==============================] - 0s 32us/sample - loss: 0.3445 - val_loss: 0.3459\nEpoch 54/100\n11610/11610 [==============================] - 0s 31us/sample - loss: 0.3441 - val_loss: 0.3380\nEpoch 55/100\n11610/11610 [==============================] - 0s 32us/sample - loss: 0.3436 - val_loss: 0.3432\nEpoch 56/100\n11610/11610 [==============================] - 0s 35us/sample - loss: 0.3432 - val_loss: 0.3415\nEpoch 57/100\n11610/11610 [==============================] - 0s 31us/sample - loss: 0.3426 - val_loss: 0.3346\nEpoch 58/100\n11610/11610 [==============================] - 0s 32us/sample - loss: 0.3421 - val_loss: 0.3486\n5160/5160 [==============================] - 0s 17us/sample - loss: 0.3466\n"
],
[
"class PrintValTrainRatioCallback(keras.callbacks.Callback):\n def on_epoch_end(self, epoch, logs):\n print(\"\\nval/train: {:.2f}\".format(logs[\"val_loss\"] / logs[\"loss\"]))",
"_____no_output_____"
],
[
"val_train_ratio_cb = PrintValTrainRatioCallback()\nhistory = model.fit(X_train, y_train, epochs=1,\n validation_data=(X_valid, y_valid),\n callbacks=[val_train_ratio_cb])",
"Train on 11610 samples, validate on 3870 samples\n10144/11610 [=========================>....] - ETA: 0s - loss: 0.3427\nval/train: 0.98\n11610/11610 [==============================] - 0s 32us/sample - loss: 0.3467 - val_loss: 0.3384\n"
]
],
[
[
"# TensorBoard",
"_____no_output_____"
]
],
[
[
"root_logdir = os.path.join(os.curdir, \"my_logs\")",
"_____no_output_____"
],
[
"def get_run_logdir():\n import time\n run_id = time.strftime(\"run_%Y_%m_%d-%H_%M_%S\")\n return os.path.join(root_logdir, run_id)\n\nrun_logdir = get_run_logdir()\nrun_logdir",
"_____no_output_____"
],
[
"keras.backend.clear_session()\nnp.random.seed(42)\ntf.random.set_seed(42)",
"_____no_output_____"
],
[
"model = keras.models.Sequential([\n keras.layers.Dense(30, activation=\"relu\", input_shape=[8]),\n keras.layers.Dense(30, activation=\"relu\"),\n keras.layers.Dense(1)\n]) \nmodel.compile(loss=\"mse\", optimizer=keras.optimizers.SGD(lr=1e-3))",
"_____no_output_____"
],
[
"tensorboard_cb = keras.callbacks.TensorBoard(run_logdir)\nhistory = model.fit(X_train, y_train, epochs=30,\n validation_data=(X_valid, y_valid),\n callbacks=[checkpoint_cb, tensorboard_cb])",
"Train on 11610 samples, validate on 3870 samples\nEpoch 1/30\n11610/11610 [==============================] - 1s 45us/sample - loss: 1.8807 - val_loss: 0.7701\nEpoch 2/30\n11610/11610 [==============================] - 0s 33us/sample - loss: 0.6570 - val_loss: 0.7268\nEpoch 3/30\n11610/11610 [==============================] - 0s 33us/sample - loss: 0.5932 - val_loss: 0.6569\nEpoch 4/30\n11610/11610 [==============================] - 0s 32us/sample - loss: 0.5547 - val_loss: 0.5244\nEpoch 5/30\n11610/11610 [==============================] - 0s 33us/sample - loss: 0.5269 - val_loss: 0.5068\nEpoch 6/30\n11610/11610 [==============================] - 0s 32us/sample - loss: 0.5035 - val_loss: 0.5028\nEpoch 7/30\n11610/11610 [==============================] - 0s 33us/sample - loss: 0.4852 - val_loss: 0.4924\nEpoch 8/30\n11610/11610 [==============================] - 0s 32us/sample - loss: 0.4709 - val_loss: 0.4534\nEpoch 9/30\n11610/11610 [==============================] - 0s 32us/sample - loss: 0.4578 - val_loss: 0.4403\nEpoch 10/30\n11610/11610 [==============================] - 0s 32us/sample - loss: 0.4473 - val_loss: 0.4496\nEpoch 11/30\n11610/11610 [==============================] - 0s 32us/sample - loss: 0.4399 - val_loss: 0.4261\nEpoch 12/30\n11610/11610 [==============================] - 0s 37us/sample - loss: 0.4320 - val_loss: 0.4039\nEpoch 13/30\n11610/11610 [==============================] - 0s 34us/sample - loss: 0.4258 - val_loss: 0.4002\nEpoch 14/30\n11610/11610 [==============================] - 0s 32us/sample - loss: 0.4207 - val_loss: 0.3980\nEpoch 15/30\n11610/11610 [==============================] - 0s 35us/sample - loss: 0.4158 - val_loss: 0.3915\nEpoch 16/30\n11610/11610 [==============================] - 0s 32us/sample - loss: 0.4118 - val_loss: 0.3851\nEpoch 17/30\n11610/11610 [==============================] - 0s 33us/sample - loss: 0.4077 - val_loss: 0.3845\nEpoch 18/30\n11610/11610 [==============================] - 0s 32us/sample - loss: 0.4042 - val_loss: 0.3784\nEpoch 19/30\n11610/11610 [==============================] - 0s 33us/sample - loss: 0.4008 - val_loss: 0.3749\nEpoch 20/30\n11610/11610 [==============================] - 0s 34us/sample - loss: 0.3979 - val_loss: 0.3728\nEpoch 21/30\n11610/11610 [==============================] - 0s 32us/sample - loss: 0.3952 - val_loss: 0.3700\nEpoch 22/30\n11610/11610 [==============================] - 0s 32us/sample - loss: 0.3924 - val_loss: 0.3739\nEpoch 23/30\n11610/11610 [==============================] - 0s 33us/sample - loss: 0.3899 - val_loss: 0.3665\nEpoch 24/30\n11610/11610 [==============================] - 0s 32us/sample - loss: 0.3874 - val_loss: 0.3659\nEpoch 25/30\n11610/11610 [==============================] - 0s 32us/sample - loss: 0.3850 - val_loss: 0.3605\nEpoch 26/30\n11610/11610 [==============================] - 0s 32us/sample - loss: 0.3828 - val_loss: 0.3594\nEpoch 27/30\n11610/11610 [==============================] - 0s 32us/sample - loss: 0.3809 - val_loss: 0.3562\nEpoch 28/30\n11610/11610 [==============================] - 0s 32us/sample - loss: 0.3786 - val_loss: 0.3586\nEpoch 29/30\n11610/11610 [==============================] - 0s 32us/sample - loss: 0.3765 - val_loss: 0.3689\nEpoch 30/30\n11610/11610 [==============================] - 0s 32us/sample - loss: 0.3749 - val_loss: 0.3510\n"
]
],
[
[
"To start the TensorBoard server, one option is to open a terminal, if needed activate the virtualenv where you installed TensorBoard, go to this notebook's directory, then type:\n\n```bash\n$ tensorboard --logdir=./my_logs --port=6006\n```\n\nYou can then open your web browser to [localhost:6006](http://localhost:6006) and use TensorBoard. Once you are done, press Ctrl-C in the terminal window, this will shutdown the TensorBoard server.\n\nAlternatively, you can load TensorBoard's Jupyter extension and run it like this:",
"_____no_output_____"
]
],
[
[
"%load_ext tensorboard\n%tensorboard --logdir=./my_logs --port=6006",
"_____no_output_____"
],
[
"run_logdir2 = get_run_logdir()\nrun_logdir2",
"_____no_output_____"
],
[
"keras.backend.clear_session()\nnp.random.seed(42)\ntf.random.set_seed(42)",
"_____no_output_____"
],
[
"model = keras.models.Sequential([\n keras.layers.Dense(30, activation=\"relu\", input_shape=[8]),\n keras.layers.Dense(30, activation=\"relu\"),\n keras.layers.Dense(1)\n]) \nmodel.compile(loss=\"mse\", optimizer=keras.optimizers.SGD(lr=0.05))",
"_____no_output_____"
],
[
"tensorboard_cb = keras.callbacks.TensorBoard(run_logdir2)\nhistory = model.fit(X_train, y_train, epochs=30,\n validation_data=(X_valid, y_valid),\n callbacks=[checkpoint_cb, tensorboard_cb])",
"Train on 11610 samples, validate on 3870 samples\nEpoch 1/30\n11610/11610 [==============================] - 1s 43us/sample - loss: 0.5259 - val_loss: 0.4834\nEpoch 2/30\n11610/11610 [==============================] - 0s 33us/sample - loss: 0.4069 - val_loss: 0.3912\nEpoch 3/30\n11610/11610 [==============================] - 0s 33us/sample - loss: 0.3744 - val_loss: 7.5517\nEpoch 4/30\n11610/11610 [==============================] - 0s 32us/sample - loss: 0.3656 - val_loss: 1.1554\nEpoch 5/30\n11610/11610 [==============================] - 0s 38us/sample - loss: 0.3524 - val_loss: 0.4531\nEpoch 6/30\n11610/11610 [==============================] - 0s 34us/sample - loss: 0.3455 - val_loss: 0.3197\nEpoch 7/30\n11610/11610 [==============================] - 0s 33us/sample - loss: 0.3355 - val_loss: 0.3217\nEpoch 8/30\n11610/11610 [==============================] - 0s 33us/sample - loss: 0.3326 - val_loss: 0.3118\nEpoch 9/30\n11610/11610 [==============================] - 0s 34us/sample - loss: 0.3219 - val_loss: 0.3636\nEpoch 10/30\n11610/11610 [==============================] - 0s 35us/sample - loss: 0.3210 - val_loss: 0.3265\nEpoch 11/30\n11610/11610 [==============================] - 0s 32us/sample - loss: 0.3148 - val_loss: 0.3178\nEpoch 12/30\n11610/11610 [==============================] - 0s 35us/sample - loss: 0.3121 - val_loss: 0.3077\nEpoch 13/30\n11610/11610 [==============================] - 0s 35us/sample - loss: 0.3098 - val_loss: 0.3166\nEpoch 14/30\n11610/11610 [==============================] - 0s 36us/sample - loss: 0.3054 - val_loss: 0.3673\nEpoch 15/30\n11610/11610 [==============================] - 0s 37us/sample - loss: 0.3110 - val_loss: 0.5225\nEpoch 16/30\n11610/11610 [==============================] - 0s 32us/sample - loss: 0.3086 - val_loss: 0.2886\nEpoch 17/30\n11610/11610 [==============================] - 0s 33us/sample - loss: 0.3044 - val_loss: 0.3345\nEpoch 18/30\n11610/11610 [==============================] - 0s 31us/sample - loss: 0.3019 - val_loss: 0.2997\nEpoch 19/30\n11610/11610 [==============================] - 0s 32us/sample - loss: 0.2979 - val_loss: 0.2767\nEpoch 20/30\n11610/11610 [==============================] - 0s 32us/sample - loss: 0.2977 - val_loss: 0.3261\nEpoch 21/30\n11610/11610 [==============================] - 0s 31us/sample - loss: 0.2967 - val_loss: 0.2839\nEpoch 22/30\n11610/11610 [==============================] - 0s 32us/sample - loss: 0.2948 - val_loss: 0.2765\nEpoch 23/30\n11610/11610 [==============================] - 0s 31us/sample - loss: 0.2918 - val_loss: 0.3468\nEpoch 24/30\n11610/11610 [==============================] - 0s 31us/sample - loss: 0.2928 - val_loss: 0.2745\nEpoch 25/30\n11610/11610 [==============================] - 0s 32us/sample - loss: 0.2894 - val_loss: 0.3026\nEpoch 26/30\n11610/11610 [==============================] - 0s 32us/sample - loss: 0.2905 - val_loss: 0.2874\nEpoch 27/30\n11610/11610 [==============================] - 0s 32us/sample - loss: 0.2860 - val_loss: 0.2851\nEpoch 28/30\n11610/11610 [==============================] - 0s 34us/sample - loss: 0.2877 - val_loss: 0.5629\nEpoch 29/30\n11610/11610 [==============================] - 0s 33us/sample - loss: 0.2861 - val_loss: 0.2808\nEpoch 30/30\n11610/11610 [==============================] - 0s 34us/sample - loss: 0.2841 - val_loss: 0.2780\n"
]
],
[
[
"Notice how TensorBoard now sees two runs, and you can compare the learning curves.",
"_____no_output_____"
],
[
"Check out the other available logging options:",
"_____no_output_____"
]
],
[
[
"help(keras.callbacks.TensorBoard.__init__)",
"Help on function __init__ in module tensorflow.python.keras.callbacks:\n\n__init__(self, log_dir='logs', histogram_freq=0, write_graph=True, write_images=False, update_freq='epoch', profile_batch=2, embeddings_freq=0, embeddings_metadata=None, **kwargs)\n Initialize self. See help(type(self)) for accurate signature.\n\n"
]
],
[
[
"# Hyperparameter Tuning",
"_____no_output_____"
]
],
[
[
"keras.backend.clear_session()\nnp.random.seed(42)\ntf.random.set_seed(42)",
"_____no_output_____"
],
[
"def build_model(n_hidden=1, n_neurons=30, learning_rate=3e-3, input_shape=[8]):\n model = keras.models.Sequential()\n model.add(keras.layers.InputLayer(input_shape=input_shape))\n for layer in range(n_hidden):\n model.add(keras.layers.Dense(n_neurons, activation=\"relu\"))\n model.add(keras.layers.Dense(1))\n optimizer = keras.optimizers.SGD(lr=learning_rate)\n model.compile(loss=\"mse\", optimizer=optimizer)\n return model",
"_____no_output_____"
],
[
"keras_reg = keras.wrappers.scikit_learn.KerasRegressor(build_model)",
"_____no_output_____"
],
[
"keras_reg.fit(X_train, y_train, epochs=100,\n validation_data=(X_valid, y_valid),\n callbacks=[keras.callbacks.EarlyStopping(patience=10)])",
"Train on 11610 samples, validate on 3870 samples\nEpoch 1/100\n11610/11610 [==============================] - 0s 41us/sample - loss: 1.0910 - val_loss: 21.1892\nEpoch 2/100\n11610/11610 [==============================] - 0s 35us/sample - loss: 0.7607 - val_loss: 4.8909\nEpoch 3/100\n11610/11610 [==============================] - 0s 31us/sample - loss: 0.5401 - val_loss: 0.5721\nEpoch 4/100\n11610/11610 [==============================] - 0s 31us/sample - loss: 0.4752 - val_loss: 0.4433\nEpoch 5/100\n11610/11610 [==============================] - 0s 34us/sample - loss: 0.4499 - val_loss: 0.4270\nEpoch 6/100\n11610/11610 [==============================] - 0s 31us/sample - loss: 0.4348 - val_loss: 0.4165\nEpoch 7/100\n11610/11610 [==============================] - 0s 36us/sample - loss: 0.4240 - val_loss: 0.4060\nEpoch 8/100\n11610/11610 [==============================] - 0s 33us/sample - loss: 0.4166 - val_loss: 0.4030\nEpoch 9/100\n11610/11610 [==============================] - 0s 34us/sample - loss: 0.4109 - val_loss: 0.4069\nEpoch 10/100\n11610/11610 [==============================] - 0s 34us/sample - loss: 0.4055 - val_loss: 0.4220\nEpoch 11/100\n11610/11610 [==============================] - 0s 30us/sample - loss: 0.4021 - val_loss: 0.4191\nEpoch 12/100\n11610/11610 [==============================] - 0s 33us/sample - loss: 0.3982 - val_loss: 0.3941\nEpoch 13/100\n11610/11610 [==============================] - 0s 32us/sample - loss: 0.3950 - val_loss: 0.4145\nEpoch 14/100\n11610/11610 [==============================] - 0s 34us/sample - loss: 0.3923 - val_loss: 0.4155\nEpoch 15/100\n11610/11610 [==============================] - 0s 30us/sample - loss: 0.3897 - val_loss: 0.3774\nEpoch 16/100\n11610/11610 [==============================] - 0s 33us/sample - loss: 0.3873 - val_loss: 0.3828\nEpoch 17/100\n11610/11610 [==============================] - 0s 35us/sample - loss: 0.3847 - val_loss: 0.4104\n<<49 more lines>>\n11610/11610 [==============================] - 0s 29us/sample - loss: 0.3553 - val_loss: 0.3677\nEpoch 43/100\n11610/11610 [==============================] - 0s 29us/sample - loss: 0.3553 - val_loss: 0.3358\nEpoch 44/100\n11610/11610 [==============================] - 0s 30us/sample - loss: 0.3544 - val_loss: 0.3434\nEpoch 45/100\n11610/11610 [==============================] - 0s 29us/sample - loss: 0.3539 - val_loss: 0.4161\nEpoch 46/100\n11610/11610 [==============================] - 0s 29us/sample - loss: 0.3540 - val_loss: 0.3337\nEpoch 47/100\n11610/11610 [==============================] - 0s 30us/sample - loss: 0.3526 - val_loss: 0.3837\nEpoch 48/100\n11610/11610 [==============================] - 1s 44us/sample - loss: 0.3526 - val_loss: 0.3353\nEpoch 49/100\n11610/11610 [==============================] - 0s 29us/sample - loss: 0.3519 - val_loss: 0.3319\nEpoch 50/100\n11610/11610 [==============================] - 0s 29us/sample - loss: 0.3510 - val_loss: 0.3782\nEpoch 51/100\n11610/11610 [==============================] - 0s 29us/sample - loss: 0.3512 - val_loss: 0.3412\nEpoch 52/100\n11610/11610 [==============================] - 0s 29us/sample - loss: 0.3502 - val_loss: 0.4134\nEpoch 53/100\n11610/11610 [==============================] - 0s 30us/sample - loss: 0.3501 - val_loss: 0.3335\nEpoch 54/100\n11610/11610 [==============================] - 0s 30us/sample - loss: 0.3490 - val_loss: 0.3411\nEpoch 55/100\n11610/11610 [==============================] - 0s 29us/sample - loss: 0.3485 - val_loss: 0.3700\nEpoch 56/100\n11610/11610 [==============================] - 0s 29us/sample - loss: 0.3483 - val_loss: 0.3793\nEpoch 57/100\n11610/11610 [==============================] - 0s 29us/sample - loss: 0.3479 - val_loss: 0.4734\nEpoch 58/100\n11610/11610 [==============================] - 0s 30us/sample - loss: 0.3480 - val_loss: 0.4231\nEpoch 59/100\n11610/11610 [==============================] - 0s 29us/sample - loss: 0.3470 - val_loss: 0.6592\n"
],
[
"mse_test = keras_reg.score(X_test, y_test)",
"5160/5160 [==============================] - 0s 15us/sample - loss: 0.3504\n"
],
[
"y_pred = keras_reg.predict(X_new)",
"_____no_output_____"
],
[
"np.random.seed(42)\ntf.random.set_seed(42)",
"_____no_output_____"
],
[
"from scipy.stats import reciprocal\nfrom sklearn.model_selection import RandomizedSearchCV\n\nparam_distribs = {\n \"n_hidden\": [0, 1, 2, 3],\n \"n_neurons\": np.arange(1, 100),\n \"learning_rate\": reciprocal(3e-4, 3e-2),\n}\n\nrnd_search_cv = RandomizedSearchCV(keras_reg, param_distribs, n_iter=10, cv=3, verbose=2)\nrnd_search_cv.fit(X_train, y_train, epochs=100,\n validation_data=(X_valid, y_valid),\n callbacks=[keras.callbacks.EarlyStopping(patience=10)])",
"Fitting 3 folds for each of 10 candidates, totalling 30 fits\n[CV] learning_rate=0.001683454924600351, n_hidden=0, n_neurons=15 ....\nTrain on 7740 samples, validate on 3870 samples\nEpoch 1/100\n"
],
[
"rnd_search_cv.best_params_",
"_____no_output_____"
],
[
"rnd_search_cv.best_score_",
"_____no_output_____"
],
[
"rnd_search_cv.best_estimator_",
"_____no_output_____"
],
[
"rnd_search_cv.score(X_test, y_test)",
"5160/5160 [==============================] - 0s 17us/sample - loss: 0.3159\n"
],
[
"model = rnd_search_cv.best_estimator_.model\nmodel",
"_____no_output_____"
],
[
"model.evaluate(X_test, y_test)",
"5160/5160==============================] - 0s 22us/sample - loss: 0.3252\n"
]
],
[
[
"# Exercise solutions",
"_____no_output_____"
],
[
"## 1. to 9.",
"_____no_output_____"
],
[
"See appendix A.",
"_____no_output_____"
],
[
"## 10.",
"_____no_output_____"
],
[
"TODO",
"_____no_output_____"
]
]
] | [
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown"
] | [
[
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code",
"code",
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown",
"markdown",
"markdown",
"markdown",
"markdown"
]
] |
d0fea63a59ac4198638a33ec6303b40bd5131bc2 | 3,931 | ipynb | Jupyter Notebook | concatenate-csv-partials.ipynb | kiem-group/GT-reference-clustering | ae04100fa2817291492b154d1c62281dad9642ab | [
"MIT"
] | null | null | null | concatenate-csv-partials.ipynb | kiem-group/GT-reference-clustering | ae04100fa2817291492b154d1c62281dad9642ab | [
"MIT"
] | null | null | null | concatenate-csv-partials.ipynb | kiem-group/GT-reference-clustering | ae04100fa2817291492b154d1c62281dad9642ab | [
"MIT"
] | null | null | null | 20.056122 | 124 | 0.512338 | [
[
[
"Notebook to concatenate all partial csv files for the dataset into a single file. Only the essential columns are kept.",
"_____no_output_____"
]
],
[
[
"import pandas as pd \nimport os",
"_____no_output_____"
],
[
"data_dir = \"data/\"",
"_____no_output_____"
],
[
"selected_fields = ['seed_ref', 'seed_ref_id', 'ref', 'ref_id', 'TRUE']",
"_____no_output_____"
],
[
"dfs = []\npartials_dir = os.path.join(data_dir, 'partials')\nfor file in os.listdir(partials_dir):\n if \".tsv\" in file:\n #print(file)\n tmp_df = pd.read_csv(os.path.join(partials_dir, file), sep='\\t')[selected_fields]\n tmp_df['cluster_file'] = file\n dfs.append(tmp_df)",
"_____no_output_____"
],
[
"allinone_df = pd.concat(dfs, ignore_index=True)",
"_____no_output_____"
],
[
"allinone_df.rename(columns={'TRUE':'same-bibl-entity'}, inplace=True)",
"_____no_output_____"
],
[
"allinone_df.to_csv(os.path.join(data_dir, 'dataset.tsv'), sep='\\t')",
"_____no_output_____"
],
[
"# number of negative ref pairs\nallinone_df[allinone_df['same-bibl-entity']==0].shape[0]",
"_____no_output_____"
],
[
"# number of positive ref pairs\nallinone_df[allinone_df['same-bibl-entity']==1].shape[0]",
"_____no_output_____"
],
[
"# number of partly positive ref pairs\nallinone_df[allinone_df['same-bibl-entity']==0.5].shape[0]",
"_____no_output_____"
],
[
"# total number of ref pairs\nallinone_df.shape[0]",
"_____no_output_____"
]
]
] | [
"markdown",
"code"
] | [
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
d0fec7973607f32808d822bb316687073b071cc3 | 106,214 | ipynb | Jupyter Notebook | Scripts/property_type_adoption.ipynb | Minjae97/Airbnb-Trend-Resaerch | 48234bbf5b6dc66db11a80e22e34a788b017f7d0 | [
"CC0-1.0"
] | null | null | null | Scripts/property_type_adoption.ipynb | Minjae97/Airbnb-Trend-Resaerch | 48234bbf5b6dc66db11a80e22e34a788b017f7d0 | [
"CC0-1.0"
] | null | null | null | Scripts/property_type_adoption.ipynb | Minjae97/Airbnb-Trend-Resaerch | 48234bbf5b6dc66db11a80e22e34a788b017f7d0 | [
"CC0-1.0"
] | null | null | null | 47.480554 | 5,744 | 0.483354 | [
[
[
"import math\nimport pandas as pd\nfrom langdetect import detect\nimport numpy as np\nimport nltk\nfrom nltk.stem import WordNetLemmatizer\nimport string\nfrom sklearn.feature_extraction.text import CountVectorizer\nimport math\nimport matplotlib.pyplot as plt",
"_____no_output_____"
],
[
"lem = WordNetLemmatizer() #create lemmatizer",
"_____no_output_____"
],
[
"import ssl\n\ntry:\n _create_unverified_https_context = ssl._create_unverified_context\nexcept AttributeError:\n pass\nelse:\n ssl._create_default_https_context = _create_unverified_https_context\n\nnltk.download('wordnet')",
"[nltk_data] Downloading package wordnet to\n[nltk_data] /Users/shirinharandi/nltk_data...\n[nltk_data] Package wordnet is already up-to-date!\n"
],
[
"df = pd.read_csv('/Users/shirinharandi/Desktop/COMP0031/Data/en_reviews/tokyo_en.csv')\nlistings = pd.read_csv('/Users/shirinharandi/Desktop/COMP0031/Data/listings/tokyo_listings.csv')\nlistings",
"/usr/local/lib/python3.7/site-packages/IPython/core/interactiveshell.py:3051: DtypeWarning: Columns (61,62) have mixed types. Specify dtype option on import or set low_memory=False.\n interactivity=interactivity, compiler=compiler, result=result)\n"
],
[
"df",
"_____no_output_____"
],
[
"listings = listings[['id', 'room_type', 'calculated_host_listings_count']].copy()\nlistings",
"_____no_output_____"
],
[
"listings['is_superhost'] = np.where(listings['calculated_host_listings_count'] >= 10, 'Yes', listings['calculated_host_listings_count'])\nlistings['is_superhost'] = np.where(listings['calculated_host_listings_count'] < 10, 'No', listings['is_superhost'])\nlistings = listings.rename(columns={\"id\": \"listing_id\"})\nlistings",
"_____no_output_____"
],
[
"out = pd.merge(df, listings, on='listing_id')\nout",
"_____no_output_____"
],
[
"# out.to_csv(r'property_type_and_superhosts/tokyo_type.csv')",
"_____no_output_____"
],
[
"dictionary = pd.read_csv('../data/processedDict.csv')\ndictionary['word'] = dictionary['word'].apply(lambda x: lem.lemmatize(x, pos='n'))",
"_____no_output_____"
],
[
"# filepath = '../data/en_reviews/Manchester.csv'\n# reviews = pd.read_csv(filepath)\n\n# reviews = reviews['date']\n# reviews\n\nreviews = out \ntable = str.maketrans('', '', string.punctuation) #mapping to strip punctuation in review\n\n#strip punct of each review -> lemmatise -> output is list of words so join into sentences\nreviews['comments'] = reviews.comments.apply(lambda review: ' '.join(map(str, [lem.lemmatize(word.translate(table), pos='n') for word in review.lower().split()])))\nreviews\n\nreviews['date'] = pd.to_datetime(reviews['date'])",
"_____no_output_____"
],
[
"#### DELETE THIS LATER ###\nmask = (reviews['date'] >= '2014-01-01') & (reviews['date'] < '2017-01-01')\nreviews = reviews.loc[mask].copy()\nreviews",
"_____no_output_____"
],
[
"reviews",
"_____no_output_____"
],
[
"def get_trends_nice(category, subcats, reviews):\n years = [2010, 2011, 2012, 2013, 2014, 2015, 2016, 2017, 2018, 2019]\n allwords = reviews['comments'].tolist()\n allwords = \" \".join(allwords)\n unique_words = set(allwords.split(' '))\n len(unique_words)\n\n unique_words = list(unique_words)\n unique_words = [string for string in unique_words if string != \"\"]\n# len(unique_words)\n\n ls = []\n for word in unique_words:\n word = ''.join([i for i in word if not i.isdigit()])\n ls += [word]\n unique_words= ls\n unique_words = [string for string in unique_words if string != \"\"]\n unique_words = list(dict.fromkeys(unique_words))\n\n def countWords(word, review):\n count = 0\n for i in review:\n if i == word:\n count+=1\n return count\n\n def getDenom(review, unique_words):\n count = 0\n den = 0\n ls = []\n review = review.split()\n for word in review:\n kmp = countWords(word, review)\n if (kmp > 0 and word not in ls):\n ls += [word]\n den += math.log(1 + kmp)\n return den\n\n reviews['den'] = reviews['comments'].apply(lambda x: getDenom(x, unique_words))\n\n def getNom(category, review, dictionary, cat_levl=\"cat_lev1\"):\n nom = 0\n review = review.split()\n dictionaryWords = dictionary[(dictionary[cat_levl] == category)]\n dictionaryWords = dictionaryWords['word']\n for word in dictionaryWords:\n nom += math.log(1 + review.count(word))\n return nom\n\n reviews['temp'] = reviews['comments'].apply(lambda x: getNom(category, x, dictionary))\n reviews[category] = reviews['temp']*100/reviews['den']\n\n k = {}\n for subcat in subcats:\n temp = reviews['comments'].apply(lambda x: getNom(subcat, x, dictionary, cat_levl=\"cat_lev1\"))\n reviews[subcat] = temp * 100 / reviews[\"den\"]\n k[subcat] = reviews[subcat].loc[reviews[subcat] > 0].min()\n \n print(k)\n \n k_business= reviews[category].loc[reviews[category] > 0]\n k_business = k_business.min()\n print(k_business)\n\n def adoptionForSetOfReviews(category, setOfReviews, dictionary, startDate, endDate, k):\n adoption = 1\n mask = (setOfReviews['date'] >= startDate) & (setOfReviews['date'] < endDate)\n setOfReviews = setOfReviews.loc[mask]\n setOfReviews = setOfReviews[category]\n if (len(setOfReviews) == 0):\n return 0\n else:\n b = 1/len(setOfReviews)\n for review in setOfReviews:\n adoption *= math.pow((review + k),b)\n adoption = adoption - k\n return adoption\n\n d2 = {'year' : years, 'value':0.0}\n out = pd.DataFrame(data=d2)\n \n for i in range(len(years)):\n out.at[i, \"value_{}\".format(category)] = adoptionForSetOfReviews(category, reviews, dictionary, \"{}-01-01\".format(years[i]), \"{}-01-01\".format(years[i] + 1), k_business)\n# for subcat in subcats:\n# out.at[i, \"value_{}_{}\".format(category, subcat)] = adoptionForSetOfReviews(subcat, reviews, dictionary, \"{}-01-01\".format(years[i]), \"{}-01-01\".format(years[i] + 1), k[subcat])\n\n return out",
"_____no_output_____"
],
[
"house_types = [\"Private room\", \"Entire home/apt\", \"Shared room\"]",
"_____no_output_____"
],
[
"r = reviews.loc[reviews[\"room_type\"] == house_types[0]].copy()\n\ntemp = get_trends_nice(\"social\", \"social\", r)\nplt.bar(temp[\"year\"], temp[\"value_social\"])",
"{'s': nan, 'o': nan, 'c': nan, 'i': nan, 'a': nan, 'l': nan}\n0.3469604658318127\n"
],
[
"r = reviews.loc[reviews[\"room_type\"] == house_types[1]].copy()\n\ntemp = get_trends_nice(\"social\", \"social\", r)\nplt.bar(temp[\"year\"], temp[\"value_social\"])",
"{'s': nan, 'o': nan, 'c': nan, 'i': nan, 'a': nan, 'l': nan}\n0.34456163927256545\n"
],
[
"r = reviews.loc[reviews[\"room_type\"] == house_types[2]].copy()\n\ntemp = get_trends_nice(\"social\", \"social\", r)\nplt.bar(temp[\"year\"], temp[\"value_social\"])",
"{'s': nan, 'o': nan, 'c': nan, 'i': nan, 'a': nan, 'l': nan}\n0.5392772461704942\n"
],
[
"r = reviews.loc[reviews[\"room_type\"] == house_types[0]].copy()\n\ntemp = get_trends_nice(\"business\", \"business\", r)\nplt.bar(temp[\"year\"], temp[\"value_business\"])",
"{'b': nan, 'u': nan, 's': nan, 'i': nan, 'n': nan, 'e': nan}\n1.33633200713629\n"
],
[
"r = reviews.loc[reviews[\"room_type\"] == house_types[1]].copy()\n\ntemp = get_trends_nice(\"business\", \"business\", r)\nplt.bar(temp[\"year\"], temp[\"value_business\"])",
"_____no_output_____"
],
[
"r = reviews.loc[reviews[\"room_type\"] == house_types[2]].copy()\n\ntemp = get_trends_nice(\"business\", \"business\", r)\nplt.bar(temp[\"year\"], temp[\"value_business\"])",
"_____no_output_____"
]
]
] | [
"code"
] | [
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
d0fed399b603b4e8ca2a7dc8c78963d0992374e3 | 9,245 | ipynb | Jupyter Notebook | cn/.ipynb_checkpoints/sicp-3-34-checkpoint.ipynb | DamonDeng/sicp_exercise | 189bb880c1205fc43394eaf67a1fa01dbc34c1b0 | [
"MIT"
] | null | null | null | cn/.ipynb_checkpoints/sicp-3-34-checkpoint.ipynb | DamonDeng/sicp_exercise | 189bb880c1205fc43394eaf67a1fa01dbc34c1b0 | [
"MIT"
] | null | null | null | cn/.ipynb_checkpoints/sicp-3-34-checkpoint.ipynb | DamonDeng/sicp_exercise | 189bb880c1205fc43394eaf67a1fa01dbc34c1b0 | [
"MIT"
] | 1 | 2021-12-17T09:49:17.000Z | 2021-12-17T09:49:17.000Z | 30.511551 | 104 | 0.481882 | [
[
[
"## title",
"_____no_output_____"
]
],
[
[
"(define (constant value connector)\n (define (me request)\n (error \"Unknow request -- CONSTANT\" request))\n (connect connector me)\n (set-value! connector value me)\n me)\n\n(define (probe name connector)\n (define (print-probe value)\n (newline)\n (display \"Probe: \")\n (display name)\n (display \" = \")\n (display value))\n (define (process-new-value)\n (print-probe (get-value connector)))\n (define (process-forget-value)\n (print-probe \"?\"))\n (define (me request)\n (cond ((eq? request 'I-have-a-value)\n\t (process-new-value))\n\t ((eq? request 'I-lost-my-value)\n\t (process-forget-value))\n\t (else\n\t (error \"Unknow request -- PROBE \" request))))\n (connect connector me)\n me)\n\n(define (make-connector)\n (let ((value false) (informant false) (constraints '()))\n (define (set-my-value newval setter)\n (cond ((not (has-value? me))\n\t (set! value newval)\n\t (set! informant setter)\n\t (for-each-except setter\n\t\t\t inform-about-value\n\t\t\t constraints))\n\t ((not (= value newval))\n\t (error \"Constradiction\" (list value newval)))\n\t (else 'ignored)))\n (define (forget-my-value retractor)\n (if (eq? retractor informant)\n\t (begin (set! informant false)\n\t\t (for-each-except retractor\n\t\t\t\t inform-about-no-value\n\t\t\t\t constraints))\n\t 'ignored))\n (define (connect new-constraint)\n (if (not (memq new-constraint constraints))\n\t (set! constraints \n\t\t(cons new-constraint constraints)))\n (if (has-value? me)\n\t (inform-about-value new-constraint))\n 'done)\n (define (me request)\n (cond ((eq? request 'has-value?)\n\t (if informant true false))\n\t ((eq? request 'value ) value)\n\t ((eq? request 'set-value!) set-my-value)\n\t ((eq? request 'forget) forget-my-value)\n\t ((eq? request 'connect) connect)\n\t (else (error \"Unknown operation -- CONNECTOR\"\n\t\t\t request))))\n me))\n\n(define (inform-about-value constraint)\n (constraint 'I-have-a-value))\n\n(define (inform-about-no-value constraint)\n (constraint 'I-lost-my-value))\n\n(define (for-each-except exception procedure list)\n (define (loop items)\n (cond ((null? items) 'done)\n\t ((eq? (car items) exception ) (loop ( cdr items)))\n\t (else (procedure (car items))\n\t\t(loop (cdr items)))))\n (loop list))\n\n(define (has-value? connector)\n (connector 'has-value?))\n(define (get-value connector)\n (connector 'value))\n(define (set-value! connector new-value informant)\n ((connector 'set-value!) new-value informant))\n(define (forget-value! connector retractor)\n ((connector 'forget) retractor))\n(define (connect connector new-constraint)\n ((connector 'connect) new-constraint))\n\n(define (adder a1 a2 sum)\n (define (process-new-value)\n (cond ((and (has-value? a1) (has-value? a2))\n\t (set-value! sum\n\t\t (+ (get-value a1) (get-value a2))\n\t\t me))\n\t ((and (has-value? a1) (has-value? sum))\n\t (set-value! a2\n\t\t (- (get-value sum) (get-value a1))\n\t\t me))\n\t ((and (has-value? a2) (has-value? sum))\n\t (set-value! a1\n\t\t (- (get-value sum) (get-value a2))\n\t\t me))))\n (define (process-forget-value)\n (forget-value! sum me)\n (forget-value! a1 me)\n (forget-value! a2 me)\n (process-new-value))\n (define (me request)\n (cond ((eq? request 'I-have-a-value)\n\t (process-new-value))\n\t ((eq? request 'I-lost-my-value)\n\t (process-forget-value))\n\t (else\n\t (error \"Unknown request -- ADDER\" request))))\n (connect a1 me)\n (connect a2 me)\n (connect sum me)\n me)\n\n(define (multiplier m1 m2 product)\n (define (process-new-value)\n (cond ((or (and (has-value? m1) (= (get-value m1) 0))\n\t (and (has-value? m2) (= (get-value m2) 0)))\n\t (set-value! product 0 me))\n\t ((and (has-value? m1) (has-value? m2))\n\t (set-value! product\n\t\t (* (get-value m1) (get-value m2))\n\t\t me))\n\t ((and (has-value? product) (has-value? m1))\n\t (set-value! m2\n\t\t (/ (get-value product) (get-value m1))\n\t\t me))\n\t ((and (has-value? product) (has-value? m2))\n\t (set-value! m1\n\t\t (/ (get-value product) (get-value m2))\n\t\t me))))\n (define (process-forget-value)\n (forget-value! product me)\n (forget-value! m1 me)\n (forget-value! m2 me)\n (process-new-value))\n (define (me request)\n (cond ((eq? request 'I-have-a-value)\n\t (process-new-value))\n\t ((eq? request 'I-lost-my-value)\n\t (process-forget-value))\n\t (else\n\t (error \"Unknown request -- MULTIPLIER \" request))))\n (connect m1 me)\n (connect m2 me)\n (connect product me)\n me)\n\n(define (start-unit-test-adder)\n (define value1 (make-connector))\n (define value2 (make-connector))\n (define my-sum (make-connector))\n\n (adder value1 value2 my-sum)\n\n (probe 'value1 value1)\n (probe 'value2 value2)\n (probe 'my-sum my-sum)\n\n (set-value! value1 1 'user)\n (set-value! value2 2 'user)\n\n (forget-value! value1 'user)\n; (forget-value! value2 'user)\n\n (set-value! value1 4 'user)\n\n (forget-value! value1 'user)\n (forget-value! my-sum 'user)\n \n (set-value! my-sum 19 'user)\n \n )\n\n\n(define (start-unit-test-multiplier)\n (define value1 (make-connector))\n (define value2 (make-connector))\n (define my-product (make-connector))\n\n (multiplier value1 value2 my-product)\n\n (probe 'value1 value1)\n (probe 'value2 value2)\n (probe 'my-product my-product)\n\n (set-value! value1 1 'user)\n (set-value! value2 2 'user)\n\n (forget-value! value1 'user)\n; (forget-value! value2 'user)\n\n (set-value! value1 4 'user)\n\n (forget-value! value1 'user)\n (forget-value! my-product 'user)\n \n (set-value! my-product 19 'user)\n \n )\n\n(define (averager a b c)\n (define number-2 (make-connector))\n (define sum-value (make-connector))\n (adder a b sum-value)\n\n (multiplier c number-2 sum-value)\n (constant 2 number-2)\n 'ok)\n\n(define (start-test-3-33)\n (define a (make-connector))\n (define b (make-connector))\n (define c (make-connector))\n \n (averager a b c)\n\n (probe 'a a)\n (probe 'b b)\n (probe 'c c)\n\n (set-value! a 3 'user)\n (set-value! b 5 'user)\n\n)\n\n(define (squarer a b)\n (multiplier a a b))\n \n(define (start-test-3-34)\n (display \"the issue in this scenario is that we can't use same connector in one constrant\")\n (newline)\n\n)\n\n\n\n\n\n\n\n\n\n\n\n\n\n\n\n\n\n\n\n\n\n\n\n",
"_____no_output_____"
]
]
] | [
"markdown",
"code"
] | [
[
"markdown"
],
[
"code"
]
] |
d0fedb760ba1f2acdbef4b620b9c56006522c1ff | 3,777 | ipynb | Jupyter Notebook | PyPoll/Resources/Untitled.ipynb | gianx1/Python-Challenge | 0ad721c23876313ce1394d52a23e2317b1d13c51 | [
"ADSL"
] | null | null | null | PyPoll/Resources/Untitled.ipynb | gianx1/Python-Challenge | 0ad721c23876313ce1394d52a23e2317b1d13c51 | [
"ADSL"
] | null | null | null | PyPoll/Resources/Untitled.ipynb | gianx1/Python-Challenge | 0ad721c23876313ce1394d52a23e2317b1d13c51 | [
"ADSL"
] | null | null | null | 29.507813 | 155 | 0.472597 | [
[
[
"#import os module and csv file \nimport os\nimport csv\n\n\nelection_data_csv_path = os.path.join(\"election_data.csv\")\n\ncandidates = []\npercent_of_candidates = []\ntotal_num_votes_per_candidate = []\n\n\nwith open(election_data_csv_path) as csvfile:\n csvreader = csv.reader(csvfile, delimiter=\",\")\n first_row = next(csvreader)\n \n \n # get total number of votes *which is just the total number of rows*\n for row in csvreader:\n \n # list of candidates who received votes\n \n if row[2] not in candidates:\n candidates.append(row[2])\n total_num_votes_per_candidate.append(1)\n else:\n cand = candidates.index(row[2])\n total_num_votes_per_candidate[cand] = total_num_votes_per_candidate[cand] + 1\n \n total_votes = sum(total_num_votes_per_candidate)\n \n \n max_votes = max(total_num_votes_per_candidate)\n\n index_winner = total_num_votes_per_candidate.index(max_votes)\n\n winner = candidates[index_winner]\n \n#results\nprint(\"Election Results\")\nprint(\"-------------------------\")\nprint(\"Total Votes: \" + str(total_votes))\nprint(\"-------------------------\") \nfor i in range(len(candidates)):\n print(f\"{candidates[i]}: {str(round(total_num_votes_per_candidate[i]/total_votes * 100, 4))} % ({total_num_votes_per_candidate[i]})\") \nprint(\"-------------------------\")\nprint(\"Winner: \" + str(winner))\nprint(\"-------------------------\") \n\noutput = open(\"main.txt\", \"w\")\noutput.write(\"Election Results\")\noutput.write(\"----------------------------\")\noutput.write(\"Total Votes: \" + str(total_votes))\noutput.write(\"-------------------------\")\nfor i in range(len(candidates)):\n output.write(f\"{candidates[i]}: {str(round(total_num_votes_per_candidate[i]/total_votes * 100, 4))} % ({total_num_votes_per_candidate[i]})\")\noutput.write(\"-------------------------\")\noutput.write(\"Winner: \" + str(winner))\noutput.write(\"-------------------------\")",
"Election Results\n-------------------------\nTotal Votes: 3521001\n-------------------------\nKhan: 63.0 % (2218231)\nCorrey: 20.0 % (704200)\nLi: 14.0 % (492940)\nO'Tooley: 3.0 % (105630)\n-------------------------\nWinner: Khan\n-------------------------\n"
]
]
] | [
"code"
] | [
[
"code"
]
] |
d0ff002e4ff46118b9f805a518d1a911d62d3342 | 9,573 | ipynb | Jupyter Notebook | 04 Python Teil 2/05 Dateien.ipynb | JonnyGrafico/00_Git_Projekte | 5df548ecd43a7071f60ce26d06eceb33f8a11815 | [
"MIT"
] | null | null | null | 04 Python Teil 2/05 Dateien.ipynb | JonnyGrafico/00_Git_Projekte | 5df548ecd43a7071f60ce26d06eceb33f8a11815 | [
"MIT"
] | null | null | null | 04 Python Teil 2/05 Dateien.ipynb | JonnyGrafico/00_Git_Projekte | 5df548ecd43a7071f60ce26d06eceb33f8a11815 | [
"MIT"
] | null | null | null | 22.471831 | 137 | 0.486159 | [
[
[
"# Dateien",
"_____no_output_____"
],
[
"## Eine Textdatei lesen und ihren Inhalt ausgeben",
"_____no_output_____"
]
],
[
[
"# Wir öffnen die Datei lesen.txt zum Lesen (\"r\") und speichern ihren Inhalt in die Variable file\nfile = open(\"lesen.txt\", \"r\")\n\n# Wir gehen alle Zeilen nacheinander durch\n# In der txt-Datei stehen für uns nicht sichtbare Zeilenumbruchszeichen, durch die jeweils das Ende einer Zeile markiert ist\nfor line in file:\n # Eine Zeile ohne Zeilenumbruch ausgeben\n print(line.strip())\nfile.close()",
"Hallo Leute.\nIch bin eine wunderschöne weitere Zeile. Gruss von Jürg\n"
],
[
"# Wir öffnen die Datei lesen.txt zum Lesen (\"r\") und speichern ihren Inhalt in die Variable file\nfile = open(\"lesen-Kopie-Juerg.txt\", \"r\")\n\n# Wir gehen alle Zeilen nacheinander durch\n# In der txt-Datei stehen für uns nicht sichtbare Zeilenumbruchszeichen, durch die jeweils das Ende einer Zeile markiert ist\nfor line in file:\n # Eine Zeile ohne Zeilenumbruch ausgeben\n print(line.strip())\nfile.close()",
"Das ist eine Kopie von Jürg\n"
]
],
[
[
"## In eine Textdatei schreiben",
"_____no_output_____"
]
],
[
[
"# Wir öffnen eine Datei zum Schreiben (\"w\": write)\nfile = open(\"schreiben.txt\", \"w\")\n\nstudents = [\"Max\", \"Monika\", \"Erik\", \"Franziska\", \"Juerg\", \"Peter\"]\n\n# Wir loopen mit einer for-Schleife durch die Liste students\nfor student in students:\n # Mit der write-Methode schreiben wir den aktuellen String student und einen Zeilenumbruch in das file-Objekt\n file.write(student + \"\\n\")\n\n# Abschließend müssen wir die Datei wieder schließen\nfile.close()",
"_____no_output_____"
],
[
"# Wir öffnen eine Datei zum Schreiben (\"w\": write)\nfile = open(\"schreiben.txt\", \"w\")\n\nstudents = [\"Max\", \"Monika\", \"Erik\", \"Franziska\", \"Juerg\", \"Peter\"]\n\n# Wir loopen mit einer for-Schleife durch die Liste students\nfor student in students:\n # Mit der write-Methode schreiben wir den aktuellen String student und einen Zeilenumbruch in das file-Objekt\n file.write(student + \"\\n\")\n\n# Abschließend müssen wir die Datei wieder schließen\nfile.close()",
"_____no_output_____"
]
],
[
[
"## Dateien öffnen mit with\nWenn wir Dateien mit einer with-Konstruktion öffnen, dann brauchen wir sie nicht mehr explizit mit der close()-Methode schließen.",
"_____no_output_____"
]
],
[
[
"with open(\"lesen.txt\", \"r\") as file:\n for line in file:\n print(line)",
"Hallo Leute.\n\nIch bin eine wunderschöne weitere Zeile.\n"
],
[
"with open(\"lesen.txt\", \"r\") as file:\n for line in file:\n print(line)\n \n \n ",
"_____no_output_____"
]
],
[
[
"## CSV-Datei lesen\ncsv steht für comma separated values. Auch solche csv-Dateien können wir mit Python auslesen.",
"_____no_output_____"
]
],
[
[
"with open(\"datei.csv\") as file:\n for line in file:\n data = line.strip().split(\";\")\n print(data[0] + \": \" + data[1])",
"Muenchen: 1800000\nBerlin: 3000000\nBudapest: 2000000\n"
]
],
[
[
"## CSV-Datei lesen (und Daten überspringen)\n\nIn dieser Lektion lernst du:\n\n- Wie du eine CSV-Datei einliest, und Zeilen überspringen kannst.",
"_____no_output_____"
]
],
[
[
"with open(\"datei.csv\") as file:\n for line in file:\n data = line.strip().split(\";\")\n \n if (data[0]) == \"Aaron\":\n print(data[0])\n \n if data[2] == \"BUD\":\n continue\n \n print(data)\n \n #if data[2] == \"BER\" or data[2] == \"BUD\":\n # print(data[2])\n # print(data)",
"['Berlin', '3000000', 'BER']\n"
],
[
"sum = 0\nwith open(\"20151001_hundenamen.csv\") as file:\n \n for line in file:\n data = line.strip().split(\",\")\n \n if (data[0]) == '\"Aaron\"':\n \n if int(data[1]) < 2012:\n sum = sum + 1 \nprint(sum)\n ",
"4\n"
],
[
"with open(\"datei.csv\") as file:\n for line in file:\n data = line.strip().split(\";\")\n print(data[1] + \" \" + data[2])",
"1800000 MUC\n3000000 BER\n2000000 BUD\n"
]
],
[
[
"## Übung\n- Besorgt euch die datei https://data.stadt-zuerich.ch/dataset/pd-stapo-hundenamen/resource/8bf2127d-c354-4834-8590-9666cbd6e160\n- Ihr findet sie auch im Ordner 20151001_hundenamen.csv\n- Findet heraus wie oft der Hundename \"Aaron\" zwischen 2000 - 2012 gebraucht wurde. ",
"_____no_output_____"
]
],
[
[
"n = \"1975\"\nprint(int(n) < 1990)",
"True\n"
],
[
"jahre = [\"Year\", \"1990\", \"1992\"]\n\nfor jahr in jahre:\n if jahr == \"Year\":\n continue\n print(int(jahr))",
"1990\n1992\n"
],
[
"### Euer code hier",
"_____no_output_____"
]
]
] | [
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] | [
[
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
]
] |
d0ff06791d0dcc169a23e9dcba6a7a046600814c | 22,545 | ipynb | Jupyter Notebook | julia/.ipynb_checkpoints/julia_introduction-checkpoint.ipynb | BKC1995/intro-to-computing | f5c7ba0b881d9fc1fd2056dcbefd60fd722a63ef | [
"MIT"
] | 5 | 2019-09-26T09:30:55.000Z | 2021-02-14T10:15:26.000Z | julia/.ipynb_checkpoints/julia_introduction-checkpoint.ipynb | BKC1995/intro-to-computing | f5c7ba0b881d9fc1fd2056dcbefd60fd722a63ef | [
"MIT"
] | null | null | null | julia/.ipynb_checkpoints/julia_introduction-checkpoint.ipynb | BKC1995/intro-to-computing | f5c7ba0b881d9fc1fd2056dcbefd60fd722a63ef | [
"MIT"
] | 9 | 2019-09-26T13:30:19.000Z | 2019-09-26T20:25:27.000Z | 27.527473 | 442 | 0.555068 | [
[
[
"empty"
]
]
] | [
"empty"
] | [
[
"empty"
]
] |
d0ff0a1edbfcb0fa00937d47d34b8e7887dab33b | 24,273 | ipynb | Jupyter Notebook | module3-permutation-boosting/LS_DS_233.ipynb | ilEnzio/DS-Unit-2-Applied-Modeling | 8072235a6be561c7231a8416b7477eea6dab3bdf | [
"MIT"
] | null | null | null | module3-permutation-boosting/LS_DS_233.ipynb | ilEnzio/DS-Unit-2-Applied-Modeling | 8072235a6be561c7231a8416b7477eea6dab3bdf | [
"MIT"
] | null | null | null | module3-permutation-boosting/LS_DS_233.ipynb | ilEnzio/DS-Unit-2-Applied-Modeling | 8072235a6be561c7231a8416b7477eea6dab3bdf | [
"MIT"
] | null | null | null | 38.345972 | 1,022 | 0.629959 | [
[
[
"Lambda School Data Science\n\n*Unit 2, Sprint 3, Module 3*\n\n---",
"_____no_output_____"
],
[
"# Permutation & Boosting\n\n- Get **permutation importances** for model interpretation and feature selection\n- Use xgboost for **gradient boosting**",
"_____no_output_____"
],
[
"### Setup\n\nRun the code cell below. You can work locally (follow the [local setup instructions](https://lambdaschool.github.io/ds/unit2/local/)) or on Colab.\n\nLibraries:\n\n- category_encoders\n- [**eli5**](https://eli5.readthedocs.io/en/latest/)\n- matplotlib\n- numpy\n- pandas\n- scikit-learn\n- [**xgboost**](https://xgboost.readthedocs.io/en/latest/)",
"_____no_output_____"
]
],
[
[
"%%capture\nimport sys\n\n# If you're on Colab:\nif 'google.colab' in sys.modules:\n DATA_PATH = 'https://raw.githubusercontent.com/LambdaSchool/DS-Unit-2-Applied-Modeling/master/data/'\n !pip install category_encoders==2.*\n !pip install eli5\n\n# If you're working locally:\nelse:\n DATA_PATH = '../data/'",
"_____no_output_____"
]
],
[
[
"We'll go back to Tanzania Waterpumps for this lesson.",
"_____no_output_____"
]
],
[
[
"import numpy as np\nimport pandas as pd\nfrom sklearn.model_selection import train_test_split\n\n# Merge train_features.csv & train_labels.csv\ntrain = pd.merge(pd.read_csv(DATA_PATH+'waterpumps/train_features.csv'), \n pd.read_csv(DATA_PATH+'waterpumps/train_labels.csv'))\n\n# Read test_features.csv & sample_submission.csv\ntest = pd.read_csv(DATA_PATH+'waterpumps/test_features.csv')\nsample_submission = pd.read_csv(DATA_PATH+'waterpumps/sample_submission.csv')\n\n\n# Split train into train & val\ntrain, val = train_test_split(train, train_size=0.80, test_size=0.20, \n stratify=train['status_group'], random_state=42)\n\n\ndef wrangle(X):\n \"\"\"Wrangle train, validate, and test sets in the same way\"\"\"\n \n # Prevent SettingWithCopyWarning\n X = X.copy()\n \n # About 3% of the time, latitude has small values near zero,\n # outside Tanzania, so we'll treat these values like zero.\n X['latitude'] = X['latitude'].replace(-2e-08, 0)\n \n # When columns have zeros and shouldn't, they are like null values.\n # So we will replace the zeros with nulls, and impute missing values later.\n # Also create a \"missing indicator\" column, because the fact that\n # values are missing may be a predictive signal.\n cols_with_zeros = ['longitude', 'latitude', 'construction_year', \n 'gps_height', 'population']\n for col in cols_with_zeros:\n X[col] = X[col].replace(0, np.nan)\n X[col+'_MISSING'] = X[col].isnull()\n \n # Drop duplicate columns\n duplicates = ['quantity_group', 'payment_type']\n X = X.drop(columns=duplicates)\n \n # Drop recorded_by (never varies) and id (always varies, random)\n unusable_variance = ['recorded_by', 'id']\n X = X.drop(columns=unusable_variance)\n \n # Convert date_recorded to datetime\n X['date_recorded'] = pd.to_datetime(X['date_recorded'], infer_datetime_format=True)\n \n # Extract components from date_recorded, then drop the original column\n X['year_recorded'] = X['date_recorded'].dt.year\n X['month_recorded'] = X['date_recorded'].dt.month\n X['day_recorded'] = X['date_recorded'].dt.day\n X = X.drop(columns='date_recorded')\n \n # Engineer feature: how many years from construction_year to date_recorded\n X['years'] = X['year_recorded'] - X['construction_year']\n X['years_MISSING'] = X['years'].isnull()\n \n # return the wrangled dataframe\n return X\n\ntrain = wrangle(train)\nval = wrangle(val)\ntest = wrangle(test)",
"_____no_output_____"
],
[
"# Arrange data into X features matrix and y target vector\ntarget = 'status_group'\nX_train = train.drop(columns=target)\ny_train = train[target]\nX_val = val.drop(columns=target)\ny_val = val[target]\nX_test = test",
"_____no_output_____"
],
[
"import category_encoders as ce\nfrom sklearn.impute import SimpleImputer\nfrom sklearn.ensemble import RandomForestClassifier\nfrom sklearn.model_selection import train_test_split\nfrom sklearn.pipeline import make_pipeline\n\npipeline = make_pipeline(\n ce.OrdinalEncoder(), \n SimpleImputer(strategy='median'), \n RandomForestClassifier(n_estimators=100, random_state=42, n_jobs=-1)\n)\n\n# Fit on train, score on val\npipeline.fit(X_train, y_train)\nprint('Validation Accuracy', pipeline.score(X_val, y_val))",
"_____no_output_____"
]
],
[
[
"# Get permutation importances for model interpretation and feature selection",
"_____no_output_____"
],
[
"## Overview",
"_____no_output_____"
],
[
"Default Feature Importances are fast, but Permutation Importances may be more accurate.\n\nThese links go deeper with explanations and examples:\n\n- Permutation Importances\n - [Kaggle / Dan Becker: Machine Learning Explainability](https://www.kaggle.com/dansbecker/permutation-importance)\n - [Christoph Molnar: Interpretable Machine Learning](https://christophm.github.io/interpretable-ml-book/feature-importance.html)\n- (Default) Feature Importances\n - [Ando Saabas: Selecting good features, Part 3, Random Forests](https://blog.datadive.net/selecting-good-features-part-iii-random-forests/)\n - [Terence Parr, et al: Beware Default Random Forest Importances](https://explained.ai/rf-importance/index.html)",
"_____no_output_____"
],
[
"There are three types of feature importances:",
"_____no_output_____"
],
[
"### 1. (Default) Feature Importances\n\nFastest, good for first estimates, but be aware:\n\n\n\n>**When the dataset has two (or more) correlated features, then from the point of view of the model, any of these correlated features can be used as the predictor, with no concrete preference of one over the others.** But once one of them is used, the importance of others is significantly reduced since effectively the impurity they can remove is already removed by the first feature. As a consequence, they will have a lower reported importance. This is not an issue when we want to use feature selection to reduce overfitting, since it makes sense to remove features that are mostly duplicated by other features. But when interpreting the data, it can lead to the incorrect conclusion that one of the variables is a strong predictor while the others in the same group are unimportant, while actually they are very close in terms of their relationship with the response variable. — [Selecting good features – Part III: random forests](https://blog.datadive.net/selecting-good-features-part-iii-random-forests/) \n\n\n \n > **The scikit-learn Random Forest feature importance ... tends to inflate the importance of continuous or high-cardinality categorical variables.** ... Breiman and Cutler, the inventors of Random Forests, indicate that this method of “adding up the gini decreases for each individual variable over all trees in the forest gives a **fast** variable importance that is often very consistent with the permutation importance measure.” — [Beware Default Random Forest Importances](https://explained.ai/rf-importance/index.html)\n\n \n",
"_____no_output_____"
]
],
[
[
"# Get feature importances\nrf = pipeline.named_steps['randomforestclassifier']\nimportances = pd.Series(rf.feature_importances_, X_train.columns)\n\n# Plot feature importances\n%matplotlib inline\nimport matplotlib.pyplot as plt\n\nn = 20\nplt.figure(figsize=(10,n/2))\nplt.title(f'Top {n} features')\nimportances.sort_values()[-n:].plot.barh(color='grey');",
"_____no_output_____"
]
],
[
[
"### 2. Drop-Column Importance\n\nThe best in theory, but too slow in practice",
"_____no_output_____"
]
],
[
[
"column = 'quantity'\n\n# Fit without column\npipeline = make_pipeline(\n ce.OrdinalEncoder(), \n SimpleImputer(strategy='median'), \n RandomForestClassifier(n_estimators=100, random_state=42, n_jobs=-1)\n)\npipeline.fit(X_train.drop(columns=column), y_train)\nscore_without = pipeline.score(X_val.drop(columns=column), y_val)\nprint(f'Validation Accuracy without {column}: {score_without}')\n\n# Fit with column\npipeline = make_pipeline(\n ce.OrdinalEncoder(), \n SimpleImputer(strategy='median'), \n RandomForestClassifier(n_estimators=100, random_state=42, n_jobs=-1)\n)\npipeline.fit(X_train, y_train)\nscore_with = pipeline.score(X_val, y_val)\nprint(f'Validation Accuracy with {column}: {score_with}')\n\n# Compare the error with & without column\nprint(f'Drop-Column Importance for {column}: {score_with - score_without}')",
"_____no_output_____"
]
],
[
[
"### 3. Permutation Importance\n\nPermutation Importance is a good compromise between Feature Importance based on impurity reduction (which is the fastest) and Drop Column Importance (which is the \"best.\")\n\n[The ELI5 library documentation explains,](https://eli5.readthedocs.io/en/latest/blackbox/permutation_importance.html)\n\n> Importance can be measured by looking at how much the score (accuracy, F1, R^2, etc. - any score we’re interested in) decreases when a feature is not available.\n>\n> To do that one can remove feature from the dataset, re-train the estimator and check the score. But it requires re-training an estimator for each feature, which can be computationally intensive. ...\n>\n>To avoid re-training the estimator we can remove a feature only from the test part of the dataset, and compute score without using this feature. It doesn’t work as-is, because estimators expect feature to be present. So instead of removing a feature we can replace it with random noise - feature column is still there, but it no longer contains useful information. This method works if noise is drawn from the same distribution as original feature values (as otherwise estimator may fail). The simplest way to get such noise is to shuffle values for a feature, i.e. use other examples’ feature values - this is how permutation importance is computed.\n>\n>The method is most suitable for computing feature importances when a number of columns (features) is not huge; it can be resource-intensive otherwise.",
"_____no_output_____"
],
[
"### Do-It-Yourself way, for intuition",
"_____no_output_____"
],
[
"### With eli5 library\n\nFor more documentation on using this library, see:\n- [eli5.sklearn.PermutationImportance](https://eli5.readthedocs.io/en/latest/autodocs/sklearn.html#eli5.sklearn.permutation_importance.PermutationImportance)\n- [eli5.show_weights](https://eli5.readthedocs.io/en/latest/autodocs/eli5.html#eli5.show_weights)\n- [scikit-learn user guide, `scoring` parameter](https://scikit-learn.org/stable/modules/model_evaluation.html#the-scoring-parameter-defining-model-evaluation-rules)\n\neli5 doesn't work with pipelines.",
"_____no_output_____"
]
],
[
[
"# Ignore warnings\n",
"_____no_output_____"
]
],
[
[
"### We can use importances for feature selection\n\nFor example, we can remove features with zero importance. The model trains faster and the score does not decrease.",
"_____no_output_____"
],
[
"# Use xgboost for gradient boosting",
"_____no_output_____"
],
[
"## Overview",
"_____no_output_____"
],
[
"In the Random Forest lesson, you learned this advice:\n\n#### Try Tree Ensembles when you do machine learning with labeled, tabular data\n- \"Tree Ensembles\" means Random Forest or **Gradient Boosting** models. \n- [Tree Ensembles often have the best predictive accuracy](https://arxiv.org/abs/1708.05070) with labeled, tabular data.\n- Why? Because trees can fit non-linear, non-[monotonic](https://en.wikipedia.org/wiki/Monotonic_function) relationships, and [interactions](https://christophm.github.io/interpretable-ml-book/interaction.html) between features.\n- A single decision tree, grown to unlimited depth, will [overfit](http://www.r2d3.us/visual-intro-to-machine-learning-part-1/). We solve this problem by ensembling trees, with bagging (Random Forest) or **[boosting](https://www.youtube.com/watch?v=GM3CDQfQ4sw)** (Gradient Boosting).\n- Random Forest's advantage: may be less sensitive to hyperparameters. **Gradient Boosting's advantage:** may get better predictive accuracy.",
"_____no_output_____"
],
[
"Like Random Forest, Gradient Boosting uses ensembles of trees. But the details of the ensembling technique are different:\n\n### Understand the difference between boosting & bagging\n\nBoosting (used by Gradient Boosting) is different than Bagging (used by Random Forests). \n\nHere's an excerpt from [_An Introduction to Statistical Learning_](http://www-bcf.usc.edu/~gareth/ISL/ISLR%20Seventh%20Printing.pdf) Chapter 8.2.3, Boosting:\n\n>Recall that bagging involves creating multiple copies of the original training data set using the bootstrap, fitting a separate decision tree to each copy, and then combining all of the trees in order to create a single predictive model.\n>\n>**Boosting works in a similar way, except that the trees are grown _sequentially_: each tree is grown using information from previously grown trees.**\n>\n>Unlike fitting a single large decision tree to the data, which amounts to _fitting the data hard_ and potentially overfitting, the boosting approach instead _learns slowly._ Given the current model, we fit a decision tree to the residuals from the model.\n>\n>We then add this new decision tree into the fitted function in order to update the residuals. Each of these trees can be rather small, with just a few terminal nodes. **By fitting small trees to the residuals, we slowly improve fˆ in areas where it does not perform well.**\n>\n>Note that in boosting, unlike in bagging, the construction of each tree depends strongly on the trees that have already been grown.\n\nThis high-level overview is all you need to know for now. If you want to go deeper, we recommend you watch the StatQuest videos on gradient boosting!",
"_____no_output_____"
],
[
"Let's write some code. We have lots of options for which libraries to use:\n\n#### Python libraries for Gradient Boosting\n- [scikit-learn Gradient Tree Boosting](https://scikit-learn.org/stable/modules/ensemble.html#gradient-boosting) — slower than other libraries, but [the new version may be better](https://twitter.com/amuellerml/status/1129443826945396737)\n - Anaconda: already installed\n - Google Colab: already installed\n- [xgboost](https://xgboost.readthedocs.io/en/latest/) — can accept missing values and enforce [monotonic constraints](https://xiaoxiaowang87.github.io/monotonicity_constraint/)\n - Anaconda, Mac/Linux: `conda install -c conda-forge xgboost`\n - Windows: `conda install -c anaconda py-xgboost`\n - Google Colab: already installed\n- [LightGBM](https://lightgbm.readthedocs.io/en/latest/) — can accept missing values and enforce [monotonic constraints](https://blog.datadive.net/monotonicity-constraints-in-machine-learning/)\n - Anaconda: `conda install -c conda-forge lightgbm`\n - Google Colab: already installed\n- [CatBoost](https://catboost.ai/) — can accept missing values and use [categorical features](https://catboost.ai/docs/concepts/algorithm-main-stages_cat-to-numberic.html) without preprocessing\n - Anaconda: `conda install -c conda-forge catboost`\n - Google Colab: `pip install catboost`",
"_____no_output_____"
],
[
"In this lesson, you'll use a new library, xgboost — But it has an API that's almost the same as scikit-learn, so it won't be a hard adjustment!\n\n#### [XGBoost Python API Reference: Scikit-Learn API](https://xgboost.readthedocs.io/en/latest/python/python_api.html#module-xgboost.sklearn)",
"_____no_output_____"
],
[
"#### [Avoid Overfitting By Early Stopping With XGBoost In Python](https://machinelearningmastery.com/avoid-overfitting-by-early-stopping-with-xgboost-in-python/)\n\nWhy is early stopping better than a For loop, or GridSearchCV, to optimize `n_estimators`?\n\nWith early stopping, if `n_iterations` is our number of iterations, then we fit `n_iterations` decision trees.\n\nWith a for loop, or GridSearchCV, we'd fit `sum(range(1,n_rounds+1))` trees.\n\nBut it doesn't work well with pipelines. You may need to re-run multiple times with different values of other parameters such as `max_depth` and `learning_rate`.\n\n#### XGBoost parameters\n- [Notes on parameter tuning](https://xgboost.readthedocs.io/en/latest/tutorials/param_tuning.html)\n- [Parameters documentation](https://xgboost.readthedocs.io/en/latest/parameter.html)\n",
"_____no_output_____"
],
[
"### Try adjusting these hyperparameters\n\n#### Random Forest\n- class_weight (for imbalanced classes)\n- max_depth (usually high, can try decreasing)\n- n_estimators (too low underfits, too high wastes time)\n- min_samples_leaf (increase if overfitting)\n- max_features (decrease for more diverse trees)\n\n#### Xgboost\n- scale_pos_weight (for imbalanced classes)\n- max_depth (usually low, can try increasing)\n- n_estimators (too low underfits, too high wastes time/overfits) — Use Early Stopping!\n- learning_rate (too low underfits, too high overfits)\n\nFor more ideas, see [Notes on Parameter Tuning](https://xgboost.readthedocs.io/en/latest/tutorials/param_tuning.html) and [DART booster](https://xgboost.readthedocs.io/en/latest/tutorials/dart.html).",
"_____no_output_____"
],
[
"## Challenge\n\nYou will use your portfolio project dataset for all assignments this sprint. Complete these tasks for your project, and document your work.\n\n- Continue to clean and explore your data. Make exploratory visualizations.\n- Fit a model. Does it beat your baseline?\n- Try xgboost.\n- Get your model's permutation importances.\n\nYou should try to complete an initial model today, because the rest of the week, we're making model interpretation visualizations.\n\nBut, if you aren't ready to try xgboost and permutation importances with your dataset today, you can practice with another dataset instead. You may choose any dataset you've worked with previously.",
"_____no_output_____"
]
]
] | [
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown"
] | [
[
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown",
"markdown",
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown"
]
] |
d0ff0bf5fc4262fa7ebfb27c9f07ba82c6f9507b | 40,789 | ipynb | Jupyter Notebook | Credit Risk Modeling/Credit Risk Modeling - LGD and EAD Models - With Comments - 12-1.ipynb | autodidact-m/Projects | f4c0473adba42f3a629b62eb09d3b1df91982f46 | [
"Apache-2.0"
] | null | null | null | Credit Risk Modeling/Credit Risk Modeling - LGD and EAD Models - With Comments - 12-1.ipynb | autodidact-m/Projects | f4c0473adba42f3a629b62eb09d3b1df91982f46 | [
"Apache-2.0"
] | null | null | null | Credit Risk Modeling/Credit Risk Modeling - LGD and EAD Models - With Comments - 12-1.ipynb | autodidact-m/Projects | f4c0473adba42f3a629b62eb09d3b1df91982f46 | [
"Apache-2.0"
] | null | null | null | 30.371556 | 448 | 0.608718 | [
[
[
"# Import Libraries",
"_____no_output_____"
]
],
[
[
"import numpy as np\nimport pandas as pd",
"_____no_output_____"
]
],
[
[
"# Import Data",
"_____no_output_____"
]
],
[
[
"# Import data.\nloan_data_preprocessed_backup = pd.read_csv('loan_data_2007_2014_preprocessed.csv')",
"_____no_output_____"
]
],
[
[
"# Explore Data",
"_____no_output_____"
]
],
[
[
"loan_data_preprocessed = loan_data_preprocessed_backup.copy()",
"_____no_output_____"
],
[
"loan_data_preprocessed.columns.values\n# Displays all column names.",
"_____no_output_____"
],
[
"loan_data_preprocessed.head()",
"_____no_output_____"
],
[
"loan_data_preprocessed.tail()",
"_____no_output_____"
],
[
"loan_data_defaults = loan_data_preprocessed[loan_data_preprocessed['loan_status'].isin(['Charged Off','Does not meet the credit policy. Status:Charged Off'])]\n# Here we take only the accounts that were charged-off (written-off).",
"_____no_output_____"
],
[
"loan_data_defaults.shape",
"_____no_output_____"
],
[
"pd.options.display.max_rows = None\n# Sets the pandas dataframe options to display all columns/ rows.",
"_____no_output_____"
],
[
"loan_data_defaults.isnull().sum()",
"_____no_output_____"
]
],
[
[
"# Independent Variables",
"_____no_output_____"
]
],
[
[
"loan_data_defaults['mths_since_last_delinq'].fillna(0, inplace = True)\n# We fill the missing values with zeroes.",
"_____no_output_____"
],
[
"#loan_data_defaults['mths_since_last_delinq'].fillna(loan_data_defaults['mths_since_last_delinq'].max() + 12, inplace=True)",
"_____no_output_____"
],
[
"loan_data_defaults['mths_since_last_record'].fillna(0, inplace=True)\n# We fill the missing values with zeroes.",
"_____no_output_____"
]
],
[
[
"# Dependent Variables",
"_____no_output_____"
]
],
[
[
"loan_data_defaults['recovery_rate'] = loan_data_defaults['recoveries'] / loan_data_defaults['funded_amnt']\n# We calculate the dependent variable for the LGD model: recovery rate.\n# It is the ratio of recoveries and funded amount.",
"_____no_output_____"
],
[
"loan_data_defaults['recovery_rate'].describe()\n# Shows some descriptive statisics for the values of a column.",
"_____no_output_____"
],
[
"loan_data_defaults['recovery_rate'] = np.where(loan_data_defaults['recovery_rate'] > 1, 1, loan_data_defaults['recovery_rate'])\nloan_data_defaults['recovery_rate'] = np.where(loan_data_defaults['recovery_rate'] < 0, 0, loan_data_defaults['recovery_rate'])\n# We set recovery rates that are greater than 1 to 1 and recovery rates that are less than 0 to 0.",
"_____no_output_____"
],
[
"loan_data_defaults['recovery_rate'].describe()\n# Shows some descriptive statisics for the values of a column.",
"_____no_output_____"
],
[
"loan_data_defaults['CCF'] = (loan_data_defaults['funded_amnt'] - loan_data_defaults['total_rec_prncp']) / loan_data_defaults['funded_amnt']\n# We calculate the dependent variable for the EAD model: credit conversion factor.\n# It is the ratio of the difference of the amount used at the moment of default to the total funded amount.",
"_____no_output_____"
],
[
"loan_data_defaults['CCF'].describe()\n# Shows some descriptive statisics for the values of a column.",
"_____no_output_____"
],
[
"loan_data_defaults.to_csv('loan_data_defaults.csv')\n# We save the data to a CSV file.",
"_____no_output_____"
]
],
[
[
"# Explore Dependent Variables",
"_____no_output_____"
]
],
[
[
"import matplotlib.pyplot as plt\nimport seaborn as sns\nsns.set()",
"_____no_output_____"
],
[
"plt.hist(loan_data_defaults['recovery_rate'], bins = 100)\n# We plot a histogram of a variable with 100 bins.",
"_____no_output_____"
],
[
"plt.hist(loan_data_defaults['recovery_rate'], bins = 50)\n# We plot a histogram of a variable with 50 bins.",
"_____no_output_____"
],
[
"plt.hist(loan_data_defaults['CCF'], bins = 100)\n# We plot a histogram of a variable with 100 bins.",
"_____no_output_____"
],
[
"loan_data_defaults['recovery_rate_0_1'] = np.where(loan_data_defaults['recovery_rate'] == 0, 0, 1)\n# We create a new variable which is 0 if recovery rate is 0 and 1 otherwise.",
"_____no_output_____"
],
[
"loan_data_defaults['recovery_rate_0_1']",
"_____no_output_____"
]
],
[
[
"# LGD Model",
"_____no_output_____"
],
[
"### Splitting Data",
"_____no_output_____"
]
],
[
[
"from sklearn.model_selection import train_test_split",
"_____no_output_____"
],
[
"# LGD model stage 1 datasets: recovery rate 0 or greater than 0.\nlgd_inputs_stage_1_train, lgd_inputs_stage_1_test, lgd_targets_stage_1_train, lgd_targets_stage_1_test = train_test_split(loan_data_defaults.drop(['good_bad', 'recovery_rate','recovery_rate_0_1', 'CCF'], axis = 1), loan_data_defaults['recovery_rate_0_1'], test_size = 0.2, random_state = 42)\n# Takes a set of inputs and a set of targets as arguments. Splits the inputs and the targets into four dataframes:\n# Inputs - Train, Inputs - Test, Targets - Train, Targets - Test.",
"_____no_output_____"
]
],
[
[
"### Preparing the Inputs",
"_____no_output_____"
]
],
[
[
"features_all = ['grade:A',\n'grade:B',\n'grade:C',\n'grade:D',\n'grade:E',\n'grade:F',\n'grade:G',\n'home_ownership:MORTGAGE',\n'home_ownership:NONE',\n'home_ownership:OTHER',\n'home_ownership:OWN',\n'home_ownership:RENT',\n'verification_status:Not Verified',\n'verification_status:Source Verified',\n'verification_status:Verified',\n'purpose:car',\n'purpose:credit_card',\n'purpose:debt_consolidation',\n'purpose:educational',\n'purpose:home_improvement',\n'purpose:house',\n'purpose:major_purchase',\n'purpose:medical',\n'purpose:moving',\n'purpose:other',\n'purpose:renewable_energy',\n'purpose:small_business',\n'purpose:vacation',\n'purpose:wedding',\n'initial_list_status:f',\n'initial_list_status:w',\n'term_int',\n'emp_length_int',\n'mths_since_issue_d',\n'mths_since_earliest_cr_line',\n'funded_amnt',\n'int_rate',\n'installment',\n'annual_inc',\n'dti',\n'delinq_2yrs',\n'inq_last_6mths',\n'mths_since_last_delinq',\n'mths_since_last_record',\n'open_acc',\n'pub_rec',\n'total_acc',\n'acc_now_delinq',\n'total_rev_hi_lim']\n# List of all independent variables for the models.",
"_____no_output_____"
],
[
"features_reference_cat = ['grade:G',\n'home_ownership:RENT',\n'verification_status:Verified',\n'purpose:credit_card',\n'initial_list_status:f']\n# List of the dummy variable reference categories. ",
"_____no_output_____"
],
[
"lgd_inputs_stage_1_train = lgd_inputs_stage_1_train[features_all]\n# Here we keep only the variables we need for the model.",
"_____no_output_____"
],
[
"lgd_inputs_stage_1_train = lgd_inputs_stage_1_train.drop(features_reference_cat, axis = 1)\n# Here we remove the dummy variable reference categories.",
"_____no_output_____"
],
[
"lgd_inputs_stage_1_train.isnull().sum()\n# Check for missing values. We check whether the value of each row for each column is missing or not,\n# then sum accross columns.",
"_____no_output_____"
]
],
[
[
"### Estimating the Model",
"_____no_output_____"
]
],
[
[
"# P values for sklearn logistic regression.\n\n# Class to display p-values for logistic regression in sklearn.\n\nfrom sklearn import linear_model\nimport scipy.stats as stat\n\nclass LogisticRegression_with_p_values:\n \n def __init__(self,*args,**kwargs):#,**kwargs):\n self.model = linear_model.LogisticRegression(*args,**kwargs)#,**args)\n\n def fit(self,X,y):\n self.model.fit(X,y)\n \n #### Get p-values for the fitted model ####\n denom = (2.0 * (1.0 + np.cosh(self.model.decision_function(X))))\n denom = np.tile(denom,(X.shape[1],1)).T\n F_ij = np.dot((X / denom).T,X) ## Fisher Information Matrix\n Cramer_Rao = np.linalg.inv(F_ij) ## Inverse Information Matrix\n sigma_estimates = np.sqrt(np.diagonal(Cramer_Rao))\n z_scores = self.model.coef_[0] / sigma_estimates # z-score for eaach model coefficient\n p_values = [stat.norm.sf(abs(x)) * 2 for x in z_scores] ### two tailed test for p-values\n \n self.coef_ = self.model.coef_\n self.intercept_ = self.model.intercept_\n #self.z_scores = z_scores\n self.p_values = p_values\n #self.sigma_estimates = sigma_estimates\n #self.F_ij = F_ij",
"_____no_output_____"
],
[
"reg_lgd_st_1 = LogisticRegression_with_p_values()\n# We create an instance of an object from the 'LogisticRegression' class.\nreg_lgd_st_1.fit(lgd_inputs_stage_1_train, lgd_targets_stage_1_train)\n# Estimates the coefficients of the object from the 'LogisticRegression' class\n# with inputs (independent variables) contained in the first dataframe\n# and targets (dependent variables) contained in the second dataframe.",
"_____no_output_____"
],
[
"feature_name = lgd_inputs_stage_1_train.columns.values\n# Stores the names of the columns of a dataframe in a variable.",
"_____no_output_____"
],
[
"summary_table = pd.DataFrame(columns = ['Feature name'], data = feature_name)\n# Creates a dataframe with a column titled 'Feature name' and row values contained in the 'feature_name' variable.\nsummary_table['Coefficients'] = np.transpose(reg_lgd_st_1.coef_)\n# Creates a new column in the dataframe, called 'Coefficients',\n# with row values the transposed coefficients from the 'LogisticRegression' object.\nsummary_table.index = summary_table.index + 1\n# Increases the index of every row of the dataframe with 1.\nsummary_table.loc[0] = ['Intercept', reg_lgd_st_1.intercept_[0]]\n# Assigns values of the row with index 0 of the dataframe.\nsummary_table = summary_table.sort_index()\n# Sorts the dataframe by index.\np_values = reg_lgd_st_1.p_values\n# We take the result of the newly added method 'p_values' and store it in a variable 'p_values'.\np_values = np.append(np.nan,np.array(p_values))\n# We add the value 'NaN' in the beginning of the variable with p-values.\nsummary_table['p_values'] = p_values\n# In the 'summary_table' dataframe, we add a new column, called 'p_values', containing the values from the 'p_values' variable.\nsummary_table",
"_____no_output_____"
],
[
"summary_table = pd.DataFrame(columns = ['Feature name'], data = feature_name)\nsummary_table['Coefficients'] = np.transpose(reg_lgd_st_1.coef_)\nsummary_table.index = summary_table.index + 1\nsummary_table.loc[0] = ['Intercept', reg_lgd_st_1.intercept_[0]]\nsummary_table = summary_table.sort_index()\np_values = reg_lgd_st_1.p_values\np_values = np.append(np.nan,np.array(p_values))\nsummary_table['p_values'] = p_values\nsummary_table",
"_____no_output_____"
]
],
[
[
"### Testing the Model",
"_____no_output_____"
]
],
[
[
"lgd_inputs_stage_1_test = lgd_inputs_stage_1_test[features_all]\n# Here we keep only the variables we need for the model.",
"_____no_output_____"
],
[
"lgd_inputs_stage_1_test = lgd_inputs_stage_1_test.drop(features_reference_cat, axis = 1)\n# Here we remove the dummy variable reference categories.",
"_____no_output_____"
],
[
"y_hat_test_lgd_stage_1 = reg_lgd_st_1.model.predict(lgd_inputs_stage_1_test)\n# Calculates the predicted values for the dependent variable (targets)\n# based on the values of the independent variables (inputs) supplied as an argument.",
"_____no_output_____"
],
[
"y_hat_test_lgd_stage_1",
"_____no_output_____"
],
[
"y_hat_test_proba_lgd_stage_1 = reg_lgd_st_1.model.predict_proba(lgd_inputs_stage_1_test)\n# Calculates the predicted probability values for the dependent variable (targets)\n# based on the values of the independent variables (inputs) supplied as an argument.",
"_____no_output_____"
],
[
"y_hat_test_proba_lgd_stage_1\n# This is an array of arrays of predicted class probabilities for all classes.\n# In this case, the first value of every sub-array is the probability for the observation to belong to the first class, i.e. 0,\n# and the second value is the probability for the observation to belong to the first class, i.e. 1.",
"_____no_output_____"
],
[
"y_hat_test_proba_lgd_stage_1 = y_hat_test_proba_lgd_stage_1[: ][: , 1]\n# Here we take all the arrays in the array, and from each array, we take all rows, and only the element with index 1,\n# that is, the second element.\n# In other words, we take only the probabilities for being 1.",
"_____no_output_____"
],
[
"y_hat_test_proba_lgd_stage_1",
"_____no_output_____"
],
[
"lgd_targets_stage_1_test_temp = lgd_targets_stage_1_test",
"_____no_output_____"
],
[
"lgd_targets_stage_1_test_temp.reset_index(drop = True, inplace = True)\n# We reset the index of a dataframe.",
"_____no_output_____"
],
[
"df_actual_predicted_probs = pd.concat([lgd_targets_stage_1_test_temp, pd.DataFrame(y_hat_test_proba_lgd_stage_1)], axis = 1)\n# Concatenates two dataframes.",
"_____no_output_____"
],
[
"df_actual_predicted_probs.columns = ['lgd_targets_stage_1_test', 'y_hat_test_proba_lgd_stage_1']",
"_____no_output_____"
],
[
"df_actual_predicted_probs.index = lgd_inputs_stage_1_test.index\n# Makes the index of one dataframe equal to the index of another dataframe.",
"_____no_output_____"
],
[
"df_actual_predicted_probs.head()",
"_____no_output_____"
]
],
[
[
"### Estimating the Аccuracy of the Мodel",
"_____no_output_____"
]
],
[
[
"tr = 0.5\n# We create a new column with an indicator,\n# where every observation that has predicted probability greater than the threshold has a value of 1,\n# and every observation that has predicted probability lower than the threshold has a value of 0.\ndf_actual_predicted_probs['y_hat_test_lgd_stage_1'] = np.where(df_actual_predicted_probs['y_hat_test_proba_lgd_stage_1'] > tr, 1, 0)",
"_____no_output_____"
],
[
"pd.crosstab(df_actual_predicted_probs['lgd_targets_stage_1_test'], df_actual_predicted_probs['y_hat_test_lgd_stage_1'], rownames = ['Actual'], colnames = ['Predicted'])\n# Creates a cross-table where the actual values are displayed by rows and the predicted values by columns.\n# This table is known as a Confusion Matrix.",
"_____no_output_____"
],
[
"pd.crosstab(df_actual_predicted_probs['lgd_targets_stage_1_test'], df_actual_predicted_probs['y_hat_test_lgd_stage_1'], rownames = ['Actual'], colnames = ['Predicted']) / df_actual_predicted_probs.shape[0]\n# Here we divide each value of the table by the total number of observations,\n# thus getting percentages, or, rates.",
"_____no_output_____"
],
[
"(pd.crosstab(df_actual_predicted_probs['lgd_targets_stage_1_test'], df_actual_predicted_probs['y_hat_test_lgd_stage_1'], rownames = ['Actual'], colnames = ['Predicted']) / df_actual_predicted_probs.shape[0]).iloc[0, 0] + (pd.crosstab(df_actual_predicted_probs['lgd_targets_stage_1_test'], df_actual_predicted_probs['y_hat_test_lgd_stage_1'], rownames = ['Actual'], colnames = ['Predicted']) / df_actual_predicted_probs.shape[0]).iloc[1, 1]\n# Here we calculate Accuracy of the model, which is the sum of the diagonal rates.",
"_____no_output_____"
],
[
"from sklearn.metrics import roc_curve, roc_auc_score",
"_____no_output_____"
],
[
"fpr, tpr, thresholds = roc_curve(df_actual_predicted_probs['lgd_targets_stage_1_test'], df_actual_predicted_probs['y_hat_test_proba_lgd_stage_1'])\n# Returns the Receiver Operating Characteristic (ROC) Curve from a set of actual values and their predicted probabilities.\n# As a result, we get three arrays: the false positive rates, the true positive rates, and the thresholds.\n# we store each of the three arrays in a separate variable.",
"_____no_output_____"
],
[
"plt.plot(fpr, tpr)\n# We plot the false positive rate along the x-axis and the true positive rate along the y-axis,\n# thus plotting the ROC curve.\nplt.plot(fpr, fpr, linestyle = '--', color = 'k')\n# We plot a seconary diagonal line, with dashed line style and black color.\nplt.xlabel('False positive rate')\n# We name the x-axis \"False positive rate\".\nplt.ylabel('True positive rate')\n# We name the x-axis \"True positive rate\".\nplt.title('ROC curve')\n# We name the graph \"ROC curve\".",
"_____no_output_____"
],
[
"AUROC = roc_auc_score(df_actual_predicted_probs['lgd_targets_stage_1_test'], df_actual_predicted_probs['y_hat_test_proba_lgd_stage_1'])\n# Calculates the Area Under the Receiver Operating Characteristic Curve (AUROC)\n# from a set of actual values and their predicted probabilities.\nAUROC",
"_____no_output_____"
]
],
[
[
"### Saving the Model",
"_____no_output_____"
]
],
[
[
"import pickle",
"_____no_output_____"
],
[
"pickle.dump(reg_lgd_st_1, open('lgd_model_stage_1.sav', 'wb'))\n# Here we export our model to a 'SAV' file with file name 'lgd_model_stage_1.sav'.",
"_____no_output_____"
]
],
[
[
"### Stage 2 – Linear Regression",
"_____no_output_____"
]
],
[
[
"lgd_stage_2_data = loan_data_defaults[loan_data_defaults['recovery_rate_0_1'] == 1]\n# Here we take only rows where the original recovery rate variable is greater than one,\n# i.e. where the indicator variable we created is equal to 1.",
"_____no_output_____"
],
[
"# LGD model stage 2 datasets: how much more than 0 is the recovery rate\nlgd_inputs_stage_2_train, lgd_inputs_stage_2_test, lgd_targets_stage_2_train, lgd_targets_stage_2_test = train_test_split(lgd_stage_2_data.drop(['good_bad', 'recovery_rate','recovery_rate_0_1', 'CCF'], axis = 1), lgd_stage_2_data['recovery_rate'], test_size = 0.2, random_state = 42)\n# Takes a set of inputs and a set of targets as arguments. Splits the inputs and the targets into four dataframes:\n# Inputs - Train, Inputs - Test, Targets - Train, Targets - Test.",
"_____no_output_____"
],
[
"from sklearn import linear_model\nfrom sklearn.metrics import mean_squared_error, r2_score",
"_____no_output_____"
],
[
"# Since the p-values are obtained through certain statistics, we need the 'stat' module from scipy.stats\nimport scipy.stats as stat\n\n# Since we are using an object oriented language such as Python, we can simply define our own \n# LinearRegression class (the same one from sklearn)\n# By typing the code below we will ovewrite a part of the class with one that includes p-values\n# Here's the full source code of the ORIGINAL class: https://github.com/scikit-learn/scikit-learn/blob/7b136e9/sklearn/linear_model/base.py#L362\n\n\nclass LinearRegression(linear_model.LinearRegression):\n \"\"\"\n LinearRegression class after sklearn's, but calculate t-statistics\n and p-values for model coefficients (betas).\n Additional attributes available after .fit()\n are `t` and `p` which are of the shape (y.shape[1], X.shape[1])\n which is (n_features, n_coefs)\n This class sets the intercept to 0 by default, since usually we include it\n in X.\n \"\"\"\n \n # nothing changes in __init__\n def __init__(self, fit_intercept=True, normalize=False, copy_X=True,\n n_jobs=1):\n self.fit_intercept = fit_intercept\n self.normalize = normalize\n self.copy_X = copy_X\n self.n_jobs = n_jobs\n\n \n def fit(self, X, y, n_jobs=1):\n self = super(LinearRegression, self).fit(X, y, n_jobs)\n \n # Calculate SSE (sum of squared errors)\n # and SE (standard error)\n sse = np.sum((self.predict(X) - y) ** 2, axis=0) / float(X.shape[0] - X.shape[1])\n se = np.array([np.sqrt(np.diagonal(sse * np.linalg.inv(np.dot(X.T, X))))])\n\n # compute the t-statistic for each feature\n self.t = self.coef_ / se\n # find the p-value for each feature\n self.p = np.squeeze(2 * (1 - stat.t.cdf(np.abs(self.t), y.shape[0] - X.shape[1])))\n return self",
"_____no_output_____"
],
[
"import scipy.stats as stat\n\nclass LinearRegression(linear_model.LinearRegression):\n def __init__(self, fit_intercept=True, normalize=False, copy_X=True,\n n_jobs=1):\n self.fit_intercept = fit_intercept\n self.normalize = normalize\n self.copy_X = copy_X\n self.n_jobs = n_jobs\n def fit(self, X, y, n_jobs=1):\n self = super(LinearRegression, self).fit(X, y, n_jobs)\n sse = np.sum((self.predict(X) - y) ** 2, axis=0) / float(X.shape[0] - X.shape[1])\n se = np.array([np.sqrt(np.diagonal(sse * np.linalg.inv(np.dot(X.T, X))))])\n self.t = self.coef_ / se\n self.p = np.squeeze(2 * (1 - stat.t.cdf(np.abs(self.t), y.shape[0] - X.shape[1])))\n return self",
"_____no_output_____"
],
[
"lgd_inputs_stage_2_train = lgd_inputs_stage_2_train[features_all]\n# Here we keep only the variables we need for the model.",
"_____no_output_____"
],
[
"lgd_inputs_stage_2_train = lgd_inputs_stage_2_train.drop(features_reference_cat, axis = 1)\n# Here we remove the dummy variable reference categories.",
"_____no_output_____"
],
[
"reg_lgd_st_2 = LinearRegression()\n# We create an instance of an object from the 'LogisticRegression' class.\nreg_lgd_st_2.fit(lgd_inputs_stage_2_train, lgd_targets_stage_2_train)\n# Estimates the coefficients of the object from the 'LogisticRegression' class\n# with inputs (independent variables) contained in the first dataframe\n# and targets (dependent variables) contained in the second dataframe.",
"_____no_output_____"
],
[
"feature_name = lgd_inputs_stage_2_train.columns.values\n# Stores the names of the columns of a dataframe in a variable.",
"_____no_output_____"
],
[
"summary_table = pd.DataFrame(columns = ['Feature name'], data = feature_name)\n# Creates a dataframe with a column titled 'Feature name' and row values contained in the 'feature_name' variable.\nsummary_table['Coefficients'] = np.transpose(reg_lgd_st_2.coef_)\n# Creates a new column in the dataframe, called 'Coefficients',\n# with row values the transposed coefficients from the 'LogisticRegression' object.\nsummary_table.index = summary_table.index + 1\n# Increases the index of every row of the dataframe with 1.\nsummary_table.loc[0] = ['Intercept', reg_lgd_st_2.intercept_]\n# Assigns values of the row with index 0 of the dataframe.\nsummary_table = summary_table.sort_index()\n# Sorts the dataframe by index.\np_values = reg_lgd_st_2.p\n# We take the result of the newly added method 'p_values' and store it in a variable 'p_values'.\np_values = np.append(np.nan,np.array(p_values))\n# We add the value 'NaN' in the beginning of the variable with p-values.\nsummary_table['p_values'] = p_values.round(3)\n# In the 'summary_table' dataframe, we add a new column, called 'p_values', containing the values from the 'p_values' variable.\nsummary_table",
"_____no_output_____"
],
[
"summary_table = pd.DataFrame(columns = ['Feature name'], data = feature_name)\nsummary_table['Coefficients'] = np.transpose(reg_lgd_st_2.coef_)\nsummary_table.index = summary_table.index + 1\nsummary_table.loc[0] = ['Intercept', reg_lgd_st_2.intercept_]\nsummary_table = summary_table.sort_index()\np_values = reg_lgd_st_2.p\np_values = np.append(np.nan,np.array(p_values))\nsummary_table['p_values'] = p_values.round(3)\nsummary_table",
"_____no_output_____"
]
],
[
[
"### Stage 2 – Linear Regression Evaluation",
"_____no_output_____"
]
],
[
[
"lgd_inputs_stage_2_test = lgd_inputs_stage_2_test[features_all]\n# Here we keep only the variables we need for the model.",
"_____no_output_____"
],
[
"lgd_inputs_stage_2_test = lgd_inputs_stage_2_test.drop(features_reference_cat, axis = 1)\n# Here we remove the dummy variable reference categories.",
"_____no_output_____"
],
[
"lgd_inputs_stage_2_test.columns.values\n# Calculates the predicted values for the dependent variable (targets)\n# based on the values of the independent variables (inputs) supplied as an argument.",
"_____no_output_____"
],
[
"y_hat_test_lgd_stage_2 = reg_lgd_st_2.predict(lgd_inputs_stage_2_test)\n# Calculates the predicted values for the dependent variable (targets)\n# based on the values of the independent variables (inputs) supplied as an argument.",
"_____no_output_____"
],
[
"lgd_targets_stage_2_test_temp = lgd_targets_stage_2_test",
"_____no_output_____"
],
[
"lgd_targets_stage_2_test_temp = lgd_targets_stage_2_test_temp.reset_index(drop = True)\n# We reset the index of a dataframe.",
"_____no_output_____"
],
[
"pd.concat([lgd_targets_stage_2_test_temp, pd.DataFrame(y_hat_test_lgd_stage_2)], axis = 1).corr()\n# We calculate the correlation between actual and predicted values.",
"_____no_output_____"
],
[
"sns.distplot(lgd_targets_stage_2_test - y_hat_test_lgd_stage_2)\n# We plot the distribution of the residuals.",
"_____no_output_____"
],
[
"pickle.dump(reg_lgd_st_2, open('lgd_model_stage_2.sav', 'wb'))\n# Here we export our model to a 'SAV' file with file name 'lgd_model_stage_1.sav'.",
"_____no_output_____"
]
],
[
[
"### Combining Stage 1 and Stage 2",
"_____no_output_____"
]
],
[
[
"y_hat_test_lgd_stage_2_all = reg_lgd_st_2.predict(lgd_inputs_stage_1_test)",
"_____no_output_____"
],
[
"y_hat_test_lgd_stage_2_all",
"_____no_output_____"
],
[
"y_hat_test_lgd = y_hat_test_lgd_stage_1 * y_hat_test_lgd_stage_2_all\n# Here we combine the predictions of the models from the two stages.",
"_____no_output_____"
],
[
"pd.DataFrame(y_hat_test_lgd).describe()\n# Shows some descriptive statisics for the values of a column.",
"_____no_output_____"
],
[
"y_hat_test_lgd = np.where(y_hat_test_lgd < 0, 0, y_hat_test_lgd)\ny_hat_test_lgd = np.where(y_hat_test_lgd > 1, 1, y_hat_test_lgd)\n# We set predicted values that are greater than 1 to 1 and predicted values that are less than 0 to 0.",
"_____no_output_____"
],
[
"pd.DataFrame(y_hat_test_lgd).describe()\n# Shows some descriptive statisics for the values of a column.",
"_____no_output_____"
]
],
[
[
"# EAD Model",
"_____no_output_____"
],
[
"### Estimation and Interpretation",
"_____no_output_____"
]
],
[
[
"# EAD model datasets\nead_inputs_train, ead_inputs_test, ead_targets_train, ead_targets_test = train_test_split(loan_data_defaults.drop(['good_bad', 'recovery_rate','recovery_rate_0_1', 'CCF'], axis = 1), loan_data_defaults['CCF'], test_size = 0.2, random_state = 42)\n# Takes a set of inputs and a set of targets as arguments. Splits the inputs and the targets into four dataframes:\n# Inputs - Train, Inputs - Test, Targets - Train, Targets - Test.",
"_____no_output_____"
],
[
"ead_inputs_train.columns.values",
"_____no_output_____"
],
[
"ead_inputs_train = ead_inputs_train[features_all]\n# Here we keep only the variables we need for the model.",
"_____no_output_____"
],
[
"ead_inputs_train = ead_inputs_train.drop(features_reference_cat, axis = 1)\n# Here we remove the dummy variable reference categories.",
"_____no_output_____"
],
[
"reg_ead = LinearRegression()\n# We create an instance of an object from the 'LogisticRegression' class.\nreg_ead.fit(ead_inputs_train, ead_targets_train)\n# Estimates the coefficients of the object from the 'LogisticRegression' class\n# with inputs (independent variables) contained in the first dataframe\n# and targets (dependent variables) contained in the second dataframe.",
"_____no_output_____"
],
[
"feature_name = ead_inputs_train.columns.values",
"_____no_output_____"
],
[
"summary_table = pd.DataFrame(columns = ['Feature name'], data = feature_name)\n# Creates a dataframe with a column titled 'Feature name' and row values contained in the 'feature_name' variable.\nsummary_table['Coefficients'] = np.transpose(reg_ead.coef_)\n# Creates a new column in the dataframe, called 'Coefficients',\n# with row values the transposed coefficients from the 'LogisticRegression' object.\nsummary_table.index = summary_table.index + 1\n# Increases the index of every row of the dataframe with 1.\nsummary_table.loc[0] = ['Intercept', reg_ead.intercept_]\n# Assigns values of the row with index 0 of the dataframe.\nsummary_table = summary_table.sort_index()\n# Sorts the dataframe by index.\np_values = reg_ead.p\n# We take the result of the newly added method 'p_values' and store it in a variable 'p_values'.\np_values = np.append(np.nan,np.array(p_values))\n# We add the value 'NaN' in the beginning of the variable with p-values.\nsummary_table['p_values'] = p_values\n# In the 'summary_table' dataframe, we add a new column, called 'p_values', containing the values from the 'p_values' variable.\nsummary_table",
"_____no_output_____"
]
]
] | [
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] | [
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
d0ff1bb01e3e9d4d5ab04fe15fd39602ff5333c9 | 754,224 | ipynb | Jupyter Notebook | examples/.ipynb_checkpoints/maryetal_visu-checkpoint.ipynb | BenjMy/pycathy_wrapper | 8639713acbb5bc5c5cd293b94bdafe017d03f199 | [
"BSD-3-Clause"
] | null | null | null | examples/.ipynb_checkpoints/maryetal_visu-checkpoint.ipynb | BenjMy/pycathy_wrapper | 8639713acbb5bc5c5cd293b94bdafe017d03f199 | [
"BSD-3-Clause"
] | 8 | 2021-10-14T13:07:46.000Z | 2022-03-02T03:32:05.000Z | examples/.ipynb_checkpoints/maryetal_visu-checkpoint.ipynb | BenjMy/pycathy_wrapper | 8639713acbb5bc5c5cd293b94bdafe017d03f199 | [
"BSD-3-Clause"
] | null | null | null | 86.28578 | 261,224 | 0.613547 | [
[
[
"# Rhizotron lab experiment",
"_____no_output_____"
]
],
[
[
"# We first need to load the pyCATHY package\n# pyCATHY is calling fortran codes via python subroutines and ease the process\n#!pip install -i https://test.pypi.org/simple/ pycathy-v001==0.0.1\n!pip install git+https://github.com/BenjMy/pycathy_wrapper.git\n#!git clone https://github.com/BenjMy/pycathy_wrapper.git",
"Collecting git+https://github.com/BenjMy/pycathy_wrapper.git\n Cloning https://github.com/BenjMy/pycathy_wrapper.git to /tmp/pip-req-build-4rkp75hc\n Running command git clone -q https://github.com/BenjMy/pycathy_wrapper.git /tmp/pip-req-build-4rkp75hc\n Resolved https://github.com/BenjMy/pycathy_wrapper.git to commit bf26543e2781b210e368d4d0aae535f79874e800\n Installing build dependencies ... \u001b[?25ldone\n\u001b[?25h Getting requirements to build wheel ... \u001b[?25ldone\n\u001b[?25h Preparing wheel metadata ... \u001b[?25ldone\n\u001b[?25h"
],
[
"import os\nimport numpy as np\nfrom pyCATHY import cathy_tools \nfrom pyCATHY import plot_tools as cplt \nfrom pyCATHY import rhizo_tools\n\n#import cathy_tools as CATHY\nos.getcwd()",
"_____no_output_____"
]
],
[
[
"We initiate a CATHY object; if the the CATHY src files are not included within the 'path2prj', they are automatically fetched from the gitbucket (If the notebook is initiate locally with an internet connection)",
"_____no_output_____"
]
],
[
[
"path2prj ='notebooks'\nprj = 'rhizo_prj2'\n#os.chdir(path2prj)\nsimu = cathy_tools.CATHY(dirName=path2prj,prjName=prj,notebook=True)\nsimu # show the CATHY object",
"init CATHY object\n"
],
[
"simu.workdir",
"_____no_output_____"
]
],
[
[
"## Rhizotron DEM",
"_____no_output_____"
],
[
"### Build the grid 3d with arbitrary values",
"_____no_output_____"
],
[
"### Run preprocessor",
"_____no_output_____"
],
[
"## Simulation inputs",
"_____no_output_____"
],
[
"### 1- Initial conditions",
"_____no_output_____"
],
[
"### 2- Irrigation schedule",
"_____no_output_____"
],
[
"### 3- Boundary conditions",
"_____no_output_____"
],
[
"### 4- Soil and roots inputs",
"_____no_output_____"
],
[
"### run processor ",
"_____no_output_____"
],
[
"## Explore outputs",
"_____no_output_____"
]
],
[
[
"step = 1",
"_____no_output_____"
],
[
"cplt.showvtk(unit='pressure',timeStep=1,notebook=True, path='./' + prj + '/vtk/')\ncplt.showvtk(unit='pressure',timeStep=2,notebook=True, path='./' + prj + '/vtk/')\ncplt.showvtk(unit='pressure',timeStep=3,notebook=True, path='./' + prj + '/vtk/')\ncplt.showvtk(unit='pressure',timeStep=4,notebook=True, path='./' + prj + '/vtk/')\n\ncplt.showvtk(unit='saturation',timeStep=1,notebook=True, path='./' + prj + '/vtk/')\ncplt.showvtk(unit='saturation',timeStep=2,notebook=True, path='./' + prj + '/vtk/')\ncplt.showvtk(unit='saturation',timeStep=3,notebook=True, path='./' + prj + '/vtk/')\ncplt.showvtk(unit='saturation',timeStep=4,notebook=True, path='./' + prj + '/vtk/')\n",
"_____no_output_____"
],
[
"import glob\nimport subprocess",
"_____no_output_____"
]
],
[
[
"We run the preprocessor with the default files inputs (weill et al example). \nThis step create all the prepro files required for the processor later on.",
"_____no_output_____"
]
],
[
[
"simu.run_preprocessor(verbose=True)",
"0\n1\nrun preprocessor\n\n wbb...\n\n searching the dtm_13.val input file...\n assigned nodata value = -9999.0000000000000 \n\n number of processed cells = 400\n\n ...wbb completed\n\n rn...\n csort I...\n ...completed\n\n depit...\n dem modifications = 0\n dem modifications = 0 (total)\n ...completed\n\n csort II...\n ...completed\n\n cca...\n\n contour curvature threshold value = 9.99999996E+11\n ...completed\n\n smean...\n mean (min,max) facet slope = 0.052056253 ( 0.020000000, 0.053851648)\n ...completed\n\n dsf...\n the drainage direction of the outlet cell ( 8 ) is used\n ...completed\n\n hg...\n ...completed\n\n saving the data in the basin_b/basin_i files...\n\n ...rn completed\n\n mrbb...\n\n\n Select the header type:\n 0) None\n 1) ESRI ascii file\n 2) GRASS ascii file\n (Ctrl C to exit)\n\n -> \n Select the nodata value:\n (Ctrl C to exit)\n\n -> \n Select the pointer system:\n 1) HAP system\n 2) Arc/Gis system\n (Ctrl C to exit)\n\n -> ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~\n\n dem file\n\n min value = 0.335000E+00\n max value = 0.100000E+01\n number of cells = 400\n mean value = 0.667500E+00\n\n writing the output file...\n\n ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~\n\n lakes_map file\n\n min value = 0\n max value = 0\n number of cells = 400\n mean value = 0.000000\n\n writing the output file...\n\n ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~\n\n zone file\n\n min value = 1\n max value = 1\n number of cells = 400\n mean value = 1.000000\n\n writing the output file...\n\n ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~\n\n dtm_w_1 file\n\n min value = 0.515524E+00\n max value = 0.100000E+01\n number of cells = 400\n mean value = 0.607575E+00\n\n writing the output file...\n\n ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~\n\n dtm_w_2 file\n\n min value = 0.000000E+00\n max value = 0.484476E+00\n number of cells = 400\n mean value = 0.392425E+00\n\n writing the output file...\n\n ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~\n\n dtm_p_outflow_1 file\n\n min value = 4\n max value = 8\n number of cells = 400\n mean value = 4.200000\n\n writing the output file...\n\n ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~\n\n dtm_p_outflow_2 file\n\n min value = 0\n max value = 9\n number of cells = 400\n mean value = 6.792500\n\n writing the output file...\n\n ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~\n\n A_inflow file\n\n min value = 0.000000000000E+00\n max value = 0.997499787031E+02\n number of cells = 400\n mean value = 0.388447785378E+01\n\n writing the output file...\n\n ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~\n\n dtm_local_slope_1 file\n\n min value = 0.200000E-01\n max value = 0.500000E-01\n number of cells = 400\n mean value = 0.485000E-01\n\n writing the output file...\n\n ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~\n\n dtm_local_slope_2 file\n\n min value = 0.000000E+00\n max value = 0.494975E-01\n number of cells = 400\n mean value = 0.400930E-01\n\n writing the output file...\n\n ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~\n\n dtm_epl_1 file\n\n min value = 0.500000E+00\n max value = 0.500000E+00\n number of cells = 400\n mean value = 0.500000E+00\n\n writing the output file...\n\n ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~\n\n dtm_epl_2 file\n\n min value = 0.000000E+00\n max value = 0.707107E+00\n number of cells = 400\n mean value = 0.572757E+00\n\n writing the output file...\n\n ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~\n\n dtm_kSs1_sf_1 file\n\n min value = 0.240040E+02\n max value = 0.240040E+02\n number of cells = 400\n mean value = 0.240040E+02\n\n writing the output file...\n\n ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~\n\n dtm_kSs1_sf_2 file\n\n min value = 0.000000E+00\n max value = 0.240040E+02\n number of cells = 400\n mean value = 0.194432E+02\n\n writing the output file...\n\n ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~\n\n dtm_Ws1_sf file\n\n min value = 0.100000E+01\n max value = 0.100000E+01\n number of cells = 400\n mean value = 0.100000E+01\n\n writing the output file...\n\n ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~\n\n dtm_Ws1_sf_2 file\n\n min value = 0.000000E+00\n max value = 0.100000E+01\n number of cells = 400\n mean value = 0.810000E+00\n\n writing the output file...\n\n ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~\n\n dtm_b1_sf file\n\n min value = 0.000000E+00\n max value = 0.000000E+00\n number of cells = 400\n mean value = 0.000000E+00\n\n writing the output file...\n\n ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~\n\n dtm_y1_sf file\n\n min value = 0.000000E+00\n max value = 0.000000E+00\n number of cells = 400\n mean value = 0.000000E+00\n\n writing the output file...\n\n ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~\n\n dtm_hcID file\n\n min value = 0\n max value = 0\n number of cells = 400\n mean value = 0.000000\n\n writing the output file...\n\n ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~\n\n dtm_q_output file\n\n min value = 0\n max value = 0\n number of cells = 400\n mean value = 0.000000\n\n writing the output file...\n\n ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~\n\n dtm_nrc file\n\n min value = 0.100000E+01\n max value = 0.100000E+01\n number of cells = 400\n mean value = 0.100000E+01\n\n writing the output file...\n\n ...mrbb completed\n\n bb2shp...\n\n writing file river_net.shp\n\n\n"
],
[
"# survey.hap_in # hap.in parameters are saved in a dict\n# survey.hap_in['Ncells'] # access and check the number of cell from the CATHY object",
"_____no_output_____"
]
],
[
[
"We run the processor with the default files inputs (weill et al example)",
"_____no_output_____"
]
],
[
[
"simu.run_processor(verbose=True)",
"recompile\n"
]
],
[
[
"Plot results",
"_____no_output_____"
]
],
[
[
"cplt.showvtk(unit='pressure',timeStep=1,notebook=True, path='./' + prj + '/vtk/')\n",
"plot pressure\n"
]
]
] | [
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] | [
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
]
] |
d0ff1f8189e174fc287eaac10fb2d04e3ff07222 | 12,612 | ipynb | Jupyter Notebook | Day5LoopingOverTasks.ipynb | pvanheus/python-novice-gapminder | 2c33906fe927e77784cff64761f6bcd8c0e9ded0 | [
"CC-BY-4.0"
] | null | null | null | Day5LoopingOverTasks.ipynb | pvanheus/python-novice-gapminder | 2c33906fe927e77784cff64761f6bcd8c0e9ded0 | [
"CC-BY-4.0"
] | null | null | null | Day5LoopingOverTasks.ipynb | pvanheus/python-novice-gapminder | 2c33906fe927e77784cff64761f6bcd8c0e9ded0 | [
"CC-BY-4.0"
] | 1 | 2018-09-28T12:00:54.000Z | 2018-09-28T12:00:54.000Z | 19.462963 | 278 | 0.480019 | [
[
[
"import pandas",
"_____no_output_____"
],
[
"mydata = pandas.read_csv('data/gapminder_gdp_africa.csv', index_col='country')",
"_____no_output_____"
],
[
"mydata.ix[:,'gdpPercap_1952'].mean()",
"_____no_output_____"
],
[
"%%bash\nls data/",
"asia_gdp_per_capita.csv\ngapminder_all.csv\ngapminder_gdp_africa.csv\ngapminder_gdp_americas.csv\ngapminder_gdp_asia.csv\ngapminder_gdp_europe.csv\ngapminder_gdp_oceania.csv\n"
],
[
"continents = ['africa', 'asia', 'americas', 'oceania', 'europe']\nfor continent in continents:\n filename = 'data/gapminder_gdp_' + continent + '.csv'\n mydata = pandas.read_csv(filename, index_col='country')\n mean = mydata.ix[:,'gdpPercap_1952'].mean()\n print(continent, round(mean, 2))",
"africa 1252.57\nasia 5195.48\namericas 4079.06\noceania 10298.09\neurope 5661.06\n"
],
[
"round(1252.5724658211536, 2)",
"_____no_output_____"
],
[
"help(round)",
"Help on built-in function round in module builtins:\n\nround(...)\n round(number[, ndigits]) -> number\n \n Round a number to a given precision in decimal digits (default 0 digits).\n This returns an int when called with one argument, otherwise the\n same type as the number. ndigits may be negative.\n\n"
],
[
"islow = 1252.57 < 2000",
"_____no_output_____"
],
[
"print(islow)",
"True\n"
],
[
"islow = 5661.06 < 2000",
"_____no_output_____"
],
[
"print(islow)",
"False\n"
]
],
[
[
"Comparison operators:\n \n < # less than\n > # greater than\n <= # less than or equal\n >= # greater than or equal\n == # equal - this is **not** the same as the = (assignment) operator\n != # not equal",
"_____no_output_____"
]
],
[
[
"1 != 2",
"_____no_output_____"
],
[
"1 = 2",
"_____no_output_____"
],
[
"1 == 2",
"_____no_output_____"
],
[
"number = 5",
"_____no_output_____"
],
[
"number < 10 and number > 3",
"_____no_output_____"
],
[
"3 < number < 10 < 12 > 0",
"_____no_output_____"
],
[
"number < 10 and > 3",
"_____no_output_____"
],
[
"number < 10 or number > 3",
"_____no_output_____"
],
[
"not True",
"_____no_output_____"
],
[
"print(number)\nnot (number < 3 and number > 10)",
"5\n"
]
],
[
[
"The logical operators\n\n and # True if both sides are True\n or # True if either side is true\n not # flip True to False or False to True",
"_____no_output_____"
]
],
[
[
"sillyval = print(\"returns None\")\nprint(sillyval is None)\nprint(sillyval is not None)",
"returns None\nTrue\nFalse\n"
],
[
"number = 1\nif number > 3:\n print(\"the number is > 3\")",
"_____no_output_____"
],
[
"number = 5\nif number > 0:\n print(\"the number is positive\")\nelse:\n print(\"the number is not positive\")",
"the number is positive\n"
],
[
"number = 5\nif number > 0:\n print(\"the number is positive\")\nelif number < 0:\n print(\"the number is negative\")\nelse:\n print(\"the number is zero\")",
"the number is positive\n"
],
[
"number = -1\nif number > 0:\n print(\"the number is positive\")\nelif number < 0:\n print(\"the number is negative\")\nelse:\n print(\"the number is zero\")",
"the number is negative\n"
],
[
"number = 13\nif number > 0:\n print(\"the number is positive\")\n if number >= 10:\n print(\"the number has more than 1 digit\")\n",
"the number is positive\nthe number has more than 1 digit\n"
],
[
"continents = ['africa', 'asia', 'americas', 'oceania', 'europe']\nfor continent in continents:\n filename = 'data/gapminder_gdp_' + continent + '.csv'\n mydata = pandas.read_csv(filename, index_col='country')\n mean = mydata.ix[:,'gdpPercap_1952'].mean()\n if mean < 2000:\n print(continent, \"income is low:\", round(mean, 2))\n elif mean >= 2000 and mean <= 6000:\n print(continent, \"income is moderate:\", round(mean, 2))\n else:\n print(continent, \"income is high:\", round(mean, 2))\n# print(continent, round(mean, 2))",
"africa income is low: 1252.57\nasia income is moderate: 5195.48\namericas income is moderate: 4079.06\noceania income is high: 10298.09\neurope income is moderate: 5661.06\n"
]
]
] | [
"code",
"markdown",
"code",
"markdown",
"code"
] | [
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
d0ff2f30462d76158d00b4a12446974973bc9190 | 18,320 | ipynb | Jupyter Notebook | principal_component_analysis/principal_component_analysis.ipynb | kcmckee/reference_guides | 12ae7c5cd149feaa0c584b069d3d789784f78b62 | [
"MIT"
] | null | null | null | principal_component_analysis/principal_component_analysis.ipynb | kcmckee/reference_guides | 12ae7c5cd149feaa0c584b069d3d789784f78b62 | [
"MIT"
] | null | null | null | principal_component_analysis/principal_component_analysis.ipynb | kcmckee/reference_guides | 12ae7c5cd149feaa0c584b069d3d789784f78b62 | [
"MIT"
] | null | null | null | 29.885808 | 948 | 0.531714 | [
[
[
"# Principal Component Analysis (PCA) in Python #",
"_____no_output_____"
],
[
"Killian McKee",
"_____no_output_____"
],
[
"### Overview ###\n\n1. [What is PCA?](#section1)\n2. [Key Terms](#section2) \n3. [Pros and Cons of PCA](#section3)\n4. [When to use PCA](#section4)\n5. [Key Parameters](#section5)\n6. [Walkthrough: PCA for data visualization](#section6)\n7. [Walkthrough: PCA w/ Random Forest](#section7)\n7. [Additional Reading](#section8)\n8. [Conclusion](#section9)\n9. [Sources](#section10)",
"_____no_output_____"
],
[
"<a id='section1'></a>",
"_____no_output_____"
],
[
"### What is Principal Component Analysis? ###\n\nPrincipal component analysis is a non-parametric data science tool that allows us to identify the most important variables in a data set consisting of many correlated variables. In more technical terms, pca helps us reduce the dimensionality of our feature space by highlighting the most important variables (principal components) of a dataset via orth0gonalization. Pca is typically done before a model is built to decide which variables to include and to eliminate those which are overly correlated with one another. Principal component analysis provides two primary benefits; firstly, it can help our models avoid overfitting by eliminating extraneous variables that are most likely only pertinent (if at all) for our training data, but not the new data it would see in the real world. Secondly, performing pca can drastically improve model training speed in high dimensional data settings (when there are lots of features in a dataset). ",
"_____no_output_____"
],
[
"<a id='section2'></a>",
"_____no_output_____"
],
[
"### Key Terms ### \n\n1. **Dimensionality**: the number of features in a dataset (represented by more columns in a tidy dataset). Pca aims to reduce excessive dimensionality in a dataset to improve model performance. \n2. **Correlation**: A measure of closeness between two variables, ranging from -1 to +1. A negative correlation indicates that when one variable goes up, the other goes down (and a posistive correlation indicates they both move in the same direction). PCA helps us eliminate redundant correlated variables.\n3. **Orthagonal**: Uncorrelated to one another i.e. they have a correlation of 0. PCA seeks to find an orthgonalized subset of the data that still captures most/all of the important information for our model. \n4. **Covariance Matrix**: A matrix we can generate to show how correlated variables are with one another. This can be a helpul tool to visualize what features PCA may or may not eliminate. ",
"_____no_output_____"
],
[
"<a id='section3'></a>",
"_____no_output_____"
],
[
"### Pros and Cons of PCA ###\n\nThere are no real cons of PCA, but it does have some limitations: \n\n**Pros**: \n\n1. Reduces model noise\n2. Easy to implement with python packages like pandas and scikit-learn \n3. Improves model training time\n\n**Limitations**: \n\n1. Linearity: pca assumes the principle components are a linear combination of the original dataset features. \n2. Variance measure: pca uses variance as the measure of dimension importance. This can mean axes with high variance can be treated as principle components and those with low variance can be cut out as noise. \n3. Orthogonality: pca assumes the principle components are orthogonal, and won't produce meaningful results otherwise. ",
"_____no_output_____"
],
[
"<a id='section4'></a>",
"_____no_output_____"
],
[
"### When to use Principal Component Analysis ### \n\nOne should consider using PCA when the following conditions are true: \n\n1. The linearity, variance, and orthogonality limitations specified above are satisfied. \n2. Your dataset contains many features \n3. You are interested in reducing the noise of your dataset or improving model training time",
"_____no_output_____"
],
[
"<a id='section5'></a>",
"_____no_output_____"
],
[
"### Key Parameters ###\n\nThe number of features to keep post pca (typically denoted by n_components) is the only major parameter for PCA. ",
"_____no_output_____"
],
[
"<a id='section6'></a>",
"_____no_output_____"
],
[
"### PCA Walkthrough: Data Visualization ### \n\nWe will be modifying scikit-learn's tutorial on fitting PCA for visualization using the iris [dataset](https://en.wikipedia.org/wiki/Iris_flower_data_set) (contains different species of flowers). ",
"_____no_output_____"
]
],
[
[
"# import the necessary packages \n\nimport numpy as np\nimport matplotlib.pyplot as plt\nfrom mpl_toolkits.mplot3d import Axes3D\nfrom sklearn import decomposition\nfrom sklearn import datasets\n\n\n#specify graph parameters for iris and load the dataset \n\ncenters = [[1, 1], [-1, -1], [1, -1]]\niris = datasets.load_iris()\n\n\n# set features and target \n\nX = iris.data\ny = iris.target\n\n\n# create the chart \n\nfig = plt.figure(1, figsize=(4, 3))\nplt.clf()\nax = Axes3D(fig, rect=[0, 0, .95, 1], elev=48, azim=134)\n\n\n# fit our PCA \n\nplt.cla()\npca = decomposition.PCA(n_components=3)\npca.fit(X)\nX = pca.transform(X)\n\n\n# plot our data\n\nfor name, label in [('Setosa', 0), ('Versicolour', 1), ('Virginica', 2)]:\n ax.text3D(X[y == label, 0].mean(),\n X[y == label, 1].mean() + 1.5,\n X[y == label, 2].mean(), name,\n horizontalalignment='center',\n bbox=dict(alpha=.5, edgecolor='w', facecolor='w'))\n \n \n# Reorder the labels to have colors matching the cluster results\n\ny = np.choose(y, [1, 2, 0]).astype(np.float)\nax.scatter(X[:, 0], X[:, 1], X[:, 2], c=y, cmap=plt.cm.nipy_spectral,\n edgecolor='k')\n\nax.w_xaxis.set_ticklabels([])\nax.w_yaxis.set_ticklabels([])\nax.w_zaxis.set_ticklabels([])\n\nplt.show()\n\n#we can clearly see the three species within the iris dataset and how the differ from one another",
"_____no_output_____"
]
],
[
[
"<a id='section7'></a>",
"_____no_output_____"
],
[
"### Walkthrough: PCA w/ Random Forest ### \n\nIn this tutorial we will be walking through the typical workflow to improve model speed with PCA, then fitting a random forest. We will be working with the iris dataset again, but we will load it into a pandas dataframe ",
"_____no_output_____"
]
],
[
[
"# import necessary packages \n\nimport numpy as np \nimport pandas as pd \nfrom sklearn import datasets\nfrom sklearn.model_selection import train_test_split\nfrom sklearn.preprocessing import StandardScaler\nfrom sklearn.decomposition import PCA\nfrom sklearn.ensemble import RandomForestClassifier\nfrom sklearn.metrics import confusion_matrix \nfrom sklearn.metrics import accuracy_score\n",
"_____no_output_____"
],
[
"# download the data \n\ndata = datasets.load_iris()\ndf = pd.DataFrame(data['data'], columns=data['feature_names'])\ndf['target'] = data['target']\ndf.head()",
"_____no_output_____"
],
[
"# split the data into features and target \n\nX = df.drop('target', 1) \ny = df['target'] ",
"_____no_output_____"
],
[
"#creating training and test splits \n\nX_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=0) ",
"_____no_output_____"
],
[
"# scaling the data \n# since pca uses variance as a measure, it is best to scale the data \n\nsc = StandardScaler() \nX_train = sc.fit_transform(X_train) \nX_test = sc.transform(X_test) ",
"_____no_output_____"
],
[
"# apply and fit the pca \n# play around with the n_components value to see how the model does \n\npca = PCA(n_components=4) \nX_train = pca.fit_transform(X_train) \nX_test = pca.transform(X_test)",
"_____no_output_____"
],
[
"# generate the explained variance, which shows us how much variance is caused by each variable \n# we can see from the example below that more than 96% of the data can be explained by the first two principle components\n\nexplained_variance = pca.explained_variance_ratio_ \nexplained_variance",
"_____no_output_____"
],
[
"# now lets fit a random forest so we can see how the accuracy changes with different levels of components \n# this model has all the components \n\nclassifier = RandomForestClassifier(max_depth=2, random_state=0) \nclassifier.fit(X_train, y_train)\n\n# Predicting the Test set results\ny_pred = classifier.predict(X_test) ",
"_____no_output_____"
],
[
"# all component model accuracy \n# we can see it achieves an accuracy of 93% \n\ncm = confusion_matrix(y_test, y_pred) \nprint(cm) \nprint('Accuracy', accuracy_score(y_test, y_pred)) ",
"[[11 0 0]\n [ 0 10 3]\n [ 0 1 5]]\nAccuracy 0.8666666666666667\n"
],
[
"# Now lets see how the model does with only 2 components \n# our accuracy decreases by about 3%, but we can see how this might be useful if we had 100s of components\n\npca = PCA(n_components=2) \nX_train = pca.fit_transform(X_train) \nX_test = pca.transform(X_test) ",
"_____no_output_____"
],
[
"classifier = RandomForestClassifier(max_depth=2, random_state=0) \nclassifier.fit(X_train, y_train)\n\n# Predicting the Test set results\ny_pred = classifier.predict(X_test) ",
"_____no_output_____"
],
[
"cm = confusion_matrix(y_test, y_pred) \nprint(cm) \nprint('Accuracy', accuracy_score(y_test, y_pred)) ",
"[[11 0 0]\n [ 0 10 3]\n [ 0 2 4]]\nAccuracy 0.8333333333333334\n"
]
],
[
[
"<a id='section8'></a>",
"_____no_output_____"
],
[
"### Additional Reading ###\n\n1. Going into much greater depth on [PCA](https://www-bcf.usc.edu/~gareth/ISL/ISLR%20First%20Printing.pdf)\n2. Visualizing [PCA](http://setosa.io/ev/principal-component-analysis/)",
"_____no_output_____"
],
[
"<a id='section9'></a>",
"_____no_output_____"
],
[
"### Conclusion ### \n\nThis guide explained how principal component analysis helps reduce noise in our dataset and improve model speed via a simplified feature space. Next, we looked at some of the key components and limitations of PCA, namely the number of preserved components and the linearity, othogonality, and variance requirements, respectively. Lastly, we stepped through two examples of how to implement PCA; the first covered visualization, while the second tackled PCA as a preprocessing step with random forests. ",
"_____no_output_____"
],
[
"<a id='section10'></a>",
"_____no_output_____"
],
[
"### Sources ### \n\n1. https://arxiv.org/pdf/1404.1100.pdf?utm_campaign=buffer&utm_content=bufferb37df&utm_medium=social&utm_source=facebook.com \n2. https://stackabuse.com/implementing-pca-in-python-with-scikit-learn/\n3. https://www-bcf.usc.edu/~gareth/ISL/ISLR%20First%20Printing.pdf\n4. http://setosa.io/ev/principal-component-analysis/\n\n",
"_____no_output_____"
]
]
] | [
"markdown",
"code",
"markdown",
"code",
"markdown"
] | [
[
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown"
]
] |
d0ff30285d457ebc277238511c3ae0c29affdd65 | 705 | ipynb | Jupyter Notebook | lezione1/testo-esercizi/ipynb/ES8-VerificaTriangoloRettangolo-testo.ipynb | DavideMaspero/python-alfabetizzazione-docenti | b7edf8291b2e7999cbc0ffe69ac5e046733c1a7e | [
"MIT"
] | null | null | null | lezione1/testo-esercizi/ipynb/ES8-VerificaTriangoloRettangolo-testo.ipynb | DavideMaspero/python-alfabetizzazione-docenti | b7edf8291b2e7999cbc0ffe69ac5e046733c1a7e | [
"MIT"
] | null | null | null | lezione1/testo-esercizi/ipynb/ES8-VerificaTriangoloRettangolo-testo.ipynb | DavideMaspero/python-alfabetizzazione-docenti | b7edf8291b2e7999cbc0ffe69ac5e046733c1a7e | [
"MIT"
] | 1 | 2019-03-26T11:14:33.000Z | 2019-03-26T11:14:33.000Z | 20.142857 | 117 | 0.570213 | [
[
[
"### ESERCIZIO 8 - Verifica di triangolo rettangolo\n\nDati tre valori (interi o decimali) verificare se possono essere le misure dei lati di un triangolo rettangolo.",
"_____no_output_____"
]
]
] | [
"markdown"
] | [
[
"markdown"
]
] |
d0ff4403f0e62eb450fa4038dbca69c7d5339ff8 | 128,682 | ipynb | Jupyter Notebook | .ipynb_checkpoints/w209_final_project_eda_don_irwin-checkpoint.ipynb | tuneman7/w209_spring_2022_thu_4_pm_team_4_web | b2707d850ea2fab3032f86533b247a585eb3b131 | [
"Apache-2.0"
] | null | null | null | .ipynb_checkpoints/w209_final_project_eda_don_irwin-checkpoint.ipynb | tuneman7/w209_spring_2022_thu_4_pm_team_4_web | b2707d850ea2fab3032f86533b247a585eb3b131 | [
"Apache-2.0"
] | 39 | 2022-03-01T14:30:36.000Z | 2022-03-31T23:59:36.000Z | .ipynb_checkpoints/w209_final_project_eda_don_irwin-checkpoint.ipynb | tuneman7/w209_spring_2022_thu_4_pm_team_4_web | b2707d850ea2fab3032f86533b247a585eb3b131 | [
"Apache-2.0"
] | null | null | null | 274.375267 | 81,997 | 0.529188 | [
[
[
"from libraries.import_export_data_objects import import_export_data as Import_Export_Data\nfrom libraries.altair_renderings import AltairRenderings\nfrom libraries.utility import Utility\nimport os\nimport altair as alt\nmy_altair = AltairRenderings()\nfrom IPython.core.display import display, HTML\ndisplay(HTML(\n '<style>'\n '#notebook { padding-top:0px !important; } ' \n '.container { width:100% !important; } '\n '.end_space { min-height:0px !important; } '\n '.end_space { min-height:0px !important; } '\n '.prompt {width: 0px; min-width: 0px; visibility: collapse } '\n '</style>'\n))",
"_____no_output_____"
]
],
[
[
"# <center>Exploratory Data Analysis</center>\n## <center>Don Irwin</center>\n### <center>U.C. Berkeley MIDS</center>\n### <center>W209 - Spring 2022 Thursday 4 P.M.</center>",
"_____no_output_____"
],
[
"### Background:\n\nMIDS students taking W209, Data Visualization, are expected to demonstrate an ability to source datasets, analyze datasets, and present visualizations in the form of web-based applications, to convey insights from those datasets to users.\n\nOur team will communicate trade trends over the past 6 years, for the world's top 20 trading nations.\n\n### Data Sources, and description of those data:\n\nOur data is sourced from the World Trade Organization, and Bloomberg. These data consist primarily of datasets which contain total trade, total imports, and total exports, time-series, by any one of the top 20 trading nations with another. These data also contain trade categories, of each of these nations with the other.\n\n### Limitations of data set:\n\nThis data set is limited to 20 countries for 6 years. This is due to the difficulty of harvesting, and pre-processing data. These data are sufficiently feature rich to demonstrate hypothesis. However, they lack longtitudinal depth require to draw conclusions about world trade trends over a long term. They do, however, provide a view into trade between the world's 20 largest trading nations over the past six years.\n\n\n## Hypothesis 1:\n\nWhile not a country, the European Union, has been the world's largest trading block. It has been so for the past 6 years.\n",
"_____no_output_____"
]
],
[
[
"my_altair.get_import_export_balance_top_five(\"World\")\n#my_altair.get_altaire_bar_top5_partners(\"World\")",
"_____no_output_____"
]
],
[
[
"### What is informative about this view:\n\nThe view above shows the top 5 trading nations or blocks in the world with a slider for the years between 2014 and 2020.\n\nUtilizing the slider, a user can see that for each of the years the European Union is consistently the top trading block.\n\nWhile not directly related to the hypothesis, we see that nations with net negative trade values (related to the world) are nations with which the world is importing more things from. We see that the world is consistently importing more things from China, while consistently exporting more things to the United States. This means china is a net retrainer of trade dollars, while the United States is a net exporter, so to speak, of trade dollars.\n\n### What could be improved about this view:\n\nThis view is bordering on being a bit busy, and may be better as a stacked bar chart, or ordered from highest trade, to lowest trade.",
"_____no_output_____"
],
[
"## Hypothesis 2:\n\nChina dominates trade with the Middle East, South Asia, Southeast Asia, and Asia Pacific nations. China has done so for the past 6 years with, although, moving the slider, below will show that the European Union was the top trading partner of Saudi Arabia and India for a few of the years prior to 2016.",
"_____no_output_____"
]
],
[
[
"indo = my_altair.get_altaire_bar_top5_partners_for_matrix(\"Indonesia\")\nindo\nsaudi = my_altair.get_altaire_bar_top5_partners_for_matrix(\"Saudi Arabia\")\nsaudi\niran = my_altair.get_altaire_bar_top5_partners_for_matrix(\"Iran\")\niran\nsk = my_altair.get_altaire_bar_top5_partners_for_matrix(\"South Korea\")\nsk\njap = my_altair.get_altaire_bar_top5_partners_for_matrix(\"Japan\")\nindia = my_altair.get_altaire_bar_top5_partners_for_matrix(\"India\")\n\nrow_1 = (indo | saudi )\nrow_2 = (iran | india )\nrow_3 = (sk | jap )\nmy_chart = (row_1 & row_2 & row_3).configure_axis(\n grid=False\n ).configure_view(\n strokeWidth=0\n )\n\nmy_chart\n#indo.properties(width=700,height=200)",
"_____no_output_____"
]
],
[
[
"### What is informative about this view:\n\nThe view above shows 6 nations in Asia, from the Middle East, to the Far East, and it provides a slider to compare different years. We see, that for almost all of these nations, for almost all years in the visualization, China is the top trading partner. In the case of India and Saudi Arabia, we see that in 2014, their leading trading partner block was the European Union, but it switched to China.\n\n### What could be improved about this view:\n\nThis view sometimes \"wiggles\" because of the labels pushing out from side to side. This could be improved.\n",
"_____no_output_____"
],
[
"## Hypothesis 3\n\nGeopolitical events can have a drastic impact on a nation's trade, and that nation's ability to buy and sell with the rest of the world.\n\n",
"_____no_output_____"
]
],
[
[
"sk = my_altair.get_altaire_line_chart_county_trade_for_matrix(\"South Korea\",\"Iran\")\nspain = my_altair.get_altaire_line_chart_county_trade_for_matrix(\"Spain\",\"Iran\")\nusa = my_altair.get_altaire_line_chart_county_trade_for_matrix(\"United States\",\"Iran\")\njap = my_altair.get_altaire_line_chart_county_trade_for_matrix(\"Indonesia\",\"Iran\")\n\n\nrow_1 = (sk | spain )\nrow_2 = (usa | jap )\nmy_chart = (row_1 & row_2).configure_axis(\n grid=False\n ).configure_view(\n strokeWidth=0\n )\n\nmy_chart\n#indo.properties(width=700,height=200)",
"_____no_output_____"
]
],
[
[
"### What is informative about this view:\n\nThe View above clearly shows the trade impact on Iran as the result of the Trump Administration's withdrawl from the JCPOA, also known as the \"Iran Nuclear Deal\".\n\nWe can see that generally from 2014 (the time of the JCPOA) until 2016-2017 trade between Iran and other nations was increasing drastically.\n\nOnce Trump imposed secondary sanctions on people trading with Iran, there was a massive crash in trade with Iran.\n\n### What could be improved about this view:\n\nLines should be thicker, and the tool tip should be improved. The charts could be changed to indicate in some fashion, the date of the cancellation of the JCPOA.",
"_____no_output_____"
],
[
"## Conclusion:\n\nThe data do appear to support the various hypothensis presented.\n\nThe challenge with this project is to create a Summary-Zoom-Detail paradign within the application that can communicate effectively to a user.\n\nAt this stage, we are still struggling with how to represent different datasets and datum effectively.\n\nThank you for your instruction during this project.\n\nBest regards,\n\nDon Irwin\n",
"_____no_output_____"
]
]
] | [
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown"
] | [
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
]
] |
d0ff493e868e18ea4b129e200664ea77e5331ca2 | 280,361 | ipynb | Jupyter Notebook | final_dlnd_face_generation.ipynb | LKona/Generate-Faces-Using-GANs | e4bfa83f1feb7e7975bb6c090bbe72f594c71d23 | [
"MIT"
] | null | null | null | final_dlnd_face_generation.ipynb | LKona/Generate-Faces-Using-GANs | e4bfa83f1feb7e7975bb6c090bbe72f594c71d23 | [
"MIT"
] | null | null | null | final_dlnd_face_generation.ipynb | LKona/Generate-Faces-Using-GANs | e4bfa83f1feb7e7975bb6c090bbe72f594c71d23 | [
"MIT"
] | null | null | null | 174.462352 | 97,460 | 0.855643 | [
[
[
"# Face Generation\n\nIn this project, you'll define and train a DCGAN on a dataset of faces. Your goal is to get a generator network to generate *new* images of faces that look as realistic as possible!\n\nThe project will be broken down into a series of tasks from **loading in data to defining and training adversarial networks**. At the end of the notebook, you'll be able to visualize the results of your trained Generator to see how it performs; your generated samples should look like fairly realistic faces with small amounts of noise.\n\n### Get the Data\n\nYou'll be using the [CelebFaces Attributes Dataset (CelebA)](http://mmlab.ie.cuhk.edu.hk/projects/CelebA.html) to train your adversarial networks.\n\nThis dataset is more complex than the number datasets (like MNIST or SVHN) you've been working with, and so, you should prepare to define deeper networks and train them for a longer time to get good results. It is suggested that you utilize a GPU for training.\n\n### Pre-processed Data\n\nSince the project's main focus is on building the GANs, we've done *some* of the pre-processing for you. Each of the CelebA images has been cropped to remove parts of the image that don't include a face, then resized down to 64x64x3 NumPy images. Some sample data is show below.\n\n<img src='assets/processed_face_data.png' width=60% />\n\n> If you are working locally, you can download this data [by clicking here](https://s3.amazonaws.com/video.udacity-data.com/topher/2018/November/5be7eb6f_processed-celeba-small/processed-celeba-small.zip)\n\nThis is a zip file that you'll need to extract in the home directory of this notebook for further loading and processing. After extracting the data, you should be left with a directory of data `processed_celeba_small/`",
"_____no_output_____"
]
],
[
[
"# can comment out after executing\n# !unzip processed_celeba_small.zip",
"_____no_output_____"
],
[
"import os\nthedir = './processed_celeba_small/celeba/'\n# [ name for name in os.listdir(thedir) if os.path.isdir(os.path.join(thedir, name)) ]\nprint (len([name for name in os.listdir(thedir) if os.path.isfile(os.path.join(thedir, name))]))",
"89932\n"
],
[
"data_dir = 'processed_celeba_small/'\n\n\"\"\"\nDON'T MODIFY ANYTHING IN THIS CELL\n\"\"\"\nimport pickle as pkl\nimport matplotlib.pyplot as plt\nimport numpy as np\nimport problem_unittests as tests\n#import helper\n\n%matplotlib inline",
"_____no_output_____"
]
],
[
[
"## Visualize the CelebA Data\n\nThe [CelebA](http://mmlab.ie.cuhk.edu.hk/projects/CelebA.html) dataset contains over 200,000 celebrity images with annotations. Since you're going to be generating faces, you won't need the annotations, you'll only need the images. Note that these are color images with [3 color channels (RGB)](https://en.wikipedia.org/wiki/Channel_(digital_image)#RGB_Images) each.\n\n### Pre-process and Load the Data\n\nSince the project's main focus is on building the GANs, we've done *some* of the pre-processing for you. Each of the CelebA images has been cropped to remove parts of the image that don't include a face, then resized down to 64x64x3 NumPy images. This *pre-processed* dataset is a smaller subset of the very large CelebA data.\n\n> There are a few other steps that you'll need to **transform** this data and create a **DataLoader**.\n\n#### Exercise: Complete the following `get_dataloader` function, such that it satisfies these requirements:\n\n* Your images should be square, Tensor images of size `image_size x image_size` in the x and y dimension.\n* Your function should return a DataLoader that shuffles and batches these Tensor images.\n\n#### ImageFolder\n\nTo create a dataset given a directory of images, it's recommended that you use PyTorch's [ImageFolder](https://pytorch.org/docs/stable/torchvision/datasets.html#imagefolder) wrapper, with a root directory `processed_celeba_small/` and data transformation passed in.",
"_____no_output_____"
]
],
[
[
"# necessary imports\nimport torch\nimport os\nfrom torchvision import datasets\nfrom torchvision import transforms\nfrom torch.utils.data import DataLoader\n",
"_____no_output_____"
],
[
"# import shutil\n# import os\n\n# image_path = './' + data_dir\n# source = os.path.join(image_path,'celeba/New Folder With Items/')\n# print(source)\n# # source = './processed_celeba_small/celeba/New Folder With Items'\n# dest1 = './processed_celeba_small/celeba/'\n\n\n# files = os.listdir(source)\n\n# for f in files:\n# shutil.move(source+f, dest1)",
"_____no_output_____"
],
[
"def get_dataloader(batch_size, image_size, data_dir='processed_celeba_small/'):\n \"\"\"\n Batch the neural network data using DataLoader\n :param batch_size: The size of each batch; the number of images in a batch\n :param img_size: The square size of the image data (x, y)\n :param data_dir: Directory where image data is located\n :return: DataLoader with batched data\n \"\"\"\n \n # TODO: Implement function and return a dataloader\n # resize and normalize the images\n transform = transforms.Compose([transforms.Resize(image_size), transforms.ToTensor()])\n\n # get training directories\n train_path = './' + data_dir\n# train_path = os.path.join(image_path,'celeba')\n print(train_path)\n\n # define datasets using ImageFolder\n train_dataset = datasets.ImageFolder(train_path, transform)\n\n # create and return DataLoaders\n num_workers = 0\n train_loader = DataLoader(dataset=train_dataset, batch_size=batch_size, shuffle=True, num_workers=num_workers)\n \n return train_loader",
"_____no_output_____"
]
],
[
[
"## Create a DataLoader\n\n#### Exercise: Create a DataLoader `celeba_train_loader` with appropriate hyperparameters.\n\nCall the above function and create a dataloader to view images. \n* You can decide on any reasonable `batch_size` parameter\n* Your `image_size` **must be** `32`. Resizing the data to a smaller size will make for faster training, while still creating convincing images of faces!",
"_____no_output_____"
]
],
[
[
"# Define function hyperparameters\nbatch_size = 128\nimg_size = 32\n\n\"\"\"\nDON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE\n\"\"\"\n# Call your function and get a dataloader\nceleba_train_loader = get_dataloader(batch_size, img_size)\n\nprint(len(celeba_train_loader.dataset))\n",
"./processed_celeba_small/\n89931\n"
]
],
[
[
"Next, you can view some images! You should seen square images of somewhat-centered faces.\n\nNote: You'll need to convert the Tensor images into a NumPy type and transpose the dimensions to correctly display an image, suggested `imshow` code is below, but it may not be perfect.",
"_____no_output_____"
]
],
[
[
"# helper display function\ndef imshow(img):\n npimg = img.numpy()\n plt.imshow(np.transpose(npimg, (1, 2, 0)))\n\n\"\"\"\nDON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE\n\"\"\"\n# obtain one batch of training images\ndataiter = iter(celeba_train_loader)\nimages, _ = dataiter.next() # _ for no labels\n\n# plot the images in the batch, along with the corresponding labels\nfig = plt.figure(figsize=(20, 4))\nplot_size=20\nfor idx in np.arange(plot_size):\n ax = fig.add_subplot(2, plot_size/2, idx+1, xticks=[], yticks=[])\n imshow(images[idx])",
"_____no_output_____"
]
],
[
[
"#### Exercise: Pre-process your image data and scale it to a pixel range of -1 to 1\n\nYou need to do a bit of pre-processing; you know that the output of a `tanh` activated generator will contain pixel values in a range from -1 to 1, and so, we need to rescale our training images to a range of -1 to 1. (Right now, they are in a range from 0-1.)",
"_____no_output_____"
]
],
[
[
"# TODO: Complete the scale function\ndef scale(x, feature_range=(-1, 1)):\n ''' Scale takes in an image x and returns that image, scaled\n with a feature_range of pixel values from -1 to 1. \n This function assumes that the input x is already scaled from 0-1.'''\n # assume x is scaled to (0, 1)\n # scale to feature_range and return scaled x\n min, max = feature_range\n x = x * (max - min) + min\n return x\n",
"_____no_output_____"
],
[
"\"\"\"\nDON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE\n\"\"\"\n# check scaled range\n# should be close to -1 to 1\nimg = images[0]\nscaled_img = scale(img)\n\nprint('Min: ', scaled_img.min())\nprint('Max: ', scaled_img.max())",
"Min: tensor(-0.9922)\nMax: tensor(0.8275)\n"
]
],
[
[
"---\n# Define the Model\n\nA GAN is comprised of two adversarial networks, a discriminator and a generator.\n\n## Discriminator\n\nYour first task will be to define the discriminator. This is a convolutional classifier like you've built before, only without any maxpooling layers. To deal with this complex data, it's suggested you use a deep network with **normalization**. You are also allowed to create any helper functions that may be useful.\n\n#### Exercise: Complete the Discriminator class\n* The inputs to the discriminator are 32x32x3 tensor images\n* The output should be a single value that will indicate whether a given image is real or fake\n",
"_____no_output_____"
]
],
[
[
"import torch.nn as nn\nimport torch.nn.functional as F",
"_____no_output_____"
],
[
"# helper conv function\ndef conv(in_channels, out_channels, kernel_size, stride=2, padding=1, batch_norm=True):\n \"\"\"Creates a convolutional layer, with optional batch normalization.\n \"\"\"\n layers = []\n conv_layer = nn.Conv2d(in_channels, out_channels, \n kernel_size, stride, padding, bias=False)\n \n # append conv layer\n layers.append(conv_layer)\n\n if batch_norm:\n # append batchnorm layer\n layers.append(nn.BatchNorm2d(out_channels))\n \n # using Sequential container\n return nn.Sequential(*layers)",
"_____no_output_____"
],
[
"class Discriminator(nn.Module):\n\n def __init__(self, conv_dim):\n \"\"\"\n Initialize the Discriminator Module\n :param conv_dim: The depth of the first convolutional layer\n \"\"\"\n super(Discriminator, self).__init__()\n\n # complete init function\n self.conv_dim = conv_dim\n\n # 32x32 input\n self.conv1 = conv(3, conv_dim, 4, batch_norm=False) # first layer, no batch_norm\n self.conv2 = conv(conv_dim, conv_dim*2, 4)\n self.conv3 = conv(conv_dim*2, conv_dim*4, 4)\n \n \n # final, fully-connected layer\n self.fc = nn.Linear(conv_dim*4*4*4, 1)\n \n\n def forward(self, x):\n \"\"\"\n Forward propagation of the neural network\n :param x: The input to the neural network \n :return: Discriminator logits; the output of the neural network\n \"\"\"\n # define feedforward behavior\n out = F.leaky_relu(self.conv1(x), 0.2)\n out = F.leaky_relu(self.conv2(out), 0.2)\n out = F.leaky_relu(self.conv3(out), 0.2)\n\n # flatten\n out = out.view(-1, self.conv_dim*4*4*4)\n \n # final output layer\n out = self.fc(out) \n return out\n\n\n\"\"\"\nDON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE\n\"\"\"\ntests.test_discriminator(Discriminator)",
"Tests Passed\n"
]
],
[
[
"## Generator\n\nThe generator should upsample an input and generate a *new* image of the same size as our training data `32x32x3`. This should be mostly transpose convolutional layers with normalization applied to the outputs.\n\n#### Exercise: Complete the Generator class\n* The inputs to the generator are vectors of some length `z_size`\n* The output should be a image of shape `32x32x3`",
"_____no_output_____"
]
],
[
[
"def deconv(in_channels, out_channels, kernel_size, stride=2, padding=1, batch_norm=True):\n \"\"\"Creates a transposed-convolutional layer, with optional batch normalization.\n \"\"\"\n # create a sequence of transpose + optional batch norm layers\n layers = []\n transpose_conv_layer = nn.ConvTranspose2d(in_channels, out_channels, \n kernel_size, stride, padding, bias=False)\n # append transpose convolutional layer\n layers.append(transpose_conv_layer)\n \n if batch_norm:\n # append batchnorm layer\n layers.append(nn.BatchNorm2d(out_channels))\n \n return nn.Sequential(*layers)",
"_____no_output_____"
],
[
"class Generator(nn.Module):\n \n def __init__(self, z_size, conv_dim):\n \"\"\"\n Initialize the Generator Module\n :param z_size: The length of the input latent vector, z\n :param conv_dim: The depth of the inputs to the *last* transpose convolutional layer\n \"\"\"\n super(Generator, self).__init__()\n\n # complete init function\n self.conv_dim = conv_dim\n \n # first, fully-connected layer\n self.fc = nn.Linear(z_size, conv_dim*4*4*4)\n\n # transpose conv layers\n self.t_conv1 = deconv(conv_dim*4, conv_dim*2, 4)\n self.t_conv2 = deconv(conv_dim*2, conv_dim, 4)\n self.t_conv3 = deconv(conv_dim, 3, 4, batch_norm=False)\n \n\n def forward(self, x):\n \"\"\"\n Forward propagation of the neural network\n :param x: The input to the neural network \n :return: A 32x32x3 Tensor image as output\n \"\"\"\n # define feedforward behavior\n # fully-connected + reshape \n out = self.fc(x)\n out = out.view(-1, self.conv_dim*4, 4, 4) # (batch_size, depth, 4, 4)\n # hidden transpose conv layers + relu\n out = F.relu(self.t_conv1(out))\n out = F.relu(self.t_conv2(out))\n \n # last layer + tanh activation\n out = self.t_conv3(out)\n out = F.tanh(out)\n return out\n\n\"\"\"\nDON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE\n\"\"\"\ntests.test_generator(Generator)",
"Tests Passed\n"
]
],
[
[
"## Initialize the weights of your networks\n\nTo help your models converge, you should initialize the weights of the convolutional and linear layers in your model. From reading the [original DCGAN paper](https://arxiv.org/pdf/1511.06434.pdf), they say:\n> All weights were initialized from a zero-centered Normal distribution with standard deviation 0.02.\n\nSo, your next task will be to define a weight initialization function that does just this!\n\nYou can refer back to the lesson on weight initialization or even consult existing model code, such as that from [the `networks.py` file in CycleGAN Github repository](https://github.com/junyanz/pytorch-CycleGAN-and-pix2pix/blob/master/models/networks.py) to help you complete this function.\n\n#### Exercise: Complete the weight initialization function\n\n* This should initialize only **convolutional** and **linear** layers\n* Initialize the weights to a normal distribution, centered around 0, with a standard deviation of 0.02.\n* The bias terms, if they exist, may be left alone or set to 0.",
"_____no_output_____"
]
],
[
[
"from torch.nn import init\n\ndef weights_init_normal(m):\n \"\"\"\n Applies initial weights to certain layers in a model .\n The weights are taken from a normal distribution \n with mean = 0, std dev = 0.02.\n :param m: A module or layer in a network \n \"\"\"\n # classname will be something like:\n # `Conv`, `BatchNorm2d`, `Linear`, etc.\n classname = m.__class__.__name__\n \n # TODO: Apply initial weights to convolutional and linear layers\n if classname.find('Conv') != -1:\n nn.init.normal_(m.weight.data, 0.0, 0.02)\n elif classname.find('BatchNorm') != -1:\n nn.init.normal_(m.weight.data, 1.0, 0.02)\n nn.init.constant_(m.bias.data, 0)\n# print('initialize network with %s' % init_type)\n \n ",
"_____no_output_____"
]
],
[
[
"## Build complete network\n\nDefine your models' hyperparameters and instantiate the discriminator and generator from the classes defined above. Make sure you've passed in the correct input arguments.",
"_____no_output_____"
]
],
[
[
"\"\"\"\nDON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE\n\"\"\"\ndef build_network(d_conv_dim, g_conv_dim, z_size):\n # define discriminator and generator\n D = Discriminator(d_conv_dim)\n G = Generator(z_size=z_size, conv_dim=g_conv_dim)\n\n # initialize model weights\n D.apply(weights_init_normal)\n G.apply(weights_init_normal)\n\n print(D)\n print()\n print(G)\n \n return D, G\n",
"_____no_output_____"
]
],
[
[
"#### Exercise: Define model hyperparameters",
"_____no_output_____"
]
],
[
[
"# Define model hyperparams\nd_conv_dim = 32\ng_conv_dim = 32\nz_size = 200\n\n\"\"\"\nDON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE\n\"\"\"\nD, G = build_network(d_conv_dim, g_conv_dim, z_size)",
"Discriminator(\n (conv1): Sequential(\n (0): Conv2d(3, 32, kernel_size=(4, 4), stride=(2, 2), padding=(1, 1), bias=False)\n )\n (conv2): Sequential(\n (0): Conv2d(32, 64, kernel_size=(4, 4), stride=(2, 2), padding=(1, 1), bias=False)\n (1): BatchNorm2d(64, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)\n )\n (conv3): Sequential(\n (0): Conv2d(64, 128, kernel_size=(4, 4), stride=(2, 2), padding=(1, 1), bias=False)\n (1): BatchNorm2d(128, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)\n )\n (fc): Linear(in_features=2048, out_features=1, bias=True)\n)\n\nGenerator(\n (fc): Linear(in_features=200, out_features=2048, bias=True)\n (t_conv1): Sequential(\n (0): ConvTranspose2d(128, 64, kernel_size=(4, 4), stride=(2, 2), padding=(1, 1), bias=False)\n (1): BatchNorm2d(64, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)\n )\n (t_conv2): Sequential(\n (0): ConvTranspose2d(64, 32, kernel_size=(4, 4), stride=(2, 2), padding=(1, 1), bias=False)\n (1): BatchNorm2d(32, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)\n )\n (t_conv3): Sequential(\n (0): ConvTranspose2d(32, 3, kernel_size=(4, 4), stride=(2, 2), padding=(1, 1), bias=False)\n )\n)\n"
]
],
[
[
"### Training on GPU\n\nCheck if you can train on GPU. Here, we'll set this as a boolean variable `train_on_gpu`. Later, you'll be responsible for making sure that \n>* Models,\n* Model inputs, and\n* Loss function arguments\n\nAre moved to GPU, where appropriate.",
"_____no_output_____"
]
],
[
[
"\"\"\"\nDON'T MODIFY ANYTHING IN THIS CELL\n\"\"\"\nimport torch\n\n# Check for a GPU\ntrain_on_gpu = torch.cuda.is_available()\nif not train_on_gpu:\n print('No GPU found. Please use a GPU to train your neural network.')\nelse:\n print('Training on GPU!')",
"Training on GPU!\n"
]
],
[
[
"---\n## Discriminator and Generator Losses\n\nNow we need to calculate the losses for both types of adversarial networks.\n\n### Discriminator Losses\n\n> * For the discriminator, the total loss is the sum of the losses for real and fake images, `d_loss = d_real_loss + d_fake_loss`. \n* Remember that we want the discriminator to output 1 for real images and 0 for fake images, so we need to set up the losses to reflect that.\n\n\n### Generator Loss\n\nThe generator loss will look similar only with flipped labels. The generator's goal is to get the discriminator to *think* its generated images are *real*.\n\n#### Exercise: Complete real and fake loss functions\n\n**You may choose to use either cross entropy or a least squares error loss to complete the following `real_loss` and `fake_loss` functions.**",
"_____no_output_____"
]
],
[
[
"def real_loss(D_out, smooth=False):\n '''Calculates how close discriminator outputs are to being real.\n param, D_out: discriminator logits\n return: real loss'''\n batch_size = D_out.size(0)\n # label smoothing\n if smooth:\n # smooth, real labels = 0.9\n labels = torch.ones(batch_size)*0.9\n else:\n labels = torch.ones(batch_size) # real labels = 1\n # move labels to GPU if available \n if train_on_gpu:\n labels = labels.cuda()\n # binary cross entropy with logits loss\n criterion = nn.BCEWithLogitsLoss()\n # calculate loss\n loss = criterion(D_out.squeeze(), labels)\n return loss\n\ndef fake_loss(D_out):\n '''Calculates how close discriminator outputs are to being fake.\n param, D_out: discriminator logits\n return: fake loss'''\n batch_size = D_out.size(0)\n labels = torch.zeros(batch_size) # fake labels = 0\n if train_on_gpu:\n labels = labels.cuda()\n criterion = nn.BCEWithLogitsLoss()\n # calculate loss\n loss = criterion(D_out.squeeze(), labels)\n return loss",
"_____no_output_____"
]
],
[
[
"## Optimizers\n\n#### Exercise: Define optimizers for your Discriminator (D) and Generator (G)\n\nDefine optimizers for your models with appropriate hyperparameters.",
"_____no_output_____"
]
],
[
[
"import torch.optim as optim\n\n# params\nlr = 0.0002\nbeta1=0.5\nbeta2=0.999 # default value\n\n# Create optimizers for the discriminator D and generator G\nd_optimizer = optim.Adam(D.parameters(), lr, [beta1, beta2])\ng_optimizer = optim.Adam(G.parameters(), lr, [beta1, beta2])",
"_____no_output_____"
]
],
[
[
"---\n## Training\n\nTraining will involve alternating between training the discriminator and the generator. You'll use your functions `real_loss` and `fake_loss` to help you calculate the discriminator losses.\n\n* You should train the discriminator by alternating on real and fake images\n* Then the generator, which tries to trick the discriminator and should have an opposing loss function\n\n\n#### Saving Samples\n\nYou've been given some code to print out some loss statistics and save some generated \"fake\" samples.",
"_____no_output_____"
],
[
"#### Exercise: Complete the training function\n\nKeep in mind that, if you've moved your models to GPU, you'll also have to move any model inputs to GPU.",
"_____no_output_____"
]
],
[
[
"def train(D, G, n_epochs, print_every=50):\n '''Trains adversarial networks for some number of epochs\n param, D: the discriminator network\n param, G: the generator network\n param, n_epochs: number of epochs to train for\n param, print_every: when to print and record the models' losses\n return: D and G losses'''\n \n # move models to GPU\n if train_on_gpu:\n D.cuda()\n G.cuda()\n\n # keep track of loss and generated, \"fake\" samples\n samples = []\n losses = []\n\n # Get some fixed data for sampling. These are images that are held\n # constant throughout training, and allow us to inspect the model's performance\n sample_size=16\n fixed_z = np.random.uniform(-1, 1, size=(sample_size, z_size))\n fixed_z = torch.from_numpy(fixed_z).float()\n # move z to GPU if available\n if train_on_gpu:\n fixed_z = fixed_z.cuda()\n\n # epoch training loop\n for epoch in range(n_epochs):\n\n # batch training loop\n for batch_i, (real_images, _) in enumerate(celeba_train_loader):\n\n batch_size = real_images.size(0)\n real_images = scale(real_images)\n\n # ===============================================\n # YOUR CODE HERE: TRAIN THE NETWORKS\n # ===============================================\n \n # ============================================\n # TRAIN THE DISCRIMINATOR\n # ============================================\n \n d_optimizer.zero_grad()\n\n # 1. Train with real images\n\n # Compute the discriminator losses on real images \n if train_on_gpu:\n real_images = real_images.cuda()\n\n D_real = D(real_images)\n d_real_loss = real_loss(D_real)\n\n # 2. Train with fake images\n\n # Generate fake images\n z = np.random.uniform(-1, 1, size=(batch_size, z_size))\n z = torch.from_numpy(z).float()\n # move x to GPU, if available\n if train_on_gpu:\n z = z.cuda()\n fake_images = G(z)\n\n # Compute the discriminator losses on fake images \n D_fake = D(fake_images)\n d_fake_loss = fake_loss(D_fake)\n\n # add up loss and perform backprop\n d_loss = d_real_loss + d_fake_loss\n d_loss.backward()\n d_optimizer.step()\n\n\n # =========================================\n # TRAIN THE GENERATOR\n # =========================================\n g_optimizer.zero_grad()\n\n # 1. Train with fake images and flipped labels\n\n # Generate fake images\n z = np.random.uniform(-1, 1, size=(batch_size, z_size))\n z = torch.from_numpy(z).float()\n if train_on_gpu:\n z = z.cuda()\n fake_images = G(z)\n\n # Compute the discriminator losses on fake images \n # using flipped labels!\n D_fake = D(fake_images)\n g_loss = real_loss(D_fake) # use real loss to flip labels\n\n # perform backprop\n g_loss.backward()\n g_optimizer.step()\n \n # 1. Train the discriminator on real and fake images\n \n # ===============================================\n # END OF YOUR CODE\n # ===============================================\n\n # Print some loss stats\n if batch_i % print_every == 0:\n # append discriminator loss and generator loss\n losses.append((d_loss.item(), g_loss.item()))\n # print discriminator and generator loss\n print('Epoch [{:5d}/{:5d}] | d_loss: {:6.4f} | g_loss: {:6.4f}'.format(\n epoch+1, n_epochs, d_loss.item(), g_loss.item()))\n\n\n ## AFTER EACH EPOCH## \n # this code assumes your generator is named G, feel free to change the name\n # generate and save sample, fake images\n G.eval() # for generating samples\n samples_z = G(fixed_z)\n samples.append(samples_z)\n G.train() # back to training mode\n\n # Save training generator samples\n with open('train_samples.pkl', 'wb') as f:\n pkl.dump(samples, f)\n \n # finally return losses\n return losses",
"_____no_output_____"
]
],
[
[
"Set your number of training epochs and train your GAN!",
"_____no_output_____"
]
],
[
[
"# set number of epochs \nn_epochs = 30\n\n\n\"\"\"\nDON'T MODIFY ANYTHING IN THIS CELL\n\"\"\"\n# call training function\nlosses = train(D, G, n_epochs=n_epochs)",
"Epoch [ 1/ 30] | d_loss: 1.4314 | g_loss: 0.9460\nEpoch [ 1/ 30] | d_loss: 0.1727 | g_loss: 3.4594\nEpoch [ 1/ 30] | d_loss: 0.2041 | g_loss: 4.0258\nEpoch [ 1/ 30] | d_loss: 0.2896 | g_loss: 3.3163\nEpoch [ 1/ 30] | d_loss: 0.1577 | g_loss: 3.8655\nEpoch [ 1/ 30] | d_loss: 1.1076 | g_loss: 6.7513\nEpoch [ 1/ 30] | d_loss: 0.4166 | g_loss: 2.2213\nEpoch [ 1/ 30] | d_loss: 0.8505 | g_loss: 4.1158\nEpoch [ 1/ 30] | d_loss: 0.4360 | g_loss: 2.8230\nEpoch [ 1/ 30] | d_loss: 0.5173 | g_loss: 1.8475\nEpoch [ 1/ 30] | d_loss: 0.5994 | g_loss: 2.2314\nEpoch [ 1/ 30] | d_loss: 0.7012 | g_loss: 1.5644\nEpoch [ 1/ 30] | d_loss: 0.6658 | g_loss: 1.7771\nEpoch [ 1/ 30] | d_loss: 0.7777 | g_loss: 2.6704\nEpoch [ 1/ 30] | d_loss: 0.7911 | g_loss: 1.4424\nEpoch [ 2/ 30] | d_loss: 0.8650 | g_loss: 1.3959\nEpoch [ 2/ 30] | d_loss: 0.9371 | g_loss: 2.0485\nEpoch [ 2/ 30] | d_loss: 0.7884 | g_loss: 2.1352\nEpoch [ 2/ 30] | d_loss: 0.9461 | g_loss: 1.4666\nEpoch [ 2/ 30] | d_loss: 0.9289 | g_loss: 0.9274\nEpoch [ 2/ 30] | d_loss: 0.9289 | g_loss: 1.6948\nEpoch [ 2/ 30] | d_loss: 1.0173 | g_loss: 0.9131\nEpoch [ 2/ 30] | d_loss: 0.9509 | g_loss: 0.7993\nEpoch [ 2/ 30] | d_loss: 1.0011 | g_loss: 1.1105\nEpoch [ 2/ 30] | d_loss: 0.8487 | g_loss: 1.1870\nEpoch [ 2/ 30] | d_loss: 1.1135 | g_loss: 0.6568\nEpoch [ 2/ 30] | d_loss: 0.8575 | g_loss: 1.1643\nEpoch [ 2/ 30] | d_loss: 1.2244 | g_loss: 0.9746\nEpoch [ 2/ 30] | d_loss: 0.8273 | g_loss: 1.1185\nEpoch [ 2/ 30] | d_loss: 1.3302 | g_loss: 1.5740\nEpoch [ 3/ 30] | d_loss: 1.0233 | g_loss: 1.7974\nEpoch [ 3/ 30] | d_loss: 0.9511 | g_loss: 1.5634\nEpoch [ 3/ 30] | d_loss: 0.9333 | g_loss: 1.7265\nEpoch [ 3/ 30] | d_loss: 0.5957 | g_loss: 1.5651\nEpoch [ 3/ 30] | d_loss: 0.9729 | g_loss: 1.1097\nEpoch [ 3/ 30] | d_loss: 0.9091 | g_loss: 0.6549\nEpoch [ 3/ 30] | d_loss: 0.6871 | g_loss: 1.0341\nEpoch [ 3/ 30] | d_loss: 1.0139 | g_loss: 1.4646\nEpoch [ 3/ 30] | d_loss: 0.8280 | g_loss: 0.9359\nEpoch [ 3/ 30] | d_loss: 0.9145 | g_loss: 1.5236\nEpoch [ 3/ 30] | d_loss: 1.3221 | g_loss: 2.8362\nEpoch [ 3/ 30] | d_loss: 0.8141 | g_loss: 2.1691\nEpoch [ 3/ 30] | d_loss: 0.9754 | g_loss: 1.6181\nEpoch [ 3/ 30] | d_loss: 0.9496 | g_loss: 1.1102\nEpoch [ 3/ 30] | d_loss: 0.7953 | g_loss: 1.2024\nEpoch [ 4/ 30] | d_loss: 1.3427 | g_loss: 2.4101\nEpoch [ 4/ 30] | d_loss: 1.2492 | g_loss: 1.8530\nEpoch [ 4/ 30] | d_loss: 1.0383 | g_loss: 1.6050\nEpoch [ 4/ 30] | d_loss: 0.7109 | g_loss: 1.3619\nEpoch [ 4/ 30] | d_loss: 0.9251 | g_loss: 0.8993\nEpoch [ 4/ 30] | d_loss: 0.9559 | g_loss: 1.3648\nEpoch [ 4/ 30] | d_loss: 0.8086 | g_loss: 1.5625\nEpoch [ 4/ 30] | d_loss: 0.9770 | g_loss: 1.6552\nEpoch [ 4/ 30] | d_loss: 0.6254 | g_loss: 1.6068\nEpoch [ 4/ 30] | d_loss: 0.8895 | g_loss: 1.6580\nEpoch [ 4/ 30] | d_loss: 1.1438 | g_loss: 0.7908\nEpoch [ 4/ 30] | d_loss: 0.8810 | g_loss: 1.1632\nEpoch [ 4/ 30] | d_loss: 0.9508 | g_loss: 0.8951\nEpoch [ 4/ 30] | d_loss: 0.8732 | g_loss: 1.3214\nEpoch [ 4/ 30] | d_loss: 0.8695 | g_loss: 1.7069\nEpoch [ 5/ 30] | d_loss: 0.7958 | g_loss: 1.2823\nEpoch [ 5/ 30] | d_loss: 0.8891 | g_loss: 1.3614\nEpoch [ 5/ 30] | d_loss: 0.8359 | g_loss: 1.5579\nEpoch [ 5/ 30] | d_loss: 0.8299 | g_loss: 0.9029\nEpoch [ 5/ 30] | d_loss: 0.8484 | g_loss: 1.4349\nEpoch [ 5/ 30] | d_loss: 0.8530 | g_loss: 1.2869\nEpoch [ 5/ 30] | d_loss: 1.3352 | g_loss: 0.9732\nEpoch [ 5/ 30] | d_loss: 0.8846 | g_loss: 1.8866\nEpoch [ 5/ 30] | d_loss: 0.7713 | g_loss: 1.2161\nEpoch [ 5/ 30] | d_loss: 0.8679 | g_loss: 1.4212\nEpoch [ 5/ 30] | d_loss: 0.8685 | g_loss: 1.4538\nEpoch [ 5/ 30] | d_loss: 1.0422 | g_loss: 0.8797\nEpoch [ 5/ 30] | d_loss: 1.0469 | g_loss: 2.3212\nEpoch [ 5/ 30] | d_loss: 0.6393 | g_loss: 1.9280\nEpoch [ 5/ 30] | d_loss: 0.8076 | g_loss: 1.0924\nEpoch [ 6/ 30] | d_loss: 0.7578 | g_loss: 1.7646\nEpoch [ 6/ 30] | d_loss: 0.8444 | g_loss: 1.8256\nEpoch [ 6/ 30] | d_loss: 0.7876 | g_loss: 0.9602\nEpoch [ 6/ 30] | d_loss: 0.7913 | g_loss: 1.4749\nEpoch [ 6/ 30] | d_loss: 0.8066 | g_loss: 1.8355\nEpoch [ 6/ 30] | d_loss: 0.7671 | g_loss: 1.3449\nEpoch [ 6/ 30] | d_loss: 0.8980 | g_loss: 1.3209\nEpoch [ 6/ 30] | d_loss: 0.8955 | g_loss: 1.3761\nEpoch [ 6/ 30] | d_loss: 0.6755 | g_loss: 1.5079\nEpoch [ 6/ 30] | d_loss: 0.6722 | g_loss: 1.7588\nEpoch [ 6/ 30] | d_loss: 0.8347 | g_loss: 1.2574\nEpoch [ 6/ 30] | d_loss: 0.8308 | g_loss: 2.1989\nEpoch [ 6/ 30] | d_loss: 0.6364 | g_loss: 0.8909\nEpoch [ 6/ 30] | d_loss: 0.9095 | g_loss: 1.7526\nEpoch [ 6/ 30] | d_loss: 0.7221 | g_loss: 2.0978\nEpoch [ 7/ 30] | d_loss: 0.6180 | g_loss: 1.8142\nEpoch [ 7/ 30] | d_loss: 0.7876 | g_loss: 1.2128\nEpoch [ 7/ 30] | d_loss: 0.6289 | g_loss: 1.6999\nEpoch [ 7/ 30] | d_loss: 0.4171 | g_loss: 1.9900\nEpoch [ 7/ 30] | d_loss: 0.7863 | g_loss: 1.2914\nEpoch [ 7/ 30] | d_loss: 0.8362 | g_loss: 1.5628\nEpoch [ 7/ 30] | d_loss: 0.5695 | g_loss: 2.0649\nEpoch [ 7/ 30] | d_loss: 0.9378 | g_loss: 1.0925\nEpoch [ 7/ 30] | d_loss: 0.7325 | g_loss: 1.8479\nEpoch [ 7/ 30] | d_loss: 0.6043 | g_loss: 1.6244\nEpoch [ 7/ 30] | d_loss: 1.0610 | g_loss: 0.5256\nEpoch [ 7/ 30] | d_loss: 0.7614 | g_loss: 1.3420\nEpoch [ 7/ 30] | d_loss: 1.0011 | g_loss: 0.6830\nEpoch [ 7/ 30] | d_loss: 0.8486 | g_loss: 1.7501\nEpoch [ 7/ 30] | d_loss: 0.6130 | g_loss: 1.8921\nEpoch [ 8/ 30] | d_loss: 0.9941 | g_loss: 0.7807\nEpoch [ 8/ 30] | d_loss: 0.7445 | g_loss: 1.5742\nEpoch [ 8/ 30] | d_loss: 0.9395 | g_loss: 1.2083\nEpoch [ 8/ 30] | d_loss: 0.6197 | g_loss: 2.0345\nEpoch [ 8/ 30] | d_loss: 0.7392 | g_loss: 1.7101\nEpoch [ 8/ 30] | d_loss: 0.6650 | g_loss: 0.9930\nEpoch [ 8/ 30] | d_loss: 0.8118 | g_loss: 1.6423\nEpoch [ 8/ 30] | d_loss: 0.8995 | g_loss: 0.8767\nEpoch [ 8/ 30] | d_loss: 1.3873 | g_loss: 2.3569\nEpoch [ 8/ 30] | d_loss: 0.8254 | g_loss: 1.6270\nEpoch [ 8/ 30] | d_loss: 0.6790 | g_loss: 1.7464\nEpoch [ 8/ 30] | d_loss: 0.7097 | g_loss: 1.2610\nEpoch [ 8/ 30] | d_loss: 1.1101 | g_loss: 1.8495\nEpoch [ 8/ 30] | d_loss: 0.7909 | g_loss: 1.2873\nEpoch [ 8/ 30] | d_loss: 0.6739 | g_loss: 1.1903\nEpoch [ 9/ 30] | d_loss: 0.6801 | g_loss: 2.4278\nEpoch [ 9/ 30] | d_loss: 0.7235 | g_loss: 2.0136\nEpoch [ 9/ 30] | d_loss: 1.1647 | g_loss: 0.6122\nEpoch [ 9/ 30] | d_loss: 0.7412 | g_loss: 1.2126\nEpoch [ 9/ 30] | d_loss: 0.6972 | g_loss: 1.7692\nEpoch [ 9/ 30] | d_loss: 1.1085 | g_loss: 1.1144\nEpoch [ 9/ 30] | d_loss: 0.5925 | g_loss: 2.7443\nEpoch [ 9/ 30] | d_loss: 0.9724 | g_loss: 1.3077\nEpoch [ 9/ 30] | d_loss: 0.5134 | g_loss: 1.7196\nEpoch [ 9/ 30] | d_loss: 0.7310 | g_loss: 1.4732\nEpoch [ 9/ 30] | d_loss: 0.7552 | g_loss: 1.4220\nEpoch [ 9/ 30] | d_loss: 0.7520 | g_loss: 0.8131\nEpoch [ 9/ 30] | d_loss: 0.6872 | g_loss: 1.4417\nEpoch [ 9/ 30] | d_loss: 1.5636 | g_loss: 3.1376\nEpoch [ 9/ 30] | d_loss: 0.7015 | g_loss: 1.5177\nEpoch [ 10/ 30] | d_loss: 0.7837 | g_loss: 2.0870\nEpoch [ 10/ 30] | d_loss: 0.5693 | g_loss: 1.8596\nEpoch [ 10/ 30] | d_loss: 0.9249 | g_loss: 2.2192\nEpoch [ 10/ 30] | d_loss: 0.6438 | g_loss: 1.5730\nEpoch [ 10/ 30] | d_loss: 1.3485 | g_loss: 3.7460\nEpoch [ 10/ 30] | d_loss: 1.0067 | g_loss: 1.3931\nEpoch [ 10/ 30] | d_loss: 0.5351 | g_loss: 1.6588\nEpoch [ 10/ 30] | d_loss: 1.1788 | g_loss: 2.7267\nEpoch [ 10/ 30] | d_loss: 0.8234 | g_loss: 2.3913\nEpoch [ 10/ 30] | d_loss: 0.6047 | g_loss: 1.6550\nEpoch [ 10/ 30] | d_loss: 0.9745 | g_loss: 0.6605\nEpoch [ 10/ 30] | d_loss: 1.0267 | g_loss: 2.0140\nEpoch [ 10/ 30] | d_loss: 0.5936 | g_loss: 2.0369\nEpoch [ 10/ 30] | d_loss: 0.6533 | g_loss: 1.9328\nEpoch [ 10/ 30] | d_loss: 0.7414 | g_loss: 1.5102\nEpoch [ 11/ 30] | d_loss: 0.9303 | g_loss: 1.9141\nEpoch [ 11/ 30] | d_loss: 0.8352 | g_loss: 1.5197\nEpoch [ 11/ 30] | d_loss: 0.7432 | g_loss: 1.1725\nEpoch [ 11/ 30] | d_loss: 0.7125 | g_loss: 1.3522\nEpoch [ 11/ 30] | d_loss: 0.6534 | g_loss: 1.3864\nEpoch [ 11/ 30] | d_loss: 0.3060 | g_loss: 3.0363\nEpoch [ 11/ 30] | d_loss: 0.6160 | g_loss: 1.8028\nEpoch [ 11/ 30] | d_loss: 1.5896 | g_loss: 1.0561\nEpoch [ 11/ 30] | d_loss: 0.7031 | g_loss: 1.6583\nEpoch [ 11/ 30] | d_loss: 0.7459 | g_loss: 1.5477\nEpoch [ 11/ 30] | d_loss: 0.4981 | g_loss: 2.3105\nEpoch [ 11/ 30] | d_loss: 0.6857 | g_loss: 1.5724\nEpoch [ 11/ 30] | d_loss: 0.7403 | g_loss: 2.9962\nEpoch [ 11/ 30] | d_loss: 0.6803 | g_loss: 1.1301\nEpoch [ 11/ 30] | d_loss: 0.7613 | g_loss: 1.4520\nEpoch [ 12/ 30] | d_loss: 0.7079 | g_loss: 2.0324\nEpoch [ 12/ 30] | d_loss: 0.7041 | g_loss: 1.3625\nEpoch [ 12/ 30] | d_loss: 0.6362 | g_loss: 2.3640\nEpoch [ 12/ 30] | d_loss: 0.6313 | g_loss: 1.5870\nEpoch [ 12/ 30] | d_loss: 0.7057 | g_loss: 2.3751\nEpoch [ 12/ 30] | d_loss: 0.5797 | g_loss: 2.0853\nEpoch [ 12/ 30] | d_loss: 0.8887 | g_loss: 2.3426\nEpoch [ 12/ 30] | d_loss: 0.7609 | g_loss: 2.5219\nEpoch [ 12/ 30] | d_loss: 0.6884 | g_loss: 1.4588\nEpoch [ 12/ 30] | d_loss: 0.9216 | g_loss: 0.9844\nEpoch [ 12/ 30] | d_loss: 0.6312 | g_loss: 1.9144\nEpoch [ 12/ 30] | d_loss: 0.6354 | g_loss: 1.5764\nEpoch [ 12/ 30] | d_loss: 0.6510 | g_loss: 1.7851\nEpoch [ 12/ 30] | d_loss: 0.6314 | g_loss: 1.3376\nEpoch [ 12/ 30] | d_loss: 0.4935 | g_loss: 2.4079\nEpoch [ 13/ 30] | d_loss: 0.4287 | g_loss: 2.8181\nEpoch [ 13/ 30] | d_loss: 0.5462 | g_loss: 2.0479\nEpoch [ 13/ 30] | d_loss: 0.8311 | g_loss: 1.3864\nEpoch [ 13/ 30] | d_loss: 0.9235 | g_loss: 1.2160\nEpoch [ 13/ 30] | d_loss: 0.5888 | g_loss: 1.5458\nEpoch [ 13/ 30] | d_loss: 0.7615 | g_loss: 1.9504\nEpoch [ 13/ 30] | d_loss: 0.6898 | g_loss: 1.9105\nEpoch [ 13/ 30] | d_loss: 0.6458 | g_loss: 2.1635\nEpoch [ 13/ 30] | d_loss: 0.7348 | g_loss: 0.9890\nEpoch [ 13/ 30] | d_loss: 0.7202 | g_loss: 1.8480\nEpoch [ 13/ 30] | d_loss: 0.7972 | g_loss: 2.3036\nEpoch [ 13/ 30] | d_loss: 0.8759 | g_loss: 1.3286\nEpoch [ 13/ 30] | d_loss: 0.7247 | g_loss: 1.3945\nEpoch [ 13/ 30] | d_loss: 1.0758 | g_loss: 3.0929\nEpoch [ 13/ 30] | d_loss: 0.6960 | g_loss: 1.5643\nEpoch [ 14/ 30] | d_loss: 0.5979 | g_loss: 1.4264\nEpoch [ 14/ 30] | d_loss: 0.5545 | g_loss: 2.0683\nEpoch [ 14/ 30] | d_loss: 0.5697 | g_loss: 1.9833\nEpoch [ 14/ 30] | d_loss: 0.6994 | g_loss: 2.3122\nEpoch [ 14/ 30] | d_loss: 1.0690 | g_loss: 0.7694\nEpoch [ 14/ 30] | d_loss: 0.7119 | g_loss: 1.5692\nEpoch [ 14/ 30] | d_loss: 0.6420 | g_loss: 2.1430\nEpoch [ 14/ 30] | d_loss: 0.5274 | g_loss: 1.4330\nEpoch [ 14/ 30] | d_loss: 0.5784 | g_loss: 2.0117\nEpoch [ 14/ 30] | d_loss: 0.7762 | g_loss: 2.0491\nEpoch [ 14/ 30] | d_loss: 0.6158 | g_loss: 1.0900\nEpoch [ 14/ 30] | d_loss: 0.5262 | g_loss: 2.0739\nEpoch [ 14/ 30] | d_loss: 0.6864 | g_loss: 2.5591\nEpoch [ 14/ 30] | d_loss: 0.4090 | g_loss: 2.9574\nEpoch [ 14/ 30] | d_loss: 0.5823 | g_loss: 1.9607\nEpoch [ 15/ 30] | d_loss: 0.3979 | g_loss: 2.1985\nEpoch [ 15/ 30] | d_loss: 0.6133 | g_loss: 2.1405\nEpoch [ 15/ 30] | d_loss: 0.6361 | g_loss: 1.7988\nEpoch [ 15/ 30] | d_loss: 0.5247 | g_loss: 1.3331\nEpoch [ 15/ 30] | d_loss: 0.7407 | g_loss: 1.7587\nEpoch [ 15/ 30] | d_loss: 0.5080 | g_loss: 2.4057\nEpoch [ 15/ 30] | d_loss: 0.5568 | g_loss: 1.5972\nEpoch [ 15/ 30] | d_loss: 0.6365 | g_loss: 1.3802\nEpoch [ 15/ 30] | d_loss: 0.5128 | g_loss: 2.2809\nEpoch [ 15/ 30] | d_loss: 0.6258 | g_loss: 1.9247\nEpoch [ 15/ 30] | d_loss: 0.6968 | g_loss: 1.9209\nEpoch [ 15/ 30] | d_loss: 0.7507 | g_loss: 2.8442\nEpoch [ 15/ 30] | d_loss: 0.4992 | g_loss: 2.6931\nEpoch [ 15/ 30] | d_loss: 0.4745 | g_loss: 2.0748\nEpoch [ 15/ 30] | d_loss: 0.6182 | g_loss: 2.1427\nEpoch [ 16/ 30] | d_loss: 0.5251 | g_loss: 1.7848\nEpoch [ 16/ 30] | d_loss: 0.3451 | g_loss: 2.0850\nEpoch [ 16/ 30] | d_loss: 0.6450 | g_loss: 2.6483\nEpoch [ 16/ 30] | d_loss: 0.5058 | g_loss: 1.6648\nEpoch [ 16/ 30] | d_loss: 0.7584 | g_loss: 1.7105\nEpoch [ 16/ 30] | d_loss: 0.7268 | g_loss: 2.1054\nEpoch [ 16/ 30] | d_loss: 0.5758 | g_loss: 2.3122\nEpoch [ 16/ 30] | d_loss: 1.2829 | g_loss: 5.2258\nEpoch [ 16/ 30] | d_loss: 0.5789 | g_loss: 2.3232\nEpoch [ 16/ 30] | d_loss: 0.7772 | g_loss: 1.1027\nEpoch [ 16/ 30] | d_loss: 0.5930 | g_loss: 1.2272\nEpoch [ 16/ 30] | d_loss: 0.4110 | g_loss: 2.0537\nEpoch [ 16/ 30] | d_loss: 0.6153 | g_loss: 1.6840\nEpoch [ 16/ 30] | d_loss: 0.8289 | g_loss: 2.7737\nEpoch [ 16/ 30] | d_loss: 0.4774 | g_loss: 2.8046\nEpoch [ 17/ 30] | d_loss: 0.4004 | g_loss: 2.0718\nEpoch [ 17/ 30] | d_loss: 0.8870 | g_loss: 3.6319\nEpoch [ 17/ 30] | d_loss: 0.9262 | g_loss: 2.5571\nEpoch [ 17/ 30] | d_loss: 0.5489 | g_loss: 2.5156\nEpoch [ 17/ 30] | d_loss: 0.6366 | g_loss: 2.9944\nEpoch [ 17/ 30] | d_loss: 0.3850 | g_loss: 1.8714\nEpoch [ 17/ 30] | d_loss: 0.4545 | g_loss: 2.0131\nEpoch [ 17/ 30] | d_loss: 1.5676 | g_loss: 3.4330\nEpoch [ 17/ 30] | d_loss: 0.5726 | g_loss: 1.5250\nEpoch [ 17/ 30] | d_loss: 0.4701 | g_loss: 2.1812\nEpoch [ 17/ 30] | d_loss: 0.5639 | g_loss: 1.7902\nEpoch [ 17/ 30] | d_loss: 0.3588 | g_loss: 1.7634\nEpoch [ 17/ 30] | d_loss: 0.6023 | g_loss: 2.9187\nEpoch [ 17/ 30] | d_loss: 0.6150 | g_loss: 1.1369\nEpoch [ 17/ 30] | d_loss: 0.4944 | g_loss: 2.0992\nEpoch [ 18/ 30] | d_loss: 0.4691 | g_loss: 2.4029\nEpoch [ 18/ 30] | d_loss: 0.3530 | g_loss: 2.6878\nEpoch [ 18/ 30] | d_loss: 0.5206 | g_loss: 1.7949\nEpoch [ 18/ 30] | d_loss: 0.3608 | g_loss: 2.3984\nEpoch [ 18/ 30] | d_loss: 0.4232 | g_loss: 2.0650\nEpoch [ 18/ 30] | d_loss: 1.2758 | g_loss: 0.3904\nEpoch [ 18/ 30] | d_loss: 0.4594 | g_loss: 1.8284\nEpoch [ 18/ 30] | d_loss: 0.5809 | g_loss: 1.9485\nEpoch [ 18/ 30] | d_loss: 0.3564 | g_loss: 1.8635\nEpoch [ 18/ 30] | d_loss: 0.3782 | g_loss: 1.6940\nEpoch [ 18/ 30] | d_loss: 0.6269 | g_loss: 1.8278\nEpoch [ 18/ 30] | d_loss: 0.8270 | g_loss: 0.5178\nEpoch [ 18/ 30] | d_loss: 0.4582 | g_loss: 2.1388\nEpoch [ 18/ 30] | d_loss: 0.7613 | g_loss: 1.0630\nEpoch [ 18/ 30] | d_loss: 0.5573 | g_loss: 2.0574\nEpoch [ 19/ 30] | d_loss: 0.5382 | g_loss: 2.3297\nEpoch [ 19/ 30] | d_loss: 0.5411 | g_loss: 2.4908\nEpoch [ 19/ 30] | d_loss: 0.4502 | g_loss: 1.7289\nEpoch [ 19/ 30] | d_loss: 0.5277 | g_loss: 2.9410\nEpoch [ 19/ 30] | d_loss: 0.4364 | g_loss: 2.0092\nEpoch [ 19/ 30] | d_loss: 0.4675 | g_loss: 2.7906\nEpoch [ 19/ 30] | d_loss: 0.5662 | g_loss: 1.6854\nEpoch [ 19/ 30] | d_loss: 0.4590 | g_loss: 2.4875\nEpoch [ 19/ 30] | d_loss: 0.4835 | g_loss: 2.1207\nEpoch [ 19/ 30] | d_loss: 0.3629 | g_loss: 2.8688\nEpoch [ 19/ 30] | d_loss: 0.6198 | g_loss: 4.2237\nEpoch [ 19/ 30] | d_loss: 0.3589 | g_loss: 2.0915\nEpoch [ 19/ 30] | d_loss: 1.5636 | g_loss: 0.9197\nEpoch [ 19/ 30] | d_loss: 0.5597 | g_loss: 2.2806\nEpoch [ 19/ 30] | d_loss: 0.4364 | g_loss: 2.2716\nEpoch [ 20/ 30] | d_loss: 0.5508 | g_loss: 1.2444\nEpoch [ 20/ 30] | d_loss: 0.5939 | g_loss: 2.1055\nEpoch [ 20/ 30] | d_loss: 0.5943 | g_loss: 2.1396\nEpoch [ 20/ 30] | d_loss: 0.8465 | g_loss: 3.4011\nEpoch [ 20/ 30] | d_loss: 0.4697 | g_loss: 2.8818\nEpoch [ 20/ 30] | d_loss: 0.5065 | g_loss: 2.1212\nEpoch [ 20/ 30] | d_loss: 0.4908 | g_loss: 1.9426\nEpoch [ 20/ 30] | d_loss: 0.3282 | g_loss: 2.1905\nEpoch [ 20/ 30] | d_loss: 0.4438 | g_loss: 1.8376\nEpoch [ 20/ 30] | d_loss: 0.4167 | g_loss: 2.1036\nEpoch [ 20/ 30] | d_loss: 0.6362 | g_loss: 1.4069\nEpoch [ 20/ 30] | d_loss: 0.4037 | g_loss: 1.8872\nEpoch [ 20/ 30] | d_loss: 0.5257 | g_loss: 2.2197\nEpoch [ 20/ 30] | d_loss: 0.5313 | g_loss: 3.0965\nEpoch [ 20/ 30] | d_loss: 0.5338 | g_loss: 2.3459\nEpoch [ 21/ 30] | d_loss: 0.5304 | g_loss: 2.3069\nEpoch [ 21/ 30] | d_loss: 0.4814 | g_loss: 2.1149\nEpoch [ 21/ 30] | d_loss: 0.6094 | g_loss: 1.8555\nEpoch [ 21/ 30] | d_loss: 0.4614 | g_loss: 2.0587\nEpoch [ 21/ 30] | d_loss: 0.7324 | g_loss: 2.2047\nEpoch [ 21/ 30] | d_loss: 0.4101 | g_loss: 2.3126\nEpoch [ 21/ 30] | d_loss: 0.9995 | g_loss: 2.0750\nEpoch [ 21/ 30] | d_loss: 0.4927 | g_loss: 1.7313\nEpoch [ 21/ 30] | d_loss: 0.4676 | g_loss: 1.7221\nEpoch [ 21/ 30] | d_loss: 0.5605 | g_loss: 1.2959\nEpoch [ 21/ 30] | d_loss: 0.6047 | g_loss: 1.5653\nEpoch [ 21/ 30] | d_loss: 0.5869 | g_loss: 2.2616\nEpoch [ 21/ 30] | d_loss: 0.4276 | g_loss: 2.1825\nEpoch [ 21/ 30] | d_loss: 0.6513 | g_loss: 1.6240\nEpoch [ 21/ 30] | d_loss: 0.4333 | g_loss: 2.3750\nEpoch [ 22/ 30] | d_loss: 0.4138 | g_loss: 1.6811\nEpoch [ 22/ 30] | d_loss: 0.4872 | g_loss: 2.3072\nEpoch [ 22/ 30] | d_loss: 0.4178 | g_loss: 1.3897\nEpoch [ 22/ 30] | d_loss: 0.5082 | g_loss: 2.0710\nEpoch [ 22/ 30] | d_loss: 0.4212 | g_loss: 2.5157\nEpoch [ 22/ 30] | d_loss: 0.4919 | g_loss: 2.9226\nEpoch [ 22/ 30] | d_loss: 0.8077 | g_loss: 2.4910\nEpoch [ 22/ 30] | d_loss: 0.3188 | g_loss: 2.4105\nEpoch [ 22/ 30] | d_loss: 0.6110 | g_loss: 2.1661\nEpoch [ 22/ 30] | d_loss: 0.4050 | g_loss: 2.4285\nEpoch [ 22/ 30] | d_loss: 0.4281 | g_loss: 2.1288\nEpoch [ 22/ 30] | d_loss: 0.4589 | g_loss: 2.3186\nEpoch [ 22/ 30] | d_loss: 0.5547 | g_loss: 1.3391\nEpoch [ 22/ 30] | d_loss: 0.5777 | g_loss: 3.2322\nEpoch [ 22/ 30] | d_loss: 0.7990 | g_loss: 3.8977\nEpoch [ 23/ 30] | d_loss: 0.4732 | g_loss: 2.3594\nEpoch [ 23/ 30] | d_loss: 0.3833 | g_loss: 2.5015\nEpoch [ 23/ 30] | d_loss: 0.4098 | g_loss: 1.8201\nEpoch [ 23/ 30] | d_loss: 0.3216 | g_loss: 2.3112\nEpoch [ 23/ 30] | d_loss: 0.5150 | g_loss: 2.0603\nEpoch [ 23/ 30] | d_loss: 0.3562 | g_loss: 2.8514\nEpoch [ 23/ 30] | d_loss: 0.2769 | g_loss: 2.1134\nEpoch [ 23/ 30] | d_loss: 0.5669 | g_loss: 2.3579\nEpoch [ 23/ 30] | d_loss: 0.4093 | g_loss: 2.3857\nEpoch [ 23/ 30] | d_loss: 0.8305 | g_loss: 3.2275\nEpoch [ 23/ 30] | d_loss: 0.4100 | g_loss: 2.4538\nEpoch [ 23/ 30] | d_loss: 0.4883 | g_loss: 1.8107\nEpoch [ 23/ 30] | d_loss: 0.3157 | g_loss: 2.7718\nEpoch [ 23/ 30] | d_loss: 0.9923 | g_loss: 2.7018\nEpoch [ 23/ 30] | d_loss: 0.5470 | g_loss: 2.9502\nEpoch [ 24/ 30] | d_loss: 0.4453 | g_loss: 2.2249\nEpoch [ 24/ 30] | d_loss: 0.4172 | g_loss: 2.1866\nEpoch [ 24/ 30] | d_loss: 0.4395 | g_loss: 2.4455\nEpoch [ 24/ 30] | d_loss: 1.4186 | g_loss: 1.2122\nEpoch [ 24/ 30] | d_loss: 0.5332 | g_loss: 3.1165\nEpoch [ 24/ 30] | d_loss: 0.5177 | g_loss: 2.7426\nEpoch [ 24/ 30] | d_loss: 0.4693 | g_loss: 3.4348\nEpoch [ 24/ 30] | d_loss: 0.2868 | g_loss: 3.1014\nEpoch [ 24/ 30] | d_loss: 0.5197 | g_loss: 1.8435\nEpoch [ 24/ 30] | d_loss: 0.3607 | g_loss: 2.3998\nEpoch [ 24/ 30] | d_loss: 0.3242 | g_loss: 2.5870\nEpoch [ 24/ 30] | d_loss: 0.3620 | g_loss: 1.9227\nEpoch [ 24/ 30] | d_loss: 1.7177 | g_loss: 0.4907\nEpoch [ 24/ 30] | d_loss: 0.3537 | g_loss: 2.3476\nEpoch [ 24/ 30] | d_loss: 0.4408 | g_loss: 1.9916\nEpoch [ 25/ 30] | d_loss: 0.5609 | g_loss: 1.6246\nEpoch [ 25/ 30] | d_loss: 0.4828 | g_loss: 1.9648\nEpoch [ 25/ 30] | d_loss: 0.3045 | g_loss: 2.4795\nEpoch [ 25/ 30] | d_loss: 0.3963 | g_loss: 2.0227\nEpoch [ 25/ 30] | d_loss: 0.3628 | g_loss: 3.0971\nEpoch [ 25/ 30] | d_loss: 0.3837 | g_loss: 1.7916\nEpoch [ 25/ 30] | d_loss: 0.3997 | g_loss: 2.2200\nEpoch [ 25/ 30] | d_loss: 0.9150 | g_loss: 4.6989\nEpoch [ 25/ 30] | d_loss: 0.5928 | g_loss: 1.4060\nEpoch [ 25/ 30] | d_loss: 0.3325 | g_loss: 2.2855\nEpoch [ 25/ 30] | d_loss: 0.4278 | g_loss: 1.7970\nEpoch [ 25/ 30] | d_loss: 0.3145 | g_loss: 2.0559\nEpoch [ 25/ 30] | d_loss: 0.2875 | g_loss: 3.1949\nEpoch [ 25/ 30] | d_loss: 0.4414 | g_loss: 2.3271\nEpoch [ 25/ 30] | d_loss: 0.2951 | g_loss: 2.8286\nEpoch [ 26/ 30] | d_loss: 0.2724 | g_loss: 3.7023\nEpoch [ 26/ 30] | d_loss: 0.4425 | g_loss: 2.9363\nEpoch [ 26/ 30] | d_loss: 0.4563 | g_loss: 2.7358\nEpoch [ 26/ 30] | d_loss: 0.5567 | g_loss: 3.0356\nEpoch [ 26/ 30] | d_loss: 0.4627 | g_loss: 3.0882\nEpoch [ 26/ 30] | d_loss: 0.8079 | g_loss: 3.8006\nEpoch [ 26/ 30] | d_loss: 0.2950 | g_loss: 2.8960\nEpoch [ 26/ 30] | d_loss: 0.3856 | g_loss: 3.2416\nEpoch [ 26/ 30] | d_loss: 0.4347 | g_loss: 2.2718\nEpoch [ 26/ 30] | d_loss: 0.2099 | g_loss: 2.9685\nEpoch [ 26/ 30] | d_loss: 0.4712 | g_loss: 3.0447\nEpoch [ 26/ 30] | d_loss: 0.4272 | g_loss: 3.1004\nEpoch [ 26/ 30] | d_loss: 0.5921 | g_loss: 2.3868\nEpoch [ 26/ 30] | d_loss: 0.3242 | g_loss: 2.7287\nEpoch [ 26/ 30] | d_loss: 0.2957 | g_loss: 2.5311\nEpoch [ 27/ 30] | d_loss: 0.6152 | g_loss: 3.0290\nEpoch [ 27/ 30] | d_loss: 0.4221 | g_loss: 2.2556\nEpoch [ 27/ 30] | d_loss: 0.4677 | g_loss: 3.0501\nEpoch [ 27/ 30] | d_loss: 2.8618 | g_loss: 4.9644\nEpoch [ 27/ 30] | d_loss: 0.3824 | g_loss: 2.3956\nEpoch [ 27/ 30] | d_loss: 0.4689 | g_loss: 2.5165\nEpoch [ 27/ 30] | d_loss: 0.4342 | g_loss: 1.9378\nEpoch [ 27/ 30] | d_loss: 0.2807 | g_loss: 2.6148\nEpoch [ 27/ 30] | d_loss: 0.3915 | g_loss: 1.4882\nEpoch [ 27/ 30] | d_loss: 0.3102 | g_loss: 2.0836\nEpoch [ 27/ 30] | d_loss: 0.4229 | g_loss: 2.5760\nEpoch [ 27/ 30] | d_loss: 0.4256 | g_loss: 1.8691\nEpoch [ 27/ 30] | d_loss: 0.4436 | g_loss: 3.3224\nEpoch [ 27/ 30] | d_loss: 0.3826 | g_loss: 2.2536\nEpoch [ 27/ 30] | d_loss: 0.4103 | g_loss: 1.8520\nEpoch [ 28/ 30] | d_loss: 0.3645 | g_loss: 3.1014\nEpoch [ 28/ 30] | d_loss: 0.3970 | g_loss: 2.7173\nEpoch [ 28/ 30] | d_loss: 2.2226 | g_loss: 0.4895\nEpoch [ 28/ 30] | d_loss: 0.4051 | g_loss: 2.7626\nEpoch [ 28/ 30] | d_loss: 0.2715 | g_loss: 3.8289\nEpoch [ 28/ 30] | d_loss: 0.2806 | g_loss: 2.5903\nEpoch [ 28/ 30] | d_loss: 0.4260 | g_loss: 2.8684\nEpoch [ 28/ 30] | d_loss: 0.2945 | g_loss: 1.5078\nEpoch [ 28/ 30] | d_loss: 0.4544 | g_loss: 2.0769\nEpoch [ 28/ 30] | d_loss: 1.5290 | g_loss: 4.4883\nEpoch [ 28/ 30] | d_loss: 0.2125 | g_loss: 3.1354\nEpoch [ 28/ 30] | d_loss: 0.3341 | g_loss: 2.7934\nEpoch [ 28/ 30] | d_loss: 0.5580 | g_loss: 2.1050\nEpoch [ 28/ 30] | d_loss: 0.4864 | g_loss: 1.2127\nEpoch [ 28/ 30] | d_loss: 0.3274 | g_loss: 2.7572\nEpoch [ 29/ 30] | d_loss: 0.4310 | g_loss: 2.0522\nEpoch [ 29/ 30] | d_loss: 0.2594 | g_loss: 2.0715\nEpoch [ 29/ 30] | d_loss: 0.3404 | g_loss: 2.5484\nEpoch [ 29/ 30] | d_loss: 0.2448 | g_loss: 2.6435\nEpoch [ 29/ 30] | d_loss: 0.3009 | g_loss: 2.4528\nEpoch [ 29/ 30] | d_loss: 0.5283 | g_loss: 2.6000\nEpoch [ 29/ 30] | d_loss: 0.3303 | g_loss: 2.8223\nEpoch [ 29/ 30] | d_loss: 1.9013 | g_loss: 1.9974\nEpoch [ 29/ 30] | d_loss: 0.5584 | g_loss: 1.6161\nEpoch [ 29/ 30] | d_loss: 0.2464 | g_loss: 2.3172\nEpoch [ 29/ 30] | d_loss: 0.3265 | g_loss: 2.1264\nEpoch [ 29/ 30] | d_loss: 0.2290 | g_loss: 2.7484\nEpoch [ 29/ 30] | d_loss: 0.2461 | g_loss: 2.9240\nEpoch [ 29/ 30] | d_loss: 0.3551 | g_loss: 2.5152\nEpoch [ 29/ 30] | d_loss: 0.3922 | g_loss: 1.2241\nEpoch [ 30/ 30] | d_loss: 0.2428 | g_loss: 1.8990\nEpoch [ 30/ 30] | d_loss: 0.2943 | g_loss: 2.8686\nEpoch [ 30/ 30] | d_loss: 1.7308 | g_loss: 0.6362\nEpoch [ 30/ 30] | d_loss: 0.3615 | g_loss: 2.4141\nEpoch [ 30/ 30] | d_loss: 0.5358 | g_loss: 1.5713\nEpoch [ 30/ 30] | d_loss: 7.1501 | g_loss: 3.9271\nEpoch [ 30/ 30] | d_loss: 0.4100 | g_loss: 3.7686\nEpoch [ 30/ 30] | d_loss: 0.4251 | g_loss: 2.0842\nEpoch [ 30/ 30] | d_loss: 0.3061 | g_loss: 2.7882\nEpoch [ 30/ 30] | d_loss: 0.3089 | g_loss: 2.0874\nEpoch [ 30/ 30] | d_loss: 0.7876 | g_loss: 1.9367\nEpoch [ 30/ 30] | d_loss: 0.4341 | g_loss: 1.7241\nEpoch [ 30/ 30] | d_loss: 0.2160 | g_loss: 2.9003\nEpoch [ 30/ 30] | d_loss: 0.4238 | g_loss: 3.8263\nEpoch [ 30/ 30] | d_loss: 0.1706 | g_loss: 3.4528\n"
]
],
[
[
"## Training loss\n\nPlot the training losses for the generator and discriminator, recorded after each epoch.",
"_____no_output_____"
]
],
[
[
"fig, ax = plt.subplots()\nlosses = np.array(losses)\nplt.plot(losses.T[0], label='Discriminator', alpha=0.5)\nplt.plot(losses.T[1], label='Generator', alpha=0.5)\nplt.title(\"Training Losses\")\nplt.legend()",
"_____no_output_____"
]
],
[
[
"## Generator samples from training\n\nView samples of images from the generator, and answer a question about the strengths and weaknesses of your trained models.",
"_____no_output_____"
]
],
[
[
"# helper function for viewing a list of passed in sample images\ndef view_samples(epoch, samples):\n fig, axes = plt.subplots(figsize=(16,4), nrows=2, ncols=8, sharey=True, sharex=True)\n for ax, img in zip(axes.flatten(), samples[epoch]):\n img = img.detach().cpu().numpy()\n img = np.transpose(img, (1, 2, 0))\n img = ((img + 1)*255 / (2)).astype(np.uint8)\n ax.xaxis.set_visible(False)\n ax.yaxis.set_visible(False)\n im = ax.imshow(img.reshape((32,32,3)))",
"_____no_output_____"
],
[
"# Load samples from generator, taken while training\nwith open('train_samples.pkl', 'rb') as f:\n samples = pkl.load(f)",
"_____no_output_____"
],
[
"_ = view_samples(-1, samples)",
"_____no_output_____"
]
],
[
[
"### Question: What do you notice about your generated samples and how might you improve this model?\nWhen you answer this question, consider the following factors:\n* The dataset is biased; it is made of \"celebrity\" faces that are mostly white\n* Model size; larger models have the opportunity to learn more features in a data feature space\n* Optimization strategy; optimizers and number of epochs affect your final result\n",
"_____no_output_____"
],
[
"**Answer:** I think a deeper architecture esp for the Generator might have helped. I played with a lot of different values of the hyperparameters, but the losses weren't going down that much after a point. Also, the network might have learnt a bit better if there was more data. A lot of the times, the images that were generated had odd features like half a black frame on the face or a piece of a microphone etc. ",
"_____no_output_____"
],
[
"### Submitting This Project\nWhen submitting this project, make sure to run all the cells before saving the notebook. Save the notebook file as \"dlnd_face_generation.ipynb\" and save it as a HTML file under \"File\" -> \"Download as\". Include the \"problem_unittests.py\" files in your submission.",
"_____no_output_____"
]
]
] | [
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown"
] | [
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown",
"markdown",
"markdown"
]
] |
d0ff550e4442387f5cdb402f87432f5d692496d9 | 66,874 | ipynb | Jupyter Notebook | arnheim_1.ipynb | TotoArt/arnheim | 48b82d10210350c12835f0d00c7114301b406a73 | [
"Apache-2.0"
] | 186 | 2021-11-03T15:01:55.000Z | 2022-03-28T04:30:48.000Z | arnheim_1.ipynb | TotoArt/arnheim | 48b82d10210350c12835f0d00c7114301b406a73 | [
"Apache-2.0"
] | 6 | 2021-11-03T20:29:49.000Z | 2022-03-18T18:02:04.000Z | arnheim_1.ipynb | TotoArt/arnheim | 48b82d10210350c12835f0d00c7114301b406a73 | [
"Apache-2.0"
] | 29 | 2021-11-03T20:28:23.000Z | 2022-03-16T16:55:37.000Z | 44.316766 | 497 | 0.511439 | [
[
[
"Copyright 2021 DeepMind Technologies Limited\n\nLicensed under the Apache License, Version 2.0 (the \"License\");\nyou may not use this file except in compliance with the License.\nYou may obtain a copy of the License at\n\n https://www.apache.org/licenses/LICENSE-2.0\n\nUnless required by applicable law or agreed to in writing, software\ndistributed under the License is distributed on an \"AS IS\" BASIS,\nWITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.\nSee the License for the specific language governing permissions and\nlimitations under the License.",
"_____no_output_____"
],
[
"#Generative Art Using Neural Visual Grammars and Dual Encoders\n\n**Chrisantha Fernando, Piotr Mirowski, Dylan Banarse, S. M. Ali Eslami, Jean-Baptiste Alayrac, Simon Osindero**\n\nDeepMind, 2021\n\n##Arnheim 1\n###Generate paintings from text prompts.\n\n Whilst there are perhaps only a few scientific methods, there seem to be almost as many artistic methods as there are artists. Artistic processes appear to inhabit the highest order of open-endedness. To begin to understand some of the processes of art making it is helpful to try to automate them even partially.\nIn this paper, a novel algorithm for producing generative art is described which allows a user to input a text string, and which in a creative response to this string, outputs an image which interprets that string. It does so by evolving images using a hierarchical neural [Lindenmeyer system](https://en.wikipedia.org/wiki/L-system), and evaluating these images along the way using an image text dual encoder trained on billions of images and their associated text from the internet.\nIn doing so we have access to and control over an instance of an artistic process, allowing analysis of which aspects of the artistic process become the task of the algorithm, and which elements remain the responsibility of the artist.\n\nThis colab accompanies the paper [Generative Art Using Neural Visual Grammars and Dual Encoders](https://arxiv.org/abs/2105.00162)\n\n##Instructions\n\n1. Click \"Connect\" button in the top right corner of this Colab\n1. Select Runtime -> Change runtime type -> Hardware accelerator -> GPU\n1. Select High-RAM for \"Runtime shape\" option\n1. Navigate to \"Get text input\"\n1. Enter text for IMAGE_NAME\n1. Select \"Run All\" from Runtime menu\n",
"_____no_output_____"
],
[
"# Imports",
"_____no_output_____"
]
],
[
[
"#@title Set CUDA version for PyTorch\n\nimport subprocess\n\nCUDA_version = [s for s in subprocess.check_output([\"nvcc\", \"--version\"]\n ).decode(\"UTF-8\").split(\", \")\n if s.startswith(\"release\")][0].split(\" \")[-1]\nprint(\"CUDA version:\", CUDA_version)\n\nif CUDA_version == \"10.0\":\n torch_version_suffix = \"+cu100\"\nelif CUDA_version == \"10.1\":\n torch_version_suffix = \"+cu101\"\nelif CUDA_version == \"10.2\":\n torch_version_suffix = \"\"\nelse:\n torch_version_suffix = \"+cu110\"\n! nvidia-smi",
"_____no_output_____"
],
[
"#@title Install and import PyTorch and Clip\n\n! pip install torch==1.7.1{torch_version_suffix} torchvision==0.8.2{torch_version_suffix} -f https://download.pytorch.org/whl/torch_stable.html\n! pip install git+https://github.com/openai/CLIP.git --no-deps\n! pip install ftfy regex\n\nimport torch\nimport torch.nn as nn\nimport clip\nprint(\"Torch version:\", torch.__version__)",
"_____no_output_____"
],
[
"#@title Install and import ray multiprocessing\n\n! pip install -q -U ray[default]\nimport ray",
"_____no_output_____"
],
[
"#@title Import all other needed libraries\n\nimport collections\nimport copy\nimport cloudpickle\nimport time\nimport numpy as np\nimport matplotlib.pyplot as plt\nimport math\nfrom PIL import Image\nfrom PIL import ImageDraw\nfrom skimage import transform",
"_____no_output_____"
],
[
"#@title Load CLIP {vertical-output: true}\n\nCLIP_MODEL = \"ViT-B/32\"\ndevice = torch.device(\"cuda\")\nprint(f\"Downloading CLIP model {CLIP_MODEL}...\")\nmodel, _ = clip.load(CLIP_MODEL, device, jit=False)",
"_____no_output_____"
]
],
[
[
"# Neural Visual Grammar",
"_____no_output_____"
],
[
"### Drawing primitives",
"_____no_output_____"
]
],
[
[
"def to_homogeneous(p):\n r, c = p\n return np.stack((r, c, np.ones_like(p[0])), axis=0)\n\ndef from_homogeneous(p):\n p = p / p.T[:, 2]\n return p[0].astype(\"int32\"), p[1].astype(\"int32\")\n\ndef apply_scale(scale, lineh):\n return np.stack([lineh[0, :] * scale,\n lineh[1, :] * scale,\n lineh[2, :]])\n\ndef apply_translation(translation, lineh, offset_r=0, offset_c=0):\n r, c = translation\n return np.stack([lineh[0, :] + c + offset_c,\n lineh[1, :] + r + offset_r,\n lineh[2, :]])\n\ndef apply_rotation(translation, rad, lineh):\n r, c = translation\n cos_rad = np.cos(rad)\n sin_rad = np.sin(rad)\n return np.stack(\n [(lineh[0, :] - c) * cos_rad - (lineh[1, :] - r) * sin_rad + c,\n (lineh[0, :] - c) * sin_rad + (lineh[1, :] - r) * cos_rad + r,\n lineh[2, :]])",
"_____no_output_____"
],
[
"def transform_lines(line_from, line_to, translation, angle, scale,\n translation2, angle2, scale2, img_siz2):\n \"\"\"Transform lines by translation, angle and scale, twice.\n\n Args:\n line_from: Line start point.\n line_to: Line end point.\n translation: 1st translation to line.\n angle: 1st angle of rotation for line.\n scale: 1st scale for line.\n translation2: 2nd translation to line.\n angle2: 2nd angle of rotation for line.\n scale2: 2nd scale for line.\n img_siz2: Offset for 2nd translation.\n\n Returns:\n Transformed lines.\n \"\"\"\n if len(line_from.shape) == 1:\n line_from = np.expand_dims(line_from, 0)\n if len(line_to.shape) == 1:\n line_to = np.expand_dims(line_to, 0)\n\n # First transform.\n line_from_h = to_homogeneous(line_from.T)\n line_to_h = to_homogeneous(line_to.T)\n line_from_h = apply_scale(scale, line_from_h)\n line_to_h = apply_scale(scale, line_to_h)\n translated_line_from = apply_translation(translation, line_from_h)\n translated_line_to = apply_translation(translation, line_to_h)\n translated_mid_point = (translated_line_from + translated_line_to) / 2.0\n translated_mid_point = translated_mid_point[[1, 0]]\n line_from_transformed = apply_rotation(translated_mid_point,\n np.pi * angle,\n translated_line_from)\n line_to_transformed = apply_rotation(translated_mid_point,\n np.pi * angle,\n translated_line_to)\n line_from_transformed = np.array(from_homogeneous(line_from_transformed))\n line_to_transformed = np.array(from_homogeneous(line_to_transformed))\n\n # Second transform.\n line_from_h = to_homogeneous(line_from_transformed)\n line_to_h = to_homogeneous(line_to_transformed)\n line_from_h = apply_scale(scale2, line_from_h)\n line_to_h = apply_scale(scale2, line_to_h)\n translated_line_from = apply_translation(\n translation2, line_from_h, offset_r=img_siz2, offset_c=img_siz2)\n translated_line_to = apply_translation(\n translation2, line_to_h, offset_r=img_siz2, offset_c=img_siz2)\n translated_mid_point = (translated_line_from + translated_line_to) / 2.0\n translated_mid_point = translated_mid_point[[1, 0]]\n line_from_transformed = apply_rotation(translated_mid_point,\n np.pi * angle2,\n translated_line_from)\n line_to_transformed = apply_rotation(translated_mid_point,\n np.pi * angle2,\n translated_line_to)\n return np.concatenate([from_homogeneous(line_from_transformed),\n from_homogeneous(line_to_transformed)],\n axis=1)",
"_____no_output_____"
]
],
[
[
"### Hierarchical stroke painting functions",
"_____no_output_____"
]
],
[
[
"# PaintingCommand\n# origin_top: Origin of line defined by top level LSTM\n# angle_top: Angle of line defined by top level LSTM\n# scale_top: Scale for line defined by top level LSTM\n# origin_bottom: Origin of line defined by bottom level LSTM\n# angle_bottom: Angle of line defined by bottom level LSTM\n# scale_bottom: Scale for line defined by bottom level LSTM\n# position_choice: Selects between use of:\n# Origin, angle and scale from both LSTM levels\n# Origin, angle and scale just from top level LSTM\n# Origin, angle and scale just from bottom level LSTM\n# transparency: Line transparency determined by bottom level LSTM\nPaintingCommand = collections.namedtuple(\"PaintingCommand\",\n [\"origin_top\",\n \"angle_top\",\n \"scale_top\",\n \"origin_bottom\",\n \"angle_bottom\",\n \"scale_bottom\",\n \"position_choice\",\n \"transparency\"])\n\ndef paint_over_image(img, strokes, painting_commands,\n allow_strokes_beyond_image_edges, coeff_size=1):\n \"\"\"Make marks over an existing image.\n\n Args:\n img: Image to draw on.\n strokes: Stroke descriptions.\n painting_commands: Top-level painting commands with transforms for the i\n sets of strokes.\n allow_strokes_beyond_image_edges: Allow strokes beyond image boundary.\n coeff_size: Determines low res (1) or high res (10) image will be drawn.\n\n Returns:\n num_strokes: The number of strokes made.\n \"\"\"\n img_center = 112. * coeff_size\n # a, b and c: determines the stroke width distribution (see 'weights' below)\n a = 10. * coeff_size\n b = 2. * coeff_size\n c = 300. * coeff_size\n # d: extent that the strokes are allowed to go beyond the edge of the canvas\n d = 223 * coeff_size\n\n def _clip_colour(col):\n return np.clip((np.round(col * 255. + 128.)).astype(np.int32), 0, 255)\n\n # Loop over all the top level...\n t0_over = time.time()\n num_strokes = sum(len(s) for s in strokes)\n translations = np.zeros((2, num_strokes,), np.float32)\n translations2 = np.zeros((2, num_strokes,), np.float32)\n angles = np.zeros((num_strokes,), np.float32)\n angles2 = np.zeros((num_strokes,), np.float32)\n scales = np.zeros((num_strokes,), np.float32)\n scales2 = np.zeros((num_strokes,), np.float32)\n weights = np.zeros((num_strokes,), np.float32)\n lines_from = np.zeros((num_strokes, 2), np.float32)\n lines_to = np.zeros((num_strokes, 2), np.float32)\n rgbas = np.zeros((num_strokes, 4), np.float32)\n k = 0\n for i in range(len(strokes)):\n\n # Get the top-level transforms for the i-th bunch of strokes\n painting_comand = painting_commands[i]\n translation_a = painting_comand.origin_top\n angle_a = (painting_comand.angle_top + 1) / 5.0\n scale_a = 0.5 + (painting_comand.scale_top + 1) / 3.0\n translation_b = painting_comand.origin_bottom\n angle_b = (painting_comand.angle_bottom + 1) / 5.0\n scale_b = 0.5 + (painting_comand.scale_bottom + 1) / 3.0\n position_choice = painting_comand.position_choice\n solid_colour = painting_comand.transparency\n\n # Do we use origin, angle and scale from both, top or bottom LSTM levels?\n if position_choice > 0.33:\n translation = translation_a\n angle = angle_a\n scale = scale_a\n translation2 = translation_b\n angle2 = angle_b\n scale2 = scale_b\n elif position_choice > -0.33:\n translation = translation_a\n angle = angle_a\n scale = scale_a\n translation2 = [-img_center, -img_center]\n angle2 = 0.\n scale2 = 1.\n else:\n translation = translation_b\n angle = angle_b\n scale = scale_b\n translation2 = [-img_center, -img_center]\n angle2 = 0.\n scale2 = 1.\n\n # Store top-level transforms\n strokes_i = strokes[i]\n n_i = len(strokes_i)\n angles[k:(k+n_i)] = angle\n angles2[k:(k+n_i)] = angle2\n scales[k:(k+n_i)] = scale\n scales2[k:(k+n_i)] = scale2\n translations[0, k:(k+n_i)] = translation[0]\n translations[1, k:(k+n_i)] = translation[1]\n translations2[0, k:(k+n_i)] = translation2[0]\n translations2[1, k:(k+n_i)] = translation2[1]\n\n # ... and the bottom level stroke definitions.\n for j in range(n_i):\n z_ij = strokes_i[j]\n\n # Store line weight (we will process micro-strokes later)\n weights[k] = z_ij[4]\n # Store line endpoints\n lines_from[k, :] = (z_ij[0], z_ij[1])\n lines_to[k, :] = (z_ij[2], z_ij[3])\n\n # Store colour and alpha\n rgbas[k, 0] = z_ij[7]\n rgbas[k, 1] = z_ij[8]\n rgbas[k, 2] = z_ij[9]\n if solid_colour > -0.5:\n rgbas[k, 3] = 25.5\n else:\n rgbas[k, 3] = z_ij[11]\n k += 1\n\n # Draw all the strokes in a batch as sequence of length 2 * num_strokes\n t1_over = time.time()\n lines_from *= img_center/2.0\n lines_to *= img_center/2.0\n rr, cc = transform_lines(lines_from, lines_to, translations, angles, scales,\n translations2, angles2, scales2, img_center)\n if not allow_strokes_beyond_image_edges:\n rrm = np.round(np.clip(rr, 1, d-1)).astype(int)\n ccm = np.round(np.clip(cc, 1, d-1)).astype(int)\n else:\n rrm = np.round(rr).astype(int)\n ccm = np.round(cc).astype(int)\n\n # Plot all the strokes\n t2_over = time.time()\n img_pil = Image.fromarray(img)\n canvas = ImageDraw.Draw(img_pil, \"RGBA\")\n rgbas[:, :3] = _clip_colour(rgbas[:, :3])\n rgbas[:, 3] = (np.clip(5.0 * np.abs(rgbas[:, 3]), 0, 255)).astype(np.int32)\n weights = (np.clip(np.round(weights * b + a), 2, c)).astype(np.int32)\n for k in range(num_strokes):\n canvas.line((rrm[k], ccm[k], rrm[k+num_strokes], ccm[k+num_strokes]),\n fill=tuple(rgbas[k]), width=weights[k])\n img[:] = np.asarray(img_pil)[:]\n t3_over = time.time()\n if VERBOSE_CODE:\n print(\"{:.2f}s to store {} stroke defs, {:.4f}s to \"\n \"compute them, {:.4f}s to plot them\".format(\n t1_over - t0_over, num_strokes, t2_over - t1_over,\n t3_over - t2_over))\n return num_strokes\n",
"_____no_output_____"
]
],
[
[
"### Recurrent Neural Network Layer Generator",
"_____no_output_____"
]
],
[
[
"# DrawingLSTMSpec - parameters defining the LSTM architecture\n# input_spec_size: Size if sequence elements\n# num_lstms: Number of LSTMs at each layer\n# net_lstm_hiddens: Number of hidden LSTM units\n# net_mlp_hiddens: Number of hidden units in MLP layer\nDrawingLSTMSpec = collections.namedtuple(\"DrawingLSTMSpec\",\n [\"input_spec_size\",\n \"num_lstms\",\n \"net_lstm_hiddens\",\n \"net_mlp_hiddens\"])\n\n\nclass MakeGeneratorLstm(nn.Module):\n \"\"\"Block of parallel LSTMs with MLP output heads.\"\"\"\n\n def __init__(self, drawing_lstm_spec, output_size):\n \"\"\"Build drawing LSTM architecture using spec.\n\n Args:\n drawing_lstm_spec: DrawingLSTMSpec with architecture parameters\n output_size: Number of outputs for the MLP head layer\n \"\"\"\n super(MakeGeneratorLstm, self).__init__()\n self._num_lstms = drawing_lstm_spec.num_lstms\n self._input_layer = nn.Sequential(\n nn.Linear(drawing_lstm_spec.input_spec_size,\n drawing_lstm_spec.net_lstm_hiddens),\n torch.nn.LeakyReLU(0.2, inplace=True))\n lstms = []\n heads = []\n for _ in range(self._num_lstms):\n lstm_layer = nn.LSTM(\n input_size=drawing_lstm_spec.net_lstm_hiddens,\n hidden_size=drawing_lstm_spec.net_lstm_hiddens,\n num_layers=2, batch_first=True, bias=True)\n head_layer = nn.Sequential(\n nn.Linear(drawing_lstm_spec.net_lstm_hiddens,\n drawing_lstm_spec.net_mlp_hiddens),\n torch.nn.LeakyReLU(0.2, inplace=True),\n nn.Linear(drawing_lstm_spec.net_mlp_hiddens, output_size))\n lstms.append(lstm_layer)\n heads.append(head_layer)\n self._lstms = nn.ModuleList(lstms)\n self._heads = nn.ModuleList(heads)\n\n def forward(self, x):\n pred = []\n x = self._input_layer(x)*10.0\n for i in range(self._num_lstms):\n y, _ = self._lstms[i](x)\n y = self._heads[i](y)\n pred.append(y)\n return pred",
"_____no_output_____"
]
],
[
[
"### DrawingLSTM - A Drawing Recurrent Neural Network",
"_____no_output_____"
]
],
[
[
"Genotype = collections.namedtuple(\"Genotype\",\n [\"top_lstm\",\n \"bottom_lstm\",\n \"input_sequence\",\n \"initial_img\"])\n\nclass DrawingLSTM:\n \"\"\"LSTM for processing input sequences and generating resultant drawings.\n\n Comprised of two LSTM layers.\n \"\"\"\n\n def __init__(self, drawing_lstm_spec, allow_strokes_beyond_image_edges):\n \"\"\"Create DrawingLSTM to interpret input sequences and paint an image.\n\n Args:\n drawing_lstm_spec: DrawingLSTMSpec with LSTM architecture parameters\n allow_strokes_beyond_image_edges: Draw lines outside image boundary\n \"\"\"\n self._input_spec_size = drawing_lstm_spec.input_spec_size\n self._num_lstms = drawing_lstm_spec.num_lstms\n self._allow_strokes_beyond_image_edges = allow_strokes_beyond_image_edges\n with torch.no_grad():\n self.top_lstm = MakeGeneratorLstm(drawing_lstm_spec,\n self._input_spec_size)\n self.bottom_lstm = MakeGeneratorLstm(drawing_lstm_spec, 12)\n self._init_all(self.top_lstm, torch.nn.init.normal_, mean=0., std=0.2)\n self._init_all(self.bottom_lstm, torch.nn.init.normal_, mean=0., std=0.2)\n\n def _init_all(self, a_model, init_func, *params, **kwargs):\n \"\"\"Method for initialising model with given init_func, params and kwargs.\"\"\"\n for p in a_model.parameters():\n init_func(p, *params, **kwargs)\n\n def _feed_top_lstm(self, input_seq):\n \"\"\"Feed all input sequences input_seq into the LSTM models.\"\"\"\n\n x_in = input_seq.reshape((len(input_seq), 1, self._input_spec_size))\n x_in = np.tile(x_in, (SEQ_LENGTH, 1))\n x_torch = torch.from_numpy(x_in).type(torch.FloatTensor)\n y_torch = self.top_lstm(x_torch)\n y_torch = [y_torch_k.detach().numpy() for y_torch_k in y_torch]\n del x_in\n del x_torch\n\n # There are multiple LSTM heads. For each sequence, read out the head and\n # length of intermediary output to keep and return intermediary outputs.\n readouts_top = np.clip(\n np.round(self._num_lstms/2.0 * (1 + input_seq[:, 1])).astype(np.int32),\n 0, self._num_lstms-1)\n lengths_top = np.clip(\n np.round(10.0 * (1 + input_seq[:, 0])).astype(np.int32),\n 0, SEQ_LENGTH) + 1\n intermediate_strings = []\n for i in range(len(readouts_top)):\n y_torch_i = y_torch[readouts_top[i]][i]\n intermediate_strings.append(y_torch_i[0:lengths_top[i], :])\n return intermediate_strings\n\n def _feed_bottom_lstm(self, intermediate_strings, input_seq, coeff_size=1):\n \"\"\"Feed all input sequences into the LSTM models.\n\n Args:\n intermediate_strings: top level strings\n input_seq: input sequences fed to the top LSTM\n coeff_size: sets centre origin\n\n Returns:\n strokes: Painting strokes.\n painting_commands: Top-level painting commands with origin, angle and scale\n information, as well as transparency.\n \"\"\"\n img_center = 112. * coeff_size\n coeff_origin = 100. * coeff_size\n top_lengths = []\n for i in range(len(intermediate_strings)):\n top_lengths.append(len(intermediate_strings[i]))\n y_flat = np.concatenate(intermediate_strings, axis=0)\n tiled_y_flat = y_flat.reshape((len(y_flat), 1, self._input_spec_size))\n tiled_y_flat = np.tile(tiled_y_flat, (SEQ_LENGTH, 1))\n y_torch = torch.from_numpy(tiled_y_flat).type(torch.FloatTensor)\n z_torch = self.bottom_lstm(y_torch)\n z_torch = [z_torch_k.detach().numpy() for z_torch_k in z_torch]\n del tiled_y_flat\n del y_torch\n\n # There are multiple LSTM heads. For each sequence, read out the head and\n # length of intermediary output to keep and return intermediary outputs.\n readouts = np.clip(np.round(\n NUM_LSTMS/2.0 * (1 + y_flat[:, 0])).astype(np.int32), 0, NUM_LSTMS-1)\n lengths_bottom = np.clip(\n np.round(10.0 * (1 + y_flat[:, 1])).astype(np.int32), 0, SEQ_LENGTH) + 1\n strokes = []\n painting_commands = []\n offset = 0\n for i in range(len(intermediate_strings)):\n origin_top = [(1+input_seq[i, 2]) * img_center,\n (1+input_seq[i, 3]) * img_center]\n angle_top = input_seq[i, 4]\n scale_top = input_seq[i, 5]\n for j in range(len(intermediate_strings[i])):\n k = j + offset\n z_torch_ij = z_torch[readouts[k]][k]\n strokes.append(z_torch_ij[0:lengths_bottom[k], :])\n y_ij = y_flat[k]\n origin_bottom = [y_ij[2] * coeff_origin, y_ij[3] * coeff_origin]\n angle_bottom = y_ij[4]\n scale_bottom = y_ij[5]\n position_choice = y_ij[6]\n transparency = y_ij[7]\n painting_command = PaintingCommand(\n origin_top, angle_top, scale_top, origin_bottom, angle_bottom,\n scale_bottom, position_choice, transparency)\n painting_commands.append(painting_command)\n offset += top_lengths[i]\n del y_flat\n return strokes, painting_commands\n\n def make_initial_genotype(self, initial_img, sequence_length,\n input_spec_size):\n \"\"\"Make and return initial DNA weights for LSTMs, input sequence, and image.\n\n Args:\n initial_img: Image (to be appended to the genotype)\n sequence_length: Length of the input sequence (i.e. number of strokes)\n input_spec_size: Number of inputs for each element in the input sequences\n Returns:\n Genotype NamedTuple with fields: [parameters of network 0,\n parameters of network 1,\n input sequence,\n initial_img]\n \"\"\"\n dna_top = []\n with torch.no_grad():\n for _, params in self.top_lstm.named_parameters():\n dna_top.append(params.clone())\n param_size = params.numpy().shape\n dna_top[-1] = np.random.uniform(\n 0.1 * DNA_SCALE, 0.3\n * DNA_SCALE) * np.random.normal(size=param_size)\n dna_bottom = []\n with torch.no_grad():\n for _, params in self.bottom_lstm.named_parameters():\n dna_bottom.append(params.clone())\n param_size = params.numpy().shape\n dna_bottom[-1] = np.random.uniform(\n 0.1 * DNA_SCALE, 0.3\n * DNA_SCALE) * np.random.normal(size=param_size)\n input_sequence = np.random.uniform(\n -1, 1, size=(sequence_length, input_spec_size))\n return Genotype(dna_top, dna_bottom, input_sequence, initial_img)\n\n def draw(self, img, genotype):\n \"\"\"Add to the image using the latest genotype and get latest input sequence.\n\n Args:\n img: image to add to.\n genotype: as created by make_initial_genotype.\n\n Returns:\n image with new strokes added.\n \"\"\"\n t0_draw = time.time()\n img = img + genotype.initial_img\n input_sequence = genotype.input_sequence\n\n # Generate the strokes for drawing in batch mode.\n # input_sequence is between 10 and 20 but is evolved, can go to 200.\n intermediate_strings = self._feed_top_lstm(input_sequence)\n strokes, painting_commands = self._feed_bottom_lstm(\n intermediate_strings, input_sequence)\n del intermediate_strings\n\n # Now we can go through the output strings producing the strokes.\n t1_draw = time.time()\n num_strokes = paint_over_image(\n img, strokes, painting_commands, self._allow_strokes_beyond_image_edges,\n coeff_size=1)\n\n t2_draw = time.time()\n if VERBOSE_CODE:\n print(\n \"Draw {:.2f}s (net {:.2f}s plot {:.2f}s {:.1f}ms/strk {}\".format(\n t2_draw - t0_draw, t1_draw - t0_draw, t2_draw - t1_draw,\n (t2_draw - t1_draw) / num_strokes * 1000, num_strokes))\n return img",
"_____no_output_____"
]
],
[
[
"## DrawingGenerator",
"_____no_output_____"
]
],
[
[
"class DrawingGenerator:\n \"\"\"Creates a drawing using a DrawingLSTM.\"\"\"\n\n def __init__(self, image_size, drawing_lstm_spec,\n allow_strokes_beyond_image_edges):\n self.primitives = [\"c\", \"r\", \"l\", \"b\", \"p\", \"j\"]\n self.pop = []\n self.size = image_size\n self.fitnesses = np.zeros(1)\n self.noise = 2\n self.mutation_std = 0.0004\n # input_spec_size, num_lstms, net_lstm_hiddens,\n # net_mlp_hiddens, output_size, allow_strokes_beyond_image_edges\n self.drawing_lstm = DrawingLSTM(drawing_lstm_spec,\n allow_strokes_beyond_image_edges)\n\n def make_initial_genotype(self, initial_img, sequence_length, input_spec_size):\n \"\"\"Use drawing_lstm to create initial genotypye.\"\"\"\n\n self.genotype = self.drawing_lstm.make_initial_genotype(\n initial_img, sequence_length, input_spec_size)\n return self.genotype\n\n\n def _copy_genotype_to_generator(self, genotype):\n \"\"\"Copy genotype's data into generator's parameters.\n\n Copies the parameters in genotype (genotype.top_lstm[:] and\n genotype.bottom_lstm[:]) into the parmaters for the drawing network so it\n can be used to evaluate the genotype.\n\n Args:\n genotype: as created by make_initial_genotype.\n\n Returns:\n None\n \"\"\"\n self.genotype = copy.deepcopy(genotype)\n i = 0\n with torch.no_grad():\n for _, param in self.drawing_lstm.top_lstm.named_parameters():\n param.copy_(torch.tensor(self.genotype.top_lstm[i]))\n i = i + 1\n i = 0\n with torch.no_grad():\n for _, param in self.drawing_lstm.bottom_lstm.named_parameters():\n param.copy_(torch.tensor(self.genotype.bottom_lstm[i]))\n i = i + 1\n\n def _interpret_genotype(self, genotype):\n img = np.zeros((self.size, self.size, 3), dtype=np.uint8)\n img = self.drawing_lstm.draw(img, genotype)\n return img\n\n def draw_from_genotype(self, genotype):\n \"\"\"Copy input sequence and LSTM weights from `genotype`, run and draw.\"\"\"\n self._copy_genotype_to_generator(genotype)\n return self._interpret_genotype(self.genotype)\n\n def visualize_genotype(self, genotype):\n \"\"\"Plot histograms of genotype\"s data.\"\"\"\n\n plt.show()\n inp_seq = np.array(genotype.input_sequence).flatten()\n plt.title(\"input seq\")\n plt.hist(inp_seq)\n plt.show()\n\n inp_seq = np.array(genotype.top_lstm).flatten()\n plt.title(\"LSTM top\")\n plt.hist(inp_seq)\n plt.show()\n\n inp_seq = np.array(genotype.bottom_lstm).flatten()\n plt.title(\"LSTM bottom\")\n plt.hist(inp_seq)\n\n plt.show()\n\n def mutate(self, genotype):\n \"\"\"Mutates `genotype`. This function is static.\n\n Args:\n genotype: genotype structure to mutate parameters of.\n\n Returns:\n new_genotype: Mutated copy of supplied genotype.\n \"\"\"\n\n new_genotype = copy.deepcopy(genotype)\n new_input_seq = new_genotype.input_sequence\n n = len(new_input_seq)\n\n if np.random.uniform() < 1.0:\n\n # Standard gaussian small mutation of input sequence.\n if np.random.uniform() > 0.5:\n new_input_seq += (\n np.random.uniform(0.001, 0.2) * np.random.normal(\n size=new_input_seq.shape))\n\n # Low frequency large mutation of individual parts of the input sequence.\n for i in range(n):\n if np.random.uniform() < 2.0/n:\n for j in range(len(new_input_seq[i])):\n if np.random.uniform() < 2.0/len(new_input_seq[i]):\n new_input_seq[i][j] = new_input_seq[i][j] + 0.5*np.random.normal()\n\n # Adding and deleting elements from the input sequence.\n if np.random.uniform() < 0.01:\n if VERBOSE_MUTATION:\n print(\"Mutation: adding\")\n a = np.random.uniform(-1, 1, size=(1, INPUT_SPEC_SIZE))\n pos = np.random.randint(1, len(new_input_seq))\n new_input_seq = np.insert(new_input_seq, pos, a, axis=0)\n if np.random.uniform() < 0.02:\n if VERBOSE_MUTATION:\n print(\"Mutation: deleting\")\n pos = np.random.randint(1, len(new_input_seq))\n new_input_seq = np.delete(new_input_seq, pos, axis=0)\n n = len(new_input_seq)\n\n # Swapping two elements in the input sequence.\n if np.random.uniform() < 0.01:\n element1 = np.random.randint(0, n)\n element2 = np.random.randint(0, n)\n while element1 == element2:\n element2 = np.random.randint(0, n)\n temp = copy.deepcopy(new_input_seq[element1])\n new_input_seq[element1] = copy.deepcopy(new_input_seq[element2])\n new_input_seq[element2] = temp\n\n # Duplicate an element in the input sequence (with some mutation).\n if np.random.uniform() < 0.01:\n if VERBOSE_MUTATION:\n print(\"Mutation: duplicating\")\n element1 = np.random.randint(0, n)\n element2 = np.random.randint(0, n)\n while element1 == element2:\n element2 = np.random.randint(0, n)\n new_input_seq[element1] = copy.deepcopy(new_input_seq[element2])\n noise = 0.05 * np.random.normal(size=new_input_seq[element1].shape)\n new_input_seq[element1] += noise\n\n # Ensure that the input sequence is always between -1 and 1\n # so that positions make sense.\n new_genotype = new_genotype._replace(\n input_sequence=np.clip(new_input_seq, -1.0, 1.0))\n\n # Mutates dna of networks.\n if np.random.uniform() < 1.0:\n for net in range(2):\n for layer in range(len(new_genotype[net])):\n weights = new_genotype[net][layer]\n if np.random.uniform() < 0.5:\n noise = 0.00001 * np.random.standard_cauchy(size=weights.shape)\n weights += noise\n else:\n noise = np.random.normal(size=weights.shape)\n noise *= np.random.uniform(0.0001, 0.006)\n weights += noise\n\n if np.random.uniform() < 0.01:\n noise = np.random.normal(size=weights.shape)\n noise *= np.random.uniform(0.1, 0.3)\n weights = noise\n\n # Ensure weights are between -10 and 10.\n weights = np.clip(weights, -1.0, 1.0)\n new_genotype[net][layer] = weights\n\n return new_genotype",
"_____no_output_____"
]
],
[
[
"## Evaluator",
"_____no_output_____"
]
],
[
[
"class Evaluator:\n \"\"\"Evaluator for a drawing.\"\"\"\n\n def __init__(self, image_size, drawing_lstm_spec,\n allow_strokes_beyond_image_edges):\n self.drawing_generator = DrawingGenerator(image_size, drawing_lstm_spec,\n allow_strokes_beyond_image_edges)\n self.calls = 0\n\n def make_initial_genotype(self, img, sequence_length, input_spec_size):\n return self.drawing_generator.make_initial_genotype(img, sequence_length,\n input_spec_size)\n\n def evaluate_genotype(self, pickled_genotype, id_num):\n \"\"\"Evaluate genotype and return genotype's image.\n\n Args:\n pickled_genotype: pickled genotype to be evaluated.\n id_num: ID number of genotype.\n\n Returns:\n dict: drawing and id_num.\n \"\"\"\n\n genotype = cloudpickle.loads(pickled_genotype)\n drawing = self.drawing_generator.draw_from_genotype(genotype)\n self.calls += 1\n return {\"drawing\": drawing, \"id\": id_num}\n\n def mutate(self, genotype):\n \"\"\"Create a mutated version of genotype.\"\"\"\n return self.drawing_generator.mutate(genotype)",
"_____no_output_____"
]
],
[
[
"# Evolution",
"_____no_output_____"
],
[
"## Fitness calculation, tournament, and crossover",
"_____no_output_____"
]
],
[
[
"IMAGE_MEAN = torch.tensor([0.48145466, 0.4578275, 0.40821073]).cuda()\nIMAGE_STD = torch.tensor([0.26862954, 0.26130258, 0.27577711]).cuda()\n\ndef get_fitness(pictures, use_projective_transform,\n projective_transform_coefficient):\n \"\"\"Run CLIP on a batch of `pictures` and return `fitnesses`.\n\n Args:\n pictures: batch if images to evaluate\n use_projective_transform: Add transformed versions of the image\n projective_transform_coefficient: Degree of transform\n\n Returns:\n Similarities between images and the text\n \"\"\"\n\n # Do we use projective transforms of images before CLIP eval?\n t0 = time.time()\n pictures_trans = np.swapaxes(np.array(pictures), 1, 3) / 244.0\n if use_projective_transform:\n for i in range(len(pictures_trans)):\n matrix = np.eye(3) + (\n projective_transform_coefficient * np.random.normal(size=(3, 3)))\n tform = transform.ProjectiveTransform(matrix=matrix)\n pictures_trans[i] = transform.warp(pictures_trans[i], tform.inverse)\n\n # Run the CLIP evaluator.\n t1 = time.time()\n image_input = torch.tensor(np.stack(pictures_trans)).cuda()\n image_input -= IMAGE_MEAN[:, None, None]\n image_input /= IMAGE_STD[:, None, None]\n with torch.no_grad():\n image_features = model.encode_image(image_input).float()\n t2 = time.time()\n similarity = torch.cosine_similarity(\n text_features, image_features, dim=1).cpu().numpy()\n t3 = time.time()\n if VERBOSE_CODE:\n print(f\"get_fitness init {t1-t0:.4f}s, CLIP {t2-t1:.4f}s, sim {t3-t2:.4f}s\")\n return similarity\n\n\ndef crossover(dna_winner, dna_loser, crossover_prob):\n \"\"\"Create new genotype by combining two genotypes.\n\n Randomly replaces parts of the genotype 'dna_winner' with parts of dna_loser\n to create a new genotype based mostly on on both 'parents'.\n\n Args:\n dna_winner: The high-fitness parent genotype - gets replaced with child.\n dna_loser: The lower-fitness parent genotype.\n crossover_prob: Probability of crossover between winner and loser.\n\n Returns:\n dna_winner: The result of crossover from parents.\n \"\"\"\n\n # Copy single input signals\n for i in range(len(dna_winner[2])):\n if i < len(dna_loser[2]):\n if np.random.uniform() < crossover_prob:\n dna_winner[2][i] = copy.deepcopy(dna_loser[2][i])\n\n # Copy whole modules\n for i in range(len(dna_winner[0])):\n if i < len(dna_loser[0]):\n if np.random.uniform() < crossover_prob:\n dna_winner[0][i] = copy.deepcopy(dna_loser[0][i])\n\n # Copy whole modules\n for i in range(len(dna_winner[1])):\n if i < len(dna_loser[1]):\n if np.random.uniform() < crossover_prob:\n dna_winner[1][i] = copy.deepcopy(dna_loser[1][i])\n\n return dna_winner\n\n\ndef truncation_selection(population, fitnesses, evaluator, use_crossover,\n crossover_prob):\n \"\"\"Create new population using truncation selection.\n\n Creates a new population by copying across the best 50% genotypes and\n filling the rest with (for use_crossover==False) a mutated copy of each\n genotype or (for use_crossover==True) with children created through crossover\n between each winner and a genotype in the bottom 50%.\n\n Args:\n population: list of current population genotypes.\n fitnesses: list of evaluated fitnesses.\n evaluator: class that evaluates a draw generator.\n use_crossover: Whether to use crossover between winner and loser.\n crossover_prob: Probability of crossover between winner and loser.\n\n Returns:\n new_pop: the new population.\n best: genotype.\n \"\"\"\n\n fitnesses = np.array(-fitnesses)\n ordered_fitness_ids = fitnesses.argsort()\n best = copy.deepcopy(population[ordered_fitness_ids[0]])\n pop_size = len(population)\n\n if not use_crossover:\n new_pop = []\n for i in range(int(pop_size/2)):\n new_pop.append(copy.deepcopy(population[ordered_fitness_ids[i]]))\n for i in range(int(pop_size/2)):\n new_pop.append(evaluator.mutate(\n copy.deepcopy(population[ordered_fitness_ids[i]])))\n else:\n new_pop = []\n for i in range(int(pop_size/2)):\n new_pop.append(copy.deepcopy(population[ordered_fitness_ids[i]]))\n for i in range(int(pop_size/2)):\n new_pop.append(evaluator.mutate(crossover(\n copy.deepcopy(population[ordered_fitness_ids[i]]),\n population[ordered_fitness_ids[int(pop_size/2) + i]], crossover_prob\n )))\n\n return new_pop, best",
"_____no_output_____"
]
],
[
[
"##Remote workers",
"_____no_output_____"
]
],
[
[
"VERBOSE_DURATION = False\n\[email protected]\nclass Worker(object):\n \"\"\"Takes a pickled dna and evaluates it, returning result.\"\"\"\n\n def __init__(self, image_size, drawing_lstm_spec,\n allow_strokes_beyond_image_edges):\n self.evaluator = Evaluator(image_size, drawing_lstm_spec,\n allow_strokes_beyond_image_edges)\n\n def compute(self, dna_pickle, genotype_id):\n if VERBOSE_DURATION:\n t0 = time.time()\n res = self.evaluator.evaluate_genotype(dna_pickle, genotype_id)\n if VERBOSE_DURATION:\n duration = time.time() - t0\n print(f\"Worker {genotype_id} evaluated params in {duration:.1f}sec\")\n return res\n\n\ndef create_workers(num_workers, image_size, drawing_lstm_spec,\n allow_strokes_beyond_image_edges):\n \"\"\"Create the workers.\n\n Args:\n num_workers: Number of parallel workers for evaluation.\n image_size: Length of side of (square) image\n drawing_lstm_spec: DrawingLSTMSpec for LSTM network\n allow_strokes_beyond_image_edges: Whether to draw outside the edges\n Returns:\n List of workers.\n \"\"\"\n worker_pool = []\n for w_i in range(num_workers):\n print(\"Creating worker\", w_i, flush=True)\n worker_pool.append(Worker.remote(image_size, drawing_lstm_spec,\n allow_strokes_beyond_image_edges))\n return worker_pool\n",
"_____no_output_____"
]
],
[
[
"##Plotting",
"_____no_output_____"
]
],
[
[
"def plot_training_res(batch_drawings, fitness_history, idx=None):\n \"\"\"Plot fitnesses and timings.\n\n Args:\n batch_drawings: Drawings\n fitness_history: History of fitnesses\n idx: Index of drawing to show, default is highest fitness\n \"\"\"\n _, (ax1, ax2) = plt.subplots(1, 2, figsize=(10, 5))\n if idx is None:\n idx = np.argmax(fitness_history[-1])\n ax1.plot(fitness_history, \".\")\n ax1.set_title(\"Fitnesses\")\n ax2.imshow(batch_drawings[idx])\n ax2.set_title(f\"{PROMPT} (fit: {fitness_history[-1][idx]:.3f})\")\n plt.show()\n\ndef plot_samples(batch_drawings, num_samples=16):\n \"\"\"Plot sample of drawings.\n\n Args:\n batch_drawings: Batch of drawings to sample from\n num_samples: Number to displa\n \"\"\"\n num_samples = min(len(batch_drawings), num_samples)\n num_rows = int(math.floor(np.sqrt(num_samples)))\n num_cols = int(math.ceil(num_samples / num_rows))\n row_images = []\n for c in range(0, num_samples, num_cols):\n if c + num_cols <= num_samples:\n row_images.append(np.concatenate(batch_drawings[c:(c+num_cols)], axis=1))\n composite_image = np.concatenate(row_images, axis=0)\n _, ax = plt.subplots(1, 1, figsize=(20, 20))\n ax.imshow(composite_image)\n ax.set_title(PROMPT)",
"_____no_output_____"
]
],
[
[
"## Population and evolution main loop",
"_____no_output_____"
]
],
[
[
"def make_population(pop_size, evaluator, image_size, input_spec_size,\n sequence_length):\n \"\"\"Make initial population.\n\n Args:\n pop_size: number of genotypes in population.\n evaluator: An Evaluator class instance for generating initial genotype.\n image_size: Size of initial image for genotype to draw on.\n input_spec_size: Sequence element size\n sequence_length: Initial length of sequences\n\n Returns:\n Initialised population.\n \"\"\"\n print(f\"Creating initial population of size {pop_size}\")\n pop = []\n for _ in range(pop_size):\n a_genotype = evaluator.make_initial_genotype(\n img=np.zeros((image_size, image_size, 3), dtype=np.uint8),\n sequence_length=sequence_length,\n input_spec_size=input_spec_size)\n pop.append(a_genotype)\n return pop\n\ndef evolution_loop(population, worker_pool, evaluator, num_generations,\n use_crossover, crossover_prob,\n use_projective_transform, projective_transform_coefficient,\n plot_every, plot_batch):\n \"\"\"Create population and run evolution.\n\n Args:\n population: Initial population of genotypes\n worker_pool: List of workers of parallel evaluations\n evaluator: image evaluator to calculate fitnesses\n num_generations: number of generations to run\n use_crossover: Whether crossover is used for offspring\n crossover_prob: Probability that crossover takes place\n use_projective_transform: Use projective transforms in evaluation\n projective_transform_coefficient: Degree of projective transform\n plot_every: number of generations between new plots\n plot_batch: whether to show all samples in the batch then plotting\n \"\"\"\n population_size = len(population)\n num_workers = len(worker_pool)\n print(\"Population of {} genotypes being evaluated by {} workers\".format(\n population_size, num_workers))\n drawings = {}\n fitness_history = []\n init_gen = len(fitness_history)\n print(f\"(Re)starting evolution at generation {init_gen}\")\n for gen in range(init_gen, num_generations):\n\n # Drawing\n t0_loop = time.time()\n futures = []\n for j in range(0, population_size, num_workers):\n for i in range(num_workers):\n futures.append(worker_pool[i].compute.remote(\n cloudpickle.dumps(population[i+j]), i+j))\n data = ray.get(futures)\n for i in range(num_workers):\n drawings[data[i+j][\"id\"]] = data[j+i][\"drawing\"]\n batch_drawings = []\n for i in range(population_size):\n batch_drawings.append(drawings[i])\n\n # Fitness evaluation using CLIP\n t1_loop = time.time()\n fitnesses = get_fitness(batch_drawings, use_projective_transform,\n projective_transform_coefficient)\n fitness_history.append(copy.deepcopy(fitnesses))\n\n # Tournament\n t2_loop = time.time()\n population, best_genotype = truncation_selection(\n population, fitnesses, evaluator, use_crossover, crossover_prob)\n t3_loop = time.time()\n duration_draw = t1_loop - t0_loop\n duration_fit = t2_loop - t1_loop\n duration_tournament = t3_loop - t2_loop\n duration_total = t3_loop - t0_loop\n if gen % plot_every == 0:\n if VISUALIZE_GENOTYPE:\n evaluator.drawing_generator.visualize_genotype(best_genotype)\n print(\"Draw: {:.2f}s fit: {:.2f}s evol: {:.2f}s total: {:.2f}s\".format(\n duration_draw, duration_fit, duration_tournament, duration_total))\n plot_training_res(batch_drawings, fitness_history)\n if plot_batch:\n num_samples_to_plot = int(math.pow(\n math.floor(np.sqrt(population_size)), 2))\n plot_samples(batch_drawings, num_samples=num_samples_to_plot)\n",
"_____no_output_____"
]
],
[
[
"# Configure and Generate",
"_____no_output_____"
]
],
[
[
"#@title Hyperparameters\n\n#@markdown Evolution parameters: population size and number of generations.\nPOPULATION_SIZE = 10 #@param {type:\"slider\", min:4, max:100, step:2}\nNUM_GENERATIONS = 5000 #@param {type:\"integer\", min:100}\n#@markdown Number of workers working in parallel (should be equal to or smaller than the population size).\nNUM_WORKERS = 10 #@param {type:\"slider\", min:4, max:100, step:2}\n#@markdown Crossover in evolution.\nUSE_CROSSOVER = True #@param {type:\"boolean\"}\nCROSSOVER_PROB = 0.01 #@param {type:\"number\"}\n#@markdown Number of LSTMs, each one encoding a group of strokes.\nNUM_LSTMS = 5 #@param {type:\"integer\", min:1, max:5}\n#@markdown Number of inputs for each element in the input sequences.\nINPUT_SPEC_SIZE = 10 #@param {type:\"integer\"}\n#@markdown Length of the input sequence fed to the LSTMs (determines number of strokes).\nSEQ_LENGTH = 20 #@param {type:\"integer\", min:20, max:200}\n#@markdown Rendering parameter.\nALLOW_STROKES_BEYOND_IMAGE_EDGES = True #@param {type:\"boolean\"}\n#@markdown CLIP evaluation: do we use projective transforms of images?\nUSE_PROJECTIVE_TRANSFORM = True #@param {type:\"boolean\"}\nPROJECTIVE_TRANSFORM_COEFFICIENT = 0.000001 #@param {type:\"number\"}\n#@markdown These parameters should be edited mostly only for debugging reasons.\nNET_LSTM_HIDDENS = 40 #@param {type:\"integer\"}\nNET_MLP_HIDDENS = 20 #@param {type:\"integer\"}\n# Scales the values used in genotype's initialisation.\nDNA_SCALE = 1.0 #@param {type:\"number\"}\nIMAGE_SIZE = 224 #@param {type:\"integer\"}\nVERBOSE_CODE = False #@param {type:\"boolean\"}\nVISUALIZE_GENOTYPE = False #@param {type:\"boolean\"}\nVERBOSE_MUTATION = False #@param {type:\"boolean\"}\n#@markdown Number of generations between new plots.\nPLOT_EVERY_NUM_GENS = 5 #@param {type:\"integer\"}\n#@markdown Whether to show all samples in the batch when plotting.\nPLOT_BATCH = True # @param {type:\"boolean\"}\n\nassert POPULATION_SIZE % NUM_WORKERS == 0, \"POPULATION_SIZE not multiple of NUM_WORKERS\"",
"_____no_output_____"
]
],
[
[
"#Running the original evolutionary algorithm\nThis is the original inefficient version of Arnheim which uses a genetic algorithm to optimize the picture. It takes at least 12 hours to produce an image using 50 workers. In our paper we used 500-1000 GPUs which speeded things up considerably. Refer to Arnheim 2 for a far more efficient way to generate images with a similar architecture.\n\nTry prompts like “A photorealistic chicken”. Feel free to modify this colab to include your own way of generating and evolving images like we did in figure 2 here https://arxiv.org/pdf/2105.00162.pdf.",
"_____no_output_____"
]
],
[
[
"# @title Get text input and run evolution\nPROMPT = \"an apple\" #@param {type:\"string\"}\n\n# Tokenize prompts and coompute CLIP features.\ntext_input = clip.tokenize(PROMPT).to(device)\nwith torch.no_grad():\n text_features = model.encode_text(text_input)\n\nray.shutdown()\nray.init()\n\ndrawing_lstm_arch = DrawingLSTMSpec(INPUT_SPEC_SIZE,\n NUM_LSTMS,\n NET_LSTM_HIDDENS,\n NET_MLP_HIDDENS)\n\nworkers = create_workers(NUM_WORKERS, IMAGE_SIZE, drawing_lstm_arch,\n ALLOW_STROKES_BEYOND_IMAGE_EDGES)\n\n\ndrawing_evaluator = Evaluator(IMAGE_SIZE, drawing_lstm_arch,\n ALLOW_STROKES_BEYOND_IMAGE_EDGES)\n\ndrawing_population = make_population(POPULATION_SIZE, drawing_evaluator,\n IMAGE_SIZE, INPUT_SPEC_SIZE, SEQ_LENGTH)\n\nevolution_loop(drawing_population, workers, drawing_evaluator, NUM_GENERATIONS,\n USE_CROSSOVER, CROSSOVER_PROB,\n USE_PROJECTIVE_TRANSFORM, PROJECTIVE_TRANSFORM_COEFFICIENT,\n PLOT_EVERY_NUM_GENS, PLOT_BATCH)",
"_____no_output_____"
]
]
] | [
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] | [
[
"markdown",
"markdown",
"markdown"
],
[
"code",
"code",
"code",
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
]
] |
d0ff69c2b0c3ddf00c15128fbd4e7372c1fd6d94 | 87,432 | ipynb | Jupyter Notebook | LV_analysis/1_get_multiplier_LV_coverage.ipynb | jjc2718/generic-expression-patterns | 99961ac3647d2447268ca73a94cab8b09ee08237 | [
"BSD-3-Clause"
] | null | null | null | LV_analysis/1_get_multiplier_LV_coverage.ipynb | jjc2718/generic-expression-patterns | 99961ac3647d2447268ca73a94cab8b09ee08237 | [
"BSD-3-Clause"
] | null | null | null | LV_analysis/1_get_multiplier_LV_coverage.ipynb | jjc2718/generic-expression-patterns | 99961ac3647d2447268ca73a94cab8b09ee08237 | [
"BSD-3-Clause"
] | null | null | null | 92.422833 | 20,612 | 0.834672 | [
[
[
"# Coverage of MultiPLIER LV\n\nThe goal of this notebook is to examine why genes were found to be generic. Specifically, this notebook is trying to answer the question: Are generic genes found in more multiplier latent variables compared to specific genes?\n\nThe PLIER model performs a matrix factorization of gene expression data to get two matrices: loadings (Z) and latent matrix (B). The loadings (Z) are constrained to aligned with curated pathways and gene sets specified by prior knowledge [Figure 1B of Taroni et. al.](https://www.cell.com/cell-systems/pdfExtended/S2405-4712(19)30119-X). This ensure that some but not all latent variables capture known biology. The way PLIER does this is by applying a penalty such that the individual latent variables represent a few gene sets in order to make the latent variables more interpretable. Ideally there would be one latent variable associated with one gene set unambiguously.\n\nWhile the PLIER model was trained on specific datasets, MultiPLIER extended this approach to all of recount2, where the latent variables should correspond to specific pathways or gene sets of interest. Therefore, we will look at the coverage of generic genes versus other genes across these MultiPLIER latent variables, which represent biological patterns.\n\n**Definitions:**\n* Generic genes: Are genes that are consistently differentially expressed across multiple simulated experiments.\n\n* Other genes: These are all other non-generic genes. These genes include those that are not consistently differentially expressed across simulated experiments - i.e. the genes are specifically changed in an experiment. It could also indicate genes that are consistently unchanged (i.e. housekeeping genes)",
"_____no_output_____"
]
],
[
[
"%load_ext autoreload\n%autoreload 2\n\nimport os\nimport random\nimport textwrap\nimport scipy\nimport pandas as pd\nimport seaborn as sns\nimport matplotlib.pyplot as plt\nfrom sklearn.preprocessing import MinMaxScaler\n\nimport rpy2.robjects as ro\nfrom rpy2.robjects import pandas2ri\nfrom rpy2.robjects.conversion import localconverter\n\nfrom ponyo import utils\nfrom generic_expression_patterns_modules import lv",
"/home/alexandra/anaconda3/envs/generic_expression/lib/python3.7/site-packages/matplotlib/__init__.py:886: MatplotlibDeprecationWarning: \nexamples.directory is deprecated; in the future, examples will be found relative to the 'datapath' directory.\n \"found relative to the 'datapath' directory.\".format(key))\n"
],
[
"# Get data directory containing gene summary data\nbase_dir = os.path.abspath(os.path.join(os.getcwd(), \"../\"))\ndata_dir = os.path.join(base_dir, \"human_general_analysis\")\n\n# Read in config variables\nconfig_filename = os.path.abspath(\n os.path.join(base_dir, \"configs\", \"config_human_general.tsv\")\n)\n\nparams = utils.read_config(config_filename)\n\nlocal_dir = params[\"local_dir\"]\n\nproject_id = params[\"project_id\"]\nquantile_threshold = 0.98",
"_____no_output_____"
],
[
"# Output file\nnonzero_figure_filename = \"nonzero_LV_coverage.svg\"\nhighweight_figure_filename = \"highweight_LV_coverage.svg\"",
"_____no_output_____"
]
],
[
[
"## Load data",
"_____no_output_____"
]
],
[
[
"# Get gene summary file\nsummary_data_filename = os.path.join(data_dir, f\"generic_gene_summary_{project_id}.tsv\")",
"_____no_output_____"
],
[
"# Load gene summary data\ndata = pd.read_csv(summary_data_filename, sep=\"\\t\", index_col=0, header=0)\n\n# Check that genes are unique since we will be using them as dictionary keys below\nassert data.shape[0] == len(data[\"Gene ID\"].unique())",
"_____no_output_____"
],
[
"# Load multiplier models\n# Converted formatted pickle files (loaded using phenoplier environment) from\n# https://github.com/greenelab/phenoplier/blob/master/nbs/01_preprocessing/005-multiplier_recount2_models.ipynb\n# into .tsv files\nmultiplier_model_z = pd.read_csv(\n \"multiplier_model_z.tsv\", sep=\"\\t\", index_col=0, header=0\n)",
"_____no_output_____"
],
[
"# Get a rough sense for how many genes contribute to a given LV\n# (i.e. how many genes have a value != 0 for a given LV)\n# Notice that multiPLIER is a sparse model\n(multiplier_model_z != 0).sum().sort_values(ascending=True)",
"_____no_output_____"
]
],
[
[
"## Get gene data\n\nDefine generic genes based on simulated gene ranking. Refer to [figure](https://github.com/greenelab/generic-expression-patterns/blob/master/human_general_analysis/gene_ranking_log2FoldChange.svg) as a guide.\n\n**Definitions:**\n* Generic genes: `Percentile (simulated) >= 60`\n\n(Having a high rank indicates that these genes are consistently changed across simulated experiments.)\n\n* Other genes: `Percentile (simulated) < 60`\n\n(Having a lower rank indicates that these genes are not consistently changed across simulated experiments - i.e. the genes are specifically changed in an experiment. It could also indicate genes that are consistently unchanged.)",
"_____no_output_____"
]
],
[
[
"generic_threshold = 60\ndict_genes = lv.get_generic_specific_genes(data, generic_threshold)",
"(17755, 13)\nNo. of generic genes: 7102\nNo. of other genes: 10653\n"
],
[
"# Check overlap between multiplier genes and our genes\nmultiplier_genes = list(multiplier_model_z.index)\nour_genes = list(data.index)\nshared_genes = set(our_genes).intersection(multiplier_genes)\n\nprint(len(our_genes))\nprint(len(shared_genes))",
"17755\n6374\n"
],
[
"# Drop gene ids not used in multiplier analysis\nprocessed_dict_genes = lv.process_generic_specific_gene_lists(\n dict_genes, multiplier_model_z\n)",
"_____no_output_____"
],
[
"# Check numbers add up\nassert len(shared_genes) == len(processed_dict_genes[\"generic\"]) + len(\n processed_dict_genes[\"other\"]\n)",
"_____no_output_____"
]
],
[
[
"## Get coverage of LVs\n\nFor each gene (generic or other) we want to find:\n1. The number of LVs that gene is present\n2. The number of LVs that the gene contributes a lot to (i.e. the gene is highly weighted within that LV)",
"_____no_output_____"
],
[
"### Nonzero LV coverage",
"_____no_output_____"
]
],
[
[
"dict_nonzero_coverage = lv.get_nonzero_LV_coverage(\n processed_dict_genes, multiplier_model_z\n)",
"_____no_output_____"
],
[
"# Check genes mapped correctly\nassert processed_dict_genes[\"generic\"][0] in dict_nonzero_coverage[\"generic\"].index\nassert len(dict_nonzero_coverage[\"generic\"]) == len(processed_dict_genes[\"generic\"])\nassert len(dict_nonzero_coverage[\"other\"]) == len(processed_dict_genes[\"other\"])",
"_____no_output_____"
]
],
[
[
"### High weight LV coverage",
"_____no_output_____"
]
],
[
[
"# Quick look at the distribution of gene weights per LV\nsns.distplot(multiplier_model_z[\"LV3\"], kde=False)\nplt.yscale(\"log\")",
"_____no_output_____"
],
[
"dict_highweight_coverage = lv.get_highweight_LV_coverage(\n processed_dict_genes, multiplier_model_z, quantile_threshold\n)",
"_____no_output_____"
],
[
"# Check genes mapped correctly\nassert processed_dict_genes[\"generic\"][0] in dict_highweight_coverage[\"generic\"].index\nassert len(dict_highweight_coverage[\"generic\"]) == len(processed_dict_genes[\"generic\"])\nassert len(dict_highweight_coverage[\"other\"]) == len(processed_dict_genes[\"other\"])",
"_____no_output_____"
]
],
[
[
"### Assemble LV coverage and plot",
"_____no_output_____"
]
],
[
[
"all_coverage = []\nfor gene_label in dict_genes.keys():\n merged_df = pd.DataFrame(\n dict_nonzero_coverage[gene_label], columns=[\"nonzero LV coverage\"]\n ).merge(\n pd.DataFrame(\n dict_highweight_coverage[gene_label], columns=[\"highweight LV coverage\"]\n ),\n left_index=True,\n right_index=True,\n )\n merged_df[\"gene type\"] = gene_label\n all_coverage.append(merged_df)\n\nall_coverage_df = pd.concat(all_coverage)",
"_____no_output_____"
],
[
"all_coverage_df = lv.assemble_coverage_df(\n processed_dict_genes, dict_nonzero_coverage, dict_highweight_coverage\n)\nall_coverage_df.head()",
"_____no_output_____"
],
[
"# Plot coverage distribution given list of generic coverage, specific coverage\nnonzero_fig = sns.boxplot(\n data=all_coverage_df,\n x=\"gene type\",\n y=\"nonzero LV coverage\",\n notch=True,\n palette=[\"#2c7fb8\", \"lightgrey\"],\n)\nnonzero_fig.set_xlabel(None)\nnonzero_fig.set_xticklabels(\n [\"generic genes\", \"other genes\"], fontsize=14, fontname=\"Verdana\"\n)\nnonzero_fig.set_ylabel(\n textwrap.fill(\"Number of LVs\", width=30), fontsize=14, fontname=\"Verdana\"\n)\nnonzero_fig.tick_params(labelsize=14)\nnonzero_fig.set_title(\n \"Number of LVs genes are present in\", fontsize=16, fontname=\"Verdana\"\n)",
"_____no_output_____"
],
[
"# Plot coverage distribution given list of generic coverage, specific coverage\nhighweight_fig = sns.boxplot(\n data=all_coverage_df,\n x=\"gene type\",\n y=\"highweight LV coverage\",\n notch=True,\n palette=[\"#2c7fb8\", \"lightgrey\"],\n)\nhighweight_fig.set_xlabel(None)\nhighweight_fig.set_xticklabels(\n [\"generic genes\", \"other genes\"], fontsize=14, fontname=\"Verdana\"\n)\nhighweight_fig.set_ylabel(\n textwrap.fill(\"Number of LVs\", width=30), fontsize=14, fontname=\"Verdana\"\n)\nhighweight_fig.tick_params(labelsize=14)\nhighweight_fig.set_title(\n \"Number of LVs genes contribute highly to\", fontsize=16, fontname=\"Verdana\"\n)",
"_____no_output_____"
]
],
[
[
"## Calculate statistics\n* Is the reduction in generic coverage significant?\n* Is the difference between generic versus other genes signficant?",
"_____no_output_____"
]
],
[
[
"# Test: mean number of LVs generic genes present in vs mean number of LVs that generic gene is high weight in\n# (compare two blue boxes between plots)\ngeneric_nonzero = all_coverage_df[all_coverage_df[\"gene type\"] == \"generic\"][\n \"nonzero LV coverage\"\n].values\ngeneric_highweight = all_coverage_df[all_coverage_df[\"gene type\"] == \"generic\"][\n \"highweight LV coverage\"\n].values\n\n(stats, pvalue) = scipy.stats.ttest_ind(generic_nonzero, generic_highweight)\nprint(pvalue)",
"0.0\n"
],
[
"# Test: mean number of LVs generic genes present in vs mean number of LVs other genes high weight in\n# (compare blue and grey boxes in high weight plot)\nother_highweight = all_coverage_df[all_coverage_df[\"gene type\"] == \"other\"][\n \"highweight LV coverage\"\n].values\ngeneric_highweight = all_coverage_df[all_coverage_df[\"gene type\"] == \"generic\"][\n \"highweight LV coverage\"\n].values\n\n(stats, pvalue) = scipy.stats.ttest_ind(other_highweight, generic_highweight)\nprint(pvalue)",
"6.307987998525766e-119\n"
],
[
"# Check that coverage of other and generic genes across all LVs is NOT signficantly different\n# (compare blue and grey boxes in nonzero weight plot)\nother_nonzero = all_coverage_df[all_coverage_df[\"gene type\"] == \"other\"][\n \"nonzero LV coverage\"\n].values\ngeneric_nonzero = all_coverage_df[all_coverage_df[\"gene type\"] == \"generic\"][\n \"nonzero LV coverage\"\n].values\n\n(stats, pvalue) = scipy.stats.ttest_ind(other_nonzero, generic_nonzero)\nprint(pvalue)",
"0.23947472582519233\n"
]
],
[
[
"## Get LVs that generic genes are highly weighted in\n\nSince we are using quantiles to get high weight genes per LV, each LV has the same number of high weight genes. For each set of high weight genes, we will get the proportion of generic vs other genes. We will select the LVs that have a high proportion of generic genes to examine.",
"_____no_output_____"
]
],
[
[
"# Get proportion of generic genes per LV\nprop_highweight_generic_dict = lv.get_prop_highweight_generic_genes(\n processed_dict_genes, multiplier_model_z, quantile_threshold\n)",
"135\n"
],
[
"# Return selected rows from summary matrix\nmultiplier_model_summary = pd.read_csv(\n \"multiplier_model_summary.tsv\", sep=\"\\t\", index_col=0, header=0\n)\nlv.create_LV_df(\n prop_highweight_generic_dict,\n multiplier_model_summary,\n 0.5,\n \"Generic_LV_summary_table.tsv\",\n)",
"LV2 0.5185185185185185\nLV3 0.6444444444444445\nLV7 0.562962962962963\nLV11 0.5851851851851851\nLV17 0.5703703703703704\nLV18 0.5333333333333333\nLV22 0.5185185185185185\nLV26 0.5111111111111111\nLV32 0.5481481481481482\nLV34 0.5333333333333333\nLV54 0.562962962962963\nLV57 0.5259259259259259\nLV58 0.5703703703703704\nLV61 0.6222222222222222\nLV68 0.6296296296296297\nLV101 0.5481481481481482\nLV135 0.5185185185185185\nLV473 0.5481481481481482\nLV524 0.6074074074074074\nLV542 0.5259259259259259\nLV603 0.5185185185185185\nLV719 0.6074074074074074\nLV728 0.6074074074074074\nLV765 0.6370370370370371\nLV767 0.5333333333333333\nLV787 0.5037037037037037\nLV823 0.6222222222222222\nLV913 0.5777777777777777\nLV920 0.6074074074074074\nLV932 0.5185185185185185\nLV958 0.5333333333333333\nLV960 0.5333333333333333\nLV977 0.5481481481481482\n"
],
[
"# Plot distribution of weights for these nodes\nnode = \"LV61\"\nlv.plot_dist_weights(\n node,\n multiplier_model_z,\n shared_genes,\n 20,\n all_coverage_df,\n f\"weight_dist_{node}.svg\",\n)",
"0 6.540359\n1 6.527677\n2 2.738563\n3 1.470443\n4 1.272303\n5 0.982001\n6 0.842076\n7 0.715623\n8 0.714085\n9 0.615867\n10 0.567222\n11 0.535279\n12 0.532891\n13 0.525537\n14 0.522841\n15 0.483024\n16 0.475159\n17 0.469341\n18 0.462943\n19 0.439901\nName: LV61, dtype: float64\n"
]
],
[
[
"## Save",
"_____no_output_____"
]
],
[
[
"# Save plot\nnonzero_fig.figure.savefig(\n nonzero_figure_filename,\n format=\"svg\",\n bbox_inches=\"tight\",\n transparent=True,\n pad_inches=0,\n dpi=300,\n)\n\n# Save plot\nhighweight_fig.figure.savefig(\n highweight_figure_filename,\n format=\"svg\",\n bbox_inches=\"tight\",\n transparent=True,\n pad_inches=0,\n dpi=300,\n)",
"_____no_output_____"
]
],
[
[
"**Takeaway:**\n* In the first nonzero boxplot, generic and other genes are present in a similar number of LVs. This isn't surprising since the number of genes that contribute to each LV is <1000.\n* In the second highweight boxplot, other genes are highly weighted in more LVs compared to generic genes. This would indicate that generic genes contribute alot to few LVs.\n\nThis is the opposite trend found using [_P. aeruginosa_ data](1_get_eADAGE_LV_coverage.ipynb). Perhaps this indicates that generic genes have different behavior/roles depending on the organism. In humans, perhaps these generic genes are related to a few hyper-responsive pathways, whereas in _P. aeruginosa_ perhaps generic genes are associated with many pathways, acting as *gene hubs*.\n\n* There are a number of LVs that contain a high proportion of generic genes can be found in [table](Generic_LV_summary_table.tsv). By quick visual inspection, it looks like many LVs are associated with immune response, signaling and metabolism. Which are consistent with the hypothesis that these generic genes are related to hyper-responsive pathways.",
"_____no_output_____"
]
]
] | [
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown"
] | [
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
]
] |
d0ff73244d3c6641018427145fa6435227431b8a | 161,880 | ipynb | Jupyter Notebook | Gradient Boosting for molecular properties/pmp-lightgbm-inference.ipynb | dhananjayraut/ML_projects | e7b5008c4039bfa057cc6f7d991224fd2d268eb6 | [
"MIT"
] | null | null | null | Gradient Boosting for molecular properties/pmp-lightgbm-inference.ipynb | dhananjayraut/ML_projects | e7b5008c4039bfa057cc6f7d991224fd2d268eb6 | [
"MIT"
] | null | null | null | Gradient Boosting for molecular properties/pmp-lightgbm-inference.ipynb | dhananjayraut/ML_projects | e7b5008c4039bfa057cc6f7d991224fd2d268eb6 | [
"MIT"
] | null | null | null | 71.691763 | 17,640 | 0.716277 | [
[
[
"# This Python 3 environment comes with many helpful analytics libraries installed\n# It is defined by the kaggle/python docker image: https://github.com/kaggle/docker-python\n# For example, here's several helpful packages to load in \nimport os\nimport random\nimport xgboost\nimport lightgbm as lgb\nimport numpy as np # linear algebra\nimport pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)\nimport matplotlib.pyplot as plt\nfrom os import listdir\nfrom os.path import isfile, join\nfrom multiprocessing import Pool\nfrom sklearn.model_selection import train_test_split\nfrom sklearn.metrics import mean_absolute_error\nfrom sklearn.model_selection import GridSearchCV \n#from sklearn.model_selection import cross_val_score,KFold\n# Input data files are available in the \"../input/\" directory.\n# For example, running this (by clicking run or pressing Shift+Enter) will list the files in the input directory\n# Any results you write to the current directory are saved as output.",
"_____no_output_____"
],
[
"''' from here https://www.kaggle.com/c/imet-2019-fgvc6/discussion/87675#latest-516375'''\ndef seed_everything(seed=1234):\n random.seed(seed)\n os.environ['PYTHONHASHSEED'] = str(seed)\n np.random.seed(seed)\nseed_everything()",
"_____no_output_____"
],
[
"train = pd.read_csv('../input/pmp-data-train/train_.csv')\ntrain.drop('molecule_name', axis=1,inplace=True)\ntrain.drop('atom_index_0' , axis=1,inplace=True)\ntrain.drop('atom_index_1' , axis=1,inplace=True)\ntrain.drop('id' , axis=1,inplace=True)\ntrain.drop(columns=\"Unnamed: 0\",inplace=True)",
"_____no_output_____"
],
[
"train.head()",
"_____no_output_____"
],
[
"def group_mean_log_mae(y_true, y_pred, floor=1e-9):\n maes = np.absolute(y_true-y_pred).mean()\n return np.log(np.maximum(maes,floor).mean())",
"_____no_output_____"
],
[
"def mean_log_mae( y_pred, dataset, floor=1e-9):\n y_true = dataset.get_label()\n maes = np.absolute(y_true-y_pred).mean()\n return 'mean_log_mae', np.log(np.maximum(maes,floor).mean()), False",
"_____no_output_____"
],
[
"def show_predictions(true, pred):\n # print(' actual | prediction | differnece')\n # for i in range(len(true)):\n # print('{:10.5f} | {:10.5f} | {:10.5f}'.format(true[i], pred[i], abs(true[i] - pred[i]) ))\n plt.scatter(true,pred)\n plt.show()",
"_____no_output_____"
],
[
"??plt.scatter()",
"Object `plt.scatter()` not found.\n"
],
[
"files = [ '../input/pmp-lightgbm/model_1JHC.txt',\n '../input/pmp-lightgbm/model_2JHH.txt',\n '../input/pmp-lightgbm/model_1JHN.txt',\n '../input/pmp-lightgbm/model_2JHN.txt',\n '../input/fork-of-pmp-lightgbm/model_2JHC.txt',\n '../input/fork-of-pmp-lightgbm/model_3JHH.txt',\n '../input/fork-of-pmp-lightgbm/model_3JHC.txt',\n '../input/fork-of-pmp-lightgbm/model_3JHN.txt',]",
"_____no_output_____"
],
[
"data_fr = {}\nmodels = {}\ntype_list = ['1JHC', '2JHH', '1JHN', '2JHN', '2JHC', '3JHH', '3JHC', '3JHN']\nscores = []\nfor i,type_ in enumerate(type_list):\n data_fr = train.loc[train['type'] == type_].copy()\n data_fr.reset_index(inplace=True)\n data_fr.drop('type', axis=1,inplace=True)\n data_fr.drop('index', axis=1,inplace=True)\n y = data_fr.scalar_coupling_constant.values\n data_fr.drop('scalar_coupling_constant', axis=1,inplace=True)\n X = data_fr.astype(float).values\n features = data_fr.columns\n print('Dataset for type {} has shape {}'.format(type_,X.shape))\n X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.05, random_state=42)\n reg = lgb.Booster(model_file=files[i])\n show_predictions(y_test[0:500],reg.predict(X_test[0:500]).tolist())\n score = group_mean_log_mae(y_test,reg.predict(X_test))\n scores.append(score)\n models[type_] = reg\n print('score for type {} is --------> {}'.format(type_,score))",
"Dataset for type 1JHC has shape (709416, 53)\n"
],
[
"print('Final score is {}'.format(np.array(scores).mean()))\nfor i,type_ in enumerate(type_list): print('score of type {} is {}'.format(type_,scores[i]))",
"Final score is -1.7321338657617118\nscore of type 1JHC is -0.5625430329149439\nscore of type 2JHH is -2.061194960115891\nscore of type 1JHN is -1.5026866055361172\nscore of type 2JHN is -2.3744295726617057\nscore of type 2JHC is -1.4289135609371875\nscore of type 3JHH is -1.926762765385606\nscore of type 3JHC is -1.3504152515798082\nscore of type 3JHN is -2.6501251769624345\n"
],
[
"plt.figure(figsize = (12,6))\nplt.bar(range(len(reg.feature_importance())), reg.feature_importance())\nplt.xticks(range(len(reg.feature_importance())), features, rotation='vertical')\nplt.show()",
"_____no_output_____"
],
[
"test = pd.read_csv('../input/pmp-data-test/test_.csv')\ntest.drop('molecule_name', axis=1,inplace=True)\ntest.drop('atom_index_0' , axis=1,inplace=True)\ntest.drop('atom_index_1' , axis=1,inplace=True)\ntest.drop('id' , axis=1,inplace=True)\ntest.drop(\"Unnamed: 0\" , axis=1,inplace=True)",
"_____no_output_____"
],
[
"test.head()",
"_____no_output_____"
],
[
"test_ = test.copy()\ntest_.drop('type', axis=1,inplace=True)\nX = test_.astype(float).values\nfeatures = test_.columns\nprint(X.shape)\ntest_.head()",
"(2505542, 53)\n"
],
[
"sub = pd.read_csv('../input/champs-scalar-coupling/sample_submission.csv')\nsub['scalar_coupling_constant'] = sub['scalar_coupling_constant'].astype(float)",
"_____no_output_____"
],
[
"sub.head()",
"_____no_output_____"
],
[
"%%time\nfrom tqdm import tnrange\nvalues = np.zeros(sub.shape[0])\nsub['scalar_coupling_constant'] = sub['scalar_coupling_constant'].astype(float)\nfor i in tnrange(sub.shape[0]):\n type_ = test['type'][i]\n values[i] = models[type_].predict(np.expand_dims(X[i], axis=0))\nsub['scalar_coupling_constant'] = pd.DataFrame(np.array(values))",
"_____no_output_____"
],
[
"print(sub.head())\nsub.to_csv('submission.csv',index=None)",
" id scalar_coupling_constant\n0 4658147 18.002834\n1 4658148 193.190463\n2 4658149 9.227583\n3 4658150 193.190463\n4 4658151 18.002834\n"
]
],
[
[
"<a href=\"submission.csv\"> Download File </a>",
"_____no_output_____"
]
],
[
[
"# def tt(j):\n# type_ = test['type'][j]\n# return models[type_].predict(np.expand_dims(X[j], axis=0)) \n# def get_dataframe():\n# with Pool(8) as p:\n# a = p.map(tt,[i for i in range(sub.shape[0])])\n# return pd.DataFrame(np.array(a))",
"_____no_output_____"
],
[
"# %%time\n# sub['scalar_coupling_constant'] = get_dataframe()",
"_____no_output_____"
],
[
"# sam['scalar_coupling_constant'] = sam['scalar_coupling_constant'].astype(float)\n# for i in range(sam.shape[0]):\n# type_ = test['type'][i]\n# value = preds[type_][i]\n# sam.at[i, 'scalar_coupling_constant'] = value",
"_____no_output_____"
],
[
"# parameters_for_testing = {\n# 'gamma':[0,0.03,0.1,0.3],\n# 'min_child_weight':[1.5,6,10],\n# 'learning_rate':[0.07,0.1,0.5,1],\n# 'max_depth':[3,4,5,6],\n# 'n_estimators':[10],\n# }\n\n# xgb_model = xgboost.XGBRegressor(learning_rate =0.1, n_estimators=10, max_depth=6,\n# min_child_weight=1, gamma=0, subsample=0.8,\n# colsample_bytree=0.8, nthread=6, \n# scale_pos_weight=1, seed=42)\n\n# gsearch = GridSearchCV(estimator = xgb_model, cv=3, param_grid=parameters_for_testing, n_jobs=10,iid=False, verbose=10, scoring='neg_mean_squared_error')\n# gsearch.fit(X_train,y_train)\n# print('best params')\n# print (gsearch.best_params_)\n# print('best score')\n# print (gsearch.best_score_)",
"_____no_output_____"
],
[
"# parameters_for_testing = {\n# 'reg_alpha':[0,0.03,0.1,0.3],\n# 'reg_labda':[1.5,6,10],\n# 'n_estimators':[100],\n# }\n\n# xgb_model = xgboost.XGBRegressor(learning_rate=0.5, \n# n_estimators=100,\n# max_depth=6,\n# min_child_weight=1.5,\n# gamma=0,\n# nthread=6, \n# scale_pos_weight=1,\n# seed=42)\n\n# gsearch1 = GridSearchCV(estimator = xgb_model, cv=3, param_grid=parameters_for_testing, n_jobs=10,iid=False, verbose=10, scoring='neg_mean_squared_error')\n# gsearch1.fit(X_train,y_train)\n# print('best params')\n# print (gsearch1.best_params_)\n# print('best score')\n# print (gsearch1.best_score_)",
"_____no_output_____"
],
[
"# reg = xgboost.XGBRegressor(\n# learning_rate=0.5, \n# n_estimators=100,\n# max_depth=6,\n# min_child_weight=1.5,\n# gamma=0,\n# nthread=6, \n# scale_pos_weight=1,\n# reg_alpha=0.3,\n# reg_lambda=1.5,\n# seed=42\n# )\n# reg.fit(X_train,y_train);\n# print('score is {}'.format(group_mean_log_mae(pd.DataFrame(y_test),pd.DataFrame(reg.predict(X_test)))))",
"_____no_output_____"
],
[
"# plt.figure(figsize = (12,6))\n# plt.bar(range(len(reg.feature_importances_)), reg.feature_importances_)\n# plt.xticks(range(len(reg.feature_importances_)), fea, rotation='vertical')\n# plt.show()",
"_____no_output_____"
],
[
"# from sklearn import linear_model\n# reg = linear_model.RidgeCV()\n# reg.fit(X_train, y_train)\n# group_mean_log_mae(pd.DataFrame(y_test),pd.DataFrame(reg.predict(X_test)))\n# print(y_train[0:5])\n# print(reg.predict(X_train[0:5]).tolist())",
"_____no_output_____"
],
[
"# from sklearn.svm import SVR\n# reg = SVR(gamma='scale', kernel='rbf', max_iter=2000)\n# reg.fit(X_train, y_train)\n# group_mean_log_mae(pd.DataFrame(y_test),pd.DataFrame(reg.predict(X_test)))",
"_____no_output_____"
],
[
"# import torch\n# X_trainc = torch.tensor(X_train,dtype=torch.float).cuda()\n# Y_trainc = torch.tensor(y_train,dtype=torch.float).cuda() \n# X_testc = torch.tensor(X_test,dtype=torch.float).cuda()\n# Y_testc = torch.tensor(y_test,dtype=torch.float).cuda()\n# class Net(torch.nn.Module):\n# def __init__(self, D_in, D_out):\n# super(Net, self).__init__()\n# self.linear1 = torch.nn.Linear(D_in, 100)\n# self.linear2 = torch.nn.Linear(100, 20)\n# self.linear3 = torch.nn.Linear(20, D_out)\n# self.drop1 = torch.nn.Dropout(p=0.45, inplace=False)\n# self.drop2 = torch.nn.Dropout(p=0.45, inplace=False)\n# self.relu = torch.nn.ReLU()\n# def forward(self, x):\n# x = self.relu(self.linear1(x))\n# x = self.drop1(x)\n# x = self.relu(self.linear2(x))\n# x = self.drop2(x)\n# y_pred = self.linear3(x)\n# return y_pred\n# model = Net(44,1).cuda()\n# criterion = torch.nn.MSELoss()\n# optimizer = torch.optim.Adam(model.parameters(), lr=1e-3)\n# step = torch.optim.lr_scheduler.StepLR(optimizer,100, gamma = 0.5, last_epoch=-1)\n# def acc(model):\n# y_p = model(X_testc).detach().cpu().numpy()\n# print(group_mean_log_mae(pd.DataFrame(y_test),pd.DataFrame(y_p)))\n# acc(model)\n# model.train()\n# for i in range(1501):\n# if i < 310: step.step()\n# model.train()\n# optimizer.zero_grad()\n# y_pred = model(X_testc)\n# loss = criterion(y_pred, Y_testc)\n# loss.backward()\n# optimizer.step()\n# if i%30 == 0: acc(model)",
"_____no_output_____"
],
[
"# lgb_train = lgb.Dataset(X_train, y_train)\n# lgb_eval = lgb.Dataset(X_test, y_test, reference=lgb_train)\n\n# def mean_log_mae( y_pred, dataset, floor=1e-9):\n# y_true = dataset.get_label()\n# maes = (pd.DataFrame(y_true)-pd.DataFrame(y_pred)).abs().mean()\n# return 'mean_log_mae', np.log(maes.map(lambda x: max(x, floor))).mean(), False\n\n# def mean_log_mae( y_pred, dataset, floor=1e-9):\n# y_true = dataset.get_label()\n# maes = np.absolute(y_true-y_pred).mean()\n# return np.log(np.maximum(maes,floor).mean())\n\n# params = {\n# 'num_leaves' : [40],\n# 'learning_rate' : [0.2, 0.3, 0.4],\n# 'num_boost_round' : [4000]\n# }\n\n# print('Starting training...')\n# # train\n# reg = lgb.LGBMRegressor(boosting_type ='gbdt',\n# objective = 'regression',\n# verbose_eval = 200)\n\n\n# gbm = GridSearchCV(estimator = reg, cv=3, param_grid=params, n_jobs=6, iid=False, verbose=10, scoring='neg_mean_squared_error')\n\n# gbm.fit(X_train, y_train)\n# print('best params')\n# print (gbm.best_params_)\n# print('best score')\n# print (gbm.best_score_)",
"_____no_output_____"
]
]
] | [
"code",
"markdown",
"code"
] | [
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
d0ff75b136a1b2e831b44b27f7e476b2ef697a72 | 11,183 | ipynb | Jupyter Notebook | II Machine Learning & Deep Learning/02_Decision Tree. A Supervised Classification Model/02session_decision-tree.ipynb | Blanes3/resolving-python-data-science | 6c97821d09fe55a3069825f0a3913b5fd29280fe | [
"MIT"
] | null | null | null | II Machine Learning & Deep Learning/02_Decision Tree. A Supervised Classification Model/02session_decision-tree.ipynb | Blanes3/resolving-python-data-science | 6c97821d09fe55a3069825f0a3913b5fd29280fe | [
"MIT"
] | null | null | null | II Machine Learning & Deep Learning/02_Decision Tree. A Supervised Classification Model/02session_decision-tree.ipynb | Blanes3/resolving-python-data-science | 6c97821d09fe55a3069825f0a3913b5fd29280fe | [
"MIT"
] | null | null | null | 23.592827 | 271 | 0.429223 | [
[
[
"<font size=\"+5\">#02 | Decision Tree. A Supervised Classification Model</font>",
"_____no_output_____"
],
[
"- Subscribe to my [Blog ↗](https://blog.pythonassembly.com/)\n- Let's keep in touch on [LinkedIn ↗](www.linkedin.com/in/jsulopz) 😄",
"_____no_output_____"
],
[
"# Discipline to Search Solutions in Google",
"_____no_output_____"
],
[
"> Apply the following steps when **looking for solutions in Google**:\n>\n> 1. **Necesity**: How to load an Excel in Python?\n> 2. **Search in Google**: by keywords\n> - `load excel python`\n> - ~~how to load excel in python~~\n> 3. **Solution**: What's the `function()` that loads an Excel in Python?\n> - A Function to Programming is what the Atom to Phisics.\n> - Every time you want to do something in programming\n> - **You will need a `function()`** to make it\n> - Theferore, you must **detect parenthesis `()`**\n> - Out of all the words that you see in a website\n> - Because they indicate the presence of a `function()`.",
"_____no_output_____"
],
[
"# Load the Data",
"_____no_output_____"
],
[
"> Load the Titanic dataset with the below commands\n> - This dataset **people** (rows) aboard the Titanic\n> - And their **sociological characteristics** (columns)\n> - The aim of this dataset is to predict the probability to `survive`\n> - Based on the social demographic characteristics.",
"_____no_output_____"
]
],
[
[
"import seaborn as sns\n\nsns.load_dataset(name='titanic').iloc[:, :4]",
"_____no_output_____"
]
],
[
[
"# `DecisionTreeClassifier()` Model in Python",
"_____no_output_____"
],
[
"## Build the Model",
"_____no_output_____"
],
[
"> 1. **Necesity**: Build Model\n> 2. **Google**: How do you search for the solution?\n> 3. **Solution**: Find the `function()` that makes it happen",
"_____no_output_____"
],
[
"## Code Thinking\n\n> Which function computes the Model?\n> - `fit()`\n>\n> How could can you **import the function in Python**?",
"_____no_output_____"
],
[
"### Separate Variables for the Model\n\n> Regarding their role:\n> 1. **Target Variable `y`**\n>\n> - [ ] What would you like **to predict**?\n>\n> 2. **Explanatory Variable `X`**\n>\n> - [ ] Which variable will you use **to explain** the target?",
"_____no_output_____"
]
],
[
[
"explanatory = ?\ntarget = ?",
"_____no_output_____"
]
],
[
[
"### Finally `fit()` the Model",
"_____no_output_____"
],
[
"## Calculate a Prediction with the Model",
"_____no_output_____"
],
[
"> - `model.predict_proba()`",
"_____no_output_____"
],
[
"## Model Visualization",
"_____no_output_____"
],
[
"> - `tree.plot_tree()`",
"_____no_output_____"
],
[
"## Model Interpretation",
"_____no_output_____"
],
[
"> Why `sex` is the most important column? What has to do with **EDA** (Exploratory Data Analysis)?",
"_____no_output_____"
]
],
[
[
"%%HTML\n\n<iframe width=\"560\" height=\"315\" src=\"https://www.youtube.com/embed/7VeUPuFGJHk\" title=\"YouTube video player\" frameborder=\"0\" allow=\"accelerometer; autoplay; clipboard-write; encrypted-media; gyroscope; picture-in-picture\" allowfullscreen></iframe>",
"_____no_output_____"
]
],
[
[
"# Prediction vs Reality",
"_____no_output_____"
],
[
"> How good is our model?",
"_____no_output_____"
],
[
"## Precision",
"_____no_output_____"
],
[
"> - `model.score()`",
"_____no_output_____"
],
[
"## Confusion Matrix",
"_____no_output_____"
],
[
"> 1. **Sensitivity** (correct prediction on positive value, $y=1$)\n> 2. **Specificity** (correct prediction on negative value $y=0$).",
"_____no_output_____"
],
[
"## ROC Curve",
"_____no_output_____"
],
[
"> A way to summarise all the metrics (score, sensitivity & specificity)",
"_____no_output_____"
]
]
] | [
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown"
] | [
[
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown",
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown"
]
] |
d0ff782a9e491f45bec93afc3a7d4f4d1194d348 | 182,562 | ipynb | Jupyter Notebook | use_PaddleOCR_colab.ipynb | hieu28022000/PaddleOCR | a4ea255edf07405ec8ae64c1ded9109dae56d94f | [
"Apache-2.0"
] | null | null | null | use_PaddleOCR_colab.ipynb | hieu28022000/PaddleOCR | a4ea255edf07405ec8ae64c1ded9109dae56d94f | [
"Apache-2.0"
] | null | null | null | use_PaddleOCR_colab.ipynb | hieu28022000/PaddleOCR | a4ea255edf07405ec8ae64c1ded9109dae56d94f | [
"Apache-2.0"
] | null | null | null | 182,562 | 182,562 | 0.794788 | [
[
[
"from google.colab import drive\ndrive.mount('/content/drive')",
"Mounted at /content/drive\n"
],
[
"import os\nos.chdir('/content/drive/My Drive/Reg_text/PaddleOCR/')\nos.getcwd()",
"_____no_output_____"
],
[
"!pip install paddlepaddle",
"Collecting paddlepaddle\n\u001b[?25l Downloading https://files.pythonhosted.org/packages/2a/12/273ec0cdf3164e610aa197425cfcaffcd4b32aa2b7a29ba628b5f29e95c9/paddlepaddle-2.1.0-cp37-cp37m-manylinux1_x86_64.whl (108.8MB)\n\u001b[K |████████████████████████████████| 108.8MB 91kB/s \n\u001b[?25hRequirement already satisfied: numpy>=1.13; python_version >= \"3.5\" and platform_system != \"Windows\" in /usr/local/lib/python3.7/dist-packages (from paddlepaddle) (1.19.5)\nRequirement already satisfied: protobuf>=3.1.0 in /usr/local/lib/python3.7/dist-packages (from paddlepaddle) (3.12.4)\nRequirement already satisfied: six in /usr/local/lib/python3.7/dist-packages (from paddlepaddle) (1.15.0)\nRequirement already satisfied: gast>=0.3.3; platform_system != \"Windows\" in /usr/local/lib/python3.7/dist-packages (from paddlepaddle) (0.4.0)\nRequirement already satisfied: astor in /usr/local/lib/python3.7/dist-packages (from paddlepaddle) (0.8.1)\nRequirement already satisfied: Pillow in /usr/local/lib/python3.7/dist-packages (from paddlepaddle) (7.1.2)\nRequirement already satisfied: decorator==4.4.2 in /usr/local/lib/python3.7/dist-packages (from paddlepaddle) (4.4.2)\nRequirement already satisfied: requests>=2.20.0 in /usr/local/lib/python3.7/dist-packages (from paddlepaddle) (2.23.0)\nRequirement already satisfied: setuptools in /usr/local/lib/python3.7/dist-packages (from protobuf>=3.1.0->paddlepaddle) (56.1.0)\nRequirement already satisfied: chardet<4,>=3.0.2 in /usr/local/lib/python3.7/dist-packages (from requests>=2.20.0->paddlepaddle) (3.0.4)\nRequirement already satisfied: certifi>=2017.4.17 in /usr/local/lib/python3.7/dist-packages (from requests>=2.20.0->paddlepaddle) (2020.12.5)\nRequirement already satisfied: idna<3,>=2.5 in /usr/local/lib/python3.7/dist-packages (from requests>=2.20.0->paddlepaddle) (2.10)\nRequirement already satisfied: urllib3!=1.25.0,!=1.25.1,<1.26,>=1.21.1 in /usr/local/lib/python3.7/dist-packages (from requests>=2.20.0->paddlepaddle) (1.24.3)\nInstalling collected packages: paddlepaddle\nSuccessfully installed paddlepaddle-2.1.0\n"
],
[
"!pip install \"paddleocr==2.0.6\" # 2.0.6 version is recommended",
"Collecting paddleocr==2.0.6\n\u001b[?25l Downloading https://files.pythonhosted.org/packages/1b/76/3ae5a1d95798039eaf3e3371b391906539da6e1cdad2a16a496a4a4f361d/paddleocr-2.0.6-py3-none-any.whl (189kB)\n\u001b[K |████████████████████████████████| 194kB 8.7MB/s \n\u001b[?25hCollecting imgaug==0.4.0\n\u001b[?25l Downloading https://files.pythonhosted.org/packages/66/b1/af3142c4a85cba6da9f4ebb5ff4e21e2616309552caca5e8acefe9840622/imgaug-0.4.0-py2.py3-none-any.whl (948kB)\n\u001b[K |████████████████████████████████| 952kB 17.1MB/s \n\u001b[?25hCollecting python-Levenshtein\n\u001b[?25l Downloading https://files.pythonhosted.org/packages/2a/dc/97f2b63ef0fa1fd78dcb7195aca577804f6b2b51e712516cc0e902a9a201/python-Levenshtein-0.12.2.tar.gz (50kB)\n\u001b[K |████████████████████████████████| 51kB 8.3MB/s \n\u001b[?25hRequirement already satisfied: opencv-contrib-python in /usr/local/lib/python3.7/dist-packages (from paddleocr==2.0.6) (4.1.2.30)\nRequirement already satisfied: shapely in /usr/local/lib/python3.7/dist-packages (from paddleocr==2.0.6) (1.7.1)\nRequirement already satisfied: numpy in /usr/local/lib/python3.7/dist-packages (from paddleocr==2.0.6) (1.19.5)\nRequirement already satisfied: lmdb in /usr/local/lib/python3.7/dist-packages (from paddleocr==2.0.6) (0.99)\nRequirement already satisfied: tqdm in /usr/local/lib/python3.7/dist-packages (from paddleocr==2.0.6) (4.41.1)\nCollecting scikit-image==0.17.2\n\u001b[?25l Downloading https://files.pythonhosted.org/packages/d7/ee/753ea56fda5bc2a5516a1becb631bf5ada593a2dd44f21971a13a762d4db/scikit_image-0.17.2-cp37-cp37m-manylinux1_x86_64.whl (12.5MB)\n\u001b[K |████████████████████████████████| 12.5MB 15.3MB/s \n\u001b[?25hCollecting visualdl\n\u001b[?25l Downloading https://files.pythonhosted.org/packages/31/99/f5f50d035006b0d9304700facd9e1c843af8e02569474996d1b6a79529f6/visualdl-2.2.0-py3-none-any.whl (2.7MB)\n\u001b[K |████████████████████████████████| 2.7MB 46.8MB/s \n\u001b[?25hCollecting pyclipper\n\u001b[?25l Downloading https://files.pythonhosted.org/packages/69/5b/92df65d3e1e5c5623e67feeac92a18d28b0bf11bdd44d200245611b0fbb8/pyclipper-1.2.1-cp37-cp37m-manylinux1_x86_64.whl (126kB)\n\u001b[K |████████████████████████████████| 133kB 57.7MB/s \n\u001b[?25hCollecting opencv-python==4.2.0.32\n\u001b[?25l Downloading https://files.pythonhosted.org/packages/34/a3/403dbaef909fee9f9f6a8eaff51d44085a14e5bb1a1ff7257117d744986a/opencv_python-4.2.0.32-cp37-cp37m-manylinux1_x86_64.whl (28.2MB)\n\u001b[K |████████████████████████████████| 28.2MB 159kB/s \n\u001b[?25hRequirement already satisfied: six in /usr/local/lib/python3.7/dist-packages (from imgaug==0.4.0->paddleocr==2.0.6) (1.15.0)\nRequirement already satisfied: scipy in /usr/local/lib/python3.7/dist-packages (from imgaug==0.4.0->paddleocr==2.0.6) (1.4.1)\nRequirement already satisfied: Pillow in /usr/local/lib/python3.7/dist-packages (from imgaug==0.4.0->paddleocr==2.0.6) (7.1.2)\nRequirement already satisfied: matplotlib in /usr/local/lib/python3.7/dist-packages (from imgaug==0.4.0->paddleocr==2.0.6) (3.2.2)\nRequirement already satisfied: imageio in /usr/local/lib/python3.7/dist-packages (from imgaug==0.4.0->paddleocr==2.0.6) (2.4.1)\nRequirement already satisfied: setuptools in /usr/local/lib/python3.7/dist-packages (from python-Levenshtein->paddleocr==2.0.6) (56.1.0)\nRequirement already satisfied: networkx>=2.0 in /usr/local/lib/python3.7/dist-packages (from scikit-image==0.17.2->paddleocr==2.0.6) (2.5.1)\nRequirement already satisfied: tifffile>=2019.7.26 in /usr/local/lib/python3.7/dist-packages (from scikit-image==0.17.2->paddleocr==2.0.6) (2021.4.8)\nRequirement already satisfied: PyWavelets>=1.1.1 in /usr/local/lib/python3.7/dist-packages (from scikit-image==0.17.2->paddleocr==2.0.6) (1.1.1)\nRequirement already satisfied: pandas in /usr/local/lib/python3.7/dist-packages (from visualdl->paddleocr==2.0.6) (1.1.5)\nRequirement already satisfied: protobuf>=3.11.0 in /usr/local/lib/python3.7/dist-packages (from visualdl->paddleocr==2.0.6) (3.12.4)\nCollecting shellcheck-py\n\u001b[?25l Downloading https://files.pythonhosted.org/packages/77/c9/6f84db444de69e1bdbfd168ae6f3d5af72107c16cd9db3c0a1f8e41204d1/shellcheck_py-0.7.2.1-py2.py3-none-manylinux1_x86_64.whl (2.0MB)\n\u001b[K |████████████████████████████████| 2.0MB 38.4MB/s \n\u001b[?25hRequirement already satisfied: flask>=1.1.1 in /usr/local/lib/python3.7/dist-packages (from visualdl->paddleocr==2.0.6) (1.1.4)\nRequirement already satisfied: requests in /usr/local/lib/python3.7/dist-packages (from visualdl->paddleocr==2.0.6) (2.23.0)\nCollecting bce-python-sdk\n\u001b[?25l Downloading https://files.pythonhosted.org/packages/09/c2/6336452f3cd267c525e1142d6f20f70dd0abc4504105d8a04e79df16b54a/bce_python_sdk-0.8.60-py3-none-any.whl (197kB)\n\u001b[K |████████████████████████████████| 204kB 50.4MB/s \n\u001b[?25hCollecting flake8>=3.7.9\n\u001b[?25l Downloading https://files.pythonhosted.org/packages/fc/80/35a0716e5d5101e643404dabd20f07f5528a21f3ef4032d31a49c913237b/flake8-3.9.2-py2.py3-none-any.whl (73kB)\n\u001b[K |████████████████████████████████| 81kB 11.6MB/s \n\u001b[?25hCollecting Flask-Babel>=1.0.0\n Downloading https://files.pythonhosted.org/packages/ab/3e/02331179ffab8b79e0383606a028b6a60fb1b4419b84935edd43223406a0/Flask_Babel-2.0.0-py3-none-any.whl\nCollecting pre-commit\n\u001b[?25l Downloading https://files.pythonhosted.org/packages/b2/38/3e5475cbd6921cd8208a06c113384a88bb19ce0cda1155f33d4f19183de1/pre_commit-2.13.0-py2.py3-none-any.whl (190kB)\n\u001b[K |████████████████████████████████| 194kB 37.4MB/s \n\u001b[?25hRequirement already satisfied: pyparsing!=2.0.4,!=2.1.2,!=2.1.6,>=2.0.1 in /usr/local/lib/python3.7/dist-packages (from matplotlib->imgaug==0.4.0->paddleocr==2.0.6) (2.4.7)\nRequirement already satisfied: python-dateutil>=2.1 in /usr/local/lib/python3.7/dist-packages (from matplotlib->imgaug==0.4.0->paddleocr==2.0.6) (2.8.1)\nRequirement already satisfied: cycler>=0.10 in /usr/local/lib/python3.7/dist-packages (from matplotlib->imgaug==0.4.0->paddleocr==2.0.6) (0.10.0)\nRequirement already satisfied: kiwisolver>=1.0.1 in /usr/local/lib/python3.7/dist-packages (from matplotlib->imgaug==0.4.0->paddleocr==2.0.6) (1.3.1)\nRequirement already satisfied: decorator<5,>=4.3 in /usr/local/lib/python3.7/dist-packages (from networkx>=2.0->scikit-image==0.17.2->paddleocr==2.0.6) (4.4.2)\nRequirement already satisfied: pytz>=2017.2 in /usr/local/lib/python3.7/dist-packages (from pandas->visualdl->paddleocr==2.0.6) (2018.9)\nRequirement already satisfied: Jinja2<3.0,>=2.10.1 in /usr/local/lib/python3.7/dist-packages (from flask>=1.1.1->visualdl->paddleocr==2.0.6) (2.11.3)\nRequirement already satisfied: click<8.0,>=5.1 in /usr/local/lib/python3.7/dist-packages (from flask>=1.1.1->visualdl->paddleocr==2.0.6) (7.1.2)\nRequirement already satisfied: Werkzeug<2.0,>=0.15 in /usr/local/lib/python3.7/dist-packages (from flask>=1.1.1->visualdl->paddleocr==2.0.6) (1.0.1)\nRequirement already satisfied: itsdangerous<2.0,>=0.24 in /usr/local/lib/python3.7/dist-packages (from flask>=1.1.1->visualdl->paddleocr==2.0.6) (1.1.0)\nRequirement already satisfied: urllib3!=1.25.0,!=1.25.1,<1.26,>=1.21.1 in /usr/local/lib/python3.7/dist-packages (from requests->visualdl->paddleocr==2.0.6) (1.24.3)\nRequirement already satisfied: idna<3,>=2.5 in /usr/local/lib/python3.7/dist-packages (from requests->visualdl->paddleocr==2.0.6) (2.10)\nRequirement already satisfied: certifi>=2017.4.17 in /usr/local/lib/python3.7/dist-packages (from requests->visualdl->paddleocr==2.0.6) (2020.12.5)\nRequirement already satisfied: chardet<4,>=3.0.2 in /usr/local/lib/python3.7/dist-packages (from requests->visualdl->paddleocr==2.0.6) (3.0.4)\nCollecting pycryptodome>=3.8.0\n\u001b[?25l Downloading https://files.pythonhosted.org/packages/ad/16/9627ab0493894a11c68e46000dbcc82f578c8ff06bc2980dcd016aea9bd3/pycryptodome-3.10.1-cp35-abi3-manylinux2010_x86_64.whl (1.9MB)\n\u001b[K |████████████████████████████████| 1.9MB 36.3MB/s \n\u001b[?25hRequirement already satisfied: future>=0.6.0 in /usr/local/lib/python3.7/dist-packages (from bce-python-sdk->visualdl->paddleocr==2.0.6) (0.16.0)\nCollecting mccabe<0.7.0,>=0.6.0\n Downloading https://files.pythonhosted.org/packages/87/89/479dc97e18549e21354893e4ee4ef36db1d237534982482c3681ee6e7b57/mccabe-0.6.1-py2.py3-none-any.whl\nCollecting pycodestyle<2.8.0,>=2.7.0\n\u001b[?25l Downloading https://files.pythonhosted.org/packages/de/cc/227251b1471f129bc35e966bb0fceb005969023926d744139642d847b7ae/pycodestyle-2.7.0-py2.py3-none-any.whl (41kB)\n\u001b[K |████████████████████████████████| 51kB 8.5MB/s \n\u001b[?25hRequirement already satisfied: importlib-metadata; python_version < \"3.8\" in /usr/local/lib/python3.7/dist-packages (from flake8>=3.7.9->visualdl->paddleocr==2.0.6) (4.0.1)\nCollecting pyflakes<2.4.0,>=2.3.0\n\u001b[?25l Downloading https://files.pythonhosted.org/packages/6c/11/2a745612f1d3cbbd9c69ba14b1b43a35a2f5c3c81cd0124508c52c64307f/pyflakes-2.3.1-py2.py3-none-any.whl (68kB)\n\u001b[K |████████████████████████████████| 71kB 11.2MB/s \n\u001b[?25hRequirement already satisfied: Babel>=2.3 in /usr/local/lib/python3.7/dist-packages (from Flask-Babel>=1.0.0->visualdl->paddleocr==2.0.6) (2.9.1)\nCollecting identify>=1.0.0\n\u001b[?25l Downloading https://files.pythonhosted.org/packages/d5/e8/8cda16c3288ebe5cad40cbb84590caf72d1aec1d4fa2e61bed4ee306c532/identify-2.2.7-py2.py3-none-any.whl (98kB)\n\u001b[K |████████████████████████████████| 102kB 13.6MB/s \n\u001b[?25hRequirement already satisfied: toml in /usr/local/lib/python3.7/dist-packages (from pre-commit->visualdl->paddleocr==2.0.6) (0.10.2)\nCollecting pyyaml>=5.1\n\u001b[?25l Downloading https://files.pythonhosted.org/packages/7a/a5/393c087efdc78091afa2af9f1378762f9821c9c1d7a22c5753fb5ac5f97a/PyYAML-5.4.1-cp37-cp37m-manylinux1_x86_64.whl (636kB)\n\u001b[K |████████████████████████████████| 645kB 37.7MB/s \n\u001b[?25hCollecting nodeenv>=0.11.1\n Downloading https://files.pythonhosted.org/packages/54/73/56c89b343befb9c63e8117294d265458f0ff726fa2abcdc6bb5ec5e66a1a/nodeenv-1.6.0-py2.py3-none-any.whl\nCollecting virtualenv>=20.0.8\n\u001b[?25l Downloading https://files.pythonhosted.org/packages/03/08/f819421002e85a71d58368f7bffbe0b1921325e0e8ca7857cb5fb0e1f7c1/virtualenv-20.4.7-py2.py3-none-any.whl (7.2MB)\n\u001b[K |████████████████████████████████| 7.2MB 31.7MB/s \n\u001b[?25hCollecting cfgv>=2.0.0\n Downloading https://files.pythonhosted.org/packages/49/54/83bf9b6ba673bf7d5ebe3846a5f6d3a53925cfd331aa29ec6b5b9c42a850/cfgv-3.3.0-py2.py3-none-any.whl\nRequirement already satisfied: MarkupSafe>=0.23 in /usr/local/lib/python3.7/dist-packages (from Jinja2<3.0,>=2.10.1->flask>=1.1.1->visualdl->paddleocr==2.0.6) (2.0.1)\nRequirement already satisfied: typing-extensions>=3.6.4; python_version < \"3.8\" in /usr/local/lib/python3.7/dist-packages (from importlib-metadata; python_version < \"3.8\"->flake8>=3.7.9->visualdl->paddleocr==2.0.6) (3.7.4.3)\nRequirement already satisfied: zipp>=0.5 in /usr/local/lib/python3.7/dist-packages (from importlib-metadata; python_version < \"3.8\"->flake8>=3.7.9->visualdl->paddleocr==2.0.6) (3.4.1)\nCollecting distlib<1,>=0.3.1\n\u001b[?25l Downloading https://files.pythonhosted.org/packages/87/26/f6a23dd3e578132cf924e0dd5d4e055af0cd4ab43e2a9f10b7568bfb39d9/distlib-0.3.2-py2.py3-none-any.whl (338kB)\n\u001b[K |████████████████████████████████| 348kB 57.8MB/s \n\u001b[?25hRequirement already satisfied: filelock<4,>=3.0.0 in /usr/local/lib/python3.7/dist-packages (from virtualenv>=20.0.8->pre-commit->visualdl->paddleocr==2.0.6) (3.0.12)\nRequirement already satisfied: appdirs<2,>=1.4.3 in /usr/local/lib/python3.7/dist-packages (from virtualenv>=20.0.8->pre-commit->visualdl->paddleocr==2.0.6) (1.4.4)\nBuilding wheels for collected packages: python-Levenshtein\n Building wheel for python-Levenshtein (setup.py) ... \u001b[?25l\u001b[?25hdone\n Created wheel for python-Levenshtein: filename=python_Levenshtein-0.12.2-cp37-cp37m-linux_x86_64.whl size=149819 sha256=7eb69b6bded8ea935307277fdaeb2865b8faff32102c140de4944b2a9e41743d\n Stored in directory: /root/.cache/pip/wheels/b3/26/73/4b48503bac73f01cf18e52cd250947049a7f339e940c5df8fc\nSuccessfully built python-Levenshtein\n\u001b[31mERROR: albumentations 0.1.12 has requirement imgaug<0.2.7,>=0.2.5, but you'll have imgaug 0.4.0 which is incompatible.\u001b[0m\nInstalling collected packages: scikit-image, opencv-python, imgaug, python-Levenshtein, shellcheck-py, pycryptodome, bce-python-sdk, mccabe, pycodestyle, pyflakes, flake8, Flask-Babel, identify, pyyaml, nodeenv, distlib, virtualenv, cfgv, pre-commit, visualdl, pyclipper, paddleocr\n Found existing installation: scikit-image 0.16.2\n Uninstalling scikit-image-0.16.2:\n Successfully uninstalled scikit-image-0.16.2\n Found existing installation: opencv-python 4.1.2.30\n Uninstalling opencv-python-4.1.2.30:\n Successfully uninstalled opencv-python-4.1.2.30\n Found existing installation: imgaug 0.2.9\n Uninstalling imgaug-0.2.9:\n Successfully uninstalled imgaug-0.2.9\n Found existing installation: PyYAML 3.13\n Uninstalling PyYAML-3.13:\n Successfully uninstalled PyYAML-3.13\nSuccessfully installed Flask-Babel-2.0.0 bce-python-sdk-0.8.60 cfgv-3.3.0 distlib-0.3.2 flake8-3.9.2 identify-2.2.7 imgaug-0.4.0 mccabe-0.6.1 nodeenv-1.6.0 opencv-python-4.2.0.32 paddleocr-2.0.6 pre-commit-2.13.0 pyclipper-1.2.1 pycodestyle-2.7.0 pycryptodome-3.10.1 pyflakes-2.3.1 python-Levenshtein-0.12.2 pyyaml-5.4.1 scikit-image-0.17.2 shellcheck-py-0.7.2.1 virtualenv-20.4.7 visualdl-2.2.0\n"
],
[
"# !git clone https://github.com/PaddlePaddle/PaddleOCR",
"_____no_output_____"
],
[
"%cd PaddleOCR/",
"/content/drive/My Drive/Reg_text/PaddleOCR/PaddleOCR\n"
],
[
"!python3 setup.py bdist_wheel",
"running bdist_wheel\nrunning build\nrunning build_py\ncopying paddleocr.py -> build/lib/paddleocr\nrunning egg_info\nwriting paddleocr.egg-info/PKG-INFO\nwriting dependency_links to paddleocr.egg-info/dependency_links.txt\nwriting entry points to paddleocr.egg-info/entry_points.txt\nwriting requirements to paddleocr.egg-info/requires.txt\nwriting top-level names to paddleocr.egg-info/top_level.txt\nadding license file 'LICENSE' (matched pattern 'LICEN[CS]E*')\nreading manifest template 'MANIFEST.in'\nwarning: no files found matching 'LICENSE.txt'\nwriting manifest file 'paddleocr.egg-info/SOURCES.txt'\ncopying paddleocr.egg-info/PKG-INFO -> build/lib/paddleocr/paddleocr.egg-info\ncopying paddleocr.egg-info/SOURCES.txt -> build/lib/paddleocr/paddleocr.egg-info\ncopying paddleocr.egg-info/dependency_links.txt -> build/lib/paddleocr/paddleocr.egg-info\ncopying paddleocr.egg-info/entry_points.txt -> build/lib/paddleocr/paddleocr.egg-info\ncopying paddleocr.egg-info/requires.txt -> build/lib/paddleocr/paddleocr.egg-info\ncopying paddleocr.egg-info/top_level.txt -> build/lib/paddleocr/paddleocr.egg-info\ninstalling to build/bdist.linux-x86_64/wheel\nrunning install\nrunning install_lib\ncreating build/bdist.linux-x86_64/wheel\ncreating build/bdist.linux-x86_64/wheel/paddleocr\ncopying build/lib/paddleocr/.style.yapf -> build/bdist.linux-x86_64/wheel/paddleocr\ncopying build/lib/paddleocr/.pre-commit-config.yaml -> build/bdist.linux-x86_64/wheel/paddleocr\ncopying build/lib/paddleocr/.gitignore -> build/bdist.linux-x86_64/wheel/paddleocr\ncopying build/lib/paddleocr/.clang_format.hook -> build/bdist.linux-x86_64/wheel/paddleocr\ncopying build/lib/paddleocr/MANIFEST.in -> build/bdist.linux-x86_64/wheel/paddleocr\ncopying build/lib/paddleocr/LICENSE -> build/bdist.linux-x86_64/wheel/paddleocr\ncopying build/lib/paddleocr/README.md -> build/bdist.linux-x86_64/wheel/paddleocr\ncopying build/lib/paddleocr/__init__.py -> build/bdist.linux-x86_64/wheel/paddleocr\ncopying build/lib/paddleocr/README_ch.md -> build/bdist.linux-x86_64/wheel/paddleocr\ncopying build/lib/paddleocr/requirements.txt -> build/bdist.linux-x86_64/wheel/paddleocr\ncopying build/lib/paddleocr/train.sh -> build/bdist.linux-x86_64/wheel/paddleocr\ncreating build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel\ncopying build/lib/paddleocr/PPOCRLabel/README_ch.md -> build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel\ncopying build/lib/paddleocr/PPOCRLabel/README.md -> build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel\ncopying build/lib/paddleocr/PPOCRLabel/combobox.py -> build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel\ncopying build/lib/paddleocr/PPOCRLabel/PPOCRLabel.py -> build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel\ncopying build/lib/paddleocr/PPOCRLabel/Makefile -> build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel\ncopying build/lib/paddleocr/PPOCRLabel/__init__.py -> build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel\ncopying build/lib/paddleocr/PPOCRLabel/setup.py -> build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel\ncopying build/lib/paddleocr/PPOCRLabel/resources.qrc -> build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel\ncopying build/lib/paddleocr/PPOCRLabel/setup.cfg -> build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel\ncreating build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel/data\ncopying build/lib/paddleocr/PPOCRLabel/data/paddle.png -> build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel/data\ncopying build/lib/paddleocr/PPOCRLabel/data/predefined_classes.txt -> build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel/data\ncreating build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel/data/gif\ncopying build/lib/paddleocr/PPOCRLabel/data/gif/steps_en.gif -> build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel/data/gif\ncopying build/lib/paddleocr/PPOCRLabel/data/gif/steps.gif -> build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel/data/gif\ncreating build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel/libs\ncopying build/lib/paddleocr/PPOCRLabel/libs/autoDialog.py -> build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel/libs\ncopying build/lib/paddleocr/PPOCRLabel/libs/editinlist.py -> build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel/libs\ncopying build/lib/paddleocr/PPOCRLabel/libs/constants.py -> build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel/libs\ncopying build/lib/paddleocr/PPOCRLabel/libs/__init__.py -> build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel/libs\ncopying build/lib/paddleocr/PPOCRLabel/libs/create_ml_io.py -> build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel/libs\ncopying build/lib/paddleocr/PPOCRLabel/libs/canvas.py -> build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel/libs\ncopying build/lib/paddleocr/PPOCRLabel/libs/colorDialog.py -> build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel/libs\ncopying build/lib/paddleocr/PPOCRLabel/libs/ustr.py -> build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel/libs\ncopying build/lib/paddleocr/PPOCRLabel/libs/utils.py -> build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel/libs\ncopying build/lib/paddleocr/PPOCRLabel/libs/labelDialog.py -> build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel/libs\ncopying build/lib/paddleocr/PPOCRLabel/libs/stringBundle.py -> build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel/libs\ncopying build/lib/paddleocr/PPOCRLabel/libs/zoomWidget.py -> build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel/libs\ncopying build/lib/paddleocr/PPOCRLabel/libs/hashableQListWidgetItem.py -> build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel/libs\ncopying build/lib/paddleocr/PPOCRLabel/libs/shape.py -> build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel/libs\ncopying build/lib/paddleocr/PPOCRLabel/libs/toolBar.py -> build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel/libs\ncopying build/lib/paddleocr/PPOCRLabel/libs/resources.py -> build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel/libs\ncopying build/lib/paddleocr/PPOCRLabel/libs/settings.py -> build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel/libs\ncreating build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel/resources\ncreating build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel/resources/icons\ncopying build/lib/paddleocr/PPOCRLabel/resources/icons/feBlend-icon.png -> build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel/resources/icons\ncopying build/lib/paddleocr/PPOCRLabel/resources/icons/close.png -> build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel/resources/icons\ncopying build/lib/paddleocr/PPOCRLabel/resources/icons/zoom-out.png -> build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel/resources/icons\ncopying build/lib/paddleocr/PPOCRLabel/resources/icons/prev.png -> build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel/resources/icons\ncopying build/lib/paddleocr/PPOCRLabel/resources/icons/reRec.png -> build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel/resources/icons\ncopying build/lib/paddleocr/PPOCRLabel/resources/icons/new.png -> build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel/resources/icons\ncopying build/lib/paddleocr/PPOCRLabel/resources/icons/delete.png -> build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel/resources/icons\ncopying build/lib/paddleocr/PPOCRLabel/resources/icons/app.icns -> build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel/resources/icons\ncopying build/lib/paddleocr/PPOCRLabel/resources/icons/quit.png -> build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel/resources/icons\ncopying build/lib/paddleocr/PPOCRLabel/resources/icons/zoom.png -> build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel/resources/icons\ncopying build/lib/paddleocr/PPOCRLabel/resources/icons/color.png -> build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel/resources/icons\ncopying build/lib/paddleocr/PPOCRLabel/resources/icons/done.svg -> build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel/resources/icons\ncopying build/lib/paddleocr/PPOCRLabel/resources/icons/open.svg -> build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel/resources/icons\ncopying build/lib/paddleocr/PPOCRLabel/resources/icons/cancel.png -> build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel/resources/icons\ncopying build/lib/paddleocr/PPOCRLabel/resources/icons/format_voc.png -> build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel/resources/icons\ncopying build/lib/paddleocr/PPOCRLabel/resources/icons/file.png -> build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel/resources/icons\ncopying build/lib/paddleocr/PPOCRLabel/resources/icons/fit.png -> build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel/resources/icons\ncopying build/lib/paddleocr/PPOCRLabel/resources/icons/save-as.svg -> build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel/resources/icons\ncopying build/lib/paddleocr/PPOCRLabel/resources/icons/app.png -> build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel/resources/icons\ncopying build/lib/paddleocr/PPOCRLabel/resources/icons/save-as.png -> build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel/resources/icons\ncopying build/lib/paddleocr/PPOCRLabel/resources/icons/done.png -> build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel/resources/icons\ncopying build/lib/paddleocr/PPOCRLabel/resources/icons/edit.png -> build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel/resources/icons\ncopying build/lib/paddleocr/PPOCRLabel/resources/icons/copy.png -> build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel/resources/icons\ncopying build/lib/paddleocr/PPOCRLabel/resources/icons/fit-width.png -> build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel/resources/icons\ncopying build/lib/paddleocr/PPOCRLabel/resources/icons/labels.png -> build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel/resources/icons\ncopying build/lib/paddleocr/PPOCRLabel/resources/icons/expert1.png -> build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel/resources/icons\ncopying build/lib/paddleocr/PPOCRLabel/resources/icons/format_yolo.png -> build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel/resources/icons\ncopying build/lib/paddleocr/PPOCRLabel/resources/icons/format_createml.png -> build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel/resources/icons\ncopying build/lib/paddleocr/PPOCRLabel/resources/icons/app.svg -> build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel/resources/icons\ncopying build/lib/paddleocr/PPOCRLabel/resources/icons/open.png -> build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel/resources/icons\ncopying build/lib/paddleocr/PPOCRLabel/resources/icons/next.png -> build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel/resources/icons\ncopying build/lib/paddleocr/PPOCRLabel/resources/icons/undo.png -> build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel/resources/icons\ncopying build/lib/paddleocr/PPOCRLabel/resources/icons/color_line.png -> build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel/resources/icons\ncopying build/lib/paddleocr/PPOCRLabel/resources/icons/save.png -> build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel/resources/icons\ncopying build/lib/paddleocr/PPOCRLabel/resources/icons/Auto.png -> build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel/resources/icons\ncopying build/lib/paddleocr/PPOCRLabel/resources/icons/objects.png -> build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel/resources/icons\ncopying build/lib/paddleocr/PPOCRLabel/resources/icons/save.svg -> build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel/resources/icons\ncopying build/lib/paddleocr/PPOCRLabel/resources/icons/expert2.png -> build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel/resources/icons\ncopying build/lib/paddleocr/PPOCRLabel/resources/icons/resetall.png -> build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel/resources/icons\ncopying build/lib/paddleocr/PPOCRLabel/resources/icons/zoom-in.png -> build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel/resources/icons\ncopying build/lib/paddleocr/PPOCRLabel/resources/icons/help.png -> build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel/resources/icons\ncopying build/lib/paddleocr/PPOCRLabel/resources/icons/undo-cross.png -> build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel/resources/icons\ncopying build/lib/paddleocr/PPOCRLabel/resources/icons/fit-window.png -> build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel/resources/icons\ncopying build/lib/paddleocr/PPOCRLabel/resources/icons/labels.svg -> build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel/resources/icons\ncopying build/lib/paddleocr/PPOCRLabel/resources/icons/verify.png -> build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel/resources/icons\ncopying build/lib/paddleocr/PPOCRLabel/resources/icons/eye.png -> build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel/resources/icons\ncreating build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel/resources/strings\ncopying build/lib/paddleocr/PPOCRLabel/resources/strings/strings.properties -> build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel/resources/strings\ncopying build/lib/paddleocr/PPOCRLabel/resources/strings/strings-zh-CN.properties -> build/bdist.linux-x86_64/wheel/paddleocr/PPOCRLabel/resources/strings\ncreating build/bdist.linux-x86_64/wheel/paddleocr/StyleText\ncopying build/lib/paddleocr/StyleText/README.md -> build/bdist.linux-x86_64/wheel/paddleocr/StyleText\ncopying build/lib/paddleocr/StyleText/README_ch.md -> build/bdist.linux-x86_64/wheel/paddleocr/StyleText\ncopying build/lib/paddleocr/StyleText/__init__.py -> build/bdist.linux-x86_64/wheel/paddleocr/StyleText\ncreating build/bdist.linux-x86_64/wheel/paddleocr/StyleText/arch\ncopying build/lib/paddleocr/StyleText/arch/spectral_norm.py -> build/bdist.linux-x86_64/wheel/paddleocr/StyleText/arch\ncopying build/lib/paddleocr/StyleText/arch/style_text_rec.py -> build/bdist.linux-x86_64/wheel/paddleocr/StyleText/arch\ncopying build/lib/paddleocr/StyleText/arch/__init__.py -> build/bdist.linux-x86_64/wheel/paddleocr/StyleText/arch\ncopying build/lib/paddleocr/StyleText/arch/encoder.py -> build/bdist.linux-x86_64/wheel/paddleocr/StyleText/arch\ncopying build/lib/paddleocr/StyleText/arch/base_module.py -> build/bdist.linux-x86_64/wheel/paddleocr/StyleText/arch\ncopying build/lib/paddleocr/StyleText/arch/decoder.py -> build/bdist.linux-x86_64/wheel/paddleocr/StyleText/arch\ncreating build/bdist.linux-x86_64/wheel/paddleocr/StyleText/configs\ncopying build/lib/paddleocr/StyleText/configs/dataset_config.yml -> build/bdist.linux-x86_64/wheel/paddleocr/StyleText/configs\ncopying build/lib/paddleocr/StyleText/configs/config.yml -> build/bdist.linux-x86_64/wheel/paddleocr/StyleText/configs\ncreating build/bdist.linux-x86_64/wheel/paddleocr/StyleText/doc\ncreating build/bdist.linux-x86_64/wheel/paddleocr/StyleText/doc/images\ncopying build/lib/paddleocr/StyleText/doc/images/7.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/StyleText/doc/images\ncopying build/lib/paddleocr/StyleText/doc/images/8.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/StyleText/doc/images\ncopying build/lib/paddleocr/StyleText/doc/images/9.png -> build/bdist.linux-x86_64/wheel/paddleocr/StyleText/doc/images\ncopying build/lib/paddleocr/StyleText/doc/images/10.png -> build/bdist.linux-x86_64/wheel/paddleocr/StyleText/doc/images\ncopying build/lib/paddleocr/StyleText/doc/images/3.png -> build/bdist.linux-x86_64/wheel/paddleocr/StyleText/doc/images\ncopying build/lib/paddleocr/StyleText/doc/images/2.png -> build/bdist.linux-x86_64/wheel/paddleocr/StyleText/doc/images\ncopying build/lib/paddleocr/StyleText/doc/images/6.png -> build/bdist.linux-x86_64/wheel/paddleocr/StyleText/doc/images\ncopying build/lib/paddleocr/StyleText/doc/images/12.png -> build/bdist.linux-x86_64/wheel/paddleocr/StyleText/doc/images\ncopying build/lib/paddleocr/StyleText/doc/images/5.png -> build/bdist.linux-x86_64/wheel/paddleocr/StyleText/doc/images\ncopying build/lib/paddleocr/StyleText/doc/images/1.png -> build/bdist.linux-x86_64/wheel/paddleocr/StyleText/doc/images\ncopying build/lib/paddleocr/StyleText/doc/images/4.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/StyleText/doc/images\ncopying build/lib/paddleocr/StyleText/doc/images/11.png -> build/bdist.linux-x86_64/wheel/paddleocr/StyleText/doc/images\ncreating build/bdist.linux-x86_64/wheel/paddleocr/StyleText/engine\ncopying build/lib/paddleocr/StyleText/engine/corpus_generators.py -> build/bdist.linux-x86_64/wheel/paddleocr/StyleText/engine\ncopying build/lib/paddleocr/StyleText/engine/style_samplers.py -> build/bdist.linux-x86_64/wheel/paddleocr/StyleText/engine\ncopying build/lib/paddleocr/StyleText/engine/writers.py -> build/bdist.linux-x86_64/wheel/paddleocr/StyleText/engine\ncopying build/lib/paddleocr/StyleText/engine/__init__.py -> build/bdist.linux-x86_64/wheel/paddleocr/StyleText/engine\ncopying build/lib/paddleocr/StyleText/engine/predictors.py -> build/bdist.linux-x86_64/wheel/paddleocr/StyleText/engine\ncopying build/lib/paddleocr/StyleText/engine/synthesisers.py -> build/bdist.linux-x86_64/wheel/paddleocr/StyleText/engine\ncopying build/lib/paddleocr/StyleText/engine/text_drawers.py -> build/bdist.linux-x86_64/wheel/paddleocr/StyleText/engine\ncreating build/bdist.linux-x86_64/wheel/paddleocr/StyleText/examples\ncopying build/lib/paddleocr/StyleText/examples/image_list.txt -> build/bdist.linux-x86_64/wheel/paddleocr/StyleText/examples\ncreating build/bdist.linux-x86_64/wheel/paddleocr/StyleText/examples/corpus\ncopying build/lib/paddleocr/StyleText/examples/corpus/example.txt -> build/bdist.linux-x86_64/wheel/paddleocr/StyleText/examples/corpus\ncreating build/bdist.linux-x86_64/wheel/paddleocr/StyleText/examples/style_images\ncopying build/lib/paddleocr/StyleText/examples/style_images/1.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/StyleText/examples/style_images\ncopying build/lib/paddleocr/StyleText/examples/style_images/2.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/StyleText/examples/style_images\ncreating build/bdist.linux-x86_64/wheel/paddleocr/StyleText/fonts\ncopying build/lib/paddleocr/StyleText/fonts/ch_standard.ttf -> build/bdist.linux-x86_64/wheel/paddleocr/StyleText/fonts\ncopying build/lib/paddleocr/StyleText/fonts/en_standard.ttf -> build/bdist.linux-x86_64/wheel/paddleocr/StyleText/fonts\ncopying build/lib/paddleocr/StyleText/fonts/ko_standard.ttf -> build/bdist.linux-x86_64/wheel/paddleocr/StyleText/fonts\ncreating build/bdist.linux-x86_64/wheel/paddleocr/StyleText/tools\ncopying build/lib/paddleocr/StyleText/tools/synth_image.py -> build/bdist.linux-x86_64/wheel/paddleocr/StyleText/tools\ncopying build/lib/paddleocr/StyleText/tools/__init__.py -> build/bdist.linux-x86_64/wheel/paddleocr/StyleText/tools\ncopying build/lib/paddleocr/StyleText/tools/synth_dataset.py -> build/bdist.linux-x86_64/wheel/paddleocr/StyleText/tools\ncreating build/bdist.linux-x86_64/wheel/paddleocr/StyleText/utils\ncopying build/lib/paddleocr/StyleText/utils/config.py -> build/bdist.linux-x86_64/wheel/paddleocr/StyleText/utils\ncopying build/lib/paddleocr/StyleText/utils/math_functions.py -> build/bdist.linux-x86_64/wheel/paddleocr/StyleText/utils\ncopying build/lib/paddleocr/StyleText/utils/__init__.py -> build/bdist.linux-x86_64/wheel/paddleocr/StyleText/utils\ncopying build/lib/paddleocr/StyleText/utils/logging.py -> build/bdist.linux-x86_64/wheel/paddleocr/StyleText/utils\ncopying build/lib/paddleocr/StyleText/utils/sys_funcs.py -> build/bdist.linux-x86_64/wheel/paddleocr/StyleText/utils\ncopying build/lib/paddleocr/StyleText/utils/load_params.py -> build/bdist.linux-x86_64/wheel/paddleocr/StyleText/utils\ncreating build/bdist.linux-x86_64/wheel/paddleocr/configs\ncreating build/bdist.linux-x86_64/wheel/paddleocr/configs/cls\ncopying build/lib/paddleocr/configs/cls/cls_mv3.yml -> build/bdist.linux-x86_64/wheel/paddleocr/configs/cls\ncreating build/bdist.linux-x86_64/wheel/paddleocr/configs/det\ncopying build/lib/paddleocr/configs/det/det_r50_vd_sast_totaltext.yml -> build/bdist.linux-x86_64/wheel/paddleocr/configs/det\ncopying build/lib/paddleocr/configs/det/det_r50_vd_db.yml -> build/bdist.linux-x86_64/wheel/paddleocr/configs/det\ncopying build/lib/paddleocr/configs/det/det_mv3_east.yml -> build/bdist.linux-x86_64/wheel/paddleocr/configs/det\ncopying build/lib/paddleocr/configs/det/det_mv3_db.yml -> build/bdist.linux-x86_64/wheel/paddleocr/configs/det\ncopying build/lib/paddleocr/configs/det/det_r50_vd_east.yml -> build/bdist.linux-x86_64/wheel/paddleocr/configs/det\ncopying build/lib/paddleocr/configs/det/det_r50_vd_sast_icdar15.yml -> build/bdist.linux-x86_64/wheel/paddleocr/configs/det\ncreating build/bdist.linux-x86_64/wheel/paddleocr/configs/det/ch_ppocr_v2.0\ncopying build/lib/paddleocr/configs/det/ch_ppocr_v2.0/ch_det_mv3_db_v2.0.yml -> build/bdist.linux-x86_64/wheel/paddleocr/configs/det/ch_ppocr_v2.0\ncopying build/lib/paddleocr/configs/det/ch_ppocr_v2.0/ch_det_res18_db_v2.0.yml -> build/bdist.linux-x86_64/wheel/paddleocr/configs/det/ch_ppocr_v2.0\ncreating build/bdist.linux-x86_64/wheel/paddleocr/configs/e2e\ncopying build/lib/paddleocr/configs/e2e/e2e_r50_vd_pg.yml -> build/bdist.linux-x86_64/wheel/paddleocr/configs/e2e\ncreating build/bdist.linux-x86_64/wheel/paddleocr/configs/rec\ncopying build/lib/paddleocr/configs/rec/rec_icdar15_train.yml -> build/bdist.linux-x86_64/wheel/paddleocr/configs/rec\ncopying build/lib/paddleocr/configs/rec/rec_r34_vd_tps_bilstm_att.yml -> build/bdist.linux-x86_64/wheel/paddleocr/configs/rec\ncopying build/lib/paddleocr/configs/rec/rec_r34_vd_tps_bilstm_ctc.yml -> build/bdist.linux-x86_64/wheel/paddleocr/configs/rec\ncopying build/lib/paddleocr/configs/rec/rec_mv3_none_none_ctc.yml -> build/bdist.linux-x86_64/wheel/paddleocr/configs/rec\ncopying build/lib/paddleocr/configs/rec/rec_mv3_none_bilstm_ctc.yml -> build/bdist.linux-x86_64/wheel/paddleocr/configs/rec\ncopying build/lib/paddleocr/configs/rec/rec_r34_vd_none_none_ctc.yml -> build/bdist.linux-x86_64/wheel/paddleocr/configs/rec\ncopying build/lib/paddleocr/configs/rec/rec_mv3_tps_bilstm_ctc.yml -> build/bdist.linux-x86_64/wheel/paddleocr/configs/rec\ncopying build/lib/paddleocr/configs/rec/rec_r50_fpn_srn.yml -> build/bdist.linux-x86_64/wheel/paddleocr/configs/rec\ncopying build/lib/paddleocr/configs/rec/rec_r34_vd_none_bilstm_ctc.yml -> build/bdist.linux-x86_64/wheel/paddleocr/configs/rec\ncopying build/lib/paddleocr/configs/rec/rec_mv3_tps_bilstm_att.yml -> build/bdist.linux-x86_64/wheel/paddleocr/configs/rec\ncreating build/bdist.linux-x86_64/wheel/paddleocr/configs/rec/ch_ppocr_v2.0\ncopying build/lib/paddleocr/configs/rec/ch_ppocr_v2.0/rec_chinese_common_train_v2.0.yml -> build/bdist.linux-x86_64/wheel/paddleocr/configs/rec/ch_ppocr_v2.0\ncopying build/lib/paddleocr/configs/rec/ch_ppocr_v2.0/rec_chinese_lite_train_v2.0.yml -> build/bdist.linux-x86_64/wheel/paddleocr/configs/rec/ch_ppocr_v2.0\ncreating build/bdist.linux-x86_64/wheel/paddleocr/configs/rec/multi_language\ncopying build/lib/paddleocr/configs/rec/multi_language/rec_en_number_lite_train.yml -> build/bdist.linux-x86_64/wheel/paddleocr/configs/rec/multi_language\ncopying build/lib/paddleocr/configs/rec/multi_language/rec_cyrillic_lite_train.yml -> build/bdist.linux-x86_64/wheel/paddleocr/configs/rec/multi_language\ncopying build/lib/paddleocr/configs/rec/multi_language/rec_devanagari_lite_train.yml -> build/bdist.linux-x86_64/wheel/paddleocr/configs/rec/multi_language\ncopying build/lib/paddleocr/configs/rec/multi_language/generate_multi_language_configs.py -> build/bdist.linux-x86_64/wheel/paddleocr/configs/rec/multi_language\ncopying build/lib/paddleocr/configs/rec/multi_language/rec_arabic_lite_train.yml -> build/bdist.linux-x86_64/wheel/paddleocr/configs/rec/multi_language\ncopying build/lib/paddleocr/configs/rec/multi_language/rec_french_lite_train.yml -> build/bdist.linux-x86_64/wheel/paddleocr/configs/rec/multi_language\ncopying build/lib/paddleocr/configs/rec/multi_language/rec_latin_lite_train.yml -> build/bdist.linux-x86_64/wheel/paddleocr/configs/rec/multi_language\ncopying build/lib/paddleocr/configs/rec/multi_language/rec_multi_language_lite_train.yml -> build/bdist.linux-x86_64/wheel/paddleocr/configs/rec/multi_language\ncopying build/lib/paddleocr/configs/rec/multi_language/rec_german_lite_train.yml -> build/bdist.linux-x86_64/wheel/paddleocr/configs/rec/multi_language\ncopying build/lib/paddleocr/configs/rec/multi_language/rec_korean_lite_train.yml -> build/bdist.linux-x86_64/wheel/paddleocr/configs/rec/multi_language\ncopying build/lib/paddleocr/configs/rec/multi_language/rec_japan_lite_train.yml -> build/bdist.linux-x86_64/wheel/paddleocr/configs/rec/multi_language\ncreating build/bdist.linux-x86_64/wheel/paddleocr/deploy\ncreating build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo\ncopying build/lib/paddleocr/deploy/android_demo/README.md -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo\ncopying build/lib/paddleocr/deploy/android_demo/.gitignore -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo\ncopying build/lib/paddleocr/deploy/android_demo/gradle.properties -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo\ncopying build/lib/paddleocr/deploy/android_demo/build.gradle -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo\ncopying build/lib/paddleocr/deploy/android_demo/gradlew.bat -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo\ncopying build/lib/paddleocr/deploy/android_demo/settings.gradle -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo\ncopying build/lib/paddleocr/deploy/android_demo/gradlew -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo\ncreating build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app\ncopying build/lib/paddleocr/deploy/android_demo/app/.gitignore -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app\ncopying build/lib/paddleocr/deploy/android_demo/app/proguard-rules.pro -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app\ncopying build/lib/paddleocr/deploy/android_demo/app/build.gradle -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app\ncreating build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src\ncreating build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/androidTest\ncreating build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/androidTest/java\ncreating build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/androidTest/java/com\ncreating build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/androidTest/java/com/baidu\ncreating build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/androidTest/java/com/baidu/paddle\ncreating build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/androidTest/java/com/baidu/paddle/lite\ncreating build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/androidTest/java/com/baidu/paddle/lite/demo\ncreating build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/androidTest/java/com/baidu/paddle/lite/demo/ocr\ncopying build/lib/paddleocr/deploy/android_demo/app/src/androidTest/java/com/baidu/paddle/lite/demo/ocr/ExampleInstrumentedTest.java -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/androidTest/java/com/baidu/paddle/lite/demo/ocr\ncreating build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main\ncopying build/lib/paddleocr/deploy/android_demo/app/src/main/AndroidManifest.xml -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main\ncreating build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main/assets\ncreating build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main/assets/images\ncopying build/lib/paddleocr/deploy/android_demo/app/src/main/assets/images/180.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main/assets/images\ncopying build/lib/paddleocr/deploy/android_demo/app/src/main/assets/images/0.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main/assets/images\ncopying build/lib/paddleocr/deploy/android_demo/app/src/main/assets/images/90.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main/assets/images\ncopying build/lib/paddleocr/deploy/android_demo/app/src/main/assets/images/270.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main/assets/images\ncreating build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main/cpp\ncopying build/lib/paddleocr/deploy/android_demo/app/src/main/cpp/ocr_clipper.cpp -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main/cpp\ncopying build/lib/paddleocr/deploy/android_demo/app/src/main/cpp/native.cpp -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main/cpp\ncopying build/lib/paddleocr/deploy/android_demo/app/src/main/cpp/ocr_cls_process.h -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main/cpp\ncopying build/lib/paddleocr/deploy/android_demo/app/src/main/cpp/CMakeLists.txt -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main/cpp\ncopying build/lib/paddleocr/deploy/android_demo/app/src/main/cpp/native.h -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main/cpp\ncopying build/lib/paddleocr/deploy/android_demo/app/src/main/cpp/ocr_clipper.hpp -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main/cpp\ncopying build/lib/paddleocr/deploy/android_demo/app/src/main/cpp/ocr_cls_process.cpp -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main/cpp\ncopying build/lib/paddleocr/deploy/android_demo/app/src/main/cpp/common.h -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main/cpp\ncopying build/lib/paddleocr/deploy/android_demo/app/src/main/cpp/ocr_crnn_process.cpp -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main/cpp\ncopying build/lib/paddleocr/deploy/android_demo/app/src/main/cpp/predictor_input.h -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main/cpp\ncopying build/lib/paddleocr/deploy/android_demo/app/src/main/cpp/ocr_crnn_process.h -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main/cpp\ncopying build/lib/paddleocr/deploy/android_demo/app/src/main/cpp/ocr_ppredictor.cpp -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main/cpp\ncopying build/lib/paddleocr/deploy/android_demo/app/src/main/cpp/ppredictor.cpp -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main/cpp\ncopying build/lib/paddleocr/deploy/android_demo/app/src/main/cpp/preprocess.cpp -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main/cpp\ncopying build/lib/paddleocr/deploy/android_demo/app/src/main/cpp/ppredictor.h -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main/cpp\ncopying build/lib/paddleocr/deploy/android_demo/app/src/main/cpp/predictor_output.h -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main/cpp\ncopying build/lib/paddleocr/deploy/android_demo/app/src/main/cpp/predictor_input.cpp -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main/cpp\ncopying build/lib/paddleocr/deploy/android_demo/app/src/main/cpp/ocr_db_post_process.cpp -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main/cpp\ncopying build/lib/paddleocr/deploy/android_demo/app/src/main/cpp/preprocess.h -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main/cpp\ncopying build/lib/paddleocr/deploy/android_demo/app/src/main/cpp/ocr_ppredictor.h -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main/cpp\ncopying build/lib/paddleocr/deploy/android_demo/app/src/main/cpp/predictor_output.cpp -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main/cpp\ncopying build/lib/paddleocr/deploy/android_demo/app/src/main/cpp/ocr_db_post_process.h -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main/cpp\ncreating build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main/java\ncreating build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main/java/com\ncreating build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main/java/com/baidu\ncreating build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main/java/com/baidu/paddle\ncreating build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main/java/com/baidu/paddle/lite\ncreating build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main/java/com/baidu/paddle/lite/demo\ncreating build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main/java/com/baidu/paddle/lite/demo/ocr\ncopying build/lib/paddleocr/deploy/android_demo/app/src/main/java/com/baidu/paddle/lite/demo/ocr/MainActivity.java -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main/java/com/baidu/paddle/lite/demo/ocr\ncopying build/lib/paddleocr/deploy/android_demo/app/src/main/java/com/baidu/paddle/lite/demo/ocr/OCRPredictorNative.java -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main/java/com/baidu/paddle/lite/demo/ocr\ncopying build/lib/paddleocr/deploy/android_demo/app/src/main/java/com/baidu/paddle/lite/demo/ocr/MiniActivity.java -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main/java/com/baidu/paddle/lite/demo/ocr\ncopying build/lib/paddleocr/deploy/android_demo/app/src/main/java/com/baidu/paddle/lite/demo/ocr/AppCompatPreferenceActivity.java -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main/java/com/baidu/paddle/lite/demo/ocr\ncopying build/lib/paddleocr/deploy/android_demo/app/src/main/java/com/baidu/paddle/lite/demo/ocr/Predictor.java -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main/java/com/baidu/paddle/lite/demo/ocr\ncopying build/lib/paddleocr/deploy/android_demo/app/src/main/java/com/baidu/paddle/lite/demo/ocr/SettingsActivity.java -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main/java/com/baidu/paddle/lite/demo/ocr\ncopying build/lib/paddleocr/deploy/android_demo/app/src/main/java/com/baidu/paddle/lite/demo/ocr/Utils.java -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main/java/com/baidu/paddle/lite/demo/ocr\ncopying build/lib/paddleocr/deploy/android_demo/app/src/main/java/com/baidu/paddle/lite/demo/ocr/OcrResultModel.java -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main/java/com/baidu/paddle/lite/demo/ocr\ncreating build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main/res\ncreating build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main/res/drawable\ncopying build/lib/paddleocr/deploy/android_demo/app/src/main/res/drawable/ic_launcher_background.xml -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main/res/drawable\ncreating build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main/res/drawable-v24\ncopying build/lib/paddleocr/deploy/android_demo/app/src/main/res/drawable-v24/ic_launcher_foreground.xml -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main/res/drawable-v24\ncreating build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main/res/layout\ncopying build/lib/paddleocr/deploy/android_demo/app/src/main/res/layout/activity_mini.xml -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main/res/layout\ncopying build/lib/paddleocr/deploy/android_demo/app/src/main/res/layout/activity_main.xml -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main/res/layout\ncreating build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main/res/menu\ncopying build/lib/paddleocr/deploy/android_demo/app/src/main/res/menu/menu_action_options.xml -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main/res/menu\ncreating build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main/res/mipmap-anydpi-v26\ncopying build/lib/paddleocr/deploy/android_demo/app/src/main/res/mipmap-anydpi-v26/ic_launcher_round.xml -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main/res/mipmap-anydpi-v26\ncopying build/lib/paddleocr/deploy/android_demo/app/src/main/res/mipmap-anydpi-v26/ic_launcher.xml -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main/res/mipmap-anydpi-v26\ncreating build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main/res/mipmap-hdpi\ncopying build/lib/paddleocr/deploy/android_demo/app/src/main/res/mipmap-hdpi/ic_launcher.png -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main/res/mipmap-hdpi\ncopying build/lib/paddleocr/deploy/android_demo/app/src/main/res/mipmap-hdpi/ic_launcher_round.png -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main/res/mipmap-hdpi\ncreating build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main/res/mipmap-mdpi\ncopying build/lib/paddleocr/deploy/android_demo/app/src/main/res/mipmap-mdpi/ic_launcher_round.png -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main/res/mipmap-mdpi\ncopying build/lib/paddleocr/deploy/android_demo/app/src/main/res/mipmap-mdpi/ic_launcher.png -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main/res/mipmap-mdpi\ncreating build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main/res/mipmap-xhdpi\ncopying build/lib/paddleocr/deploy/android_demo/app/src/main/res/mipmap-xhdpi/ic_launcher_round.png -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main/res/mipmap-xhdpi\ncopying build/lib/paddleocr/deploy/android_demo/app/src/main/res/mipmap-xhdpi/ic_launcher.png -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main/res/mipmap-xhdpi\ncreating build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main/res/mipmap-xxhdpi\ncopying build/lib/paddleocr/deploy/android_demo/app/src/main/res/mipmap-xxhdpi/ic_launcher.png -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main/res/mipmap-xxhdpi\ncopying build/lib/paddleocr/deploy/android_demo/app/src/main/res/mipmap-xxhdpi/ic_launcher_round.png -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main/res/mipmap-xxhdpi\ncreating build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main/res/mipmap-xxxhdpi\ncopying build/lib/paddleocr/deploy/android_demo/app/src/main/res/mipmap-xxxhdpi/ic_launcher_round.png -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main/res/mipmap-xxxhdpi\ncopying build/lib/paddleocr/deploy/android_demo/app/src/main/res/mipmap-xxxhdpi/ic_launcher.png -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main/res/mipmap-xxxhdpi\ncreating build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main/res/values\ncopying build/lib/paddleocr/deploy/android_demo/app/src/main/res/values/colors.xml -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main/res/values\ncopying build/lib/paddleocr/deploy/android_demo/app/src/main/res/values/styles.xml -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main/res/values\ncopying build/lib/paddleocr/deploy/android_demo/app/src/main/res/values/arrays.xml -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main/res/values\ncopying build/lib/paddleocr/deploy/android_demo/app/src/main/res/values/strings.xml -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main/res/values\ncreating build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main/res/xml\ncopying build/lib/paddleocr/deploy/android_demo/app/src/main/res/xml/file_paths.xml -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main/res/xml\ncopying build/lib/paddleocr/deploy/android_demo/app/src/main/res/xml/settings.xml -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/main/res/xml\ncreating build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/test\ncreating build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/test/java\ncreating build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/test/java/com\ncreating build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/test/java/com/baidu\ncreating build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/test/java/com/baidu/paddle\ncreating build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/test/java/com/baidu/paddle/lite\ncreating build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/test/java/com/baidu/paddle/lite/demo\ncreating build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/test/java/com/baidu/paddle/lite/demo/ocr\ncopying build/lib/paddleocr/deploy/android_demo/app/src/test/java/com/baidu/paddle/lite/demo/ocr/ExampleUnitTest.java -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/app/src/test/java/com/baidu/paddle/lite/demo/ocr\ncreating build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/gradle\ncreating build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/gradle/wrapper\ncopying build/lib/paddleocr/deploy/android_demo/gradle/wrapper/gradle-wrapper.jar -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/gradle/wrapper\ncopying build/lib/paddleocr/deploy/android_demo/gradle/wrapper/gradle-wrapper.properties -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/android_demo/gradle/wrapper\ncreating build/bdist.linux-x86_64/wheel/paddleocr/deploy/cpp_infer\ncopying build/lib/paddleocr/deploy/cpp_infer/readme.md -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/cpp_infer\ncopying build/lib/paddleocr/deploy/cpp_infer/CMakeLists.txt -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/cpp_infer\ncopying build/lib/paddleocr/deploy/cpp_infer/readme_en.md -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/cpp_infer\ncreating build/bdist.linux-x86_64/wheel/paddleocr/deploy/cpp_infer/docs\ncopying build/lib/paddleocr/deploy/cpp_infer/docs/windows_vs2019_build.md -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/cpp_infer/docs\ncreating build/bdist.linux-x86_64/wheel/paddleocr/deploy/cpp_infer/imgs\ncopying build/lib/paddleocr/deploy/cpp_infer/imgs/cpp_infer_pred_12.png -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/cpp_infer/imgs\ncreating build/bdist.linux-x86_64/wheel/paddleocr/deploy/cpp_infer/include\ncopying build/lib/paddleocr/deploy/cpp_infer/include/ocr_cls.h -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/cpp_infer/include\ncopying build/lib/paddleocr/deploy/cpp_infer/include/ocr_det.h -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/cpp_infer/include\ncopying build/lib/paddleocr/deploy/cpp_infer/include/config.h -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/cpp_infer/include\ncopying build/lib/paddleocr/deploy/cpp_infer/include/utility.h -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/cpp_infer/include\ncopying build/lib/paddleocr/deploy/cpp_infer/include/ocr_rec.h -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/cpp_infer/include\ncopying build/lib/paddleocr/deploy/cpp_infer/include/preprocess_op.h -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/cpp_infer/include\ncopying build/lib/paddleocr/deploy/cpp_infer/include/postprocess_op.h -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/cpp_infer/include\ncopying build/lib/paddleocr/deploy/cpp_infer/include/clipper.h -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/cpp_infer/include\ncreating build/bdist.linux-x86_64/wheel/paddleocr/deploy/cpp_infer/src\ncopying build/lib/paddleocr/deploy/cpp_infer/src/clipper.cpp -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/cpp_infer/src\ncopying build/lib/paddleocr/deploy/cpp_infer/src/ocr_cls.cpp -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/cpp_infer/src\ncopying build/lib/paddleocr/deploy/cpp_infer/src/preprocess_op.cpp -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/cpp_infer/src\ncopying build/lib/paddleocr/deploy/cpp_infer/src/config.cpp -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/cpp_infer/src\ncopying build/lib/paddleocr/deploy/cpp_infer/src/utility.cpp -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/cpp_infer/src\ncopying build/lib/paddleocr/deploy/cpp_infer/src/ocr_det.cpp -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/cpp_infer/src\ncopying build/lib/paddleocr/deploy/cpp_infer/src/main.cpp -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/cpp_infer/src\ncopying build/lib/paddleocr/deploy/cpp_infer/src/postprocess_op.cpp -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/cpp_infer/src\ncopying build/lib/paddleocr/deploy/cpp_infer/src/ocr_rec.cpp -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/cpp_infer/src\ncreating build/bdist.linux-x86_64/wheel/paddleocr/deploy/cpp_infer/tools\ncopying build/lib/paddleocr/deploy/cpp_infer/tools/config.txt -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/cpp_infer/tools\ncopying build/lib/paddleocr/deploy/cpp_infer/tools/build.sh -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/cpp_infer/tools\ncopying build/lib/paddleocr/deploy/cpp_infer/tools/run.sh -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/cpp_infer/tools\ncreating build/bdist.linux-x86_64/wheel/paddleocr/deploy/docker\ncreating build/bdist.linux-x86_64/wheel/paddleocr/deploy/docker/hubserving\ncopying build/lib/paddleocr/deploy/docker/hubserving/README_cn.md -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/docker/hubserving\ncopying build/lib/paddleocr/deploy/docker/hubserving/README.md -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/docker/hubserving\ncopying build/lib/paddleocr/deploy/docker/hubserving/sample_request.txt -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/docker/hubserving\ncreating build/bdist.linux-x86_64/wheel/paddleocr/deploy/docker/hubserving/cpu\ncopying build/lib/paddleocr/deploy/docker/hubserving/cpu/Dockerfile -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/docker/hubserving/cpu\ncreating build/bdist.linux-x86_64/wheel/paddleocr/deploy/docker/hubserving/gpu\ncopying build/lib/paddleocr/deploy/docker/hubserving/gpu/Dockerfile -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/docker/hubserving/gpu\ncreating build/bdist.linux-x86_64/wheel/paddleocr/deploy/hubserving\ncopying build/lib/paddleocr/deploy/hubserving/readme_en.md -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/hubserving\ncopying build/lib/paddleocr/deploy/hubserving/readme.md -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/hubserving\ncreating build/bdist.linux-x86_64/wheel/paddleocr/deploy/hubserving/ocr_cls\ncopying build/lib/paddleocr/deploy/hubserving/ocr_cls/module.py -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/hubserving/ocr_cls\ncopying build/lib/paddleocr/deploy/hubserving/ocr_cls/__init__.py -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/hubserving/ocr_cls\ncopying build/lib/paddleocr/deploy/hubserving/ocr_cls/params.py -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/hubserving/ocr_cls\ncopying build/lib/paddleocr/deploy/hubserving/ocr_cls/config.json -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/hubserving/ocr_cls\ncreating build/bdist.linux-x86_64/wheel/paddleocr/deploy/hubserving/ocr_det\ncopying build/lib/paddleocr/deploy/hubserving/ocr_det/params.py -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/hubserving/ocr_det\ncopying build/lib/paddleocr/deploy/hubserving/ocr_det/module.py -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/hubserving/ocr_det\ncopying build/lib/paddleocr/deploy/hubserving/ocr_det/config.json -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/hubserving/ocr_det\ncopying build/lib/paddleocr/deploy/hubserving/ocr_det/__init__.py -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/hubserving/ocr_det\ncreating build/bdist.linux-x86_64/wheel/paddleocr/deploy/hubserving/ocr_rec\ncopying build/lib/paddleocr/deploy/hubserving/ocr_rec/params.py -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/hubserving/ocr_rec\ncopying build/lib/paddleocr/deploy/hubserving/ocr_rec/config.json -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/hubserving/ocr_rec\ncopying build/lib/paddleocr/deploy/hubserving/ocr_rec/module.py -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/hubserving/ocr_rec\ncopying build/lib/paddleocr/deploy/hubserving/ocr_rec/__init__.py -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/hubserving/ocr_rec\ncreating build/bdist.linux-x86_64/wheel/paddleocr/deploy/hubserving/ocr_system\ncopying build/lib/paddleocr/deploy/hubserving/ocr_system/__init__.py -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/hubserving/ocr_system\ncopying build/lib/paddleocr/deploy/hubserving/ocr_system/module.py -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/hubserving/ocr_system\ncopying build/lib/paddleocr/deploy/hubserving/ocr_system/config.json -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/hubserving/ocr_system\ncopying build/lib/paddleocr/deploy/hubserving/ocr_system/params.py -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/hubserving/ocr_system\ncreating build/bdist.linux-x86_64/wheel/paddleocr/deploy/lite\ncopying build/lib/paddleocr/deploy/lite/ocr_db_crnn.cc -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/lite\ncopying build/lib/paddleocr/deploy/lite/config.txt -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/lite\ncopying build/lib/paddleocr/deploy/lite/crnn_process.h -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/lite\ncopying build/lib/paddleocr/deploy/lite/readme_en.md -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/lite\ncopying build/lib/paddleocr/deploy/lite/crnn_process.cc -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/lite\ncopying build/lib/paddleocr/deploy/lite/cls_process.cc -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/lite\ncopying build/lib/paddleocr/deploy/lite/db_post_process.h -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/lite\ncopying build/lib/paddleocr/deploy/lite/cls_process.h -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/lite\ncopying build/lib/paddleocr/deploy/lite/readme.md -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/lite\ncopying build/lib/paddleocr/deploy/lite/prepare.sh -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/lite\ncopying build/lib/paddleocr/deploy/lite/Makefile -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/lite\ncopying build/lib/paddleocr/deploy/lite/db_post_process.cc -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/lite\ncreating build/bdist.linux-x86_64/wheel/paddleocr/deploy/lite/imgs\ncopying build/lib/paddleocr/deploy/lite/imgs/lite_demo.png -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/lite/imgs\ncreating build/bdist.linux-x86_64/wheel/paddleocr/deploy/pdserving\ncopying build/lib/paddleocr/deploy/pdserving/pipeline_rpc_client.py -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/pdserving\ncopying build/lib/paddleocr/deploy/pdserving/__init__.py -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/pdserving\ncopying build/lib/paddleocr/deploy/pdserving/web_service.py -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/pdserving\ncopying build/lib/paddleocr/deploy/pdserving/pipeline_http_client.py -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/pdserving\ncopying build/lib/paddleocr/deploy/pdserving/README.md -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/pdserving\ncopying build/lib/paddleocr/deploy/pdserving/config.yml -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/pdserving\ncopying build/lib/paddleocr/deploy/pdserving/ocr_reader.py -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/pdserving\ncopying build/lib/paddleocr/deploy/pdserving/README_CN.md -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/pdserving\ncreating build/bdist.linux-x86_64/wheel/paddleocr/deploy/pdserving/imgs\ncopying build/lib/paddleocr/deploy/pdserving/imgs/start_server.png -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/pdserving/imgs\ncopying build/lib/paddleocr/deploy/pdserving/imgs/demo.png -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/pdserving/imgs\ncopying build/lib/paddleocr/deploy/pdserving/imgs/results.png -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/pdserving/imgs\ncopying build/lib/paddleocr/deploy/pdserving/imgs/cpp_infer_pred_12.png -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/pdserving/imgs\ncreating build/bdist.linux-x86_64/wheel/paddleocr/deploy/pdserving/win\ncopying build/lib/paddleocr/deploy/pdserving/win/ocr_web_server.py -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/pdserving/win\ncopying build/lib/paddleocr/deploy/pdserving/win/__init__.py -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/pdserving/win\ncopying build/lib/paddleocr/deploy/pdserving/win/ocr_web_client.py -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/pdserving/win\ncreating build/bdist.linux-x86_64/wheel/paddleocr/deploy/slim\ncreating build/bdist.linux-x86_64/wheel/paddleocr/deploy/slim/prune\ncopying build/lib/paddleocr/deploy/slim/prune/sensitivity_anal.py -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/slim/prune\ncopying build/lib/paddleocr/deploy/slim/prune/README.md -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/slim/prune\ncopying build/lib/paddleocr/deploy/slim/prune/README_en.md -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/slim/prune\ncopying build/lib/paddleocr/deploy/slim/prune/export_prune_model.py -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/slim/prune\ncreating build/bdist.linux-x86_64/wheel/paddleocr/deploy/slim/quantization\ncopying build/lib/paddleocr/deploy/slim/quantization/README_en.md -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/slim/quantization\ncopying build/lib/paddleocr/deploy/slim/quantization/export_model.py -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/slim/quantization\ncopying build/lib/paddleocr/deploy/slim/quantization/quant.py -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/slim/quantization\ncopying build/lib/paddleocr/deploy/slim/quantization/README.md -> build/bdist.linux-x86_64/wheel/paddleocr/deploy/slim/quantization\ncreating build/bdist.linux-x86_64/wheel/paddleocr/doc\ncopying build/lib/paddleocr/doc/PPOCR.pdf -> build/bdist.linux-x86_64/wheel/paddleocr/doc\ncopying build/lib/paddleocr/doc/ppocr_framework.png -> build/bdist.linux-x86_64/wheel/paddleocr/doc\ncopying build/lib/paddleocr/doc/ocr-android-easyedge.png -> build/bdist.linux-x86_64/wheel/paddleocr/doc\ncopying build/lib/paddleocr/doc/joinus.PNG -> build/bdist.linux-x86_64/wheel/paddleocr/doc\ncopying build/lib/paddleocr/doc/pgnet_framework.png -> build/bdist.linux-x86_64/wheel/paddleocr/doc\ncreating build/bdist.linux-x86_64/wheel/paddleocr/doc/datasets\ncopying build/lib/paddleocr/doc/datasets/captcha_demo.png -> build/bdist.linux-x86_64/wheel/paddleocr/doc/datasets\ncopying build/lib/paddleocr/doc/datasets/VoTT.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/datasets\ncopying build/lib/paddleocr/doc/datasets/LSVT_2.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/datasets\ncopying build/lib/paddleocr/doc/datasets/CASIA_0.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/datasets\ncopying build/lib/paddleocr/doc/datasets/ch_doc2.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/datasets\ncopying build/lib/paddleocr/doc/datasets/ArT.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/datasets\ncopying build/lib/paddleocr/doc/datasets/ch_doc3.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/datasets\ncopying build/lib/paddleocr/doc/datasets/ch_doc1.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/datasets\ncopying build/lib/paddleocr/doc/datasets/LSVT_1.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/datasets\ncopying build/lib/paddleocr/doc/datasets/ccpd_demo.png -> build/bdist.linux-x86_64/wheel/paddleocr/doc/datasets\ncopying build/lib/paddleocr/doc/datasets/rctw.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/datasets\ncopying build/lib/paddleocr/doc/datasets/ch_street_rec_2.png -> build/bdist.linux-x86_64/wheel/paddleocr/doc/datasets\ncopying build/lib/paddleocr/doc/datasets/nist_demo.png -> build/bdist.linux-x86_64/wheel/paddleocr/doc/datasets\ncopying build/lib/paddleocr/doc/datasets/labelme.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/datasets\ncopying build/lib/paddleocr/doc/datasets/cmb_demo.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/datasets\ncopying build/lib/paddleocr/doc/datasets/doc.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/datasets\ncopying build/lib/paddleocr/doc/datasets/roLabelImg.png -> build/bdist.linux-x86_64/wheel/paddleocr/doc/datasets\ncopying build/lib/paddleocr/doc/datasets/labelimg.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/datasets\ncopying build/lib/paddleocr/doc/datasets/ch_street_rec_1.png -> build/bdist.linux-x86_64/wheel/paddleocr/doc/datasets\ncreating build/bdist.linux-x86_64/wheel/paddleocr/doc/demo\ncopying build/lib/paddleocr/doc/demo/build.png -> build/bdist.linux-x86_64/wheel/paddleocr/doc/demo\ncopying build/lib/paddleocr/doc/demo/proxy.png -> build/bdist.linux-x86_64/wheel/paddleocr/doc/demo\ncopying build/lib/paddleocr/doc/demo/error.png -> build/bdist.linux-x86_64/wheel/paddleocr/doc/demo\ncreating build/bdist.linux-x86_64/wheel/paddleocr/doc/doc_ch\ncopying build/lib/paddleocr/doc/doc_ch/models_list.md -> build/bdist.linux-x86_64/wheel/paddleocr/doc/doc_ch\ncopying build/lib/paddleocr/doc/doc_ch/add_new_algorithm.md -> build/bdist.linux-x86_64/wheel/paddleocr/doc/doc_ch\ncopying build/lib/paddleocr/doc/doc_ch/detection.md -> build/bdist.linux-x86_64/wheel/paddleocr/doc/doc_ch\ncopying build/lib/paddleocr/doc/doc_ch/data_annotation.md -> build/bdist.linux-x86_64/wheel/paddleocr/doc/doc_ch\ncopying build/lib/paddleocr/doc/doc_ch/quickstart.md -> build/bdist.linux-x86_64/wheel/paddleocr/doc/doc_ch\ncopying build/lib/paddleocr/doc/doc_ch/framework.png -> build/bdist.linux-x86_64/wheel/paddleocr/doc/doc_ch\ncopying build/lib/paddleocr/doc/doc_ch/vertical_and_multilingual_datasets.md -> build/bdist.linux-x86_64/wheel/paddleocr/doc/doc_ch\ncopying build/lib/paddleocr/doc/doc_ch/tree.md -> build/bdist.linux-x86_64/wheel/paddleocr/doc/doc_ch\ncopying build/lib/paddleocr/doc/doc_ch/algorithm_overview.md -> build/bdist.linux-x86_64/wheel/paddleocr/doc/doc_ch\ncopying build/lib/paddleocr/doc/doc_ch/config.md -> build/bdist.linux-x86_64/wheel/paddleocr/doc/doc_ch\ncopying build/lib/paddleocr/doc/doc_ch/data_synthesis.md -> build/bdist.linux-x86_64/wheel/paddleocr/doc/doc_ch\ncopying build/lib/paddleocr/doc/doc_ch/FAQ.md -> build/bdist.linux-x86_64/wheel/paddleocr/doc/doc_ch\ncopying build/lib/paddleocr/doc/doc_ch/multi_languages.md -> build/bdist.linux-x86_64/wheel/paddleocr/doc/doc_ch\ncopying build/lib/paddleocr/doc/doc_ch/customize.md -> build/bdist.linux-x86_64/wheel/paddleocr/doc/doc_ch\ncopying build/lib/paddleocr/doc/doc_ch/pgnet.md -> build/bdist.linux-x86_64/wheel/paddleocr/doc/doc_ch\ncopying build/lib/paddleocr/doc/doc_ch/installation.md -> build/bdist.linux-x86_64/wheel/paddleocr/doc/doc_ch\ncopying build/lib/paddleocr/doc/doc_ch/reference.md -> build/bdist.linux-x86_64/wheel/paddleocr/doc/doc_ch\ncopying build/lib/paddleocr/doc/doc_ch/benchmark.md -> build/bdist.linux-x86_64/wheel/paddleocr/doc/doc_ch\ncopying build/lib/paddleocr/doc/doc_ch/update.md -> build/bdist.linux-x86_64/wheel/paddleocr/doc/doc_ch\ncopying build/lib/paddleocr/doc/doc_ch/recognition.md -> build/bdist.linux-x86_64/wheel/paddleocr/doc/doc_ch\ncopying build/lib/paddleocr/doc/doc_ch/inference.md -> build/bdist.linux-x86_64/wheel/paddleocr/doc/doc_ch\ncopying build/lib/paddleocr/doc/doc_ch/visualization.md -> build/bdist.linux-x86_64/wheel/paddleocr/doc/doc_ch\ncopying build/lib/paddleocr/doc/doc_ch/angle_class.md -> build/bdist.linux-x86_64/wheel/paddleocr/doc/doc_ch\ncopying build/lib/paddleocr/doc/doc_ch/whl.md -> build/bdist.linux-x86_64/wheel/paddleocr/doc/doc_ch\ncopying build/lib/paddleocr/doc/doc_ch/serving_inference.md -> build/bdist.linux-x86_64/wheel/paddleocr/doc/doc_ch\ncopying build/lib/paddleocr/doc/doc_ch/datasets.md -> build/bdist.linux-x86_64/wheel/paddleocr/doc/doc_ch\ncopying build/lib/paddleocr/doc/doc_ch/android_demo.md -> build/bdist.linux-x86_64/wheel/paddleocr/doc/doc_ch\ncopying build/lib/paddleocr/doc/doc_ch/handwritten_datasets.md -> build/bdist.linux-x86_64/wheel/paddleocr/doc/doc_ch\ncreating build/bdist.linux-x86_64/wheel/paddleocr/doc/doc_en\ncopying build/lib/paddleocr/doc/doc_en/data_synthesis_en.md -> build/bdist.linux-x86_64/wheel/paddleocr/doc/doc_en\ncopying build/lib/paddleocr/doc/doc_en/tree_en.md -> build/bdist.linux-x86_64/wheel/paddleocr/doc/doc_en\ncopying build/lib/paddleocr/doc/doc_en/whl_en.md -> build/bdist.linux-x86_64/wheel/paddleocr/doc/doc_en\ncopying build/lib/paddleocr/doc/doc_en/customize_en.md -> build/bdist.linux-x86_64/wheel/paddleocr/doc/doc_en\ncopying build/lib/paddleocr/doc/doc_en/detection_en.md -> build/bdist.linux-x86_64/wheel/paddleocr/doc/doc_en\ncopying build/lib/paddleocr/doc/doc_en/benchmark_en.md -> build/bdist.linux-x86_64/wheel/paddleocr/doc/doc_en\ncopying build/lib/paddleocr/doc/doc_en/visualization_en.md -> build/bdist.linux-x86_64/wheel/paddleocr/doc/doc_en\ncopying build/lib/paddleocr/doc/doc_en/recognition_en.md -> build/bdist.linux-x86_64/wheel/paddleocr/doc/doc_en\ncopying build/lib/paddleocr/doc/doc_en/config_en.md -> build/bdist.linux-x86_64/wheel/paddleocr/doc/doc_en\ncopying build/lib/paddleocr/doc/doc_en/installation_en.md -> build/bdist.linux-x86_64/wheel/paddleocr/doc/doc_en\ncopying build/lib/paddleocr/doc/doc_en/models_list_en.md -> build/bdist.linux-x86_64/wheel/paddleocr/doc/doc_en\ncopying build/lib/paddleocr/doc/doc_en/multi_languages_en.md -> build/bdist.linux-x86_64/wheel/paddleocr/doc/doc_en\ncopying build/lib/paddleocr/doc/doc_en/inference_en.md -> build/bdist.linux-x86_64/wheel/paddleocr/doc/doc_en\ncopying build/lib/paddleocr/doc/doc_en/add_new_algorithm_en.md -> build/bdist.linux-x86_64/wheel/paddleocr/doc/doc_en\ncopying build/lib/paddleocr/doc/doc_en/tricks_en.md -> build/bdist.linux-x86_64/wheel/paddleocr/doc/doc_en\ncopying build/lib/paddleocr/doc/doc_en/pgnet_en.md -> build/bdist.linux-x86_64/wheel/paddleocr/doc/doc_en\ncopying build/lib/paddleocr/doc/doc_en/datasets_en.md -> build/bdist.linux-x86_64/wheel/paddleocr/doc/doc_en\ncopying build/lib/paddleocr/doc/doc_en/FAQ_en.md -> build/bdist.linux-x86_64/wheel/paddleocr/doc/doc_en\ncopying build/lib/paddleocr/doc/doc_en/handwritten_datasets_en.md -> build/bdist.linux-x86_64/wheel/paddleocr/doc/doc_en\ncopying build/lib/paddleocr/doc/doc_en/algorithm_overview_en.md -> build/bdist.linux-x86_64/wheel/paddleocr/doc/doc_en\ncopying build/lib/paddleocr/doc/doc_en/data_annotation_en.md -> build/bdist.linux-x86_64/wheel/paddleocr/doc/doc_en\ncopying build/lib/paddleocr/doc/doc_en/quickstart_en.md -> build/bdist.linux-x86_64/wheel/paddleocr/doc/doc_en\ncopying build/lib/paddleocr/doc/doc_en/vertical_and_multilingual_datasets_en.md -> build/bdist.linux-x86_64/wheel/paddleocr/doc/doc_en\ncopying build/lib/paddleocr/doc/doc_en/android_demo_en.md -> build/bdist.linux-x86_64/wheel/paddleocr/doc/doc_en\ncopying build/lib/paddleocr/doc/doc_en/update_en.md -> build/bdist.linux-x86_64/wheel/paddleocr/doc/doc_en\ncopying build/lib/paddleocr/doc/doc_en/reference_en.md -> build/bdist.linux-x86_64/wheel/paddleocr/doc/doc_en\ncopying build/lib/paddleocr/doc/doc_en/angle_class_en.md -> build/bdist.linux-x86_64/wheel/paddleocr/doc/doc_en\ncreating build/bdist.linux-x86_64/wheel/paddleocr/doc/fonts\ncopying build/lib/paddleocr/doc/fonts/kannada.ttf -> build/bdist.linux-x86_64/wheel/paddleocr/doc/fonts\ncopying build/lib/paddleocr/doc/fonts/telugu.ttf -> build/bdist.linux-x86_64/wheel/paddleocr/doc/fonts\ncopying build/lib/paddleocr/doc/fonts/german.ttf -> build/bdist.linux-x86_64/wheel/paddleocr/doc/fonts\ncopying build/lib/paddleocr/doc/fonts/korean.ttf -> build/bdist.linux-x86_64/wheel/paddleocr/doc/fonts\ncopying build/lib/paddleocr/doc/fonts/tamil.ttf -> build/bdist.linux-x86_64/wheel/paddleocr/doc/fonts\ncopying build/lib/paddleocr/doc/fonts/hindi.ttf -> build/bdist.linux-x86_64/wheel/paddleocr/doc/fonts\ncopying build/lib/paddleocr/doc/fonts/simfang.ttf -> build/bdist.linux-x86_64/wheel/paddleocr/doc/fonts\ncopying build/lib/paddleocr/doc/fonts/french.ttf -> build/bdist.linux-x86_64/wheel/paddleocr/doc/fonts\ncopying build/lib/paddleocr/doc/fonts/persian.ttf -> build/bdist.linux-x86_64/wheel/paddleocr/doc/fonts\ncopying build/lib/paddleocr/doc/fonts/marathi.ttf -> build/bdist.linux-x86_64/wheel/paddleocr/doc/fonts\ncopying build/lib/paddleocr/doc/fonts/japan.ttc -> build/bdist.linux-x86_64/wheel/paddleocr/doc/fonts\ncopying build/lib/paddleocr/doc/fonts/cyrillic.ttf -> build/bdist.linux-x86_64/wheel/paddleocr/doc/fonts\ncopying build/lib/paddleocr/doc/fonts/spanish.ttf -> build/bdist.linux-x86_64/wheel/paddleocr/doc/fonts\ncopying build/lib/paddleocr/doc/fonts/urdu.ttf -> build/bdist.linux-x86_64/wheel/paddleocr/doc/fonts\ncopying build/lib/paddleocr/doc/fonts/chinese_cht.ttf -> build/bdist.linux-x86_64/wheel/paddleocr/doc/fonts\ncopying build/lib/paddleocr/doc/fonts/arabic.ttf -> build/bdist.linux-x86_64/wheel/paddleocr/doc/fonts\ncopying build/lib/paddleocr/doc/fonts/latin.ttf -> build/bdist.linux-x86_64/wheel/paddleocr/doc/fonts\ncopying build/lib/paddleocr/doc/fonts/nepali.ttf -> build/bdist.linux-x86_64/wheel/paddleocr/doc/fonts\ncopying build/lib/paddleocr/doc/fonts/uyghur.ttf -> build/bdist.linux-x86_64/wheel/paddleocr/doc/fonts\ncreating build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs\ncopying build/lib/paddleocr/doc/imgs/00111002.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs\ncopying build/lib/paddleocr/doc/imgs/00207393.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs\ncopying build/lib/paddleocr/doc/imgs/00018069.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs\ncopying build/lib/paddleocr/doc/imgs/00006737.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs\ncopying build/lib/paddleocr/doc/imgs/00015504.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs\ncopying build/lib/paddleocr/doc/imgs/00056221.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs\ncopying build/lib/paddleocr/doc/imgs/12.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs\ncopying build/lib/paddleocr/doc/imgs/11.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs\ncopying build/lib/paddleocr/doc/imgs/00009282.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs\ncopying build/lib/paddleocr/doc/imgs/1.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs\ncopying build/lib/paddleocr/doc/imgs/00077949.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs\ncopying build/lib/paddleocr/doc/imgs/00057937.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs\ncopying build/lib/paddleocr/doc/imgs/00059985.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs\ncopying build/lib/paddleocr/doc/imgs/ger_1.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs\ncopying build/lib/paddleocr/doc/imgs/japan_1.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs\ncopying build/lib/paddleocr/doc/imgs/french_0.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs\ncopying build/lib/paddleocr/doc/imgs/ger_2.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs\ncopying build/lib/paddleocr/doc/imgs/korean_1.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs\ncopying build/lib/paddleocr/doc/imgs/japan_2.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs\ncopying build/lib/paddleocr/doc/imgs/model_prod_flow_ch.png -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs\ncreating build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_en\ncopying build/lib/paddleocr/doc/imgs_en/img_12.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_en\ncopying build/lib/paddleocr/doc/imgs_en/img_195.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_en\ncopying build/lib/paddleocr/doc/imgs_en/model_prod_flow_en.png -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_en\ncopying build/lib/paddleocr/doc/imgs_en/img_10.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_en\ncopying build/lib/paddleocr/doc/imgs_en/254.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_en\ncopying build/lib/paddleocr/doc/imgs_en/img623.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_en\ncopying build/lib/paddleocr/doc/imgs_en/img_11.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_en\ncreating build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_results\ncopying build/lib/paddleocr/doc/imgs_results/det_res_00018069.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_results\ncopying build/lib/paddleocr/doc/imgs_results/det_res_img_10_db.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_results\ncopying build/lib/paddleocr/doc/imgs_results/system_res_00018069.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_results\ncopying build/lib/paddleocr/doc/imgs_results/e2e_res_img293_pgnet.png -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_results\ncopying build/lib/paddleocr/doc/imgs_results/korean.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_results\ncopying build/lib/paddleocr/doc/imgs_results/angle_class_example.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_results\ncopying build/lib/paddleocr/doc/imgs_results/det_res_img_10_east.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_results\ncopying build/lib/paddleocr/doc/imgs_results/det_res_img623_sast.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_results\ncopying build/lib/paddleocr/doc/imgs_results/e2e_res_img623_pgnet.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_results\ncopying build/lib/paddleocr/doc/imgs_results/img_10_east_starnet.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_results\ncopying build/lib/paddleocr/doc/imgs_results/french_0.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_results\ncopying build/lib/paddleocr/doc/imgs_results/e2e_res_img295_pgnet.png -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_results\ncopying build/lib/paddleocr/doc/imgs_results/det_res_img_10_sast.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_results\ncopying build/lib/paddleocr/doc/imgs_results/e2e_res_img_10_pgnet.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_results\ncreating build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_results/ch_ppocr_mobile_v2.0\ncopying build/lib/paddleocr/doc/imgs_results/ch_ppocr_mobile_v2.0/00207393.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_results/ch_ppocr_mobile_v2.0\ncopying build/lib/paddleocr/doc/imgs_results/ch_ppocr_mobile_v2.0/00059985.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_results/ch_ppocr_mobile_v2.0\ncopying build/lib/paddleocr/doc/imgs_results/ch_ppocr_mobile_v2.0/00077949.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_results/ch_ppocr_mobile_v2.0\ncopying build/lib/paddleocr/doc/imgs_results/ch_ppocr_mobile_v2.0/test_add_91.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_results/ch_ppocr_mobile_v2.0\ncopying build/lib/paddleocr/doc/imgs_results/ch_ppocr_mobile_v2.0/00006737.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_results/ch_ppocr_mobile_v2.0\ncopying build/lib/paddleocr/doc/imgs_results/ch_ppocr_mobile_v2.0/00015504.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_results/ch_ppocr_mobile_v2.0\ncopying build/lib/paddleocr/doc/imgs_results/ch_ppocr_mobile_v2.0/00111002.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_results/ch_ppocr_mobile_v2.0\ncopying build/lib/paddleocr/doc/imgs_results/ch_ppocr_mobile_v2.0/img_12.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_results/ch_ppocr_mobile_v2.0\ncopying build/lib/paddleocr/doc/imgs_results/ch_ppocr_mobile_v2.0/00009282.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_results/ch_ppocr_mobile_v2.0\ncopying build/lib/paddleocr/doc/imgs_results/ch_ppocr_mobile_v2.0/00057937.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_results/ch_ppocr_mobile_v2.0\ncopying build/lib/paddleocr/doc/imgs_results/ch_ppocr_mobile_v2.0/rotate_00052204.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_results/ch_ppocr_mobile_v2.0\ncopying build/lib/paddleocr/doc/imgs_results/ch_ppocr_mobile_v2.0/00018069.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_results/ch_ppocr_mobile_v2.0\ncopying build/lib/paddleocr/doc/imgs_results/ch_ppocr_mobile_v2.0/00056221.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_results/ch_ppocr_mobile_v2.0\ncreating build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_results/multi_lang\ncopying build/lib/paddleocr/doc/imgs_results/multi_lang/img_02.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_results/multi_lang\ncopying build/lib/paddleocr/doc/imgs_results/multi_lang/img_01.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_results/multi_lang\ncopying build/lib/paddleocr/doc/imgs_results/multi_lang/japan_2.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_results/multi_lang\ncopying build/lib/paddleocr/doc/imgs_results/multi_lang/arabic_0.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_results/multi_lang\ncopying build/lib/paddleocr/doc/imgs_results/multi_lang/korean_0.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_results/multi_lang\ncopying build/lib/paddleocr/doc/imgs_results/multi_lang/en_1.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_results/multi_lang\ncopying build/lib/paddleocr/doc/imgs_results/multi_lang/img_12.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_results/multi_lang\ncopying build/lib/paddleocr/doc/imgs_results/multi_lang/en_2.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_results/multi_lang\ncopying build/lib/paddleocr/doc/imgs_results/multi_lang/en_3.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_results/multi_lang\ncopying build/lib/paddleocr/doc/imgs_results/multi_lang/french_0.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_results/multi_lang\ncreating build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_results/whl\ncopying build/lib/paddleocr/doc/imgs_results/whl/11_det_rec.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_results/whl\ncopying build/lib/paddleocr/doc/imgs_results/whl/12_det_rec.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_results/whl\ncopying build/lib/paddleocr/doc/imgs_results/whl/12_det.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_results/whl\ncopying build/lib/paddleocr/doc/imgs_results/whl/11_det.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_results/whl\ncreating build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words\ncreating build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/arabic\ncopying build/lib/paddleocr/doc/imgs_words/arabic/ar_2.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/arabic\ncopying build/lib/paddleocr/doc/imgs_words/arabic/ar_1.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/arabic\ncreating build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/belarusian\ncopying build/lib/paddleocr/doc/imgs_words/belarusian/be_1.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/belarusian\ncopying build/lib/paddleocr/doc/imgs_words/belarusian/be_2.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/belarusian\ncreating build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/bulgarian\ncopying build/lib/paddleocr/doc/imgs_words/bulgarian/bg_1.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/bulgarian\ncopying build/lib/paddleocr/doc/imgs_words/bulgarian/bg_2.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/bulgarian\ncreating build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/ch\ncopying build/lib/paddleocr/doc/imgs_words/ch/word_5.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/ch\ncopying build/lib/paddleocr/doc/imgs_words/ch/word_2.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/ch\ncopying build/lib/paddleocr/doc/imgs_words/ch/word_3.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/ch\ncopying build/lib/paddleocr/doc/imgs_words/ch/word_4.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/ch\ncopying build/lib/paddleocr/doc/imgs_words/ch/word_1.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/ch\ncreating build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/chinese_traditional\ncopying build/lib/paddleocr/doc/imgs_words/chinese_traditional/chinese_cht_1.png -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/chinese_traditional\ncopying build/lib/paddleocr/doc/imgs_words/chinese_traditional/chinese_cht_2.png -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/chinese_traditional\ncreating build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/en\ncopying build/lib/paddleocr/doc/imgs_words/en/word_5.png -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/en\ncopying build/lib/paddleocr/doc/imgs_words/en/word_4.png -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/en\ncopying build/lib/paddleocr/doc/imgs_words/en/word_1.png -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/en\ncopying build/lib/paddleocr/doc/imgs_words/en/word_3.png -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/en\ncopying build/lib/paddleocr/doc/imgs_words/en/word_2.png -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/en\ncreating build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/french\ncopying build/lib/paddleocr/doc/imgs_words/french/2.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/french\ncopying build/lib/paddleocr/doc/imgs_words/french/1.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/french\ncreating build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/german\ncopying build/lib/paddleocr/doc/imgs_words/german/1.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/german\ncreating build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/hindi\ncopying build/lib/paddleocr/doc/imgs_words/hindi/hi_1.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/hindi\ncopying build/lib/paddleocr/doc/imgs_words/hindi/hi_2.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/hindi\ncreating build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/italian\ncopying build/lib/paddleocr/doc/imgs_words/italian/it_1.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/italian\ncopying build/lib/paddleocr/doc/imgs_words/italian/it_2.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/italian\ncreating build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/japan\ncopying build/lib/paddleocr/doc/imgs_words/japan/1.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/japan\ncreating build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/kannada\ncopying build/lib/paddleocr/doc/imgs_words/kannada/ka_1.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/kannada\ncopying build/lib/paddleocr/doc/imgs_words/kannada/ka_2.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/kannada\ncreating build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/korean\ncopying build/lib/paddleocr/doc/imgs_words/korean/1.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/korean\ncopying build/lib/paddleocr/doc/imgs_words/korean/2.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/korean\ncreating build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/marathi\ncopying build/lib/paddleocr/doc/imgs_words/marathi/mr_1.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/marathi\ncopying build/lib/paddleocr/doc/imgs_words/marathi/mr_2.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/marathi\ncreating build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/nepali\ncopying build/lib/paddleocr/doc/imgs_words/nepali/ne_2.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/nepali\ncopying build/lib/paddleocr/doc/imgs_words/nepali/ne_1.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/nepali\ncreating build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/occitan\ncopying build/lib/paddleocr/doc/imgs_words/occitan/oc_1.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/occitan\ncopying build/lib/paddleocr/doc/imgs_words/occitan/oc_2.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/occitan\ncreating build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/persian\ncopying build/lib/paddleocr/doc/imgs_words/persian/fa_1.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/persian\ncopying build/lib/paddleocr/doc/imgs_words/persian/fa_2.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/persian\ncreating build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/portuguese\ncopying build/lib/paddleocr/doc/imgs_words/portuguese/pu_1.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/portuguese\ncopying build/lib/paddleocr/doc/imgs_words/portuguese/pu_2.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/portuguese\ncreating build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/russia\ncopying build/lib/paddleocr/doc/imgs_words/russia/ru_2.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/russia\ncopying build/lib/paddleocr/doc/imgs_words/russia/ru_1.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/russia\ncreating build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/serbian_cyrillic\ncopying build/lib/paddleocr/doc/imgs_words/serbian_cyrillic/rsc_2.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/serbian_cyrillic\ncopying build/lib/paddleocr/doc/imgs_words/serbian_cyrillic/rsc_1.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/serbian_cyrillic\ncreating build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/serbian_latin\ncopying build/lib/paddleocr/doc/imgs_words/serbian_latin/rs_2.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/serbian_latin\ncopying build/lib/paddleocr/doc/imgs_words/serbian_latin/rs_1.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/serbian_latin\ncreating build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/spanish\ncopying build/lib/paddleocr/doc/imgs_words/spanish/xi_2.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/spanish\ncopying build/lib/paddleocr/doc/imgs_words/spanish/xi_1.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/spanish\ncreating build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/tamil\ncopying build/lib/paddleocr/doc/imgs_words/tamil/ta_1.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/tamil\ncopying build/lib/paddleocr/doc/imgs_words/tamil/ta_2.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/tamil\ncreating build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/telugu\ncopying build/lib/paddleocr/doc/imgs_words/telugu/te_2.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/telugu\ncopying build/lib/paddleocr/doc/imgs_words/telugu/te_1.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/telugu\ncreating build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/ukranian\ncopying build/lib/paddleocr/doc/imgs_words/ukranian/uk_2.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/ukranian\ncopying build/lib/paddleocr/doc/imgs_words/ukranian/uk_1.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/ukranian\ncreating build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/urdu\ncopying build/lib/paddleocr/doc/imgs_words/urdu/ur_1.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/urdu\ncopying build/lib/paddleocr/doc/imgs_words/urdu/ur_2.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/urdu\ncreating build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/uyghur\ncopying build/lib/paddleocr/doc/imgs_words/uyghur/ug_1.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/uyghur\ncopying build/lib/paddleocr/doc/imgs_words/uyghur/ug_2.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words/uyghur\ncreating build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words_en\ncopying build/lib/paddleocr/doc/imgs_words_en/word_336.png -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words_en\ncopying build/lib/paddleocr/doc/imgs_words_en/word_116.png -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words_en\ncopying build/lib/paddleocr/doc/imgs_words_en/word_545.png -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words_en\ncopying build/lib/paddleocr/doc/imgs_words_en/word_401.png -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words_en\ncopying build/lib/paddleocr/doc/imgs_words_en/word_10.png -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words_en\ncopying build/lib/paddleocr/doc/imgs_words_en/word_201.png -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words_en\ncopying build/lib/paddleocr/doc/imgs_words_en/word_308.png -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words_en\ncopying build/lib/paddleocr/doc/imgs_words_en/word_461.png -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words_en\ncopying build/lib/paddleocr/doc/imgs_words_en/word_52.png -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words_en\ncopying build/lib/paddleocr/doc/imgs_words_en/word_19.png -> build/bdist.linux-x86_64/wheel/paddleocr/doc/imgs_words_en\ncreating build/bdist.linux-x86_64/wheel/paddleocr/doc/tricks\ncopying build/lib/paddleocr/doc/tricks/long_text_examples.jpg -> build/bdist.linux-x86_64/wheel/paddleocr/doc/tricks\ncreating build/bdist.linux-x86_64/wheel/paddleocr/paddleocr.egg-info\ncopying build/lib/paddleocr/paddleocr.egg-info/PKG-INFO -> build/bdist.linux-x86_64/wheel/paddleocr/paddleocr.egg-info\ncopying build/lib/paddleocr/paddleocr.egg-info/requires.txt -> build/bdist.linux-x86_64/wheel/paddleocr/paddleocr.egg-info\ncopying build/lib/paddleocr/paddleocr.egg-info/top_level.txt -> build/bdist.linux-x86_64/wheel/paddleocr/paddleocr.egg-info\ncopying build/lib/paddleocr/paddleocr.egg-info/SOURCES.txt -> build/bdist.linux-x86_64/wheel/paddleocr/paddleocr.egg-info\ncopying build/lib/paddleocr/paddleocr.egg-info/dependency_links.txt -> build/bdist.linux-x86_64/wheel/paddleocr/paddleocr.egg-info\ncopying build/lib/paddleocr/paddleocr.egg-info/entry_points.txt -> build/bdist.linux-x86_64/wheel/paddleocr/paddleocr.egg-info\ncreating build/bdist.linux-x86_64/wheel/paddleocr/ppocr\ncopying build/lib/paddleocr/ppocr/__init__.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr\ncreating build/bdist.linux-x86_64/wheel/paddleocr/ppocr/data\ncopying build/lib/paddleocr/ppocr/data/lmdb_dataset.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/data\ncopying build/lib/paddleocr/ppocr/data/pgnet_dataset.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/data\ncopying build/lib/paddleocr/ppocr/data/simple_dataset.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/data\ncopying build/lib/paddleocr/ppocr/data/__init__.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/data\ncreating build/bdist.linux-x86_64/wheel/paddleocr/ppocr/data/imaug\ncopying build/lib/paddleocr/ppocr/data/imaug/__init__.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/data/imaug\ncopying build/lib/paddleocr/ppocr/data/imaug/label_ops.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/data/imaug\ncopying build/lib/paddleocr/ppocr/data/imaug/make_border_map.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/data/imaug\ncopying build/lib/paddleocr/ppocr/data/imaug/rec_img_aug.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/data/imaug\ncopying build/lib/paddleocr/ppocr/data/imaug/iaa_augment.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/data/imaug\ncopying build/lib/paddleocr/ppocr/data/imaug/sast_process.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/data/imaug\ncopying build/lib/paddleocr/ppocr/data/imaug/east_process.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/data/imaug\ncopying build/lib/paddleocr/ppocr/data/imaug/make_shrink_map.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/data/imaug\ncopying build/lib/paddleocr/ppocr/data/imaug/random_crop_data.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/data/imaug\ncopying build/lib/paddleocr/ppocr/data/imaug/operators.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/data/imaug\ncopying build/lib/paddleocr/ppocr/data/imaug/pg_process.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/data/imaug\ncopying build/lib/paddleocr/ppocr/data/imaug/randaugment.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/data/imaug\ncreating build/bdist.linux-x86_64/wheel/paddleocr/ppocr/data/imaug/text_image_aug\ncopying build/lib/paddleocr/ppocr/data/imaug/text_image_aug/__init__.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/data/imaug/text_image_aug\ncopying build/lib/paddleocr/ppocr/data/imaug/text_image_aug/warp_mls.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/data/imaug/text_image_aug\ncopying build/lib/paddleocr/ppocr/data/imaug/text_image_aug/augment.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/data/imaug/text_image_aug\ncreating build/bdist.linux-x86_64/wheel/paddleocr/ppocr/losses\ncopying build/lib/paddleocr/ppocr/losses/det_sast_loss.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/losses\ncopying build/lib/paddleocr/ppocr/losses/rec_att_loss.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/losses\ncopying build/lib/paddleocr/ppocr/losses/rec_srn_loss.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/losses\ncopying build/lib/paddleocr/ppocr/losses/e2e_pg_loss.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/losses\ncopying build/lib/paddleocr/ppocr/losses/rec_ctc_loss.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/losses\ncopying build/lib/paddleocr/ppocr/losses/__init__.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/losses\ncopying build/lib/paddleocr/ppocr/losses/det_basic_loss.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/losses\ncopying build/lib/paddleocr/ppocr/losses/det_east_loss.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/losses\ncopying build/lib/paddleocr/ppocr/losses/cls_loss.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/losses\ncopying build/lib/paddleocr/ppocr/losses/det_db_loss.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/losses\ncreating build/bdist.linux-x86_64/wheel/paddleocr/ppocr/metrics\ncopying build/lib/paddleocr/ppocr/metrics/cls_metric.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/metrics\ncopying build/lib/paddleocr/ppocr/metrics/e2e_metric.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/metrics\ncopying build/lib/paddleocr/ppocr/metrics/__init__.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/metrics\ncopying build/lib/paddleocr/ppocr/metrics/det_metric.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/metrics\ncopying build/lib/paddleocr/ppocr/metrics/rec_metric.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/metrics\ncopying build/lib/paddleocr/ppocr/metrics/eval_det_iou.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/metrics\ncreating build/bdist.linux-x86_64/wheel/paddleocr/ppocr/modeling\ncreating build/bdist.linux-x86_64/wheel/paddleocr/ppocr/modeling/architectures\ncopying build/lib/paddleocr/ppocr/modeling/architectures/base_model.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/modeling/architectures\ncopying build/lib/paddleocr/ppocr/modeling/architectures/__init__.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/modeling/architectures\ncreating build/bdist.linux-x86_64/wheel/paddleocr/ppocr/modeling/backbones\ncopying build/lib/paddleocr/ppocr/modeling/backbones/rec_resnet_fpn.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/modeling/backbones\ncopying build/lib/paddleocr/ppocr/modeling/backbones/det_resnet_vd.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/modeling/backbones\ncopying build/lib/paddleocr/ppocr/modeling/backbones/__init__.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/modeling/backbones\ncopying build/lib/paddleocr/ppocr/modeling/backbones/det_mobilenet_v3.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/modeling/backbones\ncopying build/lib/paddleocr/ppocr/modeling/backbones/det_resnet_vd_sast.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/modeling/backbones\ncopying build/lib/paddleocr/ppocr/modeling/backbones/e2e_resnet_vd_pg.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/modeling/backbones\ncopying build/lib/paddleocr/ppocr/modeling/backbones/rec_mobilenet_v3.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/modeling/backbones\ncopying build/lib/paddleocr/ppocr/modeling/backbones/rec_resnet_vd.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/modeling/backbones\ncreating build/bdist.linux-x86_64/wheel/paddleocr/ppocr/modeling/heads\ncopying build/lib/paddleocr/ppocr/modeling/heads/cls_head.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/modeling/heads\ncopying build/lib/paddleocr/ppocr/modeling/heads/self_attention.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/modeling/heads\ncopying build/lib/paddleocr/ppocr/modeling/heads/rec_att_head.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/modeling/heads\ncopying build/lib/paddleocr/ppocr/modeling/heads/e2e_pg_head.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/modeling/heads\ncopying build/lib/paddleocr/ppocr/modeling/heads/det_east_head.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/modeling/heads\ncopying build/lib/paddleocr/ppocr/modeling/heads/det_db_head.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/modeling/heads\ncopying build/lib/paddleocr/ppocr/modeling/heads/rec_srn_head.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/modeling/heads\ncopying build/lib/paddleocr/ppocr/modeling/heads/__init__.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/modeling/heads\ncopying build/lib/paddleocr/ppocr/modeling/heads/det_sast_head.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/modeling/heads\ncopying build/lib/paddleocr/ppocr/modeling/heads/rec_ctc_head.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/modeling/heads\ncreating build/bdist.linux-x86_64/wheel/paddleocr/ppocr/modeling/necks\ncopying build/lib/paddleocr/ppocr/modeling/necks/db_fpn.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/modeling/necks\ncopying build/lib/paddleocr/ppocr/modeling/necks/sast_fpn.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/modeling/necks\ncopying build/lib/paddleocr/ppocr/modeling/necks/pg_fpn.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/modeling/necks\ncopying build/lib/paddleocr/ppocr/modeling/necks/east_fpn.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/modeling/necks\ncopying build/lib/paddleocr/ppocr/modeling/necks/rnn.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/modeling/necks\ncopying build/lib/paddleocr/ppocr/modeling/necks/__init__.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/modeling/necks\ncreating build/bdist.linux-x86_64/wheel/paddleocr/ppocr/modeling/transforms\ncopying build/lib/paddleocr/ppocr/modeling/transforms/__init__.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/modeling/transforms\ncopying build/lib/paddleocr/ppocr/modeling/transforms/tps.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/modeling/transforms\ncreating build/bdist.linux-x86_64/wheel/paddleocr/ppocr/optimizer\ncopying build/lib/paddleocr/ppocr/optimizer/__init__.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/optimizer\ncopying build/lib/paddleocr/ppocr/optimizer/optimizer.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/optimizer\ncopying build/lib/paddleocr/ppocr/optimizer/learning_rate.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/optimizer\ncopying build/lib/paddleocr/ppocr/optimizer/regularizer.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/optimizer\ncopying build/lib/paddleocr/ppocr/optimizer/lr_scheduler.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/optimizer\ncreating build/bdist.linux-x86_64/wheel/paddleocr/ppocr/postprocess\ncopying build/lib/paddleocr/ppocr/postprocess/locality_aware_nms.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/postprocess\ncopying build/lib/paddleocr/ppocr/postprocess/__init__.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/postprocess\ncopying build/lib/paddleocr/ppocr/postprocess/cls_postprocess.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/postprocess\ncopying build/lib/paddleocr/ppocr/postprocess/rec_postprocess.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/postprocess\ncopying build/lib/paddleocr/ppocr/postprocess/east_postprocess.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/postprocess\ncopying build/lib/paddleocr/ppocr/postprocess/sast_postprocess.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/postprocess\ncopying build/lib/paddleocr/ppocr/postprocess/pg_postprocess.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/postprocess\ncopying build/lib/paddleocr/ppocr/postprocess/db_postprocess.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/postprocess\ncreating build/bdist.linux-x86_64/wheel/paddleocr/ppocr/utils\ncopying build/lib/paddleocr/ppocr/utils/__init__.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/utils\ncopying build/lib/paddleocr/ppocr/utils/logging.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/utils\ncopying build/lib/paddleocr/ppocr/utils/ic15_dict.txt -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/utils\ncopying build/lib/paddleocr/ppocr/utils/stats.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/utils\ncopying build/lib/paddleocr/ppocr/utils/ppocr_keys_v1.txt -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/utils\ncopying build/lib/paddleocr/ppocr/utils/utility.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/utils\ncopying build/lib/paddleocr/ppocr/utils/gen_label.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/utils\ncopying build/lib/paddleocr/ppocr/utils/save_load.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/utils\ncopying build/lib/paddleocr/ppocr/utils/en_dict.txt -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/utils\ncreating build/bdist.linux-x86_64/wheel/paddleocr/ppocr/utils/dict\ncopying build/lib/paddleocr/ppocr/utils/dict/en_dict.txt -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/utils/dict\ncopying build/lib/paddleocr/ppocr/utils/dict/devanagari_dict.txt -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/utils/dict\ncopying build/lib/paddleocr/ppocr/utils/dict/ar_dict.txt -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/utils/dict\ncopying build/lib/paddleocr/ppocr/utils/dict/fa_dict.txt -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/utils/dict\ncopying build/lib/paddleocr/ppocr/utils/dict/bg_dict.txt -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/utils/dict\ncopying build/lib/paddleocr/ppocr/utils/dict/arabic_dict.txt -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/utils/dict\ncopying build/lib/paddleocr/ppocr/utils/dict/be_dict.txt -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/utils/dict\ncopying build/lib/paddleocr/ppocr/utils/dict/cyrillic_dict.txt -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/utils/dict\ncopying build/lib/paddleocr/ppocr/utils/dict/chinese_cht_dict.txt -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/utils/dict\ncopying build/lib/paddleocr/ppocr/utils/dict/latin_dict.txt -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/utils/dict\ncopying build/lib/paddleocr/ppocr/utils/dict/ka_dict.txt -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/utils/dict\ncopying build/lib/paddleocr/ppocr/utils/dict/german_dict.txt -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/utils/dict\ncopying build/lib/paddleocr/ppocr/utils/dict/rs_dict.txt -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/utils/dict\ncopying build/lib/paddleocr/ppocr/utils/dict/uk_dict.txt -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/utils/dict\ncopying build/lib/paddleocr/ppocr/utils/dict/french_dict.txt -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/utils/dict\ncopying build/lib/paddleocr/ppocr/utils/dict/ne_dict.txt -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/utils/dict\ncopying build/lib/paddleocr/ppocr/utils/dict/ta_dict.txt -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/utils/dict\ncopying build/lib/paddleocr/ppocr/utils/dict/it_dict.txt -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/utils/dict\ncopying build/lib/paddleocr/ppocr/utils/dict/pu_dict.txt -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/utils/dict\ncopying build/lib/paddleocr/ppocr/utils/dict/japan_dict.txt -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/utils/dict\ncopying build/lib/paddleocr/ppocr/utils/dict/ru_dict.txt -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/utils/dict\ncopying build/lib/paddleocr/ppocr/utils/dict/oc_dict.txt -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/utils/dict\ncopying build/lib/paddleocr/ppocr/utils/dict/mr_dict.txt -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/utils/dict\ncopying build/lib/paddleocr/ppocr/utils/dict/te_dict.txt -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/utils/dict\ncopying build/lib/paddleocr/ppocr/utils/dict/hi_dict.txt -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/utils/dict\ncopying build/lib/paddleocr/ppocr/utils/dict/ug_dict.txt -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/utils/dict\ncopying build/lib/paddleocr/ppocr/utils/dict/ur_dict.txt -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/utils/dict\ncopying build/lib/paddleocr/ppocr/utils/dict/xi_dict.txt -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/utils/dict\ncopying build/lib/paddleocr/ppocr/utils/dict/korean_dict.txt -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/utils/dict\ncopying build/lib/paddleocr/ppocr/utils/dict/rsc_dict.txt -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/utils/dict\ncreating build/bdist.linux-x86_64/wheel/paddleocr/ppocr/utils/e2e_metric\ncopying build/lib/paddleocr/ppocr/utils/e2e_metric/polygon_fast.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/utils/e2e_metric\ncopying build/lib/paddleocr/ppocr/utils/e2e_metric/Deteval.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/utils/e2e_metric\ncreating build/bdist.linux-x86_64/wheel/paddleocr/ppocr/utils/e2e_utils\ncopying build/lib/paddleocr/ppocr/utils/e2e_utils/extract_textpoint_fast.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/utils/e2e_utils\ncopying build/lib/paddleocr/ppocr/utils/e2e_utils/pgnet_pp_utils.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/utils/e2e_utils\ncopying build/lib/paddleocr/ppocr/utils/e2e_utils/extract_textpoint_slow.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/utils/e2e_utils\ncopying build/lib/paddleocr/ppocr/utils/e2e_utils/visual.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/utils/e2e_utils\ncopying build/lib/paddleocr/ppocr/utils/e2e_utils/extract_batchsize.py -> build/bdist.linux-x86_64/wheel/paddleocr/ppocr/utils/e2e_utils\ncreating build/bdist.linux-x86_64/wheel/paddleocr/tools\ncopying build/lib/paddleocr/tools/test_hubserving.py -> build/bdist.linux-x86_64/wheel/paddleocr/tools\ncopying build/lib/paddleocr/tools/infer_det.py -> build/bdist.linux-x86_64/wheel/paddleocr/tools\ncopying build/lib/paddleocr/tools/eval.py -> build/bdist.linux-x86_64/wheel/paddleocr/tools\ncopying build/lib/paddleocr/tools/infer_e2e.py -> build/bdist.linux-x86_64/wheel/paddleocr/tools\ncopying build/lib/paddleocr/tools/infer_cls.py -> build/bdist.linux-x86_64/wheel/paddleocr/tools\ncopying build/lib/paddleocr/tools/export_model.py -> build/bdist.linux-x86_64/wheel/paddleocr/tools\ncopying build/lib/paddleocr/tools/train.py -> build/bdist.linux-x86_64/wheel/paddleocr/tools\ncopying build/lib/paddleocr/tools/infer_rec.py -> build/bdist.linux-x86_64/wheel/paddleocr/tools\ncopying build/lib/paddleocr/tools/program.py -> build/bdist.linux-x86_64/wheel/paddleocr/tools\ncreating build/bdist.linux-x86_64/wheel/paddleocr/tools/infer\ncopying build/lib/paddleocr/tools/infer/predict_det.py -> build/bdist.linux-x86_64/wheel/paddleocr/tools/infer\ncopying build/lib/paddleocr/tools/infer/predict_rec.py -> build/bdist.linux-x86_64/wheel/paddleocr/tools/infer\ncopying build/lib/paddleocr/tools/infer/predict_e2e.py -> build/bdist.linux-x86_64/wheel/paddleocr/tools/infer\ncopying build/lib/paddleocr/tools/infer/utility.py -> build/bdist.linux-x86_64/wheel/paddleocr/tools/infer\ncopying build/lib/paddleocr/tools/infer/predict_cls.py -> build/bdist.linux-x86_64/wheel/paddleocr/tools/infer\ncopying build/lib/paddleocr/tools/infer/predict_system.py -> build/bdist.linux-x86_64/wheel/paddleocr/tools/infer\ncopying build/lib/paddleocr/paddleocr.py -> build/bdist.linux-x86_64/wheel/paddleocr\nrunning install_egg_info\nCopying paddleocr.egg-info to build/bdist.linux-x86_64/wheel/paddleocr-2.0.6-py3.7.egg-info\nrunning install_scripts\nadding license file \"LICENSE\" (matched pattern \"LICEN[CS]E*\")\ncreating build/bdist.linux-x86_64/wheel/paddleocr-2.0.6.dist-info/WHEEL\ncreating 'dist/paddleocr-2.0.6-py3-none-any.whl' and adding 'build/bdist.linux-x86_64/wheel' to it\nadding 'paddleocr/.clang_format.hook'\nadding 'paddleocr/.gitignore'\nadding 'paddleocr/.pre-commit-config.yaml'\nadding 'paddleocr/.style.yapf'\nadding 'paddleocr/LICENSE'\nadding 'paddleocr/MANIFEST.in'\nadding 'paddleocr/README.md'\nadding 'paddleocr/README_ch.md'\nadding 'paddleocr/__init__.py'\nadding 'paddleocr/paddleocr.py'\nadding 'paddleocr/requirements.txt'\nadding 'paddleocr/train.sh'\nadding 'paddleocr/PPOCRLabel/Makefile'\nadding 'paddleocr/PPOCRLabel/PPOCRLabel.py'\nadding 'paddleocr/PPOCRLabel/README.md'\nadding 'paddleocr/PPOCRLabel/README_ch.md'\nadding 'paddleocr/PPOCRLabel/__init__.py'\nadding 'paddleocr/PPOCRLabel/combobox.py'\nadding 'paddleocr/PPOCRLabel/resources.qrc'\nadding 'paddleocr/PPOCRLabel/setup.cfg'\nadding 'paddleocr/PPOCRLabel/setup.py'\nadding 'paddleocr/PPOCRLabel/data/paddle.png'\nadding 'paddleocr/PPOCRLabel/data/predefined_classes.txt'\nadding 'paddleocr/PPOCRLabel/data/gif/steps.gif'\nadding 'paddleocr/PPOCRLabel/data/gif/steps_en.gif'\nadding 'paddleocr/PPOCRLabel/libs/__init__.py'\nadding 'paddleocr/PPOCRLabel/libs/autoDialog.py'\nadding 'paddleocr/PPOCRLabel/libs/canvas.py'\nadding 'paddleocr/PPOCRLabel/libs/colorDialog.py'\nadding 'paddleocr/PPOCRLabel/libs/constants.py'\nadding 'paddleocr/PPOCRLabel/libs/create_ml_io.py'\nadding 'paddleocr/PPOCRLabel/libs/editinlist.py'\nadding 'paddleocr/PPOCRLabel/libs/hashableQListWidgetItem.py'\nadding 'paddleocr/PPOCRLabel/libs/labelDialog.py'\nadding 'paddleocr/PPOCRLabel/libs/resources.py'\nadding 'paddleocr/PPOCRLabel/libs/settings.py'\nadding 'paddleocr/PPOCRLabel/libs/shape.py'\nadding 'paddleocr/PPOCRLabel/libs/stringBundle.py'\nadding 'paddleocr/PPOCRLabel/libs/toolBar.py'\nadding 'paddleocr/PPOCRLabel/libs/ustr.py'\nadding 'paddleocr/PPOCRLabel/libs/utils.py'\nadding 'paddleocr/PPOCRLabel/libs/zoomWidget.py'\nadding 'paddleocr/PPOCRLabel/resources/icons/Auto.png'\nadding 'paddleocr/PPOCRLabel/resources/icons/app.icns'\nadding 'paddleocr/PPOCRLabel/resources/icons/app.png'\nadding 'paddleocr/PPOCRLabel/resources/icons/app.svg'\nadding 'paddleocr/PPOCRLabel/resources/icons/cancel.png'\nadding 'paddleocr/PPOCRLabel/resources/icons/close.png'\nadding 'paddleocr/PPOCRLabel/resources/icons/color.png'\nadding 'paddleocr/PPOCRLabel/resources/icons/color_line.png'\nadding 'paddleocr/PPOCRLabel/resources/icons/copy.png'\nadding 'paddleocr/PPOCRLabel/resources/icons/delete.png'\nadding 'paddleocr/PPOCRLabel/resources/icons/done.png'\nadding 'paddleocr/PPOCRLabel/resources/icons/done.svg'\nadding 'paddleocr/PPOCRLabel/resources/icons/edit.png'\nadding 'paddleocr/PPOCRLabel/resources/icons/expert1.png'\nadding 'paddleocr/PPOCRLabel/resources/icons/expert2.png'\nadding 'paddleocr/PPOCRLabel/resources/icons/eye.png'\nadding 'paddleocr/PPOCRLabel/resources/icons/feBlend-icon.png'\nadding 'paddleocr/PPOCRLabel/resources/icons/file.png'\nadding 'paddleocr/PPOCRLabel/resources/icons/fit-width.png'\nadding 'paddleocr/PPOCRLabel/resources/icons/fit-window.png'\nadding 'paddleocr/PPOCRLabel/resources/icons/fit.png'\nadding 'paddleocr/PPOCRLabel/resources/icons/format_createml.png'\nadding 'paddleocr/PPOCRLabel/resources/icons/format_voc.png'\nadding 'paddleocr/PPOCRLabel/resources/icons/format_yolo.png'\nadding 'paddleocr/PPOCRLabel/resources/icons/help.png'\nadding 'paddleocr/PPOCRLabel/resources/icons/labels.png'\nadding 'paddleocr/PPOCRLabel/resources/icons/labels.svg'\nadding 'paddleocr/PPOCRLabel/resources/icons/new.png'\nadding 'paddleocr/PPOCRLabel/resources/icons/next.png'\nadding 'paddleocr/PPOCRLabel/resources/icons/objects.png'\nadding 'paddleocr/PPOCRLabel/resources/icons/open.png'\nadding 'paddleocr/PPOCRLabel/resources/icons/open.svg'\nadding 'paddleocr/PPOCRLabel/resources/icons/prev.png'\nadding 'paddleocr/PPOCRLabel/resources/icons/quit.png'\nadding 'paddleocr/PPOCRLabel/resources/icons/reRec.png'\nadding 'paddleocr/PPOCRLabel/resources/icons/resetall.png'\nadding 'paddleocr/PPOCRLabel/resources/icons/save-as.png'\nadding 'paddleocr/PPOCRLabel/resources/icons/save-as.svg'\nadding 'paddleocr/PPOCRLabel/resources/icons/save.png'\nadding 'paddleocr/PPOCRLabel/resources/icons/save.svg'\nadding 'paddleocr/PPOCRLabel/resources/icons/undo-cross.png'\nadding 'paddleocr/PPOCRLabel/resources/icons/undo.png'\nadding 'paddleocr/PPOCRLabel/resources/icons/verify.png'\nadding 'paddleocr/PPOCRLabel/resources/icons/zoom-in.png'\nadding 'paddleocr/PPOCRLabel/resources/icons/zoom-out.png'\nadding 'paddleocr/PPOCRLabel/resources/icons/zoom.png'\nadding 'paddleocr/PPOCRLabel/resources/strings/strings-zh-CN.properties'\nadding 'paddleocr/PPOCRLabel/resources/strings/strings.properties'\nadding 'paddleocr/StyleText/README.md'\nadding 'paddleocr/StyleText/README_ch.md'\nadding 'paddleocr/StyleText/__init__.py'\nadding 'paddleocr/StyleText/arch/__init__.py'\nadding 'paddleocr/StyleText/arch/base_module.py'\nadding 'paddleocr/StyleText/arch/decoder.py'\nadding 'paddleocr/StyleText/arch/encoder.py'\nadding 'paddleocr/StyleText/arch/spectral_norm.py'\nadding 'paddleocr/StyleText/arch/style_text_rec.py'\nadding 'paddleocr/StyleText/configs/config.yml'\nadding 'paddleocr/StyleText/configs/dataset_config.yml'\nadding 'paddleocr/StyleText/doc/images/1.png'\nadding 'paddleocr/StyleText/doc/images/10.png'\nadding 'paddleocr/StyleText/doc/images/11.png'\nadding 'paddleocr/StyleText/doc/images/12.png'\nadding 'paddleocr/StyleText/doc/images/2.png'\nadding 'paddleocr/StyleText/doc/images/3.png'\nadding 'paddleocr/StyleText/doc/images/4.jpg'\nadding 'paddleocr/StyleText/doc/images/5.png'\nadding 'paddleocr/StyleText/doc/images/6.png'\nadding 'paddleocr/StyleText/doc/images/7.jpg'\nadding 'paddleocr/StyleText/doc/images/8.jpg'\nadding 'paddleocr/StyleText/doc/images/9.png'\nadding 'paddleocr/StyleText/engine/__init__.py'\nadding 'paddleocr/StyleText/engine/corpus_generators.py'\nadding 'paddleocr/StyleText/engine/predictors.py'\nadding 'paddleocr/StyleText/engine/style_samplers.py'\nadding 'paddleocr/StyleText/engine/synthesisers.py'\nadding 'paddleocr/StyleText/engine/text_drawers.py'\nadding 'paddleocr/StyleText/engine/writers.py'\nadding 'paddleocr/StyleText/examples/image_list.txt'\nadding 'paddleocr/StyleText/examples/corpus/example.txt'\nadding 'paddleocr/StyleText/examples/style_images/1.jpg'\nadding 'paddleocr/StyleText/examples/style_images/2.jpg'\nadding 'paddleocr/StyleText/fonts/ch_standard.ttf'\nadding 'paddleocr/StyleText/fonts/en_standard.ttf'\nadding 'paddleocr/StyleText/fonts/ko_standard.ttf'\nadding 'paddleocr/StyleText/tools/__init__.py'\nadding 'paddleocr/StyleText/tools/synth_dataset.py'\nadding 'paddleocr/StyleText/tools/synth_image.py'\nadding 'paddleocr/StyleText/utils/__init__.py'\nadding 'paddleocr/StyleText/utils/config.py'\nadding 'paddleocr/StyleText/utils/load_params.py'\nadding 'paddleocr/StyleText/utils/logging.py'\nadding 'paddleocr/StyleText/utils/math_functions.py'\nadding 'paddleocr/StyleText/utils/sys_funcs.py'\nadding 'paddleocr/configs/cls/cls_mv3.yml'\nadding 'paddleocr/configs/det/det_mv3_db.yml'\nadding 'paddleocr/configs/det/det_mv3_east.yml'\nadding 'paddleocr/configs/det/det_r50_vd_db.yml'\nadding 'paddleocr/configs/det/det_r50_vd_east.yml'\nadding 'paddleocr/configs/det/det_r50_vd_sast_icdar15.yml'\nadding 'paddleocr/configs/det/det_r50_vd_sast_totaltext.yml'\nadding 'paddleocr/configs/det/ch_ppocr_v2.0/ch_det_mv3_db_v2.0.yml'\nadding 'paddleocr/configs/det/ch_ppocr_v2.0/ch_det_res18_db_v2.0.yml'\nadding 'paddleocr/configs/e2e/e2e_r50_vd_pg.yml'\nadding 'paddleocr/configs/rec/rec_icdar15_train.yml'\nadding 'paddleocr/configs/rec/rec_mv3_none_bilstm_ctc.yml'\nadding 'paddleocr/configs/rec/rec_mv3_none_none_ctc.yml'\nadding 'paddleocr/configs/rec/rec_mv3_tps_bilstm_att.yml'\nadding 'paddleocr/configs/rec/rec_mv3_tps_bilstm_ctc.yml'\nadding 'paddleocr/configs/rec/rec_r34_vd_none_bilstm_ctc.yml'\nadding 'paddleocr/configs/rec/rec_r34_vd_none_none_ctc.yml'\nadding 'paddleocr/configs/rec/rec_r34_vd_tps_bilstm_att.yml'\nadding 'paddleocr/configs/rec/rec_r34_vd_tps_bilstm_ctc.yml'\nadding 'paddleocr/configs/rec/rec_r50_fpn_srn.yml'\nadding 'paddleocr/configs/rec/ch_ppocr_v2.0/rec_chinese_common_train_v2.0.yml'\nadding 'paddleocr/configs/rec/ch_ppocr_v2.0/rec_chinese_lite_train_v2.0.yml'\nadding 'paddleocr/configs/rec/multi_language/generate_multi_language_configs.py'\nadding 'paddleocr/configs/rec/multi_language/rec_arabic_lite_train.yml'\nadding 'paddleocr/configs/rec/multi_language/rec_cyrillic_lite_train.yml'\nadding 'paddleocr/configs/rec/multi_language/rec_devanagari_lite_train.yml'\nadding 'paddleocr/configs/rec/multi_language/rec_en_number_lite_train.yml'\nadding 'paddleocr/configs/rec/multi_language/rec_french_lite_train.yml'\nadding 'paddleocr/configs/rec/multi_language/rec_german_lite_train.yml'\nadding 'paddleocr/configs/rec/multi_language/rec_japan_lite_train.yml'\nadding 'paddleocr/configs/rec/multi_language/rec_korean_lite_train.yml'\nadding 'paddleocr/configs/rec/multi_language/rec_latin_lite_train.yml'\nadding 'paddleocr/configs/rec/multi_language/rec_multi_language_lite_train.yml'\nadding 'paddleocr/deploy/android_demo/.gitignore'\nadding 'paddleocr/deploy/android_demo/README.md'\nadding 'paddleocr/deploy/android_demo/build.gradle'\nadding 'paddleocr/deploy/android_demo/gradle.properties'\nadding 'paddleocr/deploy/android_demo/gradlew'\nadding 'paddleocr/deploy/android_demo/gradlew.bat'\nadding 'paddleocr/deploy/android_demo/settings.gradle'\nadding 'paddleocr/deploy/android_demo/app/.gitignore'\nadding 'paddleocr/deploy/android_demo/app/build.gradle'\nadding 'paddleocr/deploy/android_demo/app/proguard-rules.pro'\nadding 'paddleocr/deploy/android_demo/app/src/androidTest/java/com/baidu/paddle/lite/demo/ocr/ExampleInstrumentedTest.java'\nadding 'paddleocr/deploy/android_demo/app/src/main/AndroidManifest.xml'\nadding 'paddleocr/deploy/android_demo/app/src/main/assets/images/0.jpg'\nadding 'paddleocr/deploy/android_demo/app/src/main/assets/images/180.jpg'\nadding 'paddleocr/deploy/android_demo/app/src/main/assets/images/270.jpg'\nadding 'paddleocr/deploy/android_demo/app/src/main/assets/images/90.jpg'\nadding 'paddleocr/deploy/android_demo/app/src/main/cpp/CMakeLists.txt'\nadding 'paddleocr/deploy/android_demo/app/src/main/cpp/common.h'\nadding 'paddleocr/deploy/android_demo/app/src/main/cpp/native.cpp'\nadding 'paddleocr/deploy/android_demo/app/src/main/cpp/native.h'\nadding 'paddleocr/deploy/android_demo/app/src/main/cpp/ocr_clipper.cpp'\nadding 'paddleocr/deploy/android_demo/app/src/main/cpp/ocr_clipper.hpp'\nadding 'paddleocr/deploy/android_demo/app/src/main/cpp/ocr_cls_process.cpp'\nadding 'paddleocr/deploy/android_demo/app/src/main/cpp/ocr_cls_process.h'\nadding 'paddleocr/deploy/android_demo/app/src/main/cpp/ocr_crnn_process.cpp'\nadding 'paddleocr/deploy/android_demo/app/src/main/cpp/ocr_crnn_process.h'\nadding 'paddleocr/deploy/android_demo/app/src/main/cpp/ocr_db_post_process.cpp'\nadding 'paddleocr/deploy/android_demo/app/src/main/cpp/ocr_db_post_process.h'\nadding 'paddleocr/deploy/android_demo/app/src/main/cpp/ocr_ppredictor.cpp'\nadding 'paddleocr/deploy/android_demo/app/src/main/cpp/ocr_ppredictor.h'\nadding 'paddleocr/deploy/android_demo/app/src/main/cpp/ppredictor.cpp'\nadding 'paddleocr/deploy/android_demo/app/src/main/cpp/ppredictor.h'\nadding 'paddleocr/deploy/android_demo/app/src/main/cpp/predictor_input.cpp'\nadding 'paddleocr/deploy/android_demo/app/src/main/cpp/predictor_input.h'\nadding 'paddleocr/deploy/android_demo/app/src/main/cpp/predictor_output.cpp'\nadding 'paddleocr/deploy/android_demo/app/src/main/cpp/predictor_output.h'\nadding 'paddleocr/deploy/android_demo/app/src/main/cpp/preprocess.cpp'\nadding 'paddleocr/deploy/android_demo/app/src/main/cpp/preprocess.h'\nadding 'paddleocr/deploy/android_demo/app/src/main/java/com/baidu/paddle/lite/demo/ocr/AppCompatPreferenceActivity.java'\nadding 'paddleocr/deploy/android_demo/app/src/main/java/com/baidu/paddle/lite/demo/ocr/MainActivity.java'\nadding 'paddleocr/deploy/android_demo/app/src/main/java/com/baidu/paddle/lite/demo/ocr/MiniActivity.java'\nadding 'paddleocr/deploy/android_demo/app/src/main/java/com/baidu/paddle/lite/demo/ocr/OCRPredictorNative.java'\nadding 'paddleocr/deploy/android_demo/app/src/main/java/com/baidu/paddle/lite/demo/ocr/OcrResultModel.java'\nadding 'paddleocr/deploy/android_demo/app/src/main/java/com/baidu/paddle/lite/demo/ocr/Predictor.java'\nadding 'paddleocr/deploy/android_demo/app/src/main/java/com/baidu/paddle/lite/demo/ocr/SettingsActivity.java'\nadding 'paddleocr/deploy/android_demo/app/src/main/java/com/baidu/paddle/lite/demo/ocr/Utils.java'\nadding 'paddleocr/deploy/android_demo/app/src/main/res/drawable/ic_launcher_background.xml'\nadding 'paddleocr/deploy/android_demo/app/src/main/res/drawable-v24/ic_launcher_foreground.xml'\nadding 'paddleocr/deploy/android_demo/app/src/main/res/layout/activity_main.xml'\nadding 'paddleocr/deploy/android_demo/app/src/main/res/layout/activity_mini.xml'\nadding 'paddleocr/deploy/android_demo/app/src/main/res/menu/menu_action_options.xml'\nadding 'paddleocr/deploy/android_demo/app/src/main/res/mipmap-anydpi-v26/ic_launcher.xml'\nadding 'paddleocr/deploy/android_demo/app/src/main/res/mipmap-anydpi-v26/ic_launcher_round.xml'\nadding 'paddleocr/deploy/android_demo/app/src/main/res/mipmap-hdpi/ic_launcher.png'\nadding 'paddleocr/deploy/android_demo/app/src/main/res/mipmap-hdpi/ic_launcher_round.png'\nadding 'paddleocr/deploy/android_demo/app/src/main/res/mipmap-mdpi/ic_launcher.png'\nadding 'paddleocr/deploy/android_demo/app/src/main/res/mipmap-mdpi/ic_launcher_round.png'\nadding 'paddleocr/deploy/android_demo/app/src/main/res/mipmap-xhdpi/ic_launcher.png'\nadding 'paddleocr/deploy/android_demo/app/src/main/res/mipmap-xhdpi/ic_launcher_round.png'\nadding 'paddleocr/deploy/android_demo/app/src/main/res/mipmap-xxhdpi/ic_launcher.png'\nadding 'paddleocr/deploy/android_demo/app/src/main/res/mipmap-xxhdpi/ic_launcher_round.png'\nadding 'paddleocr/deploy/android_demo/app/src/main/res/mipmap-xxxhdpi/ic_launcher.png'\nadding 'paddleocr/deploy/android_demo/app/src/main/res/mipmap-xxxhdpi/ic_launcher_round.png'\nadding 'paddleocr/deploy/android_demo/app/src/main/res/values/arrays.xml'\nadding 'paddleocr/deploy/android_demo/app/src/main/res/values/colors.xml'\nadding 'paddleocr/deploy/android_demo/app/src/main/res/values/strings.xml'\nadding 'paddleocr/deploy/android_demo/app/src/main/res/values/styles.xml'\nadding 'paddleocr/deploy/android_demo/app/src/main/res/xml/file_paths.xml'\nadding 'paddleocr/deploy/android_demo/app/src/main/res/xml/settings.xml'\nadding 'paddleocr/deploy/android_demo/app/src/test/java/com/baidu/paddle/lite/demo/ocr/ExampleUnitTest.java'\nadding 'paddleocr/deploy/android_demo/gradle/wrapper/gradle-wrapper.jar'\nadding 'paddleocr/deploy/android_demo/gradle/wrapper/gradle-wrapper.properties'\nadding 'paddleocr/deploy/cpp_infer/CMakeLists.txt'\nadding 'paddleocr/deploy/cpp_infer/readme.md'\nadding 'paddleocr/deploy/cpp_infer/readme_en.md'\nadding 'paddleocr/deploy/cpp_infer/docs/windows_vs2019_build.md'\nadding 'paddleocr/deploy/cpp_infer/imgs/cpp_infer_pred_12.png'\nadding 'paddleocr/deploy/cpp_infer/include/clipper.h'\nadding 'paddleocr/deploy/cpp_infer/include/config.h'\nadding 'paddleocr/deploy/cpp_infer/include/ocr_cls.h'\nadding 'paddleocr/deploy/cpp_infer/include/ocr_det.h'\nadding 'paddleocr/deploy/cpp_infer/include/ocr_rec.h'\nadding 'paddleocr/deploy/cpp_infer/include/postprocess_op.h'\nadding 'paddleocr/deploy/cpp_infer/include/preprocess_op.h'\nadding 'paddleocr/deploy/cpp_infer/include/utility.h'\nadding 'paddleocr/deploy/cpp_infer/src/clipper.cpp'\nadding 'paddleocr/deploy/cpp_infer/src/config.cpp'\nadding 'paddleocr/deploy/cpp_infer/src/main.cpp'\nadding 'paddleocr/deploy/cpp_infer/src/ocr_cls.cpp'\nadding 'paddleocr/deploy/cpp_infer/src/ocr_det.cpp'\nadding 'paddleocr/deploy/cpp_infer/src/ocr_rec.cpp'\nadding 'paddleocr/deploy/cpp_infer/src/postprocess_op.cpp'\nadding 'paddleocr/deploy/cpp_infer/src/preprocess_op.cpp'\nadding 'paddleocr/deploy/cpp_infer/src/utility.cpp'\nadding 'paddleocr/deploy/cpp_infer/tools/build.sh'\nadding 'paddleocr/deploy/cpp_infer/tools/config.txt'\nadding 'paddleocr/deploy/cpp_infer/tools/run.sh'\nadding 'paddleocr/deploy/docker/hubserving/README.md'\nadding 'paddleocr/deploy/docker/hubserving/README_cn.md'\nadding 'paddleocr/deploy/docker/hubserving/sample_request.txt'\nadding 'paddleocr/deploy/docker/hubserving/cpu/Dockerfile'\nadding 'paddleocr/deploy/docker/hubserving/gpu/Dockerfile'\nadding 'paddleocr/deploy/hubserving/readme.md'\nadding 'paddleocr/deploy/hubserving/readme_en.md'\nadding 'paddleocr/deploy/hubserving/ocr_cls/__init__.py'\nadding 'paddleocr/deploy/hubserving/ocr_cls/config.json'\nadding 'paddleocr/deploy/hubserving/ocr_cls/module.py'\nadding 'paddleocr/deploy/hubserving/ocr_cls/params.py'\nadding 'paddleocr/deploy/hubserving/ocr_det/__init__.py'\nadding 'paddleocr/deploy/hubserving/ocr_det/config.json'\nadding 'paddleocr/deploy/hubserving/ocr_det/module.py'\nadding 'paddleocr/deploy/hubserving/ocr_det/params.py'\nadding 'paddleocr/deploy/hubserving/ocr_rec/__init__.py'\nadding 'paddleocr/deploy/hubserving/ocr_rec/config.json'\nadding 'paddleocr/deploy/hubserving/ocr_rec/module.py'\nadding 'paddleocr/deploy/hubserving/ocr_rec/params.py'\nadding 'paddleocr/deploy/hubserving/ocr_system/__init__.py'\nadding 'paddleocr/deploy/hubserving/ocr_system/config.json'\nadding 'paddleocr/deploy/hubserving/ocr_system/module.py'\nadding 'paddleocr/deploy/hubserving/ocr_system/params.py'\nadding 'paddleocr/deploy/lite/Makefile'\nadding 'paddleocr/deploy/lite/cls_process.cc'\nadding 'paddleocr/deploy/lite/cls_process.h'\nadding 'paddleocr/deploy/lite/config.txt'\nadding 'paddleocr/deploy/lite/crnn_process.cc'\nadding 'paddleocr/deploy/lite/crnn_process.h'\nadding 'paddleocr/deploy/lite/db_post_process.cc'\nadding 'paddleocr/deploy/lite/db_post_process.h'\nadding 'paddleocr/deploy/lite/ocr_db_crnn.cc'\nadding 'paddleocr/deploy/lite/prepare.sh'\nadding 'paddleocr/deploy/lite/readme.md'\nadding 'paddleocr/deploy/lite/readme_en.md'\nadding 'paddleocr/deploy/lite/imgs/lite_demo.png'\nadding 'paddleocr/deploy/pdserving/README.md'\nadding 'paddleocr/deploy/pdserving/README_CN.md'\nadding 'paddleocr/deploy/pdserving/__init__.py'\nadding 'paddleocr/deploy/pdserving/config.yml'\nadding 'paddleocr/deploy/pdserving/ocr_reader.py'\nadding 'paddleocr/deploy/pdserving/pipeline_http_client.py'\nadding 'paddleocr/deploy/pdserving/pipeline_rpc_client.py'\nadding 'paddleocr/deploy/pdserving/web_service.py'\nadding 'paddleocr/deploy/pdserving/imgs/cpp_infer_pred_12.png'\nadding 'paddleocr/deploy/pdserving/imgs/demo.png'\nadding 'paddleocr/deploy/pdserving/imgs/results.png'\nadding 'paddleocr/deploy/pdserving/imgs/start_server.png'\nadding 'paddleocr/deploy/pdserving/win/__init__.py'\nadding 'paddleocr/deploy/pdserving/win/ocr_web_client.py'\nadding 'paddleocr/deploy/pdserving/win/ocr_web_server.py'\nadding 'paddleocr/deploy/slim/prune/README.md'\nadding 'paddleocr/deploy/slim/prune/README_en.md'\nadding 'paddleocr/deploy/slim/prune/export_prune_model.py'\nadding 'paddleocr/deploy/slim/prune/sensitivity_anal.py'\nadding 'paddleocr/deploy/slim/quantization/README.md'\nadding 'paddleocr/deploy/slim/quantization/README_en.md'\nadding 'paddleocr/deploy/slim/quantization/export_model.py'\nadding 'paddleocr/deploy/slim/quantization/quant.py'\nadding 'paddleocr/doc/PPOCR.pdf'\nadding 'paddleocr/doc/joinus.PNG'\nadding 'paddleocr/doc/ocr-android-easyedge.png'\nadding 'paddleocr/doc/pgnet_framework.png'\nadding 'paddleocr/doc/ppocr_framework.png'\nadding 'paddleocr/doc/datasets/ArT.jpg'\nadding 'paddleocr/doc/datasets/CASIA_0.jpg'\nadding 'paddleocr/doc/datasets/LSVT_1.jpg'\nadding 'paddleocr/doc/datasets/LSVT_2.jpg'\nadding 'paddleocr/doc/datasets/VoTT.jpg'\nadding 'paddleocr/doc/datasets/captcha_demo.png'\nadding 'paddleocr/doc/datasets/ccpd_demo.png'\nadding 'paddleocr/doc/datasets/ch_doc1.jpg'\nadding 'paddleocr/doc/datasets/ch_doc2.jpg'\nadding 'paddleocr/doc/datasets/ch_doc3.jpg'\nadding 'paddleocr/doc/datasets/ch_street_rec_1.png'\nadding 'paddleocr/doc/datasets/ch_street_rec_2.png'\nadding 'paddleocr/doc/datasets/cmb_demo.jpg'\nadding 'paddleocr/doc/datasets/doc.jpg'\nadding 'paddleocr/doc/datasets/labelimg.jpg'\nadding 'paddleocr/doc/datasets/labelme.jpg'\nadding 'paddleocr/doc/datasets/nist_demo.png'\nadding 'paddleocr/doc/datasets/rctw.jpg'\nadding 'paddleocr/doc/datasets/roLabelImg.png'\nadding 'paddleocr/doc/demo/build.png'\nadding 'paddleocr/doc/demo/error.png'\nadding 'paddleocr/doc/demo/proxy.png'\nadding 'paddleocr/doc/doc_ch/FAQ.md'\nadding 'paddleocr/doc/doc_ch/add_new_algorithm.md'\nadding 'paddleocr/doc/doc_ch/algorithm_overview.md'\nadding 'paddleocr/doc/doc_ch/android_demo.md'\nadding 'paddleocr/doc/doc_ch/angle_class.md'\nadding 'paddleocr/doc/doc_ch/benchmark.md'\nadding 'paddleocr/doc/doc_ch/config.md'\nadding 'paddleocr/doc/doc_ch/customize.md'\nadding 'paddleocr/doc/doc_ch/data_annotation.md'\nadding 'paddleocr/doc/doc_ch/data_synthesis.md'\nadding 'paddleocr/doc/doc_ch/datasets.md'\nadding 'paddleocr/doc/doc_ch/detection.md'\nadding 'paddleocr/doc/doc_ch/framework.png'\nadding 'paddleocr/doc/doc_ch/handwritten_datasets.md'\nadding 'paddleocr/doc/doc_ch/inference.md'\nadding 'paddleocr/doc/doc_ch/installation.md'\nadding 'paddleocr/doc/doc_ch/models_list.md'\nadding 'paddleocr/doc/doc_ch/multi_languages.md'\nadding 'paddleocr/doc/doc_ch/pgnet.md'\nadding 'paddleocr/doc/doc_ch/quickstart.md'\nadding 'paddleocr/doc/doc_ch/recognition.md'\nadding 'paddleocr/doc/doc_ch/reference.md'\nadding 'paddleocr/doc/doc_ch/serving_inference.md'\nadding 'paddleocr/doc/doc_ch/tree.md'\nadding 'paddleocr/doc/doc_ch/update.md'\nadding 'paddleocr/doc/doc_ch/vertical_and_multilingual_datasets.md'\nadding 'paddleocr/doc/doc_ch/visualization.md'\nadding 'paddleocr/doc/doc_ch/whl.md'\nadding 'paddleocr/doc/doc_en/FAQ_en.md'\nadding 'paddleocr/doc/doc_en/add_new_algorithm_en.md'\nadding 'paddleocr/doc/doc_en/algorithm_overview_en.md'\nadding 'paddleocr/doc/doc_en/android_demo_en.md'\nadding 'paddleocr/doc/doc_en/angle_class_en.md'\nadding 'paddleocr/doc/doc_en/benchmark_en.md'\nadding 'paddleocr/doc/doc_en/config_en.md'\nadding 'paddleocr/doc/doc_en/customize_en.md'\nadding 'paddleocr/doc/doc_en/data_annotation_en.md'\nadding 'paddleocr/doc/doc_en/data_synthesis_en.md'\nadding 'paddleocr/doc/doc_en/datasets_en.md'\nadding 'paddleocr/doc/doc_en/detection_en.md'\nadding 'paddleocr/doc/doc_en/handwritten_datasets_en.md'\nadding 'paddleocr/doc/doc_en/inference_en.md'\nadding 'paddleocr/doc/doc_en/installation_en.md'\nadding 'paddleocr/doc/doc_en/models_list_en.md'\nadding 'paddleocr/doc/doc_en/multi_languages_en.md'\nadding 'paddleocr/doc/doc_en/pgnet_en.md'\nadding 'paddleocr/doc/doc_en/quickstart_en.md'\nadding 'paddleocr/doc/doc_en/recognition_en.md'\nadding 'paddleocr/doc/doc_en/reference_en.md'\nadding 'paddleocr/doc/doc_en/tree_en.md'\nadding 'paddleocr/doc/doc_en/tricks_en.md'\nadding 'paddleocr/doc/doc_en/update_en.md'\nadding 'paddleocr/doc/doc_en/vertical_and_multilingual_datasets_en.md'\nadding 'paddleocr/doc/doc_en/visualization_en.md'\nadding 'paddleocr/doc/doc_en/whl_en.md'\nadding 'paddleocr/doc/fonts/arabic.ttf'\nadding 'paddleocr/doc/fonts/chinese_cht.ttf'\nadding 'paddleocr/doc/fonts/cyrillic.ttf'\nadding 'paddleocr/doc/fonts/french.ttf'\nadding 'paddleocr/doc/fonts/german.ttf'\nadding 'paddleocr/doc/fonts/hindi.ttf'\nadding 'paddleocr/doc/fonts/japan.ttc'\nadding 'paddleocr/doc/fonts/kannada.ttf'\nadding 'paddleocr/doc/fonts/korean.ttf'\nadding 'paddleocr/doc/fonts/latin.ttf'\nadding 'paddleocr/doc/fonts/marathi.ttf'\nadding 'paddleocr/doc/fonts/nepali.ttf'\nadding 'paddleocr/doc/fonts/persian.ttf'\nadding 'paddleocr/doc/fonts/simfang.ttf'\nadding 'paddleocr/doc/fonts/spanish.ttf'\nadding 'paddleocr/doc/fonts/tamil.ttf'\nadding 'paddleocr/doc/fonts/telugu.ttf'\nadding 'paddleocr/doc/fonts/urdu.ttf'\nadding 'paddleocr/doc/fonts/uyghur.ttf'\nadding 'paddleocr/doc/imgs/00006737.jpg'\nadding 'paddleocr/doc/imgs/00009282.jpg'\nadding 'paddleocr/doc/imgs/00015504.jpg'\nadding 'paddleocr/doc/imgs/00018069.jpg'\nadding 'paddleocr/doc/imgs/00056221.jpg'\nadding 'paddleocr/doc/imgs/00057937.jpg'\nadding 'paddleocr/doc/imgs/00059985.jpg'\nadding 'paddleocr/doc/imgs/00077949.jpg'\nadding 'paddleocr/doc/imgs/00111002.jpg'\nadding 'paddleocr/doc/imgs/00207393.jpg'\nadding 'paddleocr/doc/imgs/1.jpg'\nadding 'paddleocr/doc/imgs/11.jpg'\nadding 'paddleocr/doc/imgs/12.jpg'\nadding 'paddleocr/doc/imgs/french_0.jpg'\nadding 'paddleocr/doc/imgs/ger_1.jpg'\nadding 'paddleocr/doc/imgs/ger_2.jpg'\nadding 'paddleocr/doc/imgs/japan_1.jpg'\nadding 'paddleocr/doc/imgs/japan_2.jpg'\nadding 'paddleocr/doc/imgs/korean_1.jpg'\nadding 'paddleocr/doc/imgs/model_prod_flow_ch.png'\nadding 'paddleocr/doc/imgs_en/254.jpg'\nadding 'paddleocr/doc/imgs_en/img623.jpg'\nadding 'paddleocr/doc/imgs_en/img_10.jpg'\nadding 'paddleocr/doc/imgs_en/img_11.jpg'\nadding 'paddleocr/doc/imgs_en/img_12.jpg'\nadding 'paddleocr/doc/imgs_en/img_195.jpg'\nadding 'paddleocr/doc/imgs_en/model_prod_flow_en.png'\nadding 'paddleocr/doc/imgs_results/angle_class_example.jpg'\nadding 'paddleocr/doc/imgs_results/det_res_00018069.jpg'\nadding 'paddleocr/doc/imgs_results/det_res_img623_sast.jpg'\nadding 'paddleocr/doc/imgs_results/det_res_img_10_db.jpg'\nadding 'paddleocr/doc/imgs_results/det_res_img_10_east.jpg'\nadding 'paddleocr/doc/imgs_results/det_res_img_10_sast.jpg'\nadding 'paddleocr/doc/imgs_results/e2e_res_img293_pgnet.png'\nadding 'paddleocr/doc/imgs_results/e2e_res_img295_pgnet.png'\nadding 'paddleocr/doc/imgs_results/e2e_res_img623_pgnet.jpg'\nadding 'paddleocr/doc/imgs_results/e2e_res_img_10_pgnet.jpg'\nadding 'paddleocr/doc/imgs_results/french_0.jpg'\nadding 'paddleocr/doc/imgs_results/img_10_east_starnet.jpg'\nadding 'paddleocr/doc/imgs_results/korean.jpg'\nadding 'paddleocr/doc/imgs_results/system_res_00018069.jpg'\nadding 'paddleocr/doc/imgs_results/ch_ppocr_mobile_v2.0/00006737.jpg'\nadding 'paddleocr/doc/imgs_results/ch_ppocr_mobile_v2.0/00009282.jpg'\nadding 'paddleocr/doc/imgs_results/ch_ppocr_mobile_v2.0/00015504.jpg'\nadding 'paddleocr/doc/imgs_results/ch_ppocr_mobile_v2.0/00018069.jpg'\nadding 'paddleocr/doc/imgs_results/ch_ppocr_mobile_v2.0/00056221.jpg'\nadding 'paddleocr/doc/imgs_results/ch_ppocr_mobile_v2.0/00057937.jpg'\nadding 'paddleocr/doc/imgs_results/ch_ppocr_mobile_v2.0/00059985.jpg'\nadding 'paddleocr/doc/imgs_results/ch_ppocr_mobile_v2.0/00077949.jpg'\nadding 'paddleocr/doc/imgs_results/ch_ppocr_mobile_v2.0/00111002.jpg'\nadding 'paddleocr/doc/imgs_results/ch_ppocr_mobile_v2.0/00207393.jpg'\nadding 'paddleocr/doc/imgs_results/ch_ppocr_mobile_v2.0/img_12.jpg'\nadding 'paddleocr/doc/imgs_results/ch_ppocr_mobile_v2.0/rotate_00052204.jpg'\nadding 'paddleocr/doc/imgs_results/ch_ppocr_mobile_v2.0/test_add_91.jpg'\nadding 'paddleocr/doc/imgs_results/multi_lang/arabic_0.jpg'\nadding 'paddleocr/doc/imgs_results/multi_lang/en_1.jpg'\nadding 'paddleocr/doc/imgs_results/multi_lang/en_2.jpg'\nadding 'paddleocr/doc/imgs_results/multi_lang/en_3.jpg'\nadding 'paddleocr/doc/imgs_results/multi_lang/french_0.jpg'\nadding 'paddleocr/doc/imgs_results/multi_lang/img_01.jpg'\nadding 'paddleocr/doc/imgs_results/multi_lang/img_02.jpg'\nadding 'paddleocr/doc/imgs_results/multi_lang/img_12.jpg'\nadding 'paddleocr/doc/imgs_results/multi_lang/japan_2.jpg'\nadding 'paddleocr/doc/imgs_results/multi_lang/korean_0.jpg'\nadding 'paddleocr/doc/imgs_results/whl/11_det.jpg'\nadding 'paddleocr/doc/imgs_results/whl/11_det_rec.jpg'\nadding 'paddleocr/doc/imgs_results/whl/12_det.jpg'\nadding 'paddleocr/doc/imgs_results/whl/12_det_rec.jpg'\nadding 'paddleocr/doc/imgs_words/arabic/ar_1.jpg'\nadding 'paddleocr/doc/imgs_words/arabic/ar_2.jpg'\nadding 'paddleocr/doc/imgs_words/belarusian/be_1.jpg'\nadding 'paddleocr/doc/imgs_words/belarusian/be_2.jpg'\nadding 'paddleocr/doc/imgs_words/bulgarian/bg_1.jpg'\nadding 'paddleocr/doc/imgs_words/bulgarian/bg_2.jpg'\nadding 'paddleocr/doc/imgs_words/ch/word_1.jpg'\nadding 'paddleocr/doc/imgs_words/ch/word_2.jpg'\nadding 'paddleocr/doc/imgs_words/ch/word_3.jpg'\nadding 'paddleocr/doc/imgs_words/ch/word_4.jpg'\nadding 'paddleocr/doc/imgs_words/ch/word_5.jpg'\nadding 'paddleocr/doc/imgs_words/chinese_traditional/chinese_cht_1.png'\nadding 'paddleocr/doc/imgs_words/chinese_traditional/chinese_cht_2.png'\nadding 'paddleocr/doc/imgs_words/en/word_1.png'\nadding 'paddleocr/doc/imgs_words/en/word_2.png'\nadding 'paddleocr/doc/imgs_words/en/word_3.png'\nadding 'paddleocr/doc/imgs_words/en/word_4.png'\nadding 'paddleocr/doc/imgs_words/en/word_5.png'\nadding 'paddleocr/doc/imgs_words/french/1.jpg'\nadding 'paddleocr/doc/imgs_words/french/2.jpg'\nadding 'paddleocr/doc/imgs_words/german/1.jpg'\nadding 'paddleocr/doc/imgs_words/hindi/hi_1.jpg'\nadding 'paddleocr/doc/imgs_words/hindi/hi_2.jpg'\nadding 'paddleocr/doc/imgs_words/italian/it_1.jpg'\nadding 'paddleocr/doc/imgs_words/italian/it_2.jpg'\nadding 'paddleocr/doc/imgs_words/japan/1.jpg'\nadding 'paddleocr/doc/imgs_words/kannada/ka_1.jpg'\nadding 'paddleocr/doc/imgs_words/kannada/ka_2.jpg'\nadding 'paddleocr/doc/imgs_words/korean/1.jpg'\nadding 'paddleocr/doc/imgs_words/korean/2.jpg'\nadding 'paddleocr/doc/imgs_words/marathi/mr_1.jpg'\nadding 'paddleocr/doc/imgs_words/marathi/mr_2.jpg'\nadding 'paddleocr/doc/imgs_words/nepali/ne_1.jpg'\nadding 'paddleocr/doc/imgs_words/nepali/ne_2.jpg'\nadding 'paddleocr/doc/imgs_words/occitan/oc_1.jpg'\nadding 'paddleocr/doc/imgs_words/occitan/oc_2.jpg'\nadding 'paddleocr/doc/imgs_words/persian/fa_1.jpg'\nadding 'paddleocr/doc/imgs_words/persian/fa_2.jpg'\nadding 'paddleocr/doc/imgs_words/portuguese/pu_1.jpg'\nadding 'paddleocr/doc/imgs_words/portuguese/pu_2.jpg'\nadding 'paddleocr/doc/imgs_words/russia/ru_1.jpg'\nadding 'paddleocr/doc/imgs_words/russia/ru_2.jpg'\nadding 'paddleocr/doc/imgs_words/serbian_cyrillic/rsc_1.jpg'\nadding 'paddleocr/doc/imgs_words/serbian_cyrillic/rsc_2.jpg'\nadding 'paddleocr/doc/imgs_words/serbian_latin/rs_1.jpg'\nadding 'paddleocr/doc/imgs_words/serbian_latin/rs_2.jpg'\nadding 'paddleocr/doc/imgs_words/spanish/xi_1.jpg'\nadding 'paddleocr/doc/imgs_words/spanish/xi_2.jpg'\nadding 'paddleocr/doc/imgs_words/tamil/ta_1.jpg'\nadding 'paddleocr/doc/imgs_words/tamil/ta_2.jpg'\nadding 'paddleocr/doc/imgs_words/telugu/te_1.jpg'\nadding 'paddleocr/doc/imgs_words/telugu/te_2.jpg'\nadding 'paddleocr/doc/imgs_words/ukranian/uk_1.jpg'\nadding 'paddleocr/doc/imgs_words/ukranian/uk_2.jpg'\nadding 'paddleocr/doc/imgs_words/urdu/ur_1.jpg'\nadding 'paddleocr/doc/imgs_words/urdu/ur_2.jpg'\nadding 'paddleocr/doc/imgs_words/uyghur/ug_1.jpg'\nadding 'paddleocr/doc/imgs_words/uyghur/ug_2.jpg'\nadding 'paddleocr/doc/imgs_words_en/word_10.png'\nadding 'paddleocr/doc/imgs_words_en/word_116.png'\nadding 'paddleocr/doc/imgs_words_en/word_19.png'\nadding 'paddleocr/doc/imgs_words_en/word_201.png'\nadding 'paddleocr/doc/imgs_words_en/word_308.png'\nadding 'paddleocr/doc/imgs_words_en/word_336.png'\nadding 'paddleocr/doc/imgs_words_en/word_401.png'\nadding 'paddleocr/doc/imgs_words_en/word_461.png'\nadding 'paddleocr/doc/imgs_words_en/word_52.png'\nadding 'paddleocr/doc/imgs_words_en/word_545.png'\nadding 'paddleocr/doc/tricks/long_text_examples.jpg'\nadding 'paddleocr/paddleocr.egg-info/PKG-INFO'\nadding 'paddleocr/paddleocr.egg-info/SOURCES.txt'\nadding 'paddleocr/paddleocr.egg-info/dependency_links.txt'\nadding 'paddleocr/paddleocr.egg-info/entry_points.txt'\nadding 'paddleocr/paddleocr.egg-info/requires.txt'\nadding 'paddleocr/paddleocr.egg-info/top_level.txt'\nadding 'paddleocr/ppocr/__init__.py'\nadding 'paddleocr/ppocr/data/__init__.py'\nadding 'paddleocr/ppocr/data/lmdb_dataset.py'\nadding 'paddleocr/ppocr/data/pgnet_dataset.py'\nadding 'paddleocr/ppocr/data/simple_dataset.py'\nadding 'paddleocr/ppocr/data/imaug/__init__.py'\nadding 'paddleocr/ppocr/data/imaug/east_process.py'\nadding 'paddleocr/ppocr/data/imaug/iaa_augment.py'\nadding 'paddleocr/ppocr/data/imaug/label_ops.py'\nadding 'paddleocr/ppocr/data/imaug/make_border_map.py'\nadding 'paddleocr/ppocr/data/imaug/make_shrink_map.py'\nadding 'paddleocr/ppocr/data/imaug/operators.py'\nadding 'paddleocr/ppocr/data/imaug/pg_process.py'\nadding 'paddleocr/ppocr/data/imaug/randaugment.py'\nadding 'paddleocr/ppocr/data/imaug/random_crop_data.py'\nadding 'paddleocr/ppocr/data/imaug/rec_img_aug.py'\nadding 'paddleocr/ppocr/data/imaug/sast_process.py'\nadding 'paddleocr/ppocr/data/imaug/text_image_aug/__init__.py'\nadding 'paddleocr/ppocr/data/imaug/text_image_aug/augment.py'\nadding 'paddleocr/ppocr/data/imaug/text_image_aug/warp_mls.py'\nadding 'paddleocr/ppocr/losses/__init__.py'\nadding 'paddleocr/ppocr/losses/cls_loss.py'\nadding 'paddleocr/ppocr/losses/det_basic_loss.py'\nadding 'paddleocr/ppocr/losses/det_db_loss.py'\nadding 'paddleocr/ppocr/losses/det_east_loss.py'\nadding 'paddleocr/ppocr/losses/det_sast_loss.py'\nadding 'paddleocr/ppocr/losses/e2e_pg_loss.py'\nadding 'paddleocr/ppocr/losses/rec_att_loss.py'\nadding 'paddleocr/ppocr/losses/rec_ctc_loss.py'\nadding 'paddleocr/ppocr/losses/rec_srn_loss.py'\nadding 'paddleocr/ppocr/metrics/__init__.py'\nadding 'paddleocr/ppocr/metrics/cls_metric.py'\nadding 'paddleocr/ppocr/metrics/det_metric.py'\nadding 'paddleocr/ppocr/metrics/e2e_metric.py'\nadding 'paddleocr/ppocr/metrics/eval_det_iou.py'\nadding 'paddleocr/ppocr/metrics/rec_metric.py'\nadding 'paddleocr/ppocr/modeling/architectures/__init__.py'\nadding 'paddleocr/ppocr/modeling/architectures/base_model.py'\nadding 'paddleocr/ppocr/modeling/backbones/__init__.py'\nadding 'paddleocr/ppocr/modeling/backbones/det_mobilenet_v3.py'\nadding 'paddleocr/ppocr/modeling/backbones/det_resnet_vd.py'\nadding 'paddleocr/ppocr/modeling/backbones/det_resnet_vd_sast.py'\nadding 'paddleocr/ppocr/modeling/backbones/e2e_resnet_vd_pg.py'\nadding 'paddleocr/ppocr/modeling/backbones/rec_mobilenet_v3.py'\nadding 'paddleocr/ppocr/modeling/backbones/rec_resnet_fpn.py'\nadding 'paddleocr/ppocr/modeling/backbones/rec_resnet_vd.py'\nadding 'paddleocr/ppocr/modeling/heads/__init__.py'\nadding 'paddleocr/ppocr/modeling/heads/cls_head.py'\nadding 'paddleocr/ppocr/modeling/heads/det_db_head.py'\nadding 'paddleocr/ppocr/modeling/heads/det_east_head.py'\nadding 'paddleocr/ppocr/modeling/heads/det_sast_head.py'\nadding 'paddleocr/ppocr/modeling/heads/e2e_pg_head.py'\nadding 'paddleocr/ppocr/modeling/heads/rec_att_head.py'\nadding 'paddleocr/ppocr/modeling/heads/rec_ctc_head.py'\nadding 'paddleocr/ppocr/modeling/heads/rec_srn_head.py'\nadding 'paddleocr/ppocr/modeling/heads/self_attention.py'\nadding 'paddleocr/ppocr/modeling/necks/__init__.py'\nadding 'paddleocr/ppocr/modeling/necks/db_fpn.py'\nadding 'paddleocr/ppocr/modeling/necks/east_fpn.py'\nadding 'paddleocr/ppocr/modeling/necks/pg_fpn.py'\nadding 'paddleocr/ppocr/modeling/necks/rnn.py'\nadding 'paddleocr/ppocr/modeling/necks/sast_fpn.py'\nadding 'paddleocr/ppocr/modeling/transforms/__init__.py'\nadding 'paddleocr/ppocr/modeling/transforms/tps.py'\nadding 'paddleocr/ppocr/optimizer/__init__.py'\nadding 'paddleocr/ppocr/optimizer/learning_rate.py'\nadding 'paddleocr/ppocr/optimizer/lr_scheduler.py'\nadding 'paddleocr/ppocr/optimizer/optimizer.py'\nadding 'paddleocr/ppocr/optimizer/regularizer.py'\nadding 'paddleocr/ppocr/postprocess/__init__.py'\nadding 'paddleocr/ppocr/postprocess/cls_postprocess.py'\nadding 'paddleocr/ppocr/postprocess/db_postprocess.py'\nadding 'paddleocr/ppocr/postprocess/east_postprocess.py'\nadding 'paddleocr/ppocr/postprocess/locality_aware_nms.py'\nadding 'paddleocr/ppocr/postprocess/pg_postprocess.py'\nadding 'paddleocr/ppocr/postprocess/rec_postprocess.py'\nadding 'paddleocr/ppocr/postprocess/sast_postprocess.py'\nadding 'paddleocr/ppocr/utils/__init__.py'\nadding 'paddleocr/ppocr/utils/en_dict.txt'\nadding 'paddleocr/ppocr/utils/gen_label.py'\nadding 'paddleocr/ppocr/utils/ic15_dict.txt'\nadding 'paddleocr/ppocr/utils/logging.py'\nadding 'paddleocr/ppocr/utils/ppocr_keys_v1.txt'\nadding 'paddleocr/ppocr/utils/save_load.py'\nadding 'paddleocr/ppocr/utils/stats.py'\nadding 'paddleocr/ppocr/utils/utility.py'\nadding 'paddleocr/ppocr/utils/dict/ar_dict.txt'\nadding 'paddleocr/ppocr/utils/dict/arabic_dict.txt'\nadding 'paddleocr/ppocr/utils/dict/be_dict.txt'\nadding 'paddleocr/ppocr/utils/dict/bg_dict.txt'\nadding 'paddleocr/ppocr/utils/dict/chinese_cht_dict.txt'\nadding 'paddleocr/ppocr/utils/dict/cyrillic_dict.txt'\nadding 'paddleocr/ppocr/utils/dict/devanagari_dict.txt'\nadding 'paddleocr/ppocr/utils/dict/en_dict.txt'\nadding 'paddleocr/ppocr/utils/dict/fa_dict.txt'\nadding 'paddleocr/ppocr/utils/dict/french_dict.txt'\nadding 'paddleocr/ppocr/utils/dict/german_dict.txt'\nadding 'paddleocr/ppocr/utils/dict/hi_dict.txt'\nadding 'paddleocr/ppocr/utils/dict/it_dict.txt'\nadding 'paddleocr/ppocr/utils/dict/japan_dict.txt'\nadding 'paddleocr/ppocr/utils/dict/ka_dict.txt'\nadding 'paddleocr/ppocr/utils/dict/korean_dict.txt'\nadding 'paddleocr/ppocr/utils/dict/latin_dict.txt'\nadding 'paddleocr/ppocr/utils/dict/mr_dict.txt'\nadding 'paddleocr/ppocr/utils/dict/ne_dict.txt'\nadding 'paddleocr/ppocr/utils/dict/oc_dict.txt'\nadding 'paddleocr/ppocr/utils/dict/pu_dict.txt'\nadding 'paddleocr/ppocr/utils/dict/rs_dict.txt'\nadding 'paddleocr/ppocr/utils/dict/rsc_dict.txt'\nadding 'paddleocr/ppocr/utils/dict/ru_dict.txt'\nadding 'paddleocr/ppocr/utils/dict/ta_dict.txt'\nadding 'paddleocr/ppocr/utils/dict/te_dict.txt'\nadding 'paddleocr/ppocr/utils/dict/ug_dict.txt'\nadding 'paddleocr/ppocr/utils/dict/uk_dict.txt'\nadding 'paddleocr/ppocr/utils/dict/ur_dict.txt'\nadding 'paddleocr/ppocr/utils/dict/xi_dict.txt'\nadding 'paddleocr/ppocr/utils/e2e_metric/Deteval.py'\nadding 'paddleocr/ppocr/utils/e2e_metric/polygon_fast.py'\nadding 'paddleocr/ppocr/utils/e2e_utils/extract_batchsize.py'\nadding 'paddleocr/ppocr/utils/e2e_utils/extract_textpoint_fast.py'\nadding 'paddleocr/ppocr/utils/e2e_utils/extract_textpoint_slow.py'\nadding 'paddleocr/ppocr/utils/e2e_utils/pgnet_pp_utils.py'\nadding 'paddleocr/ppocr/utils/e2e_utils/visual.py'\nadding 'paddleocr/tools/eval.py'\nadding 'paddleocr/tools/export_model.py'\nadding 'paddleocr/tools/infer_cls.py'\nadding 'paddleocr/tools/infer_det.py'\nadding 'paddleocr/tools/infer_e2e.py'\nadding 'paddleocr/tools/infer_rec.py'\nadding 'paddleocr/tools/program.py'\nadding 'paddleocr/tools/test_hubserving.py'\nadding 'paddleocr/tools/train.py'\nadding 'paddleocr/tools/infer/predict_cls.py'\nadding 'paddleocr/tools/infer/predict_det.py'\nadding 'paddleocr/tools/infer/predict_e2e.py'\nadding 'paddleocr/tools/infer/predict_rec.py'\nadding 'paddleocr/tools/infer/predict_system.py'\nadding 'paddleocr/tools/infer/utility.py'\nadding 'paddleocr-2.0.6.dist-info/LICENSE'\nadding 'paddleocr-2.0.6.dist-info/METADATA'\nadding 'paddleocr-2.0.6.dist-info/WHEEL'\nadding 'paddleocr-2.0.6.dist-info/entry_points.txt'\nadding 'paddleocr-2.0.6.dist-info/top_level.txt'\nadding 'paddleocr-2.0.6.dist-info/RECORD'\nremoving build/bdist.linux-x86_64/wheel\n"
],
[
"!pip3 install dist/paddleocr-2.0.6-py3-none-any.whl # x.x.x is the version number of paddleocr",
"Requirement already satisfied: paddleocr==2.0.6 from file:///content/drive/My%20Drive/Reg_text/PaddleOCR/PaddleOCR/dist/paddleocr-2.0.6-py3-none-any.whl in /usr/local/lib/python3.7/dist-packages (2.0.6)\nRequirement already satisfied: visualdl in /usr/local/lib/python3.7/dist-packages (from paddleocr==2.0.6) (2.2.0)\nRequirement already satisfied: numpy in /usr/local/lib/python3.7/dist-packages (from paddleocr==2.0.6) (1.19.5)\nRequirement already satisfied: lmdb in /usr/local/lib/python3.7/dist-packages (from paddleocr==2.0.6) (0.99)\nRequirement already satisfied: opencv-contrib-python in /usr/local/lib/python3.7/dist-packages (from paddleocr==2.0.6) (4.1.2.30)\nRequirement already satisfied: imgaug==0.4.0 in /usr/local/lib/python3.7/dist-packages (from paddleocr==2.0.6) (0.4.0)\nRequirement already satisfied: shapely in /usr/local/lib/python3.7/dist-packages (from paddleocr==2.0.6) (1.7.1)\nRequirement already satisfied: opencv-python==4.2.0.32 in /usr/local/lib/python3.7/dist-packages (from paddleocr==2.0.6) (4.2.0.32)\nRequirement already satisfied: scikit-image==0.17.2 in /usr/local/lib/python3.7/dist-packages (from paddleocr==2.0.6) (0.17.2)\nRequirement already satisfied: python-Levenshtein in /usr/local/lib/python3.7/dist-packages (from paddleocr==2.0.6) (0.12.2)\nRequirement already satisfied: pyclipper in /usr/local/lib/python3.7/dist-packages (from paddleocr==2.0.6) (1.2.1)\nRequirement already satisfied: tqdm in /usr/local/lib/python3.7/dist-packages (from paddleocr==2.0.6) (4.41.1)\nRequirement already satisfied: flask>=1.1.1 in /usr/local/lib/python3.7/dist-packages (from visualdl->paddleocr==2.0.6) (1.1.4)\nRequirement already satisfied: shellcheck-py in /usr/local/lib/python3.7/dist-packages (from visualdl->paddleocr==2.0.6) (0.7.2.1)\nRequirement already satisfied: Pillow>=7.0.0 in /usr/local/lib/python3.7/dist-packages (from visualdl->paddleocr==2.0.6) (7.1.2)\nRequirement already satisfied: matplotlib in /usr/local/lib/python3.7/dist-packages (from visualdl->paddleocr==2.0.6) (3.2.2)\nRequirement already satisfied: requests in /usr/local/lib/python3.7/dist-packages (from visualdl->paddleocr==2.0.6) (2.23.0)\nRequirement already satisfied: six>=1.14.0 in /usr/local/lib/python3.7/dist-packages (from visualdl->paddleocr==2.0.6) (1.15.0)\nRequirement already satisfied: flake8>=3.7.9 in /usr/local/lib/python3.7/dist-packages (from visualdl->paddleocr==2.0.6) (3.9.2)\nRequirement already satisfied: bce-python-sdk in /usr/local/lib/python3.7/dist-packages (from visualdl->paddleocr==2.0.6) (0.8.60)\nRequirement already satisfied: Flask-Babel>=1.0.0 in /usr/local/lib/python3.7/dist-packages (from visualdl->paddleocr==2.0.6) (2.0.0)\nRequirement already satisfied: protobuf>=3.11.0 in /usr/local/lib/python3.7/dist-packages (from visualdl->paddleocr==2.0.6) (3.12.4)\nRequirement already satisfied: pre-commit in /usr/local/lib/python3.7/dist-packages (from visualdl->paddleocr==2.0.6) (2.13.0)\nRequirement already satisfied: pandas in /usr/local/lib/python3.7/dist-packages (from visualdl->paddleocr==2.0.6) (1.1.5)\nRequirement already satisfied: imageio in /usr/local/lib/python3.7/dist-packages (from imgaug==0.4.0->paddleocr==2.0.6) (2.4.1)\nRequirement already satisfied: scipy in /usr/local/lib/python3.7/dist-packages (from imgaug==0.4.0->paddleocr==2.0.6) (1.4.1)\nRequirement already satisfied: PyWavelets>=1.1.1 in /usr/local/lib/python3.7/dist-packages (from scikit-image==0.17.2->paddleocr==2.0.6) (1.1.1)\nRequirement already satisfied: tifffile>=2019.7.26 in /usr/local/lib/python3.7/dist-packages (from scikit-image==0.17.2->paddleocr==2.0.6) (2021.4.8)\nRequirement already satisfied: networkx>=2.0 in /usr/local/lib/python3.7/dist-packages (from scikit-image==0.17.2->paddleocr==2.0.6) (2.5.1)\nRequirement already satisfied: setuptools in /usr/local/lib/python3.7/dist-packages (from python-Levenshtein->paddleocr==2.0.6) (56.1.0)\nRequirement already satisfied: Werkzeug<2.0,>=0.15 in /usr/local/lib/python3.7/dist-packages (from flask>=1.1.1->visualdl->paddleocr==2.0.6) (1.0.1)\nRequirement already satisfied: itsdangerous<2.0,>=0.24 in /usr/local/lib/python3.7/dist-packages (from flask>=1.1.1->visualdl->paddleocr==2.0.6) (1.1.0)\nRequirement already satisfied: Jinja2<3.0,>=2.10.1 in /usr/local/lib/python3.7/dist-packages (from flask>=1.1.1->visualdl->paddleocr==2.0.6) (2.11.3)\nRequirement already satisfied: click<8.0,>=5.1 in /usr/local/lib/python3.7/dist-packages (from flask>=1.1.1->visualdl->paddleocr==2.0.6) (7.1.2)\nRequirement already satisfied: cycler>=0.10 in /usr/local/lib/python3.7/dist-packages (from matplotlib->visualdl->paddleocr==2.0.6) (0.10.0)\nRequirement already satisfied: pyparsing!=2.0.4,!=2.1.2,!=2.1.6,>=2.0.1 in /usr/local/lib/python3.7/dist-packages (from matplotlib->visualdl->paddleocr==2.0.6) (2.4.7)\nRequirement already satisfied: kiwisolver>=1.0.1 in /usr/local/lib/python3.7/dist-packages (from matplotlib->visualdl->paddleocr==2.0.6) (1.3.1)\nRequirement already satisfied: python-dateutil>=2.1 in /usr/local/lib/python3.7/dist-packages (from matplotlib->visualdl->paddleocr==2.0.6) (2.8.1)\nRequirement already satisfied: chardet<4,>=3.0.2 in /usr/local/lib/python3.7/dist-packages (from requests->visualdl->paddleocr==2.0.6) (3.0.4)\nRequirement already satisfied: urllib3!=1.25.0,!=1.25.1,<1.26,>=1.21.1 in /usr/local/lib/python3.7/dist-packages (from requests->visualdl->paddleocr==2.0.6) (1.24.3)\nRequirement already satisfied: idna<3,>=2.5 in /usr/local/lib/python3.7/dist-packages (from requests->visualdl->paddleocr==2.0.6) (2.10)\nRequirement already satisfied: certifi>=2017.4.17 in /usr/local/lib/python3.7/dist-packages (from requests->visualdl->paddleocr==2.0.6) (2020.12.5)\nRequirement already satisfied: mccabe<0.7.0,>=0.6.0 in /usr/local/lib/python3.7/dist-packages (from flake8>=3.7.9->visualdl->paddleocr==2.0.6) (0.6.1)\nRequirement already satisfied: pyflakes<2.4.0,>=2.3.0 in /usr/local/lib/python3.7/dist-packages (from flake8>=3.7.9->visualdl->paddleocr==2.0.6) (2.3.1)\nRequirement already satisfied: pycodestyle<2.8.0,>=2.7.0 in /usr/local/lib/python3.7/dist-packages (from flake8>=3.7.9->visualdl->paddleocr==2.0.6) (2.7.0)\nRequirement already satisfied: importlib-metadata; python_version < \"3.8\" in /usr/local/lib/python3.7/dist-packages (from flake8>=3.7.9->visualdl->paddleocr==2.0.6) (4.0.1)\nRequirement already satisfied: future>=0.6.0 in /usr/local/lib/python3.7/dist-packages (from bce-python-sdk->visualdl->paddleocr==2.0.6) (0.16.0)\nRequirement already satisfied: pycryptodome>=3.8.0 in /usr/local/lib/python3.7/dist-packages (from bce-python-sdk->visualdl->paddleocr==2.0.6) (3.10.1)\nRequirement already satisfied: Babel>=2.3 in /usr/local/lib/python3.7/dist-packages (from Flask-Babel>=1.0.0->visualdl->paddleocr==2.0.6) (2.9.1)\nRequirement already satisfied: pytz in /usr/local/lib/python3.7/dist-packages (from Flask-Babel>=1.0.0->visualdl->paddleocr==2.0.6) (2018.9)\nRequirement already satisfied: pyyaml>=5.1 in /usr/local/lib/python3.7/dist-packages (from pre-commit->visualdl->paddleocr==2.0.6) (5.4.1)\nRequirement already satisfied: virtualenv>=20.0.8 in /usr/local/lib/python3.7/dist-packages (from pre-commit->visualdl->paddleocr==2.0.6) (20.4.7)\nRequirement already satisfied: identify>=1.0.0 in /usr/local/lib/python3.7/dist-packages (from pre-commit->visualdl->paddleocr==2.0.6) (2.2.7)\nRequirement already satisfied: cfgv>=2.0.0 in /usr/local/lib/python3.7/dist-packages (from pre-commit->visualdl->paddleocr==2.0.6) (3.3.0)\nRequirement already satisfied: toml in /usr/local/lib/python3.7/dist-packages (from pre-commit->visualdl->paddleocr==2.0.6) (0.10.2)\nRequirement already satisfied: nodeenv>=0.11.1 in /usr/local/lib/python3.7/dist-packages (from pre-commit->visualdl->paddleocr==2.0.6) (1.6.0)\nRequirement already satisfied: decorator<5,>=4.3 in /usr/local/lib/python3.7/dist-packages (from networkx>=2.0->scikit-image==0.17.2->paddleocr==2.0.6) (4.4.2)\nRequirement already satisfied: MarkupSafe>=0.23 in /usr/local/lib/python3.7/dist-packages (from Jinja2<3.0,>=2.10.1->flask>=1.1.1->visualdl->paddleocr==2.0.6) (2.0.1)\nRequirement already satisfied: typing-extensions>=3.6.4; python_version < \"3.8\" in /usr/local/lib/python3.7/dist-packages (from importlib-metadata; python_version < \"3.8\"->flake8>=3.7.9->visualdl->paddleocr==2.0.6) (3.7.4.3)\nRequirement already satisfied: zipp>=0.5 in /usr/local/lib/python3.7/dist-packages (from importlib-metadata; python_version < \"3.8\"->flake8>=3.7.9->visualdl->paddleocr==2.0.6) (3.4.1)\nRequirement already satisfied: distlib<1,>=0.3.1 in /usr/local/lib/python3.7/dist-packages (from virtualenv>=20.0.8->pre-commit->visualdl->paddleocr==2.0.6) (0.3.2)\nRequirement already satisfied: filelock<4,>=3.0.0 in /usr/local/lib/python3.7/dist-packages (from virtualenv>=20.0.8->pre-commit->visualdl->paddleocr==2.0.6) (3.0.12)\nRequirement already satisfied: appdirs<2,>=1.4.3 in /usr/local/lib/python3.7/dist-packages (from virtualenv>=20.0.8->pre-commit->visualdl->paddleocr==2.0.6) (1.4.4)\n"
]
],
[
[
"Copy paddleocr.py to cp_pdocr.py \n",
"_____no_output_____"
]
],
[
[
"!python cp_pdocr.py --image_dir doc/imgs_en/can-ho.jpg --lang=en",
"Namespace(cls_batch_num=6, cls_image_shape='3, 48, 192', cls_model_dir='/root/.paddleocr/cls', cls_thresh=0.9, det=True, det_algorithm='DB', det_db_box_thresh=0.5, det_db_score_mode='fast', det_db_thresh=0.3, det_db_unclip_ratio=1.6, det_east_cover_thresh=0.1, det_east_nms_thresh=0.2, det_east_score_thresh=0.8, det_limit_side_len=960, det_limit_type='max', det_model_dir='/root/.paddleocr/2.1/det/en', drop_score=0.5, enable_mkldnn=False, gpu_mem=8000, image_dir='doc/imgs_en/can-ho.jpg', ir_optim=True, label_list=['0', '180'], lang='en', max_text_length=25, rec=True, rec_algorithm='CRNN', rec_batch_num=6, rec_char_dict_path='./ppocr/utils/en_dict.txt', rec_char_type='ch', rec_image_shape='3, 32, 320', rec_model_dir='/root/.paddleocr/2.1/rec/en', use_angle_cls=False, use_dilation=False, use_gpu=True, use_pdserving=False, use_space_char=True, use_tensorrt=False, use_zero_copy_run=False)\nI0602 07:49:46.983633 763 analysis_config.cc:424] use_dlnne_:0\nE0602 07:49:46.983685 763 analysis_config.cc:81] Please compile with gpu to EnableGpu()\nI0602 07:49:46.983705 763 analysis_config.cc:424] use_dlnne_:0\nI0602 07:49:46.983726 763 analysis_config.cc:424] use_dlnne_:0\nI0602 07:49:46.983745 763 analysis_config.cc:424] use_dlnne_:0\n\u001b[37m--- Fused 0 subgraphs into layer_norm op.\u001b[0m\nE0602 07:49:47.092418 763 analysis_config.cc:81] Please compile with gpu to EnableGpu()\n\u001b[37m--- Fused 0 subgraphs into layer_norm op.\u001b[0m\n[2021/06/02 07:49:47] root INFO: **********doc/imgs_en/can-ho.jpg**********\n[2021/06/02 07:49:47] root INFO: dt_boxes num : 20, elapse : 0.7077724933624268\n[2021/06/02 07:49:48] root INFO: rec_res num : 20, elapse : 0.38051676750183105\n[[98.0, 29.0], [232.0, 32.0], [232.0, 56.0], [97.0, 53.0]]\n[[532.0, 26.0], [669.0, 29.0], [668.0, 57.0], [531.0, 54.0]]\n[[100.0, 67.0], [255.0, 67.0], [255.0, 80.0], [100.0, 80.0]]\n[[267.0, 67.0], [329.0, 67.0], [329.0, 80.0], [267.0, 80.0]]\n[[533.0, 65.0], [758.0, 66.0], [758.0, 80.0], [533.0, 79.0]]\n[[100.0, 82.0], [321.0, 82.0], [321.0, 95.0], [100.0, 95.0]]\n[[533.0, 81.0], [760.0, 81.0], [760.0, 97.0], [533.0, 97.0]]\n[[743.0, 128.0], [763.0, 128.0], [763.0, 135.0], [743.0, 135.0]]\n[[137.0, 144.0], [159.0, 144.0], [159.0, 154.0], [137.0, 154.0]]\n[[114.0, 463.0], [250.0, 466.0], [250.0, 497.0], [113.0, 494.0]]\n[[535.0, 465.0], [673.0, 467.0], [673.0, 497.0], [534.0, 495.0]]\n[[117.0, 506.0], [253.0, 506.0], [253.0, 519.0], [117.0, 519.0]]\n[[257.0, 506.0], [320.0, 506.0], [320.0, 519.0], [257.0, 519.0]]\n[[537.0, 507.0], [693.0, 507.0], [693.0, 520.0], [537.0, 520.0]]\n[[693.0, 507.0], [768.0, 507.0], [768.0, 520.0], [693.0, 520.0]]\n[[116.0, 521.0], [321.0, 521.0], [321.0, 537.0], [116.0, 537.0]]\n[[535.0, 519.0], [758.0, 519.0], [758.0, 536.0], [535.0, 536.0]]\n"
]
]
] | [
"code",
"markdown",
"code"
] | [
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
]
] |
d0ff8383e31e5fe79d38ecfad99b5805195f3a0c | 1,109 | ipynb | Jupyter Notebook | Everything_Python.ipynb | gaurav997/PROGRAMMING-PYTHON | 21e2b59b5a66528384d76ed48f7b84fff22faf5a | [
"MIT"
] | null | null | null | Everything_Python.ipynb | gaurav997/PROGRAMMING-PYTHON | 21e2b59b5a66528384d76ed48f7b84fff22faf5a | [
"MIT"
] | null | null | null | Everything_Python.ipynb | gaurav997/PROGRAMMING-PYTHON | 21e2b59b5a66528384d76ed48f7b84fff22faf5a | [
"MIT"
] | null | null | null | 21.745098 | 133 | 0.557259 | [
[
[
"Hello World!",
"_____no_output_____"
],
[
"print(\"Hello World!\")",
"_____no_output_____"
]
]
] | [
"code"
] | [
[
"code",
"code"
]
] |
d0ff8921c4c45a5a8857164885955ed4b6a6cbf6 | 483,320 | ipynb | Jupyter Notebook | mortality_model.ipynb | cosgriffc/bst261-crrt | c2b515b3827367cad1adae23819bdb8a0a5a9455 | [
"MIT"
] | 1 | 2019-01-01T12:57:34.000Z | 2019-01-01T12:57:34.000Z | mortality_model.ipynb | cosgriffc/bst261-crrt | c2b515b3827367cad1adae23819bdb8a0a5a9455 | [
"MIT"
] | 1 | 2019-01-02T07:35:44.000Z | 2019-01-04T22:54:22.000Z | mortality_model.ipynb | cosgriffc/bst261-crrt | c2b515b3827367cad1adae23819bdb8a0a5a9455 | [
"MIT"
] | null | null | null | 196.311942 | 24,732 | 0.723947 | [
[
[
"# CRRT Mortality Prediction\n## Model Construction\n### Christopher V. Cosgriff, David Sasson, Colby Wilkinson, Kanhua Yin\n\n\nThe purpose of this notebook is to build a deep learning model that predicts ICU mortality in the CRRT population. The data is extracted in the `extract_cohort_and_features` notebook and stored in the `data` folder. This model will be mult-input and use GRUs to model sequence data. See the extraction file for a full description of the data extraction.",
"_____no_output_____"
],
[
"## Step 0: Envrionment Setup",
"_____no_output_____"
]
],
[
[
"import numpy as np\nimport matplotlib as mpl\nimport matplotlib.pyplot as plt\nfrom IPython.display import SVG\nimport os\n\nfrom keras.optimizers import Adam, SGD, rmsprop\nfrom keras.models import Sequential,Model\nfrom keras.layers import Dense, Activation, Dropout, Input, Dropout, concatenate\nfrom keras.layers.recurrent import GRU\nfrom keras.utils import plot_model\nfrom keras.utils.vis_utils import model_to_dot\n\nfrom sklearn.model_selection import train_test_split\nfrom sklearn.metrics import roc_auc_score, roc_curve\n\n# for saving images\nfig_fp = os.path.join('./', 'figures')\nif not os.path.isdir(fig_fp):\n os.mkdir(fig_fp)\n\n%matplotlib inline",
"/home/cosgriffc/env/ML/lib/python3.5/site-packages/h5py/__init__.py:36: FutureWarning: Conversion of the second argument of issubdtype from `float` to `np.floating` is deprecated. In future, it will be treated as `np.float64 == np.dtype(float).type`.\n from ._conv import register_converters as _register_converters\nUsing TensorFlow backend.\n"
]
],
[
[
"## Step 1: Load and Prepare Data\n\nHere will we load in the data, and create train, validation, and testing splits.",
"_____no_output_____"
]
],
[
[
"# set tensors to float 32 as this is what GPUs expect\nfeatures_sequence = np.load('./features_sequence.npy').astype(np.float32)\nfeatures_static = np.load('./features_static.npy').astype(np.float32)\nlabels = np.load('./labels.npy').astype(np.float32)\n\nx_seq_full_train, x_seq_test, x_static_full_train, x_static_test, y_full_train, y_test = train_test_split(\n features_sequence, features_static, labels, test_size = 0.20, random_state = 42)\n\nx_seq_train, x_seq_val, x_static_train, x_static_val, y_train, y_val = train_test_split(\n x_seq_full_train, x_static_full_train, y_full_train, test_size = 0.10, random_state = 42)",
"_____no_output_____"
]
],
[
[
"Next we need to remove NANs from the data; we'll impute the trianing population mean, the simplest method suggested by David Sontag.",
"_____no_output_____"
]
],
[
[
"def impute_mean(source_data, input_data):\n '''\n Takes the source data, and uses it to determine means for all\n features; it then applies them to the input data.\n \n inputs:\n source_data: a tensor to provide means\n input_data: the data to fill in NA for\n \n output:\n output_data: data with nans imputed for each feature\n \n '''\n \n output_data = input_data.copy()\n \n for feature in range(source_data.shape[1]):\n feature_mean = np.nanmean(source_data[:, feature, :][np.where(source_data[:, feature, :] != 0)])\n ind_output_data = np.where(np.isnan(output_data[:, feature, :]))\n output_data[:, feature, :][ind_output_data] = feature_mean\n \n return output_data\n\nx_seq_train_original = x_seq_train.copy()\nx_seq_train = impute_mean(x_seq_train_original, x_seq_train)\nx_seq_val = impute_mean(x_seq_train_original, x_seq_val)\nx_seq_test = impute_mean(x_seq_train_original, x_seq_test)",
"_____no_output_____"
]
],
[
[
"## Step 2: Build Model\n### Model 1\n\nBase model, no regularization.",
"_____no_output_____"
]
],
[
[
"# Define inputs\nsequence_input = Input(shape = (x_seq_train.shape[1], x_seq_train.shape[2], ), dtype = 'float32', name = 'sequence_input')\nstatic_input = Input(shape = (x_static_train.shape[1], ), name = 'static_input')\n\n# Network architecture\nseq_x = GRU(units = 128)(sequence_input)\n\n# Seperate output for the GRU later\nseq_aux_output = Dense(1, activation='sigmoid', name='aux_output')(seq_x)\n\n# Merge dual inputs\nx = concatenate([seq_x, static_input])\n\n# We stack a deep fully-connected network on the merged inputs\nx = Dense(128, activation = 'relu')(x)\nx = Dense(128, activation = 'relu')(x)\nx = Dense(128, activation = 'relu')(x)\nx = Dense(128, activation = 'relu')(x)\n\n# Sigmoid output layer\nmain_output = Dense(1, activation='sigmoid', name='main_output')(x)\n\n# optimizer\nopt = rmsprop(lr = 0.00001)\n\n# build model\nmodel = Model(inputs = [sequence_input, static_input], outputs = [main_output, seq_aux_output])\nmodel.compile(optimizer = opt, loss = 'binary_crossentropy', metrics = ['accuracy'], loss_weights = [1, 0.1])\n\n# save a plot of the model\nplot_model(model, to_file='experiment_GRU-base.svg')\n\n# fit the model\nhistory = model.fit([x_seq_train, x_static_train], [y_train, y_train], epochs = 500, batch_size = 128,\\\n validation_data=([x_seq_val, x_static_val], [y_val, y_val]),)",
"Train on 924 samples, validate on 103 samples\nEpoch 1/500\n924/924 [==============================] - 2s 2ms/step - loss: 0.7793 - main_output_loss: 0.7098 - aux_output_loss: 0.6952 - main_output_acc: 0.4913 - aux_output_acc: 0.5032 - val_loss: 0.7621 - val_main_output_loss: 0.6941 - val_aux_output_loss: 0.6799 - val_main_output_acc: 0.5534 - val_aux_output_acc: 0.5243\nEpoch 2/500\n924/924 [==============================] - 0s 247us/step - loss: 0.7749 - main_output_loss: 0.7056 - aux_output_loss: 0.6922 - main_output_acc: 0.4903 - aux_output_acc: 0.4989 - val_loss: 0.7551 - val_main_output_loss: 0.6872 - val_aux_output_loss: 0.6790 - val_main_output_acc: 0.5728 - val_aux_output_acc: 0.5243\nEpoch 3/500\n924/924 [==============================] - 0s 234us/step - loss: 0.7725 - main_output_loss: 0.7034 - aux_output_loss: 0.6910 - main_output_acc: 0.5097 - aux_output_acc: 0.4989 - val_loss: 0.7520 - val_main_output_loss: 0.6842 - val_aux_output_loss: 0.6784 - val_main_output_acc: 0.5922 - val_aux_output_acc: 0.5146\nEpoch 4/500\n924/924 [==============================] - 0s 239us/step - loss: 0.7694 - main_output_loss: 0.7005 - aux_output_loss: 0.6897 - main_output_acc: 0.5195 - aux_output_acc: 0.4989 - val_loss: 0.7512 - val_main_output_loss: 0.6834 - val_aux_output_loss: 0.6779 - val_main_output_acc: 0.6117 - val_aux_output_acc: 0.5146\nEpoch 5/500\n924/924 [==============================] - 0s 229us/step - loss: 0.7677 - main_output_loss: 0.6989 - aux_output_loss: 0.6884 - main_output_acc: 0.5141 - aux_output_acc: 0.4978 - val_loss: 0.7450 - val_main_output_loss: 0.6773 - val_aux_output_loss: 0.6774 - val_main_output_acc: 0.6019 - val_aux_output_acc: 0.5049\nEpoch 6/500\n924/924 [==============================] - 0s 232us/step - loss: 0.7660 - main_output_loss: 0.6973 - aux_output_loss: 0.6876 - main_output_acc: 0.5195 - aux_output_acc: 0.4924 - val_loss: 0.7417 - val_main_output_loss: 0.6740 - val_aux_output_loss: 0.6769 - val_main_output_acc: 0.5922 - val_aux_output_acc: 0.4951\nEpoch 7/500\n924/924 [==============================] - 0s 228us/step - loss: 0.7635 - main_output_loss: 0.6948 - aux_output_loss: 0.6868 - main_output_acc: 0.5314 - aux_output_acc: 0.4946 - val_loss: 0.7393 - val_main_output_loss: 0.6716 - val_aux_output_loss: 0.6764 - val_main_output_acc: 0.6117 - val_aux_output_acc: 0.4854\nEpoch 8/500\n924/924 [==============================] - 0s 231us/step - loss: 0.7620 - main_output_loss: 0.6934 - aux_output_loss: 0.6862 - main_output_acc: 0.5303 - aux_output_acc: 0.4968 - val_loss: 0.7375 - val_main_output_loss: 0.6699 - val_aux_output_loss: 0.6760 - val_main_output_acc: 0.6019 - val_aux_output_acc: 0.4951\nEpoch 9/500\n924/924 [==============================] - 0s 226us/step - loss: 0.7605 - main_output_loss: 0.6920 - aux_output_loss: 0.6856 - main_output_acc: 0.5281 - aux_output_acc: 0.5043 - val_loss: 0.7347 - val_main_output_loss: 0.6671 - val_aux_output_loss: 0.6756 - val_main_output_acc: 0.6117 - val_aux_output_acc: 0.5049\nEpoch 10/500\n924/924 [==============================] - 0s 234us/step - loss: 0.7591 - main_output_loss: 0.6906 - aux_output_loss: 0.6850 - main_output_acc: 0.5249 - aux_output_acc: 0.5000 - val_loss: 0.7344 - val_main_output_loss: 0.6668 - val_aux_output_loss: 0.6752 - val_main_output_acc: 0.6408 - val_aux_output_acc: 0.5049\nEpoch 11/500\n924/924 [==============================] - 0s 236us/step - loss: 0.7577 - main_output_loss: 0.6893 - aux_output_loss: 0.6842 - main_output_acc: 0.5444 - aux_output_acc: 0.5011 - val_loss: 0.7343 - val_main_output_loss: 0.6668 - val_aux_output_loss: 0.6749 - val_main_output_acc: 0.6117 - val_aux_output_acc: 0.5243\nEpoch 12/500\n924/924 [==============================] - 0s 232us/step - loss: 0.7564 - main_output_loss: 0.6881 - aux_output_loss: 0.6834 - main_output_acc: 0.5422 - aux_output_acc: 0.5130 - val_loss: 0.7352 - val_main_output_loss: 0.6678 - val_aux_output_loss: 0.6745 - val_main_output_acc: 0.6214 - val_aux_output_acc: 0.5340\nEpoch 13/500\n924/924 [==============================] - 0s 238us/step - loss: 0.7557 - main_output_loss: 0.6874 - aux_output_loss: 0.6827 - main_output_acc: 0.5422 - aux_output_acc: 0.5238 - val_loss: 0.7321 - val_main_output_loss: 0.6647 - val_aux_output_loss: 0.6741 - val_main_output_acc: 0.6117 - val_aux_output_acc: 0.5340\nEpoch 14/500\n924/924 [==============================] - 0s 233us/step - loss: 0.7543 - main_output_loss: 0.6861 - aux_output_loss: 0.6820 - main_output_acc: 0.5422 - aux_output_acc: 0.5368 - val_loss: 0.7339 - val_main_output_loss: 0.6665 - val_aux_output_loss: 0.6738 - val_main_output_acc: 0.6408 - val_aux_output_acc: 0.5340\nEpoch 15/500\n924/924 [==============================] - 0s 232us/step - loss: 0.7537 - main_output_loss: 0.6856 - aux_output_loss: 0.6814 - main_output_acc: 0.5357 - aux_output_acc: 0.5455 - val_loss: 0.7288 - val_main_output_loss: 0.6615 - val_aux_output_loss: 0.6731 - val_main_output_acc: 0.6117 - val_aux_output_acc: 0.5437\nEpoch 16/500\n924/924 [==============================] - 0s 230us/step - loss: 0.7526 - main_output_loss: 0.6845 - aux_output_loss: 0.6809 - main_output_acc: 0.5498 - aux_output_acc: 0.5422 - val_loss: 0.7274 - val_main_output_loss: 0.6601 - val_aux_output_loss: 0.6727 - val_main_output_acc: 0.6117 - val_aux_output_acc: 0.5437\nEpoch 17/500\n924/924 [==============================] - 0s 241us/step - loss: 0.7515 - main_output_loss: 0.6834 - aux_output_loss: 0.6803 - main_output_acc: 0.5530 - aux_output_acc: 0.5465 - val_loss: 0.7267 - val_main_output_loss: 0.6595 - val_aux_output_loss: 0.6724 - val_main_output_acc: 0.6214 - val_aux_output_acc: 0.5437\nEpoch 18/500\n924/924 [==============================] - 0s 227us/step - loss: 0.7503 - main_output_loss: 0.6823 - aux_output_loss: 0.6797 - main_output_acc: 0.5574 - aux_output_acc: 0.5617 - val_loss: 0.7256 - val_main_output_loss: 0.6584 - val_aux_output_loss: 0.6720 - val_main_output_acc: 0.6214 - val_aux_output_acc: 0.5534\nEpoch 19/500\n924/924 [==============================] - 0s 235us/step - loss: 0.7490 - main_output_loss: 0.6811 - aux_output_loss: 0.6791 - main_output_acc: 0.5671 - aux_output_acc: 0.5682 - val_loss: 0.7224 - val_main_output_loss: 0.6552 - val_aux_output_loss: 0.6714 - val_main_output_acc: 0.6117 - val_aux_output_acc: 0.5340\nEpoch 20/500\n924/924 [==============================] - 0s 227us/step - loss: 0.7481 - main_output_loss: 0.6803 - aux_output_loss: 0.6787 - main_output_acc: 0.5584 - aux_output_acc: 0.5703 - val_loss: 0.7252 - val_main_output_loss: 0.6581 - val_aux_output_loss: 0.6711 - val_main_output_acc: 0.6311 - val_aux_output_acc: 0.5728\nEpoch 21/500\n924/924 [==============================] - 0s 229us/step - loss: 0.7471 - main_output_loss: 0.6793 - aux_output_loss: 0.6780 - main_output_acc: 0.5639 - aux_output_acc: 0.5639 - val_loss: 0.7201 - val_main_output_loss: 0.6531 - val_aux_output_loss: 0.6704 - val_main_output_acc: 0.6311 - val_aux_output_acc: 0.5728\nEpoch 22/500\n924/924 [==============================] - 0s 232us/step - loss: 0.7464 - main_output_loss: 0.6787 - aux_output_loss: 0.6775 - main_output_acc: 0.5628 - aux_output_acc: 0.5725 - val_loss: 0.7181 - val_main_output_loss: 0.6511 - val_aux_output_loss: 0.6697 - val_main_output_acc: 0.6214 - val_aux_output_acc: 0.5631\nEpoch 23/500\n924/924 [==============================] - 0s 236us/step - loss: 0.7467 - main_output_loss: 0.6790 - aux_output_loss: 0.6769 - main_output_acc: 0.5541 - aux_output_acc: 0.5736 - val_loss: 0.7177 - val_main_output_loss: 0.6508 - val_aux_output_loss: 0.6689 - val_main_output_acc: 0.6117 - val_aux_output_acc: 0.5534\nEpoch 24/500\n924/924 [==============================] - 0s 226us/step - loss: 0.7462 - main_output_loss: 0.6786 - aux_output_loss: 0.6764 - main_output_acc: 0.5563 - aux_output_acc: 0.5747 - val_loss: 0.7172 - val_main_output_loss: 0.6503 - val_aux_output_loss: 0.6682 - val_main_output_acc: 0.6117 - val_aux_output_acc: 0.5437\nEpoch 25/500\n924/924 [==============================] - 0s 233us/step - loss: 0.7447 - main_output_loss: 0.6771 - aux_output_loss: 0.6760 - main_output_acc: 0.5649 - aux_output_acc: 0.5660 - val_loss: 0.7184 - val_main_output_loss: 0.6516 - val_aux_output_loss: 0.6678 - val_main_output_acc: 0.6214 - val_aux_output_acc: 0.5534\nEpoch 26/500\n924/924 [==============================] - 0s 225us/step - loss: 0.7436 - main_output_loss: 0.6761 - aux_output_loss: 0.6754 - main_output_acc: 0.5714 - aux_output_acc: 0.5693 - val_loss: 0.7175 - val_main_output_loss: 0.6508 - val_aux_output_loss: 0.6671 - val_main_output_acc: 0.6214 - val_aux_output_acc: 0.5534\nEpoch 27/500\n924/924 [==============================] - 0s 229us/step - loss: 0.7430 - main_output_loss: 0.6755 - aux_output_loss: 0.6749 - main_output_acc: 0.5703 - aux_output_acc: 0.5877 - val_loss: 0.7149 - val_main_output_loss: 0.6483 - val_aux_output_loss: 0.6668 - val_main_output_acc: 0.6311 - val_aux_output_acc: 0.5728\nEpoch 28/500\n924/924 [==============================] - 0s 222us/step - loss: 0.7427 - main_output_loss: 0.6753 - aux_output_loss: 0.6745 - main_output_acc: 0.5714 - aux_output_acc: 0.5844 - val_loss: 0.7147 - val_main_output_loss: 0.6481 - val_aux_output_loss: 0.6663 - val_main_output_acc: 0.6311 - val_aux_output_acc: 0.5728\nEpoch 29/500\n924/924 [==============================] - 0s 229us/step - loss: 0.7420 - main_output_loss: 0.6746 - aux_output_loss: 0.6740 - main_output_acc: 0.5617 - aux_output_acc: 0.5801 - val_loss: 0.7149 - val_main_output_loss: 0.6483 - val_aux_output_loss: 0.6657 - val_main_output_acc: 0.6408 - val_aux_output_acc: 0.5631\nEpoch 30/500\n924/924 [==============================] - 0s 228us/step - loss: 0.7410 - main_output_loss: 0.6737 - aux_output_loss: 0.6735 - main_output_acc: 0.5703 - aux_output_acc: 0.5866 - val_loss: 0.7197 - val_main_output_loss: 0.6531 - val_aux_output_loss: 0.6654 - val_main_output_acc: 0.6311 - val_aux_output_acc: 0.6019\nEpoch 31/500\n924/924 [==============================] - 0s 240us/step - loss: 0.7404 - main_output_loss: 0.6732 - aux_output_loss: 0.6728 - main_output_acc: 0.5801 - aux_output_acc: 0.5887 - val_loss: 0.7188 - val_main_output_loss: 0.6523 - val_aux_output_loss: 0.6649 - val_main_output_acc: 0.6214 - val_aux_output_acc: 0.5922\nEpoch 32/500\n924/924 [==============================] - 0s 235us/step - loss: 0.7389 - main_output_loss: 0.6717 - aux_output_loss: 0.6723 - main_output_acc: 0.5812 - aux_output_acc: 0.5920 - val_loss: 0.7151 - val_main_output_loss: 0.6487 - val_aux_output_loss: 0.6642 - val_main_output_acc: 0.6117 - val_aux_output_acc: 0.5922\nEpoch 33/500\n924/924 [==============================] - 0s 223us/step - loss: 0.7404 - main_output_loss: 0.6732 - aux_output_loss: 0.6718 - main_output_acc: 0.5584 - aux_output_acc: 0.5909 - val_loss: 0.7179 - val_main_output_loss: 0.6515 - val_aux_output_loss: 0.6637 - val_main_output_acc: 0.6408 - val_aux_output_acc: 0.5922\nEpoch 34/500\n924/924 [==============================] - 0s 235us/step - loss: 0.7386 - main_output_loss: 0.6715 - aux_output_loss: 0.6713 - main_output_acc: 0.5747 - aux_output_acc: 0.5963 - val_loss: 0.7161 - val_main_output_loss: 0.6498 - val_aux_output_loss: 0.6634 - val_main_output_acc: 0.6408 - val_aux_output_acc: 0.5922\nEpoch 35/500\n924/924 [==============================] - 0s 242us/step - loss: 0.7378 - main_output_loss: 0.6707 - aux_output_loss: 0.6708 - main_output_acc: 0.5812 - aux_output_acc: 0.6006 - val_loss: 0.7134 - val_main_output_loss: 0.6471 - val_aux_output_loss: 0.6629 - val_main_output_acc: 0.6117 - val_aux_output_acc: 0.5922\nEpoch 36/500\n924/924 [==============================] - 0s 233us/step - loss: 0.7371 - main_output_loss: 0.6700 - aux_output_loss: 0.6702 - main_output_acc: 0.5758 - aux_output_acc: 0.5952 - val_loss: 0.7142 - val_main_output_loss: 0.6480 - val_aux_output_loss: 0.6624 - val_main_output_acc: 0.6408 - val_aux_output_acc: 0.5825\nEpoch 37/500\n924/924 [==============================] - 0s 227us/step - loss: 0.7377 - main_output_loss: 0.6707 - aux_output_loss: 0.6698 - main_output_acc: 0.5779 - aux_output_acc: 0.6039 - val_loss: 0.7138 - val_main_output_loss: 0.6476 - val_aux_output_loss: 0.6619 - val_main_output_acc: 0.6408 - val_aux_output_acc: 0.5825\nEpoch 38/500\n924/924 [==============================] - 0s 230us/step - loss: 0.7365 - main_output_loss: 0.6696 - aux_output_loss: 0.6692 - main_output_acc: 0.5812 - aux_output_acc: 0.6017 - val_loss: 0.7138 - val_main_output_loss: 0.6477 - val_aux_output_loss: 0.6615 - val_main_output_acc: 0.6311 - val_aux_output_acc: 0.5825\nEpoch 39/500\n924/924 [==============================] - 0s 229us/step - loss: 0.7364 - main_output_loss: 0.6695 - aux_output_loss: 0.6687 - main_output_acc: 0.5812 - aux_output_acc: 0.6071 - val_loss: 0.7128 - val_main_output_loss: 0.6467 - val_aux_output_loss: 0.6611 - val_main_output_acc: 0.6214 - val_aux_output_acc: 0.5825\nEpoch 40/500\n924/924 [==============================] - 0s 228us/step - loss: 0.7350 - main_output_loss: 0.6682 - aux_output_loss: 0.6680 - main_output_acc: 0.5823 - aux_output_acc: 0.6082 - val_loss: 0.7121 - val_main_output_loss: 0.6461 - val_aux_output_loss: 0.6605 - val_main_output_acc: 0.6117 - val_aux_output_acc: 0.5825\nEpoch 41/500\n924/924 [==============================] - 0s 226us/step - loss: 0.7353 - main_output_loss: 0.6685 - aux_output_loss: 0.6677 - main_output_acc: 0.5714 - aux_output_acc: 0.6061 - val_loss: 0.7108 - val_main_output_loss: 0.6448 - val_aux_output_loss: 0.6600 - val_main_output_acc: 0.6117 - val_aux_output_acc: 0.5825\nEpoch 42/500\n924/924 [==============================] - 0s 232us/step - loss: 0.7348 - main_output_loss: 0.6681 - aux_output_loss: 0.6673 - main_output_acc: 0.5714 - aux_output_acc: 0.6115 - val_loss: 0.7106 - val_main_output_loss: 0.6447 - val_aux_output_loss: 0.6595 - val_main_output_acc: 0.6117 - val_aux_output_acc: 0.5825\nEpoch 43/500\n924/924 [==============================] - 0s 230us/step - loss: 0.7334 - main_output_loss: 0.6667 - aux_output_loss: 0.6669 - main_output_acc: 0.5747 - aux_output_acc: 0.6082 - val_loss: 0.7136 - val_main_output_loss: 0.6477 - val_aux_output_loss: 0.6592 - val_main_output_acc: 0.6408 - val_aux_output_acc: 0.5728\nEpoch 44/500\n924/924 [==============================] - 0s 231us/step - loss: 0.7331 - main_output_loss: 0.6665 - aux_output_loss: 0.6663 - main_output_acc: 0.5898 - aux_output_acc: 0.6136 - val_loss: 0.7122 - val_main_output_loss: 0.6463 - val_aux_output_loss: 0.6589 - val_main_output_acc: 0.6311 - val_aux_output_acc: 0.5728\nEpoch 45/500\n924/924 [==============================] - 0s 226us/step - loss: 0.7326 - main_output_loss: 0.6660 - aux_output_loss: 0.6659 - main_output_acc: 0.6061 - aux_output_acc: 0.6147 - val_loss: 0.7118 - val_main_output_loss: 0.6459 - val_aux_output_loss: 0.6588 - val_main_output_acc: 0.6408 - val_aux_output_acc: 0.5728\nEpoch 46/500\n924/924 [==============================] - 0s 228us/step - loss: 0.7320 - main_output_loss: 0.6655 - aux_output_loss: 0.6655 - main_output_acc: 0.6028 - aux_output_acc: 0.6126 - val_loss: 0.7141 - val_main_output_loss: 0.6482 - val_aux_output_loss: 0.6586 - val_main_output_acc: 0.6311 - val_aux_output_acc: 0.5825\nEpoch 47/500\n924/924 [==============================] - 0s 235us/step - loss: 0.7318 - main_output_loss: 0.6653 - aux_output_loss: 0.6650 - main_output_acc: 0.5823 - aux_output_acc: 0.6266 - val_loss: 0.7098 - val_main_output_loss: 0.6440 - val_aux_output_loss: 0.6582 - val_main_output_acc: 0.6408 - val_aux_output_acc: 0.5825\nEpoch 48/500\n924/924 [==============================] - 0s 229us/step - loss: 0.7307 - main_output_loss: 0.6643 - aux_output_loss: 0.6647 - main_output_acc: 0.5866 - aux_output_acc: 0.6223 - val_loss: 0.7090 - val_main_output_loss: 0.6432 - val_aux_output_loss: 0.6579 - val_main_output_acc: 0.6408 - val_aux_output_acc: 0.5825\nEpoch 49/500\n924/924 [==============================] - 0s 229us/step - loss: 0.7306 - main_output_loss: 0.6642 - aux_output_loss: 0.6643 - main_output_acc: 0.5963 - aux_output_acc: 0.6255 - val_loss: 0.7061 - val_main_output_loss: 0.6403 - val_aux_output_loss: 0.6574 - val_main_output_acc: 0.6019 - val_aux_output_acc: 0.5728\nEpoch 50/500\n924/924 [==============================] - 0s 226us/step - loss: 0.7302 - main_output_loss: 0.6637 - aux_output_loss: 0.6641 - main_output_acc: 0.5801 - aux_output_acc: 0.6255 - val_loss: 0.7065 - val_main_output_loss: 0.6408 - val_aux_output_loss: 0.6570 - val_main_output_acc: 0.6505 - val_aux_output_acc: 0.5922\nEpoch 51/500\n924/924 [==============================] - 0s 224us/step - loss: 0.7295 - main_output_loss: 0.6632 - aux_output_loss: 0.6636 - main_output_acc: 0.6017 - aux_output_acc: 0.6277 - val_loss: 0.7063 - val_main_output_loss: 0.6407 - val_aux_output_loss: 0.6566 - val_main_output_acc: 0.6214 - val_aux_output_acc: 0.5922\nEpoch 52/500\n924/924 [==============================] - 0s 224us/step - loss: 0.7288 - main_output_loss: 0.6625 - aux_output_loss: 0.6632 - main_output_acc: 0.5866 - aux_output_acc: 0.6299 - val_loss: 0.7081 - val_main_output_loss: 0.6424 - val_aux_output_loss: 0.6564 - val_main_output_acc: 0.6408 - val_aux_output_acc: 0.6019\nEpoch 53/500\n924/924 [==============================] - 0s 229us/step - loss: 0.7295 - main_output_loss: 0.6632 - aux_output_loss: 0.6629 - main_output_acc: 0.6017 - aux_output_acc: 0.6331 - val_loss: 0.7066 - val_main_output_loss: 0.6410 - val_aux_output_loss: 0.6560 - val_main_output_acc: 0.6408 - val_aux_output_acc: 0.5922\nEpoch 54/500\n924/924 [==============================] - 0s 240us/step - loss: 0.7276 - main_output_loss: 0.6614 - aux_output_loss: 0.6623 - main_output_acc: 0.6050 - aux_output_acc: 0.6374 - val_loss: 0.7047 - val_main_output_loss: 0.6391 - val_aux_output_loss: 0.6556 - val_main_output_acc: 0.6117 - val_aux_output_acc: 0.6019\nEpoch 55/500\n924/924 [==============================] - 0s 232us/step - loss: 0.7279 - main_output_loss: 0.6617 - aux_output_loss: 0.6619 - main_output_acc: 0.5844 - aux_output_acc: 0.6374 - val_loss: 0.7060 - val_main_output_loss: 0.6405 - val_aux_output_loss: 0.6555 - val_main_output_acc: 0.6505 - val_aux_output_acc: 0.6019\nEpoch 56/500\n924/924 [==============================] - 0s 223us/step - loss: 0.7269 - main_output_loss: 0.6608 - aux_output_loss: 0.6615 - main_output_acc: 0.6039 - aux_output_acc: 0.6418 - val_loss: 0.7046 - val_main_output_loss: 0.6391 - val_aux_output_loss: 0.6552 - val_main_output_acc: 0.6214 - val_aux_output_acc: 0.6019\nEpoch 57/500\n924/924 [==============================] - 0s 224us/step - loss: 0.7266 - main_output_loss: 0.6605 - aux_output_loss: 0.6611 - main_output_acc: 0.5898 - aux_output_acc: 0.6418 - val_loss: 0.7037 - val_main_output_loss: 0.6382 - val_aux_output_loss: 0.6547 - val_main_output_acc: 0.6214 - val_aux_output_acc: 0.6019\nEpoch 58/500\n924/924 [==============================] - 0s 231us/step - loss: 0.7260 - main_output_loss: 0.6599 - aux_output_loss: 0.6608 - main_output_acc: 0.6028 - aux_output_acc: 0.6385 - val_loss: 0.7041 - val_main_output_loss: 0.6386 - val_aux_output_loss: 0.6544 - val_main_output_acc: 0.6505 - val_aux_output_acc: 0.6019\nEpoch 59/500\n924/924 [==============================] - 0s 237us/step - loss: 0.7253 - main_output_loss: 0.6592 - aux_output_loss: 0.6604 - main_output_acc: 0.5963 - aux_output_acc: 0.6396 - val_loss: 0.7024 - val_main_output_loss: 0.6370 - val_aux_output_loss: 0.6539 - val_main_output_acc: 0.6214 - val_aux_output_acc: 0.6019\nEpoch 60/500\n924/924 [==============================] - 0s 237us/step - loss: 0.7252 - main_output_loss: 0.6592 - aux_output_loss: 0.6600 - main_output_acc: 0.6017 - aux_output_acc: 0.6396 - val_loss: 0.7031 - val_main_output_loss: 0.6377 - val_aux_output_loss: 0.6537 - val_main_output_acc: 0.6117 - val_aux_output_acc: 0.6019\nEpoch 61/500\n924/924 [==============================] - 0s 220us/step - loss: 0.7243 - main_output_loss: 0.6583 - aux_output_loss: 0.6596 - main_output_acc: 0.6061 - aux_output_acc: 0.6429 - val_loss: 0.7042 - val_main_output_loss: 0.6388 - val_aux_output_loss: 0.6536 - val_main_output_acc: 0.6505 - val_aux_output_acc: 0.6019\nEpoch 62/500\n924/924 [==============================] - 0s 227us/step - loss: 0.7236 - main_output_loss: 0.6577 - aux_output_loss: 0.6593 - main_output_acc: 0.6006 - aux_output_acc: 0.6483 - val_loss: 0.7063 - val_main_output_loss: 0.6410 - val_aux_output_loss: 0.6535 - val_main_output_acc: 0.6602 - val_aux_output_acc: 0.6311\nEpoch 63/500\n924/924 [==============================] - 0s 223us/step - loss: 0.7232 - main_output_loss: 0.6574 - aux_output_loss: 0.6589 - main_output_acc: 0.6158 - aux_output_acc: 0.6483 - val_loss: 0.7015 - val_main_output_loss: 0.6362 - val_aux_output_loss: 0.6533 - val_main_output_acc: 0.6117 - val_aux_output_acc: 0.6019\nEpoch 64/500\n924/924 [==============================] - 0s 224us/step - loss: 0.7228 - main_output_loss: 0.6570 - aux_output_loss: 0.6585 - main_output_acc: 0.5931 - aux_output_acc: 0.6472 - val_loss: 0.7013 - val_main_output_loss: 0.6360 - val_aux_output_loss: 0.6532 - val_main_output_acc: 0.6311 - val_aux_output_acc: 0.6117\nEpoch 65/500\n924/924 [==============================] - 0s 224us/step - loss: 0.7222 - main_output_loss: 0.6564 - aux_output_loss: 0.6582 - main_output_acc: 0.5996 - aux_output_acc: 0.6494 - val_loss: 0.7015 - val_main_output_loss: 0.6362 - val_aux_output_loss: 0.6530 - val_main_output_acc: 0.6311 - val_aux_output_acc: 0.6117\nEpoch 66/500\n924/924 [==============================] - 0s 227us/step - loss: 0.7221 - main_output_loss: 0.6563 - aux_output_loss: 0.6579 - main_output_acc: 0.6050 - aux_output_acc: 0.6494 - val_loss: 0.6999 - val_main_output_loss: 0.6346 - val_aux_output_loss: 0.6526 - val_main_output_acc: 0.6311 - val_aux_output_acc: 0.5922\nEpoch 67/500\n924/924 [==============================] - 0s 227us/step - loss: 0.7213 - main_output_loss: 0.6555 - aux_output_loss: 0.6576 - main_output_acc: 0.6006 - aux_output_acc: 0.6472 - val_loss: 0.6991 - val_main_output_loss: 0.6338 - val_aux_output_loss: 0.6525 - val_main_output_acc: 0.6214 - val_aux_output_acc: 0.6117\nEpoch 68/500\n924/924 [==============================] - 0s 228us/step - loss: 0.7210 - main_output_loss: 0.6552 - aux_output_loss: 0.6574 - main_output_acc: 0.6115 - aux_output_acc: 0.6472 - val_loss: 0.7003 - val_main_output_loss: 0.6351 - val_aux_output_loss: 0.6523 - val_main_output_acc: 0.6214 - val_aux_output_acc: 0.6019\nEpoch 69/500\n924/924 [==============================] - 0s 227us/step - loss: 0.7202 - main_output_loss: 0.6545 - aux_output_loss: 0.6570 - main_output_acc: 0.6006 - aux_output_acc: 0.6483 - val_loss: 0.7033 - val_main_output_loss: 0.6381 - val_aux_output_loss: 0.6522 - val_main_output_acc: 0.6602 - val_aux_output_acc: 0.5922\nEpoch 70/500\n924/924 [==============================] - 0s 222us/step - loss: 0.7199 - main_output_loss: 0.6542 - aux_output_loss: 0.6566 - main_output_acc: 0.6201 - aux_output_acc: 0.6569 - val_loss: 0.7000 - val_main_output_loss: 0.6348 - val_aux_output_loss: 0.6518 - val_main_output_acc: 0.6311 - val_aux_output_acc: 0.5922\nEpoch 71/500\n924/924 [==============================] - 0s 231us/step - loss: 0.7194 - main_output_loss: 0.6537 - aux_output_loss: 0.6563 - main_output_acc: 0.6082 - aux_output_acc: 0.6515 - val_loss: 0.7040 - val_main_output_loss: 0.6388 - val_aux_output_loss: 0.6519 - val_main_output_acc: 0.6699 - val_aux_output_acc: 0.6311\nEpoch 72/500\n924/924 [==============================] - 0s 229us/step - loss: 0.7192 - main_output_loss: 0.6536 - aux_output_loss: 0.6559 - main_output_acc: 0.6169 - aux_output_acc: 0.6558 - val_loss: 0.7013 - val_main_output_loss: 0.6362 - val_aux_output_loss: 0.6517 - val_main_output_acc: 0.6699 - val_aux_output_acc: 0.6311\nEpoch 73/500\n924/924 [==============================] - 0s 229us/step - loss: 0.7181 - main_output_loss: 0.6526 - aux_output_loss: 0.6555 - main_output_acc: 0.6310 - aux_output_acc: 0.6569 - val_loss: 0.7017 - val_main_output_loss: 0.6365 - val_aux_output_loss: 0.6517 - val_main_output_acc: 0.6602 - val_aux_output_acc: 0.6408\nEpoch 74/500\n924/924 [==============================] - 0s 230us/step - loss: 0.7178 - main_output_loss: 0.6522 - aux_output_loss: 0.6552 - main_output_acc: 0.6190 - aux_output_acc: 0.6602 - val_loss: 0.6979 - val_main_output_loss: 0.6327 - val_aux_output_loss: 0.6513 - val_main_output_acc: 0.6408 - val_aux_output_acc: 0.6408\nEpoch 75/500\n924/924 [==============================] - 0s 227us/step - loss: 0.7171 - main_output_loss: 0.6517 - aux_output_loss: 0.6548 - main_output_acc: 0.6158 - aux_output_acc: 0.6591 - val_loss: 0.6975 - val_main_output_loss: 0.6324 - val_aux_output_loss: 0.6511 - val_main_output_acc: 0.6214 - val_aux_output_acc: 0.6408\nEpoch 76/500\n924/924 [==============================] - 0s 222us/step - loss: 0.7170 - main_output_loss: 0.6515 - aux_output_loss: 0.6546 - main_output_acc: 0.6050 - aux_output_acc: 0.6591 - val_loss: 0.6983 - val_main_output_loss: 0.6333 - val_aux_output_loss: 0.6510 - val_main_output_acc: 0.6311 - val_aux_output_acc: 0.6408\nEpoch 77/500\n924/924 [==============================] - 0s 230us/step - loss: 0.7162 - main_output_loss: 0.6508 - aux_output_loss: 0.6542 - main_output_acc: 0.6169 - aux_output_acc: 0.6634 - val_loss: 0.6964 - val_main_output_loss: 0.6313 - val_aux_output_loss: 0.6506 - val_main_output_acc: 0.6311 - val_aux_output_acc: 0.6311\nEpoch 78/500\n924/924 [==============================] - 0s 227us/step - loss: 0.7160 - main_output_loss: 0.6506 - aux_output_loss: 0.6539 - main_output_acc: 0.6115 - aux_output_acc: 0.6602 - val_loss: 0.6976 - val_main_output_loss: 0.6325 - val_aux_output_loss: 0.6505 - val_main_output_acc: 0.6408 - val_aux_output_acc: 0.6408\nEpoch 79/500\n924/924 [==============================] - 0s 229us/step - loss: 0.7150 - main_output_loss: 0.6497 - aux_output_loss: 0.6535 - main_output_acc: 0.6169 - aux_output_acc: 0.6569 - val_loss: 0.6951 - val_main_output_loss: 0.6300 - val_aux_output_loss: 0.6503 - val_main_output_acc: 0.6408 - val_aux_output_acc: 0.6311\nEpoch 80/500\n924/924 [==============================] - 0s 231us/step - loss: 0.7146 - main_output_loss: 0.6493 - aux_output_loss: 0.6532 - main_output_acc: 0.6158 - aux_output_acc: 0.6645 - val_loss: 0.6968 - val_main_output_loss: 0.6318 - val_aux_output_loss: 0.6500 - val_main_output_acc: 0.6214 - val_aux_output_acc: 0.6214\nEpoch 81/500\n924/924 [==============================] - 0s 222us/step - loss: 0.7149 - main_output_loss: 0.6496 - aux_output_loss: 0.6530 - main_output_acc: 0.6115 - aux_output_acc: 0.6613 - val_loss: 0.6957 - val_main_output_loss: 0.6307 - val_aux_output_loss: 0.6497 - val_main_output_acc: 0.6214 - val_aux_output_acc: 0.6214\nEpoch 82/500\n924/924 [==============================] - 0s 221us/step - loss: 0.7140 - main_output_loss: 0.6487 - aux_output_loss: 0.6525 - main_output_acc: 0.6104 - aux_output_acc: 0.6645 - val_loss: 0.6940 - val_main_output_loss: 0.6291 - val_aux_output_loss: 0.6494 - val_main_output_acc: 0.6311 - val_aux_output_acc: 0.6408\nEpoch 83/500\n924/924 [==============================] - 0s 228us/step - loss: 0.7136 - main_output_loss: 0.6484 - aux_output_loss: 0.6522 - main_output_acc: 0.6223 - aux_output_acc: 0.6667 - val_loss: 0.6938 - val_main_output_loss: 0.6289 - val_aux_output_loss: 0.6493 - val_main_output_acc: 0.6408 - val_aux_output_acc: 0.6408\nEpoch 84/500\n924/924 [==============================] - 0s 225us/step - loss: 0.7131 - main_output_loss: 0.6479 - aux_output_loss: 0.6519 - main_output_acc: 0.6245 - aux_output_acc: 0.6688 - val_loss: 0.6935 - val_main_output_loss: 0.6286 - val_aux_output_loss: 0.6491 - val_main_output_acc: 0.6408 - val_aux_output_acc: 0.6311\nEpoch 85/500\n924/924 [==============================] - 0s 227us/step - loss: 0.7126 - main_output_loss: 0.6475 - aux_output_loss: 0.6516 - main_output_acc: 0.6136 - aux_output_acc: 0.6667 - val_loss: 0.6966 - val_main_output_loss: 0.6317 - val_aux_output_loss: 0.6490 - val_main_output_acc: 0.6699 - val_aux_output_acc: 0.6408\nEpoch 86/500\n924/924 [==============================] - 0s 227us/step - loss: 0.7121 - main_output_loss: 0.6470 - aux_output_loss: 0.6512 - main_output_acc: 0.6223 - aux_output_acc: 0.6699 - val_loss: 0.6966 - val_main_output_loss: 0.6317 - val_aux_output_loss: 0.6491 - val_main_output_acc: 0.6893 - val_aux_output_acc: 0.6408\nEpoch 87/500\n924/924 [==============================] - 0s 220us/step - loss: 0.7116 - main_output_loss: 0.6465 - aux_output_loss: 0.6508 - main_output_acc: 0.6374 - aux_output_acc: 0.6667 - val_loss: 0.6952 - val_main_output_loss: 0.6303 - val_aux_output_loss: 0.6490 - val_main_output_acc: 0.6699 - val_aux_output_acc: 0.6408\nEpoch 88/500\n924/924 [==============================] - 0s 219us/step - loss: 0.7114 - main_output_loss: 0.6464 - aux_output_loss: 0.6504 - main_output_acc: 0.6266 - aux_output_acc: 0.6699 - val_loss: 0.6936 - val_main_output_loss: 0.6288 - val_aux_output_loss: 0.6488 - val_main_output_acc: 0.6699 - val_aux_output_acc: 0.6505\nEpoch 89/500\n924/924 [==============================] - 0s 223us/step - loss: 0.7107 - main_output_loss: 0.6457 - aux_output_loss: 0.6500 - main_output_acc: 0.6288 - aux_output_acc: 0.6710 - val_loss: 0.6937 - val_main_output_loss: 0.6289 - val_aux_output_loss: 0.6488 - val_main_output_acc: 0.6796 - val_aux_output_acc: 0.6602\nEpoch 90/500\n924/924 [==============================] - 0s 229us/step - loss: 0.7098 - main_output_loss: 0.6448 - aux_output_loss: 0.6497 - main_output_acc: 0.6353 - aux_output_acc: 0.6602 - val_loss: 0.6912 - val_main_output_loss: 0.6264 - val_aux_output_loss: 0.6484 - val_main_output_acc: 0.6505 - val_aux_output_acc: 0.6602\nEpoch 91/500\n924/924 [==============================] - 0s 227us/step - loss: 0.7097 - main_output_loss: 0.6448 - aux_output_loss: 0.6494 - main_output_acc: 0.6158 - aux_output_acc: 0.6645 - val_loss: 0.6901 - val_main_output_loss: 0.6253 - val_aux_output_loss: 0.6481 - val_main_output_acc: 0.6311 - val_aux_output_acc: 0.6408\nEpoch 92/500\n924/924 [==============================] - 0s 224us/step - loss: 0.7089 - main_output_loss: 0.6440 - aux_output_loss: 0.6490 - main_output_acc: 0.6364 - aux_output_acc: 0.6721 - val_loss: 0.6904 - val_main_output_loss: 0.6256 - val_aux_output_loss: 0.6480 - val_main_output_acc: 0.6311 - val_aux_output_acc: 0.6602\nEpoch 93/500\n924/924 [==============================] - 0s 221us/step - loss: 0.7090 - main_output_loss: 0.6441 - aux_output_loss: 0.6488 - main_output_acc: 0.6223 - aux_output_acc: 0.6732 - val_loss: 0.6925 - val_main_output_loss: 0.6277 - val_aux_output_loss: 0.6479 - val_main_output_acc: 0.6699 - val_aux_output_acc: 0.6602\nEpoch 94/500\n924/924 [==============================] - 0s 222us/step - loss: 0.7082 - main_output_loss: 0.6434 - aux_output_loss: 0.6484 - main_output_acc: 0.6331 - aux_output_acc: 0.6699 - val_loss: 0.6944 - val_main_output_loss: 0.6296 - val_aux_output_loss: 0.6480 - val_main_output_acc: 0.6893 - val_aux_output_acc: 0.6602\nEpoch 95/500\n924/924 [==============================] - 0s 231us/step - loss: 0.7073 - main_output_loss: 0.6426 - aux_output_loss: 0.6479 - main_output_acc: 0.6472 - aux_output_acc: 0.6623 - val_loss: 0.6936 - val_main_output_loss: 0.6288 - val_aux_output_loss: 0.6479 - val_main_output_acc: 0.6990 - val_aux_output_acc: 0.6602\nEpoch 96/500\n924/924 [==============================] - 0s 227us/step - loss: 0.7072 - main_output_loss: 0.6424 - aux_output_loss: 0.6475 - main_output_acc: 0.6407 - aux_output_acc: 0.6613 - val_loss: 0.6898 - val_main_output_loss: 0.6250 - val_aux_output_loss: 0.6476 - val_main_output_acc: 0.6311 - val_aux_output_acc: 0.6602\nEpoch 97/500\n924/924 [==============================] - 0s 221us/step - loss: 0.7065 - main_output_loss: 0.6417 - aux_output_loss: 0.6473 - main_output_acc: 0.6245 - aux_output_acc: 0.6677 - val_loss: 0.6892 - val_main_output_loss: 0.6245 - val_aux_output_loss: 0.6475 - val_main_output_acc: 0.6602 - val_aux_output_acc: 0.6602\nEpoch 98/500\n924/924 [==============================] - 0s 231us/step - loss: 0.7066 - main_output_loss: 0.6419 - aux_output_loss: 0.6471 - main_output_acc: 0.6288 - aux_output_acc: 0.6677 - val_loss: 0.6933 - val_main_output_loss: 0.6285 - val_aux_output_loss: 0.6476 - val_main_output_acc: 0.6796 - val_aux_output_acc: 0.6602\nEpoch 99/500\n924/924 [==============================] - 0s 216us/step - loss: 0.7057 - main_output_loss: 0.6410 - aux_output_loss: 0.6466 - main_output_acc: 0.6656 - aux_output_acc: 0.6613 - val_loss: 0.6882 - val_main_output_loss: 0.6235 - val_aux_output_loss: 0.6474 - val_main_output_acc: 0.6602 - val_aux_output_acc: 0.6699\nEpoch 100/500\n924/924 [==============================] - 0s 208us/step - loss: 0.7052 - main_output_loss: 0.6405 - aux_output_loss: 0.6464 - main_output_acc: 0.6374 - aux_output_acc: 0.6634 - val_loss: 0.6866 - val_main_output_loss: 0.6219 - val_aux_output_loss: 0.6473 - val_main_output_acc: 0.6602 - val_aux_output_acc: 0.6602\nEpoch 101/500\n924/924 [==============================] - 0s 210us/step - loss: 0.7042 - main_output_loss: 0.6396 - aux_output_loss: 0.6461 - main_output_acc: 0.6342 - aux_output_acc: 0.6699 - val_loss: 0.6903 - val_main_output_loss: 0.6256 - val_aux_output_loss: 0.6473 - val_main_output_acc: 0.6990 - val_aux_output_acc: 0.6602\nEpoch 102/500\n924/924 [==============================] - 0s 209us/step - loss: 0.7040 - main_output_loss: 0.6394 - aux_output_loss: 0.6457 - main_output_acc: 0.6526 - aux_output_acc: 0.6634 - val_loss: 0.6853 - val_main_output_loss: 0.6206 - val_aux_output_loss: 0.6468 - val_main_output_acc: 0.6602 - val_aux_output_acc: 0.6602\nEpoch 103/500\n924/924 [==============================] - 0s 207us/step - loss: 0.7033 - main_output_loss: 0.6388 - aux_output_loss: 0.6454 - main_output_acc: 0.6418 - aux_output_acc: 0.6656 - val_loss: 0.6844 - val_main_output_loss: 0.6197 - val_aux_output_loss: 0.6466 - val_main_output_acc: 0.6505 - val_aux_output_acc: 0.6602\nEpoch 104/500\n924/924 [==============================] - 0s 210us/step - loss: 0.7030 - main_output_loss: 0.6385 - aux_output_loss: 0.6453 - main_output_acc: 0.6374 - aux_output_acc: 0.6667 - val_loss: 0.6869 - val_main_output_loss: 0.6222 - val_aux_output_loss: 0.6468 - val_main_output_acc: 0.6893 - val_aux_output_acc: 0.6602\nEpoch 105/500\n924/924 [==============================] - 0s 211us/step - loss: 0.7021 - main_output_loss: 0.6376 - aux_output_loss: 0.6450 - main_output_acc: 0.6461 - aux_output_acc: 0.6656 - val_loss: 0.6904 - val_main_output_loss: 0.6257 - val_aux_output_loss: 0.6467 - val_main_output_acc: 0.6990 - val_aux_output_acc: 0.6699\nEpoch 106/500\n924/924 [==============================] - 0s 215us/step - loss: 0.7019 - main_output_loss: 0.6374 - aux_output_loss: 0.6446 - main_output_acc: 0.6450 - aux_output_acc: 0.6677 - val_loss: 0.6859 - val_main_output_loss: 0.6213 - val_aux_output_loss: 0.6465 - val_main_output_acc: 0.6699 - val_aux_output_acc: 0.6699\nEpoch 107/500\n924/924 [==============================] - 0s 212us/step - loss: 0.7014 - main_output_loss: 0.6369 - aux_output_loss: 0.6443 - main_output_acc: 0.6461 - aux_output_acc: 0.6699 - val_loss: 0.6888 - val_main_output_loss: 0.6242 - val_aux_output_loss: 0.6467 - val_main_output_acc: 0.6893 - val_aux_output_acc: 0.6505\nEpoch 108/500\n924/924 [==============================] - 0s 212us/step - loss: 0.7011 - main_output_loss: 0.6367 - aux_output_loss: 0.6439 - main_output_acc: 0.6483 - aux_output_acc: 0.6699 - val_loss: 0.6879 - val_main_output_loss: 0.6233 - val_aux_output_loss: 0.6464 - val_main_output_acc: 0.7184 - val_aux_output_acc: 0.6505\nEpoch 109/500\n924/924 [==============================] - 0s 211us/step - loss: 0.7001 - main_output_loss: 0.6358 - aux_output_loss: 0.6436 - main_output_acc: 0.6418 - aux_output_acc: 0.6710 - val_loss: 0.6888 - val_main_output_loss: 0.6241 - val_aux_output_loss: 0.6464 - val_main_output_acc: 0.6990 - val_aux_output_acc: 0.6505\nEpoch 110/500\n924/924 [==============================] - 0s 220us/step - loss: 0.6999 - main_output_loss: 0.6356 - aux_output_loss: 0.6433 - main_output_acc: 0.6656 - aux_output_acc: 0.6710 - val_loss: 0.6853 - val_main_output_loss: 0.6207 - val_aux_output_loss: 0.6461 - val_main_output_acc: 0.6699 - val_aux_output_acc: 0.6602\nEpoch 111/500\n924/924 [==============================] - 0s 209us/step - loss: 0.6992 - main_output_loss: 0.6349 - aux_output_loss: 0.6431 - main_output_acc: 0.6515 - aux_output_acc: 0.6721 - val_loss: 0.6864 - val_main_output_loss: 0.6218 - val_aux_output_loss: 0.6459 - val_main_output_acc: 0.6699 - val_aux_output_acc: 0.6602\nEpoch 112/500\n924/924 [==============================] - 0s 215us/step - loss: 0.6983 - main_output_loss: 0.6340 - aux_output_loss: 0.6427 - main_output_acc: 0.6558 - aux_output_acc: 0.6732 - val_loss: 0.6851 - val_main_output_loss: 0.6205 - val_aux_output_loss: 0.6458 - val_main_output_acc: 0.6699 - val_aux_output_acc: 0.6505\nEpoch 113/500\n924/924 [==============================] - 0s 209us/step - loss: 0.6980 - main_output_loss: 0.6338 - aux_output_loss: 0.6425 - main_output_acc: 0.6472 - aux_output_acc: 0.6742 - val_loss: 0.6889 - val_main_output_loss: 0.6244 - val_aux_output_loss: 0.6458 - val_main_output_acc: 0.6990 - val_aux_output_acc: 0.6505\nEpoch 114/500\n924/924 [==============================] - 0s 215us/step - loss: 0.6971 - main_output_loss: 0.6329 - aux_output_loss: 0.6420 - main_output_acc: 0.6623 - aux_output_acc: 0.6732 - val_loss: 0.6817 - val_main_output_loss: 0.6172 - val_aux_output_loss: 0.6453 - val_main_output_acc: 0.6796 - val_aux_output_acc: 0.6602\nEpoch 115/500\n924/924 [==============================] - 0s 208us/step - loss: 0.6966 - main_output_loss: 0.6324 - aux_output_loss: 0.6418 - main_output_acc: 0.6429 - aux_output_acc: 0.6775 - val_loss: 0.6816 - val_main_output_loss: 0.6171 - val_aux_output_loss: 0.6454 - val_main_output_acc: 0.6699 - val_aux_output_acc: 0.6602\nEpoch 116/500\n924/924 [==============================] - 0s 207us/step - loss: 0.6964 - main_output_loss: 0.6323 - aux_output_loss: 0.6415 - main_output_acc: 0.6580 - aux_output_acc: 0.6786 - val_loss: 0.6799 - val_main_output_loss: 0.6154 - val_aux_output_loss: 0.6450 - val_main_output_acc: 0.6699 - val_aux_output_acc: 0.6602\nEpoch 117/500\n924/924 [==============================] - 0s 210us/step - loss: 0.6959 - main_output_loss: 0.6318 - aux_output_loss: 0.6412 - main_output_acc: 0.6537 - aux_output_acc: 0.6742 - val_loss: 0.6798 - val_main_output_loss: 0.6154 - val_aux_output_loss: 0.6448 - val_main_output_acc: 0.6699 - val_aux_output_acc: 0.6602\nEpoch 118/500\n924/924 [==============================] - 0s 210us/step - loss: 0.6955 - main_output_loss: 0.6314 - aux_output_loss: 0.6410 - main_output_acc: 0.6526 - aux_output_acc: 0.6753 - val_loss: 0.6783 - val_main_output_loss: 0.6139 - val_aux_output_loss: 0.6445 - val_main_output_acc: 0.6699 - val_aux_output_acc: 0.6699\nEpoch 119/500\n924/924 [==============================] - 0s 215us/step - loss: 0.6954 - main_output_loss: 0.6313 - aux_output_loss: 0.6407 - main_output_acc: 0.6461 - aux_output_acc: 0.6742 - val_loss: 0.6787 - val_main_output_loss: 0.6142 - val_aux_output_loss: 0.6445 - val_main_output_acc: 0.6699 - val_aux_output_acc: 0.6602\nEpoch 120/500\n924/924 [==============================] - 0s 214us/step - loss: 0.6938 - main_output_loss: 0.6297 - aux_output_loss: 0.6405 - main_output_acc: 0.6537 - aux_output_acc: 0.6753 - val_loss: 0.6803 - val_main_output_loss: 0.6159 - val_aux_output_loss: 0.6445 - val_main_output_acc: 0.7087 - val_aux_output_acc: 0.6602\nEpoch 121/500\n924/924 [==============================] - 0s 207us/step - loss: 0.6932 - main_output_loss: 0.6292 - aux_output_loss: 0.6402 - main_output_acc: 0.6580 - aux_output_acc: 0.6764 - val_loss: 0.6805 - val_main_output_loss: 0.6160 - val_aux_output_loss: 0.6447 - val_main_output_acc: 0.7184 - val_aux_output_acc: 0.6505\nEpoch 122/500\n924/924 [==============================] - 0s 212us/step - loss: 0.6927 - main_output_loss: 0.6287 - aux_output_loss: 0.6399 - main_output_acc: 0.6602 - aux_output_acc: 0.6775 - val_loss: 0.6809 - val_main_output_loss: 0.6164 - val_aux_output_loss: 0.6446 - val_main_output_acc: 0.7087 - val_aux_output_acc: 0.6408\nEpoch 123/500\n924/924 [==============================] - 0s 214us/step - loss: 0.6923 - main_output_loss: 0.6284 - aux_output_loss: 0.6396 - main_output_acc: 0.6613 - aux_output_acc: 0.6807 - val_loss: 0.6760 - val_main_output_loss: 0.6116 - val_aux_output_loss: 0.6441 - val_main_output_acc: 0.6699 - val_aux_output_acc: 0.6505\nEpoch 124/500\n924/924 [==============================] - 0s 208us/step - loss: 0.6919 - main_output_loss: 0.6280 - aux_output_loss: 0.6392 - main_output_acc: 0.6613 - aux_output_acc: 0.6818 - val_loss: 0.6794 - val_main_output_loss: 0.6150 - val_aux_output_loss: 0.6442 - val_main_output_acc: 0.7184 - val_aux_output_acc: 0.6505\nEpoch 125/500\n924/924 [==============================] - 0s 208us/step - loss: 0.6907 - main_output_loss: 0.6268 - aux_output_loss: 0.6389 - main_output_acc: 0.6537 - aux_output_acc: 0.6753 - val_loss: 0.6792 - val_main_output_loss: 0.6148 - val_aux_output_loss: 0.6440 - val_main_output_acc: 0.7087 - val_aux_output_acc: 0.6505\nEpoch 126/500\n924/924 [==============================] - 0s 210us/step - loss: 0.6907 - main_output_loss: 0.6268 - aux_output_loss: 0.6386 - main_output_acc: 0.6688 - aux_output_acc: 0.6742 - val_loss: 0.6769 - val_main_output_loss: 0.6125 - val_aux_output_loss: 0.6437 - val_main_output_acc: 0.7184 - val_aux_output_acc: 0.6505\nEpoch 127/500\n924/924 [==============================] - 0s 206us/step - loss: 0.6898 - main_output_loss: 0.6260 - aux_output_loss: 0.6383 - main_output_acc: 0.6623 - aux_output_acc: 0.6753 - val_loss: 0.6790 - val_main_output_loss: 0.6146 - val_aux_output_loss: 0.6438 - val_main_output_acc: 0.7087 - val_aux_output_acc: 0.6505\nEpoch 128/500\n924/924 [==============================] - 0s 207us/step - loss: 0.6898 - main_output_loss: 0.6260 - aux_output_loss: 0.6380 - main_output_acc: 0.6710 - aux_output_acc: 0.6732 - val_loss: 0.6750 - val_main_output_loss: 0.6107 - val_aux_output_loss: 0.6437 - val_main_output_acc: 0.7184 - val_aux_output_acc: 0.6505\nEpoch 129/500\n924/924 [==============================] - 0s 207us/step - loss: 0.6887 - main_output_loss: 0.6250 - aux_output_loss: 0.6377 - main_output_acc: 0.6732 - aux_output_acc: 0.6721 - val_loss: 0.6722 - val_main_output_loss: 0.6078 - val_aux_output_loss: 0.6435 - val_main_output_acc: 0.6699 - val_aux_output_acc: 0.6505\nEpoch 130/500\n924/924 [==============================] - 0s 207us/step - loss: 0.6878 - main_output_loss: 0.6240 - aux_output_loss: 0.6374 - main_output_acc: 0.6688 - aux_output_acc: 0.6710 - val_loss: 0.6762 - val_main_output_loss: 0.6118 - val_aux_output_loss: 0.6434 - val_main_output_acc: 0.7087 - val_aux_output_acc: 0.6408\nEpoch 131/500\n924/924 [==============================] - 0s 205us/step - loss: 0.6882 - main_output_loss: 0.6244 - aux_output_loss: 0.6372 - main_output_acc: 0.6699 - aux_output_acc: 0.6721 - val_loss: 0.6757 - val_main_output_loss: 0.6114 - val_aux_output_loss: 0.6436 - val_main_output_acc: 0.7087 - val_aux_output_acc: 0.6408\nEpoch 132/500\n924/924 [==============================] - 0s 207us/step - loss: 0.6869 - main_output_loss: 0.6232 - aux_output_loss: 0.6368 - main_output_acc: 0.6688 - aux_output_acc: 0.6721 - val_loss: 0.6720 - val_main_output_loss: 0.6077 - val_aux_output_loss: 0.6432 - val_main_output_acc: 0.6893 - val_aux_output_acc: 0.6408\nEpoch 133/500\n924/924 [==============================] - 0s 208us/step - loss: 0.6865 - main_output_loss: 0.6228 - aux_output_loss: 0.6367 - main_output_acc: 0.6645 - aux_output_acc: 0.6699 - val_loss: 0.6724 - val_main_output_loss: 0.6081 - val_aux_output_loss: 0.6430 - val_main_output_acc: 0.7184 - val_aux_output_acc: 0.6408\nEpoch 134/500\n924/924 [==============================] - 0s 211us/step - loss: 0.6857 - main_output_loss: 0.6221 - aux_output_loss: 0.6363 - main_output_acc: 0.6699 - aux_output_acc: 0.6786 - val_loss: 0.6695 - val_main_output_loss: 0.6052 - val_aux_output_loss: 0.6426 - val_main_output_acc: 0.6699 - val_aux_output_acc: 0.6408\nEpoch 135/500\n924/924 [==============================] - 0s 206us/step - loss: 0.6849 - main_output_loss: 0.6213 - aux_output_loss: 0.6360 - main_output_acc: 0.6613 - aux_output_acc: 0.6797 - val_loss: 0.6769 - val_main_output_loss: 0.6126 - val_aux_output_loss: 0.6427 - val_main_output_acc: 0.7087 - val_aux_output_acc: 0.6408\nEpoch 136/500\n924/924 [==============================] - 0s 208us/step - loss: 0.6848 - main_output_loss: 0.6212 - aux_output_loss: 0.6357 - main_output_acc: 0.6840 - aux_output_acc: 0.6721 - val_loss: 0.6684 - val_main_output_loss: 0.6042 - val_aux_output_loss: 0.6422 - val_main_output_acc: 0.6699 - val_aux_output_acc: 0.6408\nEpoch 137/500\n924/924 [==============================] - 0s 215us/step - loss: 0.6845 - main_output_loss: 0.6210 - aux_output_loss: 0.6355 - main_output_acc: 0.6742 - aux_output_acc: 0.6753 - val_loss: 0.6682 - val_main_output_loss: 0.6040 - val_aux_output_loss: 0.6420 - val_main_output_acc: 0.6796 - val_aux_output_acc: 0.6505\nEpoch 138/500\n924/924 [==============================] - 0s 207us/step - loss: 0.6836 - main_output_loss: 0.6201 - aux_output_loss: 0.6353 - main_output_acc: 0.6645 - aux_output_acc: 0.6797 - val_loss: 0.6679 - val_main_output_loss: 0.6037 - val_aux_output_loss: 0.6418 - val_main_output_acc: 0.6796 - val_aux_output_acc: 0.6602\nEpoch 139/500\n924/924 [==============================] - 0s 205us/step - loss: 0.6827 - main_output_loss: 0.6192 - aux_output_loss: 0.6349 - main_output_acc: 0.6580 - aux_output_acc: 0.6775 - val_loss: 0.6670 - val_main_output_loss: 0.6029 - val_aux_output_loss: 0.6416 - val_main_output_acc: 0.6796 - val_aux_output_acc: 0.6505\nEpoch 140/500\n924/924 [==============================] - 0s 207us/step - loss: 0.6826 - main_output_loss: 0.6191 - aux_output_loss: 0.6348 - main_output_acc: 0.6742 - aux_output_acc: 0.6764 - val_loss: 0.6670 - val_main_output_loss: 0.6029 - val_aux_output_loss: 0.6413 - val_main_output_acc: 0.6893 - val_aux_output_acc: 0.6602\nEpoch 141/500\n924/924 [==============================] - 0s 214us/step - loss: 0.6810 - main_output_loss: 0.6176 - aux_output_loss: 0.6343 - main_output_acc: 0.6753 - aux_output_acc: 0.6775 - val_loss: 0.6662 - val_main_output_loss: 0.6021 - val_aux_output_loss: 0.6410 - val_main_output_acc: 0.6699 - val_aux_output_acc: 0.6699\nEpoch 142/500\n924/924 [==============================] - 0s 209us/step - loss: 0.6811 - main_output_loss: 0.6177 - aux_output_loss: 0.6340 - main_output_acc: 0.6775 - aux_output_acc: 0.6764 - val_loss: 0.6654 - val_main_output_loss: 0.6014 - val_aux_output_loss: 0.6405 - val_main_output_acc: 0.6893 - val_aux_output_acc: 0.6602\nEpoch 143/500\n924/924 [==============================] - 0s 207us/step - loss: 0.6808 - main_output_loss: 0.6174 - aux_output_loss: 0.6339 - main_output_acc: 0.6699 - aux_output_acc: 0.6786 - val_loss: 0.6666 - val_main_output_loss: 0.6026 - val_aux_output_loss: 0.6403 - val_main_output_acc: 0.6990 - val_aux_output_acc: 0.6699\nEpoch 144/500\n924/924 [==============================] - 0s 205us/step - loss: 0.6804 - main_output_loss: 0.6171 - aux_output_loss: 0.6334 - main_output_acc: 0.6786 - aux_output_acc: 0.6807 - val_loss: 0.6658 - val_main_output_loss: 0.6018 - val_aux_output_loss: 0.6402 - val_main_output_acc: 0.7087 - val_aux_output_acc: 0.6699\nEpoch 145/500\n924/924 [==============================] - 0s 211us/step - loss: 0.6793 - main_output_loss: 0.6160 - aux_output_loss: 0.6331 - main_output_acc: 0.6753 - aux_output_acc: 0.6775 - val_loss: 0.6644 - val_main_output_loss: 0.6004 - val_aux_output_loss: 0.6398 - val_main_output_acc: 0.6893 - val_aux_output_acc: 0.6699\nEpoch 146/500\n924/924 [==============================] - 0s 208us/step - loss: 0.6785 - main_output_loss: 0.6153 - aux_output_loss: 0.6327 - main_output_acc: 0.6710 - aux_output_acc: 0.6764 - val_loss: 0.6625 - val_main_output_loss: 0.5986 - val_aux_output_loss: 0.6394 - val_main_output_acc: 0.6990 - val_aux_output_acc: 0.6699\nEpoch 147/500\n924/924 [==============================] - 0s 211us/step - loss: 0.6784 - main_output_loss: 0.6151 - aux_output_loss: 0.6324 - main_output_acc: 0.6742 - aux_output_acc: 0.6732 - val_loss: 0.6617 - val_main_output_loss: 0.5978 - val_aux_output_loss: 0.6391 - val_main_output_acc: 0.6990 - val_aux_output_acc: 0.6699\nEpoch 148/500\n924/924 [==============================] - 0s 213us/step - loss: 0.6773 - main_output_loss: 0.6141 - aux_output_loss: 0.6322 - main_output_acc: 0.6677 - aux_output_acc: 0.6797 - val_loss: 0.6628 - val_main_output_loss: 0.5990 - val_aux_output_loss: 0.6387 - val_main_output_acc: 0.6990 - val_aux_output_acc: 0.6796\nEpoch 149/500\n924/924 [==============================] - 0s 209us/step - loss: 0.6769 - main_output_loss: 0.6137 - aux_output_loss: 0.6318 - main_output_acc: 0.6742 - aux_output_acc: 0.6753 - val_loss: 0.6598 - val_main_output_loss: 0.5960 - val_aux_output_loss: 0.6383 - val_main_output_acc: 0.6990 - val_aux_output_acc: 0.6796\nEpoch 150/500\n924/924 [==============================] - 0s 210us/step - loss: 0.6760 - main_output_loss: 0.6129 - aux_output_loss: 0.6315 - main_output_acc: 0.6861 - aux_output_acc: 0.6818 - val_loss: 0.6593 - val_main_output_loss: 0.5955 - val_aux_output_loss: 0.6380 - val_main_output_acc: 0.6893 - val_aux_output_acc: 0.6796\nEpoch 151/500\n924/924 [==============================] - 0s 206us/step - loss: 0.6759 - main_output_loss: 0.6128 - aux_output_loss: 0.6313 - main_output_acc: 0.6829 - aux_output_acc: 0.6807 - val_loss: 0.6615 - val_main_output_loss: 0.5978 - val_aux_output_loss: 0.6379 - val_main_output_acc: 0.7379 - val_aux_output_acc: 0.6796\nEpoch 152/500\n924/924 [==============================] - 0s 207us/step - loss: 0.6749 - main_output_loss: 0.6118 - aux_output_loss: 0.6308 - main_output_acc: 0.6840 - aux_output_acc: 0.6764 - val_loss: 0.6569 - val_main_output_loss: 0.5931 - val_aux_output_loss: 0.6376 - val_main_output_acc: 0.6893 - val_aux_output_acc: 0.6796\nEpoch 153/500\n924/924 [==============================] - 0s 207us/step - loss: 0.6748 - main_output_loss: 0.6117 - aux_output_loss: 0.6307 - main_output_acc: 0.6818 - aux_output_acc: 0.6818 - val_loss: 0.6568 - val_main_output_loss: 0.5931 - val_aux_output_loss: 0.6372 - val_main_output_acc: 0.7184 - val_aux_output_acc: 0.6796\nEpoch 154/500\n924/924 [==============================] - 0s 207us/step - loss: 0.6742 - main_output_loss: 0.6111 - aux_output_loss: 0.6305 - main_output_acc: 0.6710 - aux_output_acc: 0.6851 - val_loss: 0.6593 - val_main_output_loss: 0.5956 - val_aux_output_loss: 0.6371 - val_main_output_acc: 0.7184 - val_aux_output_acc: 0.6796\nEpoch 155/500\n924/924 [==============================] - 0s 210us/step - loss: 0.6728 - main_output_loss: 0.6098 - aux_output_loss: 0.6301 - main_output_acc: 0.6861 - aux_output_acc: 0.6764 - val_loss: 0.6556 - val_main_output_loss: 0.5919 - val_aux_output_loss: 0.6367 - val_main_output_acc: 0.7087 - val_aux_output_acc: 0.6796\nEpoch 156/500\n924/924 [==============================] - 0s 209us/step - loss: 0.6729 - main_output_loss: 0.6099 - aux_output_loss: 0.6298 - main_output_acc: 0.6786 - aux_output_acc: 0.6851 - val_loss: 0.6587 - val_main_output_loss: 0.5951 - val_aux_output_loss: 0.6364 - val_main_output_acc: 0.7087 - val_aux_output_acc: 0.6796\nEpoch 157/500\n924/924 [==============================] - 0s 209us/step - loss: 0.6720 - main_output_loss: 0.6090 - aux_output_loss: 0.6294 - main_output_acc: 0.6937 - aux_output_acc: 0.6775 - val_loss: 0.6579 - val_main_output_loss: 0.5943 - val_aux_output_loss: 0.6363 - val_main_output_acc: 0.7087 - val_aux_output_acc: 0.6796\nEpoch 158/500\n924/924 [==============================] - 0s 211us/step - loss: 0.6718 - main_output_loss: 0.6089 - aux_output_loss: 0.6292 - main_output_acc: 0.6851 - aux_output_acc: 0.6818 - val_loss: 0.6569 - val_main_output_loss: 0.5932 - val_aux_output_loss: 0.6363 - val_main_output_acc: 0.7184 - val_aux_output_acc: 0.6796\nEpoch 159/500\n924/924 [==============================] - 0s 209us/step - loss: 0.6709 - main_output_loss: 0.6080 - aux_output_loss: 0.6289 - main_output_acc: 0.6818 - aux_output_acc: 0.6786 - val_loss: 0.6552 - val_main_output_loss: 0.5916 - val_aux_output_loss: 0.6360 - val_main_output_acc: 0.6990 - val_aux_output_acc: 0.6796\nEpoch 160/500\n924/924 [==============================] - 0s 209us/step - loss: 0.6703 - main_output_loss: 0.6075 - aux_output_loss: 0.6286 - main_output_acc: 0.6937 - aux_output_acc: 0.6829 - val_loss: 0.6589 - val_main_output_loss: 0.5953 - val_aux_output_loss: 0.6359 - val_main_output_acc: 0.7476 - val_aux_output_acc: 0.6699\nEpoch 161/500\n924/924 [==============================] - 0s 206us/step - loss: 0.6696 - main_output_loss: 0.6068 - aux_output_loss: 0.6281 - main_output_acc: 0.6883 - aux_output_acc: 0.6807 - val_loss: 0.6609 - val_main_output_loss: 0.5973 - val_aux_output_loss: 0.6358 - val_main_output_acc: 0.7282 - val_aux_output_acc: 0.6602\nEpoch 162/500\n924/924 [==============================] - 0s 209us/step - loss: 0.6692 - main_output_loss: 0.6064 - aux_output_loss: 0.6278 - main_output_acc: 0.6926 - aux_output_acc: 0.6753 - val_loss: 0.6559 - val_main_output_loss: 0.5923 - val_aux_output_loss: 0.6354 - val_main_output_acc: 0.7184 - val_aux_output_acc: 0.6699\nEpoch 163/500\n924/924 [==============================] - 0s 208us/step - loss: 0.6689 - main_output_loss: 0.6061 - aux_output_loss: 0.6275 - main_output_acc: 0.6981 - aux_output_acc: 0.6797 - val_loss: 0.6578 - val_main_output_loss: 0.5943 - val_aux_output_loss: 0.6352 - val_main_output_acc: 0.7573 - val_aux_output_acc: 0.6699\nEpoch 164/500\n924/924 [==============================] - 0s 208us/step - loss: 0.6675 - main_output_loss: 0.6048 - aux_output_loss: 0.6271 - main_output_acc: 0.6981 - aux_output_acc: 0.6764 - val_loss: 0.6542 - val_main_output_loss: 0.5907 - val_aux_output_loss: 0.6348 - val_main_output_acc: 0.7087 - val_aux_output_acc: 0.6699\nEpoch 165/500\n924/924 [==============================] - 0s 208us/step - loss: 0.6687 - main_output_loss: 0.6060 - aux_output_loss: 0.6271 - main_output_acc: 0.6872 - aux_output_acc: 0.6797 - val_loss: 0.6517 - val_main_output_loss: 0.5883 - val_aux_output_loss: 0.6343 - val_main_output_acc: 0.7087 - val_aux_output_acc: 0.6699\nEpoch 166/500\n924/924 [==============================] - 0s 209us/step - loss: 0.6665 - main_output_loss: 0.6038 - aux_output_loss: 0.6265 - main_output_acc: 0.6959 - aux_output_acc: 0.6807 - val_loss: 0.6519 - val_main_output_loss: 0.5885 - val_aux_output_loss: 0.6341 - val_main_output_acc: 0.7282 - val_aux_output_acc: 0.6699\nEpoch 167/500\n924/924 [==============================] - 0s 210us/step - loss: 0.6660 - main_output_loss: 0.6034 - aux_output_loss: 0.6262 - main_output_acc: 0.7002 - aux_output_acc: 0.6818 - val_loss: 0.6544 - val_main_output_loss: 0.5910 - val_aux_output_loss: 0.6340 - val_main_output_acc: 0.7476 - val_aux_output_acc: 0.6699\nEpoch 168/500\n924/924 [==============================] - 0s 208us/step - loss: 0.6652 - main_output_loss: 0.6026 - aux_output_loss: 0.6258 - main_output_acc: 0.7035 - aux_output_acc: 0.6818 - val_loss: 0.6529 - val_main_output_loss: 0.5895 - val_aux_output_loss: 0.6338 - val_main_output_acc: 0.7476 - val_aux_output_acc: 0.6699\nEpoch 169/500\n924/924 [==============================] - 0s 213us/step - loss: 0.6654 - main_output_loss: 0.6029 - aux_output_loss: 0.6256 - main_output_acc: 0.6905 - aux_output_acc: 0.6786 - val_loss: 0.6541 - val_main_output_loss: 0.5907 - val_aux_output_loss: 0.6337 - val_main_output_acc: 0.7282 - val_aux_output_acc: 0.6699\nEpoch 170/500\n924/924 [==============================] - 0s 210us/step - loss: 0.6642 - main_output_loss: 0.6017 - aux_output_loss: 0.6251 - main_output_acc: 0.6981 - aux_output_acc: 0.6807 - val_loss: 0.6476 - val_main_output_loss: 0.5843 - val_aux_output_loss: 0.6332 - val_main_output_acc: 0.7184 - val_aux_output_acc: 0.6699\nEpoch 171/500\n924/924 [==============================] - 0s 210us/step - loss: 0.6639 - main_output_loss: 0.6014 - aux_output_loss: 0.6249 - main_output_acc: 0.7056 - aux_output_acc: 0.6797 - val_loss: 0.6465 - val_main_output_loss: 0.5832 - val_aux_output_loss: 0.6328 - val_main_output_acc: 0.7184 - val_aux_output_acc: 0.6699\nEpoch 172/500\n924/924 [==============================] - 0s 213us/step - loss: 0.6632 - main_output_loss: 0.6007 - aux_output_loss: 0.6245 - main_output_acc: 0.6916 - aux_output_acc: 0.6818 - val_loss: 0.6522 - val_main_output_loss: 0.5889 - val_aux_output_loss: 0.6329 - val_main_output_acc: 0.7379 - val_aux_output_acc: 0.6699\nEpoch 173/500\n924/924 [==============================] - 0s 207us/step - loss: 0.6625 - main_output_loss: 0.6001 - aux_output_loss: 0.6241 - main_output_acc: 0.7045 - aux_output_acc: 0.6807 - val_loss: 0.6466 - val_main_output_loss: 0.5834 - val_aux_output_loss: 0.6324 - val_main_output_acc: 0.7476 - val_aux_output_acc: 0.6699\nEpoch 174/500\n924/924 [==============================] - 0s 204us/step - loss: 0.6612 - main_output_loss: 0.5988 - aux_output_loss: 0.6237 - main_output_acc: 0.7045 - aux_output_acc: 0.6829 - val_loss: 0.6464 - val_main_output_loss: 0.5831 - val_aux_output_loss: 0.6321 - val_main_output_acc: 0.7282 - val_aux_output_acc: 0.6699\nEpoch 175/500\n924/924 [==============================] - 0s 215us/step - loss: 0.6606 - main_output_loss: 0.5982 - aux_output_loss: 0.6233 - main_output_acc: 0.7067 - aux_output_acc: 0.6818 - val_loss: 0.6491 - val_main_output_loss: 0.5859 - val_aux_output_loss: 0.6320 - val_main_output_acc: 0.7476 - val_aux_output_acc: 0.6602\nEpoch 176/500\n924/924 [==============================] - 0s 208us/step - loss: 0.6609 - main_output_loss: 0.5986 - aux_output_loss: 0.6230 - main_output_acc: 0.7056 - aux_output_acc: 0.6851 - val_loss: 0.6481 - val_main_output_loss: 0.5849 - val_aux_output_loss: 0.6316 - val_main_output_acc: 0.7476 - val_aux_output_acc: 0.6699\nEpoch 177/500\n924/924 [==============================] - 0s 208us/step - loss: 0.6602 - main_output_loss: 0.5979 - aux_output_loss: 0.6227 - main_output_acc: 0.7078 - aux_output_acc: 0.6840 - val_loss: 0.6432 - val_main_output_loss: 0.5801 - val_aux_output_loss: 0.6310 - val_main_output_acc: 0.7184 - val_aux_output_acc: 0.6796\nEpoch 178/500\n924/924 [==============================] - 0s 213us/step - loss: 0.6592 - main_output_loss: 0.5969 - aux_output_loss: 0.6223 - main_output_acc: 0.6937 - aux_output_acc: 0.6840 - val_loss: 0.6444 - val_main_output_loss: 0.5813 - val_aux_output_loss: 0.6307 - val_main_output_acc: 0.7379 - val_aux_output_acc: 0.6796\nEpoch 179/500\n924/924 [==============================] - 0s 213us/step - loss: 0.6583 - main_output_loss: 0.5961 - aux_output_loss: 0.6220 - main_output_acc: 0.7067 - aux_output_acc: 0.6829 - val_loss: 0.6461 - val_main_output_loss: 0.5830 - val_aux_output_loss: 0.6305 - val_main_output_acc: 0.7476 - val_aux_output_acc: 0.6699\nEpoch 180/500\n924/924 [==============================] - 0s 206us/step - loss: 0.6580 - main_output_loss: 0.5958 - aux_output_loss: 0.6215 - main_output_acc: 0.7045 - aux_output_acc: 0.6818 - val_loss: 0.6480 - val_main_output_loss: 0.5849 - val_aux_output_loss: 0.6303 - val_main_output_acc: 0.7476 - val_aux_output_acc: 0.6699\nEpoch 181/500\n924/924 [==============================] - 0s 208us/step - loss: 0.6577 - main_output_loss: 0.5956 - aux_output_loss: 0.6211 - main_output_acc: 0.7067 - aux_output_acc: 0.6818 - val_loss: 0.6426 - val_main_output_loss: 0.5796 - val_aux_output_loss: 0.6296 - val_main_output_acc: 0.7476 - val_aux_output_acc: 0.6796\nEpoch 182/500\n924/924 [==============================] - 0s 209us/step - loss: 0.6565 - main_output_loss: 0.5944 - aux_output_loss: 0.6208 - main_output_acc: 0.7045 - aux_output_acc: 0.6851 - val_loss: 0.6406 - val_main_output_loss: 0.5777 - val_aux_output_loss: 0.6290 - val_main_output_acc: 0.7087 - val_aux_output_acc: 0.6796\nEpoch 183/500\n924/924 [==============================] - 0s 205us/step - loss: 0.6565 - main_output_loss: 0.5944 - aux_output_loss: 0.6205 - main_output_acc: 0.6872 - aux_output_acc: 0.6861 - val_loss: 0.6446 - val_main_output_loss: 0.5817 - val_aux_output_loss: 0.6290 - val_main_output_acc: 0.7476 - val_aux_output_acc: 0.6796\nEpoch 184/500\n924/924 [==============================] - 0s 213us/step - loss: 0.6550 - main_output_loss: 0.5930 - aux_output_loss: 0.6200 - main_output_acc: 0.7154 - aux_output_acc: 0.6829 - val_loss: 0.6394 - val_main_output_loss: 0.5766 - val_aux_output_loss: 0.6284 - val_main_output_acc: 0.7573 - val_aux_output_acc: 0.6796\nEpoch 185/500\n924/924 [==============================] - 0s 212us/step - loss: 0.6548 - main_output_loss: 0.5928 - aux_output_loss: 0.6198 - main_output_acc: 0.7024 - aux_output_acc: 0.6840 - val_loss: 0.6382 - val_main_output_loss: 0.5754 - val_aux_output_loss: 0.6281 - val_main_output_acc: 0.7184 - val_aux_output_acc: 0.6796\nEpoch 186/500\n924/924 [==============================] - 0s 206us/step - loss: 0.6546 - main_output_loss: 0.5927 - aux_output_loss: 0.6195 - main_output_acc: 0.6991 - aux_output_acc: 0.6851 - val_loss: 0.6378 - val_main_output_loss: 0.5750 - val_aux_output_loss: 0.6277 - val_main_output_acc: 0.7087 - val_aux_output_acc: 0.6796\nEpoch 187/500\n924/924 [==============================] - 0s 204us/step - loss: 0.6541 - main_output_loss: 0.5922 - aux_output_loss: 0.6192 - main_output_acc: 0.6916 - aux_output_acc: 0.6840 - val_loss: 0.6378 - val_main_output_loss: 0.5751 - val_aux_output_loss: 0.6274 - val_main_output_acc: 0.7476 - val_aux_output_acc: 0.6796\nEpoch 188/500\n924/924 [==============================] - 0s 211us/step - loss: 0.6529 - main_output_loss: 0.5911 - aux_output_loss: 0.6188 - main_output_acc: 0.7165 - aux_output_acc: 0.6861 - val_loss: 0.6399 - val_main_output_loss: 0.5772 - val_aux_output_loss: 0.6273 - val_main_output_acc: 0.7379 - val_aux_output_acc: 0.6796\nEpoch 189/500\n924/924 [==============================] - 0s 210us/step - loss: 0.6523 - main_output_loss: 0.5905 - aux_output_loss: 0.6184 - main_output_acc: 0.7154 - aux_output_acc: 0.6829 - val_loss: 0.6384 - val_main_output_loss: 0.5757 - val_aux_output_loss: 0.6270 - val_main_output_acc: 0.7379 - val_aux_output_acc: 0.6796\nEpoch 190/500\n924/924 [==============================] - 0s 204us/step - loss: 0.6524 - main_output_loss: 0.5906 - aux_output_loss: 0.6180 - main_output_acc: 0.7078 - aux_output_acc: 0.6840 - val_loss: 0.6353 - val_main_output_loss: 0.5726 - val_aux_output_loss: 0.6263 - val_main_output_acc: 0.7476 - val_aux_output_acc: 0.6796\nEpoch 191/500\n924/924 [==============================] - 0s 217us/step - loss: 0.6517 - main_output_loss: 0.5900 - aux_output_loss: 0.6177 - main_output_acc: 0.7121 - aux_output_acc: 0.6840 - val_loss: 0.6341 - val_main_output_loss: 0.5715 - val_aux_output_loss: 0.6260 - val_main_output_acc: 0.7379 - val_aux_output_acc: 0.6796\nEpoch 192/500\n924/924 [==============================] - 0s 209us/step - loss: 0.6502 - main_output_loss: 0.5885 - aux_output_loss: 0.6173 - main_output_acc: 0.7110 - aux_output_acc: 0.6872 - val_loss: 0.6367 - val_main_output_loss: 0.5742 - val_aux_output_loss: 0.6258 - val_main_output_acc: 0.7476 - val_aux_output_acc: 0.6796\nEpoch 193/500\n924/924 [==============================] - 0s 210us/step - loss: 0.6497 - main_output_loss: 0.5880 - aux_output_loss: 0.6168 - main_output_acc: 0.7208 - aux_output_acc: 0.6861 - val_loss: 0.6327 - val_main_output_loss: 0.5702 - val_aux_output_loss: 0.6253 - val_main_output_acc: 0.7476 - val_aux_output_acc: 0.6796\nEpoch 194/500\n924/924 [==============================] - 0s 211us/step - loss: 0.6497 - main_output_loss: 0.5880 - aux_output_loss: 0.6166 - main_output_acc: 0.7100 - aux_output_acc: 0.6872 - val_loss: 0.6318 - val_main_output_loss: 0.5693 - val_aux_output_loss: 0.6248 - val_main_output_acc: 0.7379 - val_aux_output_acc: 0.6796\nEpoch 195/500\n924/924 [==============================] - 0s 212us/step - loss: 0.6490 - main_output_loss: 0.5874 - aux_output_loss: 0.6162 - main_output_acc: 0.7056 - aux_output_acc: 0.6872 - val_loss: 0.6326 - val_main_output_loss: 0.5702 - val_aux_output_loss: 0.6245 - val_main_output_acc: 0.7476 - val_aux_output_acc: 0.6796\nEpoch 196/500\n924/924 [==============================] - 0s 209us/step - loss: 0.6477 - main_output_loss: 0.5861 - aux_output_loss: 0.6159 - main_output_acc: 0.7154 - aux_output_acc: 0.6861 - val_loss: 0.6338 - val_main_output_loss: 0.5714 - val_aux_output_loss: 0.6242 - val_main_output_acc: 0.7476 - val_aux_output_acc: 0.6796\nEpoch 197/500\n924/924 [==============================] - 0s 205us/step - loss: 0.6471 - main_output_loss: 0.5855 - aux_output_loss: 0.6155 - main_output_acc: 0.7229 - aux_output_acc: 0.6894 - val_loss: 0.6314 - val_main_output_loss: 0.5690 - val_aux_output_loss: 0.6236 - val_main_output_acc: 0.7476 - val_aux_output_acc: 0.6796\nEpoch 198/500\n924/924 [==============================] - 0s 206us/step - loss: 0.6465 - main_output_loss: 0.5850 - aux_output_loss: 0.6150 - main_output_acc: 0.7100 - aux_output_acc: 0.6894 - val_loss: 0.6372 - val_main_output_loss: 0.5748 - val_aux_output_loss: 0.6234 - val_main_output_acc: 0.7476 - val_aux_output_acc: 0.6796\nEpoch 199/500\n924/924 [==============================] - 0s 208us/step - loss: 0.6455 - main_output_loss: 0.5841 - aux_output_loss: 0.6144 - main_output_acc: 0.7165 - aux_output_acc: 0.6894 - val_loss: 0.6311 - val_main_output_loss: 0.5688 - val_aux_output_loss: 0.6229 - val_main_output_acc: 0.7476 - val_aux_output_acc: 0.6796\nEpoch 200/500\n924/924 [==============================] - 0s 208us/step - loss: 0.6453 - main_output_loss: 0.5839 - aux_output_loss: 0.6142 - main_output_acc: 0.7110 - aux_output_acc: 0.6905 - val_loss: 0.6313 - val_main_output_loss: 0.5691 - val_aux_output_loss: 0.6226 - val_main_output_acc: 0.7670 - val_aux_output_acc: 0.6699\nEpoch 201/500\n924/924 [==============================] - 0s 209us/step - loss: 0.6447 - main_output_loss: 0.5833 - aux_output_loss: 0.6137 - main_output_acc: 0.7262 - aux_output_acc: 0.6916 - val_loss: 0.6297 - val_main_output_loss: 0.5674 - val_aux_output_loss: 0.6223 - val_main_output_acc: 0.7670 - val_aux_output_acc: 0.6796\nEpoch 202/500\n924/924 [==============================] - 0s 212us/step - loss: 0.6434 - main_output_loss: 0.5820 - aux_output_loss: 0.6133 - main_output_acc: 0.7229 - aux_output_acc: 0.6937 - val_loss: 0.6325 - val_main_output_loss: 0.5703 - val_aux_output_loss: 0.6222 - val_main_output_acc: 0.7476 - val_aux_output_acc: 0.6699\nEpoch 203/500\n924/924 [==============================] - 0s 207us/step - loss: 0.6427 - main_output_loss: 0.5815 - aux_output_loss: 0.6128 - main_output_acc: 0.7197 - aux_output_acc: 0.6959 - val_loss: 0.6342 - val_main_output_loss: 0.5720 - val_aux_output_loss: 0.6220 - val_main_output_acc: 0.7379 - val_aux_output_acc: 0.6699\nEpoch 204/500\n924/924 [==============================] - 0s 208us/step - loss: 0.6426 - main_output_loss: 0.5813 - aux_output_loss: 0.6124 - main_output_acc: 0.7100 - aux_output_acc: 0.6883 - val_loss: 0.6277 - val_main_output_loss: 0.5655 - val_aux_output_loss: 0.6215 - val_main_output_acc: 0.7670 - val_aux_output_acc: 0.6699\nEpoch 205/500\n924/924 [==============================] - 0s 211us/step - loss: 0.6421 - main_output_loss: 0.5809 - aux_output_loss: 0.6121 - main_output_acc: 0.7208 - aux_output_acc: 0.6970 - val_loss: 0.6285 - val_main_output_loss: 0.5664 - val_aux_output_loss: 0.6212 - val_main_output_acc: 0.7282 - val_aux_output_acc: 0.6699\nEpoch 206/500\n924/924 [==============================] - 0s 204us/step - loss: 0.6407 - main_output_loss: 0.5796 - aux_output_loss: 0.6115 - main_output_acc: 0.7240 - aux_output_acc: 0.6959 - val_loss: 0.6281 - val_main_output_loss: 0.5660 - val_aux_output_loss: 0.6207 - val_main_output_acc: 0.7476 - val_aux_output_acc: 0.6699\nEpoch 207/500\n924/924 [==============================] - 0s 210us/step - loss: 0.6408 - main_output_loss: 0.5797 - aux_output_loss: 0.6111 - main_output_acc: 0.7208 - aux_output_acc: 0.6981 - val_loss: 0.6285 - val_main_output_loss: 0.5665 - val_aux_output_loss: 0.6204 - val_main_output_acc: 0.7476 - val_aux_output_acc: 0.6796\nEpoch 208/500\n924/924 [==============================] - 0s 205us/step - loss: 0.6394 - main_output_loss: 0.5783 - aux_output_loss: 0.6106 - main_output_acc: 0.7262 - aux_output_acc: 0.6981 - val_loss: 0.6229 - val_main_output_loss: 0.5609 - val_aux_output_loss: 0.6199 - val_main_output_acc: 0.7670 - val_aux_output_acc: 0.6893\nEpoch 209/500\n924/924 [==============================] - 0s 209us/step - loss: 0.6382 - main_output_loss: 0.5772 - aux_output_loss: 0.6101 - main_output_acc: 0.7273 - aux_output_acc: 0.6991 - val_loss: 0.6243 - val_main_output_loss: 0.5623 - val_aux_output_loss: 0.6194 - val_main_output_acc: 0.7670 - val_aux_output_acc: 0.6796\nEpoch 210/500\n924/924 [==============================] - 0s 209us/step - loss: 0.6381 - main_output_loss: 0.5771 - aux_output_loss: 0.6097 - main_output_acc: 0.7294 - aux_output_acc: 0.6959 - val_loss: 0.6224 - val_main_output_loss: 0.5605 - val_aux_output_loss: 0.6188 - val_main_output_acc: 0.7670 - val_aux_output_acc: 0.6893\nEpoch 211/500\n924/924 [==============================] - 0s 206us/step - loss: 0.6366 - main_output_loss: 0.5756 - aux_output_loss: 0.6092 - main_output_acc: 0.7208 - aux_output_acc: 0.7013 - val_loss: 0.6225 - val_main_output_loss: 0.5606 - val_aux_output_loss: 0.6185 - val_main_output_acc: 0.7476 - val_aux_output_acc: 0.6893\nEpoch 212/500\n924/924 [==============================] - 0s 215us/step - loss: 0.6364 - main_output_loss: 0.5755 - aux_output_loss: 0.6088 - main_output_acc: 0.7175 - aux_output_acc: 0.7002 - val_loss: 0.6230 - val_main_output_loss: 0.5612 - val_aux_output_loss: 0.6181 - val_main_output_acc: 0.7476 - val_aux_output_acc: 0.6893\nEpoch 213/500\n924/924 [==============================] - 0s 206us/step - loss: 0.6353 - main_output_loss: 0.5745 - aux_output_loss: 0.6083 - main_output_acc: 0.7273 - aux_output_acc: 0.6991 - val_loss: 0.6236 - val_main_output_loss: 0.5618 - val_aux_output_loss: 0.6177 - val_main_output_acc: 0.7476 - val_aux_output_acc: 0.6893\nEpoch 214/500\n924/924 [==============================] - 0s 208us/step - loss: 0.6344 - main_output_loss: 0.5736 - aux_output_loss: 0.6077 - main_output_acc: 0.7305 - aux_output_acc: 0.6981 - val_loss: 0.6195 - val_main_output_loss: 0.5578 - val_aux_output_loss: 0.6172 - val_main_output_acc: 0.7573 - val_aux_output_acc: 0.6893\nEpoch 215/500\n924/924 [==============================] - 0s 210us/step - loss: 0.6340 - main_output_loss: 0.5732 - aux_output_loss: 0.6073 - main_output_acc: 0.7316 - aux_output_acc: 0.6970 - val_loss: 0.6188 - val_main_output_loss: 0.5572 - val_aux_output_loss: 0.6167 - val_main_output_acc: 0.7670 - val_aux_output_acc: 0.6893\nEpoch 216/500\n924/924 [==============================] - 0s 209us/step - loss: 0.6328 - main_output_loss: 0.5721 - aux_output_loss: 0.6070 - main_output_acc: 0.7273 - aux_output_acc: 0.7013 - val_loss: 0.6188 - val_main_output_loss: 0.5571 - val_aux_output_loss: 0.6163 - val_main_output_acc: 0.7476 - val_aux_output_acc: 0.6893\nEpoch 217/500\n924/924 [==============================] - 0s 206us/step - loss: 0.6319 - main_output_loss: 0.5712 - aux_output_loss: 0.6065 - main_output_acc: 0.7305 - aux_output_acc: 0.7002 - val_loss: 0.6173 - val_main_output_loss: 0.5557 - val_aux_output_loss: 0.6157 - val_main_output_acc: 0.7573 - val_aux_output_acc: 0.6990\nEpoch 218/500\n924/924 [==============================] - 0s 205us/step - loss: 0.6317 - main_output_loss: 0.5711 - aux_output_loss: 0.6060 - main_output_acc: 0.7251 - aux_output_acc: 0.7002 - val_loss: 0.6157 - val_main_output_loss: 0.5542 - val_aux_output_loss: 0.6150 - val_main_output_acc: 0.7767 - val_aux_output_acc: 0.6990\nEpoch 219/500\n924/924 [==============================] - 0s 207us/step - loss: 0.6310 - main_output_loss: 0.5705 - aux_output_loss: 0.6055 - main_output_acc: 0.7262 - aux_output_acc: 0.7013 - val_loss: 0.6188 - val_main_output_loss: 0.5573 - val_aux_output_loss: 0.6148 - val_main_output_acc: 0.7573 - val_aux_output_acc: 0.6990\nEpoch 220/500\n924/924 [==============================] - 0s 206us/step - loss: 0.6296 - main_output_loss: 0.5691 - aux_output_loss: 0.6050 - main_output_acc: 0.7348 - aux_output_acc: 0.6970 - val_loss: 0.6138 - val_main_output_loss: 0.5524 - val_aux_output_loss: 0.6140 - val_main_output_acc: 0.7476 - val_aux_output_acc: 0.6893\nEpoch 221/500\n924/924 [==============================] - 0s 216us/step - loss: 0.6299 - main_output_loss: 0.5694 - aux_output_loss: 0.6048 - main_output_acc: 0.7208 - aux_output_acc: 0.7035 - val_loss: 0.6136 - val_main_output_loss: 0.5523 - val_aux_output_loss: 0.6136 - val_main_output_acc: 0.7573 - val_aux_output_acc: 0.6990\nEpoch 222/500\n924/924 [==============================] - 0s 206us/step - loss: 0.6287 - main_output_loss: 0.5682 - aux_output_loss: 0.6045 - main_output_acc: 0.7219 - aux_output_acc: 0.7013 - val_loss: 0.6194 - val_main_output_loss: 0.5580 - val_aux_output_loss: 0.6135 - val_main_output_acc: 0.7379 - val_aux_output_acc: 0.6990\nEpoch 223/500\n924/924 [==============================] - 0s 211us/step - loss: 0.6281 - main_output_loss: 0.5677 - aux_output_loss: 0.6039 - main_output_acc: 0.7424 - aux_output_acc: 0.6970 - val_loss: 0.6129 - val_main_output_loss: 0.5516 - val_aux_output_loss: 0.6128 - val_main_output_acc: 0.7670 - val_aux_output_acc: 0.6990\nEpoch 224/500\n924/924 [==============================] - 0s 209us/step - loss: 0.6269 - main_output_loss: 0.5665 - aux_output_loss: 0.6036 - main_output_acc: 0.7240 - aux_output_acc: 0.7035 - val_loss: 0.6166 - val_main_output_loss: 0.5554 - val_aux_output_loss: 0.6126 - val_main_output_acc: 0.7573 - val_aux_output_acc: 0.6990\nEpoch 225/500\n924/924 [==============================] - 0s 207us/step - loss: 0.6262 - main_output_loss: 0.5659 - aux_output_loss: 0.6032 - main_output_acc: 0.7359 - aux_output_acc: 0.6991 - val_loss: 0.6132 - val_main_output_loss: 0.5520 - val_aux_output_loss: 0.6119 - val_main_output_acc: 0.7573 - val_aux_output_acc: 0.6990\nEpoch 226/500\n924/924 [==============================] - 0s 209us/step - loss: 0.6256 - main_output_loss: 0.5653 - aux_output_loss: 0.6028 - main_output_acc: 0.7370 - aux_output_acc: 0.7035 - val_loss: 0.6182 - val_main_output_loss: 0.5570 - val_aux_output_loss: 0.6119 - val_main_output_acc: 0.7476 - val_aux_output_acc: 0.6990\nEpoch 227/500\n924/924 [==============================] - 0s 204us/step - loss: 0.6256 - main_output_loss: 0.5654 - aux_output_loss: 0.6024 - main_output_acc: 0.7370 - aux_output_acc: 0.7035 - val_loss: 0.6131 - val_main_output_loss: 0.5520 - val_aux_output_loss: 0.6114 - val_main_output_acc: 0.7670 - val_aux_output_acc: 0.6990\nEpoch 228/500\n924/924 [==============================] - 0s 208us/step - loss: 0.6249 - main_output_loss: 0.5647 - aux_output_loss: 0.6021 - main_output_acc: 0.7338 - aux_output_acc: 0.7013 - val_loss: 0.6147 - val_main_output_loss: 0.5535 - val_aux_output_loss: 0.6112 - val_main_output_acc: 0.7476 - val_aux_output_acc: 0.6990\nEpoch 229/500\n924/924 [==============================] - 0s 205us/step - loss: 0.6237 - main_output_loss: 0.5635 - aux_output_loss: 0.6015 - main_output_acc: 0.7413 - aux_output_acc: 0.7013 - val_loss: 0.6139 - val_main_output_loss: 0.5528 - val_aux_output_loss: 0.6108 - val_main_output_acc: 0.7573 - val_aux_output_acc: 0.6990\nEpoch 230/500\n924/924 [==============================] - 0s 210us/step - loss: 0.6226 - main_output_loss: 0.5625 - aux_output_loss: 0.6011 - main_output_acc: 0.7403 - aux_output_acc: 0.6948 - val_loss: 0.6065 - val_main_output_loss: 0.5455 - val_aux_output_loss: 0.6097 - val_main_output_acc: 0.7573 - val_aux_output_acc: 0.6990\nEpoch 231/500\n924/924 [==============================] - 0s 216us/step - loss: 0.6222 - main_output_loss: 0.5621 - aux_output_loss: 0.6009 - main_output_acc: 0.7284 - aux_output_acc: 0.7035 - val_loss: 0.6110 - val_main_output_loss: 0.5500 - val_aux_output_loss: 0.6096 - val_main_output_acc: 0.7670 - val_aux_output_acc: 0.6990\nEpoch 232/500\n924/924 [==============================] - 0s 210us/step - loss: 0.6213 - main_output_loss: 0.5612 - aux_output_loss: 0.6006 - main_output_acc: 0.7392 - aux_output_acc: 0.7035 - val_loss: 0.6142 - val_main_output_loss: 0.5532 - val_aux_output_loss: 0.6093 - val_main_output_acc: 0.7476 - val_aux_output_acc: 0.6990\nEpoch 233/500\n924/924 [==============================] - 0s 216us/step - loss: 0.6219 - main_output_loss: 0.5618 - aux_output_loss: 0.6003 - main_output_acc: 0.7316 - aux_output_acc: 0.7024 - val_loss: 0.6120 - val_main_output_loss: 0.5511 - val_aux_output_loss: 0.6088 - val_main_output_acc: 0.7573 - val_aux_output_acc: 0.6990\nEpoch 234/500\n924/924 [==============================] - 0s 209us/step - loss: 0.6211 - main_output_loss: 0.5611 - aux_output_loss: 0.5998 - main_output_acc: 0.7381 - aux_output_acc: 0.7035 - val_loss: 0.6116 - val_main_output_loss: 0.5508 - val_aux_output_loss: 0.6083 - val_main_output_acc: 0.7573 - val_aux_output_acc: 0.6990\nEpoch 235/500\n924/924 [==============================] - 0s 210us/step - loss: 0.6197 - main_output_loss: 0.5597 - aux_output_loss: 0.5993 - main_output_acc: 0.7370 - aux_output_acc: 0.7013 - val_loss: 0.6050 - val_main_output_loss: 0.5443 - val_aux_output_loss: 0.6072 - val_main_output_acc: 0.7573 - val_aux_output_acc: 0.6990\nEpoch 236/500\n924/924 [==============================] - 0s 213us/step - loss: 0.6194 - main_output_loss: 0.5595 - aux_output_loss: 0.5991 - main_output_acc: 0.7424 - aux_output_acc: 0.7045 - val_loss: 0.6035 - val_main_output_loss: 0.5428 - val_aux_output_loss: 0.6067 - val_main_output_acc: 0.7670 - val_aux_output_acc: 0.6990\nEpoch 237/500\n924/924 [==============================] - 0s 209us/step - loss: 0.6187 - main_output_loss: 0.5588 - aux_output_loss: 0.5989 - main_output_acc: 0.7338 - aux_output_acc: 0.7035 - val_loss: 0.6080 - val_main_output_loss: 0.5474 - val_aux_output_loss: 0.6069 - val_main_output_acc: 0.7670 - val_aux_output_acc: 0.6990\nEpoch 238/500\n924/924 [==============================] - 0s 207us/step - loss: 0.6171 - main_output_loss: 0.5573 - aux_output_loss: 0.5984 - main_output_acc: 0.7446 - aux_output_acc: 0.7035 - val_loss: 0.6036 - val_main_output_loss: 0.5430 - val_aux_output_loss: 0.6061 - val_main_output_acc: 0.7476 - val_aux_output_acc: 0.6990\nEpoch 239/500\n924/924 [==============================] - 0s 210us/step - loss: 0.6167 - main_output_loss: 0.5568 - aux_output_loss: 0.5981 - main_output_acc: 0.7392 - aux_output_acc: 0.7024 - val_loss: 0.6058 - val_main_output_loss: 0.5452 - val_aux_output_loss: 0.6059 - val_main_output_acc: 0.7670 - val_aux_output_acc: 0.6990\nEpoch 240/500\n924/924 [==============================] - 0s 209us/step - loss: 0.6160 - main_output_loss: 0.5562 - aux_output_loss: 0.5976 - main_output_acc: 0.7424 - aux_output_acc: 0.7056 - val_loss: 0.6033 - val_main_output_loss: 0.5428 - val_aux_output_loss: 0.6051 - val_main_output_acc: 0.7670 - val_aux_output_acc: 0.6990\nEpoch 241/500\n924/924 [==============================] - 0s 208us/step - loss: 0.6155 - main_output_loss: 0.5558 - aux_output_loss: 0.5973 - main_output_acc: 0.7446 - aux_output_acc: 0.7056 - val_loss: 0.6006 - val_main_output_loss: 0.5401 - val_aux_output_loss: 0.6042 - val_main_output_acc: 0.7476 - val_aux_output_acc: 0.6990\nEpoch 242/500\n924/924 [==============================] - 0s 209us/step - loss: 0.6161 - main_output_loss: 0.5564 - aux_output_loss: 0.5972 - main_output_acc: 0.7359 - aux_output_acc: 0.7067 - val_loss: 0.6017 - val_main_output_loss: 0.5412 - val_aux_output_loss: 0.6041 - val_main_output_acc: 0.7573 - val_aux_output_acc: 0.6990\nEpoch 243/500\n924/924 [==============================] - 0s 209us/step - loss: 0.6140 - main_output_loss: 0.5543 - aux_output_loss: 0.5966 - main_output_acc: 0.7381 - aux_output_acc: 0.7035 - val_loss: 0.6019 - val_main_output_loss: 0.5416 - val_aux_output_loss: 0.6036 - val_main_output_acc: 0.7670 - val_aux_output_acc: 0.6990\nEpoch 244/500\n924/924 [==============================] - 0s 206us/step - loss: 0.6135 - main_output_loss: 0.5539 - aux_output_loss: 0.5962 - main_output_acc: 0.7446 - aux_output_acc: 0.7035 - val_loss: 0.5997 - val_main_output_loss: 0.5393 - val_aux_output_loss: 0.6031 - val_main_output_acc: 0.7670 - val_aux_output_acc: 0.6990\nEpoch 245/500\n924/924 [==============================] - 0s 207us/step - loss: 0.6133 - main_output_loss: 0.5537 - aux_output_loss: 0.5959 - main_output_acc: 0.7403 - aux_output_acc: 0.7035 - val_loss: 0.6045 - val_main_output_loss: 0.5441 - val_aux_output_loss: 0.6032 - val_main_output_acc: 0.7670 - val_aux_output_acc: 0.6893\nEpoch 246/500\n924/924 [==============================] - 0s 208us/step - loss: 0.6121 - main_output_loss: 0.5526 - aux_output_loss: 0.5954 - main_output_acc: 0.7468 - aux_output_acc: 0.7056 - val_loss: 0.5977 - val_main_output_loss: 0.5374 - val_aux_output_loss: 0.6022 - val_main_output_acc: 0.7476 - val_aux_output_acc: 0.6990\nEpoch 247/500\n924/924 [==============================] - 0s 211us/step - loss: 0.6121 - main_output_loss: 0.5526 - aux_output_loss: 0.5953 - main_output_acc: 0.7392 - aux_output_acc: 0.7067 - val_loss: 0.6015 - val_main_output_loss: 0.5413 - val_aux_output_loss: 0.6022 - val_main_output_acc: 0.7767 - val_aux_output_acc: 0.6893\nEpoch 248/500\n924/924 [==============================] - 0s 208us/step - loss: 0.6105 - main_output_loss: 0.5510 - aux_output_loss: 0.5949 - main_output_acc: 0.7424 - aux_output_acc: 0.7067 - val_loss: 0.6034 - val_main_output_loss: 0.5432 - val_aux_output_loss: 0.6020 - val_main_output_acc: 0.7767 - val_aux_output_acc: 0.6796\nEpoch 249/500\n924/924 [==============================] - 0s 208us/step - loss: 0.6101 - main_output_loss: 0.5506 - aux_output_loss: 0.5944 - main_output_acc: 0.7435 - aux_output_acc: 0.7013 - val_loss: 0.5992 - val_main_output_loss: 0.5391 - val_aux_output_loss: 0.6012 - val_main_output_acc: 0.7670 - val_aux_output_acc: 0.6893\nEpoch 250/500\n924/924 [==============================] - 0s 205us/step - loss: 0.6101 - main_output_loss: 0.5506 - aux_output_loss: 0.5943 - main_output_acc: 0.7435 - aux_output_acc: 0.7013 - val_loss: 0.6063 - val_main_output_loss: 0.5462 - val_aux_output_loss: 0.6014 - val_main_output_acc: 0.7476 - val_aux_output_acc: 0.6796\nEpoch 251/500\n924/924 [==============================] - 0s 211us/step - loss: 0.6100 - main_output_loss: 0.5506 - aux_output_loss: 0.5938 - main_output_acc: 0.7392 - aux_output_acc: 0.7045 - val_loss: 0.5992 - val_main_output_loss: 0.5391 - val_aux_output_loss: 0.6007 - val_main_output_acc: 0.7670 - val_aux_output_acc: 0.6796\nEpoch 252/500\n924/924 [==============================] - 0s 209us/step - loss: 0.6079 - main_output_loss: 0.5486 - aux_output_loss: 0.5934 - main_output_acc: 0.7435 - aux_output_acc: 0.7045 - val_loss: 0.5988 - val_main_output_loss: 0.5388 - val_aux_output_loss: 0.6004 - val_main_output_acc: 0.7767 - val_aux_output_acc: 0.6796\nEpoch 253/500\n924/924 [==============================] - 0s 214us/step - loss: 0.6083 - main_output_loss: 0.5490 - aux_output_loss: 0.5933 - main_output_acc: 0.7489 - aux_output_acc: 0.7056 - val_loss: 0.6020 - val_main_output_loss: 0.5419 - val_aux_output_loss: 0.6002 - val_main_output_acc: 0.7670 - val_aux_output_acc: 0.6796\nEpoch 254/500\n924/924 [==============================] - 0s 208us/step - loss: 0.6073 - main_output_loss: 0.5480 - aux_output_loss: 0.5929 - main_output_acc: 0.7500 - aux_output_acc: 0.7056 - val_loss: 0.5998 - val_main_output_loss: 0.5398 - val_aux_output_loss: 0.5999 - val_main_output_acc: 0.7864 - val_aux_output_acc: 0.6796\nEpoch 255/500\n924/924 [==============================] - 0s 209us/step - loss: 0.6068 - main_output_loss: 0.5476 - aux_output_loss: 0.5924 - main_output_acc: 0.7446 - aux_output_acc: 0.7067 - val_loss: 0.5979 - val_main_output_loss: 0.5379 - val_aux_output_loss: 0.5994 - val_main_output_acc: 0.7767 - val_aux_output_acc: 0.6796\nEpoch 256/500\n924/924 [==============================] - 0s 207us/step - loss: 0.6064 - main_output_loss: 0.5472 - aux_output_loss: 0.5923 - main_output_acc: 0.7413 - aux_output_acc: 0.7035 - val_loss: 0.6024 - val_main_output_loss: 0.5425 - val_aux_output_loss: 0.5993 - val_main_output_acc: 0.7573 - val_aux_output_acc: 0.6796\nEpoch 257/500\n924/924 [==============================] - 0s 212us/step - loss: 0.6057 - main_output_loss: 0.5465 - aux_output_loss: 0.5919 - main_output_acc: 0.7478 - aux_output_acc: 0.7045 - val_loss: 0.5965 - val_main_output_loss: 0.5366 - val_aux_output_loss: 0.5986 - val_main_output_acc: 0.7670 - val_aux_output_acc: 0.6796\nEpoch 258/500\n924/924 [==============================] - 0s 209us/step - loss: 0.6041 - main_output_loss: 0.5449 - aux_output_loss: 0.5915 - main_output_acc: 0.7489 - aux_output_acc: 0.7056 - val_loss: 0.5942 - val_main_output_loss: 0.5344 - val_aux_output_loss: 0.5981 - val_main_output_acc: 0.7670 - val_aux_output_acc: 0.6796\nEpoch 259/500\n924/924 [==============================] - 0s 211us/step - loss: 0.6042 - main_output_loss: 0.5451 - aux_output_loss: 0.5913 - main_output_acc: 0.7543 - aux_output_acc: 0.7067 - val_loss: 0.5940 - val_main_output_loss: 0.5342 - val_aux_output_loss: 0.5977 - val_main_output_acc: 0.7670 - val_aux_output_acc: 0.6796\nEpoch 260/500\n924/924 [==============================] - 0s 207us/step - loss: 0.6042 - main_output_loss: 0.5451 - aux_output_loss: 0.5912 - main_output_acc: 0.7500 - aux_output_acc: 0.7067 - val_loss: 0.5903 - val_main_output_loss: 0.5306 - val_aux_output_loss: 0.5969 - val_main_output_acc: 0.7767 - val_aux_output_acc: 0.6893\nEpoch 261/500\n924/924 [==============================] - 0s 207us/step - loss: 0.6025 - main_output_loss: 0.5434 - aux_output_loss: 0.5907 - main_output_acc: 0.7522 - aux_output_acc: 0.7100 - val_loss: 0.5942 - val_main_output_loss: 0.5345 - val_aux_output_loss: 0.5968 - val_main_output_acc: 0.7864 - val_aux_output_acc: 0.6796\nEpoch 262/500\n924/924 [==============================] - 0s 217us/step - loss: 0.6024 - main_output_loss: 0.5433 - aux_output_loss: 0.5905 - main_output_acc: 0.7532 - aux_output_acc: 0.7100 - val_loss: 0.5974 - val_main_output_loss: 0.5377 - val_aux_output_loss: 0.5968 - val_main_output_acc: 0.7573 - val_aux_output_acc: 0.6796\nEpoch 263/500\n924/924 [==============================] - 0s 211us/step - loss: 0.6016 - main_output_loss: 0.5426 - aux_output_loss: 0.5899 - main_output_acc: 0.7543 - aux_output_acc: 0.7100 - val_loss: 0.5901 - val_main_output_loss: 0.5305 - val_aux_output_loss: 0.5958 - val_main_output_acc: 0.7670 - val_aux_output_acc: 0.6796\nEpoch 264/500\n924/924 [==============================] - 0s 209us/step - loss: 0.6007 - main_output_loss: 0.5418 - aux_output_loss: 0.5898 - main_output_acc: 0.7511 - aux_output_acc: 0.7056 - val_loss: 0.5881 - val_main_output_loss: 0.5285 - val_aux_output_loss: 0.5954 - val_main_output_acc: 0.7767 - val_aux_output_acc: 0.6893\nEpoch 265/500\n924/924 [==============================] - 0s 211us/step - loss: 0.6000 - main_output_loss: 0.5411 - aux_output_loss: 0.5896 - main_output_acc: 0.7478 - aux_output_acc: 0.7078 - val_loss: 0.5936 - val_main_output_loss: 0.5340 - val_aux_output_loss: 0.5955 - val_main_output_acc: 0.7864 - val_aux_output_acc: 0.6796\nEpoch 266/500\n924/924 [==============================] - 0s 206us/step - loss: 0.6003 - main_output_loss: 0.5414 - aux_output_loss: 0.5892 - main_output_acc: 0.7468 - aux_output_acc: 0.7089 - val_loss: 0.5933 - val_main_output_loss: 0.5338 - val_aux_output_loss: 0.5951 - val_main_output_acc: 0.7864 - val_aux_output_acc: 0.6796\nEpoch 267/500\n924/924 [==============================] - 0s 206us/step - loss: 0.5992 - main_output_loss: 0.5403 - aux_output_loss: 0.5889 - main_output_acc: 0.7468 - aux_output_acc: 0.7121 - val_loss: 0.5891 - val_main_output_loss: 0.5296 - val_aux_output_loss: 0.5944 - val_main_output_acc: 0.7767 - val_aux_output_acc: 0.6796\nEpoch 268/500\n924/924 [==============================] - 0s 219us/step - loss: 0.5976 - main_output_loss: 0.5388 - aux_output_loss: 0.5885 - main_output_acc: 0.7522 - aux_output_acc: 0.7089 - val_loss: 0.5863 - val_main_output_loss: 0.5270 - val_aux_output_loss: 0.5936 - val_main_output_acc: 0.7670 - val_aux_output_acc: 0.6893\nEpoch 269/500\n924/924 [==============================] - 0s 205us/step - loss: 0.5982 - main_output_loss: 0.5394 - aux_output_loss: 0.5884 - main_output_acc: 0.7532 - aux_output_acc: 0.7100 - val_loss: 0.5903 - val_main_output_loss: 0.5309 - val_aux_output_loss: 0.5938 - val_main_output_acc: 0.7670 - val_aux_output_acc: 0.6796\nEpoch 270/500\n924/924 [==============================] - 0s 207us/step - loss: 0.5974 - main_output_loss: 0.5386 - aux_output_loss: 0.5881 - main_output_acc: 0.7511 - aux_output_acc: 0.7067 - val_loss: 0.5860 - val_main_output_loss: 0.5267 - val_aux_output_loss: 0.5930 - val_main_output_acc: 0.7670 - val_aux_output_acc: 0.6893\nEpoch 271/500\n924/924 [==============================] - 0s 211us/step - loss: 0.5961 - main_output_loss: 0.5373 - aux_output_loss: 0.5877 - main_output_acc: 0.7565 - aux_output_acc: 0.7078 - val_loss: 0.5913 - val_main_output_loss: 0.5320 - val_aux_output_loss: 0.5932 - val_main_output_acc: 0.7767 - val_aux_output_acc: 0.6796\nEpoch 272/500\n924/924 [==============================] - 0s 204us/step - loss: 0.5962 - main_output_loss: 0.5374 - aux_output_loss: 0.5874 - main_output_acc: 0.7511 - aux_output_acc: 0.7110 - val_loss: 0.5858 - val_main_output_loss: 0.5265 - val_aux_output_loss: 0.5927 - val_main_output_acc: 0.7767 - val_aux_output_acc: 0.6796\nEpoch 273/500\n924/924 [==============================] - 0s 210us/step - loss: 0.5947 - main_output_loss: 0.5360 - aux_output_loss: 0.5871 - main_output_acc: 0.7554 - aux_output_acc: 0.7089 - val_loss: 0.5868 - val_main_output_loss: 0.5275 - val_aux_output_loss: 0.5923 - val_main_output_acc: 0.7767 - val_aux_output_acc: 0.6796\nEpoch 274/500\n924/924 [==============================] - 0s 207us/step - loss: 0.5950 - main_output_loss: 0.5363 - aux_output_loss: 0.5869 - main_output_acc: 0.7522 - aux_output_acc: 0.7089 - val_loss: 0.5844 - val_main_output_loss: 0.5253 - val_aux_output_loss: 0.5918 - val_main_output_acc: 0.7670 - val_aux_output_acc: 0.6796\nEpoch 275/500\n924/924 [==============================] - 0s 206us/step - loss: 0.5936 - main_output_loss: 0.5349 - aux_output_loss: 0.5866 - main_output_acc: 0.7457 - aux_output_acc: 0.7089 - val_loss: 0.5842 - val_main_output_loss: 0.5251 - val_aux_output_loss: 0.5914 - val_main_output_acc: 0.7670 - val_aux_output_acc: 0.6796\nEpoch 276/500\n924/924 [==============================] - 0s 208us/step - loss: 0.5940 - main_output_loss: 0.5354 - aux_output_loss: 0.5862 - main_output_acc: 0.7457 - aux_output_acc: 0.7110 - val_loss: 0.5806 - val_main_output_loss: 0.5215 - val_aux_output_loss: 0.5908 - val_main_output_acc: 0.7767 - val_aux_output_acc: 0.6893\nEpoch 277/500\n924/924 [==============================] - 0s 213us/step - loss: 0.5929 - main_output_loss: 0.5343 - aux_output_loss: 0.5859 - main_output_acc: 0.7597 - aux_output_acc: 0.7110 - val_loss: 0.5811 - val_main_output_loss: 0.5220 - val_aux_output_loss: 0.5907 - val_main_output_acc: 0.7767 - val_aux_output_acc: 0.6796\nEpoch 278/500\n924/924 [==============================] - 0s 211us/step - loss: 0.5928 - main_output_loss: 0.5342 - aux_output_loss: 0.5857 - main_output_acc: 0.7457 - aux_output_acc: 0.7110 - val_loss: 0.5834 - val_main_output_loss: 0.5243 - val_aux_output_loss: 0.5906 - val_main_output_acc: 0.7864 - val_aux_output_acc: 0.6796\nEpoch 279/500\n924/924 [==============================] - 0s 212us/step - loss: 0.5912 - main_output_loss: 0.5327 - aux_output_loss: 0.5852 - main_output_acc: 0.7522 - aux_output_acc: 0.7121 - val_loss: 0.5836 - val_main_output_loss: 0.5245 - val_aux_output_loss: 0.5903 - val_main_output_acc: 0.7767 - val_aux_output_acc: 0.6796\nEpoch 280/500\n924/924 [==============================] - 0s 217us/step - loss: 0.5911 - main_output_loss: 0.5326 - aux_output_loss: 0.5849 - main_output_acc: 0.7478 - aux_output_acc: 0.7100 - val_loss: 0.5802 - val_main_output_loss: 0.5212 - val_aux_output_loss: 0.5896 - val_main_output_acc: 0.7767 - val_aux_output_acc: 0.6893\nEpoch 281/500\n924/924 [==============================] - 0s 210us/step - loss: 0.5905 - main_output_loss: 0.5320 - aux_output_loss: 0.5848 - main_output_acc: 0.7576 - aux_output_acc: 0.7121 - val_loss: 0.5776 - val_main_output_loss: 0.5187 - val_aux_output_loss: 0.5891 - val_main_output_acc: 0.7767 - val_aux_output_acc: 0.6893\nEpoch 282/500\n924/924 [==============================] - 0s 214us/step - loss: 0.5908 - main_output_loss: 0.5323 - aux_output_loss: 0.5847 - main_output_acc: 0.7576 - aux_output_acc: 0.7132 - val_loss: 0.5787 - val_main_output_loss: 0.5198 - val_aux_output_loss: 0.5891 - val_main_output_acc: 0.7767 - val_aux_output_acc: 0.6893\nEpoch 283/500\n924/924 [==============================] - 0s 218us/step - loss: 0.5887 - main_output_loss: 0.5303 - aux_output_loss: 0.5840 - main_output_acc: 0.7543 - aux_output_acc: 0.7110 - val_loss: 0.5752 - val_main_output_loss: 0.5163 - val_aux_output_loss: 0.5883 - val_main_output_acc: 0.7670 - val_aux_output_acc: 0.6990\nEpoch 284/500\n924/924 [==============================] - 0s 209us/step - loss: 0.5889 - main_output_loss: 0.5305 - aux_output_loss: 0.5840 - main_output_acc: 0.7522 - aux_output_acc: 0.7110 - val_loss: 0.5760 - val_main_output_loss: 0.5172 - val_aux_output_loss: 0.5880 - val_main_output_acc: 0.7767 - val_aux_output_acc: 0.6990\nEpoch 285/500\n924/924 [==============================] - 0s 207us/step - loss: 0.5876 - main_output_loss: 0.5292 - aux_output_loss: 0.5837 - main_output_acc: 0.7587 - aux_output_acc: 0.7110 - val_loss: 0.5786 - val_main_output_loss: 0.5198 - val_aux_output_loss: 0.5880 - val_main_output_acc: 0.7767 - val_aux_output_acc: 0.6893\nEpoch 286/500\n924/924 [==============================] - 0s 207us/step - loss: 0.5879 - main_output_loss: 0.5295 - aux_output_loss: 0.5835 - main_output_acc: 0.7608 - aux_output_acc: 0.7132 - val_loss: 0.5756 - val_main_output_loss: 0.5169 - val_aux_output_loss: 0.5875 - val_main_output_acc: 0.7767 - val_aux_output_acc: 0.6990\nEpoch 287/500\n924/924 [==============================] - 0s 213us/step - loss: 0.5872 - main_output_loss: 0.5289 - aux_output_loss: 0.5834 - main_output_acc: 0.7597 - aux_output_acc: 0.7110 - val_loss: 0.5765 - val_main_output_loss: 0.5178 - val_aux_output_loss: 0.5873 - val_main_output_acc: 0.7864 - val_aux_output_acc: 0.6893\nEpoch 288/500\n924/924 [==============================] - 0s 210us/step - loss: 0.5867 - main_output_loss: 0.5284 - aux_output_loss: 0.5830 - main_output_acc: 0.7576 - aux_output_acc: 0.7110 - val_loss: 0.5749 - val_main_output_loss: 0.5162 - val_aux_output_loss: 0.5868 - val_main_output_acc: 0.7670 - val_aux_output_acc: 0.6893\nEpoch 289/500\n924/924 [==============================] - 0s 214us/step - loss: 0.5854 - main_output_loss: 0.5271 - aux_output_loss: 0.5826 - main_output_acc: 0.7608 - aux_output_acc: 0.7132 - val_loss: 0.5746 - val_main_output_loss: 0.5160 - val_aux_output_loss: 0.5865 - val_main_output_acc: 0.7767 - val_aux_output_acc: 0.6990\nEpoch 290/500\n924/924 [==============================] - 0s 217us/step - loss: 0.5853 - main_output_loss: 0.5270 - aux_output_loss: 0.5824 - main_output_acc: 0.7652 - aux_output_acc: 0.7132 - val_loss: 0.5738 - val_main_output_loss: 0.5152 - val_aux_output_loss: 0.5860 - val_main_output_acc: 0.7767 - val_aux_output_acc: 0.6990\nEpoch 291/500\n924/924 [==============================] - 0s 208us/step - loss: 0.5850 - main_output_loss: 0.5267 - aux_output_loss: 0.5822 - main_output_acc: 0.7554 - aux_output_acc: 0.7110 - val_loss: 0.5777 - val_main_output_loss: 0.5191 - val_aux_output_loss: 0.5860 - val_main_output_acc: 0.7767 - val_aux_output_acc: 0.6893\nEpoch 292/500\n924/924 [==============================] - 0s 206us/step - loss: 0.5837 - main_output_loss: 0.5255 - aux_output_loss: 0.5817 - main_output_acc: 0.7608 - aux_output_acc: 0.7110 - val_loss: 0.5722 - val_main_output_loss: 0.5137 - val_aux_output_loss: 0.5854 - val_main_output_acc: 0.7767 - val_aux_output_acc: 0.6893\nEpoch 293/500\n924/924 [==============================] - 0s 210us/step - loss: 0.5840 - main_output_loss: 0.5259 - aux_output_loss: 0.5816 - main_output_acc: 0.7565 - aux_output_acc: 0.7121 - val_loss: 0.5715 - val_main_output_loss: 0.5130 - val_aux_output_loss: 0.5851 - val_main_output_acc: 0.7767 - val_aux_output_acc: 0.6990\nEpoch 294/500\n924/924 [==============================] - 0s 205us/step - loss: 0.5824 - main_output_loss: 0.5243 - aux_output_loss: 0.5813 - main_output_acc: 0.7565 - aux_output_acc: 0.7110 - val_loss: 0.5760 - val_main_output_loss: 0.5174 - val_aux_output_loss: 0.5852 - val_main_output_acc: 0.7767 - val_aux_output_acc: 0.6893\nEpoch 295/500\n924/924 [==============================] - 0s 207us/step - loss: 0.5823 - main_output_loss: 0.5242 - aux_output_loss: 0.5810 - main_output_acc: 0.7522 - aux_output_acc: 0.7132 - val_loss: 0.5733 - val_main_output_loss: 0.5148 - val_aux_output_loss: 0.5849 - val_main_output_acc: 0.7767 - val_aux_output_acc: 0.6893\nEpoch 296/500\n924/924 [==============================] - 0s 208us/step - loss: 0.5830 - main_output_loss: 0.5249 - aux_output_loss: 0.5810 - main_output_acc: 0.7576 - aux_output_acc: 0.7121 - val_loss: 0.5750 - val_main_output_loss: 0.5165 - val_aux_output_loss: 0.5849 - val_main_output_acc: 0.7767 - val_aux_output_acc: 0.6893\nEpoch 297/500\n924/924 [==============================] - 0s 207us/step - loss: 0.5810 - main_output_loss: 0.5229 - aux_output_loss: 0.5806 - main_output_acc: 0.7619 - aux_output_acc: 0.7132 - val_loss: 0.5773 - val_main_output_loss: 0.5188 - val_aux_output_loss: 0.5847 - val_main_output_acc: 0.7767 - val_aux_output_acc: 0.6990\nEpoch 298/500\n924/924 [==============================] - 0s 205us/step - loss: 0.5808 - main_output_loss: 0.5228 - aux_output_loss: 0.5802 - main_output_acc: 0.7587 - aux_output_acc: 0.7110 - val_loss: 0.5777 - val_main_output_loss: 0.5192 - val_aux_output_loss: 0.5845 - val_main_output_acc: 0.7767 - val_aux_output_acc: 0.7087\nEpoch 299/500\n924/924 [==============================] - 0s 209us/step - loss: 0.5808 - main_output_loss: 0.5228 - aux_output_loss: 0.5800 - main_output_acc: 0.7543 - aux_output_acc: 0.7110 - val_loss: 0.5690 - val_main_output_loss: 0.5106 - val_aux_output_loss: 0.5838 - val_main_output_acc: 0.7864 - val_aux_output_acc: 0.6893\nEpoch 300/500\n924/924 [==============================] - 0s 208us/step - loss: 0.5799 - main_output_loss: 0.5219 - aux_output_loss: 0.5799 - main_output_acc: 0.7576 - aux_output_acc: 0.7121 - val_loss: 0.5750 - val_main_output_loss: 0.5166 - val_aux_output_loss: 0.5838 - val_main_output_acc: 0.7767 - val_aux_output_acc: 0.7087\nEpoch 301/500\n924/924 [==============================] - 0s 207us/step - loss: 0.5800 - main_output_loss: 0.5220 - aux_output_loss: 0.5796 - main_output_acc: 0.7554 - aux_output_acc: 0.7132 - val_loss: 0.5691 - val_main_output_loss: 0.5108 - val_aux_output_loss: 0.5831 - val_main_output_acc: 0.7864 - val_aux_output_acc: 0.6990\nEpoch 302/500\n924/924 [==============================] - 0s 205us/step - loss: 0.5778 - main_output_loss: 0.5199 - aux_output_loss: 0.5791 - main_output_acc: 0.7576 - aux_output_acc: 0.7143 - val_loss: 0.5668 - val_main_output_loss: 0.5085 - val_aux_output_loss: 0.5827 - val_main_output_acc: 0.7767 - val_aux_output_acc: 0.6990\nEpoch 303/500\n924/924 [==============================] - 0s 205us/step - loss: 0.5770 - main_output_loss: 0.5191 - aux_output_loss: 0.5788 - main_output_acc: 0.7576 - aux_output_acc: 0.7110 - val_loss: 0.5773 - val_main_output_loss: 0.5190 - val_aux_output_loss: 0.5828 - val_main_output_acc: 0.7767 - val_aux_output_acc: 0.7087\nEpoch 304/500\n924/924 [==============================] - 0s 215us/step - loss: 0.5788 - main_output_loss: 0.5210 - aux_output_loss: 0.5787 - main_output_acc: 0.7608 - aux_output_acc: 0.7089 - val_loss: 0.5674 - val_main_output_loss: 0.5092 - val_aux_output_loss: 0.5822 - val_main_output_acc: 0.7864 - val_aux_output_acc: 0.6990\nEpoch 305/500\n924/924 [==============================] - 0s 207us/step - loss: 0.5770 - main_output_loss: 0.5191 - aux_output_loss: 0.5784 - main_output_acc: 0.7619 - aux_output_acc: 0.7121 - val_loss: 0.5676 - val_main_output_loss: 0.5094 - val_aux_output_loss: 0.5819 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.6990\nEpoch 306/500\n924/924 [==============================] - 0s 208us/step - loss: 0.5769 - main_output_loss: 0.5191 - aux_output_loss: 0.5783 - main_output_acc: 0.7608 - aux_output_acc: 0.7154 - val_loss: 0.5676 - val_main_output_loss: 0.5094 - val_aux_output_loss: 0.5816 - val_main_output_acc: 0.7864 - val_aux_output_acc: 0.7087\nEpoch 307/500\n924/924 [==============================] - 0s 211us/step - loss: 0.5755 - main_output_loss: 0.5177 - aux_output_loss: 0.5778 - main_output_acc: 0.7587 - aux_output_acc: 0.7089 - val_loss: 0.5675 - val_main_output_loss: 0.5093 - val_aux_output_loss: 0.5813 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7087\nEpoch 308/500\n924/924 [==============================] - 0s 203us/step - loss: 0.5756 - main_output_loss: 0.5178 - aux_output_loss: 0.5777 - main_output_acc: 0.7576 - aux_output_acc: 0.7121 - val_loss: 0.5692 - val_main_output_loss: 0.5111 - val_aux_output_loss: 0.5811 - val_main_output_acc: 0.7864 - val_aux_output_acc: 0.7087\nEpoch 309/500\n924/924 [==============================] - 0s 210us/step - loss: 0.5757 - main_output_loss: 0.5179 - aux_output_loss: 0.5774 - main_output_acc: 0.7565 - aux_output_acc: 0.7110 - val_loss: 0.5673 - val_main_output_loss: 0.5092 - val_aux_output_loss: 0.5808 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7087\nEpoch 310/500\n924/924 [==============================] - 0s 211us/step - loss: 0.5738 - main_output_loss: 0.5161 - aux_output_loss: 0.5771 - main_output_acc: 0.7608 - aux_output_acc: 0.7100 - val_loss: 0.5640 - val_main_output_loss: 0.5060 - val_aux_output_loss: 0.5801 - val_main_output_acc: 0.7767 - val_aux_output_acc: 0.7087\nEpoch 311/500\n924/924 [==============================] - 0s 232us/step - loss: 0.5744 - main_output_loss: 0.5167 - aux_output_loss: 0.5770 - main_output_acc: 0.7565 - aux_output_acc: 0.7110 - val_loss: 0.5651 - val_main_output_loss: 0.5071 - val_aux_output_loss: 0.5799 - val_main_output_acc: 0.7864 - val_aux_output_acc: 0.7087\nEpoch 312/500\n924/924 [==============================] - 0s 217us/step - loss: 0.5730 - main_output_loss: 0.5154 - aux_output_loss: 0.5767 - main_output_acc: 0.7576 - aux_output_acc: 0.7121 - val_loss: 0.5722 - val_main_output_loss: 0.5142 - val_aux_output_loss: 0.5802 - val_main_output_acc: 0.7767 - val_aux_output_acc: 0.6990\nEpoch 313/500\n924/924 [==============================] - 0s 212us/step - loss: 0.5726 - main_output_loss: 0.5150 - aux_output_loss: 0.5761 - main_output_acc: 0.7608 - aux_output_acc: 0.7110 - val_loss: 0.5651 - val_main_output_loss: 0.5072 - val_aux_output_loss: 0.5796 - val_main_output_acc: 0.7864 - val_aux_output_acc: 0.7087\nEpoch 314/500\n924/924 [==============================] - 0s 214us/step - loss: 0.5724 - main_output_loss: 0.5147 - aux_output_loss: 0.5762 - main_output_acc: 0.7630 - aux_output_acc: 0.7121 - val_loss: 0.5670 - val_main_output_loss: 0.5091 - val_aux_output_loss: 0.5795 - val_main_output_acc: 0.7864 - val_aux_output_acc: 0.7087\nEpoch 315/500\n924/924 [==============================] - 0s 209us/step - loss: 0.5711 - main_output_loss: 0.5135 - aux_output_loss: 0.5757 - main_output_acc: 0.7543 - aux_output_acc: 0.7110 - val_loss: 0.5619 - val_main_output_loss: 0.5041 - val_aux_output_loss: 0.5787 - val_main_output_acc: 0.7864 - val_aux_output_acc: 0.7087\nEpoch 316/500\n924/924 [==============================] - 0s 209us/step - loss: 0.5713 - main_output_loss: 0.5137 - aux_output_loss: 0.5758 - main_output_acc: 0.7630 - aux_output_acc: 0.7100 - val_loss: 0.5612 - val_main_output_loss: 0.5034 - val_aux_output_loss: 0.5783 - val_main_output_acc: 0.7767 - val_aux_output_acc: 0.7087\nEpoch 317/500\n924/924 [==============================] - 0s 206us/step - loss: 0.5721 - main_output_loss: 0.5145 - aux_output_loss: 0.5758 - main_output_acc: 0.7641 - aux_output_acc: 0.7143 - val_loss: 0.5611 - val_main_output_loss: 0.5033 - val_aux_output_loss: 0.5778 - val_main_output_acc: 0.7767 - val_aux_output_acc: 0.7087\nEpoch 318/500\n924/924 [==============================] - 0s 205us/step - loss: 0.5708 - main_output_loss: 0.5133 - aux_output_loss: 0.5754 - main_output_acc: 0.7597 - aux_output_acc: 0.7154 - val_loss: 0.5640 - val_main_output_loss: 0.5062 - val_aux_output_loss: 0.5779 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7087\nEpoch 319/500\n924/924 [==============================] - 0s 206us/step - loss: 0.5695 - main_output_loss: 0.5120 - aux_output_loss: 0.5750 - main_output_acc: 0.7597 - aux_output_acc: 0.7110 - val_loss: 0.5635 - val_main_output_loss: 0.5057 - val_aux_output_loss: 0.5775 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7087\nEpoch 320/500\n924/924 [==============================] - 0s 217us/step - loss: 0.5694 - main_output_loss: 0.5119 - aux_output_loss: 0.5748 - main_output_acc: 0.7576 - aux_output_acc: 0.7143 - val_loss: 0.5633 - val_main_output_loss: 0.5056 - val_aux_output_loss: 0.5773 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7087\nEpoch 321/500\n924/924 [==============================] - 0s 213us/step - loss: 0.5686 - main_output_loss: 0.5111 - aux_output_loss: 0.5745 - main_output_acc: 0.7630 - aux_output_acc: 0.7121 - val_loss: 0.5622 - val_main_output_loss: 0.5045 - val_aux_output_loss: 0.5770 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7087\nEpoch 322/500\n924/924 [==============================] - 0s 205us/step - loss: 0.5686 - main_output_loss: 0.5111 - aux_output_loss: 0.5742 - main_output_acc: 0.7576 - aux_output_acc: 0.7121 - val_loss: 0.5620 - val_main_output_loss: 0.5044 - val_aux_output_loss: 0.5767 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7087\nEpoch 323/500\n924/924 [==============================] - 0s 209us/step - loss: 0.5677 - main_output_loss: 0.5103 - aux_output_loss: 0.5738 - main_output_acc: 0.7554 - aux_output_acc: 0.7143 - val_loss: 0.5636 - val_main_output_loss: 0.5060 - val_aux_output_loss: 0.5764 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7087\nEpoch 324/500\n924/924 [==============================] - 0s 205us/step - loss: 0.5683 - main_output_loss: 0.5110 - aux_output_loss: 0.5738 - main_output_acc: 0.7587 - aux_output_acc: 0.7089 - val_loss: 0.5601 - val_main_output_loss: 0.5025 - val_aux_output_loss: 0.5759 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7184\nEpoch 325/500\n924/924 [==============================] - 0s 207us/step - loss: 0.5670 - main_output_loss: 0.5096 - aux_output_loss: 0.5735 - main_output_acc: 0.7641 - aux_output_acc: 0.7132 - val_loss: 0.5631 - val_main_output_loss: 0.5055 - val_aux_output_loss: 0.5758 - val_main_output_acc: 0.7670 - val_aux_output_acc: 0.6990\nEpoch 326/500\n924/924 [==============================] - 0s 210us/step - loss: 0.5678 - main_output_loss: 0.5105 - aux_output_loss: 0.5734 - main_output_acc: 0.7587 - aux_output_acc: 0.7132 - val_loss: 0.5635 - val_main_output_loss: 0.5060 - val_aux_output_loss: 0.5756 - val_main_output_acc: 0.7670 - val_aux_output_acc: 0.6990\nEpoch 327/500\n924/924 [==============================] - 0s 205us/step - loss: 0.5656 - main_output_loss: 0.5083 - aux_output_loss: 0.5728 - main_output_acc: 0.7608 - aux_output_acc: 0.7154 - val_loss: 0.5576 - val_main_output_loss: 0.5001 - val_aux_output_loss: 0.5749 - val_main_output_acc: 0.7864 - val_aux_output_acc: 0.7184\nEpoch 328/500\n924/924 [==============================] - 0s 216us/step - loss: 0.5665 - main_output_loss: 0.5092 - aux_output_loss: 0.5728 - main_output_acc: 0.7619 - aux_output_acc: 0.7132 - val_loss: 0.5575 - val_main_output_loss: 0.5001 - val_aux_output_loss: 0.5742 - val_main_output_acc: 0.7767 - val_aux_output_acc: 0.7184\nEpoch 329/500\n924/924 [==============================] - 0s 211us/step - loss: 0.5662 - main_output_loss: 0.5089 - aux_output_loss: 0.5726 - main_output_acc: 0.7716 - aux_output_acc: 0.7121 - val_loss: 0.5575 - val_main_output_loss: 0.5001 - val_aux_output_loss: 0.5739 - val_main_output_acc: 0.7767 - val_aux_output_acc: 0.7282\nEpoch 330/500\n924/924 [==============================] - 0s 211us/step - loss: 0.5661 - main_output_loss: 0.5088 - aux_output_loss: 0.5725 - main_output_acc: 0.7597 - aux_output_acc: 0.7121 - val_loss: 0.5578 - val_main_output_loss: 0.5004 - val_aux_output_loss: 0.5736 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7184\nEpoch 331/500\n924/924 [==============================] - 0s 217us/step - loss: 0.5640 - main_output_loss: 0.5068 - aux_output_loss: 0.5720 - main_output_acc: 0.7576 - aux_output_acc: 0.7165 - val_loss: 0.5622 - val_main_output_loss: 0.5048 - val_aux_output_loss: 0.5736 - val_main_output_acc: 0.7670 - val_aux_output_acc: 0.7184\nEpoch 332/500\n924/924 [==============================] - 0s 209us/step - loss: 0.5630 - main_output_loss: 0.5058 - aux_output_loss: 0.5717 - main_output_acc: 0.7673 - aux_output_acc: 0.7143 - val_loss: 0.5552 - val_main_output_loss: 0.4979 - val_aux_output_loss: 0.5730 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7282\nEpoch 333/500\n924/924 [==============================] - 0s 214us/step - loss: 0.5636 - main_output_loss: 0.5064 - aux_output_loss: 0.5716 - main_output_acc: 0.7641 - aux_output_acc: 0.7143 - val_loss: 0.5574 - val_main_output_loss: 0.5001 - val_aux_output_loss: 0.5729 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7184\nEpoch 334/500\n924/924 [==============================] - 0s 208us/step - loss: 0.5626 - main_output_loss: 0.5054 - aux_output_loss: 0.5713 - main_output_acc: 0.7652 - aux_output_acc: 0.7165 - val_loss: 0.5580 - val_main_output_loss: 0.5007 - val_aux_output_loss: 0.5726 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7184\nEpoch 335/500\n924/924 [==============================] - 0s 214us/step - loss: 0.5633 - main_output_loss: 0.5062 - aux_output_loss: 0.5715 - main_output_acc: 0.7630 - aux_output_acc: 0.7132 - val_loss: 0.5580 - val_main_output_loss: 0.5007 - val_aux_output_loss: 0.5723 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7184\nEpoch 336/500\n924/924 [==============================] - 0s 207us/step - loss: 0.5622 - main_output_loss: 0.5051 - aux_output_loss: 0.5709 - main_output_acc: 0.7608 - aux_output_acc: 0.7165 - val_loss: 0.5578 - val_main_output_loss: 0.5006 - val_aux_output_loss: 0.5721 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7184\nEpoch 337/500\n924/924 [==============================] - 0s 208us/step - loss: 0.5609 - main_output_loss: 0.5039 - aux_output_loss: 0.5706 - main_output_acc: 0.7630 - aux_output_acc: 0.7143 - val_loss: 0.5606 - val_main_output_loss: 0.5034 - val_aux_output_loss: 0.5721 - val_main_output_acc: 0.7670 - val_aux_output_acc: 0.7087\nEpoch 338/500\n924/924 [==============================] - 0s 205us/step - loss: 0.5605 - main_output_loss: 0.5035 - aux_output_loss: 0.5703 - main_output_acc: 0.7630 - aux_output_acc: 0.7143 - val_loss: 0.5598 - val_main_output_loss: 0.5026 - val_aux_output_loss: 0.5720 - val_main_output_acc: 0.7670 - val_aux_output_acc: 0.7087\nEpoch 339/500\n924/924 [==============================] - 0s 208us/step - loss: 0.5617 - main_output_loss: 0.5047 - aux_output_loss: 0.5703 - main_output_acc: 0.7608 - aux_output_acc: 0.7143 - val_loss: 0.5563 - val_main_output_loss: 0.4991 - val_aux_output_loss: 0.5716 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7184\nEpoch 340/500\n924/924 [==============================] - 0s 213us/step - loss: 0.5598 - main_output_loss: 0.5028 - aux_output_loss: 0.5698 - main_output_acc: 0.7630 - aux_output_acc: 0.7121 - val_loss: 0.5534 - val_main_output_loss: 0.4963 - val_aux_output_loss: 0.5709 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7282\nEpoch 341/500\n924/924 [==============================] - 0s 214us/step - loss: 0.5601 - main_output_loss: 0.5031 - aux_output_loss: 0.5698 - main_output_acc: 0.7641 - aux_output_acc: 0.7197 - val_loss: 0.5525 - val_main_output_loss: 0.4954 - val_aux_output_loss: 0.5706 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7282\nEpoch 342/500\n924/924 [==============================] - 0s 213us/step - loss: 0.5594 - main_output_loss: 0.5025 - aux_output_loss: 0.5696 - main_output_acc: 0.7576 - aux_output_acc: 0.7165 - val_loss: 0.5517 - val_main_output_loss: 0.4946 - val_aux_output_loss: 0.5703 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7282\nEpoch 343/500\n924/924 [==============================] - 0s 216us/step - loss: 0.5590 - main_output_loss: 0.5020 - aux_output_loss: 0.5695 - main_output_acc: 0.7662 - aux_output_acc: 0.7154 - val_loss: 0.5624 - val_main_output_loss: 0.5053 - val_aux_output_loss: 0.5708 - val_main_output_acc: 0.7767 - val_aux_output_acc: 0.7087\nEpoch 344/500\n924/924 [==============================] - 0s 210us/step - loss: 0.5584 - main_output_loss: 0.5015 - aux_output_loss: 0.5687 - main_output_acc: 0.7630 - aux_output_acc: 0.7143 - val_loss: 0.5546 - val_main_output_loss: 0.4976 - val_aux_output_loss: 0.5701 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7184\nEpoch 345/500\n924/924 [==============================] - 0s 208us/step - loss: 0.5580 - main_output_loss: 0.5011 - aux_output_loss: 0.5689 - main_output_acc: 0.7641 - aux_output_acc: 0.7154 - val_loss: 0.5565 - val_main_output_loss: 0.4995 - val_aux_output_loss: 0.5700 - val_main_output_acc: 0.7767 - val_aux_output_acc: 0.7087\nEpoch 346/500\n924/924 [==============================] - 0s 206us/step - loss: 0.5571 - main_output_loss: 0.5002 - aux_output_loss: 0.5687 - main_output_acc: 0.7608 - aux_output_acc: 0.7229 - val_loss: 0.5580 - val_main_output_loss: 0.5010 - val_aux_output_loss: 0.5698 - val_main_output_acc: 0.7670 - val_aux_output_acc: 0.7087\nEpoch 347/500\n924/924 [==============================] - 0s 204us/step - loss: 0.5573 - main_output_loss: 0.5004 - aux_output_loss: 0.5684 - main_output_acc: 0.7597 - aux_output_acc: 0.7165 - val_loss: 0.5615 - val_main_output_loss: 0.5046 - val_aux_output_loss: 0.5699 - val_main_output_acc: 0.7767 - val_aux_output_acc: 0.7087\nEpoch 348/500\n924/924 [==============================] - 0s 207us/step - loss: 0.5576 - main_output_loss: 0.5008 - aux_output_loss: 0.5681 - main_output_acc: 0.7597 - aux_output_acc: 0.7154 - val_loss: 0.5537 - val_main_output_loss: 0.4968 - val_aux_output_loss: 0.5690 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7282\nEpoch 349/500\n924/924 [==============================] - 0s 209us/step - loss: 0.5562 - main_output_loss: 0.4994 - aux_output_loss: 0.5679 - main_output_acc: 0.7630 - aux_output_acc: 0.7143 - val_loss: 0.5540 - val_main_output_loss: 0.4971 - val_aux_output_loss: 0.5687 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7184\nEpoch 350/500\n924/924 [==============================] - 0s 208us/step - loss: 0.5563 - main_output_loss: 0.4995 - aux_output_loss: 0.5677 - main_output_acc: 0.7641 - aux_output_acc: 0.7175 - val_loss: 0.5515 - val_main_output_loss: 0.4947 - val_aux_output_loss: 0.5683 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7282\nEpoch 351/500\n924/924 [==============================] - 0s 213us/step - loss: 0.5555 - main_output_loss: 0.4987 - aux_output_loss: 0.5675 - main_output_acc: 0.7630 - aux_output_acc: 0.7121 - val_loss: 0.5580 - val_main_output_loss: 0.5011 - val_aux_output_loss: 0.5686 - val_main_output_acc: 0.7670 - val_aux_output_acc: 0.7087\nEpoch 352/500\n924/924 [==============================] - 0s 209us/step - loss: 0.5552 - main_output_loss: 0.4985 - aux_output_loss: 0.5670 - main_output_acc: 0.7652 - aux_output_acc: 0.7165 - val_loss: 0.5545 - val_main_output_loss: 0.4978 - val_aux_output_loss: 0.5680 - val_main_output_acc: 0.7864 - val_aux_output_acc: 0.7282\nEpoch 353/500\n924/924 [==============================] - 0s 207us/step - loss: 0.5548 - main_output_loss: 0.4981 - aux_output_loss: 0.5671 - main_output_acc: 0.7662 - aux_output_acc: 0.7143 - val_loss: 0.5529 - val_main_output_loss: 0.4962 - val_aux_output_loss: 0.5675 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7282\nEpoch 354/500\n924/924 [==============================] - 0s 207us/step - loss: 0.5540 - main_output_loss: 0.4973 - aux_output_loss: 0.5669 - main_output_acc: 0.7641 - aux_output_acc: 0.7110 - val_loss: 0.5530 - val_main_output_loss: 0.4962 - val_aux_output_loss: 0.5673 - val_main_output_acc: 0.7767 - val_aux_output_acc: 0.7282\nEpoch 355/500\n924/924 [==============================] - 0s 205us/step - loss: 0.5531 - main_output_loss: 0.4965 - aux_output_loss: 0.5667 - main_output_acc: 0.7619 - aux_output_acc: 0.7175 - val_loss: 0.5504 - val_main_output_loss: 0.4937 - val_aux_output_loss: 0.5668 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7282\nEpoch 356/500\n924/924 [==============================] - 0s 208us/step - loss: 0.5536 - main_output_loss: 0.4969 - aux_output_loss: 0.5665 - main_output_acc: 0.7608 - aux_output_acc: 0.7143 - val_loss: 0.5481 - val_main_output_loss: 0.4914 - val_aux_output_loss: 0.5663 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7184\nEpoch 357/500\n924/924 [==============================] - 0s 205us/step - loss: 0.5527 - main_output_loss: 0.4960 - aux_output_loss: 0.5662 - main_output_acc: 0.7652 - aux_output_acc: 0.7154 - val_loss: 0.5515 - val_main_output_loss: 0.4949 - val_aux_output_loss: 0.5662 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7282\nEpoch 358/500\n924/924 [==============================] - 0s 207us/step - loss: 0.5515 - main_output_loss: 0.4950 - aux_output_loss: 0.5659 - main_output_acc: 0.7673 - aux_output_acc: 0.7121 - val_loss: 0.5548 - val_main_output_loss: 0.4982 - val_aux_output_loss: 0.5659 - val_main_output_acc: 0.7670 - val_aux_output_acc: 0.7282\nEpoch 359/500\n924/924 [==============================] - 0s 208us/step - loss: 0.5530 - main_output_loss: 0.4964 - aux_output_loss: 0.5658 - main_output_acc: 0.7619 - aux_output_acc: 0.7121 - val_loss: 0.5537 - val_main_output_loss: 0.4971 - val_aux_output_loss: 0.5656 - val_main_output_acc: 0.7670 - val_aux_output_acc: 0.7282\nEpoch 360/500\n924/924 [==============================] - 0s 207us/step - loss: 0.5517 - main_output_loss: 0.4951 - aux_output_loss: 0.5654 - main_output_acc: 0.7652 - aux_output_acc: 0.7165 - val_loss: 0.5491 - val_main_output_loss: 0.4926 - val_aux_output_loss: 0.5652 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7282\nEpoch 361/500\n924/924 [==============================] - 0s 207us/step - loss: 0.5520 - main_output_loss: 0.4955 - aux_output_loss: 0.5654 - main_output_acc: 0.7597 - aux_output_acc: 0.7154 - val_loss: 0.5460 - val_main_output_loss: 0.4895 - val_aux_output_loss: 0.5648 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7184\nEpoch 362/500\n924/924 [==============================] - 0s 207us/step - loss: 0.5506 - main_output_loss: 0.4941 - aux_output_loss: 0.5652 - main_output_acc: 0.7630 - aux_output_acc: 0.7132 - val_loss: 0.5451 - val_main_output_loss: 0.4886 - val_aux_output_loss: 0.5643 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7087\nEpoch 363/500\n924/924 [==============================] - 0s 207us/step - loss: 0.5517 - main_output_loss: 0.4952 - aux_output_loss: 0.5652 - main_output_acc: 0.7641 - aux_output_acc: 0.7154 - val_loss: 0.5486 - val_main_output_loss: 0.4922 - val_aux_output_loss: 0.5643 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7282\nEpoch 364/500\n924/924 [==============================] - 0s 210us/step - loss: 0.5499 - main_output_loss: 0.4934 - aux_output_loss: 0.5646 - main_output_acc: 0.7652 - aux_output_acc: 0.7110 - val_loss: 0.5490 - val_main_output_loss: 0.4926 - val_aux_output_loss: 0.5641 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7282\nEpoch 365/500\n924/924 [==============================] - 0s 205us/step - loss: 0.5494 - main_output_loss: 0.4929 - aux_output_loss: 0.5644 - main_output_acc: 0.7706 - aux_output_acc: 0.7110 - val_loss: 0.5462 - val_main_output_loss: 0.4898 - val_aux_output_loss: 0.5638 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7184\nEpoch 366/500\n924/924 [==============================] - 0s 208us/step - loss: 0.5490 - main_output_loss: 0.4925 - aux_output_loss: 0.5642 - main_output_acc: 0.7641 - aux_output_acc: 0.7132 - val_loss: 0.5450 - val_main_output_loss: 0.4886 - val_aux_output_loss: 0.5635 - val_main_output_acc: 0.7864 - val_aux_output_acc: 0.7184\nEpoch 367/500\n924/924 [==============================] - 0s 210us/step - loss: 0.5494 - main_output_loss: 0.4930 - aux_output_loss: 0.5642 - main_output_acc: 0.7565 - aux_output_acc: 0.7186 - val_loss: 0.5454 - val_main_output_loss: 0.4891 - val_aux_output_loss: 0.5632 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7184\nEpoch 368/500\n924/924 [==============================] - 0s 207us/step - loss: 0.5495 - main_output_loss: 0.4931 - aux_output_loss: 0.5641 - main_output_acc: 0.7706 - aux_output_acc: 0.7154 - val_loss: 0.5445 - val_main_output_loss: 0.4882 - val_aux_output_loss: 0.5630 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7087\nEpoch 369/500\n924/924 [==============================] - 0s 209us/step - loss: 0.5486 - main_output_loss: 0.4922 - aux_output_loss: 0.5639 - main_output_acc: 0.7641 - aux_output_acc: 0.7165 - val_loss: 0.5447 - val_main_output_loss: 0.4885 - val_aux_output_loss: 0.5629 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7087\nEpoch 370/500\n924/924 [==============================] - 0s 205us/step - loss: 0.5477 - main_output_loss: 0.4914 - aux_output_loss: 0.5635 - main_output_acc: 0.7738 - aux_output_acc: 0.7143 - val_loss: 0.5453 - val_main_output_loss: 0.4891 - val_aux_output_loss: 0.5626 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7184\nEpoch 371/500\n924/924 [==============================] - 0s 208us/step - loss: 0.5480 - main_output_loss: 0.4916 - aux_output_loss: 0.5634 - main_output_acc: 0.7684 - aux_output_acc: 0.7165 - val_loss: 0.5527 - val_main_output_loss: 0.4964 - val_aux_output_loss: 0.5629 - val_main_output_acc: 0.7670 - val_aux_output_acc: 0.7379\nEpoch 372/500\n924/924 [==============================] - 0s 204us/step - loss: 0.5475 - main_output_loss: 0.4912 - aux_output_loss: 0.5628 - main_output_acc: 0.7662 - aux_output_acc: 0.7132 - val_loss: 0.5504 - val_main_output_loss: 0.4941 - val_aux_output_loss: 0.5626 - val_main_output_acc: 0.7767 - val_aux_output_acc: 0.7379\nEpoch 373/500\n924/924 [==============================] - 0s 207us/step - loss: 0.5470 - main_output_loss: 0.4907 - aux_output_loss: 0.5626 - main_output_acc: 0.7706 - aux_output_acc: 0.7154 - val_loss: 0.5449 - val_main_output_loss: 0.4886 - val_aux_output_loss: 0.5622 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7184\nEpoch 374/500\n924/924 [==============================] - 0s 211us/step - loss: 0.5457 - main_output_loss: 0.4895 - aux_output_loss: 0.5625 - main_output_acc: 0.7716 - aux_output_acc: 0.7154 - val_loss: 0.5551 - val_main_output_loss: 0.4989 - val_aux_output_loss: 0.5626 - val_main_output_acc: 0.7573 - val_aux_output_acc: 0.7379\nEpoch 375/500\n924/924 [==============================] - 0s 220us/step - loss: 0.5468 - main_output_loss: 0.4906 - aux_output_loss: 0.5622 - main_output_acc: 0.7673 - aux_output_acc: 0.7143 - val_loss: 0.5510 - val_main_output_loss: 0.4948 - val_aux_output_loss: 0.5622 - val_main_output_acc: 0.7767 - val_aux_output_acc: 0.7379\nEpoch 376/500\n924/924 [==============================] - 0s 206us/step - loss: 0.5457 - main_output_loss: 0.4895 - aux_output_loss: 0.5619 - main_output_acc: 0.7706 - aux_output_acc: 0.7154 - val_loss: 0.5532 - val_main_output_loss: 0.4970 - val_aux_output_loss: 0.5621 - val_main_output_acc: 0.7573 - val_aux_output_acc: 0.7379\nEpoch 377/500\n924/924 [==============================] - 0s 205us/step - loss: 0.5452 - main_output_loss: 0.4891 - aux_output_loss: 0.5616 - main_output_acc: 0.7706 - aux_output_acc: 0.7121 - val_loss: 0.5518 - val_main_output_loss: 0.4956 - val_aux_output_loss: 0.5619 - val_main_output_acc: 0.7670 - val_aux_output_acc: 0.7379\nEpoch 378/500\n924/924 [==============================] - 0s 205us/step - loss: 0.5456 - main_output_loss: 0.4894 - aux_output_loss: 0.5616 - main_output_acc: 0.7706 - aux_output_acc: 0.7143 - val_loss: 0.5547 - val_main_output_loss: 0.4985 - val_aux_output_loss: 0.5618 - val_main_output_acc: 0.7573 - val_aux_output_acc: 0.7379\nEpoch 379/500\n924/924 [==============================] - 0s 207us/step - loss: 0.5457 - main_output_loss: 0.4896 - aux_output_loss: 0.5613 - main_output_acc: 0.7706 - aux_output_acc: 0.7132 - val_loss: 0.5490 - val_main_output_loss: 0.4928 - val_aux_output_loss: 0.5612 - val_main_output_acc: 0.7767 - val_aux_output_acc: 0.7379\nEpoch 380/500\n924/924 [==============================] - 0s 206us/step - loss: 0.5432 - main_output_loss: 0.4871 - aux_output_loss: 0.5610 - main_output_acc: 0.7652 - aux_output_acc: 0.7143 - val_loss: 0.5443 - val_main_output_loss: 0.4883 - val_aux_output_loss: 0.5607 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7282\nEpoch 381/500\n924/924 [==============================] - 0s 213us/step - loss: 0.5441 - main_output_loss: 0.4880 - aux_output_loss: 0.5611 - main_output_acc: 0.7695 - aux_output_acc: 0.7154 - val_loss: 0.5505 - val_main_output_loss: 0.4944 - val_aux_output_loss: 0.5608 - val_main_output_acc: 0.7670 - val_aux_output_acc: 0.7379\nEpoch 382/500\n924/924 [==============================] - 0s 209us/step - loss: 0.5428 - main_output_loss: 0.4867 - aux_output_loss: 0.5605 - main_output_acc: 0.7695 - aux_output_acc: 0.7175 - val_loss: 0.5455 - val_main_output_loss: 0.4895 - val_aux_output_loss: 0.5604 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7282\nEpoch 383/500\n924/924 [==============================] - 0s 207us/step - loss: 0.5431 - main_output_loss: 0.4871 - aux_output_loss: 0.5605 - main_output_acc: 0.7662 - aux_output_acc: 0.7143 - val_loss: 0.5425 - val_main_output_loss: 0.4864 - val_aux_output_loss: 0.5602 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7282\nEpoch 384/500\n924/924 [==============================] - 0s 213us/step - loss: 0.5433 - main_output_loss: 0.4873 - aux_output_loss: 0.5606 - main_output_acc: 0.7706 - aux_output_acc: 0.7175 - val_loss: 0.5414 - val_main_output_loss: 0.4854 - val_aux_output_loss: 0.5600 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7379\nEpoch 385/500\n924/924 [==============================] - 0s 205us/step - loss: 0.5421 - main_output_loss: 0.4861 - aux_output_loss: 0.5602 - main_output_acc: 0.7673 - aux_output_acc: 0.7154 - val_loss: 0.5412 - val_main_output_loss: 0.4852 - val_aux_output_loss: 0.5597 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7087\nEpoch 386/500\n924/924 [==============================] - 0s 209us/step - loss: 0.5423 - main_output_loss: 0.4863 - aux_output_loss: 0.5602 - main_output_acc: 0.7695 - aux_output_acc: 0.7165 - val_loss: 0.5423 - val_main_output_loss: 0.4863 - val_aux_output_loss: 0.5596 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7282\nEpoch 387/500\n924/924 [==============================] - 0s 207us/step - loss: 0.5414 - main_output_loss: 0.4854 - aux_output_loss: 0.5597 - main_output_acc: 0.7652 - aux_output_acc: 0.7143 - val_loss: 0.5465 - val_main_output_loss: 0.4906 - val_aux_output_loss: 0.5596 - val_main_output_acc: 0.7767 - val_aux_output_acc: 0.7282\nEpoch 388/500\n924/924 [==============================] - 0s 204us/step - loss: 0.5410 - main_output_loss: 0.4850 - aux_output_loss: 0.5595 - main_output_acc: 0.7706 - aux_output_acc: 0.7165 - val_loss: 0.5446 - val_main_output_loss: 0.4886 - val_aux_output_loss: 0.5595 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7282\nEpoch 389/500\n924/924 [==============================] - 0s 206us/step - loss: 0.5404 - main_output_loss: 0.4845 - aux_output_loss: 0.5594 - main_output_acc: 0.7706 - aux_output_acc: 0.7154 - val_loss: 0.5450 - val_main_output_loss: 0.4891 - val_aux_output_loss: 0.5592 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7282\nEpoch 390/500\n924/924 [==============================] - 0s 207us/step - loss: 0.5416 - main_output_loss: 0.4856 - aux_output_loss: 0.5595 - main_output_acc: 0.7641 - aux_output_acc: 0.7165 - val_loss: 0.5442 - val_main_output_loss: 0.4883 - val_aux_output_loss: 0.5592 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7282\nEpoch 391/500\n924/924 [==============================] - 0s 207us/step - loss: 0.5404 - main_output_loss: 0.4845 - aux_output_loss: 0.5591 - main_output_acc: 0.7738 - aux_output_acc: 0.7154 - val_loss: 0.5463 - val_main_output_loss: 0.4904 - val_aux_output_loss: 0.5591 - val_main_output_acc: 0.7767 - val_aux_output_acc: 0.7282\nEpoch 392/500\n924/924 [==============================] - 0s 211us/step - loss: 0.5388 - main_output_loss: 0.4830 - aux_output_loss: 0.5585 - main_output_acc: 0.7727 - aux_output_acc: 0.7175 - val_loss: 0.5416 - val_main_output_loss: 0.4857 - val_aux_output_loss: 0.5586 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7282\nEpoch 393/500\n924/924 [==============================] - 0s 205us/step - loss: 0.5399 - main_output_loss: 0.4840 - aux_output_loss: 0.5586 - main_output_acc: 0.7727 - aux_output_acc: 0.7186 - val_loss: 0.5398 - val_main_output_loss: 0.4840 - val_aux_output_loss: 0.5583 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7379\nEpoch 394/500\n924/924 [==============================] - 0s 212us/step - loss: 0.5402 - main_output_loss: 0.4843 - aux_output_loss: 0.5586 - main_output_acc: 0.7749 - aux_output_acc: 0.7197 - val_loss: 0.5393 - val_main_output_loss: 0.4835 - val_aux_output_loss: 0.5579 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7379\nEpoch 395/500\n924/924 [==============================] - 0s 211us/step - loss: 0.5382 - main_output_loss: 0.4824 - aux_output_loss: 0.5582 - main_output_acc: 0.7760 - aux_output_acc: 0.7208 - val_loss: 0.5411 - val_main_output_loss: 0.4854 - val_aux_output_loss: 0.5576 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7184\nEpoch 396/500\n924/924 [==============================] - 0s 207us/step - loss: 0.5378 - main_output_loss: 0.4821 - aux_output_loss: 0.5578 - main_output_acc: 0.7738 - aux_output_acc: 0.7186 - val_loss: 0.5424 - val_main_output_loss: 0.4867 - val_aux_output_loss: 0.5574 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7282\nEpoch 397/500\n924/924 [==============================] - 0s 206us/step - loss: 0.5372 - main_output_loss: 0.4815 - aux_output_loss: 0.5577 - main_output_acc: 0.7749 - aux_output_acc: 0.7186 - val_loss: 0.5514 - val_main_output_loss: 0.4957 - val_aux_output_loss: 0.5577 - val_main_output_acc: 0.7573 - val_aux_output_acc: 0.7282\nEpoch 398/500\n924/924 [==============================] - 0s 206us/step - loss: 0.5381 - main_output_loss: 0.4823 - aux_output_loss: 0.5573 - main_output_acc: 0.7771 - aux_output_acc: 0.7175 - val_loss: 0.5397 - val_main_output_loss: 0.4840 - val_aux_output_loss: 0.5572 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7282\nEpoch 399/500\n924/924 [==============================] - 0s 207us/step - loss: 0.5377 - main_output_loss: 0.4820 - aux_output_loss: 0.5575 - main_output_acc: 0.7738 - aux_output_acc: 0.7197 - val_loss: 0.5447 - val_main_output_loss: 0.4889 - val_aux_output_loss: 0.5572 - val_main_output_acc: 0.7767 - val_aux_output_acc: 0.7282\nEpoch 400/500\n924/924 [==============================] - 0s 219us/step - loss: 0.5373 - main_output_loss: 0.4816 - aux_output_loss: 0.5572 - main_output_acc: 0.7781 - aux_output_acc: 0.7208 - val_loss: 0.5437 - val_main_output_loss: 0.4880 - val_aux_output_loss: 0.5569 - val_main_output_acc: 0.7767 - val_aux_output_acc: 0.7282\nEpoch 401/500\n924/924 [==============================] - 0s 210us/step - loss: 0.5361 - main_output_loss: 0.4804 - aux_output_loss: 0.5568 - main_output_acc: 0.7792 - aux_output_acc: 0.7208 - val_loss: 0.5514 - val_main_output_loss: 0.4957 - val_aux_output_loss: 0.5573 - val_main_output_acc: 0.7573 - val_aux_output_acc: 0.7379\nEpoch 402/500\n924/924 [==============================] - 0s 209us/step - loss: 0.5361 - main_output_loss: 0.4805 - aux_output_loss: 0.5562 - main_output_acc: 0.7781 - aux_output_acc: 0.7208 - val_loss: 0.5398 - val_main_output_loss: 0.4841 - val_aux_output_loss: 0.5566 - val_main_output_acc: 0.8155 - val_aux_output_acc: 0.7184\nEpoch 403/500\n924/924 [==============================] - 0s 205us/step - loss: 0.5367 - main_output_loss: 0.4811 - aux_output_loss: 0.5566 - main_output_acc: 0.7781 - aux_output_acc: 0.7219 - val_loss: 0.5398 - val_main_output_loss: 0.4842 - val_aux_output_loss: 0.5562 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7184\nEpoch 404/500\n924/924 [==============================] - 0s 213us/step - loss: 0.5349 - main_output_loss: 0.4793 - aux_output_loss: 0.5561 - main_output_acc: 0.7749 - aux_output_acc: 0.7208 - val_loss: 0.5446 - val_main_output_loss: 0.4889 - val_aux_output_loss: 0.5564 - val_main_output_acc: 0.7670 - val_aux_output_acc: 0.7282\nEpoch 405/500\n924/924 [==============================] - 0s 209us/step - loss: 0.5358 - main_output_loss: 0.4802 - aux_output_loss: 0.5561 - main_output_acc: 0.7749 - aux_output_acc: 0.7197 - val_loss: 0.5471 - val_main_output_loss: 0.4915 - val_aux_output_loss: 0.5563 - val_main_output_acc: 0.7573 - val_aux_output_acc: 0.7282\nEpoch 406/500\n924/924 [==============================] - 0s 211us/step - loss: 0.5352 - main_output_loss: 0.4797 - aux_output_loss: 0.5558 - main_output_acc: 0.7749 - aux_output_acc: 0.7273 - val_loss: 0.5402 - val_main_output_loss: 0.4846 - val_aux_output_loss: 0.5558 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7282\nEpoch 407/500\n924/924 [==============================] - 0s 210us/step - loss: 0.5345 - main_output_loss: 0.4790 - aux_output_loss: 0.5554 - main_output_acc: 0.7760 - aux_output_acc: 0.7229 - val_loss: 0.5417 - val_main_output_loss: 0.4862 - val_aux_output_loss: 0.5558 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7282\nEpoch 408/500\n924/924 [==============================] - 0s 205us/step - loss: 0.5337 - main_output_loss: 0.4781 - aux_output_loss: 0.5554 - main_output_acc: 0.7781 - aux_output_acc: 0.7208 - val_loss: 0.5378 - val_main_output_loss: 0.4823 - val_aux_output_loss: 0.5554 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7379\nEpoch 409/500\n924/924 [==============================] - 0s 211us/step - loss: 0.5344 - main_output_loss: 0.4789 - aux_output_loss: 0.5554 - main_output_acc: 0.7749 - aux_output_acc: 0.7251 - val_loss: 0.5424 - val_main_output_loss: 0.4869 - val_aux_output_loss: 0.5554 - val_main_output_acc: 0.7767 - val_aux_output_acc: 0.7282\nEpoch 410/500\n924/924 [==============================] - 0s 206us/step - loss: 0.5334 - main_output_loss: 0.4779 - aux_output_loss: 0.5551 - main_output_acc: 0.7716 - aux_output_acc: 0.7219 - val_loss: 0.5393 - val_main_output_loss: 0.4838 - val_aux_output_loss: 0.5550 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7282\nEpoch 411/500\n924/924 [==============================] - 0s 217us/step - loss: 0.5334 - main_output_loss: 0.4779 - aux_output_loss: 0.5551 - main_output_acc: 0.7814 - aux_output_acc: 0.7229 - val_loss: 0.5371 - val_main_output_loss: 0.4817 - val_aux_output_loss: 0.5547 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7379\nEpoch 412/500\n924/924 [==============================] - 0s 212us/step - loss: 0.5343 - main_output_loss: 0.4788 - aux_output_loss: 0.5551 - main_output_acc: 0.7792 - aux_output_acc: 0.7229 - val_loss: 0.5369 - val_main_output_loss: 0.4815 - val_aux_output_loss: 0.5545 - val_main_output_acc: 0.8155 - val_aux_output_acc: 0.7379\nEpoch 413/500\n924/924 [==============================] - 0s 204us/step - loss: 0.5323 - main_output_loss: 0.4769 - aux_output_loss: 0.5545 - main_output_acc: 0.7825 - aux_output_acc: 0.7229 - val_loss: 0.5368 - val_main_output_loss: 0.4814 - val_aux_output_loss: 0.5543 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7379\nEpoch 414/500\n924/924 [==============================] - 0s 207us/step - loss: 0.5330 - main_output_loss: 0.4775 - aux_output_loss: 0.5549 - main_output_acc: 0.7727 - aux_output_acc: 0.7251 - val_loss: 0.5432 - val_main_output_loss: 0.4877 - val_aux_output_loss: 0.5544 - val_main_output_acc: 0.7670 - val_aux_output_acc: 0.7282\nEpoch 415/500\n924/924 [==============================] - 0s 218us/step - loss: 0.5322 - main_output_loss: 0.4768 - aux_output_loss: 0.5544 - main_output_acc: 0.7716 - aux_output_acc: 0.7240 - val_loss: 0.5449 - val_main_output_loss: 0.4894 - val_aux_output_loss: 0.5543 - val_main_output_acc: 0.7573 - val_aux_output_acc: 0.7282\nEpoch 416/500\n924/924 [==============================] - 0s 214us/step - loss: 0.5326 - main_output_loss: 0.4772 - aux_output_loss: 0.5541 - main_output_acc: 0.7684 - aux_output_acc: 0.7229 - val_loss: 0.5375 - val_main_output_loss: 0.4821 - val_aux_output_loss: 0.5540 - val_main_output_acc: 0.8155 - val_aux_output_acc: 0.7379\nEpoch 417/500\n924/924 [==============================] - 0s 206us/step - loss: 0.5310 - main_output_loss: 0.4756 - aux_output_loss: 0.5538 - main_output_acc: 0.7803 - aux_output_acc: 0.7251 - val_loss: 0.5356 - val_main_output_loss: 0.4802 - val_aux_output_loss: 0.5537 - val_main_output_acc: 0.8155 - val_aux_output_acc: 0.7379\nEpoch 418/500\n924/924 [==============================] - 0s 208us/step - loss: 0.5315 - main_output_loss: 0.4761 - aux_output_loss: 0.5539 - main_output_acc: 0.7749 - aux_output_acc: 0.7240 - val_loss: 0.5353 - val_main_output_loss: 0.4800 - val_aux_output_loss: 0.5534 - val_main_output_acc: 0.8155 - val_aux_output_acc: 0.7379\nEpoch 419/500\n924/924 [==============================] - 0s 211us/step - loss: 0.5317 - main_output_loss: 0.4763 - aux_output_loss: 0.5537 - main_output_acc: 0.7781 - aux_output_acc: 0.7229 - val_loss: 0.5408 - val_main_output_loss: 0.4854 - val_aux_output_loss: 0.5535 - val_main_output_acc: 0.7864 - val_aux_output_acc: 0.7379\nEpoch 420/500\n924/924 [==============================] - 0s 208us/step - loss: 0.5295 - main_output_loss: 0.4742 - aux_output_loss: 0.5532 - main_output_acc: 0.7738 - aux_output_acc: 0.7240 - val_loss: 0.5362 - val_main_output_loss: 0.4809 - val_aux_output_loss: 0.5530 - val_main_output_acc: 0.8155 - val_aux_output_acc: 0.7379\nEpoch 421/500\n924/924 [==============================] - 0s 211us/step - loss: 0.5312 - main_output_loss: 0.4759 - aux_output_loss: 0.5533 - main_output_acc: 0.7727 - aux_output_acc: 0.7240 - val_loss: 0.5347 - val_main_output_loss: 0.4795 - val_aux_output_loss: 0.5528 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7379\nEpoch 422/500\n924/924 [==============================] - 0s 213us/step - loss: 0.5302 - main_output_loss: 0.4749 - aux_output_loss: 0.5530 - main_output_acc: 0.7727 - aux_output_acc: 0.7262 - val_loss: 0.5350 - val_main_output_loss: 0.4798 - val_aux_output_loss: 0.5526 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7379\nEpoch 423/500\n924/924 [==============================] - 0s 209us/step - loss: 0.5302 - main_output_loss: 0.4749 - aux_output_loss: 0.5529 - main_output_acc: 0.7803 - aux_output_acc: 0.7240 - val_loss: 0.5377 - val_main_output_loss: 0.4825 - val_aux_output_loss: 0.5525 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7379\nEpoch 424/500\n924/924 [==============================] - 0s 205us/step - loss: 0.5291 - main_output_loss: 0.4738 - aux_output_loss: 0.5526 - main_output_acc: 0.7781 - aux_output_acc: 0.7229 - val_loss: 0.5441 - val_main_output_loss: 0.4889 - val_aux_output_loss: 0.5525 - val_main_output_acc: 0.7573 - val_aux_output_acc: 0.7379\nEpoch 425/500\n924/924 [==============================] - 0s 208us/step - loss: 0.5297 - main_output_loss: 0.4744 - aux_output_loss: 0.5522 - main_output_acc: 0.7716 - aux_output_acc: 0.7229 - val_loss: 0.5376 - val_main_output_loss: 0.4824 - val_aux_output_loss: 0.5521 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7379\nEpoch 426/500\n924/924 [==============================] - 0s 213us/step - loss: 0.5277 - main_output_loss: 0.4725 - aux_output_loss: 0.5521 - main_output_acc: 0.7803 - aux_output_acc: 0.7262 - val_loss: 0.5462 - val_main_output_loss: 0.4909 - val_aux_output_loss: 0.5523 - val_main_output_acc: 0.7573 - val_aux_output_acc: 0.7379\nEpoch 427/500\n924/924 [==============================] - 0s 215us/step - loss: 0.5280 - main_output_loss: 0.4729 - aux_output_loss: 0.5518 - main_output_acc: 0.7771 - aux_output_acc: 0.7229 - val_loss: 0.5395 - val_main_output_loss: 0.4843 - val_aux_output_loss: 0.5519 - val_main_output_acc: 0.7767 - val_aux_output_acc: 0.7379\nEpoch 428/500\n924/924 [==============================] - 0s 207us/step - loss: 0.5282 - main_output_loss: 0.4731 - aux_output_loss: 0.5517 - main_output_acc: 0.7825 - aux_output_acc: 0.7229 - val_loss: 0.5355 - val_main_output_loss: 0.4804 - val_aux_output_loss: 0.5515 - val_main_output_acc: 0.8155 - val_aux_output_acc: 0.7379\nEpoch 429/500\n924/924 [==============================] - 0s 204us/step - loss: 0.5273 - main_output_loss: 0.4722 - aux_output_loss: 0.5516 - main_output_acc: 0.7846 - aux_output_acc: 0.7219 - val_loss: 0.5345 - val_main_output_loss: 0.4794 - val_aux_output_loss: 0.5514 - val_main_output_acc: 0.8155 - val_aux_output_acc: 0.7379\nEpoch 430/500\n924/924 [==============================] - 0s 210us/step - loss: 0.5268 - main_output_loss: 0.4716 - aux_output_loss: 0.5513 - main_output_acc: 0.7781 - aux_output_acc: 0.7240 - val_loss: 0.5384 - val_main_output_loss: 0.4833 - val_aux_output_loss: 0.5515 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7379\nEpoch 431/500\n924/924 [==============================] - 0s 210us/step - loss: 0.5267 - main_output_loss: 0.4716 - aux_output_loss: 0.5513 - main_output_acc: 0.7727 - aux_output_acc: 0.7273 - val_loss: 0.5350 - val_main_output_loss: 0.4798 - val_aux_output_loss: 0.5514 - val_main_output_acc: 0.8155 - val_aux_output_acc: 0.7379\nEpoch 432/500\n924/924 [==============================] - 0s 209us/step - loss: 0.5270 - main_output_loss: 0.4718 - aux_output_loss: 0.5513 - main_output_acc: 0.7760 - aux_output_acc: 0.7240 - val_loss: 0.5344 - val_main_output_loss: 0.4793 - val_aux_output_loss: 0.5511 - val_main_output_acc: 0.8155 - val_aux_output_acc: 0.7379\nEpoch 433/500\n924/924 [==============================] - 0s 219us/step - loss: 0.5257 - main_output_loss: 0.4706 - aux_output_loss: 0.5510 - main_output_acc: 0.7792 - aux_output_acc: 0.7262 - val_loss: 0.5353 - val_main_output_loss: 0.4801 - val_aux_output_loss: 0.5512 - val_main_output_acc: 0.8155 - val_aux_output_acc: 0.7379\nEpoch 434/500\n924/924 [==============================] - 0s 227us/step - loss: 0.5283 - main_output_loss: 0.4732 - aux_output_loss: 0.5512 - main_output_acc: 0.7771 - aux_output_acc: 0.7273 - val_loss: 0.5336 - val_main_output_loss: 0.4785 - val_aux_output_loss: 0.5509 - val_main_output_acc: 0.8155 - val_aux_output_acc: 0.7379\nEpoch 435/500\n924/924 [==============================] - 0s 220us/step - loss: 0.5258 - main_output_loss: 0.4707 - aux_output_loss: 0.5509 - main_output_acc: 0.7803 - aux_output_acc: 0.7262 - val_loss: 0.5369 - val_main_output_loss: 0.4819 - val_aux_output_loss: 0.5507 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7379\nEpoch 436/500\n924/924 [==============================] - 0s 224us/step - loss: 0.5248 - main_output_loss: 0.4698 - aux_output_loss: 0.5504 - main_output_acc: 0.7792 - aux_output_acc: 0.7251 - val_loss: 0.5350 - val_main_output_loss: 0.4800 - val_aux_output_loss: 0.5503 - val_main_output_acc: 0.8155 - val_aux_output_acc: 0.7379\nEpoch 437/500\n924/924 [==============================] - 0s 217us/step - loss: 0.5245 - main_output_loss: 0.4695 - aux_output_loss: 0.5504 - main_output_acc: 0.7835 - aux_output_acc: 0.7262 - val_loss: 0.5332 - val_main_output_loss: 0.4782 - val_aux_output_loss: 0.5500 - val_main_output_acc: 0.8155 - val_aux_output_acc: 0.7379\nEpoch 438/500\n924/924 [==============================] - 0s 211us/step - loss: 0.5243 - main_output_loss: 0.4693 - aux_output_loss: 0.5502 - main_output_acc: 0.7857 - aux_output_acc: 0.7294 - val_loss: 0.5366 - val_main_output_loss: 0.4816 - val_aux_output_loss: 0.5500 - val_main_output_acc: 0.7767 - val_aux_output_acc: 0.7379\nEpoch 439/500\n924/924 [==============================] - 0s 212us/step - loss: 0.5241 - main_output_loss: 0.4691 - aux_output_loss: 0.5500 - main_output_acc: 0.7803 - aux_output_acc: 0.7284 - val_loss: 0.5409 - val_main_output_loss: 0.4859 - val_aux_output_loss: 0.5498 - val_main_output_acc: 0.7670 - val_aux_output_acc: 0.7379\nEpoch 440/500\n924/924 [==============================] - 0s 212us/step - loss: 0.5238 - main_output_loss: 0.4688 - aux_output_loss: 0.5493 - main_output_acc: 0.7792 - aux_output_acc: 0.7327 - val_loss: 0.5322 - val_main_output_loss: 0.4773 - val_aux_output_loss: 0.5492 - val_main_output_acc: 0.8155 - val_aux_output_acc: 0.7379\nEpoch 441/500\n924/924 [==============================] - 0s 212us/step - loss: 0.5239 - main_output_loss: 0.4689 - aux_output_loss: 0.5498 - main_output_acc: 0.7771 - aux_output_acc: 0.7316 - val_loss: 0.5403 - val_main_output_loss: 0.4853 - val_aux_output_loss: 0.5493 - val_main_output_acc: 0.7670 - val_aux_output_acc: 0.7379\nEpoch 442/500\n924/924 [==============================] - 0s 205us/step - loss: 0.5233 - main_output_loss: 0.4684 - aux_output_loss: 0.5492 - main_output_acc: 0.7792 - aux_output_acc: 0.7273 - val_loss: 0.5310 - val_main_output_loss: 0.4761 - val_aux_output_loss: 0.5487 - val_main_output_acc: 0.8155 - val_aux_output_acc: 0.7379\nEpoch 443/500\n924/924 [==============================] - 0s 210us/step - loss: 0.5231 - main_output_loss: 0.4681 - aux_output_loss: 0.5493 - main_output_acc: 0.7825 - aux_output_acc: 0.7305 - val_loss: 0.5396 - val_main_output_loss: 0.4847 - val_aux_output_loss: 0.5489 - val_main_output_acc: 0.7670 - val_aux_output_acc: 0.7379\nEpoch 444/500\n924/924 [==============================] - 0s 212us/step - loss: 0.5226 - main_output_loss: 0.4677 - aux_output_loss: 0.5489 - main_output_acc: 0.7814 - aux_output_acc: 0.7284 - val_loss: 0.5355 - val_main_output_loss: 0.4806 - val_aux_output_loss: 0.5487 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7379\nEpoch 445/500\n924/924 [==============================] - 0s 208us/step - loss: 0.5226 - main_output_loss: 0.4677 - aux_output_loss: 0.5489 - main_output_acc: 0.7814 - aux_output_acc: 0.7294 - val_loss: 0.5362 - val_main_output_loss: 0.4813 - val_aux_output_loss: 0.5485 - val_main_output_acc: 0.7767 - val_aux_output_acc: 0.7379\nEpoch 446/500\n924/924 [==============================] - 0s 207us/step - loss: 0.5215 - main_output_loss: 0.4666 - aux_output_loss: 0.5485 - main_output_acc: 0.7760 - aux_output_acc: 0.7294 - val_loss: 0.5310 - val_main_output_loss: 0.4761 - val_aux_output_loss: 0.5484 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7379\nEpoch 447/500\n924/924 [==============================] - 0s 209us/step - loss: 0.5234 - main_output_loss: 0.4685 - aux_output_loss: 0.5491 - main_output_acc: 0.7846 - aux_output_acc: 0.7316 - val_loss: 0.5306 - val_main_output_loss: 0.4758 - val_aux_output_loss: 0.5482 - val_main_output_acc: 0.8155 - val_aux_output_acc: 0.7379\nEpoch 448/500\n924/924 [==============================] - 0s 208us/step - loss: 0.5226 - main_output_loss: 0.4677 - aux_output_loss: 0.5488 - main_output_acc: 0.7825 - aux_output_acc: 0.7348 - val_loss: 0.5351 - val_main_output_loss: 0.4803 - val_aux_output_loss: 0.5482 - val_main_output_acc: 0.7767 - val_aux_output_acc: 0.7379\nEpoch 449/500\n924/924 [==============================] - 0s 206us/step - loss: 0.5213 - main_output_loss: 0.4665 - aux_output_loss: 0.5484 - main_output_acc: 0.7803 - aux_output_acc: 0.7327 - val_loss: 0.5438 - val_main_output_loss: 0.4890 - val_aux_output_loss: 0.5484 - val_main_output_acc: 0.7573 - val_aux_output_acc: 0.7379\nEpoch 450/500\n924/924 [==============================] - 0s 209us/step - loss: 0.5224 - main_output_loss: 0.4675 - aux_output_loss: 0.5481 - main_output_acc: 0.7792 - aux_output_acc: 0.7305 - val_loss: 0.5310 - val_main_output_loss: 0.4762 - val_aux_output_loss: 0.5481 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7379\nEpoch 451/500\n924/924 [==============================] - 0s 205us/step - loss: 0.5215 - main_output_loss: 0.4667 - aux_output_loss: 0.5485 - main_output_acc: 0.7868 - aux_output_acc: 0.7327 - val_loss: 0.5310 - val_main_output_loss: 0.4763 - val_aux_output_loss: 0.5477 - val_main_output_acc: 0.8155 - val_aux_output_acc: 0.7379\nEpoch 452/500\n924/924 [==============================] - 0s 206us/step - loss: 0.5201 - main_output_loss: 0.4653 - aux_output_loss: 0.5481 - main_output_acc: 0.7760 - aux_output_acc: 0.7327 - val_loss: 0.5392 - val_main_output_loss: 0.4844 - val_aux_output_loss: 0.5476 - val_main_output_acc: 0.7670 - val_aux_output_acc: 0.7379\nEpoch 453/500\n924/924 [==============================] - 0s 218us/step - loss: 0.5201 - main_output_loss: 0.4653 - aux_output_loss: 0.5476 - main_output_acc: 0.7825 - aux_output_acc: 0.7327 - val_loss: 0.5353 - val_main_output_loss: 0.4805 - val_aux_output_loss: 0.5473 - val_main_output_acc: 0.7767 - val_aux_output_acc: 0.7379\nEpoch 454/500\n924/924 [==============================] - 0s 216us/step - loss: 0.5207 - main_output_loss: 0.4659 - aux_output_loss: 0.5477 - main_output_acc: 0.7792 - aux_output_acc: 0.7348 - val_loss: 0.5322 - val_main_output_loss: 0.4775 - val_aux_output_loss: 0.5471 - val_main_output_acc: 0.8155 - val_aux_output_acc: 0.7379\nEpoch 455/500\n924/924 [==============================] - 0s 219us/step - loss: 0.5199 - main_output_loss: 0.4651 - aux_output_loss: 0.5475 - main_output_acc: 0.7814 - aux_output_acc: 0.7338 - val_loss: 0.5306 - val_main_output_loss: 0.4759 - val_aux_output_loss: 0.5467 - val_main_output_acc: 0.8155 - val_aux_output_acc: 0.7379\nEpoch 456/500\n924/924 [==============================] - 0s 226us/step - loss: 0.5201 - main_output_loss: 0.4654 - aux_output_loss: 0.5475 - main_output_acc: 0.7825 - aux_output_acc: 0.7381 - val_loss: 0.5361 - val_main_output_loss: 0.4814 - val_aux_output_loss: 0.5466 - val_main_output_acc: 0.7767 - val_aux_output_acc: 0.7476\nEpoch 457/500\n924/924 [==============================] - 0s 222us/step - loss: 0.5185 - main_output_loss: 0.4638 - aux_output_loss: 0.5469 - main_output_acc: 0.7835 - aux_output_acc: 0.7338 - val_loss: 0.5352 - val_main_output_loss: 0.4806 - val_aux_output_loss: 0.5464 - val_main_output_acc: 0.7864 - val_aux_output_acc: 0.7476\nEpoch 458/500\n924/924 [==============================] - 0s 217us/step - loss: 0.5188 - main_output_loss: 0.4641 - aux_output_loss: 0.5469 - main_output_acc: 0.7835 - aux_output_acc: 0.7338 - val_loss: 0.5382 - val_main_output_loss: 0.4836 - val_aux_output_loss: 0.5461 - val_main_output_acc: 0.7767 - val_aux_output_acc: 0.7476\nEpoch 459/500\n924/924 [==============================] - 0s 213us/step - loss: 0.5181 - main_output_loss: 0.4635 - aux_output_loss: 0.5465 - main_output_acc: 0.7803 - aux_output_acc: 0.7370 - val_loss: 0.5412 - val_main_output_loss: 0.4867 - val_aux_output_loss: 0.5458 - val_main_output_acc: 0.7573 - val_aux_output_acc: 0.7476\nEpoch 460/500\n924/924 [==============================] - 0s 218us/step - loss: 0.5193 - main_output_loss: 0.4646 - aux_output_loss: 0.5464 - main_output_acc: 0.7835 - aux_output_acc: 0.7338 - val_loss: 0.5378 - val_main_output_loss: 0.4833 - val_aux_output_loss: 0.5452 - val_main_output_acc: 0.7767 - val_aux_output_acc: 0.7476\nEpoch 461/500\n924/924 [==============================] - 0s 217us/step - loss: 0.5181 - main_output_loss: 0.4635 - aux_output_loss: 0.5463 - main_output_acc: 0.7771 - aux_output_acc: 0.7370 - val_loss: 0.5300 - val_main_output_loss: 0.4755 - val_aux_output_loss: 0.5451 - val_main_output_acc: 0.8155 - val_aux_output_acc: 0.7476\nEpoch 462/500\n924/924 [==============================] - 0s 218us/step - loss: 0.5172 - main_output_loss: 0.4626 - aux_output_loss: 0.5462 - main_output_acc: 0.7803 - aux_output_acc: 0.7392 - val_loss: 0.5306 - val_main_output_loss: 0.4761 - val_aux_output_loss: 0.5448 - val_main_output_acc: 0.8155 - val_aux_output_acc: 0.7476\nEpoch 463/500\n924/924 [==============================] - 0s 218us/step - loss: 0.5181 - main_output_loss: 0.4634 - aux_output_loss: 0.5463 - main_output_acc: 0.7803 - aux_output_acc: 0.7370 - val_loss: 0.5306 - val_main_output_loss: 0.4761 - val_aux_output_loss: 0.5447 - val_main_output_acc: 0.8155 - val_aux_output_acc: 0.7476\nEpoch 464/500\n924/924 [==============================] - 0s 219us/step - loss: 0.5160 - main_output_loss: 0.4615 - aux_output_loss: 0.5459 - main_output_acc: 0.7803 - aux_output_acc: 0.7392 - val_loss: 0.5387 - val_main_output_loss: 0.4842 - val_aux_output_loss: 0.5448 - val_main_output_acc: 0.7767 - val_aux_output_acc: 0.7476\nEpoch 465/500\n924/924 [==============================] - 0s 217us/step - loss: 0.5172 - main_output_loss: 0.4626 - aux_output_loss: 0.5456 - main_output_acc: 0.7857 - aux_output_acc: 0.7370 - val_loss: 0.5351 - val_main_output_loss: 0.4806 - val_aux_output_loss: 0.5449 - val_main_output_acc: 0.7767 - val_aux_output_acc: 0.7476\nEpoch 466/500\n924/924 [==============================] - 0s 219us/step - loss: 0.5158 - main_output_loss: 0.4613 - aux_output_loss: 0.5454 - main_output_acc: 0.7825 - aux_output_acc: 0.7370 - val_loss: 0.5290 - val_main_output_loss: 0.4745 - val_aux_output_loss: 0.5446 - val_main_output_acc: 0.8155 - val_aux_output_acc: 0.7476\nEpoch 467/500\n924/924 [==============================] - 0s 221us/step - loss: 0.5163 - main_output_loss: 0.4618 - aux_output_loss: 0.5455 - main_output_acc: 0.7825 - aux_output_acc: 0.7338 - val_loss: 0.5337 - val_main_output_loss: 0.4792 - val_aux_output_loss: 0.5447 - val_main_output_acc: 0.7767 - val_aux_output_acc: 0.7476\nEpoch 468/500\n924/924 [==============================] - 0s 222us/step - loss: 0.5157 - main_output_loss: 0.4612 - aux_output_loss: 0.5452 - main_output_acc: 0.7868 - aux_output_acc: 0.7381 - val_loss: 0.5280 - val_main_output_loss: 0.4736 - val_aux_output_loss: 0.5446 - val_main_output_acc: 0.8155 - val_aux_output_acc: 0.7476\nEpoch 469/500\n924/924 [==============================] - 0s 215us/step - loss: 0.5155 - main_output_loss: 0.4610 - aux_output_loss: 0.5452 - main_output_acc: 0.7814 - aux_output_acc: 0.7381 - val_loss: 0.5346 - val_main_output_loss: 0.4801 - val_aux_output_loss: 0.5445 - val_main_output_acc: 0.7767 - val_aux_output_acc: 0.7476\nEpoch 470/500\n924/924 [==============================] - 0s 222us/step - loss: 0.5139 - main_output_loss: 0.4595 - aux_output_loss: 0.5445 - main_output_acc: 0.7835 - aux_output_acc: 0.7403 - val_loss: 0.5278 - val_main_output_loss: 0.4734 - val_aux_output_loss: 0.5444 - val_main_output_acc: 0.8155 - val_aux_output_acc: 0.7476\nEpoch 471/500\n924/924 [==============================] - 0s 222us/step - loss: 0.5158 - main_output_loss: 0.4613 - aux_output_loss: 0.5449 - main_output_acc: 0.7857 - aux_output_acc: 0.7370 - val_loss: 0.5293 - val_main_output_loss: 0.4748 - val_aux_output_loss: 0.5444 - val_main_output_acc: 0.8155 - val_aux_output_acc: 0.7476\nEpoch 472/500\n924/924 [==============================] - 0s 214us/step - loss: 0.5137 - main_output_loss: 0.4593 - aux_output_loss: 0.5446 - main_output_acc: 0.7868 - aux_output_acc: 0.7370 - val_loss: 0.5297 - val_main_output_loss: 0.4753 - val_aux_output_loss: 0.5439 - val_main_output_acc: 0.8155 - val_aux_output_acc: 0.7476\nEpoch 473/500\n924/924 [==============================] - 0s 215us/step - loss: 0.5137 - main_output_loss: 0.4592 - aux_output_loss: 0.5444 - main_output_acc: 0.7857 - aux_output_acc: 0.7381 - val_loss: 0.5322 - val_main_output_loss: 0.4778 - val_aux_output_loss: 0.5435 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7476\nEpoch 474/500\n924/924 [==============================] - 0s 225us/step - loss: 0.5130 - main_output_loss: 0.4586 - aux_output_loss: 0.5438 - main_output_acc: 0.7857 - aux_output_acc: 0.7370 - val_loss: 0.5274 - val_main_output_loss: 0.4731 - val_aux_output_loss: 0.5430 - val_main_output_acc: 0.8155 - val_aux_output_acc: 0.7379\nEpoch 475/500\n924/924 [==============================] - 0s 218us/step - loss: 0.5147 - main_output_loss: 0.4603 - aux_output_loss: 0.5440 - main_output_acc: 0.7814 - aux_output_acc: 0.7403 - val_loss: 0.5288 - val_main_output_loss: 0.4745 - val_aux_output_loss: 0.5426 - val_main_output_acc: 0.8155 - val_aux_output_acc: 0.7476\nEpoch 476/500\n924/924 [==============================] - 0s 220us/step - loss: 0.5126 - main_output_loss: 0.4582 - aux_output_loss: 0.5434 - main_output_acc: 0.7846 - aux_output_acc: 0.7424 - val_loss: 0.5288 - val_main_output_loss: 0.4746 - val_aux_output_loss: 0.5426 - val_main_output_acc: 0.8155 - val_aux_output_acc: 0.7476\nEpoch 477/500\n924/924 [==============================] - 0s 218us/step - loss: 0.5143 - main_output_loss: 0.4600 - aux_output_loss: 0.5436 - main_output_acc: 0.7814 - aux_output_acc: 0.7370 - val_loss: 0.5344 - val_main_output_loss: 0.4801 - val_aux_output_loss: 0.5429 - val_main_output_acc: 0.7767 - val_aux_output_acc: 0.7476\nEpoch 478/500\n924/924 [==============================] - 0s 218us/step - loss: 0.5132 - main_output_loss: 0.4589 - aux_output_loss: 0.5435 - main_output_acc: 0.7803 - aux_output_acc: 0.7370 - val_loss: 0.5274 - val_main_output_loss: 0.4731 - val_aux_output_loss: 0.5431 - val_main_output_acc: 0.8155 - val_aux_output_acc: 0.7379\nEpoch 479/500\n924/924 [==============================] - 0s 223us/step - loss: 0.5140 - main_output_loss: 0.4596 - aux_output_loss: 0.5437 - main_output_acc: 0.7846 - aux_output_acc: 0.7381 - val_loss: 0.5329 - val_main_output_loss: 0.4786 - val_aux_output_loss: 0.5429 - val_main_output_acc: 0.7864 - val_aux_output_acc: 0.7476\nEpoch 480/500\n924/924 [==============================] - 0s 222us/step - loss: 0.5119 - main_output_loss: 0.4576 - aux_output_loss: 0.5431 - main_output_acc: 0.7803 - aux_output_acc: 0.7392 - val_loss: 0.5359 - val_main_output_loss: 0.4816 - val_aux_output_loss: 0.5430 - val_main_output_acc: 0.7767 - val_aux_output_acc: 0.7476\nEpoch 481/500\n924/924 [==============================] - 0s 212us/step - loss: 0.5118 - main_output_loss: 0.4575 - aux_output_loss: 0.5429 - main_output_acc: 0.7857 - aux_output_acc: 0.7413 - val_loss: 0.5278 - val_main_output_loss: 0.4735 - val_aux_output_loss: 0.5429 - val_main_output_acc: 0.8155 - val_aux_output_acc: 0.7379\nEpoch 482/500\n924/924 [==============================] - 0s 211us/step - loss: 0.5116 - main_output_loss: 0.4573 - aux_output_loss: 0.5431 - main_output_acc: 0.7846 - aux_output_acc: 0.7413 - val_loss: 0.5340 - val_main_output_loss: 0.4797 - val_aux_output_loss: 0.5428 - val_main_output_acc: 0.7767 - val_aux_output_acc: 0.7476\nEpoch 483/500\n924/924 [==============================] - 0s 207us/step - loss: 0.5114 - main_output_loss: 0.4571 - aux_output_loss: 0.5426 - main_output_acc: 0.7922 - aux_output_acc: 0.7424 - val_loss: 0.5305 - val_main_output_loss: 0.4762 - val_aux_output_loss: 0.5430 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7476\nEpoch 484/500\n924/924 [==============================] - 0s 209us/step - loss: 0.5118 - main_output_loss: 0.4575 - aux_output_loss: 0.5430 - main_output_acc: 0.7846 - aux_output_acc: 0.7392 - val_loss: 0.5301 - val_main_output_loss: 0.4758 - val_aux_output_loss: 0.5429 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7476\nEpoch 485/500\n924/924 [==============================] - 0s 208us/step - loss: 0.5109 - main_output_loss: 0.4567 - aux_output_loss: 0.5426 - main_output_acc: 0.7890 - aux_output_acc: 0.7413 - val_loss: 0.5335 - val_main_output_loss: 0.4793 - val_aux_output_loss: 0.5429 - val_main_output_acc: 0.7767 - val_aux_output_acc: 0.7476\nEpoch 486/500\n924/924 [==============================] - 0s 208us/step - loss: 0.5106 - main_output_loss: 0.4563 - aux_output_loss: 0.5424 - main_output_acc: 0.7900 - aux_output_acc: 0.7413 - val_loss: 0.5300 - val_main_output_loss: 0.4757 - val_aux_output_loss: 0.5426 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7476\nEpoch 487/500\n924/924 [==============================] - 0s 208us/step - loss: 0.5130 - main_output_loss: 0.4587 - aux_output_loss: 0.5430 - main_output_acc: 0.7890 - aux_output_acc: 0.7392 - val_loss: 0.5356 - val_main_output_loss: 0.4813 - val_aux_output_loss: 0.5425 - val_main_output_acc: 0.7767 - val_aux_output_acc: 0.7476\nEpoch 488/500\n924/924 [==============================] - 0s 212us/step - loss: 0.5119 - main_output_loss: 0.4576 - aux_output_loss: 0.5424 - main_output_acc: 0.7825 - aux_output_acc: 0.7435 - val_loss: 0.5293 - val_main_output_loss: 0.4750 - val_aux_output_loss: 0.5423 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7476\nEpoch 489/500\n924/924 [==============================] - 0s 202us/step - loss: 0.5095 - main_output_loss: 0.4553 - aux_output_loss: 0.5419 - main_output_acc: 0.7879 - aux_output_acc: 0.7413 - val_loss: 0.5268 - val_main_output_loss: 0.4726 - val_aux_output_loss: 0.5420 - val_main_output_acc: 0.8155 - val_aux_output_acc: 0.7476\nEpoch 490/500\n924/924 [==============================] - 0s 204us/step - loss: 0.5099 - main_output_loss: 0.4557 - aux_output_loss: 0.5419 - main_output_acc: 0.7900 - aux_output_acc: 0.7403 - val_loss: 0.5307 - val_main_output_loss: 0.4766 - val_aux_output_loss: 0.5417 - val_main_output_acc: 0.7864 - val_aux_output_acc: 0.7476\nEpoch 491/500\n924/924 [==============================] - 0s 210us/step - loss: 0.5085 - main_output_loss: 0.4544 - aux_output_loss: 0.5414 - main_output_acc: 0.7857 - aux_output_acc: 0.7424 - val_loss: 0.5261 - val_main_output_loss: 0.4720 - val_aux_output_loss: 0.5415 - val_main_output_acc: 0.8155 - val_aux_output_acc: 0.7476\nEpoch 492/500\n924/924 [==============================] - 0s 210us/step - loss: 0.5091 - main_output_loss: 0.4550 - aux_output_loss: 0.5414 - main_output_acc: 0.7857 - aux_output_acc: 0.7392 - val_loss: 0.5269 - val_main_output_loss: 0.4728 - val_aux_output_loss: 0.5410 - val_main_output_acc: 0.8155 - val_aux_output_acc: 0.7476\nEpoch 493/500\n924/924 [==============================] - 0s 216us/step - loss: 0.5081 - main_output_loss: 0.4540 - aux_output_loss: 0.5410 - main_output_acc: 0.7879 - aux_output_acc: 0.7424 - val_loss: 0.5289 - val_main_output_loss: 0.4748 - val_aux_output_loss: 0.5410 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7476\nEpoch 494/500\n924/924 [==============================] - 0s 210us/step - loss: 0.5083 - main_output_loss: 0.4542 - aux_output_loss: 0.5408 - main_output_acc: 0.7835 - aux_output_acc: 0.7392 - val_loss: 0.5249 - val_main_output_loss: 0.4708 - val_aux_output_loss: 0.5410 - val_main_output_acc: 0.8155 - val_aux_output_acc: 0.7379\nEpoch 495/500\n924/924 [==============================] - 0s 206us/step - loss: 0.5083 - main_output_loss: 0.4542 - aux_output_loss: 0.5410 - main_output_acc: 0.7922 - aux_output_acc: 0.7446 - val_loss: 0.5381 - val_main_output_loss: 0.4840 - val_aux_output_loss: 0.5412 - val_main_output_acc: 0.7670 - val_aux_output_acc: 0.7573\nEpoch 496/500\n924/924 [==============================] - 0s 211us/step - loss: 0.5085 - main_output_loss: 0.4544 - aux_output_loss: 0.5406 - main_output_acc: 0.7890 - aux_output_acc: 0.7446 - val_loss: 0.5383 - val_main_output_loss: 0.4842 - val_aux_output_loss: 0.5413 - val_main_output_acc: 0.7767 - val_aux_output_acc: 0.7573\nEpoch 497/500\n924/924 [==============================] - 0s 203us/step - loss: 0.5083 - main_output_loss: 0.4543 - aux_output_loss: 0.5404 - main_output_acc: 0.7825 - aux_output_acc: 0.7392 - val_loss: 0.5277 - val_main_output_loss: 0.4737 - val_aux_output_loss: 0.5407 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7476\nEpoch 498/500\n924/924 [==============================] - 0s 204us/step - loss: 0.5076 - main_output_loss: 0.4536 - aux_output_loss: 0.5404 - main_output_acc: 0.7846 - aux_output_acc: 0.7413 - val_loss: 0.5243 - val_main_output_loss: 0.4702 - val_aux_output_loss: 0.5407 - val_main_output_acc: 0.8155 - val_aux_output_acc: 0.7379\nEpoch 499/500\n924/924 [==============================] - 0s 208us/step - loss: 0.5072 - main_output_loss: 0.4532 - aux_output_loss: 0.5404 - main_output_acc: 0.7911 - aux_output_acc: 0.7392 - val_loss: 0.5280 - val_main_output_loss: 0.4740 - val_aux_output_loss: 0.5405 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7476\nEpoch 500/500\n924/924 [==============================] - 0s 206us/step - loss: 0.5064 - main_output_loss: 0.4524 - aux_output_loss: 0.5402 - main_output_acc: 0.7911 - aux_output_acc: 0.7424 - val_loss: 0.5290 - val_main_output_loss: 0.4750 - val_aux_output_loss: 0.5401 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7476\n"
],
[
"# plot the fit\npred_main, pred_aux = model.predict([x_seq_test, x_static_test])\nroc = roc_curve(y_test, pred_main)\nauc = roc_auc_score(y_test, pred_main)\nfig = plt.figure(figsize=(4, 3)) # in inches\nplt.plot(roc[0], roc[1], color = 'darkorange', label = 'ROC curve\\n(area = %0.2f)' % auc)\nplt.plot([0, 1], [0, 1], color= 'navy', linestyle = '--')\nplt.xlabel('False Positive Rate')\nplt.ylabel('True Positive Rate')\nplt.title('%s: ROC' % 'GRU-base')\nplt.legend(loc = \"lower right\")\nfig_name = 'gru-base.pdf'\nfig.savefig(os.path.join(fig_fp, fig_name), bbox_inches='tight')\nplt.show()",
"_____no_output_____"
],
[
"# plot training and validation loss and accuracy\n\nacc = history.history['main_output_acc']\nval_acc = history.history['val_main_output_acc']\nloss = history.history['main_output_loss']\nval_loss = history.history['val_loss']\n\nepochs = range(len(acc))\n\nplt.plot(epochs, acc, 'bo', label='Training acc')\nplt.plot(epochs, val_acc, 'b', label='Validation acc')\nplt.title('Training and validation accuracy')\nplt.legend()\n\nplt.figure()\nplt.plot(epochs, loss, 'bo', label='Training loss')\nplt.plot(epochs, val_loss, 'b', label='Validation loss')\nplt.title('Training and validation loss')\nplt.legend()\n\nfig_name = 'loss_svg.svg'\nfig.savefig('./loss_svg.svg', bbox_inches='tight')",
"_____no_output_____"
]
],
[
[
"### 10% Dropout",
"_____no_output_____"
]
],
[
[
"# Define inputs\nsequence_input = Input(shape = (x_seq_train.shape[1], x_seq_train.shape[2], ), dtype = 'float32', name = 'sequence_input')\nstatic_input = Input(shape = (x_static_train.shape[1], ), name = 'static_input')\n\n# Network architecture\nseq_x = GRU(units = 128)(sequence_input)\n\n# Seperate output for the GRU later\nseq_aux_output = Dense(1, activation='sigmoid', name='aux_output')(seq_x)\n\n# Merge dual inputs\nx = concatenate([seq_x, static_input])\n\n# We stack a deep fully-connected network on the merged inputs\nx = Dense(128, activation = 'relu')(x)\nx = Dense(128, activation = 'relu')(x)\nx = Dropout(0.10)(x)\nx = Dense(128, activation = 'relu')(x)\nx = Dense(128, activation = 'relu')(x)\n\n# Sigmoid output layer\nmain_output = Dense(1, activation='sigmoid', name='main_output')(x)\n\n# optimizer\nopt = rmsprop(lr = 0.00001)\n\n# build model\nmodel = Model(inputs = [sequence_input, static_input], outputs = [main_output, seq_aux_output])\nmodel.compile(optimizer = opt, loss = 'binary_crossentropy', metrics = ['accuracy'], loss_weights = [1, 0.1])\n\n# save a plot of the model\n#plot_model(model, to_file='experiment_GRU-DO.svg')\n\n# fit the model\nhistory = model.fit([x_seq_train, x_static_train], [y_train, y_train], epochs = 500, batch_size = 128,\\\n validation_data=([x_seq_val, x_static_val], [y_val, y_val]),)\n\n",
"Train on 924 samples, validate on 103 samples\nEpoch 1/500\n924/924 [==============================] - 1s 1ms/step - loss: 1.2946 - main_output_loss: 1.2198 - aux_output_loss: 0.7480 - main_output_acc: 0.5152 - aux_output_acc: 0.5087 - val_loss: 1.1543 - val_main_output_loss: 1.0797 - val_aux_output_loss: 0.7468 - val_main_output_acc: 0.5437 - val_aux_output_acc: 0.5243\nEpoch 2/500\n924/924 [==============================] - 0s 226us/step - loss: 1.1701 - main_output_loss: 1.0959 - aux_output_loss: 0.7427 - main_output_acc: 0.5152 - aux_output_acc: 0.5076 - val_loss: 1.0828 - val_main_output_loss: 1.0085 - val_aux_output_loss: 0.7431 - val_main_output_acc: 0.5437 - val_aux_output_acc: 0.5146\nEpoch 3/500\n924/924 [==============================] - 0s 218us/step - loss: 1.1137 - main_output_loss: 1.0399 - aux_output_loss: 0.7385 - main_output_acc: 0.5152 - aux_output_acc: 0.5076 - val_loss: 1.0288 - val_main_output_loss: 0.9548 - val_aux_output_loss: 0.7399 - val_main_output_acc: 0.5437 - val_aux_output_acc: 0.5146\nEpoch 4/500\n924/924 [==============================] - 0s 210us/step - loss: 1.0426 - main_output_loss: 0.9691 - aux_output_loss: 0.7347 - main_output_acc: 0.5141 - aux_output_acc: 0.5087 - val_loss: 0.9887 - val_main_output_loss: 0.9149 - val_aux_output_loss: 0.7378 - val_main_output_acc: 0.5437 - val_aux_output_acc: 0.5146\nEpoch 5/500\n924/924 [==============================] - 0s 211us/step - loss: 1.0021 - main_output_loss: 0.9289 - aux_output_loss: 0.7321 - main_output_acc: 0.5162 - aux_output_acc: 0.5065 - val_loss: 0.9493 - val_main_output_loss: 0.8758 - val_aux_output_loss: 0.7348 - val_main_output_acc: 0.5437 - val_aux_output_acc: 0.4951\nEpoch 6/500\n924/924 [==============================] - 0s 210us/step - loss: 0.9431 - main_output_loss: 0.8703 - aux_output_loss: 0.7285 - main_output_acc: 0.5184 - aux_output_acc: 0.5065 - val_loss: 0.9162 - val_main_output_loss: 0.8430 - val_aux_output_loss: 0.7319 - val_main_output_acc: 0.5437 - val_aux_output_acc: 0.4951\nEpoch 7/500\n924/924 [==============================] - 0s 216us/step - loss: 0.9128 - main_output_loss: 0.8403 - aux_output_loss: 0.7247 - main_output_acc: 0.5076 - aux_output_acc: 0.5076 - val_loss: 0.8922 - val_main_output_loss: 0.8194 - val_aux_output_loss: 0.7286 - val_main_output_acc: 0.5340 - val_aux_output_acc: 0.4951\nEpoch 8/500\n924/924 [==============================] - 0s 217us/step - loss: 0.8946 - main_output_loss: 0.8225 - aux_output_loss: 0.7204 - main_output_acc: 0.5000 - aux_output_acc: 0.5076 - val_loss: 0.8741 - val_main_output_loss: 0.8015 - val_aux_output_loss: 0.7261 - val_main_output_acc: 0.5049 - val_aux_output_acc: 0.4951\nEpoch 9/500\n924/924 [==============================] - 0s 213us/step - loss: 0.8663 - main_output_loss: 0.7947 - aux_output_loss: 0.7169 - main_output_acc: 0.4848 - aux_output_acc: 0.5054 - val_loss: 0.8627 - val_main_output_loss: 0.7903 - val_aux_output_loss: 0.7235 - val_main_output_acc: 0.4951 - val_aux_output_acc: 0.5049\nEpoch 10/500\n924/924 [==============================] - 0s 218us/step - loss: 0.8382 - main_output_loss: 0.7669 - aux_output_loss: 0.7135 - main_output_acc: 0.4913 - aux_output_acc: 0.5065 - val_loss: 0.8537 - val_main_output_loss: 0.7816 - val_aux_output_loss: 0.7211 - val_main_output_acc: 0.4854 - val_aux_output_acc: 0.5049\nEpoch 11/500\n924/924 [==============================] - 0s 223us/step - loss: 0.8244 - main_output_loss: 0.7534 - aux_output_loss: 0.7101 - main_output_acc: 0.4968 - aux_output_acc: 0.5065 - val_loss: 0.8468 - val_main_output_loss: 0.7749 - val_aux_output_loss: 0.7184 - val_main_output_acc: 0.4951 - val_aux_output_acc: 0.4951\nEpoch 12/500\n924/924 [==============================] - 0s 209us/step - loss: 0.8302 - main_output_loss: 0.7596 - aux_output_loss: 0.7063 - main_output_acc: 0.4935 - aux_output_acc: 0.5043 - val_loss: 0.8397 - val_main_output_loss: 0.7681 - val_aux_output_loss: 0.7156 - val_main_output_acc: 0.4466 - val_aux_output_acc: 0.4854\nEpoch 13/500\n924/924 [==============================] - 0s 212us/step - loss: 0.8291 - main_output_loss: 0.7588 - aux_output_loss: 0.7026 - main_output_acc: 0.4578 - aux_output_acc: 0.5032 - val_loss: 0.8354 - val_main_output_loss: 0.7641 - val_aux_output_loss: 0.7136 - val_main_output_acc: 0.4369 - val_aux_output_acc: 0.4757\nEpoch 14/500\n924/924 [==============================] - 0s 217us/step - loss: 0.8174 - main_output_loss: 0.7474 - aux_output_loss: 0.6996 - main_output_acc: 0.4654 - aux_output_acc: 0.4989 - val_loss: 0.8302 - val_main_output_loss: 0.7591 - val_aux_output_loss: 0.7118 - val_main_output_acc: 0.4369 - val_aux_output_acc: 0.4854\nEpoch 15/500\n924/924 [==============================] - 0s 232us/step - loss: 0.8337 - main_output_loss: 0.7640 - aux_output_loss: 0.6966 - main_output_acc: 0.4394 - aux_output_acc: 0.5000 - val_loss: 0.8254 - val_main_output_loss: 0.7544 - val_aux_output_loss: 0.7101 - val_main_output_acc: 0.4175 - val_aux_output_acc: 0.4660\nEpoch 16/500\n924/924 [==============================] - 0s 217us/step - loss: 0.8080 - main_output_loss: 0.7386 - aux_output_loss: 0.6940 - main_output_acc: 0.4805 - aux_output_acc: 0.4946 - val_loss: 0.8209 - val_main_output_loss: 0.7500 - val_aux_output_loss: 0.7089 - val_main_output_acc: 0.3981 - val_aux_output_acc: 0.4563\nEpoch 17/500\n924/924 [==============================] - 0s 210us/step - loss: 0.8053 - main_output_loss: 0.7361 - aux_output_loss: 0.6920 - main_output_acc: 0.4794 - aux_output_acc: 0.4903 - val_loss: 0.8169 - val_main_output_loss: 0.7461 - val_aux_output_loss: 0.7078 - val_main_output_acc: 0.3883 - val_aux_output_acc: 0.4563\nEpoch 18/500\n924/924 [==============================] - 0s 207us/step - loss: 0.7963 - main_output_loss: 0.7273 - aux_output_loss: 0.6902 - main_output_acc: 0.4913 - aux_output_acc: 0.4892 - val_loss: 0.8120 - val_main_output_loss: 0.7414 - val_aux_output_loss: 0.7067 - val_main_output_acc: 0.3883 - val_aux_output_acc: 0.4466\nEpoch 19/500\n924/924 [==============================] - 0s 210us/step - loss: 0.8086 - main_output_loss: 0.7398 - aux_output_loss: 0.6883 - main_output_acc: 0.4675 - aux_output_acc: 0.4903 - val_loss: 0.8079 - val_main_output_loss: 0.7373 - val_aux_output_loss: 0.7059 - val_main_output_acc: 0.3883 - val_aux_output_acc: 0.4466\nEpoch 20/500\n924/924 [==============================] - 0s 214us/step - loss: 0.7977 - main_output_loss: 0.7290 - aux_output_loss: 0.6869 - main_output_acc: 0.4935 - aux_output_acc: 0.4957 - val_loss: 0.8052 - val_main_output_loss: 0.7346 - val_aux_output_loss: 0.7052 - val_main_output_acc: 0.4369 - val_aux_output_acc: 0.4078\nEpoch 21/500\n924/924 [==============================] - 0s 213us/step - loss: 0.7911 - main_output_loss: 0.7226 - aux_output_loss: 0.6855 - main_output_acc: 0.4773 - aux_output_acc: 0.4924 - val_loss: 0.8005 - val_main_output_loss: 0.7301 - val_aux_output_loss: 0.7046 - val_main_output_acc: 0.4466 - val_aux_output_acc: 0.4272\nEpoch 22/500\n924/924 [==============================] - 0s 214us/step - loss: 0.7973 - main_output_loss: 0.7289 - aux_output_loss: 0.6844 - main_output_acc: 0.4740 - aux_output_acc: 0.4968 - val_loss: 0.7962 - val_main_output_loss: 0.7259 - val_aux_output_loss: 0.7038 - val_main_output_acc: 0.4563 - val_aux_output_acc: 0.4272\nEpoch 23/500\n924/924 [==============================] - 0s 222us/step - loss: 0.7961 - main_output_loss: 0.7277 - aux_output_loss: 0.6836 - main_output_acc: 0.4859 - aux_output_acc: 0.5054 - val_loss: 0.7941 - val_main_output_loss: 0.7238 - val_aux_output_loss: 0.7032 - val_main_output_acc: 0.4660 - val_aux_output_acc: 0.4563\nEpoch 24/500\n924/924 [==============================] - 0s 212us/step - loss: 0.7902 - main_output_loss: 0.7220 - aux_output_loss: 0.6827 - main_output_acc: 0.4968 - aux_output_acc: 0.5130 - val_loss: 0.7917 - val_main_output_loss: 0.7214 - val_aux_output_loss: 0.7029 - val_main_output_acc: 0.4854 - val_aux_output_acc: 0.4660\nEpoch 25/500\n924/924 [==============================] - 0s 213us/step - loss: 0.7830 - main_output_loss: 0.7148 - aux_output_loss: 0.6819 - main_output_acc: 0.4881 - aux_output_acc: 0.5325 - val_loss: 0.7875 - val_main_output_loss: 0.7172 - val_aux_output_loss: 0.7022 - val_main_output_acc: 0.4951 - val_aux_output_acc: 0.4757\nEpoch 26/500\n924/924 [==============================] - 0s 215us/step - loss: 0.7922 - main_output_loss: 0.7241 - aux_output_loss: 0.6812 - main_output_acc: 0.4773 - aux_output_acc: 0.5422 - val_loss: 0.7823 - val_main_output_loss: 0.7122 - val_aux_output_loss: 0.7015 - val_main_output_acc: 0.5437 - val_aux_output_acc: 0.4757\nEpoch 27/500\n924/924 [==============================] - 0s 212us/step - loss: 0.7989 - main_output_loss: 0.7309 - aux_output_loss: 0.6804 - main_output_acc: 0.4708 - aux_output_acc: 0.5530 - val_loss: 0.7798 - val_main_output_loss: 0.7097 - val_aux_output_loss: 0.7008 - val_main_output_acc: 0.5243 - val_aux_output_acc: 0.4951\nEpoch 28/500\n924/924 [==============================] - 0s 207us/step - loss: 0.7872 - main_output_loss: 0.7192 - aux_output_loss: 0.6798 - main_output_acc: 0.5195 - aux_output_acc: 0.5498 - val_loss: 0.7767 - val_main_output_loss: 0.7067 - val_aux_output_loss: 0.7001 - val_main_output_acc: 0.5243 - val_aux_output_acc: 0.5049\nEpoch 29/500\n924/924 [==============================] - 0s 217us/step - loss: 0.7794 - main_output_loss: 0.7115 - aux_output_loss: 0.6790 - main_output_acc: 0.5325 - aux_output_acc: 0.5606 - val_loss: 0.7735 - val_main_output_loss: 0.7035 - val_aux_output_loss: 0.6996 - val_main_output_acc: 0.5243 - val_aux_output_acc: 0.5146\nEpoch 30/500\n924/924 [==============================] - 0s 213us/step - loss: 0.7768 - main_output_loss: 0.7090 - aux_output_loss: 0.6783 - main_output_acc: 0.5130 - aux_output_acc: 0.5909 - val_loss: 0.7689 - val_main_output_loss: 0.6991 - val_aux_output_loss: 0.6985 - val_main_output_acc: 0.5728 - val_aux_output_acc: 0.5146\nEpoch 31/500\n924/924 [==============================] - 0s 215us/step - loss: 0.7841 - main_output_loss: 0.7164 - aux_output_loss: 0.6776 - main_output_acc: 0.5043 - aux_output_acc: 0.5909 - val_loss: 0.7659 - val_main_output_loss: 0.6961 - val_aux_output_loss: 0.6978 - val_main_output_acc: 0.5631 - val_aux_output_acc: 0.5049\nEpoch 32/500\n924/924 [==============================] - 0s 212us/step - loss: 0.7713 - main_output_loss: 0.7036 - aux_output_loss: 0.6769 - main_output_acc: 0.5335 - aux_output_acc: 0.5942 - val_loss: 0.7640 - val_main_output_loss: 0.6943 - val_aux_output_loss: 0.6973 - val_main_output_acc: 0.5631 - val_aux_output_acc: 0.5049\nEpoch 33/500\n924/924 [==============================] - 0s 211us/step - loss: 0.7854 - main_output_loss: 0.7177 - aux_output_loss: 0.6764 - main_output_acc: 0.4924 - aux_output_acc: 0.6028 - val_loss: 0.7611 - val_main_output_loss: 0.6915 - val_aux_output_loss: 0.6964 - val_main_output_acc: 0.5631 - val_aux_output_acc: 0.5049\nEpoch 34/500\n924/924 [==============================] - 0s 208us/step - loss: 0.7783 - main_output_loss: 0.7107 - aux_output_loss: 0.6757 - main_output_acc: 0.5087 - aux_output_acc: 0.6082 - val_loss: 0.7598 - val_main_output_loss: 0.6902 - val_aux_output_loss: 0.6961 - val_main_output_acc: 0.5728 - val_aux_output_acc: 0.5146\nEpoch 35/500\n924/924 [==============================] - 0s 210us/step - loss: 0.7791 - main_output_loss: 0.7116 - aux_output_loss: 0.6752 - main_output_acc: 0.4838 - aux_output_acc: 0.6223 - val_loss: 0.7579 - val_main_output_loss: 0.6883 - val_aux_output_loss: 0.6954 - val_main_output_acc: 0.5534 - val_aux_output_acc: 0.5534\nEpoch 36/500\n924/924 [==============================] - 0s 211us/step - loss: 0.7755 - main_output_loss: 0.7080 - aux_output_loss: 0.6746 - main_output_acc: 0.4870 - aux_output_acc: 0.6288 - val_loss: 0.7547 - val_main_output_loss: 0.6852 - val_aux_output_loss: 0.6945 - val_main_output_acc: 0.5534 - val_aux_output_acc: 0.5340\nEpoch 37/500\n924/924 [==============================] - 0s 211us/step - loss: 0.7700 - main_output_loss: 0.7026 - aux_output_loss: 0.6740 - main_output_acc: 0.5238 - aux_output_acc: 0.6245 - val_loss: 0.7537 - val_main_output_loss: 0.6843 - val_aux_output_loss: 0.6940 - val_main_output_acc: 0.5728 - val_aux_output_acc: 0.5437\nEpoch 38/500\n924/924 [==============================] - 0s 214us/step - loss: 0.7833 - main_output_loss: 0.7160 - aux_output_loss: 0.6735 - main_output_acc: 0.4784 - aux_output_acc: 0.6331 - val_loss: 0.7527 - val_main_output_loss: 0.6834 - val_aux_output_loss: 0.6936 - val_main_output_acc: 0.5534 - val_aux_output_acc: 0.5631\nEpoch 39/500\n924/924 [==============================] - 0s 214us/step - loss: 0.7602 - main_output_loss: 0.6929 - aux_output_loss: 0.6729 - main_output_acc: 0.5455 - aux_output_acc: 0.6364 - val_loss: 0.7497 - val_main_output_loss: 0.6804 - val_aux_output_loss: 0.6930 - val_main_output_acc: 0.5631 - val_aux_output_acc: 0.5534\nEpoch 40/500\n924/924 [==============================] - 0s 213us/step - loss: 0.7697 - main_output_loss: 0.7025 - aux_output_loss: 0.6724 - main_output_acc: 0.5195 - aux_output_acc: 0.6407 - val_loss: 0.7485 - val_main_output_loss: 0.6793 - val_aux_output_loss: 0.6919 - val_main_output_acc: 0.5534 - val_aux_output_acc: 0.5631\nEpoch 41/500\n924/924 [==============================] - 0s 215us/step - loss: 0.7566 - main_output_loss: 0.6894 - aux_output_loss: 0.6718 - main_output_acc: 0.5346 - aux_output_acc: 0.6396 - val_loss: 0.7465 - val_main_output_loss: 0.6774 - val_aux_output_loss: 0.6911 - val_main_output_acc: 0.5631 - val_aux_output_acc: 0.5534\nEpoch 42/500\n924/924 [==============================] - 0s 219us/step - loss: 0.7671 - main_output_loss: 0.6999 - aux_output_loss: 0.6712 - main_output_acc: 0.5455 - aux_output_acc: 0.6407 - val_loss: 0.7466 - val_main_output_loss: 0.6776 - val_aux_output_loss: 0.6901 - val_main_output_acc: 0.5728 - val_aux_output_acc: 0.5728\nEpoch 43/500\n924/924 [==============================] - 0s 211us/step - loss: 0.7713 - main_output_loss: 0.7042 - aux_output_loss: 0.6707 - main_output_acc: 0.5184 - aux_output_acc: 0.6494 - val_loss: 0.7443 - val_main_output_loss: 0.6754 - val_aux_output_loss: 0.6897 - val_main_output_acc: 0.5728 - val_aux_output_acc: 0.5728\nEpoch 44/500\n924/924 [==============================] - 0s 212us/step - loss: 0.7683 - main_output_loss: 0.7013 - aux_output_loss: 0.6701 - main_output_acc: 0.4968 - aux_output_acc: 0.6548 - val_loss: 0.7433 - val_main_output_loss: 0.6744 - val_aux_output_loss: 0.6890 - val_main_output_acc: 0.5728 - val_aux_output_acc: 0.5825\nEpoch 45/500\n924/924 [==============================] - 0s 210us/step - loss: 0.7596 - main_output_loss: 0.6927 - aux_output_loss: 0.6696 - main_output_acc: 0.5498 - aux_output_acc: 0.6558 - val_loss: 0.7416 - val_main_output_loss: 0.6728 - val_aux_output_loss: 0.6880 - val_main_output_acc: 0.5922 - val_aux_output_acc: 0.5631\nEpoch 46/500\n924/924 [==============================] - 0s 216us/step - loss: 0.7674 - main_output_loss: 0.7005 - aux_output_loss: 0.6691 - main_output_acc: 0.5087 - aux_output_acc: 0.6364 - val_loss: 0.7425 - val_main_output_loss: 0.6738 - val_aux_output_loss: 0.6877 - val_main_output_acc: 0.5437 - val_aux_output_acc: 0.5728\nEpoch 47/500\n924/924 [==============================] - 0s 214us/step - loss: 0.7606 - main_output_loss: 0.6938 - aux_output_loss: 0.6685 - main_output_acc: 0.5249 - aux_output_acc: 0.6667 - val_loss: 0.7411 - val_main_output_loss: 0.6724 - val_aux_output_loss: 0.6870 - val_main_output_acc: 0.5340 - val_aux_output_acc: 0.5728\nEpoch 48/500\n924/924 [==============================] - 0s 217us/step - loss: 0.7633 - main_output_loss: 0.6965 - aux_output_loss: 0.6680 - main_output_acc: 0.5390 - aux_output_acc: 0.6645 - val_loss: 0.7395 - val_main_output_loss: 0.6708 - val_aux_output_loss: 0.6865 - val_main_output_acc: 0.5534 - val_aux_output_acc: 0.5825\nEpoch 49/500\n924/924 [==============================] - 0s 213us/step - loss: 0.7542 - main_output_loss: 0.6874 - aux_output_loss: 0.6674 - main_output_acc: 0.5444 - aux_output_acc: 0.6688 - val_loss: 0.7393 - val_main_output_loss: 0.6707 - val_aux_output_loss: 0.6858 - val_main_output_acc: 0.5437 - val_aux_output_acc: 0.6214\nEpoch 50/500\n924/924 [==============================] - 0s 210us/step - loss: 0.7623 - main_output_loss: 0.6957 - aux_output_loss: 0.6667 - main_output_acc: 0.5271 - aux_output_acc: 0.6699 - val_loss: 0.7379 - val_main_output_loss: 0.6694 - val_aux_output_loss: 0.6853 - val_main_output_acc: 0.5631 - val_aux_output_acc: 0.6311\nEpoch 51/500\n924/924 [==============================] - 0s 213us/step - loss: 0.7544 - main_output_loss: 0.6878 - aux_output_loss: 0.6662 - main_output_acc: 0.5422 - aux_output_acc: 0.6732 - val_loss: 0.7362 - val_main_output_loss: 0.6677 - val_aux_output_loss: 0.6846 - val_main_output_acc: 0.5728 - val_aux_output_acc: 0.6214\nEpoch 52/500\n924/924 [==============================] - 0s 212us/step - loss: 0.7498 - main_output_loss: 0.6832 - aux_output_loss: 0.6656 - main_output_acc: 0.5628 - aux_output_acc: 0.6775 - val_loss: 0.7339 - val_main_output_loss: 0.6655 - val_aux_output_loss: 0.6836 - val_main_output_acc: 0.5922 - val_aux_output_acc: 0.6408\nEpoch 53/500\n924/924 [==============================] - 0s 211us/step - loss: 0.7531 - main_output_loss: 0.6866 - aux_output_loss: 0.6650 - main_output_acc: 0.5563 - aux_output_acc: 0.6764 - val_loss: 0.7324 - val_main_output_loss: 0.6641 - val_aux_output_loss: 0.6828 - val_main_output_acc: 0.5922 - val_aux_output_acc: 0.6311\nEpoch 54/500\n924/924 [==============================] - 0s 210us/step - loss: 0.7568 - main_output_loss: 0.6903 - aux_output_loss: 0.6644 - main_output_acc: 0.5584 - aux_output_acc: 0.6807 - val_loss: 0.7325 - val_main_output_loss: 0.6643 - val_aux_output_loss: 0.6821 - val_main_output_acc: 0.5922 - val_aux_output_acc: 0.6408\nEpoch 55/500\n924/924 [==============================] - 0s 212us/step - loss: 0.7554 - main_output_loss: 0.6890 - aux_output_loss: 0.6638 - main_output_acc: 0.5400 - aux_output_acc: 0.6818 - val_loss: 0.7323 - val_main_output_loss: 0.6642 - val_aux_output_loss: 0.6813 - val_main_output_acc: 0.5728 - val_aux_output_acc: 0.6311\nEpoch 56/500\n924/924 [==============================] - 0s 210us/step - loss: 0.7495 - main_output_loss: 0.6832 - aux_output_loss: 0.6633 - main_output_acc: 0.5649 - aux_output_acc: 0.6829 - val_loss: 0.7309 - val_main_output_loss: 0.6629 - val_aux_output_loss: 0.6808 - val_main_output_acc: 0.5728 - val_aux_output_acc: 0.6311\nEpoch 57/500\n924/924 [==============================] - 0s 211us/step - loss: 0.7555 - main_output_loss: 0.6892 - aux_output_loss: 0.6628 - main_output_acc: 0.5465 - aux_output_acc: 0.6786 - val_loss: 0.7290 - val_main_output_loss: 0.6610 - val_aux_output_loss: 0.6799 - val_main_output_acc: 0.5922 - val_aux_output_acc: 0.6311\nEpoch 58/500\n924/924 [==============================] - 0s 219us/step - loss: 0.7602 - main_output_loss: 0.6939 - aux_output_loss: 0.6621 - main_output_acc: 0.5281 - aux_output_acc: 0.6797 - val_loss: 0.7277 - val_main_output_loss: 0.6598 - val_aux_output_loss: 0.6791 - val_main_output_acc: 0.6019 - val_aux_output_acc: 0.6311\nEpoch 59/500\n924/924 [==============================] - 0s 218us/step - loss: 0.7521 - main_output_loss: 0.6860 - aux_output_loss: 0.6616 - main_output_acc: 0.5476 - aux_output_acc: 0.6797 - val_loss: 0.7280 - val_main_output_loss: 0.6602 - val_aux_output_loss: 0.6782 - val_main_output_acc: 0.5825 - val_aux_output_acc: 0.6311\nEpoch 60/500\n924/924 [==============================] - 0s 208us/step - loss: 0.7683 - main_output_loss: 0.7023 - aux_output_loss: 0.6609 - main_output_acc: 0.5043 - aux_output_acc: 0.6807 - val_loss: 0.7261 - val_main_output_loss: 0.6584 - val_aux_output_loss: 0.6773 - val_main_output_acc: 0.6019 - val_aux_output_acc: 0.6311\nEpoch 61/500\n924/924 [==============================] - 0s 211us/step - loss: 0.7526 - main_output_loss: 0.6866 - aux_output_loss: 0.6603 - main_output_acc: 0.5433 - aux_output_acc: 0.6840 - val_loss: 0.7263 - val_main_output_loss: 0.6587 - val_aux_output_loss: 0.6767 - val_main_output_acc: 0.5825 - val_aux_output_acc: 0.6408\nEpoch 62/500\n924/924 [==============================] - 0s 214us/step - loss: 0.7586 - main_output_loss: 0.6926 - aux_output_loss: 0.6597 - main_output_acc: 0.5422 - aux_output_acc: 0.6851 - val_loss: 0.7253 - val_main_output_loss: 0.6577 - val_aux_output_loss: 0.6760 - val_main_output_acc: 0.5825 - val_aux_output_acc: 0.6408\nEpoch 63/500\n924/924 [==============================] - 0s 212us/step - loss: 0.7532 - main_output_loss: 0.6873 - aux_output_loss: 0.6591 - main_output_acc: 0.5617 - aux_output_acc: 0.6851 - val_loss: 0.7239 - val_main_output_loss: 0.6564 - val_aux_output_loss: 0.6749 - val_main_output_acc: 0.6019 - val_aux_output_acc: 0.6311\nEpoch 64/500\n924/924 [==============================] - 0s 214us/step - loss: 0.7488 - main_output_loss: 0.6830 - aux_output_loss: 0.6586 - main_output_acc: 0.5444 - aux_output_acc: 0.6861 - val_loss: 0.7242 - val_main_output_loss: 0.6568 - val_aux_output_loss: 0.6743 - val_main_output_acc: 0.5922 - val_aux_output_acc: 0.6311\nEpoch 65/500\n924/924 [==============================] - 0s 209us/step - loss: 0.7559 - main_output_loss: 0.6901 - aux_output_loss: 0.6581 - main_output_acc: 0.5433 - aux_output_acc: 0.6883 - val_loss: 0.7248 - val_main_output_loss: 0.6574 - val_aux_output_loss: 0.6738 - val_main_output_acc: 0.5728 - val_aux_output_acc: 0.6408\nEpoch 66/500\n924/924 [==============================] - 0s 214us/step - loss: 0.7488 - main_output_loss: 0.6831 - aux_output_loss: 0.6576 - main_output_acc: 0.5584 - aux_output_acc: 0.6861 - val_loss: 0.7240 - val_main_output_loss: 0.6567 - val_aux_output_loss: 0.6731 - val_main_output_acc: 0.5825 - val_aux_output_acc: 0.6408\nEpoch 67/500\n924/924 [==============================] - 0s 220us/step - loss: 0.7392 - main_output_loss: 0.6735 - aux_output_loss: 0.6570 - main_output_acc: 0.5768 - aux_output_acc: 0.6829 - val_loss: 0.7225 - val_main_output_loss: 0.6552 - val_aux_output_loss: 0.6724 - val_main_output_acc: 0.6019 - val_aux_output_acc: 0.6408\nEpoch 68/500\n924/924 [==============================] - 0s 207us/step - loss: 0.7438 - main_output_loss: 0.6781 - aux_output_loss: 0.6565 - main_output_acc: 0.5530 - aux_output_acc: 0.6872 - val_loss: 0.7209 - val_main_output_loss: 0.6537 - val_aux_output_loss: 0.6715 - val_main_output_acc: 0.6019 - val_aux_output_acc: 0.6505\nEpoch 69/500\n924/924 [==============================] - 0s 208us/step - loss: 0.7388 - main_output_loss: 0.6732 - aux_output_loss: 0.6559 - main_output_acc: 0.5898 - aux_output_acc: 0.6926 - val_loss: 0.7208 - val_main_output_loss: 0.6536 - val_aux_output_loss: 0.6711 - val_main_output_acc: 0.6019 - val_aux_output_acc: 0.6408\nEpoch 70/500\n924/924 [==============================] - 0s 214us/step - loss: 0.7404 - main_output_loss: 0.6749 - aux_output_loss: 0.6554 - main_output_acc: 0.5790 - aux_output_acc: 0.6872 - val_loss: 0.7197 - val_main_output_loss: 0.6527 - val_aux_output_loss: 0.6703 - val_main_output_acc: 0.6117 - val_aux_output_acc: 0.6505\nEpoch 71/500\n924/924 [==============================] - 0s 215us/step - loss: 0.7590 - main_output_loss: 0.6935 - aux_output_loss: 0.6548 - main_output_acc: 0.5400 - aux_output_acc: 0.6926 - val_loss: 0.7196 - val_main_output_loss: 0.6526 - val_aux_output_loss: 0.6699 - val_main_output_acc: 0.6117 - val_aux_output_acc: 0.6505\nEpoch 72/500\n924/924 [==============================] - 0s 214us/step - loss: 0.7453 - main_output_loss: 0.6798 - aux_output_loss: 0.6544 - main_output_acc: 0.5433 - aux_output_acc: 0.6916 - val_loss: 0.7182 - val_main_output_loss: 0.6513 - val_aux_output_loss: 0.6690 - val_main_output_acc: 0.5922 - val_aux_output_acc: 0.6602\nEpoch 73/500\n924/924 [==============================] - 0s 215us/step - loss: 0.7492 - main_output_loss: 0.6838 - aux_output_loss: 0.6538 - main_output_acc: 0.5779 - aux_output_acc: 0.6959 - val_loss: 0.7182 - val_main_output_loss: 0.6514 - val_aux_output_loss: 0.6685 - val_main_output_acc: 0.6117 - val_aux_output_acc: 0.6505\nEpoch 74/500\n924/924 [==============================] - 0s 215us/step - loss: 0.7603 - main_output_loss: 0.6950 - aux_output_loss: 0.6534 - main_output_acc: 0.5227 - aux_output_acc: 0.6905 - val_loss: 0.7177 - val_main_output_loss: 0.6509 - val_aux_output_loss: 0.6677 - val_main_output_acc: 0.6019 - val_aux_output_acc: 0.6505\nEpoch 75/500\n924/924 [==============================] - 0s 225us/step - loss: 0.7524 - main_output_loss: 0.6871 - aux_output_loss: 0.6530 - main_output_acc: 0.5563 - aux_output_acc: 0.6970 - val_loss: 0.7168 - val_main_output_loss: 0.6501 - val_aux_output_loss: 0.6670 - val_main_output_acc: 0.6019 - val_aux_output_acc: 0.6505\nEpoch 76/500\n924/924 [==============================] - 0s 213us/step - loss: 0.7385 - main_output_loss: 0.6733 - aux_output_loss: 0.6526 - main_output_acc: 0.5812 - aux_output_acc: 0.6959 - val_loss: 0.7176 - val_main_output_loss: 0.6509 - val_aux_output_loss: 0.6665 - val_main_output_acc: 0.6019 - val_aux_output_acc: 0.6505\nEpoch 77/500\n924/924 [==============================] - 0s 212us/step - loss: 0.7498 - main_output_loss: 0.6846 - aux_output_loss: 0.6520 - main_output_acc: 0.5574 - aux_output_acc: 0.6948 - val_loss: 0.7180 - val_main_output_loss: 0.6514 - val_aux_output_loss: 0.6659 - val_main_output_acc: 0.5922 - val_aux_output_acc: 0.6505\nEpoch 78/500\n924/924 [==============================] - 0s 212us/step - loss: 0.7356 - main_output_loss: 0.6704 - aux_output_loss: 0.6514 - main_output_acc: 0.5985 - aux_output_acc: 0.6970 - val_loss: 0.7166 - val_main_output_loss: 0.6501 - val_aux_output_loss: 0.6652 - val_main_output_acc: 0.6214 - val_aux_output_acc: 0.6505\nEpoch 79/500\n924/924 [==============================] - 0s 206us/step - loss: 0.7491 - main_output_loss: 0.6840 - aux_output_loss: 0.6509 - main_output_acc: 0.5400 - aux_output_acc: 0.7002 - val_loss: 0.7159 - val_main_output_loss: 0.6495 - val_aux_output_loss: 0.6645 - val_main_output_acc: 0.6117 - val_aux_output_acc: 0.6505\nEpoch 80/500\n924/924 [==============================] - 0s 217us/step - loss: 0.7531 - main_output_loss: 0.6881 - aux_output_loss: 0.6503 - main_output_acc: 0.5487 - aux_output_acc: 0.6991 - val_loss: 0.7165 - val_main_output_loss: 0.6502 - val_aux_output_loss: 0.6637 - val_main_output_acc: 0.6214 - val_aux_output_acc: 0.6505\nEpoch 81/500\n924/924 [==============================] - 0s 210us/step - loss: 0.7510 - main_output_loss: 0.6860 - aux_output_loss: 0.6499 - main_output_acc: 0.5346 - aux_output_acc: 0.7013 - val_loss: 0.7174 - val_main_output_loss: 0.6510 - val_aux_output_loss: 0.6634 - val_main_output_acc: 0.5825 - val_aux_output_acc: 0.6408\nEpoch 82/500\n924/924 [==============================] - 0s 211us/step - loss: 0.7391 - main_output_loss: 0.6742 - aux_output_loss: 0.6493 - main_output_acc: 0.5639 - aux_output_acc: 0.7067 - val_loss: 0.7149 - val_main_output_loss: 0.6486 - val_aux_output_loss: 0.6625 - val_main_output_acc: 0.6408 - val_aux_output_acc: 0.6602\nEpoch 83/500\n924/924 [==============================] - 0s 205us/step - loss: 0.7467 - main_output_loss: 0.6819 - aux_output_loss: 0.6488 - main_output_acc: 0.5444 - aux_output_acc: 0.7078 - val_loss: 0.7148 - val_main_output_loss: 0.6486 - val_aux_output_loss: 0.6617 - val_main_output_acc: 0.6117 - val_aux_output_acc: 0.6602\nEpoch 84/500\n924/924 [==============================] - 0s 219us/step - loss: 0.7456 - main_output_loss: 0.6808 - aux_output_loss: 0.6483 - main_output_acc: 0.5476 - aux_output_acc: 0.7024 - val_loss: 0.7138 - val_main_output_loss: 0.6477 - val_aux_output_loss: 0.6610 - val_main_output_acc: 0.6214 - val_aux_output_acc: 0.6602\nEpoch 85/500\n924/924 [==============================] - 0s 210us/step - loss: 0.7460 - main_output_loss: 0.6812 - aux_output_loss: 0.6479 - main_output_acc: 0.5487 - aux_output_acc: 0.7035 - val_loss: 0.7139 - val_main_output_loss: 0.6479 - val_aux_output_loss: 0.6603 - val_main_output_acc: 0.6214 - val_aux_output_acc: 0.6602\nEpoch 86/500\n924/924 [==============================] - 0s 213us/step - loss: 0.7401 - main_output_loss: 0.6753 - aux_output_loss: 0.6474 - main_output_acc: 0.5628 - aux_output_acc: 0.7089 - val_loss: 0.7144 - val_main_output_loss: 0.6484 - val_aux_output_loss: 0.6599 - val_main_output_acc: 0.6214 - val_aux_output_acc: 0.6699\nEpoch 87/500\n924/924 [==============================] - 0s 217us/step - loss: 0.7463 - main_output_loss: 0.6816 - aux_output_loss: 0.6470 - main_output_acc: 0.5465 - aux_output_acc: 0.7067 - val_loss: 0.7135 - val_main_output_loss: 0.6475 - val_aux_output_loss: 0.6595 - val_main_output_acc: 0.6117 - val_aux_output_acc: 0.6796\nEpoch 88/500\n924/924 [==============================] - 0s 210us/step - loss: 0.7447 - main_output_loss: 0.6800 - aux_output_loss: 0.6466 - main_output_acc: 0.5628 - aux_output_acc: 0.7110 - val_loss: 0.7128 - val_main_output_loss: 0.6470 - val_aux_output_loss: 0.6587 - val_main_output_acc: 0.6214 - val_aux_output_acc: 0.6796\nEpoch 89/500\n924/924 [==============================] - 0s 210us/step - loss: 0.7423 - main_output_loss: 0.6777 - aux_output_loss: 0.6461 - main_output_acc: 0.5606 - aux_output_acc: 0.7089 - val_loss: 0.7121 - val_main_output_loss: 0.6462 - val_aux_output_loss: 0.6584 - val_main_output_acc: 0.6214 - val_aux_output_acc: 0.6796\nEpoch 90/500\n924/924 [==============================] - 0s 209us/step - loss: 0.7478 - main_output_loss: 0.6833 - aux_output_loss: 0.6455 - main_output_acc: 0.5444 - aux_output_acc: 0.7154 - val_loss: 0.7113 - val_main_output_loss: 0.6455 - val_aux_output_loss: 0.6577 - val_main_output_acc: 0.6214 - val_aux_output_acc: 0.6796\nEpoch 91/500\n924/924 [==============================] - 0s 213us/step - loss: 0.7418 - main_output_loss: 0.6773 - aux_output_loss: 0.6450 - main_output_acc: 0.5649 - aux_output_acc: 0.7110 - val_loss: 0.7108 - val_main_output_loss: 0.6451 - val_aux_output_loss: 0.6571 - val_main_output_acc: 0.6214 - val_aux_output_acc: 0.6699\nEpoch 92/500\n924/924 [==============================] - 0s 209us/step - loss: 0.7438 - main_output_loss: 0.6793 - aux_output_loss: 0.6446 - main_output_acc: 0.5530 - aux_output_acc: 0.7089 - val_loss: 0.7108 - val_main_output_loss: 0.6452 - val_aux_output_loss: 0.6566 - val_main_output_acc: 0.6117 - val_aux_output_acc: 0.6699\nEpoch 93/500\n924/924 [==============================] - 0s 210us/step - loss: 0.7370 - main_output_loss: 0.6726 - aux_output_loss: 0.6440 - main_output_acc: 0.5693 - aux_output_acc: 0.7143 - val_loss: 0.7118 - val_main_output_loss: 0.6462 - val_aux_output_loss: 0.6560 - val_main_output_acc: 0.6117 - val_aux_output_acc: 0.6699\nEpoch 94/500\n924/924 [==============================] - 0s 216us/step - loss: 0.7468 - main_output_loss: 0.6824 - aux_output_loss: 0.6436 - main_output_acc: 0.5628 - aux_output_acc: 0.7121 - val_loss: 0.7119 - val_main_output_loss: 0.6463 - val_aux_output_loss: 0.6558 - val_main_output_acc: 0.5825 - val_aux_output_acc: 0.6893\nEpoch 95/500\n924/924 [==============================] - 0s 212us/step - loss: 0.7354 - main_output_loss: 0.6711 - aux_output_loss: 0.6432 - main_output_acc: 0.5855 - aux_output_acc: 0.7100 - val_loss: 0.7130 - val_main_output_loss: 0.6474 - val_aux_output_loss: 0.6553 - val_main_output_acc: 0.5922 - val_aux_output_acc: 0.7087\nEpoch 96/500\n924/924 [==============================] - 0s 210us/step - loss: 0.7445 - main_output_loss: 0.6802 - aux_output_loss: 0.6428 - main_output_acc: 0.5617 - aux_output_acc: 0.7078 - val_loss: 0.7129 - val_main_output_loss: 0.6474 - val_aux_output_loss: 0.6553 - val_main_output_acc: 0.5922 - val_aux_output_acc: 0.7087\nEpoch 97/500\n924/924 [==============================] - 0s 214us/step - loss: 0.7414 - main_output_loss: 0.6772 - aux_output_loss: 0.6423 - main_output_acc: 0.5541 - aux_output_acc: 0.7078 - val_loss: 0.7121 - val_main_output_loss: 0.6466 - val_aux_output_loss: 0.6550 - val_main_output_acc: 0.6019 - val_aux_output_acc: 0.7087\nEpoch 98/500\n924/924 [==============================] - 0s 215us/step - loss: 0.7396 - main_output_loss: 0.6754 - aux_output_loss: 0.6419 - main_output_acc: 0.5541 - aux_output_acc: 0.7013 - val_loss: 0.7091 - val_main_output_loss: 0.6436 - val_aux_output_loss: 0.6541 - val_main_output_acc: 0.6019 - val_aux_output_acc: 0.7087\nEpoch 99/500\n924/924 [==============================] - 0s 207us/step - loss: 0.7347 - main_output_loss: 0.6705 - aux_output_loss: 0.6414 - main_output_acc: 0.5758 - aux_output_acc: 0.7067 - val_loss: 0.7081 - val_main_output_loss: 0.6427 - val_aux_output_loss: 0.6535 - val_main_output_acc: 0.6019 - val_aux_output_acc: 0.7087\nEpoch 100/500\n924/924 [==============================] - 0s 208us/step - loss: 0.7336 - main_output_loss: 0.6695 - aux_output_loss: 0.6409 - main_output_acc: 0.5931 - aux_output_acc: 0.7035 - val_loss: 0.7068 - val_main_output_loss: 0.6415 - val_aux_output_loss: 0.6530 - val_main_output_acc: 0.6214 - val_aux_output_acc: 0.7087\nEpoch 101/500\n924/924 [==============================] - 0s 208us/step - loss: 0.7356 - main_output_loss: 0.6715 - aux_output_loss: 0.6404 - main_output_acc: 0.5671 - aux_output_acc: 0.7056 - val_loss: 0.7058 - val_main_output_loss: 0.6405 - val_aux_output_loss: 0.6522 - val_main_output_acc: 0.6117 - val_aux_output_acc: 0.7184\nEpoch 102/500\n924/924 [==============================] - 0s 212us/step - loss: 0.7336 - main_output_loss: 0.6696 - aux_output_loss: 0.6400 - main_output_acc: 0.5584 - aux_output_acc: 0.7089 - val_loss: 0.7050 - val_main_output_loss: 0.6398 - val_aux_output_loss: 0.6516 - val_main_output_acc: 0.6311 - val_aux_output_acc: 0.7087\nEpoch 103/500\n924/924 [==============================] - 0s 223us/step - loss: 0.7380 - main_output_loss: 0.6740 - aux_output_loss: 0.6397 - main_output_acc: 0.5671 - aux_output_acc: 0.7056 - val_loss: 0.7057 - val_main_output_loss: 0.6405 - val_aux_output_loss: 0.6512 - val_main_output_acc: 0.6214 - val_aux_output_acc: 0.7184\nEpoch 104/500\n924/924 [==============================] - 0s 212us/step - loss: 0.7340 - main_output_loss: 0.6701 - aux_output_loss: 0.6392 - main_output_acc: 0.5909 - aux_output_acc: 0.7078 - val_loss: 0.7059 - val_main_output_loss: 0.6408 - val_aux_output_loss: 0.6511 - val_main_output_acc: 0.6311 - val_aux_output_acc: 0.7184\nEpoch 105/500\n924/924 [==============================] - 0s 218us/step - loss: 0.7357 - main_output_loss: 0.6718 - aux_output_loss: 0.6388 - main_output_acc: 0.5649 - aux_output_acc: 0.7100 - val_loss: 0.7033 - val_main_output_loss: 0.6383 - val_aux_output_loss: 0.6503 - val_main_output_acc: 0.6408 - val_aux_output_acc: 0.7184\nEpoch 106/500\n924/924 [==============================] - 0s 213us/step - loss: 0.7309 - main_output_loss: 0.6671 - aux_output_loss: 0.6383 - main_output_acc: 0.5768 - aux_output_acc: 0.7100 - val_loss: 0.7032 - val_main_output_loss: 0.6382 - val_aux_output_loss: 0.6500 - val_main_output_acc: 0.6311 - val_aux_output_acc: 0.7184\nEpoch 107/500\n924/924 [==============================] - 0s 220us/step - loss: 0.7444 - main_output_loss: 0.6806 - aux_output_loss: 0.6379 - main_output_acc: 0.5584 - aux_output_acc: 0.7089 - val_loss: 0.7023 - val_main_output_loss: 0.6374 - val_aux_output_loss: 0.6491 - val_main_output_acc: 0.6505 - val_aux_output_acc: 0.7184\nEpoch 108/500\n924/924 [==============================] - 0s 215us/step - loss: 0.7365 - main_output_loss: 0.6727 - aux_output_loss: 0.6375 - main_output_acc: 0.5703 - aux_output_acc: 0.7067 - val_loss: 0.7026 - val_main_output_loss: 0.6377 - val_aux_output_loss: 0.6488 - val_main_output_acc: 0.6505 - val_aux_output_acc: 0.7184\nEpoch 109/500\n924/924 [==============================] - 0s 215us/step - loss: 0.7268 - main_output_loss: 0.6631 - aux_output_loss: 0.6370 - main_output_acc: 0.5671 - aux_output_acc: 0.7110 - val_loss: 0.7014 - val_main_output_loss: 0.6366 - val_aux_output_loss: 0.6482 - val_main_output_acc: 0.6602 - val_aux_output_acc: 0.7184\nEpoch 110/500\n924/924 [==============================] - 0s 215us/step - loss: 0.7306 - main_output_loss: 0.6669 - aux_output_loss: 0.6366 - main_output_acc: 0.5812 - aux_output_acc: 0.7078 - val_loss: 0.7016 - val_main_output_loss: 0.6368 - val_aux_output_loss: 0.6479 - val_main_output_acc: 0.6214 - val_aux_output_acc: 0.7184\nEpoch 111/500\n924/924 [==============================] - 0s 208us/step - loss: 0.7373 - main_output_loss: 0.6736 - aux_output_loss: 0.6361 - main_output_acc: 0.5703 - aux_output_acc: 0.7089 - val_loss: 0.7009 - val_main_output_loss: 0.6361 - val_aux_output_loss: 0.6474 - val_main_output_acc: 0.6505 - val_aux_output_acc: 0.7184\nEpoch 112/500\n924/924 [==============================] - 0s 215us/step - loss: 0.7320 - main_output_loss: 0.6685 - aux_output_loss: 0.6358 - main_output_acc: 0.5877 - aux_output_acc: 0.7056 - val_loss: 0.7023 - val_main_output_loss: 0.6376 - val_aux_output_loss: 0.6474 - val_main_output_acc: 0.6214 - val_aux_output_acc: 0.7184\nEpoch 113/500\n924/924 [==============================] - 0s 216us/step - loss: 0.7324 - main_output_loss: 0.6689 - aux_output_loss: 0.6354 - main_output_acc: 0.5747 - aux_output_acc: 0.7110 - val_loss: 0.7020 - val_main_output_loss: 0.6373 - val_aux_output_loss: 0.6468 - val_main_output_acc: 0.6214 - val_aux_output_acc: 0.7087\nEpoch 114/500\n924/924 [==============================] - 0s 211us/step - loss: 0.7312 - main_output_loss: 0.6677 - aux_output_loss: 0.6350 - main_output_acc: 0.5963 - aux_output_acc: 0.7110 - val_loss: 0.7014 - val_main_output_loss: 0.6368 - val_aux_output_loss: 0.6462 - val_main_output_acc: 0.6117 - val_aux_output_acc: 0.7184\nEpoch 115/500\n924/924 [==============================] - 0s 213us/step - loss: 0.7325 - main_output_loss: 0.6690 - aux_output_loss: 0.6345 - main_output_acc: 0.6006 - aux_output_acc: 0.7089 - val_loss: 0.7002 - val_main_output_loss: 0.6357 - val_aux_output_loss: 0.6453 - val_main_output_acc: 0.6602 - val_aux_output_acc: 0.7087\nEpoch 116/500\n924/924 [==============================] - 0s 214us/step - loss: 0.7336 - main_output_loss: 0.6702 - aux_output_loss: 0.6342 - main_output_acc: 0.5595 - aux_output_acc: 0.7100 - val_loss: 0.6986 - val_main_output_loss: 0.6341 - val_aux_output_loss: 0.6449 - val_main_output_acc: 0.6505 - val_aux_output_acc: 0.7087\nEpoch 117/500\n924/924 [==============================] - 0s 211us/step - loss: 0.7367 - main_output_loss: 0.6733 - aux_output_loss: 0.6338 - main_output_acc: 0.5758 - aux_output_acc: 0.7110 - val_loss: 0.6991 - val_main_output_loss: 0.6347 - val_aux_output_loss: 0.6443 - val_main_output_acc: 0.6505 - val_aux_output_acc: 0.7087\nEpoch 118/500\n924/924 [==============================] - 0s 207us/step - loss: 0.7303 - main_output_loss: 0.6669 - aux_output_loss: 0.6334 - main_output_acc: 0.5768 - aux_output_acc: 0.7110 - val_loss: 0.6985 - val_main_output_loss: 0.6341 - val_aux_output_loss: 0.6439 - val_main_output_acc: 0.6602 - val_aux_output_acc: 0.7087\nEpoch 119/500\n924/924 [==============================] - 0s 210us/step - loss: 0.7245 - main_output_loss: 0.6612 - aux_output_loss: 0.6329 - main_output_acc: 0.5812 - aux_output_acc: 0.7089 - val_loss: 0.6990 - val_main_output_loss: 0.6346 - val_aux_output_loss: 0.6437 - val_main_output_acc: 0.6408 - val_aux_output_acc: 0.6990\nEpoch 120/500\n924/924 [==============================] - 0s 209us/step - loss: 0.7396 - main_output_loss: 0.6764 - aux_output_loss: 0.6325 - main_output_acc: 0.5779 - aux_output_acc: 0.7110 - val_loss: 0.6994 - val_main_output_loss: 0.6350 - val_aux_output_loss: 0.6434 - val_main_output_acc: 0.6117 - val_aux_output_acc: 0.6990\nEpoch 121/500\n924/924 [==============================] - 0s 213us/step - loss: 0.7327 - main_output_loss: 0.6695 - aux_output_loss: 0.6321 - main_output_acc: 0.5671 - aux_output_acc: 0.7078 - val_loss: 0.6992 - val_main_output_loss: 0.6349 - val_aux_output_loss: 0.6430 - val_main_output_acc: 0.6408 - val_aux_output_acc: 0.6990\nEpoch 122/500\n924/924 [==============================] - 0s 213us/step - loss: 0.7283 - main_output_loss: 0.6651 - aux_output_loss: 0.6317 - main_output_acc: 0.5855 - aux_output_acc: 0.7110 - val_loss: 0.6976 - val_main_output_loss: 0.6333 - val_aux_output_loss: 0.6422 - val_main_output_acc: 0.6505 - val_aux_output_acc: 0.7087\nEpoch 123/500\n924/924 [==============================] - 0s 214us/step - loss: 0.7264 - main_output_loss: 0.6632 - aux_output_loss: 0.6312 - main_output_acc: 0.5714 - aux_output_acc: 0.7121 - val_loss: 0.6981 - val_main_output_loss: 0.6340 - val_aux_output_loss: 0.6417 - val_main_output_acc: 0.6505 - val_aux_output_acc: 0.7087\nEpoch 124/500\n924/924 [==============================] - 0s 212us/step - loss: 0.7287 - main_output_loss: 0.6657 - aux_output_loss: 0.6309 - main_output_acc: 0.5931 - aux_output_acc: 0.7110 - val_loss: 0.6971 - val_main_output_loss: 0.6330 - val_aux_output_loss: 0.6410 - val_main_output_acc: 0.6311 - val_aux_output_acc: 0.7184\nEpoch 125/500\n924/924 [==============================] - 0s 218us/step - loss: 0.7309 - main_output_loss: 0.6679 - aux_output_loss: 0.6305 - main_output_acc: 0.5660 - aux_output_acc: 0.7132 - val_loss: 0.6953 - val_main_output_loss: 0.6313 - val_aux_output_loss: 0.6403 - val_main_output_acc: 0.6408 - val_aux_output_acc: 0.7184\nEpoch 126/500\n924/924 [==============================] - 0s 209us/step - loss: 0.7305 - main_output_loss: 0.6675 - aux_output_loss: 0.6301 - main_output_acc: 0.5823 - aux_output_acc: 0.7110 - val_loss: 0.6958 - val_main_output_loss: 0.6317 - val_aux_output_loss: 0.6403 - val_main_output_acc: 0.6408 - val_aux_output_acc: 0.7087\nEpoch 127/500\n924/924 [==============================] - 0s 209us/step - loss: 0.7293 - main_output_loss: 0.6663 - aux_output_loss: 0.6297 - main_output_acc: 0.5812 - aux_output_acc: 0.7132 - val_loss: 0.6967 - val_main_output_loss: 0.6327 - val_aux_output_loss: 0.6400 - val_main_output_acc: 0.6602 - val_aux_output_acc: 0.6990\nEpoch 128/500\n924/924 [==============================] - 0s 207us/step - loss: 0.7296 - main_output_loss: 0.6666 - aux_output_loss: 0.6294 - main_output_acc: 0.5877 - aux_output_acc: 0.7154 - val_loss: 0.6969 - val_main_output_loss: 0.6329 - val_aux_output_loss: 0.6398 - val_main_output_acc: 0.6311 - val_aux_output_acc: 0.6990\nEpoch 129/500\n924/924 [==============================] - 0s 222us/step - loss: 0.7290 - main_output_loss: 0.6661 - aux_output_loss: 0.6290 - main_output_acc: 0.5833 - aux_output_acc: 0.7132 - val_loss: 0.6952 - val_main_output_loss: 0.6312 - val_aux_output_loss: 0.6394 - val_main_output_acc: 0.6602 - val_aux_output_acc: 0.6990\nEpoch 130/500\n924/924 [==============================] - 0s 211us/step - loss: 0.7158 - main_output_loss: 0.6529 - aux_output_loss: 0.6287 - main_output_acc: 0.6028 - aux_output_acc: 0.7143 - val_loss: 0.6936 - val_main_output_loss: 0.6297 - val_aux_output_loss: 0.6388 - val_main_output_acc: 0.6505 - val_aux_output_acc: 0.6990\nEpoch 131/500\n924/924 [==============================] - 0s 207us/step - loss: 0.7208 - main_output_loss: 0.6580 - aux_output_loss: 0.6282 - main_output_acc: 0.6039 - aux_output_acc: 0.7132 - val_loss: 0.6928 - val_main_output_loss: 0.6289 - val_aux_output_loss: 0.6384 - val_main_output_acc: 0.6408 - val_aux_output_acc: 0.6990\nEpoch 132/500\n924/924 [==============================] - 0s 220us/step - loss: 0.7311 - main_output_loss: 0.6683 - aux_output_loss: 0.6279 - main_output_acc: 0.5887 - aux_output_acc: 0.7121 - val_loss: 0.6941 - val_main_output_loss: 0.6303 - val_aux_output_loss: 0.6384 - val_main_output_acc: 0.6311 - val_aux_output_acc: 0.6990\nEpoch 133/500\n924/924 [==============================] - 0s 207us/step - loss: 0.7287 - main_output_loss: 0.6659 - aux_output_loss: 0.6275 - main_output_acc: 0.5844 - aux_output_acc: 0.7121 - val_loss: 0.6926 - val_main_output_loss: 0.6288 - val_aux_output_loss: 0.6379 - val_main_output_acc: 0.6408 - val_aux_output_acc: 0.6990\nEpoch 134/500\n924/924 [==============================] - 0s 213us/step - loss: 0.7358 - main_output_loss: 0.6731 - aux_output_loss: 0.6272 - main_output_acc: 0.5693 - aux_output_acc: 0.7154 - val_loss: 0.6940 - val_main_output_loss: 0.6303 - val_aux_output_loss: 0.6374 - val_main_output_acc: 0.6602 - val_aux_output_acc: 0.6990\nEpoch 135/500\n924/924 [==============================] - 0s 213us/step - loss: 0.7179 - main_output_loss: 0.6553 - aux_output_loss: 0.6267 - main_output_acc: 0.6082 - aux_output_acc: 0.7121 - val_loss: 0.6943 - val_main_output_loss: 0.6306 - val_aux_output_loss: 0.6373 - val_main_output_acc: 0.6505 - val_aux_output_acc: 0.7184\nEpoch 136/500\n924/924 [==============================] - 0s 210us/step - loss: 0.7230 - main_output_loss: 0.6604 - aux_output_loss: 0.6265 - main_output_acc: 0.5898 - aux_output_acc: 0.7143 - val_loss: 0.6923 - val_main_output_loss: 0.6286 - val_aux_output_loss: 0.6371 - val_main_output_acc: 0.6602 - val_aux_output_acc: 0.7087\nEpoch 137/500\n924/924 [==============================] - 0s 209us/step - loss: 0.7224 - main_output_loss: 0.6598 - aux_output_loss: 0.6260 - main_output_acc: 0.5942 - aux_output_acc: 0.7143 - val_loss: 0.6896 - val_main_output_loss: 0.6260 - val_aux_output_loss: 0.6360 - val_main_output_acc: 0.6505 - val_aux_output_acc: 0.6990\nEpoch 138/500\n924/924 [==============================] - 0s 207us/step - loss: 0.7269 - main_output_loss: 0.6644 - aux_output_loss: 0.6256 - main_output_acc: 0.5790 - aux_output_acc: 0.7132 - val_loss: 0.6908 - val_main_output_loss: 0.6272 - val_aux_output_loss: 0.6357 - val_main_output_acc: 0.6408 - val_aux_output_acc: 0.6990\nEpoch 139/500\n924/924 [==============================] - 0s 210us/step - loss: 0.7154 - main_output_loss: 0.6529 - aux_output_loss: 0.6252 - main_output_acc: 0.5974 - aux_output_acc: 0.7143 - val_loss: 0.6897 - val_main_output_loss: 0.6261 - val_aux_output_loss: 0.6351 - val_main_output_acc: 0.6505 - val_aux_output_acc: 0.6990\nEpoch 140/500\n924/924 [==============================] - 0s 209us/step - loss: 0.7300 - main_output_loss: 0.6675 - aux_output_loss: 0.6249 - main_output_acc: 0.5931 - aux_output_acc: 0.7165 - val_loss: 0.6881 - val_main_output_loss: 0.6246 - val_aux_output_loss: 0.6344 - val_main_output_acc: 0.6408 - val_aux_output_acc: 0.7184\nEpoch 141/500\n924/924 [==============================] - 0s 224us/step - loss: 0.7174 - main_output_loss: 0.6550 - aux_output_loss: 0.6246 - main_output_acc: 0.5996 - aux_output_acc: 0.7165 - val_loss: 0.6884 - val_main_output_loss: 0.6250 - val_aux_output_loss: 0.6342 - val_main_output_acc: 0.6505 - val_aux_output_acc: 0.7087\nEpoch 142/500\n924/924 [==============================] - 0s 215us/step - loss: 0.7174 - main_output_loss: 0.6550 - aux_output_loss: 0.6242 - main_output_acc: 0.6050 - aux_output_acc: 0.7154 - val_loss: 0.6872 - val_main_output_loss: 0.6239 - val_aux_output_loss: 0.6339 - val_main_output_acc: 0.6505 - val_aux_output_acc: 0.7087\nEpoch 143/500\n924/924 [==============================] - 0s 216us/step - loss: 0.7300 - main_output_loss: 0.6676 - aux_output_loss: 0.6239 - main_output_acc: 0.5790 - aux_output_acc: 0.7175 - val_loss: 0.6863 - val_main_output_loss: 0.6230 - val_aux_output_loss: 0.6333 - val_main_output_acc: 0.6408 - val_aux_output_acc: 0.7087\nEpoch 144/500\n924/924 [==============================] - 0s 212us/step - loss: 0.7244 - main_output_loss: 0.6620 - aux_output_loss: 0.6236 - main_output_acc: 0.5909 - aux_output_acc: 0.7100 - val_loss: 0.6872 - val_main_output_loss: 0.6239 - val_aux_output_loss: 0.6331 - val_main_output_acc: 0.6408 - val_aux_output_acc: 0.6990\nEpoch 145/500\n924/924 [==============================] - 0s 217us/step - loss: 0.7242 - main_output_loss: 0.6619 - aux_output_loss: 0.6233 - main_output_acc: 0.5985 - aux_output_acc: 0.7110 - val_loss: 0.6869 - val_main_output_loss: 0.6236 - val_aux_output_loss: 0.6327 - val_main_output_acc: 0.6699 - val_aux_output_acc: 0.6990\nEpoch 146/500\n924/924 [==============================] - 0s 210us/step - loss: 0.7158 - main_output_loss: 0.6536 - aux_output_loss: 0.6228 - main_output_acc: 0.6039 - aux_output_acc: 0.7110 - val_loss: 0.6876 - val_main_output_loss: 0.6244 - val_aux_output_loss: 0.6327 - val_main_output_acc: 0.6505 - val_aux_output_acc: 0.7087\nEpoch 147/500\n924/924 [==============================] - 0s 210us/step - loss: 0.7169 - main_output_loss: 0.6547 - aux_output_loss: 0.6224 - main_output_acc: 0.5985 - aux_output_acc: 0.7165 - val_loss: 0.6860 - val_main_output_loss: 0.6228 - val_aux_output_loss: 0.6323 - val_main_output_acc: 0.6602 - val_aux_output_acc: 0.6990\nEpoch 148/500\n924/924 [==============================] - 0s 211us/step - loss: 0.7162 - main_output_loss: 0.6540 - aux_output_loss: 0.6221 - main_output_acc: 0.5768 - aux_output_acc: 0.7154 - val_loss: 0.6845 - val_main_output_loss: 0.6213 - val_aux_output_loss: 0.6317 - val_main_output_acc: 0.6408 - val_aux_output_acc: 0.6990\nEpoch 149/500\n924/924 [==============================] - 0s 213us/step - loss: 0.7146 - main_output_loss: 0.6524 - aux_output_loss: 0.6219 - main_output_acc: 0.6006 - aux_output_acc: 0.7154 - val_loss: 0.6846 - val_main_output_loss: 0.6215 - val_aux_output_loss: 0.6314 - val_main_output_acc: 0.6602 - val_aux_output_acc: 0.6990\nEpoch 150/500\n924/924 [==============================] - 0s 215us/step - loss: 0.7193 - main_output_loss: 0.6571 - aux_output_loss: 0.6216 - main_output_acc: 0.5996 - aux_output_acc: 0.7121 - val_loss: 0.6828 - val_main_output_loss: 0.6197 - val_aux_output_loss: 0.6310 - val_main_output_acc: 0.6505 - val_aux_output_acc: 0.7087\nEpoch 151/500\n924/924 [==============================] - 0s 209us/step - loss: 0.7192 - main_output_loss: 0.6571 - aux_output_loss: 0.6214 - main_output_acc: 0.6061 - aux_output_acc: 0.7110 - val_loss: 0.6822 - val_main_output_loss: 0.6192 - val_aux_output_loss: 0.6307 - val_main_output_acc: 0.6602 - val_aux_output_acc: 0.7087\nEpoch 152/500\n924/924 [==============================] - 0s 211us/step - loss: 0.7155 - main_output_loss: 0.6534 - aux_output_loss: 0.6209 - main_output_acc: 0.6201 - aux_output_acc: 0.7132 - val_loss: 0.6817 - val_main_output_loss: 0.6186 - val_aux_output_loss: 0.6302 - val_main_output_acc: 0.6505 - val_aux_output_acc: 0.7087\nEpoch 153/500\n924/924 [==============================] - 0s 206us/step - loss: 0.7217 - main_output_loss: 0.6596 - aux_output_loss: 0.6206 - main_output_acc: 0.5931 - aux_output_acc: 0.7143 - val_loss: 0.6804 - val_main_output_loss: 0.6175 - val_aux_output_loss: 0.6298 - val_main_output_acc: 0.6408 - val_aux_output_acc: 0.7184\nEpoch 154/500\n924/924 [==============================] - 0s 205us/step - loss: 0.7209 - main_output_loss: 0.6589 - aux_output_loss: 0.6201 - main_output_acc: 0.6071 - aux_output_acc: 0.7143 - val_loss: 0.6808 - val_main_output_loss: 0.6179 - val_aux_output_loss: 0.6294 - val_main_output_acc: 0.6505 - val_aux_output_acc: 0.7184\nEpoch 155/500\n924/924 [==============================] - 0s 213us/step - loss: 0.7152 - main_output_loss: 0.6533 - aux_output_loss: 0.6198 - main_output_acc: 0.6104 - aux_output_acc: 0.7143 - val_loss: 0.6798 - val_main_output_loss: 0.6169 - val_aux_output_loss: 0.6288 - val_main_output_acc: 0.6699 - val_aux_output_acc: 0.7184\nEpoch 156/500\n924/924 [==============================] - 0s 207us/step - loss: 0.7187 - main_output_loss: 0.6568 - aux_output_loss: 0.6195 - main_output_acc: 0.6126 - aux_output_acc: 0.7110 - val_loss: 0.6798 - val_main_output_loss: 0.6169 - val_aux_output_loss: 0.6288 - val_main_output_acc: 0.6505 - val_aux_output_acc: 0.7184\nEpoch 157/500\n924/924 [==============================] - 0s 208us/step - loss: 0.7225 - main_output_loss: 0.6605 - aux_output_loss: 0.6191 - main_output_acc: 0.5985 - aux_output_acc: 0.7143 - val_loss: 0.6795 - val_main_output_loss: 0.6167 - val_aux_output_loss: 0.6285 - val_main_output_acc: 0.6505 - val_aux_output_acc: 0.7087\nEpoch 158/500\n924/924 [==============================] - 0s 211us/step - loss: 0.6999 - main_output_loss: 0.6380 - aux_output_loss: 0.6188 - main_output_acc: 0.6364 - aux_output_acc: 0.7165 - val_loss: 0.6791 - val_main_output_loss: 0.6163 - val_aux_output_loss: 0.6281 - val_main_output_acc: 0.6602 - val_aux_output_acc: 0.7087\nEpoch 159/500\n924/924 [==============================] - 0s 209us/step - loss: 0.7152 - main_output_loss: 0.6534 - aux_output_loss: 0.6185 - main_output_acc: 0.5931 - aux_output_acc: 0.7154 - val_loss: 0.6794 - val_main_output_loss: 0.6166 - val_aux_output_loss: 0.6280 - val_main_output_acc: 0.6796 - val_aux_output_acc: 0.7087\nEpoch 160/500\n924/924 [==============================] - 0s 211us/step - loss: 0.7288 - main_output_loss: 0.6670 - aux_output_loss: 0.6182 - main_output_acc: 0.5779 - aux_output_acc: 0.7186 - val_loss: 0.6793 - val_main_output_loss: 0.6165 - val_aux_output_loss: 0.6278 - val_main_output_acc: 0.6699 - val_aux_output_acc: 0.7087\nEpoch 161/500\n924/924 [==============================] - 0s 224us/step - loss: 0.7169 - main_output_loss: 0.6551 - aux_output_loss: 0.6177 - main_output_acc: 0.6061 - aux_output_acc: 0.7186 - val_loss: 0.6800 - val_main_output_loss: 0.6172 - val_aux_output_loss: 0.6276 - val_main_output_acc: 0.6602 - val_aux_output_acc: 0.7087\nEpoch 162/500\n924/924 [==============================] - 0s 209us/step - loss: 0.7143 - main_output_loss: 0.6526 - aux_output_loss: 0.6174 - main_output_acc: 0.5952 - aux_output_acc: 0.7186 - val_loss: 0.6791 - val_main_output_loss: 0.6164 - val_aux_output_loss: 0.6271 - val_main_output_acc: 0.6699 - val_aux_output_acc: 0.7087\nEpoch 163/500\n924/924 [==============================] - 0s 212us/step - loss: 0.7120 - main_output_loss: 0.6502 - aux_output_loss: 0.6171 - main_output_acc: 0.6039 - aux_output_acc: 0.7143 - val_loss: 0.6787 - val_main_output_loss: 0.6160 - val_aux_output_loss: 0.6269 - val_main_output_acc: 0.6602 - val_aux_output_acc: 0.7087\nEpoch 164/500\n924/924 [==============================] - 0s 212us/step - loss: 0.7062 - main_output_loss: 0.6446 - aux_output_loss: 0.6167 - main_output_acc: 0.6126 - aux_output_acc: 0.7154 - val_loss: 0.6762 - val_main_output_loss: 0.6136 - val_aux_output_loss: 0.6261 - val_main_output_acc: 0.6505 - val_aux_output_acc: 0.7087\nEpoch 165/500\n924/924 [==============================] - 0s 210us/step - loss: 0.7131 - main_output_loss: 0.6515 - aux_output_loss: 0.6164 - main_output_acc: 0.5963 - aux_output_acc: 0.7165 - val_loss: 0.6766 - val_main_output_loss: 0.6140 - val_aux_output_loss: 0.6262 - val_main_output_acc: 0.6893 - val_aux_output_acc: 0.7087\nEpoch 166/500\n924/924 [==============================] - 0s 216us/step - loss: 0.7037 - main_output_loss: 0.6421 - aux_output_loss: 0.6160 - main_output_acc: 0.6245 - aux_output_acc: 0.7165 - val_loss: 0.6761 - val_main_output_loss: 0.6135 - val_aux_output_loss: 0.6259 - val_main_output_acc: 0.6796 - val_aux_output_acc: 0.7184\nEpoch 167/500\n924/924 [==============================] - 0s 210us/step - loss: 0.7118 - main_output_loss: 0.6502 - aux_output_loss: 0.6158 - main_output_acc: 0.6017 - aux_output_acc: 0.7175 - val_loss: 0.6766 - val_main_output_loss: 0.6140 - val_aux_output_loss: 0.6258 - val_main_output_acc: 0.6699 - val_aux_output_acc: 0.7087\nEpoch 168/500\n924/924 [==============================] - 0s 211us/step - loss: 0.7097 - main_output_loss: 0.6482 - aux_output_loss: 0.6154 - main_output_acc: 0.6050 - aux_output_acc: 0.7154 - val_loss: 0.6750 - val_main_output_loss: 0.6125 - val_aux_output_loss: 0.6252 - val_main_output_acc: 0.6893 - val_aux_output_acc: 0.7184\nEpoch 169/500\n924/924 [==============================] - 0s 208us/step - loss: 0.7052 - main_output_loss: 0.6437 - aux_output_loss: 0.6150 - main_output_acc: 0.6050 - aux_output_acc: 0.7154 - val_loss: 0.6737 - val_main_output_loss: 0.6112 - val_aux_output_loss: 0.6246 - val_main_output_acc: 0.6602 - val_aux_output_acc: 0.7087\nEpoch 170/500\n924/924 [==============================] - 0s 215us/step - loss: 0.7041 - main_output_loss: 0.6426 - aux_output_loss: 0.6147 - main_output_acc: 0.6136 - aux_output_acc: 0.7121 - val_loss: 0.6737 - val_main_output_loss: 0.6113 - val_aux_output_loss: 0.6241 - val_main_output_acc: 0.6602 - val_aux_output_acc: 0.7087\nEpoch 171/500\n924/924 [==============================] - 0s 218us/step - loss: 0.7060 - main_output_loss: 0.6445 - aux_output_loss: 0.6144 - main_output_acc: 0.6017 - aux_output_acc: 0.7143 - val_loss: 0.6721 - val_main_output_loss: 0.6097 - val_aux_output_loss: 0.6236 - val_main_output_acc: 0.6893 - val_aux_output_acc: 0.7087\nEpoch 172/500\n924/924 [==============================] - 0s 221us/step - loss: 0.7083 - main_output_loss: 0.6469 - aux_output_loss: 0.6141 - main_output_acc: 0.6115 - aux_output_acc: 0.7165 - val_loss: 0.6729 - val_main_output_loss: 0.6106 - val_aux_output_loss: 0.6232 - val_main_output_acc: 0.6796 - val_aux_output_acc: 0.7087\nEpoch 173/500\n924/924 [==============================] - 0s 218us/step - loss: 0.7131 - main_output_loss: 0.6517 - aux_output_loss: 0.6137 - main_output_acc: 0.6050 - aux_output_acc: 0.7154 - val_loss: 0.6727 - val_main_output_loss: 0.6104 - val_aux_output_loss: 0.6228 - val_main_output_acc: 0.6796 - val_aux_output_acc: 0.7087\nEpoch 174/500\n924/924 [==============================] - 0s 212us/step - loss: 0.7020 - main_output_loss: 0.6406 - aux_output_loss: 0.6134 - main_output_acc: 0.6201 - aux_output_acc: 0.7143 - val_loss: 0.6718 - val_main_output_loss: 0.6096 - val_aux_output_loss: 0.6224 - val_main_output_acc: 0.6893 - val_aux_output_acc: 0.7087\nEpoch 175/500\n924/924 [==============================] - 0s 213us/step - loss: 0.7089 - main_output_loss: 0.6476 - aux_output_loss: 0.6131 - main_output_acc: 0.6126 - aux_output_acc: 0.7165 - val_loss: 0.6713 - val_main_output_loss: 0.6090 - val_aux_output_loss: 0.6222 - val_main_output_acc: 0.6796 - val_aux_output_acc: 0.7087\nEpoch 176/500\n924/924 [==============================] - 0s 209us/step - loss: 0.7064 - main_output_loss: 0.6452 - aux_output_loss: 0.6128 - main_output_acc: 0.6158 - aux_output_acc: 0.7143 - val_loss: 0.6702 - val_main_output_loss: 0.6080 - val_aux_output_loss: 0.6219 - val_main_output_acc: 0.6893 - val_aux_output_acc: 0.7087\nEpoch 177/500\n924/924 [==============================] - 0s 214us/step - loss: 0.7090 - main_output_loss: 0.6478 - aux_output_loss: 0.6124 - main_output_acc: 0.6017 - aux_output_acc: 0.7165 - val_loss: 0.6724 - val_main_output_loss: 0.6102 - val_aux_output_loss: 0.6221 - val_main_output_acc: 0.6602 - val_aux_output_acc: 0.7087\nEpoch 178/500\n924/924 [==============================] - 0s 211us/step - loss: 0.7023 - main_output_loss: 0.6411 - aux_output_loss: 0.6121 - main_output_acc: 0.6104 - aux_output_acc: 0.7154 - val_loss: 0.6723 - val_main_output_loss: 0.6101 - val_aux_output_loss: 0.6220 - val_main_output_acc: 0.6699 - val_aux_output_acc: 0.7087\nEpoch 179/500\n924/924 [==============================] - 0s 209us/step - loss: 0.7076 - main_output_loss: 0.6465 - aux_output_loss: 0.6118 - main_output_acc: 0.6147 - aux_output_acc: 0.7175 - val_loss: 0.6727 - val_main_output_loss: 0.6105 - val_aux_output_loss: 0.6217 - val_main_output_acc: 0.6796 - val_aux_output_acc: 0.7184\nEpoch 180/500\n924/924 [==============================] - 0s 213us/step - loss: 0.7045 - main_output_loss: 0.6434 - aux_output_loss: 0.6114 - main_output_acc: 0.6266 - aux_output_acc: 0.7165 - val_loss: 0.6727 - val_main_output_loss: 0.6106 - val_aux_output_loss: 0.6215 - val_main_output_acc: 0.6796 - val_aux_output_acc: 0.7087\nEpoch 181/500\n924/924 [==============================] - 0s 209us/step - loss: 0.6996 - main_output_loss: 0.6385 - aux_output_loss: 0.6111 - main_output_acc: 0.6385 - aux_output_acc: 0.7143 - val_loss: 0.6709 - val_main_output_loss: 0.6088 - val_aux_output_loss: 0.6214 - val_main_output_acc: 0.6796 - val_aux_output_acc: 0.7087\nEpoch 182/500\n924/924 [==============================] - 0s 207us/step - loss: 0.7043 - main_output_loss: 0.6432 - aux_output_loss: 0.6108 - main_output_acc: 0.6136 - aux_output_acc: 0.7165 - val_loss: 0.6695 - val_main_output_loss: 0.6074 - val_aux_output_loss: 0.6208 - val_main_output_acc: 0.6699 - val_aux_output_acc: 0.7282\nEpoch 183/500\n924/924 [==============================] - 0s 210us/step - loss: 0.6907 - main_output_loss: 0.6296 - aux_output_loss: 0.6105 - main_output_acc: 0.6299 - aux_output_acc: 0.7143 - val_loss: 0.6675 - val_main_output_loss: 0.6055 - val_aux_output_loss: 0.6201 - val_main_output_acc: 0.6699 - val_aux_output_acc: 0.7087\nEpoch 184/500\n924/924 [==============================] - 0s 222us/step - loss: 0.7061 - main_output_loss: 0.6450 - aux_output_loss: 0.6101 - main_output_acc: 0.6212 - aux_output_acc: 0.7175 - val_loss: 0.6675 - val_main_output_loss: 0.6056 - val_aux_output_loss: 0.6196 - val_main_output_acc: 0.6699 - val_aux_output_acc: 0.7087\nEpoch 185/500\n924/924 [==============================] - 0s 212us/step - loss: 0.7010 - main_output_loss: 0.6400 - aux_output_loss: 0.6097 - main_output_acc: 0.6006 - aux_output_acc: 0.7186 - val_loss: 0.6676 - val_main_output_loss: 0.6056 - val_aux_output_loss: 0.6191 - val_main_output_acc: 0.6602 - val_aux_output_acc: 0.7087\nEpoch 186/500\n924/924 [==============================] - 0s 222us/step - loss: 0.7091 - main_output_loss: 0.6482 - aux_output_loss: 0.6093 - main_output_acc: 0.6061 - aux_output_acc: 0.7197 - val_loss: 0.6647 - val_main_output_loss: 0.6029 - val_aux_output_loss: 0.6184 - val_main_output_acc: 0.6893 - val_aux_output_acc: 0.7087\nEpoch 187/500\n924/924 [==============================] - 0s 218us/step - loss: 0.6950 - main_output_loss: 0.6341 - aux_output_loss: 0.6091 - main_output_acc: 0.6104 - aux_output_acc: 0.7175 - val_loss: 0.6644 - val_main_output_loss: 0.6026 - val_aux_output_loss: 0.6180 - val_main_output_acc: 0.6893 - val_aux_output_acc: 0.7087\nEpoch 188/500\n924/924 [==============================] - 0s 211us/step - loss: 0.7014 - main_output_loss: 0.6405 - aux_output_loss: 0.6088 - main_output_acc: 0.6277 - aux_output_acc: 0.7186 - val_loss: 0.6646 - val_main_output_loss: 0.6028 - val_aux_output_loss: 0.6178 - val_main_output_acc: 0.6699 - val_aux_output_acc: 0.7087\nEpoch 189/500\n924/924 [==============================] - 0s 207us/step - loss: 0.7021 - main_output_loss: 0.6413 - aux_output_loss: 0.6084 - main_output_acc: 0.6180 - aux_output_acc: 0.7208 - val_loss: 0.6634 - val_main_output_loss: 0.6016 - val_aux_output_loss: 0.6175 - val_main_output_acc: 0.6796 - val_aux_output_acc: 0.7087\nEpoch 190/500\n924/924 [==============================] - 0s 211us/step - loss: 0.7073 - main_output_loss: 0.6465 - aux_output_loss: 0.6081 - main_output_acc: 0.5996 - aux_output_acc: 0.7175 - val_loss: 0.6613 - val_main_output_loss: 0.5996 - val_aux_output_loss: 0.6170 - val_main_output_acc: 0.6893 - val_aux_output_acc: 0.7087\nEpoch 191/500\n924/924 [==============================] - 0s 211us/step - loss: 0.6976 - main_output_loss: 0.6368 - aux_output_loss: 0.6077 - main_output_acc: 0.6234 - aux_output_acc: 0.7154 - val_loss: 0.6611 - val_main_output_loss: 0.5994 - val_aux_output_loss: 0.6165 - val_main_output_acc: 0.6893 - val_aux_output_acc: 0.7087\nEpoch 192/500\n924/924 [==============================] - 0s 213us/step - loss: 0.7061 - main_output_loss: 0.6453 - aux_output_loss: 0.6075 - main_output_acc: 0.6039 - aux_output_acc: 0.7175 - val_loss: 0.6619 - val_main_output_loss: 0.6002 - val_aux_output_loss: 0.6168 - val_main_output_acc: 0.6893 - val_aux_output_acc: 0.7087\nEpoch 193/500\n924/924 [==============================] - 0s 211us/step - loss: 0.6982 - main_output_loss: 0.6374 - aux_output_loss: 0.6071 - main_output_acc: 0.6266 - aux_output_acc: 0.7154 - val_loss: 0.6613 - val_main_output_loss: 0.5997 - val_aux_output_loss: 0.6164 - val_main_output_acc: 0.6699 - val_aux_output_acc: 0.7087\nEpoch 194/500\n924/924 [==============================] - 0s 210us/step - loss: 0.6945 - main_output_loss: 0.6339 - aux_output_loss: 0.6067 - main_output_acc: 0.6288 - aux_output_acc: 0.7154 - val_loss: 0.6620 - val_main_output_loss: 0.6003 - val_aux_output_loss: 0.6167 - val_main_output_acc: 0.6893 - val_aux_output_acc: 0.7184\nEpoch 195/500\n924/924 [==============================] - 0s 209us/step - loss: 0.7003 - main_output_loss: 0.6396 - aux_output_loss: 0.6065 - main_output_acc: 0.6169 - aux_output_acc: 0.7175 - val_loss: 0.6597 - val_main_output_loss: 0.5981 - val_aux_output_loss: 0.6161 - val_main_output_acc: 0.6893 - val_aux_output_acc: 0.7087\nEpoch 196/500\n924/924 [==============================] - 0s 211us/step - loss: 0.6933 - main_output_loss: 0.6327 - aux_output_loss: 0.6061 - main_output_acc: 0.6288 - aux_output_acc: 0.7208 - val_loss: 0.6578 - val_main_output_loss: 0.5962 - val_aux_output_loss: 0.6155 - val_main_output_acc: 0.7087 - val_aux_output_acc: 0.7087\nEpoch 197/500\n924/924 [==============================] - 0s 206us/step - loss: 0.6967 - main_output_loss: 0.6361 - aux_output_loss: 0.6059 - main_output_acc: 0.6234 - aux_output_acc: 0.7186 - val_loss: 0.6571 - val_main_output_loss: 0.5956 - val_aux_output_loss: 0.6154 - val_main_output_acc: 0.7087 - val_aux_output_acc: 0.7087\nEpoch 198/500\n924/924 [==============================] - 0s 213us/step - loss: 0.6934 - main_output_loss: 0.6329 - aux_output_loss: 0.6055 - main_output_acc: 0.6190 - aux_output_acc: 0.7165 - val_loss: 0.6563 - val_main_output_loss: 0.5948 - val_aux_output_loss: 0.6149 - val_main_output_acc: 0.7087 - val_aux_output_acc: 0.7087\nEpoch 199/500\n924/924 [==============================] - 0s 210us/step - loss: 0.6992 - main_output_loss: 0.6387 - aux_output_loss: 0.6051 - main_output_acc: 0.6061 - aux_output_acc: 0.7154 - val_loss: 0.6566 - val_main_output_loss: 0.5951 - val_aux_output_loss: 0.6150 - val_main_output_acc: 0.7184 - val_aux_output_acc: 0.7087\nEpoch 200/500\n924/924 [==============================] - 0s 205us/step - loss: 0.6913 - main_output_loss: 0.6308 - aux_output_loss: 0.6049 - main_output_acc: 0.6396 - aux_output_acc: 0.7186 - val_loss: 0.6557 - val_main_output_loss: 0.5943 - val_aux_output_loss: 0.6145 - val_main_output_acc: 0.7184 - val_aux_output_acc: 0.7087\nEpoch 201/500\n924/924 [==============================] - 0s 215us/step - loss: 0.6960 - main_output_loss: 0.6356 - aux_output_loss: 0.6046 - main_output_acc: 0.6169 - aux_output_acc: 0.7175 - val_loss: 0.6568 - val_main_output_loss: 0.5953 - val_aux_output_loss: 0.6148 - val_main_output_acc: 0.6893 - val_aux_output_acc: 0.7087\nEpoch 202/500\n924/924 [==============================] - 0s 213us/step - loss: 0.6960 - main_output_loss: 0.6355 - aux_output_loss: 0.6043 - main_output_acc: 0.6266 - aux_output_acc: 0.7197 - val_loss: 0.6553 - val_main_output_loss: 0.5939 - val_aux_output_loss: 0.6144 - val_main_output_acc: 0.7087 - val_aux_output_acc: 0.7087\nEpoch 203/500\n924/924 [==============================] - 0s 212us/step - loss: 0.6961 - main_output_loss: 0.6357 - aux_output_loss: 0.6040 - main_output_acc: 0.6245 - aux_output_acc: 0.7165 - val_loss: 0.6549 - val_main_output_loss: 0.5936 - val_aux_output_loss: 0.6139 - val_main_output_acc: 0.6990 - val_aux_output_acc: 0.6990\nEpoch 204/500\n924/924 [==============================] - 0s 207us/step - loss: 0.6964 - main_output_loss: 0.6361 - aux_output_loss: 0.6035 - main_output_acc: 0.6331 - aux_output_acc: 0.7154 - val_loss: 0.6540 - val_main_output_loss: 0.5926 - val_aux_output_loss: 0.6136 - val_main_output_acc: 0.7087 - val_aux_output_acc: 0.7087\nEpoch 205/500\n924/924 [==============================] - 0s 212us/step - loss: 0.6922 - main_output_loss: 0.6319 - aux_output_loss: 0.6033 - main_output_acc: 0.6374 - aux_output_acc: 0.7186 - val_loss: 0.6547 - val_main_output_loss: 0.5933 - val_aux_output_loss: 0.6137 - val_main_output_acc: 0.7087 - val_aux_output_acc: 0.7087\nEpoch 206/500\n924/924 [==============================] - 0s 211us/step - loss: 0.6966 - main_output_loss: 0.6363 - aux_output_loss: 0.6031 - main_output_acc: 0.6201 - aux_output_acc: 0.7165 - val_loss: 0.6530 - val_main_output_loss: 0.5917 - val_aux_output_loss: 0.6133 - val_main_output_acc: 0.7184 - val_aux_output_acc: 0.6990\nEpoch 207/500\n924/924 [==============================] - 0s 211us/step - loss: 0.6911 - main_output_loss: 0.6309 - aux_output_loss: 0.6028 - main_output_acc: 0.6299 - aux_output_acc: 0.7175 - val_loss: 0.6521 - val_main_output_loss: 0.5907 - val_aux_output_loss: 0.6133 - val_main_output_acc: 0.7184 - val_aux_output_acc: 0.7087\nEpoch 208/500\n924/924 [==============================] - 0s 210us/step - loss: 0.7062 - main_output_loss: 0.6459 - aux_output_loss: 0.6025 - main_output_acc: 0.6147 - aux_output_acc: 0.7143 - val_loss: 0.6510 - val_main_output_loss: 0.5897 - val_aux_output_loss: 0.6128 - val_main_output_acc: 0.7282 - val_aux_output_acc: 0.6990\nEpoch 209/500\n924/924 [==============================] - 0s 223us/step - loss: 0.6882 - main_output_loss: 0.6280 - aux_output_loss: 0.6022 - main_output_acc: 0.6320 - aux_output_acc: 0.7219 - val_loss: 0.6516 - val_main_output_loss: 0.5903 - val_aux_output_loss: 0.6128 - val_main_output_acc: 0.7184 - val_aux_output_acc: 0.6990\nEpoch 210/500\n924/924 [==============================] - 0s 207us/step - loss: 0.6881 - main_output_loss: 0.6279 - aux_output_loss: 0.6018 - main_output_acc: 0.6418 - aux_output_acc: 0.7165 - val_loss: 0.6521 - val_main_output_loss: 0.5908 - val_aux_output_loss: 0.6127 - val_main_output_acc: 0.7087 - val_aux_output_acc: 0.7087\nEpoch 211/500\n924/924 [==============================] - 0s 217us/step - loss: 0.6894 - main_output_loss: 0.6293 - aux_output_loss: 0.6016 - main_output_acc: 0.6255 - aux_output_acc: 0.7165 - val_loss: 0.6499 - val_main_output_loss: 0.5887 - val_aux_output_loss: 0.6121 - val_main_output_acc: 0.7087 - val_aux_output_acc: 0.6990\nEpoch 212/500\n924/924 [==============================] - 0s 210us/step - loss: 0.6889 - main_output_loss: 0.6288 - aux_output_loss: 0.6013 - main_output_acc: 0.6364 - aux_output_acc: 0.7219 - val_loss: 0.6507 - val_main_output_loss: 0.5895 - val_aux_output_loss: 0.6119 - val_main_output_acc: 0.7087 - val_aux_output_acc: 0.7087\nEpoch 213/500\n924/924 [==============================] - 0s 205us/step - loss: 0.6925 - main_output_loss: 0.6324 - aux_output_loss: 0.6009 - main_output_acc: 0.6169 - aux_output_acc: 0.7154 - val_loss: 0.6513 - val_main_output_loss: 0.5901 - val_aux_output_loss: 0.6119 - val_main_output_acc: 0.7184 - val_aux_output_acc: 0.7087\nEpoch 214/500\n924/924 [==============================] - 0s 206us/step - loss: 0.6868 - main_output_loss: 0.6268 - aux_output_loss: 0.6006 - main_output_acc: 0.6580 - aux_output_acc: 0.7132 - val_loss: 0.6502 - val_main_output_loss: 0.5891 - val_aux_output_loss: 0.6114 - val_main_output_acc: 0.7184 - val_aux_output_acc: 0.7087\nEpoch 215/500\n924/924 [==============================] - 0s 207us/step - loss: 0.6834 - main_output_loss: 0.6234 - aux_output_loss: 0.6002 - main_output_acc: 0.6288 - aux_output_acc: 0.7132 - val_loss: 0.6462 - val_main_output_loss: 0.5852 - val_aux_output_loss: 0.6106 - val_main_output_acc: 0.7184 - val_aux_output_acc: 0.6990\nEpoch 216/500\n924/924 [==============================] - 0s 213us/step - loss: 0.6892 - main_output_loss: 0.6292 - aux_output_loss: 0.5999 - main_output_acc: 0.6245 - aux_output_acc: 0.7197 - val_loss: 0.6474 - val_main_output_loss: 0.5863 - val_aux_output_loss: 0.6107 - val_main_output_acc: 0.7282 - val_aux_output_acc: 0.7087\nEpoch 217/500\n924/924 [==============================] - 0s 207us/step - loss: 0.6827 - main_output_loss: 0.6227 - aux_output_loss: 0.5996 - main_output_acc: 0.6461 - aux_output_acc: 0.7165 - val_loss: 0.6473 - val_main_output_loss: 0.5862 - val_aux_output_loss: 0.6106 - val_main_output_acc: 0.7282 - val_aux_output_acc: 0.7087\nEpoch 218/500\n924/924 [==============================] - 0s 213us/step - loss: 0.6877 - main_output_loss: 0.6277 - aux_output_loss: 0.5994 - main_output_acc: 0.6407 - aux_output_acc: 0.7186 - val_loss: 0.6468 - val_main_output_loss: 0.5857 - val_aux_output_loss: 0.6104 - val_main_output_acc: 0.7379 - val_aux_output_acc: 0.7087\nEpoch 219/500\n924/924 [==============================] - 0s 216us/step - loss: 0.6894 - main_output_loss: 0.6295 - aux_output_loss: 0.5992 - main_output_acc: 0.6310 - aux_output_acc: 0.7154 - val_loss: 0.6440 - val_main_output_loss: 0.5831 - val_aux_output_loss: 0.6097 - val_main_output_acc: 0.7184 - val_aux_output_acc: 0.7087\nEpoch 220/500\n924/924 [==============================] - 0s 210us/step - loss: 0.6844 - main_output_loss: 0.6245 - aux_output_loss: 0.5988 - main_output_acc: 0.6483 - aux_output_acc: 0.7175 - val_loss: 0.6424 - val_main_output_loss: 0.5815 - val_aux_output_loss: 0.6092 - val_main_output_acc: 0.7184 - val_aux_output_acc: 0.6990\nEpoch 221/500\n924/924 [==============================] - 0s 207us/step - loss: 0.6892 - main_output_loss: 0.6293 - aux_output_loss: 0.5984 - main_output_acc: 0.6461 - aux_output_acc: 0.7219 - val_loss: 0.6432 - val_main_output_loss: 0.5823 - val_aux_output_loss: 0.6091 - val_main_output_acc: 0.7379 - val_aux_output_acc: 0.7087\nEpoch 222/500\n924/924 [==============================] - 0s 214us/step - loss: 0.6882 - main_output_loss: 0.6284 - aux_output_loss: 0.5982 - main_output_acc: 0.6299 - aux_output_acc: 0.7251 - val_loss: 0.6446 - val_main_output_loss: 0.5837 - val_aux_output_loss: 0.6093 - val_main_output_acc: 0.7379 - val_aux_output_acc: 0.7087\nEpoch 223/500\n924/924 [==============================] - 0s 210us/step - loss: 0.6822 - main_output_loss: 0.6224 - aux_output_loss: 0.5979 - main_output_acc: 0.6418 - aux_output_acc: 0.7154 - val_loss: 0.6419 - val_main_output_loss: 0.5810 - val_aux_output_loss: 0.6086 - val_main_output_acc: 0.7282 - val_aux_output_acc: 0.7087\nEpoch 224/500\n924/924 [==============================] - 0s 214us/step - loss: 0.6875 - main_output_loss: 0.6277 - aux_output_loss: 0.5976 - main_output_acc: 0.6299 - aux_output_acc: 0.7197 - val_loss: 0.6395 - val_main_output_loss: 0.5787 - val_aux_output_loss: 0.6078 - val_main_output_acc: 0.7282 - val_aux_output_acc: 0.7087\nEpoch 225/500\n924/924 [==============================] - 0s 220us/step - loss: 0.6889 - main_output_loss: 0.6292 - aux_output_loss: 0.5975 - main_output_acc: 0.6320 - aux_output_acc: 0.7229 - val_loss: 0.6392 - val_main_output_loss: 0.5785 - val_aux_output_loss: 0.6074 - val_main_output_acc: 0.7184 - val_aux_output_acc: 0.7184\nEpoch 226/500\n924/924 [==============================] - 0s 213us/step - loss: 0.6845 - main_output_loss: 0.6248 - aux_output_loss: 0.5972 - main_output_acc: 0.6396 - aux_output_acc: 0.7262 - val_loss: 0.6390 - val_main_output_loss: 0.5783 - val_aux_output_loss: 0.6072 - val_main_output_acc: 0.7282 - val_aux_output_acc: 0.7184\nEpoch 227/500\n924/924 [==============================] - 0s 218us/step - loss: 0.6791 - main_output_loss: 0.6194 - aux_output_loss: 0.5970 - main_output_acc: 0.6515 - aux_output_acc: 0.7219 - val_loss: 0.6396 - val_main_output_loss: 0.5789 - val_aux_output_loss: 0.6072 - val_main_output_acc: 0.7282 - val_aux_output_acc: 0.7087\nEpoch 228/500\n924/924 [==============================] - 0s 212us/step - loss: 0.6926 - main_output_loss: 0.6329 - aux_output_loss: 0.5966 - main_output_acc: 0.6299 - aux_output_acc: 0.7229 - val_loss: 0.6407 - val_main_output_loss: 0.5800 - val_aux_output_loss: 0.6072 - val_main_output_acc: 0.7282 - val_aux_output_acc: 0.7087\nEpoch 229/500\n924/924 [==============================] - 0s 211us/step - loss: 0.6819 - main_output_loss: 0.6222 - aux_output_loss: 0.5964 - main_output_acc: 0.6439 - aux_output_acc: 0.7197 - val_loss: 0.6419 - val_main_output_loss: 0.5812 - val_aux_output_loss: 0.6073 - val_main_output_acc: 0.7476 - val_aux_output_acc: 0.7087\nEpoch 230/500\n924/924 [==============================] - 0s 211us/step - loss: 0.6856 - main_output_loss: 0.6259 - aux_output_loss: 0.5961 - main_output_acc: 0.6396 - aux_output_acc: 0.7154 - val_loss: 0.6412 - val_main_output_loss: 0.5805 - val_aux_output_loss: 0.6071 - val_main_output_acc: 0.7476 - val_aux_output_acc: 0.7087\nEpoch 231/500\n924/924 [==============================] - 0s 215us/step - loss: 0.6761 - main_output_loss: 0.6165 - aux_output_loss: 0.5959 - main_output_acc: 0.6396 - aux_output_acc: 0.7186 - val_loss: 0.6381 - val_main_output_loss: 0.5774 - val_aux_output_loss: 0.6066 - val_main_output_acc: 0.7379 - val_aux_output_acc: 0.7087\nEpoch 232/500\n924/924 [==============================] - 0s 207us/step - loss: 0.6896 - main_output_loss: 0.6301 - aux_output_loss: 0.5956 - main_output_acc: 0.6320 - aux_output_acc: 0.7197 - val_loss: 0.6368 - val_main_output_loss: 0.5762 - val_aux_output_loss: 0.6059 - val_main_output_acc: 0.7379 - val_aux_output_acc: 0.7184\nEpoch 233/500\n924/924 [==============================] - 0s 208us/step - loss: 0.6854 - main_output_loss: 0.6259 - aux_output_loss: 0.5953 - main_output_acc: 0.6255 - aux_output_acc: 0.7154 - val_loss: 0.6362 - val_main_output_loss: 0.5756 - val_aux_output_loss: 0.6057 - val_main_output_acc: 0.7379 - val_aux_output_acc: 0.7184\nEpoch 234/500\n924/924 [==============================] - 0s 217us/step - loss: 0.6799 - main_output_loss: 0.6204 - aux_output_loss: 0.5950 - main_output_acc: 0.6331 - aux_output_acc: 0.7219 - val_loss: 0.6374 - val_main_output_loss: 0.5769 - val_aux_output_loss: 0.6057 - val_main_output_acc: 0.7282 - val_aux_output_acc: 0.7184\nEpoch 235/500\n924/924 [==============================] - 0s 223us/step - loss: 0.6869 - main_output_loss: 0.6274 - aux_output_loss: 0.5947 - main_output_acc: 0.6223 - aux_output_acc: 0.7186 - val_loss: 0.6356 - val_main_output_loss: 0.5751 - val_aux_output_loss: 0.6049 - val_main_output_acc: 0.7379 - val_aux_output_acc: 0.7184\nEpoch 236/500\n924/924 [==============================] - 0s 217us/step - loss: 0.6798 - main_output_loss: 0.6203 - aux_output_loss: 0.5945 - main_output_acc: 0.6439 - aux_output_acc: 0.7219 - val_loss: 0.6353 - val_main_output_loss: 0.5749 - val_aux_output_loss: 0.6047 - val_main_output_acc: 0.7379 - val_aux_output_acc: 0.7184\nEpoch 237/500\n924/924 [==============================] - 0s 208us/step - loss: 0.6766 - main_output_loss: 0.6171 - aux_output_loss: 0.5944 - main_output_acc: 0.6364 - aux_output_acc: 0.7208 - val_loss: 0.6354 - val_main_output_loss: 0.5750 - val_aux_output_loss: 0.6046 - val_main_output_acc: 0.7282 - val_aux_output_acc: 0.7184\nEpoch 238/500\n924/924 [==============================] - 0s 212us/step - loss: 0.6786 - main_output_loss: 0.6192 - aux_output_loss: 0.5939 - main_output_acc: 0.6537 - aux_output_acc: 0.7219 - val_loss: 0.6365 - val_main_output_loss: 0.5760 - val_aux_output_loss: 0.6048 - val_main_output_acc: 0.7476 - val_aux_output_acc: 0.7184\nEpoch 239/500\n924/924 [==============================] - 0s 215us/step - loss: 0.6757 - main_output_loss: 0.6164 - aux_output_loss: 0.5937 - main_output_acc: 0.6461 - aux_output_acc: 0.7186 - val_loss: 0.6352 - val_main_output_loss: 0.5747 - val_aux_output_loss: 0.6047 - val_main_output_acc: 0.7476 - val_aux_output_acc: 0.7184\nEpoch 240/500\n924/924 [==============================] - 0s 208us/step - loss: 0.6772 - main_output_loss: 0.6179 - aux_output_loss: 0.5933 - main_output_acc: 0.6580 - aux_output_acc: 0.7208 - val_loss: 0.6323 - val_main_output_loss: 0.5719 - val_aux_output_loss: 0.6039 - val_main_output_acc: 0.7379 - val_aux_output_acc: 0.7184\nEpoch 241/500\n924/924 [==============================] - 0s 209us/step - loss: 0.6729 - main_output_loss: 0.6136 - aux_output_loss: 0.5931 - main_output_acc: 0.6504 - aux_output_acc: 0.7219 - val_loss: 0.6312 - val_main_output_loss: 0.5709 - val_aux_output_loss: 0.6035 - val_main_output_acc: 0.7379 - val_aux_output_acc: 0.7184\nEpoch 242/500\n924/924 [==============================] - 0s 207us/step - loss: 0.6796 - main_output_loss: 0.6203 - aux_output_loss: 0.5929 - main_output_acc: 0.6407 - aux_output_acc: 0.7186 - val_loss: 0.6302 - val_main_output_loss: 0.5700 - val_aux_output_loss: 0.6028 - val_main_output_acc: 0.7282 - val_aux_output_acc: 0.7184\nEpoch 243/500\n924/924 [==============================] - 0s 209us/step - loss: 0.6861 - main_output_loss: 0.6269 - aux_output_loss: 0.5925 - main_output_acc: 0.6320 - aux_output_acc: 0.7197 - val_loss: 0.6307 - val_main_output_loss: 0.5704 - val_aux_output_loss: 0.6027 - val_main_output_acc: 0.7476 - val_aux_output_acc: 0.7184\nEpoch 244/500\n924/924 [==============================] - 0s 218us/step - loss: 0.6765 - main_output_loss: 0.6173 - aux_output_loss: 0.5922 - main_output_acc: 0.6483 - aux_output_acc: 0.7197 - val_loss: 0.6294 - val_main_output_loss: 0.5692 - val_aux_output_loss: 0.6023 - val_main_output_acc: 0.7476 - val_aux_output_acc: 0.7184\nEpoch 245/500\n924/924 [==============================] - 0s 209us/step - loss: 0.6781 - main_output_loss: 0.6189 - aux_output_loss: 0.5919 - main_output_acc: 0.6353 - aux_output_acc: 0.7197 - val_loss: 0.6299 - val_main_output_loss: 0.5697 - val_aux_output_loss: 0.6021 - val_main_output_acc: 0.7282 - val_aux_output_acc: 0.7184\nEpoch 246/500\n924/924 [==============================] - 0s 207us/step - loss: 0.6729 - main_output_loss: 0.6137 - aux_output_loss: 0.5916 - main_output_acc: 0.6548 - aux_output_acc: 0.7186 - val_loss: 0.6288 - val_main_output_loss: 0.5687 - val_aux_output_loss: 0.6014 - val_main_output_acc: 0.7379 - val_aux_output_acc: 0.7184\nEpoch 247/500\n924/924 [==============================] - 0s 208us/step - loss: 0.6714 - main_output_loss: 0.6122 - aux_output_loss: 0.5914 - main_output_acc: 0.6656 - aux_output_acc: 0.7186 - val_loss: 0.6271 - val_main_output_loss: 0.5670 - val_aux_output_loss: 0.6010 - val_main_output_acc: 0.7379 - val_aux_output_acc: 0.7184\nEpoch 248/500\n924/924 [==============================] - 0s 210us/step - loss: 0.6724 - main_output_loss: 0.6133 - aux_output_loss: 0.5910 - main_output_acc: 0.6450 - aux_output_acc: 0.7208 - val_loss: 0.6274 - val_main_output_loss: 0.5674 - val_aux_output_loss: 0.6008 - val_main_output_acc: 0.7379 - val_aux_output_acc: 0.7282\nEpoch 249/500\n924/924 [==============================] - 0s 214us/step - loss: 0.6770 - main_output_loss: 0.6179 - aux_output_loss: 0.5910 - main_output_acc: 0.6569 - aux_output_acc: 0.7219 - val_loss: 0.6292 - val_main_output_loss: 0.5691 - val_aux_output_loss: 0.6013 - val_main_output_acc: 0.7476 - val_aux_output_acc: 0.7184\nEpoch 250/500\n924/924 [==============================] - 0s 210us/step - loss: 0.6662 - main_output_loss: 0.6071 - aux_output_loss: 0.5904 - main_output_acc: 0.6364 - aux_output_acc: 0.7197 - val_loss: 0.6277 - val_main_output_loss: 0.5677 - val_aux_output_loss: 0.6005 - val_main_output_acc: 0.7476 - val_aux_output_acc: 0.7184\nEpoch 251/500\n924/924 [==============================] - 0s 206us/step - loss: 0.6716 - main_output_loss: 0.6126 - aux_output_loss: 0.5902 - main_output_acc: 0.6645 - aux_output_acc: 0.7208 - val_loss: 0.6255 - val_main_output_loss: 0.5656 - val_aux_output_loss: 0.5997 - val_main_output_acc: 0.7379 - val_aux_output_acc: 0.7184\nEpoch 252/500\n924/924 [==============================] - 0s 211us/step - loss: 0.6690 - main_output_loss: 0.6100 - aux_output_loss: 0.5899 - main_output_acc: 0.6526 - aux_output_acc: 0.7208 - val_loss: 0.6250 - val_main_output_loss: 0.5651 - val_aux_output_loss: 0.5994 - val_main_output_acc: 0.7379 - val_aux_output_acc: 0.7184\nEpoch 253/500\n924/924 [==============================] - 0s 211us/step - loss: 0.6600 - main_output_loss: 0.6010 - aux_output_loss: 0.5898 - main_output_acc: 0.6948 - aux_output_acc: 0.7251 - val_loss: 0.6252 - val_main_output_loss: 0.5652 - val_aux_output_loss: 0.5997 - val_main_output_acc: 0.7573 - val_aux_output_acc: 0.7184\nEpoch 254/500\n924/924 [==============================] - 0s 214us/step - loss: 0.6700 - main_output_loss: 0.6111 - aux_output_loss: 0.5894 - main_output_acc: 0.6602 - aux_output_acc: 0.7219 - val_loss: 0.6267 - val_main_output_loss: 0.5667 - val_aux_output_loss: 0.5999 - val_main_output_acc: 0.7476 - val_aux_output_acc: 0.7282\nEpoch 255/500\n924/924 [==============================] - 0s 214us/step - loss: 0.6713 - main_output_loss: 0.6124 - aux_output_loss: 0.5892 - main_output_acc: 0.6537 - aux_output_acc: 0.7197 - val_loss: 0.6241 - val_main_output_loss: 0.5642 - val_aux_output_loss: 0.5990 - val_main_output_acc: 0.7476 - val_aux_output_acc: 0.7184\nEpoch 256/500\n924/924 [==============================] - 0s 215us/step - loss: 0.6626 - main_output_loss: 0.6037 - aux_output_loss: 0.5888 - main_output_acc: 0.6623 - aux_output_acc: 0.7229 - val_loss: 0.6248 - val_main_output_loss: 0.5649 - val_aux_output_loss: 0.5989 - val_main_output_acc: 0.7476 - val_aux_output_acc: 0.7184\nEpoch 257/500\n924/924 [==============================] - 0s 217us/step - loss: 0.6668 - main_output_loss: 0.6079 - aux_output_loss: 0.5885 - main_output_acc: 0.6569 - aux_output_acc: 0.7208 - val_loss: 0.6240 - val_main_output_loss: 0.5641 - val_aux_output_loss: 0.5988 - val_main_output_acc: 0.7476 - val_aux_output_acc: 0.7282\nEpoch 258/500\n924/924 [==============================] - 0s 214us/step - loss: 0.6727 - main_output_loss: 0.6138 - aux_output_loss: 0.5882 - main_output_acc: 0.6504 - aux_output_acc: 0.7229 - val_loss: 0.6241 - val_main_output_loss: 0.5642 - val_aux_output_loss: 0.5987 - val_main_output_acc: 0.7476 - val_aux_output_acc: 0.7282\nEpoch 259/500\n924/924 [==============================] - 0s 211us/step - loss: 0.6732 - main_output_loss: 0.6144 - aux_output_loss: 0.5880 - main_output_acc: 0.6537 - aux_output_acc: 0.7186 - val_loss: 0.6216 - val_main_output_loss: 0.5618 - val_aux_output_loss: 0.5979 - val_main_output_acc: 0.7379 - val_aux_output_acc: 0.7184\nEpoch 260/500\n924/924 [==============================] - 0s 212us/step - loss: 0.6722 - main_output_loss: 0.6134 - aux_output_loss: 0.5877 - main_output_acc: 0.6558 - aux_output_acc: 0.7240 - val_loss: 0.6210 - val_main_output_loss: 0.5612 - val_aux_output_loss: 0.5974 - val_main_output_acc: 0.7476 - val_aux_output_acc: 0.7184\nEpoch 261/500\n924/924 [==============================] - 0s 216us/step - loss: 0.6599 - main_output_loss: 0.6011 - aux_output_loss: 0.5875 - main_output_acc: 0.6558 - aux_output_acc: 0.7262 - val_loss: 0.6192 - val_main_output_loss: 0.5595 - val_aux_output_loss: 0.5970 - val_main_output_acc: 0.7573 - val_aux_output_acc: 0.7184\nEpoch 262/500\n924/924 [==============================] - 0s 220us/step - loss: 0.6598 - main_output_loss: 0.6011 - aux_output_loss: 0.5872 - main_output_acc: 0.6688 - aux_output_acc: 0.7284 - val_loss: 0.6201 - val_main_output_loss: 0.5604 - val_aux_output_loss: 0.5970 - val_main_output_acc: 0.7282 - val_aux_output_acc: 0.7184\nEpoch 263/500\n924/924 [==============================] - 0s 213us/step - loss: 0.6663 - main_output_loss: 0.6076 - aux_output_loss: 0.5869 - main_output_acc: 0.6569 - aux_output_acc: 0.7219 - val_loss: 0.6180 - val_main_output_loss: 0.5583 - val_aux_output_loss: 0.5964 - val_main_output_acc: 0.7573 - val_aux_output_acc: 0.7282\nEpoch 264/500\n924/924 [==============================] - 0s 214us/step - loss: 0.6723 - main_output_loss: 0.6136 - aux_output_loss: 0.5867 - main_output_acc: 0.6439 - aux_output_acc: 0.7273 - val_loss: 0.6176 - val_main_output_loss: 0.5580 - val_aux_output_loss: 0.5961 - val_main_output_acc: 0.7573 - val_aux_output_acc: 0.7184\nEpoch 265/500\n924/924 [==============================] - 0s 208us/step - loss: 0.6479 - main_output_loss: 0.5893 - aux_output_loss: 0.5864 - main_output_acc: 0.6677 - aux_output_acc: 0.7273 - val_loss: 0.6181 - val_main_output_loss: 0.5584 - val_aux_output_loss: 0.5963 - val_main_output_acc: 0.7476 - val_aux_output_acc: 0.7282\nEpoch 266/500\n924/924 [==============================] - 0s 210us/step - loss: 0.6610 - main_output_loss: 0.6024 - aux_output_loss: 0.5861 - main_output_acc: 0.6645 - aux_output_acc: 0.7240 - val_loss: 0.6197 - val_main_output_loss: 0.5600 - val_aux_output_loss: 0.5967 - val_main_output_acc: 0.7864 - val_aux_output_acc: 0.7282\nEpoch 267/500\n924/924 [==============================] - 0s 234us/step - loss: 0.6690 - main_output_loss: 0.6104 - aux_output_loss: 0.5858 - main_output_acc: 0.6613 - aux_output_acc: 0.7219 - val_loss: 0.6169 - val_main_output_loss: 0.5573 - val_aux_output_loss: 0.5959 - val_main_output_acc: 0.7476 - val_aux_output_acc: 0.7282\nEpoch 268/500\n924/924 [==============================] - 0s 221us/step - loss: 0.6588 - main_output_loss: 0.6003 - aux_output_loss: 0.5855 - main_output_acc: 0.6883 - aux_output_acc: 0.7229 - val_loss: 0.6170 - val_main_output_loss: 0.5574 - val_aux_output_loss: 0.5956 - val_main_output_acc: 0.7476 - val_aux_output_acc: 0.7282\nEpoch 269/500\n924/924 [==============================] - 0s 220us/step - loss: 0.6621 - main_output_loss: 0.6036 - aux_output_loss: 0.5853 - main_output_acc: 0.6775 - aux_output_acc: 0.7219 - val_loss: 0.6177 - val_main_output_loss: 0.5581 - val_aux_output_loss: 0.5957 - val_main_output_acc: 0.7767 - val_aux_output_acc: 0.7282\nEpoch 270/500\n924/924 [==============================] - 0s 224us/step - loss: 0.6542 - main_output_loss: 0.5957 - aux_output_loss: 0.5851 - main_output_acc: 0.6634 - aux_output_acc: 0.7251 - val_loss: 0.6149 - val_main_output_loss: 0.5554 - val_aux_output_loss: 0.5949 - val_main_output_acc: 0.7379 - val_aux_output_acc: 0.7282\nEpoch 271/500\n924/924 [==============================] - 0s 215us/step - loss: 0.6611 - main_output_loss: 0.6026 - aux_output_loss: 0.5847 - main_output_acc: 0.6504 - aux_output_acc: 0.7229 - val_loss: 0.6158 - val_main_output_loss: 0.5563 - val_aux_output_loss: 0.5946 - val_main_output_acc: 0.7476 - val_aux_output_acc: 0.7282\nEpoch 272/500\n924/924 [==============================] - 0s 216us/step - loss: 0.6498 - main_output_loss: 0.5914 - aux_output_loss: 0.5846 - main_output_acc: 0.6786 - aux_output_acc: 0.7219 - val_loss: 0.6141 - val_main_output_loss: 0.5547 - val_aux_output_loss: 0.5941 - val_main_output_acc: 0.7379 - val_aux_output_acc: 0.7282\nEpoch 273/500\n924/924 [==============================] - 0s 221us/step - loss: 0.6632 - main_output_loss: 0.6048 - aux_output_loss: 0.5842 - main_output_acc: 0.6602 - aux_output_acc: 0.7251 - val_loss: 0.6132 - val_main_output_loss: 0.5538 - val_aux_output_loss: 0.5941 - val_main_output_acc: 0.7476 - val_aux_output_acc: 0.7282\nEpoch 274/500\n924/924 [==============================] - 0s 221us/step - loss: 0.6653 - main_output_loss: 0.6069 - aux_output_loss: 0.5839 - main_output_acc: 0.6548 - aux_output_acc: 0.7229 - val_loss: 0.6140 - val_main_output_loss: 0.5545 - val_aux_output_loss: 0.5942 - val_main_output_acc: 0.7767 - val_aux_output_acc: 0.7282\nEpoch 275/500\n924/924 [==============================] - 0s 229us/step - loss: 0.6608 - main_output_loss: 0.6025 - aux_output_loss: 0.5837 - main_output_acc: 0.6558 - aux_output_acc: 0.7251 - val_loss: 0.6152 - val_main_output_loss: 0.5558 - val_aux_output_loss: 0.5944 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7282\nEpoch 276/500\n924/924 [==============================] - 0s 224us/step - loss: 0.6492 - main_output_loss: 0.5909 - aux_output_loss: 0.5834 - main_output_acc: 0.6926 - aux_output_acc: 0.7229 - val_loss: 0.6127 - val_main_output_loss: 0.5533 - val_aux_output_loss: 0.5935 - val_main_output_acc: 0.7476 - val_aux_output_acc: 0.7282\nEpoch 277/500\n924/924 [==============================] - 0s 218us/step - loss: 0.6621 - main_output_loss: 0.6038 - aux_output_loss: 0.5832 - main_output_acc: 0.6623 - aux_output_acc: 0.7229 - val_loss: 0.6112 - val_main_output_loss: 0.5519 - val_aux_output_loss: 0.5929 - val_main_output_acc: 0.7379 - val_aux_output_acc: 0.7282\nEpoch 278/500\n924/924 [==============================] - 0s 222us/step - loss: 0.6437 - main_output_loss: 0.5854 - aux_output_loss: 0.5829 - main_output_acc: 0.7045 - aux_output_acc: 0.7262 - val_loss: 0.6103 - val_main_output_loss: 0.5510 - val_aux_output_loss: 0.5926 - val_main_output_acc: 0.7573 - val_aux_output_acc: 0.7282\nEpoch 279/500\n924/924 [==============================] - 0s 219us/step - loss: 0.6590 - main_output_loss: 0.6007 - aux_output_loss: 0.5828 - main_output_acc: 0.6764 - aux_output_acc: 0.7219 - val_loss: 0.6096 - val_main_output_loss: 0.5504 - val_aux_output_loss: 0.5921 - val_main_output_acc: 0.7379 - val_aux_output_acc: 0.7282\nEpoch 280/500\n924/924 [==============================] - 0s 216us/step - loss: 0.6582 - main_output_loss: 0.5999 - aux_output_loss: 0.5824 - main_output_acc: 0.6602 - aux_output_acc: 0.7273 - val_loss: 0.6096 - val_main_output_loss: 0.5504 - val_aux_output_loss: 0.5922 - val_main_output_acc: 0.7573 - val_aux_output_acc: 0.7282\nEpoch 281/500\n924/924 [==============================] - 0s 218us/step - loss: 0.6595 - main_output_loss: 0.6013 - aux_output_loss: 0.5823 - main_output_acc: 0.6634 - aux_output_acc: 0.7262 - val_loss: 0.6113 - val_main_output_loss: 0.5521 - val_aux_output_loss: 0.5924 - val_main_output_acc: 0.7767 - val_aux_output_acc: 0.7282\nEpoch 282/500\n924/924 [==============================] - 0s 220us/step - loss: 0.6495 - main_output_loss: 0.5913 - aux_output_loss: 0.5820 - main_output_acc: 0.6851 - aux_output_acc: 0.7240 - val_loss: 0.6094 - val_main_output_loss: 0.5502 - val_aux_output_loss: 0.5918 - val_main_output_acc: 0.7476 - val_aux_output_acc: 0.7282\nEpoch 283/500\n924/924 [==============================] - 0s 218us/step - loss: 0.6476 - main_output_loss: 0.5895 - aux_output_loss: 0.5817 - main_output_acc: 0.6840 - aux_output_acc: 0.7229 - val_loss: 0.6107 - val_main_output_loss: 0.5514 - val_aux_output_loss: 0.5921 - val_main_output_acc: 0.7864 - val_aux_output_acc: 0.7282\nEpoch 284/500\n924/924 [==============================] - 0s 221us/step - loss: 0.6480 - main_output_loss: 0.5899 - aux_output_loss: 0.5815 - main_output_acc: 0.7056 - aux_output_acc: 0.7219 - val_loss: 0.6082 - val_main_output_loss: 0.5490 - val_aux_output_loss: 0.5913 - val_main_output_acc: 0.7864 - val_aux_output_acc: 0.7282\nEpoch 285/500\n924/924 [==============================] - 0s 216us/step - loss: 0.6491 - main_output_loss: 0.5910 - aux_output_loss: 0.5812 - main_output_acc: 0.6775 - aux_output_acc: 0.7240 - val_loss: 0.6088 - val_main_output_loss: 0.5497 - val_aux_output_loss: 0.5914 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7282\nEpoch 286/500\n924/924 [==============================] - 0s 219us/step - loss: 0.6525 - main_output_loss: 0.5944 - aux_output_loss: 0.5809 - main_output_acc: 0.6926 - aux_output_acc: 0.7240 - val_loss: 0.6082 - val_main_output_loss: 0.5490 - val_aux_output_loss: 0.5913 - val_main_output_acc: 0.7864 - val_aux_output_acc: 0.7282\nEpoch 287/500\n924/924 [==============================] - 0s 217us/step - loss: 0.6425 - main_output_loss: 0.5844 - aux_output_loss: 0.5807 - main_output_acc: 0.7056 - aux_output_acc: 0.7219 - val_loss: 0.6067 - val_main_output_loss: 0.5476 - val_aux_output_loss: 0.5910 - val_main_output_acc: 0.7864 - val_aux_output_acc: 0.7282\nEpoch 288/500\n924/924 [==============================] - 0s 218us/step - loss: 0.6510 - main_output_loss: 0.5930 - aux_output_loss: 0.5804 - main_output_acc: 0.6894 - aux_output_acc: 0.7229 - val_loss: 0.6031 - val_main_output_loss: 0.5441 - val_aux_output_loss: 0.5903 - val_main_output_acc: 0.7573 - val_aux_output_acc: 0.7282\nEpoch 289/500\n924/924 [==============================] - 0s 215us/step - loss: 0.6561 - main_output_loss: 0.5981 - aux_output_loss: 0.5802 - main_output_acc: 0.6591 - aux_output_acc: 0.7208 - val_loss: 0.6022 - val_main_output_loss: 0.5432 - val_aux_output_loss: 0.5899 - val_main_output_acc: 0.7573 - val_aux_output_acc: 0.7282\nEpoch 290/500\n924/924 [==============================] - 0s 222us/step - loss: 0.6553 - main_output_loss: 0.5973 - aux_output_loss: 0.5800 - main_output_acc: 0.6591 - aux_output_acc: 0.7229 - val_loss: 0.6035 - val_main_output_loss: 0.5444 - val_aux_output_loss: 0.5901 - val_main_output_acc: 0.7767 - val_aux_output_acc: 0.7282\nEpoch 291/500\n924/924 [==============================] - 0s 227us/step - loss: 0.6406 - main_output_loss: 0.5826 - aux_output_loss: 0.5799 - main_output_acc: 0.6894 - aux_output_acc: 0.7229 - val_loss: 0.6042 - val_main_output_loss: 0.5452 - val_aux_output_loss: 0.5902 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7282\nEpoch 292/500\n924/924 [==============================] - 0s 233us/step - loss: 0.6467 - main_output_loss: 0.5887 - aux_output_loss: 0.5796 - main_output_acc: 0.6775 - aux_output_acc: 0.7197 - val_loss: 0.6008 - val_main_output_loss: 0.5419 - val_aux_output_loss: 0.5893 - val_main_output_acc: 0.7864 - val_aux_output_acc: 0.7282\nEpoch 293/500\n924/924 [==============================] - 0s 245us/step - loss: 0.6462 - main_output_loss: 0.5883 - aux_output_loss: 0.5794 - main_output_acc: 0.6775 - aux_output_acc: 0.7229 - val_loss: 0.5992 - val_main_output_loss: 0.5404 - val_aux_output_loss: 0.5885 - val_main_output_acc: 0.7476 - val_aux_output_acc: 0.7282\nEpoch 294/500\n924/924 [==============================] - 0s 238us/step - loss: 0.6466 - main_output_loss: 0.5887 - aux_output_loss: 0.5790 - main_output_acc: 0.6840 - aux_output_acc: 0.7240 - val_loss: 0.5980 - val_main_output_loss: 0.5392 - val_aux_output_loss: 0.5880 - val_main_output_acc: 0.7379 - val_aux_output_acc: 0.7184\nEpoch 295/500\n924/924 [==============================] - 0s 241us/step - loss: 0.6497 - main_output_loss: 0.5918 - aux_output_loss: 0.5788 - main_output_acc: 0.6775 - aux_output_acc: 0.7208 - val_loss: 0.5979 - val_main_output_loss: 0.5391 - val_aux_output_loss: 0.5883 - val_main_output_acc: 0.7767 - val_aux_output_acc: 0.7184\nEpoch 296/500\n924/924 [==============================] - 0s 226us/step - loss: 0.6422 - main_output_loss: 0.5843 - aux_output_loss: 0.5786 - main_output_acc: 0.6861 - aux_output_acc: 0.7229 - val_loss: 0.5966 - val_main_output_loss: 0.5378 - val_aux_output_loss: 0.5878 - val_main_output_acc: 0.7864 - val_aux_output_acc: 0.7282\nEpoch 297/500\n924/924 [==============================] - 0s 232us/step - loss: 0.6454 - main_output_loss: 0.5875 - aux_output_loss: 0.5784 - main_output_acc: 0.6905 - aux_output_acc: 0.7219 - val_loss: 0.5956 - val_main_output_loss: 0.5369 - val_aux_output_loss: 0.5878 - val_main_output_acc: 0.7864 - val_aux_output_acc: 0.7184\nEpoch 298/500\n924/924 [==============================] - 0s 225us/step - loss: 0.6391 - main_output_loss: 0.5813 - aux_output_loss: 0.5781 - main_output_acc: 0.6970 - aux_output_acc: 0.7229 - val_loss: 0.5968 - val_main_output_loss: 0.5380 - val_aux_output_loss: 0.5881 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7282\nEpoch 299/500\n924/924 [==============================] - 0s 221us/step - loss: 0.6536 - main_output_loss: 0.5958 - aux_output_loss: 0.5778 - main_output_acc: 0.6721 - aux_output_acc: 0.7240 - val_loss: 0.5937 - val_main_output_loss: 0.5350 - val_aux_output_loss: 0.5869 - val_main_output_acc: 0.7864 - val_aux_output_acc: 0.7282\nEpoch 300/500\n924/924 [==============================] - 0s 213us/step - loss: 0.6409 - main_output_loss: 0.5831 - aux_output_loss: 0.5776 - main_output_acc: 0.6883 - aux_output_acc: 0.7208 - val_loss: 0.5931 - val_main_output_loss: 0.5344 - val_aux_output_loss: 0.5865 - val_main_output_acc: 0.7864 - val_aux_output_acc: 0.7282\nEpoch 301/500\n924/924 [==============================] - 0s 225us/step - loss: 0.6440 - main_output_loss: 0.5863 - aux_output_loss: 0.5774 - main_output_acc: 0.6872 - aux_output_acc: 0.7208 - val_loss: 0.5954 - val_main_output_loss: 0.5367 - val_aux_output_loss: 0.5875 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7184\nEpoch 302/500\n924/924 [==============================] - 0s 217us/step - loss: 0.6370 - main_output_loss: 0.5793 - aux_output_loss: 0.5771 - main_output_acc: 0.6883 - aux_output_acc: 0.7251 - val_loss: 0.5942 - val_main_output_loss: 0.5355 - val_aux_output_loss: 0.5869 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7184\nEpoch 303/500\n924/924 [==============================] - 0s 222us/step - loss: 0.6497 - main_output_loss: 0.5920 - aux_output_loss: 0.5770 - main_output_acc: 0.6916 - aux_output_acc: 0.7229 - val_loss: 0.5939 - val_main_output_loss: 0.5352 - val_aux_output_loss: 0.5871 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7184\nEpoch 304/500\n924/924 [==============================] - 0s 221us/step - loss: 0.6416 - main_output_loss: 0.5839 - aux_output_loss: 0.5768 - main_output_acc: 0.6948 - aux_output_acc: 0.7240 - val_loss: 0.5928 - val_main_output_loss: 0.5341 - val_aux_output_loss: 0.5868 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7184\nEpoch 305/500\n924/924 [==============================] - 0s 218us/step - loss: 0.6454 - main_output_loss: 0.5878 - aux_output_loss: 0.5765 - main_output_acc: 0.6926 - aux_output_acc: 0.7229 - val_loss: 0.5937 - val_main_output_loss: 0.5350 - val_aux_output_loss: 0.5871 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7282\nEpoch 306/500\n924/924 [==============================] - 0s 220us/step - loss: 0.6388 - main_output_loss: 0.5812 - aux_output_loss: 0.5763 - main_output_acc: 0.7056 - aux_output_acc: 0.7240 - val_loss: 0.5926 - val_main_output_loss: 0.5339 - val_aux_output_loss: 0.5866 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7184\nEpoch 307/500\n924/924 [==============================] - 0s 216us/step - loss: 0.6303 - main_output_loss: 0.5727 - aux_output_loss: 0.5760 - main_output_acc: 0.7067 - aux_output_acc: 0.7240 - val_loss: 0.5891 - val_main_output_loss: 0.5305 - val_aux_output_loss: 0.5855 - val_main_output_acc: 0.7864 - val_aux_output_acc: 0.7282\nEpoch 308/500\n924/924 [==============================] - 0s 228us/step - loss: 0.6387 - main_output_loss: 0.5812 - aux_output_loss: 0.5758 - main_output_acc: 0.6905 - aux_output_acc: 0.7208 - val_loss: 0.5893 - val_main_output_loss: 0.5307 - val_aux_output_loss: 0.5858 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7184\nEpoch 309/500\n924/924 [==============================] - 0s 218us/step - loss: 0.6501 - main_output_loss: 0.5925 - aux_output_loss: 0.5756 - main_output_acc: 0.6753 - aux_output_acc: 0.7273 - val_loss: 0.5897 - val_main_output_loss: 0.5311 - val_aux_output_loss: 0.5860 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7184\nEpoch 310/500\n924/924 [==============================] - 0s 221us/step - loss: 0.6310 - main_output_loss: 0.5735 - aux_output_loss: 0.5753 - main_output_acc: 0.6970 - aux_output_acc: 0.7229 - val_loss: 0.5893 - val_main_output_loss: 0.5308 - val_aux_output_loss: 0.5853 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7184\nEpoch 311/500\n924/924 [==============================] - 0s 219us/step - loss: 0.6345 - main_output_loss: 0.5769 - aux_output_loss: 0.5751 - main_output_acc: 0.7056 - aux_output_acc: 0.7251 - val_loss: 0.5895 - val_main_output_loss: 0.5309 - val_aux_output_loss: 0.5853 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7184\nEpoch 312/500\n924/924 [==============================] - 0s 215us/step - loss: 0.6361 - main_output_loss: 0.5786 - aux_output_loss: 0.5749 - main_output_acc: 0.6851 - aux_output_acc: 0.7229 - val_loss: 0.5894 - val_main_output_loss: 0.5309 - val_aux_output_loss: 0.5854 - val_main_output_acc: 0.8155 - val_aux_output_acc: 0.7184\nEpoch 313/500\n924/924 [==============================] - 0s 221us/step - loss: 0.6312 - main_output_loss: 0.5737 - aux_output_loss: 0.5745 - main_output_acc: 0.6894 - aux_output_acc: 0.7262 - val_loss: 0.5885 - val_main_output_loss: 0.5300 - val_aux_output_loss: 0.5851 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7184\nEpoch 314/500\n924/924 [==============================] - 0s 218us/step - loss: 0.6321 - main_output_loss: 0.5747 - aux_output_loss: 0.5744 - main_output_acc: 0.6916 - aux_output_acc: 0.7273 - val_loss: 0.5858 - val_main_output_loss: 0.5274 - val_aux_output_loss: 0.5839 - val_main_output_acc: 0.7864 - val_aux_output_acc: 0.7379\nEpoch 315/500\n924/924 [==============================] - 0s 223us/step - loss: 0.6350 - main_output_loss: 0.5776 - aux_output_loss: 0.5742 - main_output_acc: 0.7035 - aux_output_acc: 0.7219 - val_loss: 0.5846 - val_main_output_loss: 0.5262 - val_aux_output_loss: 0.5838 - val_main_output_acc: 0.7864 - val_aux_output_acc: 0.7379\nEpoch 316/500\n924/924 [==============================] - 0s 217us/step - loss: 0.6343 - main_output_loss: 0.5769 - aux_output_loss: 0.5740 - main_output_acc: 0.6926 - aux_output_acc: 0.7219 - val_loss: 0.5844 - val_main_output_loss: 0.5260 - val_aux_output_loss: 0.5838 - val_main_output_acc: 0.7864 - val_aux_output_acc: 0.7282\nEpoch 317/500\n924/924 [==============================] - 0s 222us/step - loss: 0.6339 - main_output_loss: 0.5766 - aux_output_loss: 0.5736 - main_output_acc: 0.7024 - aux_output_acc: 0.7229 - val_loss: 0.5839 - val_main_output_loss: 0.5255 - val_aux_output_loss: 0.5839 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7184\nEpoch 318/500\n924/924 [==============================] - 0s 227us/step - loss: 0.6418 - main_output_loss: 0.5844 - aux_output_loss: 0.5734 - main_output_acc: 0.6818 - aux_output_acc: 0.7240 - val_loss: 0.5840 - val_main_output_loss: 0.5256 - val_aux_output_loss: 0.5834 - val_main_output_acc: 0.8155 - val_aux_output_acc: 0.7184\nEpoch 319/500\n924/924 [==============================] - 0s 217us/step - loss: 0.6408 - main_output_loss: 0.5835 - aux_output_loss: 0.5732 - main_output_acc: 0.6710 - aux_output_acc: 0.7240 - val_loss: 0.5858 - val_main_output_loss: 0.5274 - val_aux_output_loss: 0.5837 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7184\nEpoch 320/500\n924/924 [==============================] - 0s 225us/step - loss: 0.6321 - main_output_loss: 0.5748 - aux_output_loss: 0.5730 - main_output_acc: 0.6861 - aux_output_acc: 0.7251 - val_loss: 0.5834 - val_main_output_loss: 0.5251 - val_aux_output_loss: 0.5826 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7379\nEpoch 321/500\n924/924 [==============================] - 0s 218us/step - loss: 0.6347 - main_output_loss: 0.5774 - aux_output_loss: 0.5728 - main_output_acc: 0.7013 - aux_output_acc: 0.7240 - val_loss: 0.5829 - val_main_output_loss: 0.5247 - val_aux_output_loss: 0.5822 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7379\nEpoch 322/500\n924/924 [==============================] - 0s 220us/step - loss: 0.6320 - main_output_loss: 0.5747 - aux_output_loss: 0.5726 - main_output_acc: 0.6970 - aux_output_acc: 0.7240 - val_loss: 0.5824 - val_main_output_loss: 0.5242 - val_aux_output_loss: 0.5820 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7379\nEpoch 323/500\n924/924 [==============================] - 0s 221us/step - loss: 0.6363 - main_output_loss: 0.5790 - aux_output_loss: 0.5723 - main_output_acc: 0.6861 - aux_output_acc: 0.7229 - val_loss: 0.5819 - val_main_output_loss: 0.5237 - val_aux_output_loss: 0.5820 - val_main_output_acc: 0.7864 - val_aux_output_acc: 0.7379\nEpoch 324/500\n924/924 [==============================] - 0s 218us/step - loss: 0.6297 - main_output_loss: 0.5725 - aux_output_loss: 0.5720 - main_output_acc: 0.6959 - aux_output_acc: 0.7251 - val_loss: 0.5827 - val_main_output_loss: 0.5244 - val_aux_output_loss: 0.5823 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7184\nEpoch 325/500\n924/924 [==============================] - 0s 226us/step - loss: 0.6381 - main_output_loss: 0.5809 - aux_output_loss: 0.5718 - main_output_acc: 0.6861 - aux_output_acc: 0.7240 - val_loss: 0.5849 - val_main_output_loss: 0.5266 - val_aux_output_loss: 0.5824 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7184\nEpoch 326/500\n924/924 [==============================] - 0s 215us/step - loss: 0.6336 - main_output_loss: 0.5764 - aux_output_loss: 0.5715 - main_output_acc: 0.6959 - aux_output_acc: 0.7262 - val_loss: 0.5808 - val_main_output_loss: 0.5227 - val_aux_output_loss: 0.5810 - val_main_output_acc: 0.7864 - val_aux_output_acc: 0.7379\nEpoch 327/500\n924/924 [==============================] - 0s 220us/step - loss: 0.6293 - main_output_loss: 0.5721 - aux_output_loss: 0.5713 - main_output_acc: 0.6937 - aux_output_acc: 0.7262 - val_loss: 0.5819 - val_main_output_loss: 0.5238 - val_aux_output_loss: 0.5812 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7282\nEpoch 328/500\n924/924 [==============================] - 0s 216us/step - loss: 0.6205 - main_output_loss: 0.5634 - aux_output_loss: 0.5711 - main_output_acc: 0.7045 - aux_output_acc: 0.7273 - val_loss: 0.5819 - val_main_output_loss: 0.5238 - val_aux_output_loss: 0.5812 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7184\nEpoch 329/500\n924/924 [==============================] - 0s 214us/step - loss: 0.6180 - main_output_loss: 0.5609 - aux_output_loss: 0.5708 - main_output_acc: 0.7100 - aux_output_acc: 0.7294 - val_loss: 0.5793 - val_main_output_loss: 0.5213 - val_aux_output_loss: 0.5801 - val_main_output_acc: 0.7767 - val_aux_output_acc: 0.7379\nEpoch 330/500\n924/924 [==============================] - 0s 218us/step - loss: 0.6276 - main_output_loss: 0.5705 - aux_output_loss: 0.5709 - main_output_acc: 0.6894 - aux_output_acc: 0.7251 - val_loss: 0.5788 - val_main_output_loss: 0.5208 - val_aux_output_loss: 0.5802 - val_main_output_acc: 0.8155 - val_aux_output_acc: 0.7282\nEpoch 331/500\n924/924 [==============================] - 0s 219us/step - loss: 0.6304 - main_output_loss: 0.5733 - aux_output_loss: 0.5704 - main_output_acc: 0.6937 - aux_output_acc: 0.7284 - val_loss: 0.5810 - val_main_output_loss: 0.5229 - val_aux_output_loss: 0.5807 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7184\nEpoch 332/500\n924/924 [==============================] - 0s 218us/step - loss: 0.6303 - main_output_loss: 0.5733 - aux_output_loss: 0.5702 - main_output_acc: 0.6916 - aux_output_acc: 0.7284 - val_loss: 0.5781 - val_main_output_loss: 0.5202 - val_aux_output_loss: 0.5797 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7379\nEpoch 333/500\n924/924 [==============================] - 0s 217us/step - loss: 0.6227 - main_output_loss: 0.5657 - aux_output_loss: 0.5699 - main_output_acc: 0.6959 - aux_output_acc: 0.7284 - val_loss: 0.5781 - val_main_output_loss: 0.5201 - val_aux_output_loss: 0.5797 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7282\nEpoch 334/500\n924/924 [==============================] - 0s 222us/step - loss: 0.6338 - main_output_loss: 0.5768 - aux_output_loss: 0.5697 - main_output_acc: 0.6916 - aux_output_acc: 0.7294 - val_loss: 0.5780 - val_main_output_loss: 0.5199 - val_aux_output_loss: 0.5801 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7184\nEpoch 335/500\n924/924 [==============================] - 0s 221us/step - loss: 0.6238 - main_output_loss: 0.5669 - aux_output_loss: 0.5696 - main_output_acc: 0.7132 - aux_output_acc: 0.7262 - val_loss: 0.5757 - val_main_output_loss: 0.5178 - val_aux_output_loss: 0.5791 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7282\nEpoch 336/500\n924/924 [==============================] - 0s 222us/step - loss: 0.6287 - main_output_loss: 0.5718 - aux_output_loss: 0.5693 - main_output_acc: 0.6818 - aux_output_acc: 0.7273 - val_loss: 0.5753 - val_main_output_loss: 0.5174 - val_aux_output_loss: 0.5787 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7379\nEpoch 337/500\n924/924 [==============================] - 0s 218us/step - loss: 0.6261 - main_output_loss: 0.5692 - aux_output_loss: 0.5692 - main_output_acc: 0.7089 - aux_output_acc: 0.7294 - val_loss: 0.5754 - val_main_output_loss: 0.5175 - val_aux_output_loss: 0.5787 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7379\nEpoch 338/500\n924/924 [==============================] - 0s 220us/step - loss: 0.6206 - main_output_loss: 0.5637 - aux_output_loss: 0.5687 - main_output_acc: 0.7132 - aux_output_acc: 0.7273 - val_loss: 0.5769 - val_main_output_loss: 0.5190 - val_aux_output_loss: 0.5789 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7282\nEpoch 339/500\n924/924 [==============================] - 0s 220us/step - loss: 0.6151 - main_output_loss: 0.5582 - aux_output_loss: 0.5685 - main_output_acc: 0.7067 - aux_output_acc: 0.7273 - val_loss: 0.5758 - val_main_output_loss: 0.5179 - val_aux_output_loss: 0.5784 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7282\nEpoch 340/500\n924/924 [==============================] - 0s 221us/step - loss: 0.6251 - main_output_loss: 0.5683 - aux_output_loss: 0.5683 - main_output_acc: 0.7013 - aux_output_acc: 0.7294 - val_loss: 0.5729 - val_main_output_loss: 0.5152 - val_aux_output_loss: 0.5775 - val_main_output_acc: 0.7864 - val_aux_output_acc: 0.7476\nEpoch 341/500\n924/924 [==============================] - 0s 221us/step - loss: 0.6163 - main_output_loss: 0.5595 - aux_output_loss: 0.5681 - main_output_acc: 0.6959 - aux_output_acc: 0.7316 - val_loss: 0.5729 - val_main_output_loss: 0.5152 - val_aux_output_loss: 0.5775 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7476\nEpoch 342/500\n924/924 [==============================] - 0s 216us/step - loss: 0.6196 - main_output_loss: 0.5628 - aux_output_loss: 0.5678 - main_output_acc: 0.6829 - aux_output_acc: 0.7305 - val_loss: 0.5743 - val_main_output_loss: 0.5165 - val_aux_output_loss: 0.5781 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7379\nEpoch 343/500\n924/924 [==============================] - 0s 223us/step - loss: 0.6054 - main_output_loss: 0.5486 - aux_output_loss: 0.5677 - main_output_acc: 0.7251 - aux_output_acc: 0.7305 - val_loss: 0.5709 - val_main_output_loss: 0.5132 - val_aux_output_loss: 0.5769 - val_main_output_acc: 0.8155 - val_aux_output_acc: 0.7476\nEpoch 344/500\n924/924 [==============================] - 0s 214us/step - loss: 0.6144 - main_output_loss: 0.5576 - aux_output_loss: 0.5677 - main_output_acc: 0.7143 - aux_output_acc: 0.7294 - val_loss: 0.5713 - val_main_output_loss: 0.5136 - val_aux_output_loss: 0.5769 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7476\nEpoch 345/500\n924/924 [==============================] - 0s 234us/step - loss: 0.6128 - main_output_loss: 0.5561 - aux_output_loss: 0.5672 - main_output_acc: 0.7219 - aux_output_acc: 0.7316 - val_loss: 0.5698 - val_main_output_loss: 0.5122 - val_aux_output_loss: 0.5760 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7476\nEpoch 346/500\n924/924 [==============================] - 0s 221us/step - loss: 0.6080 - main_output_loss: 0.5513 - aux_output_loss: 0.5670 - main_output_acc: 0.7089 - aux_output_acc: 0.7273 - val_loss: 0.5711 - val_main_output_loss: 0.5135 - val_aux_output_loss: 0.5762 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7476\nEpoch 347/500\n924/924 [==============================] - 0s 223us/step - loss: 0.6236 - main_output_loss: 0.5669 - aux_output_loss: 0.5667 - main_output_acc: 0.7067 - aux_output_acc: 0.7294 - val_loss: 0.5700 - val_main_output_loss: 0.5125 - val_aux_output_loss: 0.5756 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7476\nEpoch 348/500\n924/924 [==============================] - 0s 223us/step - loss: 0.6184 - main_output_loss: 0.5617 - aux_output_loss: 0.5668 - main_output_acc: 0.7089 - aux_output_acc: 0.7262 - val_loss: 0.5705 - val_main_output_loss: 0.5129 - val_aux_output_loss: 0.5758 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7476\nEpoch 349/500\n924/924 [==============================] - 0s 218us/step - loss: 0.6357 - main_output_loss: 0.5791 - aux_output_loss: 0.5663 - main_output_acc: 0.6797 - aux_output_acc: 0.7305 - val_loss: 0.5697 - val_main_output_loss: 0.5122 - val_aux_output_loss: 0.5754 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7476\nEpoch 350/500\n924/924 [==============================] - 0s 222us/step - loss: 0.6182 - main_output_loss: 0.5616 - aux_output_loss: 0.5660 - main_output_acc: 0.7045 - aux_output_acc: 0.7305 - val_loss: 0.5680 - val_main_output_loss: 0.5105 - val_aux_output_loss: 0.5747 - val_main_output_acc: 0.7864 - val_aux_output_acc: 0.7476\nEpoch 351/500\n924/924 [==============================] - 0s 223us/step - loss: 0.6146 - main_output_loss: 0.5581 - aux_output_loss: 0.5658 - main_output_acc: 0.7132 - aux_output_acc: 0.7240 - val_loss: 0.5698 - val_main_output_loss: 0.5123 - val_aux_output_loss: 0.5754 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7379\nEpoch 352/500\n924/924 [==============================] - 0s 232us/step - loss: 0.6105 - main_output_loss: 0.5540 - aux_output_loss: 0.5657 - main_output_acc: 0.7143 - aux_output_acc: 0.7316 - val_loss: 0.5699 - val_main_output_loss: 0.5124 - val_aux_output_loss: 0.5749 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7476\nEpoch 353/500\n924/924 [==============================] - 0s 230us/step - loss: 0.6082 - main_output_loss: 0.5516 - aux_output_loss: 0.5654 - main_output_acc: 0.7067 - aux_output_acc: 0.7294 - val_loss: 0.5690 - val_main_output_loss: 0.5115 - val_aux_output_loss: 0.5746 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7476\nEpoch 354/500\n924/924 [==============================] - 0s 225us/step - loss: 0.6118 - main_output_loss: 0.5553 - aux_output_loss: 0.5651 - main_output_acc: 0.7165 - aux_output_acc: 0.7316 - val_loss: 0.5677 - val_main_output_loss: 0.5103 - val_aux_output_loss: 0.5741 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7476\nEpoch 355/500\n924/924 [==============================] - 0s 229us/step - loss: 0.6099 - main_output_loss: 0.5534 - aux_output_loss: 0.5650 - main_output_acc: 0.7154 - aux_output_acc: 0.7294 - val_loss: 0.5679 - val_main_output_loss: 0.5104 - val_aux_output_loss: 0.5743 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7476\nEpoch 356/500\n924/924 [==============================] - 0s 238us/step - loss: 0.6060 - main_output_loss: 0.5496 - aux_output_loss: 0.5645 - main_output_acc: 0.7175 - aux_output_acc: 0.7305 - val_loss: 0.5681 - val_main_output_loss: 0.5107 - val_aux_output_loss: 0.5740 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7476\nEpoch 357/500\n924/924 [==============================] - 0s 260us/step - loss: 0.6138 - main_output_loss: 0.5574 - aux_output_loss: 0.5644 - main_output_acc: 0.7056 - aux_output_acc: 0.7316 - val_loss: 0.5657 - val_main_output_loss: 0.5084 - val_aux_output_loss: 0.5734 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7476\nEpoch 358/500\n924/924 [==============================] - 0s 229us/step - loss: 0.6112 - main_output_loss: 0.5548 - aux_output_loss: 0.5642 - main_output_acc: 0.6991 - aux_output_acc: 0.7294 - val_loss: 0.5657 - val_main_output_loss: 0.5084 - val_aux_output_loss: 0.5734 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7476\nEpoch 359/500\n924/924 [==============================] - 0s 229us/step - loss: 0.6131 - main_output_loss: 0.5567 - aux_output_loss: 0.5640 - main_output_acc: 0.7132 - aux_output_acc: 0.7316 - val_loss: 0.5643 - val_main_output_loss: 0.5070 - val_aux_output_loss: 0.5731 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7476\nEpoch 360/500\n924/924 [==============================] - 0s 218us/step - loss: 0.6147 - main_output_loss: 0.5583 - aux_output_loss: 0.5636 - main_output_acc: 0.7175 - aux_output_acc: 0.7316 - val_loss: 0.5642 - val_main_output_loss: 0.5070 - val_aux_output_loss: 0.5726 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7476\nEpoch 361/500\n924/924 [==============================] - 0s 220us/step - loss: 0.6123 - main_output_loss: 0.5559 - aux_output_loss: 0.5635 - main_output_acc: 0.7165 - aux_output_acc: 0.7284 - val_loss: 0.5627 - val_main_output_loss: 0.5055 - val_aux_output_loss: 0.5721 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7379\nEpoch 362/500\n924/924 [==============================] - 0s 232us/step - loss: 0.6025 - main_output_loss: 0.5461 - aux_output_loss: 0.5633 - main_output_acc: 0.7229 - aux_output_acc: 0.7273 - val_loss: 0.5644 - val_main_output_loss: 0.5072 - val_aux_output_loss: 0.5723 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7476\nEpoch 363/500\n924/924 [==============================] - 0s 223us/step - loss: 0.6113 - main_output_loss: 0.5550 - aux_output_loss: 0.5631 - main_output_acc: 0.7002 - aux_output_acc: 0.7294 - val_loss: 0.5628 - val_main_output_loss: 0.5057 - val_aux_output_loss: 0.5713 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7379\nEpoch 364/500\n924/924 [==============================] - 0s 217us/step - loss: 0.6162 - main_output_loss: 0.5599 - aux_output_loss: 0.5629 - main_output_acc: 0.6970 - aux_output_acc: 0.7273 - val_loss: 0.5641 - val_main_output_loss: 0.5069 - val_aux_output_loss: 0.5717 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7379\nEpoch 365/500\n924/924 [==============================] - 0s 220us/step - loss: 0.6045 - main_output_loss: 0.5482 - aux_output_loss: 0.5626 - main_output_acc: 0.7262 - aux_output_acc: 0.7316 - val_loss: 0.5613 - val_main_output_loss: 0.5042 - val_aux_output_loss: 0.5710 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7379\nEpoch 366/500\n924/924 [==============================] - 0s 219us/step - loss: 0.6020 - main_output_loss: 0.5457 - aux_output_loss: 0.5623 - main_output_acc: 0.7251 - aux_output_acc: 0.7305 - val_loss: 0.5599 - val_main_output_loss: 0.5029 - val_aux_output_loss: 0.5702 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7476\nEpoch 367/500\n924/924 [==============================] - 0s 221us/step - loss: 0.6100 - main_output_loss: 0.5538 - aux_output_loss: 0.5625 - main_output_acc: 0.7110 - aux_output_acc: 0.7305 - val_loss: 0.5604 - val_main_output_loss: 0.5034 - val_aux_output_loss: 0.5702 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7476\nEpoch 368/500\n924/924 [==============================] - 0s 216us/step - loss: 0.6048 - main_output_loss: 0.5486 - aux_output_loss: 0.5621 - main_output_acc: 0.7121 - aux_output_acc: 0.7305 - val_loss: 0.5597 - val_main_output_loss: 0.5028 - val_aux_output_loss: 0.5698 - val_main_output_acc: 0.7864 - val_aux_output_acc: 0.7476\nEpoch 369/500\n924/924 [==============================] - 0s 217us/step - loss: 0.5927 - main_output_loss: 0.5366 - aux_output_loss: 0.5619 - main_output_acc: 0.7229 - aux_output_acc: 0.7273 - val_loss: 0.5592 - val_main_output_loss: 0.5023 - val_aux_output_loss: 0.5691 - val_main_output_acc: 0.7864 - val_aux_output_acc: 0.7476\nEpoch 370/500\n924/924 [==============================] - 0s 217us/step - loss: 0.6032 - main_output_loss: 0.5470 - aux_output_loss: 0.5619 - main_output_acc: 0.7262 - aux_output_acc: 0.7338 - val_loss: 0.5589 - val_main_output_loss: 0.5020 - val_aux_output_loss: 0.5688 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7476\nEpoch 371/500\n924/924 [==============================] - 0s 222us/step - loss: 0.6096 - main_output_loss: 0.5534 - aux_output_loss: 0.5617 - main_output_acc: 0.7100 - aux_output_acc: 0.7348 - val_loss: 0.5592 - val_main_output_loss: 0.5023 - val_aux_output_loss: 0.5688 - val_main_output_acc: 0.7864 - val_aux_output_acc: 0.7476\nEpoch 372/500\n924/924 [==============================] - 0s 222us/step - loss: 0.6027 - main_output_loss: 0.5466 - aux_output_loss: 0.5614 - main_output_acc: 0.7208 - aux_output_acc: 0.7316 - val_loss: 0.5602 - val_main_output_loss: 0.5033 - val_aux_output_loss: 0.5691 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7476\nEpoch 373/500\n924/924 [==============================] - 0s 217us/step - loss: 0.6062 - main_output_loss: 0.5500 - aux_output_loss: 0.5613 - main_output_acc: 0.7208 - aux_output_acc: 0.7316 - val_loss: 0.5599 - val_main_output_loss: 0.5030 - val_aux_output_loss: 0.5687 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7476\nEpoch 374/500\n924/924 [==============================] - 0s 218us/step - loss: 0.6056 - main_output_loss: 0.5495 - aux_output_loss: 0.5610 - main_output_acc: 0.7240 - aux_output_acc: 0.7294 - val_loss: 0.5587 - val_main_output_loss: 0.5018 - val_aux_output_loss: 0.5684 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7476\nEpoch 375/500\n924/924 [==============================] - 0s 218us/step - loss: 0.5993 - main_output_loss: 0.5433 - aux_output_loss: 0.5606 - main_output_acc: 0.7143 - aux_output_acc: 0.7327 - val_loss: 0.5602 - val_main_output_loss: 0.5034 - val_aux_output_loss: 0.5684 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7476\nEpoch 376/500\n924/924 [==============================] - 0s 220us/step - loss: 0.5928 - main_output_loss: 0.5367 - aux_output_loss: 0.5606 - main_output_acc: 0.7392 - aux_output_acc: 0.7316 - val_loss: 0.5591 - val_main_output_loss: 0.5022 - val_aux_output_loss: 0.5683 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7476\nEpoch 377/500\n924/924 [==============================] - 0s 220us/step - loss: 0.6073 - main_output_loss: 0.5512 - aux_output_loss: 0.5605 - main_output_acc: 0.7229 - aux_output_acc: 0.7327 - val_loss: 0.5571 - val_main_output_loss: 0.5003 - val_aux_output_loss: 0.5680 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7476\nEpoch 378/500\n924/924 [==============================] - 0s 229us/step - loss: 0.6008 - main_output_loss: 0.5448 - aux_output_loss: 0.5600 - main_output_acc: 0.7316 - aux_output_acc: 0.7316 - val_loss: 0.5575 - val_main_output_loss: 0.5007 - val_aux_output_loss: 0.5679 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7476\nEpoch 379/500\n924/924 [==============================] - 0s 222us/step - loss: 0.5936 - main_output_loss: 0.5376 - aux_output_loss: 0.5596 - main_output_acc: 0.7327 - aux_output_acc: 0.7305 - val_loss: 0.5550 - val_main_output_loss: 0.4982 - val_aux_output_loss: 0.5675 - val_main_output_acc: 0.7864 - val_aux_output_acc: 0.7476\nEpoch 380/500\n924/924 [==============================] - 0s 216us/step - loss: 0.5933 - main_output_loss: 0.5374 - aux_output_loss: 0.5595 - main_output_acc: 0.7294 - aux_output_acc: 0.7327 - val_loss: 0.5570 - val_main_output_loss: 0.5002 - val_aux_output_loss: 0.5680 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7476\nEpoch 381/500\n924/924 [==============================] - 0s 218us/step - loss: 0.6046 - main_output_loss: 0.5486 - aux_output_loss: 0.5595 - main_output_acc: 0.7262 - aux_output_acc: 0.7294 - val_loss: 0.5568 - val_main_output_loss: 0.5001 - val_aux_output_loss: 0.5677 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7476\nEpoch 382/500\n924/924 [==============================] - 0s 221us/step - loss: 0.6046 - main_output_loss: 0.5487 - aux_output_loss: 0.5591 - main_output_acc: 0.7175 - aux_output_acc: 0.7338 - val_loss: 0.5552 - val_main_output_loss: 0.4985 - val_aux_output_loss: 0.5672 - val_main_output_acc: 0.7864 - val_aux_output_acc: 0.7476\nEpoch 383/500\n924/924 [==============================] - 0s 225us/step - loss: 0.5991 - main_output_loss: 0.5433 - aux_output_loss: 0.5588 - main_output_acc: 0.7251 - aux_output_acc: 0.7327 - val_loss: 0.5570 - val_main_output_loss: 0.5003 - val_aux_output_loss: 0.5677 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7573\nEpoch 384/500\n924/924 [==============================] - 0s 224us/step - loss: 0.6103 - main_output_loss: 0.5545 - aux_output_loss: 0.5587 - main_output_acc: 0.7013 - aux_output_acc: 0.7316 - val_loss: 0.5559 - val_main_output_loss: 0.4992 - val_aux_output_loss: 0.5673 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7573\nEpoch 385/500\n924/924 [==============================] - 0s 217us/step - loss: 0.5881 - main_output_loss: 0.5323 - aux_output_loss: 0.5584 - main_output_acc: 0.7359 - aux_output_acc: 0.7327 - val_loss: 0.5533 - val_main_output_loss: 0.4966 - val_aux_output_loss: 0.5664 - val_main_output_acc: 0.7864 - val_aux_output_acc: 0.7476\nEpoch 386/500\n924/924 [==============================] - 0s 225us/step - loss: 0.5955 - main_output_loss: 0.5397 - aux_output_loss: 0.5582 - main_output_acc: 0.7186 - aux_output_acc: 0.7348 - val_loss: 0.5531 - val_main_output_loss: 0.4965 - val_aux_output_loss: 0.5662 - val_main_output_acc: 0.7864 - val_aux_output_acc: 0.7476\nEpoch 387/500\n924/924 [==============================] - 0s 225us/step - loss: 0.5989 - main_output_loss: 0.5431 - aux_output_loss: 0.5579 - main_output_acc: 0.7348 - aux_output_acc: 0.7348 - val_loss: 0.5512 - val_main_output_loss: 0.4947 - val_aux_output_loss: 0.5655 - val_main_output_acc: 0.7864 - val_aux_output_acc: 0.7476\nEpoch 388/500\n924/924 [==============================] - 0s 224us/step - loss: 0.5958 - main_output_loss: 0.5399 - aux_output_loss: 0.5581 - main_output_acc: 0.7273 - aux_output_acc: 0.7316 - val_loss: 0.5505 - val_main_output_loss: 0.4940 - val_aux_output_loss: 0.5652 - val_main_output_acc: 0.7864 - val_aux_output_acc: 0.7476\nEpoch 389/500\n924/924 [==============================] - 0s 220us/step - loss: 0.5924 - main_output_loss: 0.5366 - aux_output_loss: 0.5577 - main_output_acc: 0.7262 - aux_output_acc: 0.7316 - val_loss: 0.5504 - val_main_output_loss: 0.4939 - val_aux_output_loss: 0.5652 - val_main_output_acc: 0.7864 - val_aux_output_acc: 0.7476\nEpoch 390/500\n924/924 [==============================] - 0s 232us/step - loss: 0.6039 - main_output_loss: 0.5481 - aux_output_loss: 0.5574 - main_output_acc: 0.7089 - aux_output_acc: 0.7327 - val_loss: 0.5522 - val_main_output_loss: 0.4956 - val_aux_output_loss: 0.5660 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7573\nEpoch 391/500\n924/924 [==============================] - 0s 222us/step - loss: 0.5939 - main_output_loss: 0.5382 - aux_output_loss: 0.5572 - main_output_acc: 0.7132 - aux_output_acc: 0.7338 - val_loss: 0.5493 - val_main_output_loss: 0.4928 - val_aux_output_loss: 0.5652 - val_main_output_acc: 0.7864 - val_aux_output_acc: 0.7476\nEpoch 392/500\n924/924 [==============================] - 0s 218us/step - loss: 0.6023 - main_output_loss: 0.5466 - aux_output_loss: 0.5571 - main_output_acc: 0.7143 - aux_output_acc: 0.7327 - val_loss: 0.5483 - val_main_output_loss: 0.4918 - val_aux_output_loss: 0.5649 - val_main_output_acc: 0.7767 - val_aux_output_acc: 0.7476\nEpoch 393/500\n924/924 [==============================] - 0s 222us/step - loss: 0.5884 - main_output_loss: 0.5327 - aux_output_loss: 0.5568 - main_output_acc: 0.7251 - aux_output_acc: 0.7294 - val_loss: 0.5486 - val_main_output_loss: 0.4921 - val_aux_output_loss: 0.5647 - val_main_output_acc: 0.7767 - val_aux_output_acc: 0.7476\nEpoch 394/500\n924/924 [==============================] - 0s 220us/step - loss: 0.5904 - main_output_loss: 0.5347 - aux_output_loss: 0.5565 - main_output_acc: 0.7229 - aux_output_acc: 0.7359 - val_loss: 0.5502 - val_main_output_loss: 0.4937 - val_aux_output_loss: 0.5650 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7476\nEpoch 395/500\n924/924 [==============================] - 0s 218us/step - loss: 0.6023 - main_output_loss: 0.5467 - aux_output_loss: 0.5564 - main_output_acc: 0.7165 - aux_output_acc: 0.7316 - val_loss: 0.5510 - val_main_output_loss: 0.4945 - val_aux_output_loss: 0.5653 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7476\nEpoch 396/500\n924/924 [==============================] - 0s 223us/step - loss: 0.5821 - main_output_loss: 0.5265 - aux_output_loss: 0.5562 - main_output_acc: 0.7381 - aux_output_acc: 0.7348 - val_loss: 0.5514 - val_main_output_loss: 0.4948 - val_aux_output_loss: 0.5653 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7476\nEpoch 397/500\n924/924 [==============================] - 0s 223us/step - loss: 0.5887 - main_output_loss: 0.5331 - aux_output_loss: 0.5559 - main_output_acc: 0.7316 - aux_output_acc: 0.7348 - val_loss: 0.5492 - val_main_output_loss: 0.4928 - val_aux_output_loss: 0.5645 - val_main_output_acc: 0.7864 - val_aux_output_acc: 0.7476\nEpoch 398/500\n924/924 [==============================] - 0s 219us/step - loss: 0.5826 - main_output_loss: 0.5270 - aux_output_loss: 0.5558 - main_output_acc: 0.7284 - aux_output_acc: 0.7338 - val_loss: 0.5503 - val_main_output_loss: 0.4939 - val_aux_output_loss: 0.5648 - val_main_output_acc: 0.7864 - val_aux_output_acc: 0.7476\nEpoch 399/500\n924/924 [==============================] - 0s 218us/step - loss: 0.5912 - main_output_loss: 0.5356 - aux_output_loss: 0.5556 - main_output_acc: 0.7240 - aux_output_acc: 0.7359 - val_loss: 0.5480 - val_main_output_loss: 0.4916 - val_aux_output_loss: 0.5640 - val_main_output_acc: 0.7864 - val_aux_output_acc: 0.7476\nEpoch 400/500\n924/924 [==============================] - 0s 217us/step - loss: 0.5885 - main_output_loss: 0.5329 - aux_output_loss: 0.5555 - main_output_acc: 0.7262 - aux_output_acc: 0.7359 - val_loss: 0.5472 - val_main_output_loss: 0.4909 - val_aux_output_loss: 0.5633 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7476\nEpoch 401/500\n924/924 [==============================] - 0s 227us/step - loss: 0.5819 - main_output_loss: 0.5264 - aux_output_loss: 0.5552 - main_output_acc: 0.7284 - aux_output_acc: 0.7338 - val_loss: 0.5477 - val_main_output_loss: 0.4913 - val_aux_output_loss: 0.5638 - val_main_output_acc: 0.7864 - val_aux_output_acc: 0.7476\nEpoch 402/500\n924/924 [==============================] - 0s 216us/step - loss: 0.5845 - main_output_loss: 0.5290 - aux_output_loss: 0.5550 - main_output_acc: 0.7370 - aux_output_acc: 0.7348 - val_loss: 0.5476 - val_main_output_loss: 0.4912 - val_aux_output_loss: 0.5635 - val_main_output_acc: 0.7864 - val_aux_output_acc: 0.7476\nEpoch 403/500\n924/924 [==============================] - 0s 219us/step - loss: 0.5818 - main_output_loss: 0.5263 - aux_output_loss: 0.5547 - main_output_acc: 0.7392 - aux_output_acc: 0.7348 - val_loss: 0.5479 - val_main_output_loss: 0.4915 - val_aux_output_loss: 0.5638 - val_main_output_acc: 0.7864 - val_aux_output_acc: 0.7573\nEpoch 404/500\n924/924 [==============================] - 0s 217us/step - loss: 0.5980 - main_output_loss: 0.5426 - aux_output_loss: 0.5546 - main_output_acc: 0.7305 - aux_output_acc: 0.7359 - val_loss: 0.5463 - val_main_output_loss: 0.4900 - val_aux_output_loss: 0.5629 - val_main_output_acc: 0.7864 - val_aux_output_acc: 0.7476\nEpoch 405/500\n924/924 [==============================] - 0s 213us/step - loss: 0.5836 - main_output_loss: 0.5282 - aux_output_loss: 0.5543 - main_output_acc: 0.7413 - aux_output_acc: 0.7370 - val_loss: 0.5461 - val_main_output_loss: 0.4898 - val_aux_output_loss: 0.5626 - val_main_output_acc: 0.7864 - val_aux_output_acc: 0.7476\nEpoch 406/500\n924/924 [==============================] - 0s 219us/step - loss: 0.5794 - main_output_loss: 0.5240 - aux_output_loss: 0.5541 - main_output_acc: 0.7348 - aux_output_acc: 0.7370 - val_loss: 0.5460 - val_main_output_loss: 0.4897 - val_aux_output_loss: 0.5626 - val_main_output_acc: 0.7864 - val_aux_output_acc: 0.7476\nEpoch 407/500\n924/924 [==============================] - 0s 216us/step - loss: 0.5995 - main_output_loss: 0.5441 - aux_output_loss: 0.5539 - main_output_acc: 0.7208 - aux_output_acc: 0.7370 - val_loss: 0.5456 - val_main_output_loss: 0.4894 - val_aux_output_loss: 0.5626 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7476\nEpoch 408/500\n924/924 [==============================] - 0s 216us/step - loss: 0.5870 - main_output_loss: 0.5317 - aux_output_loss: 0.5535 - main_output_acc: 0.7381 - aux_output_acc: 0.7359 - val_loss: 0.5472 - val_main_output_loss: 0.4909 - val_aux_output_loss: 0.5629 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7476\nEpoch 409/500\n924/924 [==============================] - 0s 221us/step - loss: 0.5706 - main_output_loss: 0.5153 - aux_output_loss: 0.5533 - main_output_acc: 0.7403 - aux_output_acc: 0.7381 - val_loss: 0.5481 - val_main_output_loss: 0.4918 - val_aux_output_loss: 0.5629 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7476\nEpoch 410/500\n924/924 [==============================] - 0s 222us/step - loss: 0.5881 - main_output_loss: 0.5328 - aux_output_loss: 0.5531 - main_output_acc: 0.7338 - aux_output_acc: 0.7359 - val_loss: 0.5432 - val_main_output_loss: 0.4871 - val_aux_output_loss: 0.5614 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7476\nEpoch 411/500\n924/924 [==============================] - 0s 219us/step - loss: 0.5757 - main_output_loss: 0.5204 - aux_output_loss: 0.5531 - main_output_acc: 0.7348 - aux_output_acc: 0.7370 - val_loss: 0.5431 - val_main_output_loss: 0.4870 - val_aux_output_loss: 0.5613 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7476\nEpoch 412/500\n924/924 [==============================] - 0s 218us/step - loss: 0.5816 - main_output_loss: 0.5263 - aux_output_loss: 0.5527 - main_output_acc: 0.7327 - aux_output_acc: 0.7392 - val_loss: 0.5483 - val_main_output_loss: 0.4920 - val_aux_output_loss: 0.5626 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7476\nEpoch 413/500\n924/924 [==============================] - 0s 223us/step - loss: 0.5819 - main_output_loss: 0.5266 - aux_output_loss: 0.5527 - main_output_acc: 0.7327 - aux_output_acc: 0.7370 - val_loss: 0.5504 - val_main_output_loss: 0.4941 - val_aux_output_loss: 0.5628 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7476\nEpoch 414/500\n924/924 [==============================] - 0s 224us/step - loss: 0.5867 - main_output_loss: 0.5314 - aux_output_loss: 0.5527 - main_output_acc: 0.7348 - aux_output_acc: 0.7381 - val_loss: 0.5494 - val_main_output_loss: 0.4931 - val_aux_output_loss: 0.5629 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7476\nEpoch 415/500\n924/924 [==============================] - 0s 219us/step - loss: 0.5813 - main_output_loss: 0.5261 - aux_output_loss: 0.5520 - main_output_acc: 0.7446 - aux_output_acc: 0.7381 - val_loss: 0.5431 - val_main_output_loss: 0.4869 - val_aux_output_loss: 0.5614 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7476\nEpoch 416/500\n924/924 [==============================] - 0s 223us/step - loss: 0.5839 - main_output_loss: 0.5287 - aux_output_loss: 0.5519 - main_output_acc: 0.7165 - aux_output_acc: 0.7370 - val_loss: 0.5433 - val_main_output_loss: 0.4872 - val_aux_output_loss: 0.5613 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7476\nEpoch 417/500\n924/924 [==============================] - 0s 224us/step - loss: 0.5863 - main_output_loss: 0.5311 - aux_output_loss: 0.5517 - main_output_acc: 0.7251 - aux_output_acc: 0.7392 - val_loss: 0.5465 - val_main_output_loss: 0.4903 - val_aux_output_loss: 0.5618 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7476\nEpoch 418/500\n924/924 [==============================] - 0s 223us/step - loss: 0.5765 - main_output_loss: 0.5213 - aux_output_loss: 0.5516 - main_output_acc: 0.7338 - aux_output_acc: 0.7392 - val_loss: 0.5431 - val_main_output_loss: 0.4870 - val_aux_output_loss: 0.5611 - val_main_output_acc: 0.7864 - val_aux_output_acc: 0.7476\nEpoch 419/500\n924/924 [==============================] - 0s 221us/step - loss: 0.5969 - main_output_loss: 0.5418 - aux_output_loss: 0.5512 - main_output_acc: 0.7078 - aux_output_acc: 0.7392 - val_loss: 0.5412 - val_main_output_loss: 0.4852 - val_aux_output_loss: 0.5604 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7476\nEpoch 420/500\n924/924 [==============================] - 0s 212us/step - loss: 0.5752 - main_output_loss: 0.5201 - aux_output_loss: 0.5514 - main_output_acc: 0.7468 - aux_output_acc: 0.7413 - val_loss: 0.5417 - val_main_output_loss: 0.4856 - val_aux_output_loss: 0.5605 - val_main_output_acc: 0.7864 - val_aux_output_acc: 0.7379\nEpoch 421/500\n924/924 [==============================] - 0s 206us/step - loss: 0.5840 - main_output_loss: 0.5289 - aux_output_loss: 0.5510 - main_output_acc: 0.7100 - aux_output_acc: 0.7381 - val_loss: 0.5430 - val_main_output_loss: 0.4869 - val_aux_output_loss: 0.5606 - val_main_output_acc: 0.7864 - val_aux_output_acc: 0.7476\nEpoch 422/500\n924/924 [==============================] - 0s 210us/step - loss: 0.5906 - main_output_loss: 0.5355 - aux_output_loss: 0.5508 - main_output_acc: 0.7197 - aux_output_acc: 0.7392 - val_loss: 0.5426 - val_main_output_loss: 0.4866 - val_aux_output_loss: 0.5601 - val_main_output_acc: 0.7864 - val_aux_output_acc: 0.7379\nEpoch 423/500\n924/924 [==============================] - 0s 213us/step - loss: 0.5864 - main_output_loss: 0.5314 - aux_output_loss: 0.5505 - main_output_acc: 0.7273 - aux_output_acc: 0.7381 - val_loss: 0.5436 - val_main_output_loss: 0.4876 - val_aux_output_loss: 0.5599 - val_main_output_acc: 0.7864 - val_aux_output_acc: 0.7476\nEpoch 424/500\n924/924 [==============================] - 0s 210us/step - loss: 0.5840 - main_output_loss: 0.5289 - aux_output_loss: 0.5504 - main_output_acc: 0.7338 - aux_output_acc: 0.7424 - val_loss: 0.5413 - val_main_output_loss: 0.4854 - val_aux_output_loss: 0.5592 - val_main_output_acc: 0.7864 - val_aux_output_acc: 0.7476\nEpoch 425/500\n924/924 [==============================] - 0s 210us/step - loss: 0.5718 - main_output_loss: 0.5168 - aux_output_loss: 0.5500 - main_output_acc: 0.7468 - aux_output_acc: 0.7435 - val_loss: 0.5402 - val_main_output_loss: 0.4843 - val_aux_output_loss: 0.5591 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7476\nEpoch 426/500\n924/924 [==============================] - 0s 212us/step - loss: 0.5791 - main_output_loss: 0.5241 - aux_output_loss: 0.5499 - main_output_acc: 0.7457 - aux_output_acc: 0.7413 - val_loss: 0.5436 - val_main_output_loss: 0.4876 - val_aux_output_loss: 0.5595 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7476\nEpoch 427/500\n924/924 [==============================] - 0s 210us/step - loss: 0.5884 - main_output_loss: 0.5335 - aux_output_loss: 0.5495 - main_output_acc: 0.7338 - aux_output_acc: 0.7403 - val_loss: 0.5388 - val_main_output_loss: 0.4830 - val_aux_output_loss: 0.5580 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7476\nEpoch 428/500\n924/924 [==============================] - 0s 207us/step - loss: 0.5712 - main_output_loss: 0.5162 - aux_output_loss: 0.5500 - main_output_acc: 0.7500 - aux_output_acc: 0.7381 - val_loss: 0.5392 - val_main_output_loss: 0.4834 - val_aux_output_loss: 0.5582 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7476\nEpoch 429/500\n924/924 [==============================] - 0s 209us/step - loss: 0.5795 - main_output_loss: 0.5246 - aux_output_loss: 0.5492 - main_output_acc: 0.7273 - aux_output_acc: 0.7446 - val_loss: 0.5380 - val_main_output_loss: 0.4822 - val_aux_output_loss: 0.5579 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7476\nEpoch 430/500\n924/924 [==============================] - 0s 209us/step - loss: 0.5862 - main_output_loss: 0.5313 - aux_output_loss: 0.5492 - main_output_acc: 0.7273 - aux_output_acc: 0.7403 - val_loss: 0.5406 - val_main_output_loss: 0.4848 - val_aux_output_loss: 0.5586 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7476\nEpoch 431/500\n924/924 [==============================] - 0s 208us/step - loss: 0.5834 - main_output_loss: 0.5285 - aux_output_loss: 0.5489 - main_output_acc: 0.7240 - aux_output_acc: 0.7403 - val_loss: 0.5393 - val_main_output_loss: 0.4835 - val_aux_output_loss: 0.5584 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7476\nEpoch 432/500\n924/924 [==============================] - 0s 207us/step - loss: 0.5722 - main_output_loss: 0.5173 - aux_output_loss: 0.5488 - main_output_acc: 0.7294 - aux_output_acc: 0.7457 - val_loss: 0.5387 - val_main_output_loss: 0.4829 - val_aux_output_loss: 0.5580 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7476\nEpoch 433/500\n924/924 [==============================] - 0s 210us/step - loss: 0.5747 - main_output_loss: 0.5198 - aux_output_loss: 0.5483 - main_output_acc: 0.7403 - aux_output_acc: 0.7403 - val_loss: 0.5400 - val_main_output_loss: 0.4842 - val_aux_output_loss: 0.5579 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7476\nEpoch 434/500\n924/924 [==============================] - 0s 216us/step - loss: 0.5798 - main_output_loss: 0.5249 - aux_output_loss: 0.5482 - main_output_acc: 0.7403 - aux_output_acc: 0.7446 - val_loss: 0.5414 - val_main_output_loss: 0.4856 - val_aux_output_loss: 0.5583 - val_main_output_acc: 0.8155 - val_aux_output_acc: 0.7476\nEpoch 435/500\n924/924 [==============================] - 0s 213us/step - loss: 0.5812 - main_output_loss: 0.5264 - aux_output_loss: 0.5480 - main_output_acc: 0.7294 - aux_output_acc: 0.7468 - val_loss: 0.5356 - val_main_output_loss: 0.4799 - val_aux_output_loss: 0.5570 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7476\nEpoch 436/500\n924/924 [==============================] - 0s 218us/step - loss: 0.5774 - main_output_loss: 0.5226 - aux_output_loss: 0.5478 - main_output_acc: 0.7381 - aux_output_acc: 0.7468 - val_loss: 0.5349 - val_main_output_loss: 0.4791 - val_aux_output_loss: 0.5571 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7476\nEpoch 437/500\n924/924 [==============================] - 0s 206us/step - loss: 0.5820 - main_output_loss: 0.5272 - aux_output_loss: 0.5477 - main_output_acc: 0.7208 - aux_output_acc: 0.7478 - val_loss: 0.5356 - val_main_output_loss: 0.4799 - val_aux_output_loss: 0.5572 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7476\nEpoch 438/500\n924/924 [==============================] - 0s 216us/step - loss: 0.5821 - main_output_loss: 0.5273 - aux_output_loss: 0.5473 - main_output_acc: 0.7381 - aux_output_acc: 0.7413 - val_loss: 0.5363 - val_main_output_loss: 0.4805 - val_aux_output_loss: 0.5574 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7379\nEpoch 439/500\n924/924 [==============================] - 0s 211us/step - loss: 0.5669 - main_output_loss: 0.5122 - aux_output_loss: 0.5472 - main_output_acc: 0.7273 - aux_output_acc: 0.7468 - val_loss: 0.5356 - val_main_output_loss: 0.4799 - val_aux_output_loss: 0.5574 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7379\nEpoch 440/500\n924/924 [==============================] - 0s 210us/step - loss: 0.5797 - main_output_loss: 0.5250 - aux_output_loss: 0.5467 - main_output_acc: 0.7446 - aux_output_acc: 0.7435 - val_loss: 0.5331 - val_main_output_loss: 0.4774 - val_aux_output_loss: 0.5566 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7476\nEpoch 441/500\n924/924 [==============================] - 0s 209us/step - loss: 0.5776 - main_output_loss: 0.5230 - aux_output_loss: 0.5465 - main_output_acc: 0.7424 - aux_output_acc: 0.7478 - val_loss: 0.5350 - val_main_output_loss: 0.4792 - val_aux_output_loss: 0.5572 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7379\nEpoch 442/500\n924/924 [==============================] - 0s 217us/step - loss: 0.5761 - main_output_loss: 0.5215 - aux_output_loss: 0.5464 - main_output_acc: 0.7435 - aux_output_acc: 0.7478 - val_loss: 0.5397 - val_main_output_loss: 0.4839 - val_aux_output_loss: 0.5584 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7476\nEpoch 443/500\n924/924 [==============================] - 0s 216us/step - loss: 0.5730 - main_output_loss: 0.5184 - aux_output_loss: 0.5463 - main_output_acc: 0.7359 - aux_output_acc: 0.7457 - val_loss: 0.5349 - val_main_output_loss: 0.4792 - val_aux_output_loss: 0.5569 - val_main_output_acc: 0.7864 - val_aux_output_acc: 0.7379\nEpoch 444/500\n924/924 [==============================] - 0s 208us/step - loss: 0.5627 - main_output_loss: 0.5081 - aux_output_loss: 0.5460 - main_output_acc: 0.7489 - aux_output_acc: 0.7446 - val_loss: 0.5329 - val_main_output_loss: 0.4773 - val_aux_output_loss: 0.5564 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7379\nEpoch 445/500\n924/924 [==============================] - 0s 210us/step - loss: 0.5717 - main_output_loss: 0.5172 - aux_output_loss: 0.5456 - main_output_acc: 0.7424 - aux_output_acc: 0.7478 - val_loss: 0.5345 - val_main_output_loss: 0.4788 - val_aux_output_loss: 0.5566 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7379\nEpoch 446/500\n924/924 [==============================] - 0s 214us/step - loss: 0.5767 - main_output_loss: 0.5222 - aux_output_loss: 0.5455 - main_output_acc: 0.7327 - aux_output_acc: 0.7478 - val_loss: 0.5309 - val_main_output_loss: 0.4753 - val_aux_output_loss: 0.5553 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7476\nEpoch 447/500\n924/924 [==============================] - 0s 220us/step - loss: 0.5672 - main_output_loss: 0.5126 - aux_output_loss: 0.5455 - main_output_acc: 0.7348 - aux_output_acc: 0.7478 - val_loss: 0.5309 - val_main_output_loss: 0.4754 - val_aux_output_loss: 0.5553 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7476\nEpoch 448/500\n924/924 [==============================] - 0s 210us/step - loss: 0.5575 - main_output_loss: 0.5030 - aux_output_loss: 0.5450 - main_output_acc: 0.7403 - aux_output_acc: 0.7478 - val_loss: 0.5343 - val_main_output_loss: 0.4787 - val_aux_output_loss: 0.5563 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7476\nEpoch 449/500\n924/924 [==============================] - 0s 213us/step - loss: 0.5630 - main_output_loss: 0.5085 - aux_output_loss: 0.5450 - main_output_acc: 0.7511 - aux_output_acc: 0.7468 - val_loss: 0.5303 - val_main_output_loss: 0.4748 - val_aux_output_loss: 0.5550 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7476\nEpoch 450/500\n924/924 [==============================] - 0s 212us/step - loss: 0.5606 - main_output_loss: 0.5062 - aux_output_loss: 0.5448 - main_output_acc: 0.7446 - aux_output_acc: 0.7522 - val_loss: 0.5310 - val_main_output_loss: 0.4755 - val_aux_output_loss: 0.5550 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7476\nEpoch 451/500\n924/924 [==============================] - 0s 207us/step - loss: 0.5722 - main_output_loss: 0.5178 - aux_output_loss: 0.5445 - main_output_acc: 0.7392 - aux_output_acc: 0.7500 - val_loss: 0.5304 - val_main_output_loss: 0.4749 - val_aux_output_loss: 0.5547 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7476\nEpoch 452/500\n924/924 [==============================] - 0s 209us/step - loss: 0.5644 - main_output_loss: 0.5099 - aux_output_loss: 0.5443 - main_output_acc: 0.7478 - aux_output_acc: 0.7511 - val_loss: 0.5321 - val_main_output_loss: 0.4766 - val_aux_output_loss: 0.5550 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7379\nEpoch 453/500\n924/924 [==============================] - 0s 213us/step - loss: 0.5652 - main_output_loss: 0.5108 - aux_output_loss: 0.5440 - main_output_acc: 0.7478 - aux_output_acc: 0.7522 - val_loss: 0.5342 - val_main_output_loss: 0.4787 - val_aux_output_loss: 0.5552 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7379\nEpoch 454/500\n924/924 [==============================] - 0s 220us/step - loss: 0.5726 - main_output_loss: 0.5182 - aux_output_loss: 0.5437 - main_output_acc: 0.7500 - aux_output_acc: 0.7543 - val_loss: 0.5307 - val_main_output_loss: 0.4752 - val_aux_output_loss: 0.5544 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7379\nEpoch 455/500\n924/924 [==============================] - 0s 208us/step - loss: 0.5683 - main_output_loss: 0.5140 - aux_output_loss: 0.5438 - main_output_acc: 0.7500 - aux_output_acc: 0.7478 - val_loss: 0.5304 - val_main_output_loss: 0.4750 - val_aux_output_loss: 0.5542 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7476\nEpoch 456/500\n924/924 [==============================] - 0s 208us/step - loss: 0.5738 - main_output_loss: 0.5194 - aux_output_loss: 0.5434 - main_output_acc: 0.7316 - aux_output_acc: 0.7522 - val_loss: 0.5294 - val_main_output_loss: 0.4740 - val_aux_output_loss: 0.5534 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7476\nEpoch 457/500\n924/924 [==============================] - 0s 221us/step - loss: 0.5673 - main_output_loss: 0.5129 - aux_output_loss: 0.5434 - main_output_acc: 0.7359 - aux_output_acc: 0.7489 - val_loss: 0.5303 - val_main_output_loss: 0.4749 - val_aux_output_loss: 0.5535 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7476\nEpoch 458/500\n924/924 [==============================] - 0s 209us/step - loss: 0.5730 - main_output_loss: 0.5187 - aux_output_loss: 0.5431 - main_output_acc: 0.7316 - aux_output_acc: 0.7500 - val_loss: 0.5292 - val_main_output_loss: 0.4739 - val_aux_output_loss: 0.5532 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7476\nEpoch 459/500\n924/924 [==============================] - 0s 206us/step - loss: 0.5745 - main_output_loss: 0.5202 - aux_output_loss: 0.5430 - main_output_acc: 0.7381 - aux_output_acc: 0.7478 - val_loss: 0.5292 - val_main_output_loss: 0.4738 - val_aux_output_loss: 0.5531 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7476\nEpoch 460/500\n924/924 [==============================] - 0s 210us/step - loss: 0.5679 - main_output_loss: 0.5136 - aux_output_loss: 0.5426 - main_output_acc: 0.7403 - aux_output_acc: 0.7532 - val_loss: 0.5278 - val_main_output_loss: 0.4725 - val_aux_output_loss: 0.5524 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7476\nEpoch 461/500\n924/924 [==============================] - 0s 206us/step - loss: 0.5662 - main_output_loss: 0.5120 - aux_output_loss: 0.5424 - main_output_acc: 0.7413 - aux_output_acc: 0.7489 - val_loss: 0.5326 - val_main_output_loss: 0.4772 - val_aux_output_loss: 0.5534 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7379\nEpoch 462/500\n924/924 [==============================] - 0s 206us/step - loss: 0.5654 - main_output_loss: 0.5113 - aux_output_loss: 0.5417 - main_output_acc: 0.7435 - aux_output_acc: 0.7522 - val_loss: 0.5269 - val_main_output_loss: 0.4717 - val_aux_output_loss: 0.5521 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7573\nEpoch 463/500\n924/924 [==============================] - 0s 218us/step - loss: 0.5693 - main_output_loss: 0.5151 - aux_output_loss: 0.5421 - main_output_acc: 0.7435 - aux_output_acc: 0.7554 - val_loss: 0.5277 - val_main_output_loss: 0.4725 - val_aux_output_loss: 0.5521 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7573\nEpoch 464/500\n924/924 [==============================] - 0s 208us/step - loss: 0.5630 - main_output_loss: 0.5088 - aux_output_loss: 0.5418 - main_output_acc: 0.7446 - aux_output_acc: 0.7532 - val_loss: 0.5309 - val_main_output_loss: 0.4757 - val_aux_output_loss: 0.5526 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7379\nEpoch 465/500\n924/924 [==============================] - 0s 207us/step - loss: 0.5707 - main_output_loss: 0.5165 - aux_output_loss: 0.5416 - main_output_acc: 0.7316 - aux_output_acc: 0.7543 - val_loss: 0.5293 - val_main_output_loss: 0.4741 - val_aux_output_loss: 0.5520 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7379\nEpoch 466/500\n924/924 [==============================] - 0s 215us/step - loss: 0.5691 - main_output_loss: 0.5150 - aux_output_loss: 0.5413 - main_output_acc: 0.7392 - aux_output_acc: 0.7511 - val_loss: 0.5277 - val_main_output_loss: 0.4725 - val_aux_output_loss: 0.5517 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7573\nEpoch 467/500\n924/924 [==============================] - 0s 209us/step - loss: 0.5490 - main_output_loss: 0.4949 - aux_output_loss: 0.5412 - main_output_acc: 0.7608 - aux_output_acc: 0.7543 - val_loss: 0.5315 - val_main_output_loss: 0.4762 - val_aux_output_loss: 0.5525 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7379\nEpoch 468/500\n924/924 [==============================] - 0s 210us/step - loss: 0.5518 - main_output_loss: 0.4978 - aux_output_loss: 0.5408 - main_output_acc: 0.7554 - aux_output_acc: 0.7543 - val_loss: 0.5280 - val_main_output_loss: 0.4728 - val_aux_output_loss: 0.5517 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7476\nEpoch 469/500\n924/924 [==============================] - 0s 213us/step - loss: 0.5670 - main_output_loss: 0.5130 - aux_output_loss: 0.5407 - main_output_acc: 0.7554 - aux_output_acc: 0.7532 - val_loss: 0.5251 - val_main_output_loss: 0.4700 - val_aux_output_loss: 0.5508 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7476\nEpoch 470/500\n924/924 [==============================] - 0s 215us/step - loss: 0.5638 - main_output_loss: 0.5098 - aux_output_loss: 0.5408 - main_output_acc: 0.7468 - aux_output_acc: 0.7565 - val_loss: 0.5239 - val_main_output_loss: 0.4688 - val_aux_output_loss: 0.5503 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7573\nEpoch 471/500\n924/924 [==============================] - 0s 214us/step - loss: 0.5586 - main_output_loss: 0.5045 - aux_output_loss: 0.5407 - main_output_acc: 0.7597 - aux_output_acc: 0.7522 - val_loss: 0.5245 - val_main_output_loss: 0.4695 - val_aux_output_loss: 0.5503 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7476\nEpoch 472/500\n924/924 [==============================] - 0s 214us/step - loss: 0.5492 - main_output_loss: 0.4952 - aux_output_loss: 0.5406 - main_output_acc: 0.7630 - aux_output_acc: 0.7532 - val_loss: 0.5250 - val_main_output_loss: 0.4700 - val_aux_output_loss: 0.5502 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7476\nEpoch 473/500\n924/924 [==============================] - 0s 206us/step - loss: 0.5512 - main_output_loss: 0.4972 - aux_output_loss: 0.5402 - main_output_acc: 0.7565 - aux_output_acc: 0.7565 - val_loss: 0.5287 - val_main_output_loss: 0.4736 - val_aux_output_loss: 0.5510 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7476\nEpoch 474/500\n924/924 [==============================] - 0s 208us/step - loss: 0.5608 - main_output_loss: 0.5068 - aux_output_loss: 0.5398 - main_output_acc: 0.7554 - aux_output_acc: 0.7522 - val_loss: 0.5235 - val_main_output_loss: 0.4685 - val_aux_output_loss: 0.5498 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7573\nEpoch 475/500\n924/924 [==============================] - 0s 211us/step - loss: 0.5625 - main_output_loss: 0.5085 - aux_output_loss: 0.5398 - main_output_acc: 0.7370 - aux_output_acc: 0.7554 - val_loss: 0.5233 - val_main_output_loss: 0.4684 - val_aux_output_loss: 0.5497 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7476\nEpoch 476/500\n924/924 [==============================] - 0s 207us/step - loss: 0.5545 - main_output_loss: 0.5005 - aux_output_loss: 0.5395 - main_output_acc: 0.7478 - aux_output_acc: 0.7565 - val_loss: 0.5229 - val_main_output_loss: 0.4680 - val_aux_output_loss: 0.5495 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7573\nEpoch 477/500\n924/924 [==============================] - 0s 209us/step - loss: 0.5609 - main_output_loss: 0.5069 - aux_output_loss: 0.5392 - main_output_acc: 0.7457 - aux_output_acc: 0.7532 - val_loss: 0.5286 - val_main_output_loss: 0.4735 - val_aux_output_loss: 0.5508 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7476\nEpoch 478/500\n924/924 [==============================] - 0s 210us/step - loss: 0.5681 - main_output_loss: 0.5141 - aux_output_loss: 0.5395 - main_output_acc: 0.7381 - aux_output_acc: 0.7565 - val_loss: 0.5261 - val_main_output_loss: 0.4710 - val_aux_output_loss: 0.5502 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7476\nEpoch 479/500\n924/924 [==============================] - 0s 208us/step - loss: 0.5545 - main_output_loss: 0.5006 - aux_output_loss: 0.5389 - main_output_acc: 0.7424 - aux_output_acc: 0.7543 - val_loss: 0.5227 - val_main_output_loss: 0.4678 - val_aux_output_loss: 0.5493 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7476\nEpoch 480/500\n924/924 [==============================] - 0s 216us/step - loss: 0.5488 - main_output_loss: 0.4949 - aux_output_loss: 0.5388 - main_output_acc: 0.7457 - aux_output_acc: 0.7565 - val_loss: 0.5240 - val_main_output_loss: 0.4690 - val_aux_output_loss: 0.5495 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7573\nEpoch 481/500\n924/924 [==============================] - 0s 212us/step - loss: 0.5602 - main_output_loss: 0.5064 - aux_output_loss: 0.5385 - main_output_acc: 0.7522 - aux_output_acc: 0.7565 - val_loss: 0.5227 - val_main_output_loss: 0.4678 - val_aux_output_loss: 0.5492 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7476\nEpoch 482/500\n924/924 [==============================] - 0s 216us/step - loss: 0.5669 - main_output_loss: 0.5131 - aux_output_loss: 0.5385 - main_output_acc: 0.7532 - aux_output_acc: 0.7576 - val_loss: 0.5221 - val_main_output_loss: 0.4672 - val_aux_output_loss: 0.5489 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7476\nEpoch 483/500\n924/924 [==============================] - 0s 220us/step - loss: 0.5473 - main_output_loss: 0.4935 - aux_output_loss: 0.5385 - main_output_acc: 0.7695 - aux_output_acc: 0.7565 - val_loss: 0.5219 - val_main_output_loss: 0.4670 - val_aux_output_loss: 0.5486 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7573\nEpoch 484/500\n924/924 [==============================] - 0s 213us/step - loss: 0.5649 - main_output_loss: 0.5111 - aux_output_loss: 0.5383 - main_output_acc: 0.7489 - aux_output_acc: 0.7543 - val_loss: 0.5250 - val_main_output_loss: 0.4700 - val_aux_output_loss: 0.5492 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7476\nEpoch 485/500\n924/924 [==============================] - 0s 231us/step - loss: 0.5604 - main_output_loss: 0.5066 - aux_output_loss: 0.5380 - main_output_acc: 0.7554 - aux_output_acc: 0.7576 - val_loss: 0.5250 - val_main_output_loss: 0.4701 - val_aux_output_loss: 0.5492 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7476\nEpoch 486/500\n924/924 [==============================] - 0s 205us/step - loss: 0.5526 - main_output_loss: 0.4988 - aux_output_loss: 0.5377 - main_output_acc: 0.7673 - aux_output_acc: 0.7554 - val_loss: 0.5222 - val_main_output_loss: 0.4674 - val_aux_output_loss: 0.5483 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7573\nEpoch 487/500\n924/924 [==============================] - 0s 212us/step - loss: 0.5546 - main_output_loss: 0.5008 - aux_output_loss: 0.5376 - main_output_acc: 0.7500 - aux_output_acc: 0.7543 - val_loss: 0.5216 - val_main_output_loss: 0.4668 - val_aux_output_loss: 0.5480 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7573\nEpoch 488/500\n924/924 [==============================] - 0s 211us/step - loss: 0.5434 - main_output_loss: 0.4896 - aux_output_loss: 0.5376 - main_output_acc: 0.7511 - aux_output_acc: 0.7565 - val_loss: 0.5219 - val_main_output_loss: 0.4671 - val_aux_output_loss: 0.5480 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7573\nEpoch 489/500\n924/924 [==============================] - 0s 210us/step - loss: 0.5550 - main_output_loss: 0.5013 - aux_output_loss: 0.5375 - main_output_acc: 0.7522 - aux_output_acc: 0.7565 - val_loss: 0.5219 - val_main_output_loss: 0.4671 - val_aux_output_loss: 0.5477 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7573\nEpoch 490/500\n924/924 [==============================] - 0s 208us/step - loss: 0.5548 - main_output_loss: 0.5011 - aux_output_loss: 0.5371 - main_output_acc: 0.7554 - aux_output_acc: 0.7576 - val_loss: 0.5233 - val_main_output_loss: 0.4685 - val_aux_output_loss: 0.5479 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7476\nEpoch 491/500\n924/924 [==============================] - 0s 216us/step - loss: 0.5523 - main_output_loss: 0.4986 - aux_output_loss: 0.5367 - main_output_acc: 0.7500 - aux_output_acc: 0.7565 - val_loss: 0.5206 - val_main_output_loss: 0.4659 - val_aux_output_loss: 0.5471 - val_main_output_acc: 0.7961 - val_aux_output_acc: 0.7573\nEpoch 492/500\n924/924 [==============================] - 0s 227us/step - loss: 0.5642 - main_output_loss: 0.5106 - aux_output_loss: 0.5369 - main_output_acc: 0.7478 - aux_output_acc: 0.7511 - val_loss: 0.5232 - val_main_output_loss: 0.4684 - val_aux_output_loss: 0.5476 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7476\nEpoch 493/500\n924/924 [==============================] - 0s 225us/step - loss: 0.5493 - main_output_loss: 0.4957 - aux_output_loss: 0.5363 - main_output_acc: 0.7413 - aux_output_acc: 0.7576 - val_loss: 0.5212 - val_main_output_loss: 0.4665 - val_aux_output_loss: 0.5471 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7573\nEpoch 494/500\n924/924 [==============================] - 0s 233us/step - loss: 0.5538 - main_output_loss: 0.5002 - aux_output_loss: 0.5364 - main_output_acc: 0.7511 - aux_output_acc: 0.7587 - val_loss: 0.5219 - val_main_output_loss: 0.4672 - val_aux_output_loss: 0.5473 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7476\nEpoch 495/500\n924/924 [==============================] - 0s 223us/step - loss: 0.5531 - main_output_loss: 0.4995 - aux_output_loss: 0.5360 - main_output_acc: 0.7500 - aux_output_acc: 0.7565 - val_loss: 0.5228 - val_main_output_loss: 0.4681 - val_aux_output_loss: 0.5473 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7476\nEpoch 496/500\n924/924 [==============================] - 0s 223us/step - loss: 0.5506 - main_output_loss: 0.4970 - aux_output_loss: 0.5361 - main_output_acc: 0.7543 - aux_output_acc: 0.7565 - val_loss: 0.5196 - val_main_output_loss: 0.4650 - val_aux_output_loss: 0.5467 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7573\nEpoch 497/500\n924/924 [==============================] - 0s 220us/step - loss: 0.5498 - main_output_loss: 0.4962 - aux_output_loss: 0.5360 - main_output_acc: 0.7532 - aux_output_acc: 0.7565 - val_loss: 0.5185 - val_main_output_loss: 0.4639 - val_aux_output_loss: 0.5466 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7476\nEpoch 498/500\n924/924 [==============================] - 0s 219us/step - loss: 0.5515 - main_output_loss: 0.4979 - aux_output_loss: 0.5354 - main_output_acc: 0.7500 - aux_output_acc: 0.7565 - val_loss: 0.5207 - val_main_output_loss: 0.4660 - val_aux_output_loss: 0.5472 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7476\nEpoch 499/500\n924/924 [==============================] - 0s 220us/step - loss: 0.5429 - main_output_loss: 0.4894 - aux_output_loss: 0.5352 - main_output_acc: 0.7532 - aux_output_acc: 0.7554 - val_loss: 0.5202 - val_main_output_loss: 0.4655 - val_aux_output_loss: 0.5471 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7476\nEpoch 500/500\n924/924 [==============================] - 0s 225us/step - loss: 0.5431 - main_output_loss: 0.4896 - aux_output_loss: 0.5351 - main_output_acc: 0.7608 - aux_output_acc: 0.7576 - val_loss: 0.5203 - val_main_output_loss: 0.4656 - val_aux_output_loss: 0.5470 - val_main_output_acc: 0.8058 - val_aux_output_acc: 0.7476\n"
],
[
"# plot the fit\npred_main, pred_aux = model.predict([x_seq_test, x_static_test])\nroc = roc_curve(y_test, pred_main)\nauc = roc_auc_score(y_test, pred_main)\nfig = plt.figure(figsize=(4, 3)) # in inches\nplt.plot(roc[0], roc[1], color = 'darkorange', label = 'ROC curve\\n(area = %0.2f)' % auc)\nplt.plot([0, 1], [0, 1], color= 'navy', linestyle = '--')\nplt.xlabel('False Positive Rate')\nplt.ylabel('True Positive Rate')\nplt.title('%s: ROC' % 'GRU-base')\nplt.legend(loc = \"lower right\")\nfig_name = 'gru-do.pdf'\nfig.savefig(os.path.join(fig_fp, fig_name), bbox_inches='tight')\nplt.show()",
"_____no_output_____"
],
[
"# plot training and validation loss and accuracy\n\nacc = history.history['main_output_acc']\nval_acc = history.history['val_main_output_acc']\nloss = history.history['main_output_loss']\nval_loss = history.history['val_loss']\n\nepochs = range(len(acc))\n\nplt.plot(epochs, acc, 'bo', label='Training acc')\nplt.plot(epochs, val_acc, 'b', label='Validation acc')\nplt.title('Training and validation accuracy')\nplt.legend()\n\nplt.figure()\n\nplt.plot(epochs, loss, 'bo', label='Training loss')\nplt.plot(epochs, val_loss, 'b', label='Validation loss')\nplt.title('Training and validation loss')\nplt.legend()\n\nfig_name = 'do_loss_acc.pdf'\nfig.savefig(os.path.join(fig_fp, fig_name), bbox_inches='tight')",
"_____no_output_____"
]
]
] | [
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] | [
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
]
] |
d0ff8a488847edac05b92cfbd4d756afd6d50ad5 | 106,105 | ipynb | Jupyter Notebook | course/1 First Deep Learning Model.ipynb | Zhenxingzhang/zero_to_deep_learning_video | 1cd49a017ac306efd4bd6c405bf8f1523b1f08b3 | [
"MIT"
] | null | null | null | course/1 First Deep Learning Model.ipynb | Zhenxingzhang/zero_to_deep_learning_video | 1cd49a017ac306efd4bd6c405bf8f1523b1f08b3 | [
"MIT"
] | null | null | null | course/1 First Deep Learning Model.ipynb | Zhenxingzhang/zero_to_deep_learning_video | 1cd49a017ac306efd4bd6c405bf8f1523b1f08b3 | [
"MIT"
] | null | null | null | 327.484568 | 50,576 | 0.927892 | [
[
[
"# First Deep Learning Model",
"_____no_output_____"
]
],
[
[
"import numpy as np\n%matplotlib inline\nimport matplotlib.pyplot as plt",
"_____no_output_____"
],
[
"from sklearn.datasets.samples_generator import make_circles",
"_____no_output_____"
],
[
"X, y = make_circles(n_samples=1000,\n noise=0.1,\n factor=0.2,\n random_state=0)",
"_____no_output_____"
],
[
"X",
"_____no_output_____"
],
[
"X.shape",
"_____no_output_____"
],
[
"plt.figure(figsize=(5, 5))\nplt.plot(X[y==0, 0], X[y==0, 1], 'ob', alpha=0.5)\nplt.plot(X[y==1, 0], X[y==1, 1], 'xr', alpha=0.5)\nplt.xlim(-1.5, 1.5)\nplt.ylim(-1.5, 1.5)\nplt.legend(['0', '1'])\nplt.title(\"Blue circles and Red crosses\")",
"_____no_output_____"
],
[
"from tensorflow.keras.models import Sequential\nfrom tensorflow.keras.layers import Dense\nfrom tensorflow.keras.optimizers import SGD",
"_____no_output_____"
],
[
"model = Sequential()",
"_____no_output_____"
],
[
"model.add(Dense(4, input_shape=(2,), activation='tanh'))",
"_____no_output_____"
],
[
"model.add(Dense(1, activation='sigmoid'))",
"_____no_output_____"
],
[
"model.compile(SGD(lr=0.5), 'binary_crossentropy', metrics=['accuracy'])",
"_____no_output_____"
],
[
"model.fit(X, y, epochs=20)",
"Epoch 1/20\n1000/1000 [==============================] - 0s 313us/sample - loss: 0.6945 - accuracy: 0.5230\nEpoch 2/20\n1000/1000 [==============================] - 0s 52us/sample - loss: 0.6948 - accuracy: 0.4930\nEpoch 3/20\n1000/1000 [==============================] - 0s 67us/sample - loss: 0.6911 - accuracy: 0.5300\nEpoch 4/20\n1000/1000 [==============================] - 0s 62us/sample - loss: 0.6797 - accuracy: 0.6400\nEpoch 5/20\n1000/1000 [==============================] - 0s 64us/sample - loss: 0.6445 - accuracy: 0.7110\nEpoch 6/20\n1000/1000 [==============================] - 0s 56us/sample - loss: 0.5602 - accuracy: 0.8260\nEpoch 7/20\n1000/1000 [==============================] - 0s 62us/sample - loss: 0.4544 - accuracy: 0.8580\nEpoch 8/20\n1000/1000 [==============================] - 0s 59us/sample - loss: 0.3555 - accuracy: 0.8980\nEpoch 9/20\n1000/1000 [==============================] - 0s 66us/sample - loss: 0.2687 - accuracy: 0.9550\nEpoch 10/20\n1000/1000 [==============================] - 0s 74us/sample - loss: 0.2007 - accuracy: 0.9970\nEpoch 11/20\n1000/1000 [==============================] - 0s 50us/sample - loss: 0.1543 - accuracy: 1.0000\nEpoch 12/20\n1000/1000 [==============================] - 0s 50us/sample - loss: 0.1246 - accuracy: 1.0000\nEpoch 13/20\n1000/1000 [==============================] - 0s 54us/sample - loss: 0.1038 - accuracy: 1.0000\nEpoch 14/20\n1000/1000 [==============================] - 0s 48us/sample - loss: 0.0886 - accuracy: 1.0000\nEpoch 15/20\n1000/1000 [==============================] - 0s 48us/sample - loss: 0.0771 - accuracy: 1.0000\nEpoch 16/20\n1000/1000 [==============================] - 0s 48us/sample - loss: 0.0681 - accuracy: 1.0000\nEpoch 17/20\n1000/1000 [==============================] - 0s 59us/sample - loss: 0.0610 - accuracy: 1.0000\nEpoch 18/20\n1000/1000 [==============================] - 0s 62us/sample - loss: 0.0550 - accuracy: 1.0000\nEpoch 19/20\n1000/1000 [==============================] - 0s 61us/sample - loss: 0.0500 - accuracy: 1.0000\nEpoch 20/20\n1000/1000 [==============================] - 0s 55us/sample - loss: 0.0457 - accuracy: 1.0000\n"
],
[
"hticks = np.linspace(-1.5, 1.5, 101)\nvticks = np.linspace(-1.5, 1.5, 101)\naa, bb = np.meshgrid(hticks, vticks)\nab = np.c_[aa.ravel(), bb.ravel()]\nc = model.predict(ab)\ncc = c.reshape(aa.shape)",
"_____no_output_____"
],
[
"plt.figure(figsize=(5, 5))\nplt.contourf(aa, bb, cc, cmap='bwr', alpha=0.2)\nplt.plot(X[y==0, 0], X[y==0, 1], 'ob', alpha=0.5)\nplt.plot(X[y==1, 0], X[y==1, 1], 'xr', alpha=0.5)\nplt.xlim(-1.5, 1.5)\nplt.ylim(-1.5, 1.5)\nplt.legend(['0', '1'])\nplt.title(\"Blue circles and Red crosses\")",
"_____no_output_____"
]
]
] | [
"markdown",
"code"
] | [
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
d0ffa7fb8739489f601431061f3f0e96edd3fc14 | 3,619 | ipynb | Jupyter Notebook | docs_src/models.unet.ipynb | Gokkulnath/fastai_v1 | 5a24c6ebd42223d37e90463f69d32b4a52b6895c | [
"Apache-2.0"
] | 115 | 2018-07-11T06:21:32.000Z | 2018-09-25T09:16:44.000Z | docs_src/models.unet.ipynb | rsaxby/fastai_old | c8285fbc246f41066da02a74b86c917923892ba8 | [
"Apache-2.0"
] | 24 | 2018-07-10T22:18:05.000Z | 2018-09-22T00:26:54.000Z | docs_src/models.unet.ipynb | rsaxby/fastai_old | c8285fbc246f41066da02a74b86c917923892ba8 | [
"Apache-2.0"
] | 45 | 2018-10-10T01:48:03.000Z | 2022-01-12T16:50:54.000Z | 32.9 | 592 | 0.60514 | [
[
[
"# models.unet",
"_____no_output_____"
],
[
"Type an introduction of the package here.",
"_____no_output_____"
]
],
[
[
"from fastai.gen_doc.nbdoc import *\nfrom fastai.models.unet import * ",
"_____no_output_____"
]
],
[
[
"### Global Variable Definitions:",
"_____no_output_____"
]
],
[
[
"show_doc(DynamicUnet)",
"_____no_output_____"
]
],
[
[
"[`DynamicUnet`](/models.unet.html#DynamicUnet)",
"_____no_output_____"
]
],
[
[
"show_doc(UnetBlock)",
"_____no_output_____"
]
],
[
[
"[`UnetBlock`](/models.unet.html#UnetBlock)",
"_____no_output_____"
]
],
[
[
"show_doc(UnetBlock.forward)",
"_____no_output_____"
]
],
[
[
"`UnetBlock.forward`",
"_____no_output_____"
]
]
] | [
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown"
] | [
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
]
] |
d0ffaa67c400dffd6c12e8cec1995c9a8872f5be | 34,239 | ipynb | Jupyter Notebook | fitting_data.ipynb | bibliotekue/python-applied-data-analysis | 893c9e5d7de176ce3271e01e3958797c6561d551 | [
"MIT"
] | null | null | null | fitting_data.ipynb | bibliotekue/python-applied-data-analysis | 893c9e5d7de176ce3271e01e3958797c6561d551 | [
"MIT"
] | null | null | null | fitting_data.ipynb | bibliotekue/python-applied-data-analysis | 893c9e5d7de176ce3271e01e3958797c6561d551 | [
"MIT"
] | null | null | null | 77.993166 | 14,056 | 0.824703 | [
[
[
"# $\\color{black}{}$\n### 1. Fitting data\n---",
"_____no_output_____"
],
[
"### Input data",
"_____no_output_____"
]
],
[
[
"import pandas as pd\nimport seaborn as sns\n\ndata = pd.read_csv('https://milliams.com/courses/applied_data_analysis/linear.csv')\ndata.head()",
"_____no_output_____"
]
],
[
[
"Let's check how many rows we have",
"_____no_output_____"
]
],
[
[
"data.count()",
"_____no_output_____"
]
],
[
[
"We have 50 rows here. In the input data, each row is often called a sample (though sometimes also called an instance, example or observation). For example, it could be the information about a single person from a census or the measurements at a particular time from a weather station.",
"_____no_output_____"
],
[
"Let's have a look at what the data looks like when plotted",
"_____no_output_____"
]
],
[
[
"sns.scatterplot(data=data, x=\"x\", y=\"y\")",
"_____no_output_____"
]
],
[
[
"We can clearly visually see here that there is a linear relationship between the x and y values but we need to be able to extract the exact parameters programmatically.",
"_____no_output_____"
],
[
"### Setting up our model",
"_____no_output_____"
],
[
"We import the model and create an instance of it. By default the LinearRegression model will fit the y-intercept, but since we don't want to make that assumption we explicitly pass fit_intercept=True. fit_intercept is an example of a hyperparameter, which are variables or options in a model which you set up-front rather than letting them be learned from the data.",
"_____no_output_____"
]
],
[
[
"from sklearn.linear_model import LinearRegression\nmodel = LinearRegression(fit_intercept=True)",
"_____no_output_____"
]
],
[
[
"### Fitting the data",
"_____no_output_____"
],
[
"Once we have created our model, we can fit it to the data by calling the `fit()` method on it. This takes two arguments:\n\n1. The input data as a two-dimensional structure of the size ($N_{samples}$,$N_{features}$)\n\n2. The labels or targets of the data as a one-dimensional data structure of size ($N_{samples}$)\n",
"_____no_output_____"
],
[
"If we just request `data[\"x\"]` then that will be a 1D array (actually a pandas `Series`) of shape (50) so we must request the data with `data[[\"x\"]]` (which returns it as a single-column, but still two-dimensional, `DataFrame`).",
"_____no_output_____"
],
[
"If you're using pandas to store your data (as we are) then just remember that the first argument should be a `DataFrame` and the second should be a `Series`.",
"_____no_output_____"
]
],
[
[
"X = data[['x']]\ny = data['y']",
"_____no_output_____"
],
[
"model.fit(X, y)",
"_____no_output_____"
]
],
[
[
"### Making predictions using the model",
"_____no_output_____"
],
[
"We can use this to plot the fit over the original data to compare the result. By getting the predicted *__y__* values for the minimum and maximum x values, we can plot a straight line between them to visualise the model.",
"_____no_output_____"
],
[
"The `predict()` function takes an array of the same shape as the original input data (($N_{samples}$, $N_{features}$)) so we put our list of *__x__* values into a `DataFrame` before passing it to `predict()`.",
"_____no_output_____"
],
[
"We then plot the original data in the same way as before and draw the prediction line in the same plot.",
"_____no_output_____"
]
],
[
[
"x_fit = pd.DataFrame({'x': [0, 10]})\ny_pred = model.predict(x_fit)",
"_____no_output_____"
],
[
"import matplotlib.pyplot as plt\n\nplt.figure(figsize=(20,20))\n\nfig, ax = plt.subplots()\nsns.scatterplot(data=data, x='x', y='y', ax=ax)\nax.plot(x_fit['x'], y_pred, linestyle=':', color='red')\nplt.show()",
"_____no_output_____"
]
],
[
[
"As well as plotting the line in a graph, we can also extract the calculated values of the gradient and y-intercept. The gradient is available as a list of values, `model.coef_`, one for each dimension or feature. The intercept is available as `model.intercept_`.",
"_____no_output_____"
]
],
[
[
"print(f'Model gradient: {model.coef_[0]}')\nprint(f'Model intercept: {model.intercept_}')",
"Model gradient: 1.9776566003853107\nModel intercept: -4.903310725531115\n"
]
],
[
[
"The equation that we have extracted can therefore be represented as:\n __$y = 1.97x - 4.90$__",
"_____no_output_____"
],
[
"The original data was produced (with random wobble applied) from a straight line with gradient __$2$__ and y-intercept of __$−5$__. Our model has managed to predict values very close to the original.",
"_____no_output_____"
]
]
] | [
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown"
] | [
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown",
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown",
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
]
] |
d0ffb20f341b75705349b8147af6005b297e60fd | 345,624 | ipynb | Jupyter Notebook | 1_2_Convolutional_Filters_Edge_Detection/5. Canny Edge Detection.ipynb | Echo9k/CVND_Exercises | c5d84e9407fdee642ec394f5c17a92d29433b9cc | [
"MIT"
] | null | null | null | 1_2_Convolutional_Filters_Edge_Detection/5. Canny Edge Detection.ipynb | Echo9k/CVND_Exercises | c5d84e9407fdee642ec394f5c17a92d29433b9cc | [
"MIT"
] | null | null | null | 1_2_Convolutional_Filters_Edge_Detection/5. Canny Edge Detection.ipynb | Echo9k/CVND_Exercises | c5d84e9407fdee642ec394f5c17a92d29433b9cc | [
"MIT"
] | null | null | null | 1,350.09375 | 112,676 | 0.960443 | [
[
[
"import numpy as np\nimport matplotlib.pyplot as plt\nimport cv2\n\n%matplotlib inline\n\n# Read in the image\nimage = cv2.imread('images/brain_MR.jpg')\n\n# Change color to RGB (from BGR)\nimage = cv2.cvtColor(image, cv2.COLOR_BGR2RGB)\n\nplt.imshow(image)",
"_____no_output_____"
],
[
"# Convert the image to grayscale for processing\ngray = cv2.cvtColor(image, cv2.COLOR_RGB2GRAY)\n\nplt.imshow(gray, cmap='gray')",
"_____no_output_____"
]
],
[
[
"### Implement Canny edge detection",
"_____no_output_____"
]
],
[
[
"# Try Canny using \"wide\" and \"tight\" thresholds\n\nwide = cv2.Canny(gray, 30, 100)\ntight = cv2.Canny(gray, 200, 240)\n \n \n# Display the images\nf, (ax1, ax2) = plt.subplots(1, 2, figsize=(20,10))\n\nax1.set_title('wide')\nax1.imshow(wide, cmap='gray')\n\nax2.set_title('tight')\nax2.imshow(tight, cmap='gray')",
"_____no_output_____"
]
],
[
[
"### TODO: Try to find the edges of this flower\n\nSet a small enough threshold to isolate the boundary of the flower.",
"_____no_output_____"
]
],
[
[
"# Read in the image\nimage = cv2.imread('images/sunflower.jpg')\n\n# Change color to RGB (from BGR)\nimage = cv2.cvtColor(image, cv2.COLOR_BGR2RGB)\n\nplt.imshow(image)",
"_____no_output_____"
],
[
"# Convert the image to grayscale\ngray = cv2.cvtColor(image, cv2.COLOR_RGB2GRAY)\n\n## TODO: Define lower and upper thresholds for hysteresis\n# right now the threshold is so small and low that it will pick up a lot of noise\nlower = 0\nupper = 50\n\nedges = cv2.Canny(gray, lower, upper)\n\nplt.figure(figsize=(20,10))\nplt.imshow(edges, cmap='gray')",
"_____no_output_____"
]
]
] | [
"code",
"markdown",
"code",
"markdown",
"code"
] | [
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
]
] |
d0ffb499a894f883c615a7199ba13da65262748f | 62,756 | ipynb | Jupyter Notebook | PennyLane/Data Reuploading Classifier/.ipynb_checkpoints/DRC MNIST MultiClass PCA-Copy1-checkpoint.ipynb | Graciaira/quantum_image_classifier | 1e6a8ec93f51dcbfd63c2e652be5d1fcbce283ce | [
"MIT"
] | 1 | 2021-06-08T12:32:09.000Z | 2021-06-08T12:32:09.000Z | PennyLane/Data Reuploading Classifier/.ipynb_checkpoints/DRC MNIST MultiClass PCA-Copy1-checkpoint.ipynb | Graciaira/quantum_image_classifier | 1e6a8ec93f51dcbfd63c2e652be5d1fcbce283ce | [
"MIT"
] | null | null | null | PennyLane/Data Reuploading Classifier/.ipynb_checkpoints/DRC MNIST MultiClass PCA-Copy1-checkpoint.ipynb | Graciaira/quantum_image_classifier | 1e6a8ec93f51dcbfd63c2e652be5d1fcbce283ce | [
"MIT"
] | null | null | null | 43.013023 | 23,656 | 0.699933 | [
[
[
"# Mount Google Drive\nfrom google.colab import drive # import drive from google colab\n \nROOT = \"/content/drive\" # default location for the drive\nprint(ROOT) # print content of ROOT (Optional)\n \ndrive.mount(ROOT) # we mount the google drive at /content/drive",
"/content/drive\nMounted at /content/drive\n"
],
[
"!pip install pennylane\nfrom IPython.display import clear_output\nclear_output()",
"_____no_output_____"
],
[
"import os\n\ndef restart_runtime():\n os.kill(os.getpid(), 9)\nrestart_runtime()",
"_____no_output_____"
],
[
"# %matplotlib inline\nimport matplotlib.pyplot as plt\nfrom mpl_toolkits.axes_grid1 import make_axes_locatable\n\nimport numpy as np",
"_____no_output_____"
]
],
[
[
"# Loading Raw Data",
"_____no_output_____"
]
],
[
[
"import tensorflow as tf\n\n(x_train, y_train), (x_test, y_test) = tf.keras.datasets.mnist.load_data()",
"_____no_output_____"
],
[
"x_train_flatten = x_train.reshape(x_train.shape[0], x_train.shape[1]*x_train.shape[2])/255.0\nx_test_flatten = x_test.reshape(x_test.shape[0], x_test.shape[1]*x_test.shape[2])/255.0",
"_____no_output_____"
],
[
"print(x_train_flatten.shape, y_train.shape)\nprint(x_test_flatten.shape, y_test.shape)",
"(60000, 784) (60000,)\n(10000, 784) (10000,)\n"
],
[
"x_train_0 = x_train_flatten[y_train == 0]\nx_train_1 = x_train_flatten[y_train == 1]\nx_train_2 = x_train_flatten[y_train == 2]\nx_train_3 = x_train_flatten[y_train == 3]\nx_train_4 = x_train_flatten[y_train == 4]\nx_train_5 = x_train_flatten[y_train == 5]\nx_train_6 = x_train_flatten[y_train == 6]\nx_train_7 = x_train_flatten[y_train == 7]\nx_train_8 = x_train_flatten[y_train == 8]\nx_train_9 = x_train_flatten[y_train == 9]\n\nx_train_list = [x_train_0, x_train_1, x_train_2, x_train_3, x_train_4, x_train_5, x_train_6, x_train_7, x_train_8, x_train_9]\n\nprint(x_train_0.shape)\nprint(x_train_1.shape)\nprint(x_train_2.shape)\nprint(x_train_3.shape)\nprint(x_train_4.shape)\nprint(x_train_5.shape)\nprint(x_train_6.shape)\nprint(x_train_7.shape)\nprint(x_train_8.shape)\nprint(x_train_9.shape)",
"(5923, 784)\n(6742, 784)\n(5958, 784)\n(6131, 784)\n(5842, 784)\n(5421, 784)\n(5918, 784)\n(6265, 784)\n(5851, 784)\n(5949, 784)\n"
],
[
"x_test_0 = x_test_flatten[y_test == 0]\nx_test_1 = x_test_flatten[y_test == 1]\nx_test_2 = x_test_flatten[y_test == 2]\nx_test_3 = x_test_flatten[y_test == 3]\nx_test_4 = x_test_flatten[y_test == 4]\nx_test_5 = x_test_flatten[y_test == 5]\nx_test_6 = x_test_flatten[y_test == 6]\nx_test_7 = x_test_flatten[y_test == 7]\nx_test_8 = x_test_flatten[y_test == 8]\nx_test_9 = x_test_flatten[y_test == 9]\n\nx_test_list = [x_test_0, x_test_1, x_test_2, x_test_3, x_test_4, x_test_5, x_test_6, x_test_7, x_test_8, x_test_9]\n\nprint(x_test_0.shape)\nprint(x_test_1.shape)\nprint(x_test_2.shape)\nprint(x_test_3.shape)\nprint(x_test_4.shape)\nprint(x_test_5.shape)\nprint(x_test_6.shape)\nprint(x_test_7.shape)\nprint(x_test_8.shape)\nprint(x_test_9.shape)",
"(980, 784)\n(1135, 784)\n(1032, 784)\n(1010, 784)\n(982, 784)\n(892, 784)\n(958, 784)\n(1028, 784)\n(974, 784)\n(1009, 784)\n"
]
],
[
[
"# Selecting the dataset\n\nOutput: X_train, Y_train, X_test, Y_test",
"_____no_output_____"
]
],
[
[
"num_sample = 300\nn_class = 4\nmult_test = 0.25\n\nX_train = x_train_list[0][:num_sample, :]\nX_test = x_test_list[0][:int(mult_test*num_sample), :]\n\nY_train = np.zeros((n_class*X_train.shape[0],), dtype=int)\nY_test = np.zeros((n_class*X_test.shape[0],), dtype=int)\n\nfor i in range(n_class-1):\n X_train = np.concatenate((X_train, x_train_list[i+1][:num_sample, :]), axis=0)\n Y_train[num_sample*(i+1):num_sample*(i+2)] = int(i+1)\n\n X_test = np.concatenate((X_test, x_test_list[i+1][:int(mult_test*num_sample), :]), axis=0)\n Y_test[int(mult_test*num_sample*(i+1)):int(mult_test*num_sample*(i+2))] = int(i+1)\n\nprint(X_train.shape, Y_train.shape)\nprint(X_test.shape, Y_test.shape)",
"(1200, 784) (1200,)\n(300, 784) (300,)\n"
]
],
[
[
"# Dataset Preprocessing (Standardization + PCA)",
"_____no_output_____"
],
[
"## Standardization",
"_____no_output_____"
]
],
[
[
"def normalize(X, use_params=False, params=None):\n \"\"\"Normalize the given dataset X\n Args:\n X: ndarray, dataset\n \n Returns:\n (Xbar, mean, std): tuple of ndarray, Xbar is the normalized dataset\n with mean 0 and standard deviation 1; mean and std are the \n mean and standard deviation respectively.\n \n Note:\n You will encounter dimensions where the standard deviation is\n zero, for those when you do normalization the normalized data\n will be NaN. Handle this by setting using `std = 1` for those \n dimensions when doing normalization.\n \"\"\"\n if use_params:\n mu = params[0]\n std_filled = [1]\n else:\n mu = np.mean(X, axis=0)\n std = np.std(X, axis=0)\n #std_filled = std.copy()\n #std_filled[std==0] = 1.\n Xbar = (X - mu)/(std + 1e-8)\n return Xbar, mu, std\n",
"_____no_output_____"
],
[
"X_train, mu_train, std_train = normalize(X_train)\nX_train.shape, Y_train.shape",
"_____no_output_____"
],
[
"X_test = (X_test - mu_train)/(std_train + 1e-8)\nX_test.shape, Y_test.shape",
"_____no_output_____"
]
],
[
[
"## PCA",
"_____no_output_____"
]
],
[
[
"from sklearn.decomposition import PCA\nfrom matplotlib import pyplot as plt",
"_____no_output_____"
],
[
"num_component = 9\npca = PCA(n_components=num_component, svd_solver='full')",
"_____no_output_____"
],
[
"pca.fit(X_train)",
"_____no_output_____"
],
[
"np.cumsum(pca.explained_variance_ratio_)",
"_____no_output_____"
],
[
"X_train = pca.transform(X_train)\nX_test = pca.transform(X_test)",
"_____no_output_____"
],
[
"print(X_train.shape, Y_train.shape)\nprint(X_test.shape, Y_test.shape)",
"(1200, 9) (1200,)\n(300, 9) (300,)\n"
]
],
[
[
"## Norm",
"_____no_output_____"
]
],
[
[
"X_train = (X_train.T / np.sqrt(np.sum(X_train ** 2, -1))).T\nX_test = (X_test.T / np.sqrt(np.sum(X_test ** 2, -1))).T",
"_____no_output_____"
],
[
"plt.scatter(X_train[:100, 0], X_train[:100, 1])\nplt.scatter(X_train[100:200, 0], X_train[100:200, 1])\nplt.scatter(X_train[200:300, 0], X_train[200:300, 1])",
"_____no_output_____"
]
],
[
[
"# Quantum",
"_____no_output_____"
]
],
[
[
"import pennylane as qml\nfrom pennylane import numpy as np\nfrom pennylane.optimize import AdamOptimizer, GradientDescentOptimizer\n\nqml.enable_tape()\n\n\n# Set a random seed\nnp.random.seed(42)",
"_____no_output_____"
],
[
"def plot_data(x, y, fig=None, ax=None):\n \"\"\"\n Plot data with red/blue values for a binary classification.\n\n Args:\n x (array[tuple]): array of data points as tuples\n y (array[int]): array of data points as tuples\n \"\"\"\n if fig == None:\n fig, ax = plt.subplots(1, 1, figsize=(5, 5))\n reds = y == 0\n blues = y == 1\n ax.scatter(x[reds, 0], x[reds, 1], c=\"red\", s=20, edgecolor=\"k\")\n ax.scatter(x[blues, 0], x[blues, 1], c=\"blue\", s=20, edgecolor=\"k\")\n ax.set_xlabel(\"$x_1$\")\n ax.set_ylabel(\"$x_2$\")",
"_____no_output_____"
],
[
"# Define output labels as quantum state vectors\n\n# def density_matrix(state):\n# \"\"\"Calculates the density matrix representation of a state.\n\n# Args:\n# state (array[complex]): array representing a quantum state vector\n\n# Returns:\n# dm: (array[complex]): array representing the density matrix\n# \"\"\"\n# return state * np.conj(state).T\n\n\nlabel_0 = [[1], [0]]\nlabel_1 = [[0], [1]]\n\n\ndef density_matrix(state):\n \"\"\"Calculates the density matrix representation of a state.\n\n Args:\n state (array[complex]): array representing a quantum state vector\n\n Returns:\n dm: (array[complex]): array representing the density matrix\n \"\"\"\n return np.outer(state, np.conj(state))\n\n#state_labels = [label_0, label_1]\nstate_labels = np.loadtxt('./tetra_states.txt', dtype=np.complex_)",
"_____no_output_____"
],
[
"dev = qml.device(\"default.qubit\", wires=1)\n# Install any pennylane-plugin to run on some particular backend\n\n\[email protected](dev)\ndef qcircuit(params, x=None, y=None):\n \"\"\"A variational quantum circuit representing the Universal classifier.\n\n Args:\n params (array[float]): array of parameters\n x (array[float]): single input vector\n y (array[float]): single output state density matrix\n\n Returns:\n float: fidelity between output state and input\n \"\"\"\n for i in range(len(params[0])):\n for j in range(int(len(x)/3)):\n qml.Rot(*(params[0][i][3*j:3*(j+1)]*x[3*j:3*(j+1)] + params[1][i][3*j:3*(j+1)]), wires=0)\n #qml.Rot(*params[1][i][3*j:3*(j+1)], wires=0)\n return qml.expval(qml.Hermitian(y, wires=[0]))",
"_____no_output_____"
],
[
"X_train[0].shape",
"_____no_output_____"
],
[
"a = np.random.uniform(size=(2, 1, 9))\n\nqcircuit(a, X_train[0], density_matrix(state_labels[3]))",
"_____no_output_____"
],
[
"tetra_class = np.loadtxt('./tetra_class_label.txt')\n\ntetra_class",
"_____no_output_____"
],
[
"binary_class = np.array([[1, 0], [0, 1]])\nbinary_class",
"_____no_output_____"
],
[
"class_labels = tetra_class",
"_____no_output_____"
],
[
"class_labels[0][0]",
"_____no_output_____"
],
[
"dm_labels = [density_matrix(s) for s in state_labels]\n\ndef cost(params, x, y, state_labels=None):\n \"\"\"Cost function to be minimized.\n\n Args:\n params (array[float]): array of parameters\n x (array[float]): 2-d array of input vectors\n y (array[float]): 1-d array of targets\n state_labels (array[float]): array of state representations for labels\n\n Returns:\n float: loss value to be minimized\n \"\"\"\n # Compute prediction for each input in data batch\n loss = 0.0\n for i in range(len(x)):\n f = qcircuit(params, x=x[i], y=dm_labels[y[i]])\n loss = loss + (1 - f) ** 2\n return loss / len(x)\n\n\n# loss = 0.0\n# for i in range(len(x)):\n# f = 0.0\n# for j in range(len(dm_labels)):\n# f += (qcircuit(params, x=x[i], y=dm_labels[j]) - class_labels[y[i]][j])**2\n# loss = loss + f\n# return loss / len(x)",
"_____no_output_____"
],
[
"def test(params, x, y, state_labels=None):\n \"\"\"\n Tests on a given set of data.\n\n Args:\n params (array[float]): array of parameters\n x (array[float]): 2-d array of input vectors\n y (array[float]): 1-d array of targets\n state_labels (array[float]): 1-d array of state representations for labels\n\n Returns:\n predicted (array([int]): predicted labels for test data\n output_states (array[float]): output quantum states from the circuit\n \"\"\"\n fidelity_values = []\n dm_labels = [density_matrix(s) for s in state_labels]\n predicted = []\n\n for i in range(len(x)):\n fidel_function = lambda y: qcircuit(params, x=x[i], y=y)\n fidelities = [fidel_function(dm) for dm in dm_labels]\n best_fidel = np.argmax(fidelities)\n\n predicted.append(best_fidel)\n fidelity_values.append(fidelities)\n\n return np.array(predicted), np.array(fidelity_values)\n\n\ndef accuracy_score(y_true, y_pred):\n \"\"\"Accuracy score.\n\n Args:\n y_true (array[float]): 1-d array of targets\n y_predicted (array[float]): 1-d array of predictions\n state_labels (array[float]): 1-d array of state representations for labels\n\n Returns:\n score (float): the fraction of correctly classified samples\n \"\"\"\n score = y_true == y_pred\n return score.sum() / len(y_true)\n\n\ndef iterate_minibatches(inputs, targets, batch_size):\n \"\"\"\n A generator for batches of the input data\n\n Args:\n inputs (array[float]): input data\n targets (array[float]): targets\n\n Returns:\n inputs (array[float]): one batch of input data of length `batch_size`\n targets (array[float]): one batch of targets of length `batch_size`\n \"\"\"\n for start_idx in range(0, inputs.shape[0] - batch_size + 1, batch_size):\n idxs = slice(start_idx, start_idx + batch_size)\n yield inputs[idxs], targets[idxs]",
"_____no_output_____"
],
[
"# Train using Adam optimizer and evaluate the classifier\nnum_layers = 2\nlearning_rate = 0.1\nepochs = 100\nbatch_size = 32\n\nopt = AdamOptimizer(learning_rate)\n\n# initialize random weights\ntheta = np.random.uniform(size=(num_layers, 18))\nw = np.random.uniform(size=(num_layers, 18))\nparams = [w, theta]\n\npredicted_train, fidel_train = test(params, X_train, Y_train, state_labels)\naccuracy_train = accuracy_score(Y_train, predicted_train)\n\npredicted_test, fidel_test = test(params, X_test, Y_test, state_labels)\naccuracy_test = accuracy_score(Y_test, predicted_test)\n\n# save predictions with random weights for comparison\ninitial_predictions = predicted_test\n\nloss = cost(params, X_test, Y_test, state_labels)\n\nprint(\n \"Epoch: {:2d} | Loss: {:3f} | Train accuracy: {:3f} | Test Accuracy: {:3f}\".format(\n 0, loss, accuracy_train, accuracy_test\n )\n)\n\nfor it in range(epochs):\n for Xbatch, ybatch in iterate_minibatches(X_train, Y_train, batch_size=batch_size):\n params = opt.step(lambda v: cost(v, Xbatch, ybatch, state_labels), params)\n\n predicted_train, fidel_train = test(params, X_train, Y_train, state_labels)\n accuracy_train = accuracy_score(Y_train, predicted_train)\n loss = cost(params, X_train, Y_train, state_labels)\n\n predicted_test, fidel_test = test(params, X_test, Y_test, state_labels)\n accuracy_test = accuracy_score(Y_test, predicted_test)\n res = [it + 1, loss, accuracy_train, accuracy_test]\n print(\n \"Epoch: {:2d} | Loss: {:3f} | Train accuracy: {:3f} | Test accuracy: {:3f}\".format(\n *res\n )\n )",
"Epoch: 0 | Loss: 0.283926 | Train accuracy: 0.241667 | Test Accuracy: 0.226667\n"
],
[
"qml.Rot(*(params[0][0][0:3]*X_train[0, 0:3] + params[1][0][0:3]), wires=[0])",
"_____no_output_____"
],
[
"params[1][0][0:3]",
"_____no_output_____"
]
]
] | [
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] | [
[
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
d0ffb918628aa2b78c106a945bf88185bac62b35 | 4,651 | ipynb | Jupyter Notebook | Ch_Lifted_Heston.ipynb | NicolasMakaroff/implied-volatility-learning | 907dfe4496be35708881f7b40c1b543a8574d649 | [
"MIT"
] | null | null | null | Ch_Lifted_Heston.ipynb | NicolasMakaroff/implied-volatility-learning | 907dfe4496be35708881f7b40c1b543a8574d649 | [
"MIT"
] | 2 | 2020-03-28T11:36:39.000Z | 2020-04-21T14:01:12.000Z | Ch_Lifted_Heston.ipynb | NicolasMakaroff/implied-volatility-learning | 907dfe4496be35708881f7b40c1b543a8574d649 | [
"MIT"
] | 1 | 2020-03-27T17:47:36.000Z | 2020-03-27T17:47:36.000Z | 29.436709 | 125 | 0.503978 | [
[
[
"import numpy as np\nnp.seterr(divide='ignore', invalid='ignore')\nimport scipy.integrate as integrate\nfrom scipy.special import gamma\n\n# Characteristic function of the Lifted Heston model see Slides 85-87\ndef Ch_Lifted_Heston(omega,S0,T,rho,lamb,theta,nu,V0,N,rN,alpha,M):\n # omega = argument of the ch. function\n # S0 = Initial price\n # rho,lamb,theta,nu,V0 = parameters Lifted Heston\n # N = number of factors in the model\n # rN = constant used to define weights and mean-reversions\n # alpha = H+1/2 where H is the Hurst index\n # T = maturity\n # M = number of steps in the time discretization to calculate ch. function\n\n # to make sure we calculate ch. function and not moment gen. function\n i=complex(0,1)\n omega=i*omega\n \n # Definition of weights and mean reversions in the approximation\n h=np.linspace(0,N-1,N)\n rpowerN=np.power(rN,h-N/2) \n # weights\n c=(rN**(1-alpha)-1)*(rpowerN**(1-alpha))/(gamma(alpha)*gamma(2-alpha))\n # mean reversions \n gammas=((1-alpha)/(2-alpha))*((rN**(2-alpha)-1)/(rN**(1-alpha)-1))*rpowerN\n \n # Definition of the initial curve\n g = lambda t: V0+lamb*theta*np.dot(c/gammas,1-np.exp(-t*gammas))\n \n \n # Time steps for the approximation of psi \n delta = T/M;\n t=np.linspace(0,M,M+1)\n t = t * delta\n \n # Function F\n F = lambda u,v : 0.5*(u**2-u)+(rho*nu*u-lamb)*v+.5*nu**2*v**2\n \n \n # Iteration for approximation of psi - see Slide 87\n psi=np.zeros((M+1,N),dtype=complex)\n \n for k in range (1,M+1):\n psi[k,:] = (np.ones(N)/(1+delta*gammas))*(psi[k-1,:]+delta*F(omega,np.dot(c,psi[k-1,:]))*np.ones(N))\n \n \n # Invert g_0 to calculate phi - see Slide 87\n g_0=np.zeros((1,M+1))\n \n for k in range(1,M+2):\n g_0[0,k-1]=g(T-t[k-1])\n \n \n Y=np.zeros((1,M+1),dtype=complex)\n phi=0\n \n Y=F(omega,np.dot(c,psi.transpose()))*g_0\n \n \n # Trapezoid rule to calculate phi\n weights=np.ones(M+1)*delta\n weights[0]=delta/2\n weights[M]=delta/2\n phi=np.dot(weights,Y.transpose())\n \n phi=np.exp(omega*np.log(S0)+phi)\n \n return phi",
"_____no_output_____"
],
[
"def psi_Lifted_Heston(K_,r_,omega,S0,T,rho,lamb,theta,nu,V0,N,rN,alpha,M):\n k_ = np.log(K_)\n phi = Ch_Lifted_Heston(omega,S0,T,rho,lamb,theta,nu,V0,N,rN,alpha,M)\n F = phi*np.exp(-1j*omega.real*k_)\n d = (1+1j*omega.real)*(2+1j*omega.real)\n return np.exp(-r_*T-k_)/np.pi*(F/d).real",
"_____no_output_____"
],
[
"import scipy as scp\n\ndef C_Lifted_Heston(K_,r_,S0,T,rho,lamb,theta,nu,V0,N,rN,alpha,M,L_):\n I = scp.integrate.quad(lambda x: psi_Lifted_Heston(K_,r_,x-2*1j,S0,T,rho,lamb,theta,nu,V0,N,rN,alpha,M) , 0, L_)\n return I[0]",
"_____no_output_____"
],
[
"C_Lifted_Heston(90,0.03,100,0.5,-0.7,2,0.04,0.5,0.04,20,2.5,0.6,100,50)",
"_____no_output_____"
]
]
] | [
"code"
] | [
[
"code",
"code",
"code",
"code"
]
] |
d0ffbe11d67609989d6e6a2176962e84b63923c6 | 175,807 | ipynb | Jupyter Notebook | Matplotlib Pie Chart [4 exercises with solution].ipynb | bibek376/Python-With-Data-Science- | bc7c473b98d7a1f841129762c2d83597b0f8c003 | [
"MIT"
] | null | null | null | Matplotlib Pie Chart [4 exercises with solution].ipynb | bibek376/Python-With-Data-Science- | bc7c473b98d7a1f841129762c2d83597b0f8c003 | [
"MIT"
] | null | null | null | Matplotlib Pie Chart [4 exercises with solution].ipynb | bibek376/Python-With-Data-Science- | bc7c473b98d7a1f841129762c2d83597b0f8c003 | [
"MIT"
] | null | null | null | 686.746094 | 47,992 | 0.953193 | [
[
[
"#Write a Python programming to create a pie chart of the popularity of programming Languages. \nimport matplotlib.pyplot as plt\nimport numpy as np\nimport pandas as pd\nplt.figure(figsize=(8,8))\nlanguages=['Java', 'Python', 'PHP', 'JavaScript','c#','c++']\nPopularity=[22.2, 17.6, 8.8, 8, 7.7, 6.7]\nplt.pie(Popularity,labels=languages,startangle=60,explode=[0.1,0.09,0,0,0,0],autopct=\"%.f\",shadow=True)\nplt.show()",
"_____no_output_____"
],
[
"#Write a Python programming to create a pie chart with a title of the popularity of programming Languages.\nimport matplotlib.pyplot as plt\nimport numpy as np\nimport pandas as pd\nplt.figure(figsize=(8,8))\nlanguages=['Java', 'Python', 'PHP', 'JavaScript','c#','c++']\nPopularity=[22.2, 17.6, 8.8, 8, 7.7, 6.7]\nplt.pie(Popularity,labels=languages,startangle=140,explode=[0.1,0,0,0,0,0],autopct=\"%1.1f%%\",shadow=True)\nplt.title(\"Pie chart example\",bbox={'facecolor':'0.8', 'pad':5})\nplt.show()",
"_____no_output_____"
],
[
"#Write a Python programming to create a pie chart with a title of the popularity of programming Languages. \n#Make multiple wedges of the pie.\nimport matplotlib.pyplot as plt\nimport numpy as np\nimport pandas as pd\nplt.figure(figsize=(8,8))\nlanguages=['Java', 'Python', 'PHP', 'JavaScript','c#','c++']\nPopularity=[22.2, 17.6, 8.8, 8, 7.7, 6.7]\nplt.pie(Popularity,labels=languages,startangle=140,explode=[0.1,0,0,0,0,0.1],autopct=\"%1.1f%%\",shadow=True)\nplt.title(\"Pie chart example\",bbox={'facecolor':'0.8', 'pad':5})\nplt.show()\n\n",
"_____no_output_____"
],
[
"#Write a Python programming to create a pie chart of gold medal achievements of five most successful\n#countries in 2016 Summer Olympics. Read the data from a csv file.\nimport matplotlib.pyplot as plt\nimport numpy as np\nimport pandas as pd\nwith open(\"medal.csv\",\"r\") as f:\n x=f.read()\nprint(x)\n#read method is not possible because of spaces.",
"country,gold_medal\nUnited States,46\nGreat Britain,27\nChina,26\nRussia,19\nGermany,17\n"
],
[
"plt.figure(figsize=(10,6))\ndf=pd.read_csv(\"medal.csv\")\ncountry_name=df['country']\ngold_medals=df['gold_medal']\nplt.pie(gold_medals,labels=country_name,autopct=\"%.f\",explode=[0.1,0,0,0,0])\nplt.title(\"Gold medal in olympic\")\nplt.show()",
"_____no_output_____"
],
[
"import matplotlib.pyplot as plt\nwith open(\"test.txt\") as f:\n data = f.read()\nprint(data)\ndata=data.split('\\n')\nprint(\"----------\")\nprint(data)\nprint(\"----------------\")\n# for i in zip(*data):\n# print(i)\nprint(data[0])\nx=[a.split(' ')[0] for a in x]\nx",
"1 2\n2 4\n3 1\n----------\n['1 2', '2 4', '3 1']\n----------------\n1 2\n"
],
[
"import matplotlib.pyplot as plt\nimport numpy\nimport pandas as pd\nlabels = 'Frogs', 'Hogs', 'Dogs'\nsizes = numpy.array([5860, 677, 3200])\ncolors = ['yellowgreen', 'gold', 'lightskyblue']\n\np, tx, autotexts = plt.pie(sizes, labels=labels, colors=colors,\n autopct=\"\", shadow=True)\n\nfor i, a in enumerate(autotexts):\n a.set_text(\"{}\".format(sizes[i]))\n\nplt.axis('equal')\nplt.show()",
"_____no_output_____"
]
]
] | [
"code"
] | [
[
"code",
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
d0ffc0d2a64a35c2c73f2e563579bb8474cd85f4 | 70,128 | ipynb | Jupyter Notebook | structure/deep_learning_models/StellarGraph_DGCNN_Coarse.ipynb | andrewmagill/fakenews | f3fa9b783c8fa29348aef9969cc4ac176f97e5d0 | [
"Apache-2.0"
] | 1 | 2022-01-15T11:06:53.000Z | 2022-01-15T11:06:53.000Z | structure/deep_learning_models/StellarGraph_DGCNN_Coarse.ipynb | andrewmagill/fakenews | f3fa9b783c8fa29348aef9969cc4ac176f97e5d0 | [
"Apache-2.0"
] | null | null | null | structure/deep_learning_models/StellarGraph_DGCNN_Coarse.ipynb | andrewmagill/fakenews | f3fa9b783c8fa29348aef9969cc4ac176f97e5d0 | [
"Apache-2.0"
] | 1 | 2020-12-14T18:42:40.000Z | 2020-12-14T18:42:40.000Z | 114.588235 | 37,308 | 0.783638 | [
[
[
"import pandas as pd\nimport numpy as np\n\nimport stellargraph as sg\nfrom stellargraph.mapper import PaddedGraphGenerator\nfrom stellargraph.layer import DeepGraphCNN\nfrom stellargraph import StellarGraph\n\nfrom stellargraph import datasets\n\nfrom sklearn import model_selection\nfrom IPython.display import display, HTML\n\nfrom tensorflow.keras import Model\nfrom tensorflow.keras.optimizers import Adam\nfrom tensorflow.keras.layers import Dense, Conv1D, MaxPool1D, Dropout, Flatten\nfrom tensorflow.keras.losses import binary_crossentropy\nimport tensorflow as tf",
"_____no_output_____"
],
[
"conspiracy_5G_path = '/Users/maria/Desktop/twitterAnalysis/FakeNews/dataset/graphs/5g_corona_conspiracy/'\nconspiracy_other_path = '/Users/maria/Desktop/twitterAnalysis/FakeNews/dataset/graphs/non_conspiracy/'\nnon_conspiracy_path = '/Users/maria/Desktop/twitterAnalysis/FakeNews/dataset/graphs/other_conspiracy/'\ntest_graphs_path = '/Users/maria/Desktop/twitterAnalysis/FakeNews/dataset/graphs/test_graphs/'\n\nconspiracy_5G_N = 270\nconspiracy_other_N = 1660\nnon_conspiracy_N = 397\ntest_graphs_N = 1165",
"_____no_output_____"
],
[
"conspiracy_5G = list()\nfor i in range(conspiracy_5G_N):\n g_id = i+1\n nodes_path = conspiracy_5G_path + str(g_id) + '/nodes.csv'\n edges_path = conspiracy_5G_path + str(g_id) + '/edges.txt'\n g_nodes = pd.read_csv(nodes_path) \n g_nodes = g_nodes.set_index('id') \n g_edges = pd.read_csv(edges_path, header = None, sep=' ')\n g_edges = g_edges.rename(columns={0: 'source', 1: 'target'})\n g = StellarGraph(g_nodes, edges=g_edges)\n conspiracy_5G.append(g)",
"_____no_output_____"
],
[
"conspiracy_other = list()\nfor i in range(conspiracy_other_N):\n g_id = i+1\n nodes_path = conspiracy_other_path + str(g_id) + '/nodes.csv'\n edges_path = conspiracy_other_path + str(g_id) + '/edges.txt'\n g_nodes = pd.read_csv(nodes_path)\n g_nodes = g_nodes.set_index('id')\n g_edges = pd.read_csv(edges_path, header = None, sep=' ')\n g_edges = g_edges.rename(columns={0: 'source', 1: 'target'})\n g = StellarGraph(g_nodes, edges=g_edges)\n conspiracy_other.append(g)",
"_____no_output_____"
],
[
"non_conspiracy = list()\nfor i in range(non_conspiracy_N):\n g_id = i+1\n nodes_path = non_conspiracy_path + str(g_id) + '/nodes.csv'\n edges_path = non_conspiracy_path + str(g_id) + '/edges.txt'\n g_nodes = pd.read_csv(nodes_path) \n g_nodes = g_nodes.set_index('id')\n g_edges = pd.read_csv(edges_path, header = None, sep=' ')\n g_edges = g_edges.rename(columns={0: 'source', 1: 'target'})\n g = StellarGraph(g_nodes, edges=g_edges)\n non_conspiracy.append(g)",
"_____no_output_____"
],
[
"test_graphs_off = list()\nfor i in range(test_graphs_N):\n g_id = i+1\n nodes_path = test_graphs_path + str(g_id) + '/nodes.csv'\n edges_path = test_graphs_path + str(g_id) + '/edges.txt'\n g_nodes = pd.read_csv(nodes_path) \n g_nodes = g_nodes.set_index('id')\n g_edges = pd.read_csv(edges_path, header = None, sep=' ')\n g_edges = g_edges.rename(columns={0: 'source', 1: 'target'})\n g = StellarGraph(g_nodes, edges=g_edges)\n test_graphs_off.append(g)",
"_____no_output_____"
],
[
"graphs = conspiracy_5G + conspiracy_other + non_conspiracy\ngraph_labels = pd.Series(np.repeat([1, -1], [conspiracy_5G_N, conspiracy_other_N+non_conspiracy_N], axis=0))",
"_____no_output_____"
],
[
"graph_labels = pd.get_dummies(graph_labels, drop_first=True)",
"_____no_output_____"
],
[
"generator = PaddedGraphGenerator(graphs=graphs)",
"_____no_output_____"
],
[
"k = 35 # the number of rows for the output tensor\nlayer_sizes = [32, 32, 32, 1]\n\ndgcnn_model = DeepGraphCNN(\n layer_sizes=layer_sizes,\n activations=[\"tanh\", \"tanh\", \"tanh\", \"tanh\"],\n k=k,\n bias=False,\n generator=generator,\n)\nx_inp, x_out = dgcnn_model.in_out_tensors()",
"_____no_output_____"
],
[
"x_out = Conv1D(filters=16, kernel_size=sum(layer_sizes), strides=sum(layer_sizes))(x_out)\nx_out = MaxPool1D(pool_size=2)(x_out)\n\nx_out = Conv1D(filters=32, kernel_size=5, strides=1)(x_out)\n\nx_out = Flatten()(x_out)\n\nx_out = Dense(units=128, activation=\"relu\")(x_out)\nx_out = Dropout(rate=0.5)(x_out)\n\npredictions = Dense(units=1, activation=\"sigmoid\")(x_out)",
"_____no_output_____"
],
[
"model = Model(inputs=x_inp, outputs=predictions)\n\nmodel.compile(\n optimizer=Adam(lr=0.0001), loss=binary_crossentropy, metrics=[\"acc\"],\n)",
"_____no_output_____"
],
[
"train_graphs, test_graphs = model_selection.train_test_split(\n graph_labels, train_size=0.9, test_size=None, stratify=graph_labels,\n)",
"_____no_output_____"
],
[
"gen = PaddedGraphGenerator(graphs=graphs)\n\ntrain_gen = gen.flow(\n list(train_graphs.index - 1),\n targets=train_graphs.values,\n batch_size=50,\n symmetric_normalization=False,\n)\n\ntest_gen = gen.flow(\n list(test_graphs.index - 1),\n targets=test_graphs.values,\n batch_size=1,\n symmetric_normalization=False,\n)",
"_____no_output_____"
],
[
"epochs = 100\nhistory = model.fit(\n train_gen, epochs=epochs, verbose=1, validation_data=test_gen, shuffle=True,\n)",
"WARNING:tensorflow:sample_weight modes were coerced from\n ...\n to \n ['...']\nWARNING:tensorflow:sample_weight modes were coerced from\n ...\n to \n ['...']\nTrain for 42 steps, validate for 233 steps\nEpoch 1/100\n"
],
[
"sg.utils.plot_history(history)",
"_____no_output_____"
],
[
"test_gen_off = PaddedGraphGenerator(graphs=test_graphs_off)\ntest_gen_off_f = test_gen_off.flow(graphs=test_graphs_off)\npreds = model.predict(test_gen_off_f)",
"_____no_output_____"
],
[
"import matplotlib.pyplot as plt\nplt.hist(preds)",
"_____no_output_____"
],
[
"print(preds)",
"[[0.12391277]\n [0.13464499]\n [0.16950193]\n ...\n [0.25458154]\n [0.14482737]\n [0.11278567]]\n"
],
[
"# Sources\n\n# https://stellargraph.readthedocs.io/en/stable/demos/graph-classification/dgcnn-graph-classification.html",
"_____no_output_____"
]
]
] | [
"code"
] | [
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
d0ffcf2214c2f67dc9eae68c36b45cf35a763576 | 75,366 | ipynb | Jupyter Notebook | train.ipynb | TMYuan/video-interpolation | 9f52ebf519a33756939b1a6f5c4e83e3270be578 | [
"MIT"
] | null | null | null | train.ipynb | TMYuan/video-interpolation | 9f52ebf519a33756939b1a6f5c4e83e3270be578 | [
"MIT"
] | null | null | null | train.ipynb | TMYuan/video-interpolation | 9f52ebf519a33756939b1a6f5c4e83e3270be578 | [
"MIT"
] | null | null | null | 30.170536 | 134 | 0.513109 | [
[
[
"import logging\nimport torch\nimport torch.optim as optim\nimport torch.nn as nn\nimport torch.nn.functional as F\nimport argparse\nimport os\nimport random\nimport numpy as np\nfrom torch.autograd import Variable\nfrom torch.utils.data import DataLoader\nimport utils\nimport itertools\nfrom tqdm import tqdm_notebook\nimport models.dcgan_unet_64 as dcgan_unet_models\nimport models.dcgan_64 as dcgan_models\nimport models.classifiers as classifiers\nimport models.my_model as my_model\nfrom data.moving_mnist import MovingMNIST",
"_____no_output_____"
],
[
"torch.cuda.set_device(0)",
"_____no_output_____"
]
],
[
[
"Constant definition",
"_____no_output_____"
]
],
[
[
"np.random.seed(1)\nrandom.seed(1)\ntorch.manual_seed(1)\ntorch.cuda.manual_seed_all(1)\ndevice = torch.device(\"cuda\" if torch.cuda.is_available() else \"cpu\")",
"_____no_output_____"
],
[
"lr = 2e-3\nseq_len = 12\nbeta1 = 0.5\ncontent_dim = 128\npose_dim = 10\nchannels = 3\nnormalize = False\nsd_nf = 100\nimage_width = 64\nbatch_size = 100\nlog_dir = './logs/0522_my_model_CVAE_ourDisc_newPair/'\nos.makedirs(os.path.join(log_dir, 'rec'), exist_ok=True)\nos.makedirs(os.path.join(log_dir, 'analogy'), exist_ok=True)\nlogging.basicConfig(filename=os.path.join(log_dir, 'record.txt'), level=logging.DEBUG)",
"_____no_output_____"
]
],
[
[
"Data Loader",
"_____no_output_____"
]
],
[
[
"train_data = MovingMNIST(True, '../data_uni/', seq_len=seq_len)\ntest_data = MovingMNIST(False, '../data_uni/', seq_len=seq_len)\n\ntrain_loader = DataLoader(\n train_data,\n batch_size=batch_size,\n num_workers=16,\n shuffle=True,\n drop_last=True,\n pin_memory=True\n)\ntest_loader = DataLoader(\n test_data,\n batch_size=batch_size,\n num_workers=0,\n shuffle=True,\n drop_last=True,\n pin_memory=True\n)",
"_____no_output_____"
]
],
[
[
"Model definition",
"_____no_output_____"
]
],
[
[
"# # netEC = dcgan_unet_models.content_encoder(content_dim, channels).to(device)\n# netEC = dcgan_models.content_encoder(content_dim, channels).to(device)\n# netEP = dcgan_models.pose_encoder(pose_dim, channels).to(device)\n# # netD = dcgan_unet_models.decoder(content_dim, pose_dim, channels).to(device)\n# netD = dcgan_models.decoder(content_dim, pose_dim, channels).to(device)\n# netC = classifiers.scene_discriminator(pose_dim, sd_nf).to(device)\n\nnetEC = my_model.content_encoder(content_dim, channels).to(device)\nnetEP = my_model.pose_encoder(pose_dim, channels, conditional=True).to(device)\nnetD = my_model.decoder(content_dim, pose_dim, channels).to(device)\n# netC = my_model.scene_discriminator(pose_dim, sd_nf).to(device)\nnetC = my_model.Discriminator(channels).to(device)\n\nnetEC.apply(utils.weights_init)\nnetEP.apply(utils.weights_init)\nnetD.apply(utils.weights_init)\nnetC.apply(utils.weights_init)\n\nprint(netEC)\nprint(netEP)\nprint(netD)\nprint(netC)",
"content_encoder(\n (main): Sequential(\n (0): dcgan_conv(\n (main): Sequential(\n (0): Conv2d(3, 64, kernel_size=(4, 4), stride=(2, 2), padding=(1, 1), bias=False)\n (1): BatchNorm2d(64, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)\n (2): ReLU(inplace)\n )\n )\n (1): dcgan_conv(\n (main): Sequential(\n (0): Conv2d(64, 128, kernel_size=(4, 4), stride=(2, 2), padding=(1, 1), bias=False)\n (1): BatchNorm2d(128, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)\n (2): ReLU(inplace)\n )\n )\n (2): dcgan_conv(\n (main): Sequential(\n (0): Conv2d(128, 256, kernel_size=(4, 4), stride=(2, 2), padding=(1, 1), bias=False)\n (1): BatchNorm2d(256, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)\n (2): ReLU(inplace)\n )\n )\n (3): dcgan_conv(\n (main): Sequential(\n (0): Conv2d(256, 512, kernel_size=(4, 4), stride=(2, 2), padding=(1, 1), bias=False)\n (1): BatchNorm2d(512, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)\n (2): ReLU(inplace)\n )\n )\n (4): Conv2d(512, 128, kernel_size=(4, 4), stride=(1, 1))\n (5): BatchNorm2d(128, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)\n (6): Tanh()\n )\n)\npose_encoder(\n (c1): dcgan_conv(\n (main): Sequential(\n (0): Conv2d(5, 64, kernel_size=(4, 4), stride=(2, 2), padding=(1, 1), bias=False)\n (1): BatchNorm2d(64, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)\n (2): ReLU(inplace)\n )\n )\n (c2): dcgan_conv(\n (main): Sequential(\n (0): Conv2d(66, 128, kernel_size=(4, 4), stride=(2, 2), padding=(1, 1), bias=False)\n (1): BatchNorm2d(128, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)\n (2): ReLU(inplace)\n )\n )\n (c3): dcgan_conv(\n (main): Sequential(\n (0): Conv2d(130, 256, kernel_size=(4, 4), stride=(2, 2), padding=(1, 1), bias=False)\n (1): BatchNorm2d(256, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)\n (2): ReLU(inplace)\n )\n )\n (c4): dcgan_conv(\n (main): Sequential(\n (0): Conv2d(258, 512, kernel_size=(4, 4), stride=(2, 2), padding=(1, 1), bias=False)\n (1): BatchNorm2d(512, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)\n (2): ReLU(inplace)\n )\n )\n (c5): Sequential(\n (0): Conv2d(514, 10, kernel_size=(4, 4), stride=(1, 1))\n (1): BatchNorm2d(10, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)\n (2): Tanh()\n )\n)\ndecoder(\n (main): Sequential(\n (0): ConvTranspose2d(138, 512, kernel_size=(4, 4), stride=(1, 1), bias=False)\n (1): BatchNorm2d(512, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)\n (2): ReLU(inplace)\n (3): dcgan_upconv(\n (main): Sequential(\n (0): ConvTranspose2d(512, 256, kernel_size=(4, 4), stride=(2, 2), padding=(1, 1), bias=False)\n (1): BatchNorm2d(256, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)\n (2): ReLU(inplace)\n )\n )\n (4): dcgan_upconv(\n (main): Sequential(\n (0): ConvTranspose2d(256, 128, kernel_size=(4, 4), stride=(2, 2), padding=(1, 1), bias=False)\n (1): BatchNorm2d(128, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)\n (2): ReLU(inplace)\n )\n )\n (5): dcgan_upconv(\n (main): Sequential(\n (0): ConvTranspose2d(128, 64, kernel_size=(4, 4), stride=(2, 2), padding=(1, 1), bias=False)\n (1): BatchNorm2d(64, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)\n (2): ReLU(inplace)\n )\n )\n (6): ConvTranspose2d(64, 3, kernel_size=(4, 4), stride=(2, 2), padding=(1, 1))\n (7): Sigmoid()\n )\n)\nDiscriminator(\n (main): Sequential(\n (0): Conv2d(6, 64, kernel_size=(4, 4), stride=(2, 2), padding=(1, 1), bias=False)\n (1): LeakyReLU(negative_slope=0.2, inplace)\n (2): Conv2d(64, 128, kernel_size=(4, 4), stride=(2, 2), padding=(1, 1), bias=False)\n (3): BatchNorm2d(128, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)\n (4): LeakyReLU(negative_slope=0.2, inplace)\n (5): Conv2d(128, 256, kernel_size=(4, 4), stride=(2, 2), padding=(1, 1), bias=False)\n (6): BatchNorm2d(256, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)\n (7): LeakyReLU(negative_slope=0.2, inplace)\n (8): Conv2d(256, 512, kernel_size=(4, 4), stride=(2, 2), padding=(1, 1), bias=False)\n (9): BatchNorm2d(512, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)\n (10): LeakyReLU(negative_slope=0.2, inplace)\n (11): Conv2d(512, 1, kernel_size=(4, 4), stride=(1, 1), bias=False)\n (12): Sigmoid()\n )\n)\n"
],
[
"optimizerEC = optim.Adam(netEC.parameters(), lr=lr, betas=(beta1, 0.999))\noptimizerEP = optim.Adam(netEP.parameters(), lr=lr, betas=(beta1, 0.999))\noptimizerD = optim.Adam(netD.parameters(), lr=lr, betas=(beta1, 0.999))\noptimizerC = optim.Adam(netC.parameters(), lr=lr, betas=(beta1, 0.999))",
"_____no_output_____"
]
],
[
[
"Plot function",
"_____no_output_____"
]
],
[
[
"# --------- plotting funtions ------------------------------------\ndef plot_rec(x, epoch, dtype):\n x_c = x[0]\n x_p = x[np.random.randint(1, len(x))]\n\n h_c = netEC(x_c)\n h_p = netEP(x_p, h_c)\n rec = netD([h_c, h_p])\n\n x_c, x_p, rec = x_c.data, x_p.data, rec.data\n fname = '{}-{}.png'.format(dtype, epoch)\n fname = os.path.join(log_dir, 'rec', fname)\n to_plot = []\n row_sz = 5\n nplot = 20\n for i in range(0, nplot-row_sz, row_sz):\n row = [[xc, xp, xr] for xc, xp, xr in zip(x_c[i:i+row_sz], x_p[i:i+row_sz], rec[i:i+row_sz])]\n to_plot.append(list(itertools.chain(*row)))\n utils.save_tensors_image(fname, to_plot)\n\ndef plot_analogy(x, epoch, dtype):\n x_c = x[0]\n h_c = netEC(x_c)\n \n nrow = 10\n row_sz = len(x)\n to_plot = []\n row = [xi[0].data for xi in x]\n zeros = torch.zeros(channels, image_width, image_width)\n to_plot.append([zeros] + row)\n for i in range(nrow):\n to_plot.append([x[0][i].data])\n\n for j in range(0, row_sz):\n # for each time step\n h_p = netEP(x[j], h_c).data\n # first 10 pose vector, equal to first pose vector\n for i in range(nrow):\n h_p[i] = h_p[0]\n rec = netD([h_c, h_p])\n for i in range(nrow):\n to_plot[i+1].append(rec[i].data.clone())\n\n fname = '{}-{}.png'.format(dtype, epoch)\n fname = os.path.join(log_dir, 'analogy', fname)\n utils.save_tensors_image(fname, to_plot)",
"_____no_output_____"
]
],
[
[
"Training function",
"_____no_output_____"
]
],
[
[
"def train(x):\n optimizerEC.zero_grad()\n optimizerEP.zero_grad()\n optimizerD.zero_grad()\n\n# x_c1 = x[0]\n# x_c2 = x[1]\n# x_p1 = x[2]\n# x_p2 = x[3]\n x_c1 = x[np.random.randint(len(x))]\n x_c2 = x[np.random.randint(len(x))]\n x_p1 = x[np.random.randint(len(x))]\n x_p2 = x[np.random.randint(len(x))]\n\n h_c1 = netEC(x_c1)\n# h_c2 = netEC(x_c2)[0].detach()\n h_c2 = netEC(x_c2).detach()\n h_p1 = netEP(x_p1, h_c1.detach()) # used for scene discriminator\n h_p2 = netEP(x_p2, h_c1.detach())\n\n\n # similarity loss: ||h_c1 - h_c2||\n# sim_loss = F.mse_loss(h_c1[0], h_c2)\n sim_loss = F.mse_loss(h_c1, h_c2)\n\n\n # reconstruction loss: ||D(h_c1, h_p1), x_p1|| \n rec = netD([h_c1, h_p1])\n rec_loss = F.mse_loss(rec, x_p1)\n\n # scene discriminator loss: maximize entropy of output\n # target = torch.FloatTensor(batch_size, 1).fill_(0.5).to(device)\n # out = netC([h_p1, h_p2])\n # sd_loss = F.binary_cross_entropy(out, target)\n \n # Swap pose vector to train the discriminator\n target = torch.FloatTensor(batch_size, 1).fill_(1).to(device)\n idx = torch.randperm(batch_size)\n h_p2 = h_p2[idx]\n rec_swap = netD([h_c1, h_p2])\n out = netC([x_c1.detach(), rec_swap]).view(-1, 1)\n D_G_fake = out.mean().item()\n adv_loss = F.binary_cross_entropy(out, target)\n \n # full loss\n loss = sim_loss + rec_loss + 0.1 * adv_loss\n loss.backward()\n\n optimizerEC.step()\n optimizerEP.step()\n optimizerD.step()\n\n return sim_loss.item(), rec_loss.item(), adv_loss.item(), D_G_fake",
"_____no_output_____"
],
[
"def train_scene_discriminator(x):\n optimizerC.zero_grad()\n\n target = torch.FloatTensor(batch_size, 1).to(device)\n \n # condition\n h_c = netEC(x[np.random.randint(len(x))]).detach()\n \n x1 = x[0]\n x2 = x[1]\n h_p1 = netEP(x1, h_c).detach()\n h_p2 = netEP(x2, h_c).detach()\n\n half = batch_size // 2\n rp = torch.randperm(half).cuda()\n h_p2[:half] = h_p2[rp]\n target[:half] = 0\n target[half:] = 1\n\n out = netC([h_p1, h_p2])\n bce = F.binary_cross_entropy(out, target)\n\n bce.backward()\n optimizerC.step()\n\n acc =out[:half].le(0.5).sum() + out[half:].gt(0.5).sum()\n return bce.data.cpu().numpy(), acc.data.cpu().numpy() / batch_size",
"_____no_output_____"
],
[
"def train_discriminator(x):\n optimizerC.zero_grad()\n\n real_lbl = torch.FloatTensor(batch_size, 1).fill_(1).to(device)\n fake_lbl = torch.FloatTensor(batch_size, 1).fill_(0).to(device)\n \n x1 = x[np.random.randint(len(x))]\n x2 = x[np.random.randint(len(x))]\n x3 = x[np.random.randint(len(x))]\n\n # real pair\n # 1. x1\n # 2. reconstructed frames by pose(x2) and content(x1)\n h_c = netEC(x1).detach()\n h_p = netEP(x3, h_c)\n x_rec = netD([h_c, h_p]).detach()\n out_real = netC([x1, x_rec]).view(-1, 1)\n loss_real = F.binary_cross_entropy(out_real, real_lbl)\n D_real = loss_real.mean().item()\n \n # fake pair\n # 1. x1\n # 2. swapped reconstructed frames\n # by swapped pose(x3) and content(x1)\n idx = torch.randperm(batch_size)\n h_p = netEP(x3, h_c)\n h_p = h_p[idx]\n x_swap = netD([h_c, h_p]).detach()\n out_fake = netC([x1, x_swap]).view(-1, 1)\n loss_fake = F.binary_cross_entropy(out_fake, fake_lbl)\n D_fake = loss_fake.mean().item()\n \n \n bce = 0.5*loss_real + 0.5*loss_fake\n bce.backward()\n optimizerC.step()\n\n \n return bce.item(), D_real, D_fake",
"_____no_output_____"
],
[
"epoch_size = len(train_loader)\ntest_x = next(iter(test_loader))\ntest_x = torch.transpose(test_x, 0, 1)\ntest_x = test_x.to(device)\n\nfor epoch in tqdm_notebook(range(200), desc='EPOCH'):\n netEP.train()\n netEC.train()\n netD.train()\n netC.train()\n epoch_sim_loss, epoch_rec_loss, epoch_adv_loss, epoch_sd_loss = 0, 0, 0, 0\n epoch_D_real, epoch_D_fake, epoch_D_G_fake = 0, 0, 0\n \n for i, x in enumerate(tqdm_notebook(train_loader, desc='BATCH')):\n # x to device\n x = torch.transpose(x, 0, 1)\n x = x.to(device)\n \n # train scene discriminator\n # sd_loss, sd_acc = train_scene_discriminator(x)\n sd_loss, D_real, D_fake = train_discriminator(x)\n epoch_sd_loss += sd_loss\n epoch_D_real += D_real\n epoch_D_fake += D_fake\n \n # train main model\n sim_loss, rec_loss, adv_loss, D_G_fake = train(x)\n epoch_sim_loss += sim_loss\n epoch_rec_loss += rec_loss\n epoch_adv_loss += adv_loss\n epoch_D_G_fake += D_G_fake\n \n log_str='[%02d]rec loss: %.4f |sim loss: %.4f|adv loss: %.4f |sd loss: %.4f \\\n|D(real): %.2f |D(fake): %.2f |D(G(fake)): %.2f' %\\\n (epoch,\n epoch_rec_loss/epoch_size,\n epoch_sim_loss/epoch_size,\n epoch_adv_loss/epoch_size,\n epoch_sd_loss/epoch_size,\n epoch_D_real/epoch_size,\n epoch_D_fake/epoch_size,\n epoch_D_G_fake/epoch_size)\n \n print(log_str)\n logging.info(log_str)\n \n netEP.eval()\n netEC.eval()\n netD.eval()\n \n with torch.no_grad():\n plot_rec(test_x, epoch, 'test')\n plot_analogy(test_x, epoch, 'test')\n\n # save the model\n torch.save({\n 'netD': netD,\n 'netEP': netEP,\n 'netEC': netEC},\n '%s/model.pth' % log_dir)",
"_____no_output_____"
],
[
"len(train_loader)",
"_____no_output_____"
],
[
"for i, x in enumerate(train_loader):\n if i == 0:\n with torch.no_grad():\n x = torch.transpose(x, 0, 1)\n x = x.to(device)\n plot_rec(x, 200)\n plot_analogy(x, 200)",
"_____no_output_____"
]
]
] | [
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] | [
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
d0ffdd511542d810ebb6609581f05d1f5ad8f6da | 6,842 | ipynb | Jupyter Notebook | lessons/Chapter5/03_Hcurl_elliptic_problem.ipynb | pyccel/IGA-Python | e3604ba3d76a20e3d30ed3c7c952dcd2dc8147bb | [
"MIT"
] | 2 | 2022-01-21T08:51:30.000Z | 2022-03-17T12:14:02.000Z | lessons/Chapter5/03_Hcurl_elliptic_problem.ipynb | pyccel/IGA-Python | e3604ba3d76a20e3d30ed3c7c952dcd2dc8147bb | [
"MIT"
] | null | null | null | lessons/Chapter5/03_Hcurl_elliptic_problem.ipynb | pyccel/IGA-Python | e3604ba3d76a20e3d30ed3c7c952dcd2dc8147bb | [
"MIT"
] | 1 | 2022-03-01T06:41:54.000Z | 2022-03-01T06:41:54.000Z | 31.242009 | 260 | 0.539901 | [
[
[
"# $H(curl, \\Omega)$ Elliptic Problems",
"_____no_output_____"
],
[
"$\\newcommand{\\dd}{\\,{\\rm d}}$\n$\\newcommand{\\uu}{\\mathbf{u}}$\n$\\newcommand{\\vv}{\\mathbf{v}}$\n$\\newcommand{\\nn}{\\mathbf{n}}$\n$\\newcommand{\\ff}{\\mathbf{f}}$\n$\\newcommand{\\Hcurlzero}{\\mathbf{H}_0(\\mbox{curl}, \\Omega)}$\n$\\newcommand{\\Curl}{\\nabla \\times}$\n\nLet $\\Omega \\subset \\mathbb{R}^d$ be an open Liptschitz bounded set, and we look for the solution of the following problem\n\\begin{align}\n \\left\\{ \n \\begin{array}{rl}\n \\Curl \\Curl \\uu + \\mu \\uu &= \\ff, \\quad \\Omega \n \\\\\n \\uu \\times \\nn &= 0, \\quad \\partial\\Omega\n \\end{array} \\right.\n \\label{eq:elliptic_hcurl}\n\\end{align}\nwhere $\\ff \\in \\mathbf{L}^2(\\Omega)$, $\\mu \\in L^\\infty(\\Omega)$ and there exists $\\mu_0 > 0$ such that $\\mu \\geq \\mu_0$ almost everywhere.\nWe take the Hilbert space $V := \\Hcurlzero$, in which case the variational formulation corresponding to \\eqref{eq:elliptic_hcurl} writes \n\n---\nFind $\\uu \\in V$ such that\n\\begin{align}\n a(\\uu,\\vv) = l(\\vv) \\quad \\forall \\vv \\in V \n\\label{eq:abs_var_elliptic_hcurl}\n\\end{align}\nwhere \n\\begin{align}\n\\left\\{ \n\\begin{array}{rll}\na(\\uu, \\vv) &:= \\int_{\\Omega} \\Curl \\uu \\cdot \\Curl \\vv + \\int_{\\Omega} \\mu \\uu \\cdot \\vv, & \\forall \\uu, \\vv \\in V \\\\\nl(\\vv) &:= \\int_{\\Omega} \\vv \\cdot \\ff, & \\forall \\vv \\in V \n\\end{array} \\right.\n\\label{tcb:elliptic_hcurl}\n\\end{align}\n\n---\n\nWe recall that in $\\Hcurlzero$, the bilinear form $a$ is equivalent to the inner product and is therefor continuous and coercive. Hence, our abstract theory applies and there exists a unique solution to the problem \\eqref{eq:abs_var_elliptic_hcurl}.\n\n",
"_____no_output_____"
]
],
[
[
"import numpy as np\nfrom sympy import pi, cos, sin, sqrt, Matrix, Tuple, lambdify\nfrom scipy.sparse.linalg import spsolve\nfrom scipy.sparse.linalg import gmres as sp_gmres\nfrom scipy.sparse.linalg import minres as sp_minres\nfrom scipy.sparse.linalg import cg as sp_cg\nfrom scipy.sparse.linalg import bicg as sp_bicg\nfrom scipy.sparse.linalg import bicgstab as sp_bicgstab\n\nfrom sympde.calculus import grad, dot, inner, div, curl, cross\nfrom sympde.topology import NormalVector\nfrom sympde.topology import ScalarFunctionSpace, VectorFunctionSpace\nfrom sympde.topology import ProductSpace\nfrom sympde.topology import element_of, elements_of\nfrom sympde.topology import Square\nfrom sympde.expr import BilinearForm, LinearForm, integral\nfrom sympde.expr import Norm\nfrom sympde.expr import find, EssentialBC\n\nfrom psydac.fem.basic import FemField\nfrom psydac.fem.vector import ProductFemSpace\nfrom psydac.api.discretization import discretize\nfrom psydac.linalg.utilities import array_to_stencil\nfrom psydac.linalg.iterative_solvers import pcg, bicg",
"_____no_output_____"
],
[
"# ... abstract model\ndomain = Square('A')\nB_dirichlet_0 = domain.boundary\n\nx, y = domain.coordinates\n\nalpha = 1.\nuex = Tuple(sin(pi*y), sin(pi*x)*cos(pi*y))\nf = Tuple(alpha*sin(pi*y) - pi**2*sin(pi*y)*cos(pi*x) + pi**2*sin(pi*y),\n alpha*sin(pi*x)*cos(pi*y) + pi**2*sin(pi*x)*cos(pi*y))\n\nV = VectorFunctionSpace('V', domain, kind='hcurl')\n\nu = element_of(V, name='u')\nv = element_of(V, name='v')\nF = element_of(V, name='F')\n\n# Bilinear form a: V x V --> R\na = BilinearForm((u, v), integral(domain, curl(u)*curl(v) + alpha*dot(u,v)))\n\nnn = NormalVector('nn')\na_bc = BilinearForm((u, v), integral(domain.boundary, 1e30 * cross(u, nn) * cross(v, nn)))\n\n\n# Linear form l: V --> R\nl = LinearForm(v, integral(domain, dot(f,v)))\n\n# l2 error\nerror = Matrix([F[0]-uex[0],F[1]-uex[1]])\nl2norm = Norm(error, domain, kind='l2')",
"_____no_output_____"
],
[
"ncells = [2**3, 2**3]\ndegree = [2, 2]",
"_____no_output_____"
],
[
"# Create computational domain from topological domain\ndomain_h = discretize(domain, ncells=ncells)\n\n# Discrete spaces\nVh = discretize(V, domain_h, degree=degree)\n\n# Discretize bi-linear and linear form\na_h = discretize(a, domain_h, [Vh, Vh])\na_bc_h = discretize(a_bc, domain_h, [Vh, Vh])\n\nl_h = discretize(l, domain_h, Vh)\nl2_norm_h = discretize(l2norm, domain_h, Vh)\n\nM = a_h.assemble() + a_bc_h.assemble()\nb = l_h.assemble()",
"_____no_output_____"
],
[
"# Solve linear system\nsol, info = pcg(M ,b, pc='jacobi', tol=1e-8)\n\nuh = FemField( Vh, sol )\nl2_error = l2_norm_h.assemble(F=uh)\nprint(l2_error)",
"0.0029394893438220407\n"
]
]
] | [
"markdown",
"code"
] | [
[
"markdown",
"markdown"
],
[
"code",
"code",
"code",
"code",
"code"
]
] |
d0ffe39e2f574c89be60b5d6d7e5f9b5e855ce98 | 605,826 | ipynb | Jupyter Notebook | notebooks_examples/emotion classification using SVMs in scikit-learn.ipynb | virtualcharacters/DisVoice | e53e4949a6023694720ba51708ec1646dcc4a667 | [
"MIT"
] | 191 | 2017-10-17T14:21:03.000Z | 2022-03-31T01:50:48.000Z | notebooks_examples/emotion classification using SVMs in scikit-learn.ipynb | virtualcharacters/DisVoice | e53e4949a6023694720ba51708ec1646dcc4a667 | [
"MIT"
] | 21 | 2017-08-14T16:06:01.000Z | 2022-01-11T17:03:11.000Z | notebooks_examples/emotion classification using SVMs in scikit-learn.ipynb | virtualcharacters/DisVoice | e53e4949a6023694720ba51708ec1646dcc4a667 | [
"MIT"
] | 56 | 2017-08-09T14:59:30.000Z | 2022-03-14T03:56:36.000Z | 452.784753 | 28,848 | 0.9365 | [
[
[
"# Emotion recognition using Emo-DB dataset and scikit-learn\n\n### Database: Emo-DB database (free) 7 emotions\nThe data can be downloaded from http://emodb.bilderbar.info/index-1024.html\n\nCode of emotions\n\nW->Anger->Wut\n\nL->Boredom->Langeweile\n\nE->Disgust->Ekel\n\nA->Anxiety/Fear->Angst\n\nF->Happiness->Freude\n\nT->Sadness->Trauer\n\nN->Neutral\n\n\n",
"_____no_output_____"
]
],
[
[
"import requests \nimport zipfile\nimport os\nimport numpy as np\nimport matplotlib.pyplot as plt\nimport scipy.stats as st\nimport itertools\nimport sys\nsys.path.append(\"../\")\nfrom plots_examples import plot_confusion_matrix, plot_ROC, plot_histogram\n\n# disvoice imports\nfrom phonation.phonation import Phonation\nfrom articulation.articulation import Articulation\nfrom prosody.prosody import Prosody\nfrom phonological.phonological import Phonological\nfrom replearning.replearning import RepLearning\n\n# sklearn methods\nfrom sklearn.model_selection import RandomizedSearchCV, train_test_split\nfrom sklearn import preprocessing\nfrom sklearn import metrics\nfrom sklearn import svm",
"Using TensorFlow backend.\n/home/camilo/anaconda3/lib/python3.6/site-packages/tensorflow/python/framework/dtypes.py:526: FutureWarning: Passing (type, 1) or '1type' as a synonym of type is deprecated; in a future version of numpy, it will be understood as (type, (1,)) / '(1,)type'.\n _np_qint8 = np.dtype([(\"qint8\", np.int8, 1)])\n/home/camilo/anaconda3/lib/python3.6/site-packages/tensorflow/python/framework/dtypes.py:527: FutureWarning: Passing (type, 1) or '1type' as a synonym of type is deprecated; in a future version of numpy, it will be understood as (type, (1,)) / '(1,)type'.\n _np_quint8 = np.dtype([(\"quint8\", np.uint8, 1)])\n/home/camilo/anaconda3/lib/python3.6/site-packages/tensorflow/python/framework/dtypes.py:528: FutureWarning: Passing (type, 1) or '1type' as a synonym of type is deprecated; in a future version of numpy, it will be understood as (type, (1,)) / '(1,)type'.\n _np_qint16 = np.dtype([(\"qint16\", np.int16, 1)])\n/home/camilo/anaconda3/lib/python3.6/site-packages/tensorflow/python/framework/dtypes.py:529: FutureWarning: Passing (type, 1) or '1type' as a synonym of type is deprecated; in a future version of numpy, it will be understood as (type, (1,)) / '(1,)type'.\n _np_quint16 = np.dtype([(\"quint16\", np.uint16, 1)])\n/home/camilo/anaconda3/lib/python3.6/site-packages/tensorflow/python/framework/dtypes.py:530: FutureWarning: Passing (type, 1) or '1type' as a synonym of type is deprecated; in a future version of numpy, it will be understood as (type, (1,)) / '(1,)type'.\n _np_qint32 = np.dtype([(\"qint32\", np.int32, 1)])\n/home/camilo/anaconda3/lib/python3.6/site-packages/tensorflow/python/framework/dtypes.py:535: FutureWarning: Passing (type, 1) or '1type' as a synonym of type is deprecated; in a future version of numpy, it will be understood as (type, (1,)) / '(1,)type'.\n np_resource = np.dtype([(\"resource\", np.ubyte, 1)])\n"
]
],
[
[
"## Download and unzip data\n",
"_____no_output_____"
]
],
[
[
"def download_url(url, save_path, chunk_size=128):\n r = requests.get(url, stream=True)\n with open(save_path, 'wb') as fd:\n for chunk in r.iter_content(chunk_size=chunk_size):\n fd.write(chunk)\n \nPATH_data=\"http://emodb.bilderbar.info/download/download.zip\"",
"_____no_output_____"
],
[
"download_url(PATH_data, \"./download.zip\")",
"_____no_output_____"
],
[
"with zipfile.ZipFile(\"./download.zip\", 'r') as zip_ref:\n zip_ref.extractall(\"./emo-db/\")",
"_____no_output_____"
]
],
[
[
"## prepare labels from the dataset\n\nwe will get labels for two classification problems: \n\n1. high vs. low arousal emotions\n2. positive vs. negative emotions\n",
"_____no_output_____"
]
],
[
[
"PATH_AUDIO=os.path.abspath(\"./emo-db/wav\")+\"/\"\nlabelsd='WLEAFTN'\nlabelshl= [0, 1, 0, 0, 0, 1, 1] # 0 high arousal emotion, 1 low arousal emotions\nlabelspn= [0, 0, 0, 0, 1, 0, 1] # 0 negative valence emotion, 1 positive valence emotion\n\nhf=os.listdir(PATH_AUDIO)\nhf.sort()\n\nyArousal=np.zeros(len(hf))\nyValence=np.zeros(len(hf))\nfor j in range(len(hf)):\n name_file=hf[j]\n label=hf[j][5]\n poslabel=labelsd.find(label)\n yArousal[j]=labelshl[poslabel]\n yValence[j]=labelspn[poslabel]",
"_____no_output_____"
]
],
[
[
"## compute features using disvoice: phonation, articulation, prosody, phonological",
"_____no_output_____"
]
],
[
[
"phonationf=Phonation()\narticulationf=Articulation()\nprosodyf=Prosody()\nphonologicalf=Phonological()\nreplearningf=RepLearning('CAE')\n",
"WARNING:tensorflow:From /home/camilo/anaconda3/lib/python3.6/site-packages/tensorflow/python/ops/resource_variable_ops.py:435: colocate_with (from tensorflow.python.framework.ops) is deprecated and will be removed in a future version.\nInstructions for updating:\nColocations handled automatically by placer.\n"
]
],
[
[
"### phonation features",
"_____no_output_____"
]
],
[
[
"Xphonation=phonationf.extract_features_path(PATH_AUDIO, static=True, plots=False, fmt=\"npy\")\nprint(Xphonation.shape)",
"Processing 14a07Aa.wav: 69%|██████▉ | 368/535 [00:10<00:04, 40.87it/s]../phonation/phonation.py:161: WavFileWarning: Chunk (non-data) not understood, skipping it.\n fs, data_audio=read(audio)\nProcessing 16b10Wb.wav: 100%|██████████| 535/535 [00:14<00:00, 36.70it/s]"
]
],
[
[
"### articulation features",
"_____no_output_____"
]
],
[
[
"Xarticulation=articulationf.extract_features_path(PATH_AUDIO, static=True, plots=False, fmt=\"npy\")\nprint(Xarticulation.shape)",
"Processing 03a01Fa.wav: 0%| | 0/535 [00:00<?, ?it/s]/home/camilo/anaconda3/lib/python3.6/site-packages/numpy/core/_asarray.py:136: VisibleDeprecationWarning: Creating an ndarray from ragged nested sequences (which is a list-or-tuple of lists-or-tuples-or ndarrays with different lengths or shapes) is deprecated. If you meant to do this, you must specify 'dtype=object' when creating the ndarray\n return array(a, dtype, copy=False, order=order, subok=True)\nProcessing 14a07Aa.wav: 69%|██████▉ | 370/535 [02:49<01:36, 1.71it/s]../articulation/articulation.py:264: WavFileWarning: Chunk (non-data) not understood, skipping it.\n fs, data_audio=read(audio)\nProcessing 16b10Wb.wav: 100%|██████████| 535/535 [04:01<00:00, 2.21it/s]"
]
],
[
[
"### prosody features",
"_____no_output_____"
]
],
[
[
"Xprosody=prosodyf.extract_features_path(PATH_AUDIO, static=True, plots=False, fmt=\"npy\")\nprint(Xprosody.shape)",
"Processing 14a07Aa.wav: 69%|██████▉ | 370/535 [01:12<00:33, 4.96it/s]../prosody/prosody.py:300: WavFileWarning: Chunk (non-data) not understood, skipping it.\n fs, data_audio=read(audio)\nProcessing 14a07Fd.wav: 70%|██████▉ | 372/535 [01:12<00:29, 5.57it/s]../prosody/prosody.py:300: WavFileWarning: Chunk (non-data) not understood, skipping it.\n fs, data_audio=read(audio)\nProcessing 14a07Na.wav: 70%|███████ | 375/535 [01:12<00:28, 5.52it/s]../prosody/prosody.py:300: WavFileWarning: Chunk (non-data) not understood, skipping it.\n fs, data_audio=read(audio)\nProcessing 14a07Wc.wav: 70%|███████ | 377/535 [01:13<00:36, 4.32it/s]../prosody/prosody.py:300: WavFileWarning: Chunk (non-data) not understood, skipping it.\n fs, data_audio=read(audio)\nProcessing 16b10Wb.wav: 100%|██████████| 535/535 [01:46<00:00, 5.04it/s]"
]
],
[
[
"### phonological features",
"_____no_output_____"
]
],
[
[
"Xphonological=phonologicalf.extract_features_path(PATH_AUDIO, static=True, plots=False, fmt=\"npy\")\nprint(Xphonological.shape)",
"Processing 14a07Aa.wav: 69%|██████▉ | 370/535 [02:24<00:57, 2.89it/s]/home/camilo/anaconda3/lib/python3.6/site-packages/phonet/phonet.py:235: WavFileWarning: Chunk (non-data) not understood, skipping it.\n fs, signal=read(audio_file)\nProcessing 16b10Wb.wav: 100%|██████████| 535/535 [03:28<00:00, 2.57it/s]"
]
],
[
[
"### representation learning features",
"_____no_output_____"
]
],
[
[
"Xrep=replearningf.extract_features_path(PATH_AUDIO, static=True, plots=False, fmt=\"npy\")\nprint(Xrep.shape)",
"Processing 03a01Fa.wav: 0%| | 0/535 [00:00<?, ?it/s]/home/camilo/anaconda3/lib/python3.6/site-packages/torch/nn/functional.py:1639: UserWarning: nn.functional.sigmoid is deprecated. Use torch.sigmoid instead.\n warnings.warn(\"nn.functional.sigmoid is deprecated. Use torch.sigmoid instead.\")\nProcessing 08b01Wa.wav: 16%|█▌ | 83/535 [00:45<04:39, 1.62it/s]../replearning/AEspeech.py:95: UserWarning: There is Inf values in the Mel spectrogram\n warnings.warn(\"There is Inf values in the Mel spectrogram\")\nProcessing 14a07Aa.wav: 69%|██████▉ | 370/535 [03:12<01:47, 1.53it/s]../replearning/AEspeech.py:77: WavFileWarning: Chunk (non-data) not understood, skipping it.\n fs, signal=read(wav_file)\nProcessing 16b10Wb.wav: 100%|██████████| 535/535 [04:46<00:00, 1.87it/s]"
]
],
[
[
"### Emotion classification using an SVM classifier\n\n",
"_____no_output_____"
]
],
[
[
"def classify(X, y):\n \n # train test split\n Xtrain, Xtest, ytrain, ytest = train_test_split(X, y, test_size=0.30, random_state=42)\n \n # z-score standarization\n scaler = preprocessing.StandardScaler().fit(Xtrain) \n Xtrain=scaler.transform(Xtrain) \n Xtest=scaler.transform(Xtest)\n Results=[]\n \n # randomized search cross-validation to optimize hyper-parameters of SVM \n parameters = {'kernel':['rbf'], 'class_weight': ['balanced'], \n 'C':st.expon(scale=10),\n 'gamma':st.expon(scale=0.01)}\n\n svc = svm.SVC()\n\n clf=RandomizedSearchCV(svc, parameters, n_jobs=4, cv=10, verbose=1, n_iter=200, scoring='balanced_accuracy')\n\n clf.fit(Xtrain, ytrain) # train the SVM\n accDev= clf.best_score_ # validation accuracy\n Copt=clf.best_params_.get('C') # best C\n gammaopt=clf.best_params_.get('gamma') # best gamma\n \n # train the SVM with the optimal hyper-parameters\n cls=svm.SVC(kernel='rbf', C=Copt, gamma=gammaopt, class_weight='balanced') \n cls.fit(Xtrain, ytrain)\n ypred=cls.predict(Xtest) # test predictions\n \n # check the results\n acc=metrics.accuracy_score(ytest, ypred)\n score_test=cls.decision_function(Xtest)\n dfclass=metrics.classification_report(ytest, ypred,digits=4)\n\n # display the results\n \n plot_confusion_matrix(ytest, ypred, classes=[\"class 0\", \"class 1\"], normalize=True)\n plot_ROC(ytest, score_test)\n plot_histogram(ytest, score_test, name_clases=[\"class 0\", \"class 1\"])\n \n print(\"Accuracy: \", acc)\n print(dfclass)\n\n \n",
"_____no_output_____"
]
],
[
[
"## classify high vs. low arousal with the different feature sets",
"_____no_output_____"
]
],
[
[
"classify(Xphonation, yArousal)",
"Fitting 10 folds for each of 500 candidates, totalling 5000 fits\n"
],
[
"classify(Xarticulation, yArousal)",
"Fitting 10 folds for each of 500 candidates, totalling 5000 fits\n"
],
[
"classify(Xprosody, yArousal)",
"Fitting 10 folds for each of 500 candidates, totalling 5000 fits\n"
],
[
"classify(Xphonological, yArousal)",
"Fitting 10 folds for each of 500 candidates, totalling 5000 fits\n"
],
[
"classify(Xrep, yArousal)",
"Fitting 10 folds for each of 500 candidates, totalling 5000 fits\n"
]
],
[
[
"## classify positive vs. negative valence with the different feature sets",
"_____no_output_____"
]
],
[
[
"classify(Xphonation, yValence)",
"Fitting 10 folds for each of 500 candidates, totalling 5000 fits\n"
],
[
"classify(Xarticulation, yValence)",
"Fitting 10 folds for each of 500 candidates, totalling 5000 fits\n"
],
[
"classify(Xprosody, yValence)",
"Fitting 10 folds for each of 500 candidates, totalling 5000 fits\n"
],
[
"classify(Xphonological, yValence)",
"Fitting 10 folds for each of 500 candidates, totalling 5000 fits\n"
],
[
"classify(Xrep, yValence)",
"Fitting 10 folds for each of 500 candidates, totalling 5000 fits\n"
]
]
] | [
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] | [
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code"
]
] |
d0ffeacb6d20f35aca081449adbadf69beee6857 | 8,468 | ipynb | Jupyter Notebook | Keras_mnist_digit_dataset.ipynb | Nikhitha-S-Pavan/Deep-learning-examples-using-keras | f8c8e60894bd9b5a2bf9a0cdfbb46b96208d11a1 | [
"Apache-2.0"
] | null | null | null | Keras_mnist_digit_dataset.ipynb | Nikhitha-S-Pavan/Deep-learning-examples-using-keras | f8c8e60894bd9b5a2bf9a0cdfbb46b96208d11a1 | [
"Apache-2.0"
] | null | null | null | Keras_mnist_digit_dataset.ipynb | Nikhitha-S-Pavan/Deep-learning-examples-using-keras | f8c8e60894bd9b5a2bf9a0cdfbb46b96208d11a1 | [
"Apache-2.0"
] | null | null | null | 31.834586 | 273 | 0.500827 | [
[
[
"<a href=\"https://colab.research.google.com/github/Nikhitha-S-Pavan/Deep-learning-examples-using-keras/blob/main/Keras_mnist_digit_dataset.ipynb\" target=\"_parent\"><img src=\"https://colab.research.google.com/assets/colab-badge.svg\" alt=\"Open In Colab\"/></a>",
"_____no_output_____"
]
],
[
[
"!pip install keras-tuner",
"_____no_output_____"
],
[
"import tensorflow as tf\r\nfrom tensorflow import keras\r\nimport numpy as np\r\nfrom matplotlib import pyplot",
"_____no_output_____"
],
[
"mnist=keras.datasets.mnist",
"_____no_output_____"
],
[
"(train_x, train_y), (test_x, test_y) = mnist.load_data()",
"_____no_output_____"
],
[
"for i in range(9):\r\n\t# define subplot\r\n\tpyplot.subplot(330 + 1 + i)\r\n\t# plot raw pixel data\r\n\tpyplot.imshow(train_x[i], cmap=pyplot.get_cmap('gray'))\r\n# show the figure\r\npyplot.show()",
"_____no_output_____"
],
[
"train_images=train_x.reshape(len(train_x),28,28,1)\r\ntest_images=test_x.reshape(len(test_x),28,28,1)",
"_____no_output_____"
],
[
"train_x.shape",
"_____no_output_____"
],
[
"train_images=train_images/255",
"_____no_output_____"
],
[
"test_images = test_images/255",
"_____no_output_____"
],
[
"from kerastuner import RandomSearch\r\nfrom kerastuner.engine.hyperparameters import HyperParameters\r\ndef build_model(hp): \r\n model = keras.Sequential([\r\n keras.layers.Conv2D(\r\n filters=hp.Int('conv_1_filter', min_value=32, max_value=128, step=16),\r\n kernel_size=hp.Choice('conv_1_kernel', values = [3,5]),\r\n activation='relu',\r\n input_shape=(28,28,1)\r\n ),\r\n keras.layers.Conv2D(\r\n filters=hp.Int('conv_2_filter', min_value=32, max_value=64, step=16),\r\n kernel_size=hp.Choice('conv_2_kernel', values = [3,5]),\r\n activation='relu'\r\n ),\r\n keras.layers.Flatten(),\r\n keras.layers.Dense(\r\n units=hp.Int('dense_1_units', min_value=32, max_value=128, step=16),\r\n activation='relu'\r\n ),\r\n keras.layers.Dense(10, activation='softmax')\r\n ])\r\n \r\n model.compile(optimizer=keras.optimizers.Adam(hp.Choice('learning_rate', values=[1e-2, 1e-3])),\r\n loss='sparse_categorical_crossentropy',\r\n metrics=['accuracy'])\r\n model.summary()\r\n return model",
"_____no_output_____"
],
[
"tuner_search=RandomSearch(build_model,\r\n objective='val_accuracy',\r\n max_trials=2,directory='output',project_name=\"Mnist\")\r\ntuner_search.search(train_images,train_y,epochs=3,validation_split=0.1)\r\nmodel=tuner_search.get_best_models(num_models=1)[0]\r\nmodel.fit(train_images, train_y, epochs=10, validation_split=0.1, initial_epoch=3)",
"_____no_output_____"
],
[
"\"\"\"\r\nimport keras\r\nfrom keras.datasets import mnist\r\nfrom keras.models import Sequential\r\nfrom keras.layers import Dense, Dropout, Flatten\r\nfrom keras.layers import Conv2D, MaxPooling2D\r\nfrom keras import backend as K\r\nbatch_size = 128\r\nnum_classes = 10\r\nepochs = 12\r\n# input image dimensions\r\nimg_rows, img_cols = 28, 28\r\n# the data, split between train and test sets\r\n(x_train, y_train), (x_test, y_test) = mnist.load_data()\r\n\r\nx_train = x_train.reshape(x_train.shape[0], img_rows, img_cols, 1)\r\nx_test = x_test.reshape(x_test.shape[0], img_rows, img_cols, 1)\r\ninput_shape = (img_rows, img_cols, 1) \r\nx_train = x_train.astype('float32')\r\nx_test = x_test.astype('float32')\r\nx_train /= 255\r\nx_test /= 255\r\nprint('x_train shape:', x_train.shape)\r\nprint(x_train.shape[0], 'train samples')\r\nprint(x_test.shape[0], 'test samples')\r\n# convert class vectors to binary class matrices\r\ny_train = keras.utils.to_categorical(y_train, num_classes)\r\ny_test = keras.utils.to_categorical(y_test, num_classes)\r\nmodel = Sequential()\r\nmodel.add(Conv2D(32, kernel_size=(3, 3),\r\n activation='relu',\r\n input_shape=input_shape))\r\nmodel.add(Conv2D(64, (3, 3), activation='relu'))\r\nmodel.add(MaxPooling2D(pool_size=(2, 2)))\r\nmodel.add(Dropout(0.25))\r\nmodel.add(Flatten())\r\nmodel.add(Dense(128, activation='relu'))\r\nmodel.add(Dropout(0.5))\r\nmodel.add(Dense(num_classes, activation='softmax'))\r\nmodel.compile(loss=keras.losses.categorical_crossentropy,\r\n optimizer=keras.optimizers.Adadelta(),\r\n metrics=['accuracy'])\r\nmodel.fit(x_train, y_train,\r\n batch_size=batch_size,\r\n epochs=5,\r\n verbose=1,\r\n validation_data=(x_test, y_test))\r\nscore = model.evaluate(x_test, y_test, verbose=0)\r\nprint('Test loss:', score[0])\r\nprint('Test accuracy:', score[1])\r\nmodel.save_weights(\"model.h5\")\"\"\"",
"_____no_output_____"
],
[
"model.summary()",
"_____no_output_____"
]
]
] | [
"markdown",
"code"
] | [
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
d0ffec8a98bd7bfad64ac2ecacc84d83ba2eec00 | 165,878 | ipynb | Jupyter Notebook | notebooks/07_EnsembleMethods.ipynb | Tionick/AppliedDeepLearningClass | 3be1bdf666d7c2704284898e9c43ffca1078b61a | [
"MIT"
] | 33 | 2018-06-06T19:38:06.000Z | 2021-10-19T13:59:45.000Z | notebooks/07_EnsembleMethods.ipynb | Tionick/AppliedDeepLearningClass | 3be1bdf666d7c2704284898e9c43ffca1078b61a | [
"MIT"
] | null | null | null | notebooks/07_EnsembleMethods.ipynb | Tionick/AppliedDeepLearningClass | 3be1bdf666d7c2704284898e9c43ffca1078b61a | [
"MIT"
] | 30 | 2018-06-06T22:59:15.000Z | 2022-01-02T01:18:37.000Z | 66.086853 | 38,080 | 0.73885 | [
[
[
"# 07 - Ensemble Methods\n\nby [Alejandro Correa Bahnsen](http://www.albahnsen.com/) & [Iván Torroledo](http://www.ivantorroledo.com/)\n\nversion 1.3, June 2018\n\n## Part of the class [Applied Deep Learning](https://github.com/albahnsen/AppliedDeepLearningClass)\n\n\nThis notebook is licensed under a [Creative Commons Attribution-ShareAlike 3.0 Unported License](http://creativecommons.org/licenses/by-sa/3.0/deed.en_US). Special thanks goes to [Kevin Markham](https://github.com/justmarkham)",
"_____no_output_____"
],
[
"Why are we learning about ensembling?\n\n- Very popular method for improving the predictive performance of machine learning models\n- Provides a foundation for understanding more sophisticated models",
"_____no_output_____"
],
[
"## Lesson objectives\n\nStudents will be able to:\n\n- Define ensembling and its requirements\n- Identify the two basic methods of ensembling\n- Decide whether manual ensembling is a useful approach for a given problem\n- Explain bagging and how it can be applied to decision trees\n- Explain how out-of-bag error and feature importances are calculated from bagged trees\n- Explain the difference between bagged trees and Random Forests\n- Build and tune a Random Forest model in scikit-learn\n- Decide whether a decision tree or a Random Forest is a better model for a given problem",
"_____no_output_____"
],
[
"# Part 1: Introduction",
"_____no_output_____"
],
[
"Ensemble learning is a widely studied topic in the machine learning community. The main idea behind \nthe ensemble methodology is to combine several individual base classifiers in order to have a \nclassifier that outperforms each of them.\n\nNowadays, ensemble methods are one \nof the most popular and well studied machine learning techniques, and it can be \nnoted that since 2009 all the first-place and second-place winners of the KDD-Cup https://www.sigkdd.org/kddcup/ used ensemble methods. The core \nprinciple in ensemble learning, is to induce random perturbations into the learning procedure in \norder to produce several different base classifiers from a single training set, then combining the \nbase classifiers in order to make the final prediction. In order to induce the random permutations \nand therefore create the different base classifiers, several methods have been proposed, in \nparticular: \n* bagging\n* pasting\n* random forests \n* random patches \n\nFinally, after the base classifiers \nare trained, they are typically combined using either:\n* majority voting\n* weighted voting \n* stacking\n",
"_____no_output_____"
],
[
"There are three main reasons regarding why ensemble \nmethods perform better than single models: statistical, computational and representational . First, from a statistical point of view, when the learning set is too \nsmall, an algorithm can find several good models within the search space, that arise to the same \nperformance on the training set $\\mathcal{S}$. Nevertheless, without a validation set, there is \na risk of choosing the wrong model. The second reason is computational; in general, algorithms \nrely on some local search optimization and may get stuck in a local optima. Then, an ensemble may \nsolve this by focusing different algorithms to different spaces across the training set. The last \nreason is representational. In most cases, for a learning set of finite size, the true function \n$f$ cannot be represented by any of the candidate models. By combining several models in an \nensemble, it may be possible to obtain a model with a larger coverage across the space of \nrepresentable functions.",
"_____no_output_____"
],
[
"",
"_____no_output_____"
],
[
"## Example\n\nLet's pretend that instead of building a single model to solve a binary classification problem, you created **five independent models**, and each model was correct about 70% of the time. If you combined these models into an \"ensemble\" and used their majority vote as a prediction, how often would the ensemble be correct?",
"_____no_output_____"
]
],
[
[
"import numpy as np\n\n# set a seed for reproducibility\nnp.random.seed(1234)\n\n# generate 1000 random numbers (between 0 and 1) for each model, representing 1000 observations\nmod1 = np.random.rand(1000)\nmod2 = np.random.rand(1000)\nmod3 = np.random.rand(1000)\nmod4 = np.random.rand(1000)\nmod5 = np.random.rand(1000)\n\n# each model independently predicts 1 (the \"correct response\") if random number was at least 0.3\npreds1 = np.where(mod1 > 0.3, 1, 0)\npreds2 = np.where(mod2 > 0.3, 1, 0)\npreds3 = np.where(mod3 > 0.3, 1, 0)\npreds4 = np.where(mod4 > 0.3, 1, 0)\npreds5 = np.where(mod5 > 0.3, 1, 0)\n\n# print the first 20 predictions from each model\nprint(preds1[:20])\nprint(preds2[:20])\nprint(preds3[:20])\nprint(preds4[:20])\nprint(preds5[:20])",
"[0 1 1 1 1 0 0 1 1 1 1 1 1 1 1 1 1 0 1 1]\n[1 1 1 1 1 1 1 0 1 0 0 0 1 1 1 0 1 0 0 0]\n[1 1 1 1 0 1 1 0 0 1 1 1 1 1 1 1 1 0 1 1]\n[1 1 0 0 0 0 1 1 0 1 1 1 1 1 1 0 1 1 1 0]\n[0 0 1 0 0 0 1 0 1 0 0 0 1 1 1 1 1 1 1 1]\n"
],
[
"# average the predictions and then round to 0 or 1\nensemble_preds = np.round((preds1 + preds2 + preds3 + preds4 + preds5)/5.0).astype(int)\n\n# print the ensemble's first 20 predictions\nprint(ensemble_preds[:20])",
"[1 1 1 1 0 0 1 0 1 1 1 1 1 1 1 1 1 0 1 1]\n"
],
[
"# how accurate was each individual model?\nprint(preds1.mean())\nprint(preds2.mean())\nprint(preds3.mean())\nprint(preds4.mean())\nprint(preds5.mean())",
"0.713\n0.665\n0.717\n0.712\n0.687\n"
],
[
"# how accurate was the ensemble?\nprint(ensemble_preds.mean())",
"0.841\n"
]
],
[
[
"**Note:** As you add more models to the voting process, the probability of error decreases, which is known as [Condorcet's Jury Theorem](http://en.wikipedia.org/wiki/Condorcet%27s_jury_theorem).",
"_____no_output_____"
],
[
"## What is ensembling?\n\n**Ensemble learning (or \"ensembling\")** is the process of combining several predictive models in order to produce a combined model that is more accurate than any individual model.\n\n- **Regression:** take the average of the predictions\n- **Classification:** take a vote and use the most common prediction, or take the average of the predicted probabilities\n\nFor ensembling to work well, the models must have the following characteristics:\n\n- **Accurate:** they outperform the null model\n- **Independent:** their predictions are generated using different processes\n\n**The big idea:** If you have a collection of individually imperfect (and independent) models, the \"one-off\" mistakes made by each model are probably not going to be made by the rest of the models, and thus the mistakes will be discarded when averaging the models.\n\nThere are two basic **methods for ensembling:**\n\n- Manually ensemble your individual models\n- Use a model that ensembles for you",
"_____no_output_____"
],
[
"### Theoretical performance of an ensemble\n If we assume that each one of the $T$ base classifiers has a probability $\\rho$ of \n being correct, the probability of an ensemble making the correct decision, assuming independence, \n denoted by $P_c$, can be calculated using the binomial distribution\n\n$$P_c = \\sum_{j>T/2}^{T} {{T}\\choose{j}} \\rho^j(1-\\rho)^{T-j}.$$\n\n Furthermore, as shown, if $T\\ge3$ then:\n\n$$\n \\lim_{T \\to \\infty} P_c= \\begin{cases} \n 1 &\\mbox{if } \\rho>0.5 \\\\ \n 0 &\\mbox{if } \\rho<0.5 \\\\ \n 0.5 &\\mbox{if } \\rho=0.5 ,\n \\end{cases}\n$$\n\tleading to the conclusion that \n$$\n \\rho \\ge 0.5 \\quad \\text{and} \\quad T\\ge3 \\quad \\Rightarrow \\quad P_c\\ge \\rho.\n$$",
"_____no_output_____"
],
[
"# Part 2: Manual ensembling\n\nWhat makes a good manual ensemble?\n\n- Different types of **models**\n- Different combinations of **features**\n- Different **tuning parameters**",
"_____no_output_____"
],
[
"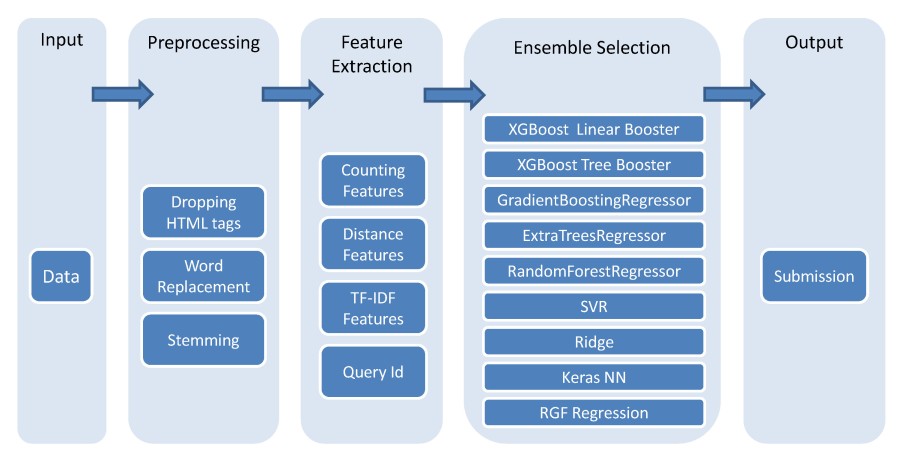\n\n*Machine learning flowchart created by the [winner](https://github.com/ChenglongChen/Kaggle_CrowdFlower) of Kaggle's [CrowdFlower competition](https://www.kaggle.com/c/crowdflower-search-relevance)*",
"_____no_output_____"
]
],
[
[
"# read in and prepare the vehicle training data\nimport zipfile\nimport pandas as pd\nwith zipfile.ZipFile('../datasets/vehicles_train.csv.zip', 'r') as z:\n f = z.open('vehicles_train.csv')\n train = pd.io.parsers.read_table(f, index_col=False, sep=',')\nwith zipfile.ZipFile('../datasets/vehicles_test.csv.zip', 'r') as z:\n f = z.open('vehicles_test.csv')\n test = pd.io.parsers.read_table(f, index_col=False, sep=',')\n\ntrain['vtype'] = train.vtype.map({'car':0, 'truck':1})\n# read in and prepare the vehicle testing data\ntest['vtype'] = test.vtype.map({'car':0, 'truck':1})",
"_____no_output_____"
],
[
"train.head()",
"_____no_output_____"
]
],
[
[
"### Train different models",
"_____no_output_____"
]
],
[
[
"from sklearn.linear_model import LinearRegression\nfrom sklearn.tree import DecisionTreeRegressor\nfrom sklearn.naive_bayes import GaussianNB\nfrom sklearn.neighbors import KNeighborsRegressor\n\nmodels = {'lr': LinearRegression(),\n 'dt': DecisionTreeRegressor(),\n 'nb': GaussianNB(),\n 'nn': KNeighborsRegressor()}",
"_____no_output_____"
],
[
"# Train all the models\nX_train = train.iloc[:, 1:]\nX_test = test.iloc[:, 1:]\ny_train = train.price\ny_test = test.price\n\nfor model in models.keys():\n models[model].fit(X_train, y_train)",
"_____no_output_____"
],
[
"# predict test for each model\ny_pred = pd.DataFrame(index=test.index, columns=models.keys())\nfor model in models.keys():\n y_pred[model] = models[model].predict(X_test)\n ",
"_____no_output_____"
],
[
"# Evaluate each model\nfrom sklearn.metrics import mean_squared_error\n\nfor model in models.keys():\n print(model,np.sqrt(mean_squared_error(y_pred[model], y_test)))",
"lr 2138.3579028745116\ndt 1414.213562373095\nnb 5477.2255750516615\nnn 1671.3268182295567\n"
]
],
[
[
"### Evaluate the error of the mean of the predictions",
"_____no_output_____"
]
],
[
[
"np.sqrt(mean_squared_error(y_pred.mean(axis=1), y_test))",
"_____no_output_____"
]
],
[
[
"## Comparing manual ensembling with a single model approach\n\n**Advantages of manual ensembling:**\n\n- Increases predictive accuracy\n- Easy to get started\n\n**Disadvantages of manual ensembling:**\n\n- Decreases interpretability\n- Takes longer to train\n- Takes longer to predict\n- More complex to automate and maintain\n- Small gains in accuracy may not be worth the added complexity",
"_____no_output_____"
],
[
"# Part 3: Bagging\n\nThe primary weakness of **decision trees** is that they don't tend to have the best predictive accuracy. This is partially due to **high variance**, meaning that different splits in the training data can lead to very different trees.\n\n**Bagging** is a general purpose procedure for reducing the variance of a machine learning method, but is particularly useful for decision trees. Bagging is short for **bootstrap aggregation**, meaning the aggregation of bootstrap samples.\n\nWhat is a **bootstrap sample**? A random sample with replacement:",
"_____no_output_____"
]
],
[
[
"# set a seed for reproducibility\nnp.random.seed(1)\n\n# create an array of 1 through 20\nnums = np.arange(1, 21)\nprint(nums)\n\n# sample that array 20 times with replacement\nprint(np.random.choice(a=nums, size=20, replace=True))",
"[ 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20]\n[ 6 12 13 9 10 12 6 16 1 17 2 13 8 14 7 19 6 19 12 11]\n"
]
],
[
[
"**How does bagging work (for decision trees)?**\n\n1. Grow B trees using B bootstrap samples from the training data.\n2. Train each tree on its bootstrap sample and make predictions.\n3. Combine the predictions:\n - Average the predictions for **regression trees**\n - Take a vote for **classification trees**\n\nNotes:\n\n- **Each bootstrap sample** should be the same size as the original training set.\n- **B** should be a large enough value that the error seems to have \"stabilized\".\n- The trees are **grown deep** so that they have low bias/high variance.\n\nBagging increases predictive accuracy by **reducing the variance**, similar to how cross-validation reduces the variance associated with train/test split (for estimating out-of-sample error) by splitting many times an averaging the results.\n",
"_____no_output_____"
]
],
[
[
"# set a seed for reproducibility\nnp.random.seed(123)\n\nn_samples = train.shape[0]\nn_B = 10\n\n# create ten bootstrap samples (will be used to select rows from the DataFrame)\nsamples = [np.random.choice(a=n_samples, size=n_samples, replace=True) for _ in range(1, n_B +1 )]\nsamples",
"_____no_output_____"
],
[
"# show the rows for the first decision tree\ntrain.iloc[samples[0], :]",
"_____no_output_____"
]
],
[
[
" Build one tree for each sample",
"_____no_output_____"
]
],
[
[
"from sklearn.tree import DecisionTreeRegressor\n\n# grow each tree deep\ntreereg = DecisionTreeRegressor(max_depth=None, random_state=123)\n\n# DataFrame for storing predicted price from each tree\ny_pred = pd.DataFrame(index=test.index, columns=[list(range(n_B))])\n\n# grow one tree for each bootstrap sample and make predictions on testing data\nfor i, sample in enumerate(samples):\n X_train = train.iloc[sample, 1:]\n y_train = train.iloc[sample, 0]\n treereg.fit(X_train, y_train)\n y_pred[i] = treereg.predict(X_test)",
"_____no_output_____"
],
[
"y_pred",
"_____no_output_____"
]
],
[
[
"Results of each tree",
"_____no_output_____"
]
],
[
[
"for i in range(n_B):\n print(i, np.sqrt(mean_squared_error(y_pred[i], y_test)))",
"0 1621.7274740226856\n1 2942.7877939124323\n2 1825.7418583505537\n3 1000.0\n4 1276.7145334803704\n5 1414.213562373095\n6 1414.213562373095\n7 1000.0\n8 1554.5631755148024\n9 1914.854215512676\n"
]
],
[
[
"Results of the ensemble",
"_____no_output_____"
]
],
[
[
"y_pred.mean(axis=1)",
"_____no_output_____"
],
[
"np.sqrt(mean_squared_error(y_test, y_pred.mean(axis=1)))",
"_____no_output_____"
]
],
[
[
"## Bagged decision trees in scikit-learn (with B=500)",
"_____no_output_____"
]
],
[
[
"# define the training and testing sets\nX_train = train.iloc[:, 1:]\ny_train = train.iloc[:, 0]\nX_test = test.iloc[:, 1:]\ny_test = test.iloc[:, 0]",
"_____no_output_____"
],
[
"# instruct BaggingRegressor to use DecisionTreeRegressor as the \"base estimator\"\nfrom sklearn.ensemble import BaggingRegressor\nbagreg = BaggingRegressor(DecisionTreeRegressor(), n_estimators=500, \n bootstrap=True, oob_score=True, random_state=1)",
"_____no_output_____"
],
[
"# fit and predict\nbagreg.fit(X_train, y_train)\ny_pred = bagreg.predict(X_test)\ny_pred",
"_____no_output_____"
],
[
"# calculate RMSE\nnp.sqrt(mean_squared_error(y_test, y_pred))",
"_____no_output_____"
]
],
[
[
"## Estimating out-of-sample error\n\nFor bagged models, out-of-sample error can be estimated without using **train/test split** or **cross-validation**!\n\nOn average, each bagged tree uses about **two-thirds** of the observations. For each tree, the **remaining observations** are called \"out-of-bag\" observations.",
"_____no_output_____"
]
],
[
[
"# show the first bootstrap sample\nsamples[0]",
"_____no_output_____"
],
[
"# show the \"in-bag\" observations for each sample\nfor sample in samples:\n print(set(sample))",
"{0, 1, 2, 3, 6, 9, 10, 11, 12, 13}\n{0, 1, 2, 3, 4, 7, 9, 13}\n{0, 2, 3, 4, 6, 7, 8, 9, 12, 13}\n{0, 1, 2, 3, 5, 6, 8, 10, 11, 12}\n{2, 3, 4, 6, 10, 11, 12, 13}\n{0, 1, 4, 5, 6, 7, 9, 10, 11}\n{0, 1, 2, 3, 4, 5, 8, 9, 12}\n{1, 2, 3, 5, 6, 7, 9, 11}\n{1, 3, 6, 7, 8, 9, 11, 12}\n{0, 1, 3, 4, 5, 6, 8, 10, 11, 13}\n"
],
[
"# show the \"out-of-bag\" observations for each sample\nfor sample in samples:\n print(sorted(set(range(n_samples)) - set(sample)))",
"[4, 5, 7, 8]\n[5, 6, 8, 10, 11, 12]\n[1, 5, 10, 11]\n[4, 7, 9, 13]\n[0, 1, 5, 7, 8, 9]\n[2, 3, 8, 12, 13]\n[6, 7, 10, 11, 13]\n[0, 4, 8, 10, 12, 13]\n[0, 2, 4, 5, 10, 13]\n[2, 7, 9, 12]\n"
]
],
[
[
"How to calculate **\"out-of-bag error\":**\n\n1. For every observation in the training data, predict its response value using **only** the trees in which that observation was out-of-bag. Average those predictions (for regression) or take a vote (for classification).\n2. Compare all predictions to the actual response values in order to compute the out-of-bag error.\n\nWhen B is sufficiently large, the **out-of-bag error** is an accurate estimate of **out-of-sample error**.",
"_____no_output_____"
]
],
[
[
"# compute the out-of-bag R-squared score (not MSE, unfortunately!) for B=500\nbagreg.oob_score_",
"_____no_output_____"
]
],
[
[
"## Estimating feature importance\n\nBagging increases **predictive accuracy**, but decreases **model interpretability** because it's no longer possible to visualize the tree to understand the importance of each feature.\n\nHowever, we can still obtain an overall summary of **feature importance** from bagged models:\n\n- **Bagged regression trees:** calculate the total amount that **MSE** is decreased due to splits over a given feature, averaged over all trees\n- **Bagged classification trees:** calculate the total amount that **Gini index** is decreased due to splits over a given feature, averaged over all trees",
"_____no_output_____"
],
[
"# Part 4: Random Forests\n\nRandom Forests is a **slight variation of bagged trees** that has even better performance:\n\n- Exactly like bagging, we create an ensemble of decision trees using bootstrapped samples of the training set.\n- However, when building each tree, each time a split is considered, a **random sample of m features** is chosen as split candidates from the **full set of p features**. The split is only allowed to use **one of those m features**.\n - A new random sample of features is chosen for **every single tree at every single split**.\n - For **classification**, m is typically chosen to be the square root of p.\n - For **regression**, m is typically chosen to be somewhere between p/3 and p.\n\nWhat's the point?\n\n- Suppose there is **one very strong feature** in the data set. When using bagged trees, most of the trees will use that feature as the top split, resulting in an ensemble of similar trees that are **highly correlated**.\n- Averaging highly correlated quantities does not significantly reduce variance (which is the entire goal of bagging).\n- By randomly leaving out candidate features from each split, **Random Forests \"decorrelates\" the trees**, such that the averaging process can reduce the variance of the resulting model.",
"_____no_output_____"
],
[
"# Part 5: Building and tuning decision trees and Random Forests\n\n- Major League Baseball player data from 1986-87: [data](https://github.com/justmarkham/DAT8/blob/master/data/hitters.csv), [data dictionary](https://cran.r-project.org/web/packages/ISLR/ISLR.pdf) (page 7)\n- Each observation represents a player\n- **Goal:** Predict player salary",
"_____no_output_____"
]
],
[
[
"# read in the data\nwith zipfile.ZipFile('../datasets/hitters.csv.zip', 'r') as z:\n f = z.open('hitters.csv')\n hitters = pd.read_csv(f, sep=',', index_col=False)\n\n# remove rows with missing values\nhitters.dropna(inplace=True)\nhitters.head()",
"_____no_output_____"
],
[
"# encode categorical variables as integers\nhitters['League'] = pd.factorize(hitters.League)[0]\nhitters['Division'] = pd.factorize(hitters.Division)[0]\nhitters['NewLeague'] = pd.factorize(hitters.NewLeague)[0]\nhitters.head()",
"_____no_output_____"
],
[
"# allow plots to appear in the notebook\n%matplotlib inline\nimport matplotlib.pyplot as plt\nplt.style.use('fivethirtyeight')",
"_____no_output_____"
],
[
"# scatter plot of Years versus Hits colored by Salary\nhitters.plot(kind='scatter', x='Years', y='Hits', c='Salary', colormap='jet', xlim=(0, 25), ylim=(0, 250))",
"_____no_output_____"
],
[
"# define features: exclude career statistics (which start with \"C\") and the response (Salary)\nfeature_cols = hitters.columns[hitters.columns.str.startswith('C') == False].drop('Salary')\nfeature_cols",
"_____no_output_____"
],
[
"# define X and y\nX = hitters[feature_cols]\ny = hitters.Salary",
"_____no_output_____"
]
],
[
[
"## Predicting salary with a decision tree\n\nFind the best **max_depth** for a decision tree using cross-validation:",
"_____no_output_____"
]
],
[
[
"# list of values to try for max_depth\nmax_depth_range = range(1, 21)\n\n# list to store the average RMSE for each value of max_depth\nRMSE_scores = []\n\n# use 10-fold cross-validation with each value of max_depth\nfrom sklearn.model_selection import cross_val_score\nfor depth in max_depth_range:\n treereg = DecisionTreeRegressor(max_depth=depth, random_state=1)\n MSE_scores = cross_val_score(treereg, X, y, cv=10, scoring='neg_mean_squared_error')\n RMSE_scores.append(np.mean(np.sqrt(-MSE_scores)))",
"_____no_output_____"
],
[
"# plot max_depth (x-axis) versus RMSE (y-axis)\nplt.plot(max_depth_range, RMSE_scores)\nplt.xlabel('max_depth')\nplt.ylabel('RMSE (lower is better)')",
"_____no_output_____"
],
[
"# show the best RMSE and the corresponding max_depth\nsorted(zip(RMSE_scores, max_depth_range))[0]",
"_____no_output_____"
],
[
"# max_depth=2 was best, so fit a tree using that parameter\ntreereg = DecisionTreeRegressor(max_depth=2, random_state=1)\ntreereg.fit(X, y)",
"_____no_output_____"
],
[
"# compute feature importances\npd.DataFrame({'feature':feature_cols, 'importance':treereg.feature_importances_}).sort_values('importance')",
"_____no_output_____"
]
],
[
[
"## Predicting salary with a Random Forest",
"_____no_output_____"
]
],
[
[
"from sklearn.ensemble import RandomForestRegressor\nrfreg = RandomForestRegressor()\nrfreg",
"_____no_output_____"
]
],
[
[
"### Tuning n_estimators\n\nOne important tuning parameter is **n_estimators**, which is the number of trees that should be grown. It should be a large enough value that the error seems to have \"stabilized\".",
"_____no_output_____"
]
],
[
[
"# list of values to try for n_estimators\nestimator_range = range(10, 310, 10)\n\n# list to store the average RMSE for each value of n_estimators\nRMSE_scores = []\n\n# use 5-fold cross-validation with each value of n_estimators (WARNING: SLOW!)\nfor estimator in estimator_range:\n rfreg = RandomForestRegressor(n_estimators=estimator, random_state=1, n_jobs=-1)\n MSE_scores = cross_val_score(rfreg, X, y, cv=5, scoring='neg_mean_squared_error')\n RMSE_scores.append(np.mean(np.sqrt(-MSE_scores)))",
"_____no_output_____"
],
[
"# plot n_estimators (x-axis) versus RMSE (y-axis)\nplt.plot(estimator_range, RMSE_scores)\nplt.xlabel('n_estimators')\nplt.ylabel('RMSE (lower is better)')",
"_____no_output_____"
]
],
[
[
"### Tuning max_features\n\nThe other important tuning parameter is **max_features**, which is the number of features that should be considered at each split.",
"_____no_output_____"
]
],
[
[
"# list of values to try for max_features\nfeature_range = range(1, len(feature_cols)+1)\n\n# list to store the average RMSE for each value of max_features\nRMSE_scores = []\n\n# use 10-fold cross-validation with each value of max_features (WARNING: SLOW!)\nfor feature in feature_range:\n rfreg = RandomForestRegressor(n_estimators=150, max_features=feature, random_state=1, n_jobs=-1)\n MSE_scores = cross_val_score(rfreg, X, y, cv=10, scoring='neg_mean_squared_error')\n RMSE_scores.append(np.mean(np.sqrt(-MSE_scores)))",
"_____no_output_____"
],
[
"# plot max_features (x-axis) versus RMSE (y-axis)\nplt.plot(feature_range, RMSE_scores)\nplt.xlabel('max_features')\nplt.ylabel('RMSE (lower is better)')",
"_____no_output_____"
],
[
"# show the best RMSE and the corresponding max_features\nsorted(zip(RMSE_scores, feature_range))[0]",
"_____no_output_____"
]
],
[
[
"### Fitting a Random Forest with the best parameters",
"_____no_output_____"
]
],
[
[
"# max_features=8 is best and n_estimators=150 is sufficiently large\nrfreg = RandomForestRegressor(n_estimators=150, max_features=8, max_depth=3, oob_score=True, random_state=1)\nrfreg.fit(X, y)",
"_____no_output_____"
],
[
"# compute feature importances\npd.DataFrame({'feature':feature_cols, 'importance':rfreg.feature_importances_}).sort_values('importance')",
"_____no_output_____"
],
[
"# compute the out-of-bag R-squared score\nrfreg.oob_score_",
"_____no_output_____"
]
],
[
[
"### Reducing X to its most important features\n",
"_____no_output_____"
]
],
[
[
"# check the shape of X\nX.shape",
"_____no_output_____"
],
[
"rfreg",
"_____no_output_____"
],
[
"# set a threshold for which features to include\nfrom sklearn.feature_selection import SelectFromModel\nprint(SelectFromModel(rfreg, threshold=0.1, prefit=True).transform(X).shape)\nprint(SelectFromModel(rfreg, threshold='mean', prefit=True).transform(X).shape)\nprint(SelectFromModel(rfreg, threshold='median', prefit=True).transform(X).shape)",
"(263, 4)\n(263, 5)\n(263, 7)\n"
],
[
"# create a new feature matrix that only includes important features\nX_important = SelectFromModel(rfreg, threshold='mean', prefit=True).transform(X)",
"_____no_output_____"
],
[
"# check the RMSE for a Random Forest that only includes important features\nrfreg = RandomForestRegressor(n_estimators=150, max_features=3, random_state=1)\nscores = cross_val_score(rfreg, X_important, y, cv=10, scoring='neg_mean_squared_error')\nnp.mean(np.sqrt(-scores))",
"_____no_output_____"
]
],
[
[
"## Comparing Random Forests with decision trees\n\n**Advantages of Random Forests:**\n\n- Performance is competitive with the best supervised learning methods\n- Provides a more reliable estimate of feature importance\n- Allows you to estimate out-of-sample error without using train/test split or cross-validation\n\n**Disadvantages of Random Forests:**\n\n- Less interpretable\n- Slower to train\n- Slower to predict",
"_____no_output_____"
]
]
] | [
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown"
] | [
[
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown"
],
[
"code",
"code",
"code",
"code"
],
[
"markdown",
"markdown",
"markdown",
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown",
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
]
] |
d0fffd1277602e164332a8b3cca9aaaacac99ed1 | 4,556 | ipynb | Jupyter Notebook | docs/examples/SolarAbund.ipynb | jchowk/linetools | 5a0eafa96ab854c52c070ce756033c0499414dde | [
"BSD-3-Clause"
] | 23 | 2015-07-09T02:24:27.000Z | 2022-01-22T15:13:31.000Z | docs/examples/SolarAbund.ipynb | jchowk/linetools | 5a0eafa96ab854c52c070ce756033c0499414dde | [
"BSD-3-Clause"
] | 491 | 2015-06-21T20:01:33.000Z | 2021-02-11T03:29:19.000Z | docs/examples/SolarAbund.ipynb | jchowk/linetools | 5a0eafa96ab854c52c070ce756033c0499414dde | [
"BSD-3-Clause"
] | 35 | 2015-05-25T00:18:59.000Z | 2021-08-30T06:53:14.000Z | 16.156028 | 76 | 0.459175 | [
[
[
"# Examples with the SolarAbund Class (v1.1)",
"_____no_output_____"
]
],
[
[
"# import\nfrom linetools.abund import solar as labsol",
"_____no_output_____"
]
],
[
[
"## Init",
"_____no_output_____"
]
],
[
[
"sol = labsol.SolarAbund()",
"Loading abundances from Asplund2009\nAbundances are relative by number on a logarithmic scale with H=12\n"
],
[
"sol",
"_____no_output_____"
]
],
[
[
"## Usage",
"_____no_output_____"
]
],
[
[
"# Simple calls\nprint(sol['C'])\nprint(sol[6])",
"8.43\n8.43\n"
],
[
"# Ratio\nprint(sol.get_ratio('C/Fe'))",
"0.98\n"
],
[
"# Multiple calls\nprint(sol[6,12,'S'])",
"[ 8.43 7.53 7.15]\n"
]
],
[
[
"## Bits and pieces",
"_____no_output_____"
]
],
[
[
"from linetools.abund import ions as laions",
"_____no_output_____"
],
[
"# Ion name\nlaions.ion_name((6,2))",
"_____no_output_____"
],
[
"# Name to ion\nlaions.name_ion('CII')",
"_____no_output_____"
],
[
"from linetools.abund.elements import ELEMENTS\n",
"_____no_output_____"
],
[
"ele = ELEMENTS['C']",
"_____no_output_____"
],
[
"ele.eleconfig_dict",
"_____no_output_____"
]
]
] | [
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] | [
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
d0fffd9f2b87ae1899c948a7908d61abeabd9528 | 135,060 | ipynb | Jupyter Notebook | .ipynb_checkpoints/tutorial_deep-checkpoint.ipynb | xriflo/thesis_deep_learning | 1d5bef42874d2b03762c004a63d0010b93eff8d0 | [
"MIT"
] | null | null | null | .ipynb_checkpoints/tutorial_deep-checkpoint.ipynb | xriflo/thesis_deep_learning | 1d5bef42874d2b03762c004a63d0010b93eff8d0 | [
"MIT"
] | null | null | null | .ipynb_checkpoints/tutorial_deep-checkpoint.ipynb | xriflo/thesis_deep_learning | 1d5bef42874d2b03762c004a63d0010b93eff8d0 | [
"MIT"
] | null | null | null | 24.804408 | 266 | 0.542892 | [
[
[
"empty"
]
]
] | [
"empty"
] | [
[
"empty"
]
] |
cb0000d6f27d6b03bf54ba5e871f88d57e17ffc1 | 298,294 | ipynb | Jupyter Notebook | covid-19-prediction-using-cnn.ipynb | AdityaIyer2609/Covid-19-Prediction-Using-CNN | 9a5e50395a58f6b101abdd2eb315bcea592c0b3f | [
"CC0-1.0"
] | 2 | 2021-08-29T14:25:34.000Z | 2021-12-05T11:26:36.000Z | covid-19-prediction-using-cnn.ipynb | AdityaIyer2609/Covid-19-Prediction-Using-CNN | 9a5e50395a58f6b101abdd2eb315bcea592c0b3f | [
"CC0-1.0"
] | null | null | null | covid-19-prediction-using-cnn.ipynb | AdityaIyer2609/Covid-19-Prediction-Using-CNN | 9a5e50395a58f6b101abdd2eb315bcea592c0b3f | [
"CC0-1.0"
] | null | null | null | 298,294 | 298,294 | 0.92967 | [
[
[
"\nimport os\nfor dirname, _, filenames in os.walk('../input/covid19-image-dataset'):\n for filename in filenames:\n print(os.path.join(dirname, filename))\n\n",
"../input/covid19-image-dataset/Covid19-dataset/test/Normal/0117.jpeg\n../input/covid19-image-dataset/Covid19-dataset/test/Normal/0101.jpeg\n../input/covid19-image-dataset/Covid19-dataset/test/Normal/0108.jpeg\n../input/covid19-image-dataset/Covid19-dataset/test/Normal/0110.jpeg\n../input/covid19-image-dataset/Covid19-dataset/test/Normal/0119.jpeg\n../input/covid19-image-dataset/Covid19-dataset/test/Normal/0115.jpeg\n../input/covid19-image-dataset/Covid19-dataset/test/Normal/0116.jpeg\n../input/covid19-image-dataset/Covid19-dataset/test/Normal/0105.jpeg\n../input/covid19-image-dataset/Covid19-dataset/test/Normal/0106.jpeg\n../input/covid19-image-dataset/Covid19-dataset/test/Normal/0121.jpeg\n../input/covid19-image-dataset/Covid19-dataset/test/Normal/0111.jpeg\n../input/covid19-image-dataset/Covid19-dataset/test/Normal/0103.jpeg\n../input/covid19-image-dataset/Covid19-dataset/test/Normal/0120.jpeg\n../input/covid19-image-dataset/Covid19-dataset/test/Normal/0122.jpeg\n../input/covid19-image-dataset/Covid19-dataset/test/Normal/0109.jpeg\n../input/covid19-image-dataset/Covid19-dataset/test/Normal/0112.jpeg\n../input/covid19-image-dataset/Covid19-dataset/test/Normal/0114.jpeg\n../input/covid19-image-dataset/Covid19-dataset/test/Normal/0107.jpeg\n../input/covid19-image-dataset/Covid19-dataset/test/Normal/0118.jpeg\n../input/covid19-image-dataset/Covid19-dataset/test/Normal/0102.jpeg\n../input/covid19-image-dataset/Covid19-dataset/test/Viral Pneumonia/0117.jpeg\n../input/covid19-image-dataset/Covid19-dataset/test/Viral Pneumonia/0101.jpeg\n../input/covid19-image-dataset/Covid19-dataset/test/Viral Pneumonia/0104.jpeg\n../input/covid19-image-dataset/Covid19-dataset/test/Viral Pneumonia/0108.jpeg\n../input/covid19-image-dataset/Covid19-dataset/test/Viral Pneumonia/0110.jpeg\n../input/covid19-image-dataset/Covid19-dataset/test/Viral Pneumonia/0119.jpeg\n../input/covid19-image-dataset/Covid19-dataset/test/Viral Pneumonia/0115.jpeg\n../input/covid19-image-dataset/Covid19-dataset/test/Viral Pneumonia/0116.jpeg\n../input/covid19-image-dataset/Covid19-dataset/test/Viral Pneumonia/0105.jpeg\n../input/covid19-image-dataset/Covid19-dataset/test/Viral Pneumonia/0106.jpeg\n../input/covid19-image-dataset/Covid19-dataset/test/Viral Pneumonia/0111.jpeg\n../input/covid19-image-dataset/Covid19-dataset/test/Viral Pneumonia/0103.jpeg\n../input/covid19-image-dataset/Covid19-dataset/test/Viral Pneumonia/0120.jpeg\n../input/covid19-image-dataset/Covid19-dataset/test/Viral Pneumonia/0109.jpeg\n../input/covid19-image-dataset/Covid19-dataset/test/Viral Pneumonia/0112.jpeg\n../input/covid19-image-dataset/Covid19-dataset/test/Viral Pneumonia/0113.jpeg\n../input/covid19-image-dataset/Covid19-dataset/test/Viral Pneumonia/0114.jpeg\n../input/covid19-image-dataset/Covid19-dataset/test/Viral Pneumonia/0107.jpeg\n../input/covid19-image-dataset/Covid19-dataset/test/Viral Pneumonia/0118.jpeg\n../input/covid19-image-dataset/Covid19-dataset/test/Viral Pneumonia/0102.jpeg\n../input/covid19-image-dataset/Covid19-dataset/test/Covid/0108.jpeg\n../input/covid19-image-dataset/Covid19-dataset/test/Covid/0119.jpeg\n../input/covid19-image-dataset/Covid19-dataset/test/Covid/0115.jpeg\n../input/covid19-image-dataset/Covid19-dataset/test/Covid/auntminnie-2020_01_31_20_24_2322_2020_01_31_x-ray_coronavirus_US.jpg\n../input/covid19-image-dataset/Covid19-dataset/test/Covid/COVID-00037.jpg\n../input/covid19-image-dataset/Covid19-dataset/test/Covid/0106.jpeg\n../input/covid19-image-dataset/Covid19-dataset/test/Covid/0111.jpg\n../input/covid19-image-dataset/Covid19-dataset/test/Covid/COVID-00022.jpg\n../input/covid19-image-dataset/Covid19-dataset/test/Covid/auntminnie-a-2020_01_28_23_51_6665_2020_01_28_Vietnam_coronavirus.jpeg\n../input/covid19-image-dataset/Covid19-dataset/test/Covid/0120.jpg\n../input/covid19-image-dataset/Covid19-dataset/test/Covid/094.png\n../input/covid19-image-dataset/Covid19-dataset/test/Covid/COVID-00003b.jpg\n../input/covid19-image-dataset/Covid19-dataset/test/Covid/0105.png\n../input/covid19-image-dataset/Covid19-dataset/test/Covid/radiopaedia-2019-novel-coronavirus-infected-pneumonia.jpg\n../input/covid19-image-dataset/Covid19-dataset/test/Covid/auntminnie-c-2020_01_28_23_51_6665_2020_01_28_Vietnam_coronavirus.jpeg\n../input/covid19-image-dataset/Covid19-dataset/test/Covid/0112.jpg\n../input/covid19-image-dataset/Covid19-dataset/test/Covid/COVID-00012.jpg\n../input/covid19-image-dataset/Covid19-dataset/test/Covid/COVID-00033.jpg\n../input/covid19-image-dataset/Covid19-dataset/test/Covid/0113.jpg\n../input/covid19-image-dataset/Covid19-dataset/test/Covid/0100.jpeg\n../input/covid19-image-dataset/Covid19-dataset/test/Covid/auntminnie-b-2020_01_28_23_51_6665_2020_01_28_Vietnam_coronavirus.jpeg\n../input/covid19-image-dataset/Covid19-dataset/test/Covid/098.jpeg\n../input/covid19-image-dataset/Covid19-dataset/test/Covid/096.png\n../input/covid19-image-dataset/Covid19-dataset/test/Covid/auntminnie-d-2020_01_28_23_51_6665_2020_01_28_Vietnam_coronavirus.jpeg\n../input/covid19-image-dataset/Covid19-dataset/test/Covid/0118.jpeg\n../input/covid19-image-dataset/Covid19-dataset/test/Covid/0102.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Normal/071.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Normal/014.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Normal/073.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Normal/05.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Normal/023.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Normal/01.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Normal/096.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Normal/060.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Normal/063.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Normal/059.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Normal/087.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Normal/064.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Normal/011.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Normal/068.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Normal/058.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Normal/020.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Normal/050.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Normal/03.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Normal/02.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Normal/093.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Normal/016.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Normal/010.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Normal/018.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Normal/067.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Normal/088.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Normal/054.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Normal/052.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Normal/084.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Normal/04.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Normal/066.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Normal/079.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Normal/077.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Normal/082.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Normal/056.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Normal/094.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Normal/080.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Normal/057.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Normal/019.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Normal/021.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Normal/074.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Normal/061.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Normal/08.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Normal/069.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Normal/062.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Normal/06.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Normal/013.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Normal/051.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Normal/097.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Normal/024.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Normal/015.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Normal/012.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Normal/081.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Normal/095.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Normal/072.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Normal/092.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Normal/055.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Normal/086.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Normal/053.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Normal/025.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Normal/022.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Normal/017.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Normal/065.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Normal/091.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Normal/076.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Normal/07.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Normal/083.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Normal/075.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Normal/070.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Normal/09.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Normal/085.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Viral Pneumonia/071.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Viral Pneumonia/035.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Viral Pneumonia/032.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Viral Pneumonia/073.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Viral Pneumonia/037.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Viral Pneumonia/05.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Viral Pneumonia/023.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Viral Pneumonia/01.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Viral Pneumonia/096.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Viral Pneumonia/063.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Viral Pneumonia/044.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Viral Pneumonia/064.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Viral Pneumonia/036.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Viral Pneumonia/011.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Viral Pneumonia/068.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Viral Pneumonia/058.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Viral Pneumonia/020.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Viral Pneumonia/03.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Viral Pneumonia/02.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Viral Pneumonia/016.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Viral Pneumonia/010.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Viral Pneumonia/018.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Viral Pneumonia/067.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Viral Pneumonia/054.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Viral Pneumonia/052.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Viral Pneumonia/084.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Viral Pneumonia/04.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Viral Pneumonia/066.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Viral Pneumonia/042.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Viral Pneumonia/077.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Viral Pneumonia/041.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Viral Pneumonia/082.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Viral Pneumonia/056.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Viral Pneumonia/094.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Viral Pneumonia/038.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Viral Pneumonia/057.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Viral Pneumonia/019.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Viral Pneumonia/021.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Viral Pneumonia/027.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Viral Pneumonia/046.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Viral Pneumonia/074.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Viral Pneumonia/061.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Viral Pneumonia/08.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Viral Pneumonia/062.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Viral Pneumonia/045.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Viral Pneumonia/033.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Viral Pneumonia/06.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Viral Pneumonia/013.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Viral Pneumonia/051.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Viral Pneumonia/078.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Viral Pneumonia/024.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Viral Pneumonia/034.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Viral Pneumonia/012.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Viral Pneumonia/081.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Viral Pneumonia/095.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Viral Pneumonia/072.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Viral Pneumonia/055.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Viral Pneumonia/043.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Viral Pneumonia/053.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Viral Pneumonia/025.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Viral Pneumonia/022.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Viral Pneumonia/065.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Viral Pneumonia/076.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Viral Pneumonia/047.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Viral Pneumonia/07.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Viral Pneumonia/083.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Viral Pneumonia/048.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Viral Pneumonia/075.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Viral Pneumonia/031.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Viral Pneumonia/09.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/032.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/01.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/COVID-00014.jpg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/080.jpg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/060.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/092.png\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/COVID-00005.jpg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/COVID-00031.jpg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/059.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/067.jpg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/044.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/COVID-00037.jpg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/069.jpg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/089.jpg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/076.jpg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/COVID-00032.jpg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/COVID-00027.jpg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/058.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/050.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/03.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/02.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/COVID-00003a.jpg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/056.jpg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/033.png\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/COVID-00016.jpg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/COVID-00017.jpg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/COVID-00022.jpg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/09.png\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/088.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/068.jpg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/COVID-00034.jpg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/064.jpg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/COVID-00029.jpg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/015.jpg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/COVID-00030.jpg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/054.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/086.jpg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/052.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/COVID-00007.jpg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/COVID-00025.jpg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/084.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/079.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/042.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/COVID-00038.jpg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/COVID-00003b.jpg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/041.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/COVID-00026.jpg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/057.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/021.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/COVID-00001.jpg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/027.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/046.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/COVID-00028.jpg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/039.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/071.jpg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/08.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/COVID-00009.jpg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/062.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/COVID-00024.jpg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/045.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/COVID-00018.jpg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/019.png\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/COVID-00015a.png\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/COVID-00020.jpg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/06.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/061.jpg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/051.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/073.jpg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/078.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/024.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/020.jpg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/040.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/COVID-00021.jpg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/012.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/COVID-00019.jpg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/COVID-00012.jpg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/081.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/COVID-00033.jpg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/072.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/COVID-00011.jpg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/074.jpg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/055.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/026.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/043.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/049.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/COVID-00023.jpg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/COVID-00036.jpg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/COVID-00013a.jpg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/053.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/025.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/COVID-00035.jpg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/082.jpg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/022.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/COVID-00006.jpg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/07.jpg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/065.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/047.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/COVID-00013b.jpg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/COVID-00010.jpg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/COVID-00015b.png\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/COVID-00004.jpg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/010.png\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/091.jpg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/090.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/083.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/048.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/COVID-00008.jpg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/031.jpeg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/04.png\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/COVID-00002.jpg\n../input/covid19-image-dataset/Covid19-dataset/train/Covid/085.jpeg\n"
],
[
"import tensorflow as tf\nimport numpy as np\nimport os\nfrom matplotlib import pyplot as plt\nimport cv2\nfrom tensorflow import keras\nfrom keras.models import Sequential\nfrom keras.layers import Conv2D #images are two dimensional. Videos are three dimension.\nfrom keras.layers import MaxPooling2D\nfrom keras.layers import Flatten\nfrom keras.layers import Dense\n\nfrom keras.preprocessing import image\nfrom tensorflow.keras.preprocessing.image import ImageDataGenerator",
"_____no_output_____"
]
],
[
[
"# Plotting Various Scan Reports",
"_____no_output_____"
],
[
"**Covid Patient**",
"_____no_output_____"
]
],
[
[
"plt.imshow(cv2.imread(\"../input/covid19-image-dataset/Covid19-dataset/train/Covid/022.jpeg\"))",
"_____no_output_____"
]
],
[
[
"**Pneumomnia Patient**",
"_____no_output_____"
]
],
[
[
"plt.imshow(cv2.imread(\"../input/covid19-image-dataset/Covid19-dataset/train/Viral Pneumonia/020.jpeg\"))",
"_____no_output_____"
]
],
[
[
"**Normal Patient**",
"_____no_output_____"
]
],
[
[
"plt.imshow(cv2.imread(\"../input/covid19-image-dataset/Covid19-dataset/train/Normal/018.jpeg\"))",
"_____no_output_____"
]
],
[
[
"# Preprocessing the images",
"_____no_output_____"
]
],
[
[
"train_datagen=ImageDataGenerator(rescale=1/255,\n shear_range=0.2,\n zoom_range=2,\n horizontal_flip=True)\ntraining_set=train_datagen.flow_from_directory('../input/covid19-image-dataset/Covid19-dataset/train',\n target_size=(224,224),\n batch_size=32)",
"Found 251 images belonging to 3 classes.\n"
],
[
"training_set.class_indices",
"_____no_output_____"
],
[
"test_datagen=ImageDataGenerator(rescale=1/255,\n shear_range=0.2,\n zoom_range=2,\n horizontal_flip=True)\ntest_set=test_datagen.flow_from_directory('../input/covid19-image-dataset/Covid19-dataset/test',\n target_size=(224,224),\n batch_size=32)",
"Found 66 images belonging to 3 classes.\n"
],
[
"test_set.class_indices",
"_____no_output_____"
]
],
[
[
"# Building VGG-16 architecture & Neural Network",
"_____no_output_____"
]
],
[
[
"model = tf.keras.models.Sequential([\n tf.keras.layers.Conv2D(32,(3,3),strides=(1, 1),activation='relu',padding='same', input_shape=(224, 224, 3)), \n tf.keras.layers.MaxPooling2D(pool_size=(2,2)),\n \n tf.keras.layers.Conv2D(64,(3,3),strides=(1, 1) ,padding='same',activation='relu'),\n tf.keras.layers.MaxPooling2D(pool_size=(2,2)),\n\n tf.keras.layers.Conv2D(128,(3,3),strides=(1, 1),padding='same', activation='relu'),\n tf.keras.layers.MaxPooling2D(pool_size=(2,2)),\n\n tf.keras.layers.Conv2D(256,(3,3),strides=(1, 1),padding='same', activation='relu'),\n tf.keras.layers.MaxPooling2D(pool_size=(2,2)),\n\n\n tf.keras.layers.Flatten(),\n\n tf.keras.layers.Dense(128, activation='relu'),\n tf.keras.layers.Dense(64, activation='relu'),\n tf.keras.layers.Dense(3, activation='softmax'),\n])",
"_____no_output_____"
],
[
"model.compile(optimizer='adam', loss='categorical_crossentropy', metrics=['accuracy'])\nmodel_fit = model.fit(training_set,\n epochs = 71,\n validation_data = test_set)",
"Epoch 1/71\n8/8 [==============================] - 16s 2s/step - loss: 1.1425 - accuracy: 0.3302 - val_loss: 1.0993 - val_accuracy: 0.4091\nEpoch 2/71\n8/8 [==============================] - 14s 2s/step - loss: 1.0259 - accuracy: 0.5257 - val_loss: 1.0013 - val_accuracy: 0.4394\nEpoch 3/71\n8/8 [==============================] - 14s 2s/step - loss: 1.0064 - accuracy: 0.4887 - val_loss: 1.0497 - val_accuracy: 0.4242\nEpoch 4/71\n8/8 [==============================] - 14s 2s/step - loss: 0.9968 - accuracy: 0.5543 - val_loss: 1.0409 - val_accuracy: 0.5152\nEpoch 5/71\n8/8 [==============================] - 15s 2s/step - loss: 0.9630 - accuracy: 0.5299 - val_loss: 1.0379 - val_accuracy: 0.5303\nEpoch 6/71\n8/8 [==============================] - 14s 2s/step - loss: 0.9084 - accuracy: 0.5945 - val_loss: 0.9447 - val_accuracy: 0.5152\nEpoch 7/71\n8/8 [==============================] - 14s 2s/step - loss: 0.8535 - accuracy: 0.5974 - val_loss: 0.7613 - val_accuracy: 0.6970\nEpoch 8/71\n8/8 [==============================] - 15s 2s/step - loss: 0.8367 - accuracy: 0.6256 - val_loss: 0.9538 - val_accuracy: 0.5909\nEpoch 9/71\n8/8 [==============================] - 14s 2s/step - loss: 0.9136 - accuracy: 0.5744 - val_loss: 0.9496 - val_accuracy: 0.5303\nEpoch 10/71\n8/8 [==============================] - 15s 2s/step - loss: 0.8363 - accuracy: 0.6322 - val_loss: 0.8672 - val_accuracy: 0.6364\nEpoch 11/71\n8/8 [==============================] - 14s 2s/step - loss: 0.7752 - accuracy: 0.6537 - val_loss: 0.8620 - val_accuracy: 0.6364\nEpoch 12/71\n8/8 [==============================] - 15s 2s/step - loss: 0.7635 - accuracy: 0.6559 - val_loss: 0.8075 - val_accuracy: 0.5606\nEpoch 13/71\n8/8 [==============================] - 14s 2s/step - loss: 0.7345 - accuracy: 0.6542 - val_loss: 0.7792 - val_accuracy: 0.6515\nEpoch 14/71\n8/8 [==============================] - 15s 2s/step - loss: 0.6317 - accuracy: 0.7171 - val_loss: 0.8167 - val_accuracy: 0.6212\nEpoch 15/71\n8/8 [==============================] - 14s 2s/step - loss: 0.7799 - accuracy: 0.6895 - val_loss: 0.9335 - val_accuracy: 0.6061\nEpoch 16/71\n8/8 [==============================] - 15s 2s/step - loss: 0.7858 - accuracy: 0.6256 - val_loss: 0.8358 - val_accuracy: 0.5606\nEpoch 17/71\n8/8 [==============================] - 15s 2s/step - loss: 0.6430 - accuracy: 0.7556 - val_loss: 0.6417 - val_accuracy: 0.7576\nEpoch 18/71\n8/8 [==============================] - 14s 2s/step - loss: 0.7289 - accuracy: 0.6405 - val_loss: 0.6528 - val_accuracy: 0.6970\nEpoch 19/71\n8/8 [==============================] - 15s 2s/step - loss: 0.6562 - accuracy: 0.7087 - val_loss: 0.9921 - val_accuracy: 0.4394\nEpoch 20/71\n8/8 [==============================] - 14s 2s/step - loss: 0.5586 - accuracy: 0.7620 - val_loss: 0.7364 - val_accuracy: 0.7121\nEpoch 21/71\n8/8 [==============================] - 15s 2s/step - loss: 0.5421 - accuracy: 0.7419 - val_loss: 1.1178 - val_accuracy: 0.5455\nEpoch 22/71\n8/8 [==============================] - 14s 2s/step - loss: 0.6322 - accuracy: 0.7552 - val_loss: 0.6772 - val_accuracy: 0.6364\nEpoch 23/71\n8/8 [==============================] - 15s 2s/step - loss: 0.7506 - accuracy: 0.6702 - val_loss: 0.9450 - val_accuracy: 0.5758\nEpoch 24/71\n8/8 [==============================] - 14s 2s/step - loss: 0.5421 - accuracy: 0.7705 - val_loss: 0.7125 - val_accuracy: 0.6818\nEpoch 25/71\n8/8 [==============================] - 15s 2s/step - loss: 0.6229 - accuracy: 0.7121 - val_loss: 0.5577 - val_accuracy: 0.7121\nEpoch 26/71\n8/8 [==============================] - 14s 2s/step - loss: 0.6555 - accuracy: 0.7290 - val_loss: 0.5884 - val_accuracy: 0.7576\nEpoch 27/71\n8/8 [==============================] - 14s 2s/step - loss: 0.5235 - accuracy: 0.7342 - val_loss: 0.5832 - val_accuracy: 0.6970\nEpoch 28/71\n8/8 [==============================] - 15s 2s/step - loss: 0.6093 - accuracy: 0.6981 - val_loss: 0.7279 - val_accuracy: 0.6970\nEpoch 29/71\n8/8 [==============================] - 14s 2s/step - loss: 0.5479 - accuracy: 0.7768 - val_loss: 0.7402 - val_accuracy: 0.6667\nEpoch 30/71\n8/8 [==============================] - 15s 2s/step - loss: 0.6048 - accuracy: 0.7199 - val_loss: 0.5727 - val_accuracy: 0.7424\nEpoch 31/71\n8/8 [==============================] - 14s 2s/step - loss: 0.5839 - accuracy: 0.7564 - val_loss: 0.6964 - val_accuracy: 0.7727\nEpoch 32/71\n8/8 [==============================] - 15s 2s/step - loss: 0.4373 - accuracy: 0.8647 - val_loss: 0.7259 - val_accuracy: 0.6970\nEpoch 33/71\n8/8 [==============================] - 14s 2s/step - loss: 0.5612 - accuracy: 0.7622 - val_loss: 0.6280 - val_accuracy: 0.7121\nEpoch 34/71\n8/8 [==============================] - 15s 2s/step - loss: 0.4673 - accuracy: 0.8049 - val_loss: 0.6012 - val_accuracy: 0.7121\nEpoch 35/71\n8/8 [==============================] - 14s 2s/step - loss: 0.5047 - accuracy: 0.7741 - val_loss: 0.4595 - val_accuracy: 0.8182\nEpoch 36/71\n8/8 [==============================] - 14s 2s/step - loss: 0.4661 - accuracy: 0.7814 - val_loss: 0.4531 - val_accuracy: 0.7879\nEpoch 37/71\n8/8 [==============================] - 15s 2s/step - loss: 0.4559 - accuracy: 0.8291 - val_loss: 0.5220 - val_accuracy: 0.8030\nEpoch 38/71\n8/8 [==============================] - 14s 2s/step - loss: 0.4248 - accuracy: 0.8028 - val_loss: 0.6106 - val_accuracy: 0.7576\nEpoch 39/71\n8/8 [==============================] - 15s 2s/step - loss: 0.4179 - accuracy: 0.8102 - val_loss: 0.7376 - val_accuracy: 0.6818\nEpoch 40/71\n8/8 [==============================] - 14s 2s/step - loss: 0.4329 - accuracy: 0.8229 - val_loss: 0.5541 - val_accuracy: 0.7273\nEpoch 41/71\n8/8 [==============================] - 15s 2s/step - loss: 0.4393 - accuracy: 0.7936 - val_loss: 0.6836 - val_accuracy: 0.6515\nEpoch 42/71\n8/8 [==============================] - 14s 2s/step - loss: 0.4967 - accuracy: 0.7738 - val_loss: 0.5089 - val_accuracy: 0.8030\nEpoch 43/71\n8/8 [==============================] - 15s 2s/step - loss: 0.4477 - accuracy: 0.8450 - val_loss: 0.6061 - val_accuracy: 0.8333\nEpoch 44/71\n8/8 [==============================] - 14s 2s/step - loss: 0.5114 - accuracy: 0.7783 - val_loss: 0.5147 - val_accuracy: 0.8636\nEpoch 45/71\n8/8 [==============================] - 14s 2s/step - loss: 0.5250 - accuracy: 0.7839 - val_loss: 0.5079 - val_accuracy: 0.8030\nEpoch 46/71\n8/8 [==============================] - 15s 2s/step - loss: 0.4791 - accuracy: 0.7805 - val_loss: 0.4951 - val_accuracy: 0.7576\nEpoch 47/71\n8/8 [==============================] - 14s 2s/step - loss: 0.4107 - accuracy: 0.8141 - val_loss: 0.5682 - val_accuracy: 0.8030\nEpoch 48/71\n8/8 [==============================] - 14s 2s/step - loss: 0.3878 - accuracy: 0.8057 - val_loss: 0.5381 - val_accuracy: 0.6970\nEpoch 49/71\n8/8 [==============================] - 14s 2s/step - loss: 0.3678 - accuracy: 0.8252 - val_loss: 0.4896 - val_accuracy: 0.7727\nEpoch 50/71\n8/8 [==============================] - 14s 2s/step - loss: 0.4083 - accuracy: 0.8162 - val_loss: 0.5547 - val_accuracy: 0.7273\nEpoch 51/71\n8/8 [==============================] - 14s 2s/step - loss: 0.4655 - accuracy: 0.7632 - val_loss: 0.5316 - val_accuracy: 0.7273\nEpoch 52/71\n8/8 [==============================] - 14s 2s/step - loss: 0.4136 - accuracy: 0.7903 - val_loss: 0.4828 - val_accuracy: 0.7727\nEpoch 53/71\n8/8 [==============================] - 14s 2s/step - loss: 0.4049 - accuracy: 0.8468 - val_loss: 0.4734 - val_accuracy: 0.7273\nEpoch 54/71\n8/8 [==============================] - 14s 2s/step - loss: 0.4032 - accuracy: 0.8278 - val_loss: 0.4956 - val_accuracy: 0.7727\nEpoch 55/71\n8/8 [==============================] - 15s 2s/step - loss: 0.4229 - accuracy: 0.7920 - val_loss: 0.6003 - val_accuracy: 0.6667\nEpoch 56/71\n8/8 [==============================] - 14s 2s/step - loss: 0.4196 - accuracy: 0.7936 - val_loss: 0.5194 - val_accuracy: 0.7727\nEpoch 57/71\n8/8 [==============================] - 15s 2s/step - loss: 0.4004 - accuracy: 0.8304 - val_loss: 0.5006 - val_accuracy: 0.7424\nEpoch 58/71\n8/8 [==============================] - 14s 2s/step - loss: 0.4789 - accuracy: 0.7841 - val_loss: 0.5009 - val_accuracy: 0.7576\nEpoch 59/71\n8/8 [==============================] - 15s 2s/step - loss: 0.3738 - accuracy: 0.8406 - val_loss: 0.6380 - val_accuracy: 0.7576\nEpoch 60/71\n8/8 [==============================] - 14s 2s/step - loss: 0.3811 - accuracy: 0.8306 - val_loss: 0.6308 - val_accuracy: 0.6818\nEpoch 61/71\n8/8 [==============================] - 14s 2s/step - loss: 0.3838 - accuracy: 0.8126 - val_loss: 0.5173 - val_accuracy: 0.7121\nEpoch 62/71\n8/8 [==============================] - 14s 2s/step - loss: 0.4291 - accuracy: 0.8262 - val_loss: 0.6995 - val_accuracy: 0.6970\nEpoch 63/71\n8/8 [==============================] - 14s 2s/step - loss: 0.4390 - accuracy: 0.8163 - val_loss: 0.6468 - val_accuracy: 0.7727\nEpoch 64/71\n8/8 [==============================] - 15s 2s/step - loss: 0.3977 - accuracy: 0.8211 - val_loss: 0.6470 - val_accuracy: 0.6667\nEpoch 65/71\n8/8 [==============================] - 14s 2s/step - loss: 0.4817 - accuracy: 0.8264 - val_loss: 0.5726 - val_accuracy: 0.7424\nEpoch 66/71\n8/8 [==============================] - 14s 2s/step - loss: 0.4996 - accuracy: 0.7769 - val_loss: 0.7133 - val_accuracy: 0.6515\nEpoch 67/71\n8/8 [==============================] - 14s 2s/step - loss: 0.4114 - accuracy: 0.8322 - val_loss: 0.5105 - val_accuracy: 0.7424\nEpoch 68/71\n8/8 [==============================] - 14s 2s/step - loss: 0.3862 - accuracy: 0.8038 - val_loss: 0.5303 - val_accuracy: 0.7424\nEpoch 69/71\n8/8 [==============================] - 14s 2s/step - loss: 0.3908 - accuracy: 0.8288 - val_loss: 0.4983 - val_accuracy: 0.8182\nEpoch 70/71\n8/8 [==============================] - 14s 2s/step - loss: 0.4291 - accuracy: 0.8018 - val_loss: 0.4024 - val_accuracy: 0.8030\nEpoch 71/71\n8/8 [==============================] - 14s 2s/step - loss: 0.3582 - accuracy: 0.8421 - val_loss: 0.3996 - val_accuracy: 0.8333\n"
]
],
[
[
"# Prediction",
"_____no_output_____"
]
],
[
[
"import numpy as np\nfrom keras.preprocessing import image\ntest_image = image.load_img('../input/covid19-image-dataset/Covid19-dataset/test/Viral Pneumonia/0112.jpeg', target_size = (224, 224))\ntest_image = image.img_to_array(test_image)\ntest_image = np.expand_dims(test_image, axis = 0)\nresult = model.predict(test_image)\ntraining_set.class_indices\nprint(result)",
"[[0. 0.00101711 0.99898285]]\n"
]
],
[
[
"[0,0,1] which means Pneumonia. Hence our model is accurate",
"_____no_output_____"
]
]
] | [
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown"
] | [
[
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
]
] |
cb000e81500076fa6eb0c221b21bfe67849c2015 | 190,104 | ipynb | Jupyter Notebook | courses/machine_learning/deepdive/10_recommend/labs/wals.ipynb | KayvanShah1/training-data-analyst | 3f778a57b8e6d2446af40ca6063b2fd9c1b4bc88 | [
"Apache-2.0"
] | 6,140 | 2016-05-23T16:09:35.000Z | 2022-03-30T19:00:46.000Z | courses/machine_learning/deepdive/10_recommend/labs/wals.ipynb | KayvanShah1/training-data-analyst | 3f778a57b8e6d2446af40ca6063b2fd9c1b4bc88 | [
"Apache-2.0"
] | 1,384 | 2016-07-08T22:26:41.000Z | 2022-03-24T16:39:43.000Z | courses/machine_learning/deepdive/10_recommend/labs/wals.ipynb | KayvanShah1/training-data-analyst | 3f778a57b8e6d2446af40ca6063b2fd9c1b4bc88 | [
"Apache-2.0"
] | 5,110 | 2016-05-27T13:45:18.000Z | 2022-03-31T18:40:42.000Z | 110.013889 | 105,688 | 0.830677 | [
[
[
"# Collaborative filtering on Google Analytics data\n\nThis notebook demonstrates how to implement a WALS matrix refactorization approach to do collaborative filtering.",
"_____no_output_____"
]
],
[
[
"import os\nPROJECT = \"cloud-training-demos\" # REPLACE WITH YOUR PROJECT ID\nBUCKET = \"cloud-training-demos-ml\" # REPLACE WITH YOUR BUCKET NAME\nREGION = \"us-central1\" # REPLACE WITH YOUR BUCKET REGION e.g. us-central1\n\n# Do not change these\nos.environ[\"PROJECT\"] = PROJECT\nos.environ[\"BUCKET\"] = BUCKET\nos.environ[\"REGION\"] = REGION\nos.environ[\"TFVERSION\"] = \"1.13\"",
"_____no_output_____"
],
[
"%%bash\ngcloud config set project $PROJECT\ngcloud config set compute/region $REGION",
"Updated property [core/project].\nUpdated property [compute/region].\n"
],
[
"import tensorflow as tf\nprint(tf.__version__)",
"1.13.1\n"
]
],
[
[
"## Create raw dataset\n<p>\nFor collaborative filtering, we don't need to know anything about either the users or the content. Essentially, all we need to know is userId, itemId, and rating that the particular user gave the particular item.\n<p>\nIn this case, we are working with newspaper articles. The company doesn't ask their users to rate the articles. However, we can use the time-spent on the page as a proxy for rating.\n<p>\nNormally, we would also add a time filter to this (\"latest 7 days\"), but our dataset is itself limited to a few days.",
"_____no_output_____"
]
],
[
[
"from google.cloud import bigquery\nbq = bigquery.Client(project = PROJECT)\n\nsql = \"\"\"\nWITH CTE_visitor_page_content AS (\n SELECT\n # Schema: https://support.google.com/analytics/answer/3437719?hl=en\n # For a completely unique visit-session ID, we combine combination of fullVisitorId and visitNumber:\n CONCAT(fullVisitorID,'-',CAST(visitNumber AS STRING)) AS visitorId,\n (SELECT MAX(IF(index=10, value, NULL)) FROM UNNEST(hits.customDimensions)) AS latestContentId, \n (LEAD(hits.time, 1) OVER (PARTITION BY fullVisitorId ORDER BY hits.time ASC) - hits.time) AS session_duration \n FROM\n `cloud-training-demos.GA360_test.ga_sessions_sample`, \n UNNEST(hits) AS hits\n WHERE \n # only include hits on pages\n hits.type = \"PAGE\"\nGROUP BY \n fullVisitorId,\n visitNumber,\n latestContentId,\n hits.time )\n-- Aggregate web stats\nSELECT \n visitorId,\n latestContentId as contentId,\n SUM(session_duration) AS session_duration\nFROM\n CTE_visitor_page_content\nWHERE\n latestContentId IS NOT NULL \nGROUP BY\n visitorId, \n latestContentId\nHAVING \n session_duration > 0\n\"\"\"\n\ndf = bq.query(sql).to_dataframe()\ndf.head()",
"_____no_output_____"
],
[
"stats = df.describe()\nstats",
"_____no_output_____"
],
[
"df[[\"session_duration\"]].plot(kind=\"hist\", logy=True, bins=100, figsize=[8,5])",
"_____no_output_____"
],
[
"# The rating is the session_duration scaled to be in the range 0-1. This will help with training.\nmedian = stats.loc[\"50%\", \"session_duration\"]\ndf[\"rating\"] = 0.3 * df[\"session_duration\"] / median\ndf.loc[df[\"rating\"] > 1, \"rating\"] = 1\ndf[[\"rating\"]].plot(kind=\"hist\", logy=True, bins=100, figsize=[8,5])",
"_____no_output_____"
],
[
"del df[\"session_duration\"]",
"_____no_output_____"
],
[
"%%bash\nrm -rf data\nmkdir data",
"_____no_output_____"
],
[
"df.to_csv(path_or_buf = \"data/collab_raw.csv\", index = False, header = False)",
"_____no_output_____"
],
[
"!head data/collab_raw.csv",
"7337153711992174438,100074831,0.2321051400452234\n5190801220865459604,100170790,1.0\n2293633612703952721,100510126,0.2481776360816793\n5874973374932455844,100510126,0.16690549004998828\n1173698801255170595,100676857,0.05464232805149575\n883397426232997550,10083328,0.9487035095774818\n1808867070685560283,100906145,1.0\n7615995624631762562,100906145,0.48418654214351925\n5519169380728479914,100915139,0.20026163722525925\n3427736932800080345,100950628,0.558924688331153\n"
]
],
[
[
"## Create dataset for WALS\n<p>\nThe raw dataset (above) won't work for WALS:\n<ol>\n<li> The userId and itemId have to be 0,1,2 ... so we need to create a mapping from visitorId (in the raw data) to userId and contentId (in the raw data) to itemId.\n<li> We will need to save the above mapping to a file because at prediction time, we'll need to know how to map the contentId in the table above to the itemId.\n<li> We'll need two files: a \"rows\" dataset where all the items for a particular user are listed; and a \"columns\" dataset where all the users for a particular item are listed.\n</ol>\n\n<p>\n\n### Mapping",
"_____no_output_____"
]
],
[
[
"import pandas as pd\nimport numpy as np\ndef create_mapping(values, filename):\n with open(filename, 'w') as ofp:\n value_to_id = {value:idx for idx, value in enumerate(values.unique())}\n for value, idx in value_to_id.items():\n ofp.write(\"{},{}\\n\".format(value, idx))\n return value_to_id\n\ndf = pd.read_csv(filepath_or_buffer = \"data/collab_raw.csv\",\n header = None,\n names = [\"visitorId\", \"contentId\", \"rating\"],\n dtype = {\"visitorId\": str, \"contentId\": str, \"rating\": np.float})\ndf.to_csv(path_or_buf = \"data/collab_raw.csv\", index = False, header = False)\nuser_mapping = create_mapping(df[\"visitorId\"], \"data/users.csv\")\nitem_mapping = create_mapping(df[\"contentId\"], \"data/items.csv\")",
"_____no_output_____"
],
[
"!head -3 data/*.csv",
"==> data/collab_raw.csv <==\n7337153711992174438,100074831,0.2321051400452234\n5190801220865459604,100170790,1.0\n2293633612703952721,100510126,0.2481776360816793\n\n==> data/items.csv <==\n727741,5272\n179038175,626\n299458287,4513\n\n==> data/users.csv <==\n6319375062712956077,33748\n7933447845885715412,47057\n5774017011910110015,76528\n"
],
[
"df[\"userId\"] = df[\"visitorId\"].map(user_mapping.get)\ndf[\"itemId\"] = df[\"contentId\"].map(item_mapping.get)",
"_____no_output_____"
],
[
"mapped_df = df[[\"userId\", \"itemId\", \"rating\"]]\nmapped_df.to_csv(path_or_buf = \"data/collab_mapped.csv\", index = False, header = False)\nmapped_df.head()",
"_____no_output_____"
]
],
[
[
"### Creating rows and columns datasets",
"_____no_output_____"
]
],
[
[
"import pandas as pd\nimport numpy as np\nmapped_df = pd.read_csv(filepath_or_buffer = \"data/collab_mapped.csv\", header = None, names = [\"userId\", \"itemId\", \"rating\"])\nmapped_df.head()",
"_____no_output_____"
],
[
"NITEMS = np.max(mapped_df[\"itemId\"]) + 1\nNUSERS = np.max(mapped_df[\"userId\"]) + 1\nmapped_df[\"rating\"] = np.round(mapped_df[\"rating\"].values, 2)\nprint(\"{} items, {} users, {} interactions\".format( NITEMS, NUSERS, len(mapped_df) ))",
"5721 items, 82902 users, 279594 interactions\n"
],
[
"grouped_by_items = mapped_df.groupby(\"itemId\")\niter = 0\nfor item, grouped in grouped_by_items:\n print(item, grouped[\"userId\"].values, grouped[\"rating\"].values)\n iter = iter + 1\n if iter > 5:\n break",
"0 [0] [0.23]\n1 [1] [1.]\n2 [2 3] [0.25 0.17]\n3 [4] [0.05]\n4 [5] [0.95]\n5 [6 7] [1. 0.48]\n"
],
[
"import tensorflow as tf\ngrouped_by_items = mapped_df.groupby(\"itemId\")\nwith tf.python_io.TFRecordWriter(\"data/users_for_item\") as ofp:\n for item, grouped in grouped_by_items:\n example = tf.train.Example(features = tf.train.Features(feature = {\n \"key\": tf.train.Feature(int64_list = tf.train.Int64List(value = [item])),\n \"indices\": tf.train.Feature(int64_list = tf.train.Int64List(value = grouped[\"userId\"].values)),\n \"values\": tf.train.Feature(float_list = tf.train.FloatList(value = grouped[\"rating\"].values))\n }))\n ofp.write(example.SerializeToString())",
"_____no_output_____"
],
[
"grouped_by_users = mapped_df.groupby(\"userId\")\nwith tf.python_io.TFRecordWriter(\"data/items_for_user\") as ofp:\n for user, grouped in grouped_by_users:\n example = tf.train.Example(features = tf.train.Features(feature = {\n \"key\": tf.train.Feature(int64_list = tf.train.Int64List(value = [user])),\n \"indices\": tf.train.Feature(int64_list = tf.train.Int64List(value = grouped[\"itemId\"].values)),\n \"values\": tf.train.Feature(float_list = tf.train.FloatList(value = grouped[\"rating\"].values))\n }))\n ofp.write(example.SerializeToString())",
"_____no_output_____"
],
[
"!ls -lrt data",
"total 31908\n-rw-r--r-- 1 jupyter jupyter 13152765 Jul 31 20:41 collab_raw.csv\n-rw-r--r-- 1 jupyter jupyter 2134511 Jul 31 20:41 users.csv\n-rw-r--r-- 1 jupyter jupyter 82947 Jul 31 20:41 items.csv\n-rw-r--r-- 1 jupyter jupyter 7812739 Jul 31 20:41 collab_mapped.csv\n-rw-r--r-- 1 jupyter jupyter 2252828 Jul 31 20:41 users_for_item\n-rw-r--r-- 1 jupyter jupyter 7217822 Jul 31 20:41 items_for_user\n"
]
],
[
[
"To summarize, we created the following data files from collab_raw.csv:\n<ol>\n<li> ```collab_mapped.csv``` is essentially the same data as in ```collab_raw.csv``` except that ```visitorId``` and ```contentId``` which are business-specific have been mapped to ```userId``` and ```itemId``` which are enumerated in 0,1,2,.... The mappings themselves are stored in ```items.csv``` and ```users.csv``` so that they can be used during inference.\n<li> ```users_for_item``` contains all the users/ratings for each item in TFExample format\n<li> ```items_for_user``` contains all the items/ratings for each user in TFExample format\n</ol>",
"_____no_output_____"
],
[
"## Train with WALS\n\nOnce you have the dataset, do matrix factorization with WALS using the [WALSMatrixFactorization](https://www.tensorflow.org/versions/master/api_docs/python/tf/contrib/factorization/WALSMatrixFactorization) in the contrib directory.\nThis is an estimator model, so it should be relatively familiar.\n<p>\nAs usual, we write an input_fn to provide the data to the model, and then create the Estimator to do train_and_evaluate.\nBecause it is in contrib and hasn't moved over to tf.estimator yet, we use tf.contrib.learn.Experiment to handle the training loop.<p>\nMake sure to replace <strong># TODO</strong> in below code.",
"_____no_output_____"
]
],
[
[
"import os\nimport tensorflow as tf\nfrom tensorflow.python.lib.io import file_io\nfrom tensorflow.contrib.factorization import WALSMatrixFactorization\n \ndef read_dataset(mode, args):\n def decode_example(protos, vocab_size):\n # TODO\n return\n \n \n def remap_keys(sparse_tensor):\n # Current indices of our SparseTensor that we need to fix\n bad_indices = sparse_tensor.indices # shape = (current_batch_size * (number_of_items/users[i] + 1), 2)\n # Current values of our SparseTensor that we need to fix\n bad_values = sparse_tensor.values # shape = (current_batch_size * (number_of_items/users[i] + 1),)\n\n # Since batch is ordered, the last value for a batch index is the user\n # Find where the batch index chages to extract the user rows\n # 1 where user, else 0\n user_mask = tf.concat(values = [bad_indices[1:,0] - bad_indices[:-1,0], tf.constant(value = [1], dtype = tf.int64)], axis = 0) # shape = (current_batch_size * (number_of_items/users[i] + 1), 2)\n\n # Mask out the user rows from the values\n good_values = tf.boolean_mask(tensor = bad_values, mask = tf.equal(x = user_mask, y = 0)) # shape = (current_batch_size * number_of_items/users[i],)\n item_indices = tf.boolean_mask(tensor = bad_indices, mask = tf.equal(x = user_mask, y = 0)) # shape = (current_batch_size * number_of_items/users[i],)\n user_indices = tf.boolean_mask(tensor = bad_indices, mask = tf.equal(x = user_mask, y = 1))[:, 1] # shape = (current_batch_size,)\n\n good_user_indices = tf.gather(params = user_indices, indices = item_indices[:,0]) # shape = (current_batch_size * number_of_items/users[i],)\n\n # User and item indices are rank 1, need to make rank 1 to concat\n good_user_indices_expanded = tf.expand_dims(input = good_user_indices, axis = -1) # shape = (current_batch_size * number_of_items/users[i], 1)\n good_item_indices_expanded = tf.expand_dims(input = item_indices[:, 1], axis = -1) # shape = (current_batch_size * number_of_items/users[i], 1)\n good_indices = tf.concat(values = [good_user_indices_expanded, good_item_indices_expanded], axis = 1) # shape = (current_batch_size * number_of_items/users[i], 2)\n\n remapped_sparse_tensor = tf.SparseTensor(indices = good_indices, values = good_values, dense_shape = sparse_tensor.dense_shape)\n return remapped_sparse_tensor\n\n \n def parse_tfrecords(filename, vocab_size):\n if mode == tf.estimator.ModeKeys.TRAIN:\n num_epochs = None # indefinitely\n else:\n num_epochs = 1 # end-of-input after this\n\n files = tf.gfile.Glob(filename = os.path.join(args[\"input_path\"], filename))\n\n # Create dataset from file list\n dataset = tf.data.TFRecordDataset(files)\n dataset = dataset.map(map_func = lambda x: decode_example(x, vocab_size))\n dataset = dataset.repeat(count = num_epochs)\n dataset = dataset.batch(batch_size = args[\"batch_size\"])\n dataset = dataset.map(map_func = lambda x: remap_keys(x))\n return dataset.make_one_shot_iterator().get_next()\n \n def _input_fn():\n features = {\n WALSMatrixFactorization.INPUT_ROWS: parse_tfrecords(\"items_for_user\", args[\"nitems\"]),\n WALSMatrixFactorization.INPUT_COLS: parse_tfrecords(\"users_for_item\", args[\"nusers\"]),\n WALSMatrixFactorization.PROJECT_ROW: tf.constant(True)\n }\n return features, None\n\n return _input_fn\n \n def input_cols():\n return parse_tfrecords(\"users_for_item\", args[\"nusers\"])\n \n return _input_fn#_subset",
"_____no_output_____"
]
],
[
[
"This code is helpful in developing the input function. You don't need it in production.",
"_____no_output_____"
]
],
[
[
"def try_out():\n with tf.Session() as sess:\n fn = read_dataset(\n mode = tf.estimator.ModeKeys.EVAL, \n args = {\"input_path\": \"data\", \"batch_size\": 4, \"nitems\": NITEMS, \"nusers\": NUSERS})\n feats, _ = fn()\n \n print(feats[\"input_rows\"].eval())\n print(feats[\"input_rows\"].eval())\n\ntry_out()",
"SparseTensorValue(indices=array([[ 0, 0],\n [ 0, 3522],\n [ 0, 3583],\n [ 1, 1],\n [ 1, 2359],\n [ 1, 3133],\n [ 1, 4864],\n [ 1, 4901],\n [ 1, 4906],\n [ 1, 5667],\n [ 2, 2],\n [ 3, 2],\n [ 3, 1467]]), values=array([0.23, 0.05, 0.18, 1. , 0.11, 0.55, 0.3 , 0.72, 0.46, 0.3 , 0.25,\n 0.17, 0.13], dtype=float32), dense_shape=array([ 4, 5721]))\nSparseTensorValue(indices=array([[ 4, 3],\n [ 5, 4],\n [ 5, 5042],\n [ 5, 5525],\n [ 5, 5553],\n [ 6, 5],\n [ 7, 5]]), values=array([0.05, 0.95, 0.63, 1. , 0.16, 1. , 0.48], dtype=float32), dense_shape=array([ 4, 5721]))\n"
],
[
"def find_top_k(user, item_factors, k):\n all_items = tf.matmul(a = tf.expand_dims(input = user, axis = 0), b = tf.transpose(a = item_factors))\n topk = tf.nn.top_k(input = all_items, k = k)\n return tf.cast(x = topk.indices, dtype = tf.int64)\n \ndef batch_predict(args):\n import numpy as np\n with tf.Session() as sess:\n estimator = tf.contrib.factorization.WALSMatrixFactorization(\n num_rows = args[\"nusers\"], \n num_cols = args[\"nitems\"],\n embedding_dimension = args[\"n_embeds\"],\n model_dir = args[\"output_dir\"])\n \n # This is how you would get the row factors for out-of-vocab user data\n # row_factors = list(estimator.get_projections(input_fn=read_dataset(tf.estimator.ModeKeys.EVAL, args)))\n # user_factors = tf.convert_to_tensor(np.array(row_factors))\n\n # But for in-vocab data, the row factors are already in the checkpoint\n user_factors = tf.convert_to_tensor(value = estimator.get_row_factors()[0]) # (nusers, nembeds)\n # In either case, we have to assume catalog doesn\"t change, so col_factors are read in\n item_factors = tf.convert_to_tensor(value = estimator.get_col_factors()[0])# (nitems, nembeds)\n\n # For each user, find the top K items\n topk = tf.squeeze(input = tf.map_fn(fn = lambda user: find_top_k(user, item_factors, args[\"topk\"]), elems = user_factors, dtype = tf.int64))\n with file_io.FileIO(os.path.join(args[\"output_dir\"], \"batch_pred.txt\"), mode = 'w') as f:\n for best_items_for_user in topk.eval():\n f.write(\",\".join(str(x) for x in best_items_for_user) + '\\n')\n\ndef train_and_evaluate(args):\n train_steps = int(0.5 + (1.0 * args[\"num_epochs\"] * args[\"nusers\"]) / args[\"batch_size\"])\n steps_in_epoch = int(0.5 + args[\"nusers\"] / args[\"batch_size\"])\n print(\"Will train for {} steps, evaluating once every {} steps\".format(train_steps, steps_in_epoch))\n def experiment_fn(output_dir):\n return tf.contrib.learn.Experiment(\n tf.contrib.factorization.WALSMatrixFactorization(\n num_rows = args[\"nusers\"], \n num_cols = args[\"nitems\"],\n embedding_dimension = args[\"n_embeds\"],\n model_dir = args[\"output_dir\"]),\n train_input_fn = read_dataset(tf.estimator.ModeKeys.TRAIN, args),\n eval_input_fn = read_dataset(tf.estimator.ModeKeys.EVAL, args),\n train_steps = train_steps,\n eval_steps = 1,\n min_eval_frequency = steps_in_epoch\n )\n\n from tensorflow.contrib.learn.python.learn import learn_runner\n learn_runner.run(experiment_fn = experiment_fn, output_dir = args[\"output_dir\"])\n \n batch_predict(args)",
"_____no_output_____"
],
[
"import shutil\nshutil.rmtree(path = \"wals_trained\", ignore_errors=True)\ntrain_and_evaluate({\n \"output_dir\": \"wals_trained\",\n \"input_path\": \"data/\",\n \"num_epochs\": 0.05,\n \"nitems\": NITEMS,\n \"nusers\": NUSERS,\n\n \"batch_size\": 512,\n \"n_embeds\": 10,\n \"topk\": 3\n })",
"Will train for 8 steps, evaluating once every 162 steps\nWARNING:tensorflow:From <ipython-input-25-4ad1e7c785ce>:49: run (from tensorflow.contrib.learn.python.learn.learn_runner) is deprecated and will be removed in a future version.\nInstructions for updating:\nUse tf.estimator.train_and_evaluate.\nWARNING:tensorflow:From /home/jupyter/.local/lib/python3.5/site-packages/tensorflow/contrib/learn/python/learn/estimators/estimator.py:1179: BaseEstimator.__init__ (from tensorflow.contrib.learn.python.learn.estimators.estimator) is deprecated and will be removed in a future version.\nInstructions for updating:\nPlease replace uses of any Estimator from tf.contrib.learn with an Estimator from tf.estimator.*\nWARNING:tensorflow:From /home/jupyter/.local/lib/python3.5/site-packages/tensorflow/contrib/learn/python/learn/estimators/estimator.py:427: RunConfig.__init__ (from tensorflow.contrib.learn.python.learn.estimators.run_config) is deprecated and will be removed in a future version.\nInstructions for updating:\nWhen switching to tf.estimator.Estimator, use tf.estimator.RunConfig instead.\nINFO:tensorflow:Using default config.\nINFO:tensorflow:Using config: {'_tf_config': gpu_options {\n per_process_gpu_memory_fraction: 1.0\n}\n, '_task_id': 0, '_is_chief': True, '_cluster_spec': <tensorflow.python.training.server_lib.ClusterSpec object at 0x7fb220516da0>, '_save_checkpoints_steps': None, '_task_type': None, '_tf_random_seed': None, '_save_summary_steps': 100, '_num_ps_replicas': 0, '_keep_checkpoint_max': 5, '_log_step_count_steps': 100, '_keep_checkpoint_every_n_hours': 10000, '_environment': 'local', '_eval_distribute': None, '_session_config': None, '_train_distribute': None, '_evaluation_master': '', '_num_worker_replicas': 0, '_device_fn': None, '_master': '', '_protocol': None, '_save_checkpoints_secs': 600, '_model_dir': 'wals_trained'}\nWARNING:tensorflow:From <ipython-input-25-4ad1e7c785ce>:45: Experiment.__init__ (from tensorflow.contrib.learn.python.learn.experiment) is deprecated and will be removed in a future version.\nInstructions for updating:\nPlease switch to tf.estimator.train_and_evaluate. You will also have to convert to a tf.estimator.Estimator.\nWARNING:tensorflow:From /home/jupyter/.local/lib/python3.5/site-packages/tensorflow/contrib/learn/python/learn/monitors.py:279: BaseMonitor.__init__ (from tensorflow.contrib.learn.python.learn.monitors) is deprecated and will be removed after 2016-12-05.\nInstructions for updating:\nMonitors are deprecated. Please use tf.train.SessionRunHook.\nWARNING:tensorflow:From /home/jupyter/.local/lib/python3.5/site-packages/tensorflow/python/framework/op_def_library.py:263: colocate_with (from tensorflow.python.framework.ops) is deprecated and will be removed in a future version.\nInstructions for updating:\nColocations handled automatically by placer.\nWARNING:tensorflow:From /home/jupyter/.local/lib/python3.5/site-packages/tensorflow/contrib/factorization/python/ops/wals.py:315: ModelFnOps.__new__ (from tensorflow.contrib.learn.python.learn.estimators.model_fn) is deprecated and will be removed in a future version.\nInstructions for updating:\nWhen switching to tf.estimator.Estimator, use tf.estimator.EstimatorSpec. You can use the `estimator_spec` method to create an equivalent one.\nINFO:tensorflow:Create CheckpointSaverHook.\nINFO:tensorflow:Graph was finalized.\nINFO:tensorflow:Running local_init_op.\nINFO:tensorflow:Done running local_init_op.\nINFO:tensorflow:Saving checkpoints for 0 into wals_trained/model.ckpt.\nINFO:tensorflow:SweepHook running init op.\nINFO:tensorflow:SweepHook running prep ops for the row sweep.\nINFO:tensorflow:Next fit step starting.\nINFO:tensorflow:loss = 96509.96, step = 1\nINFO:tensorflow:Next fit step starting.\nINFO:tensorflow:Next fit step starting.\nINFO:tensorflow:Next fit step starting.\nINFO:tensorflow:Next fit step starting.\nINFO:tensorflow:Next fit step starting.\nINFO:tensorflow:Next fit step starting.\nINFO:tensorflow:Next fit step starting.\nINFO:tensorflow:Saving checkpoints for 8 into wals_trained/model.ckpt.\nINFO:tensorflow:Loss for final step: 110142.75.\nWARNING:tensorflow:From /home/jupyter/.local/lib/python3.5/site-packages/tensorflow/python/ops/metrics_impl.py:363: to_float (from tensorflow.python.ops.math_ops) is deprecated and will be removed in a future version.\nInstructions for updating:\nUse tf.cast instead.\nINFO:tensorflow:Starting evaluation at 2019-07-31T20:43:12Z\nINFO:tensorflow:Graph was finalized.\nWARNING:tensorflow:From /home/jupyter/.local/lib/python3.5/site-packages/tensorflow/python/training/saver.py:1266: checkpoint_exists (from tensorflow.python.training.checkpoint_management) is deprecated and will be removed in a future version.\nInstructions for updating:\nUse standard file APIs to check for files with this prefix.\nINFO:tensorflow:Restoring parameters from wals_trained/model.ckpt-8\nINFO:tensorflow:Running local_init_op.\nINFO:tensorflow:Done running local_init_op.\nINFO:tensorflow:Evaluation [1/1]\nINFO:tensorflow:Finished evaluation at 2019-07-31-20:43:12\nINFO:tensorflow:Saving dict for global step 8: global_step = 8, loss = 96509.96\nINFO:tensorflow:Using default config.\nINFO:tensorflow:Using config: {'_tf_config': gpu_options {\n per_process_gpu_memory_fraction: 1.0\n}\n, '_task_id': 0, '_is_chief': True, '_cluster_spec': <tensorflow.python.training.server_lib.ClusterSpec object at 0x7fb2207064e0>, '_save_checkpoints_steps': None, '_task_type': None, '_tf_random_seed': None, '_save_summary_steps': 100, '_num_ps_replicas': 0, '_keep_checkpoint_max': 5, '_log_step_count_steps': 100, '_keep_checkpoint_every_n_hours': 10000, '_environment': 'local', '_eval_distribute': None, '_session_config': None, '_train_distribute': None, '_evaluation_master': '', '_num_worker_replicas': 0, '_device_fn': None, '_master': '', '_protocol': None, '_save_checkpoints_secs': 600, '_model_dir': 'wals_trained'}\n"
],
[
"!ls wals_trained",
"batch_pred.txt\t\t\t model.ckpt-0.index\ncheckpoint\t\t\t model.ckpt-0.meta\neval\t\t\t\t model.ckpt-8.data-00000-of-00001\nevents.out.tfevents.1564605788.r model.ckpt-8.index\ngraph.pbtxt\t\t\t model.ckpt-8.meta\nmodel.ckpt-0.data-00000-of-00001\n"
],
[
"!head wals_trained/batch_pred.txt",
"284,5609,36\n284,2754,42\n284,3168,534\n2621,5528,2694\n4409,5295,343\n5161,3267,3369\n5479,1335,55\n5479,1335,55\n4414,284,5572\n284,241,2359\n"
]
],
[
[
"## Run as a Python module\n\nLet's run it as Python module for just a few steps.",
"_____no_output_____"
]
],
[
[
"os.environ[\"NITEMS\"] = str(NITEMS)\nos.environ[\"NUSERS\"] = str(NUSERS)",
"_____no_output_____"
],
[
"%%bash\nrm -rf wals.tar.gz wals_trained\ngcloud ai-platform local train \\\n --module-name=walsmodel.task \\\n --package-path=${PWD}/walsmodel \\\n -- \\\n --output_dir=${PWD}/wals_trained \\\n --input_path=${PWD}/data \\\n --num_epochs=0.01 --nitems=${NITEMS} --nusers=${NUSERS} \\\n --job-dir=./tmp",
"Will train for 2 steps, evaluating once every 162 steps\n"
]
],
[
[
"## Run on Cloud",
"_____no_output_____"
]
],
[
[
"%%bash\ngsutil -m cp data/* gs://${BUCKET}/wals/data",
"_____no_output_____"
],
[
"%%bash\nOUTDIR=gs://${BUCKET}/wals/model_trained\nJOBNAME=wals_$(date -u +%y%m%d_%H%M%S)\necho $OUTDIR $REGION $JOBNAME\ngsutil -m rm -rf $OUTDIR\ngcloud ai-platform jobs submit training $JOBNAME \\\n --region=$REGION \\\n --module-name=walsmodel.task \\\n --package-path=${PWD}/walsmodel \\\n --job-dir=$OUTDIR \\\n --staging-bucket=gs://$BUCKET \\\n --scale-tier=BASIC_GPU \\\n --runtime-version=$TFVERSION \\\n -- \\\n --output_dir=$OUTDIR \\\n --input_path=gs://${BUCKET}/wals/data \\\n --num_epochs=10 --nitems=${NITEMS} --nusers=${NUSERS} ",
"_____no_output_____"
]
],
[
[
"This took <b>10 minutes</b> for me.",
"_____no_output_____"
],
[
"## Get row and column factors\n\nOnce you have a trained WALS model, you can get row and column factors (user and item embeddings) from the checkpoint file. We'll look at how to use these in the section on building a recommendation system using deep neural networks.",
"_____no_output_____"
]
],
[
[
"def get_factors(args):\n with tf.Session() as sess:\n estimator = tf.contrib.factorization.WALSMatrixFactorization(\n num_rows = args[\"nusers\"], \n num_cols = args[\"nitems\"],\n embedding_dimension = args[\"n_embeds\"],\n model_dir = args[\"output_dir\"])\n \n row_factors = estimator.get_row_factors()[0]\n col_factors = estimator.get_col_factors()[0]\n return row_factors, col_factors",
"_____no_output_____"
],
[
"args = {\n \"output_dir\": \"gs://{}/wals/model_trained\".format(BUCKET),\n \"nitems\": NITEMS,\n \"nusers\": NUSERS,\n \"n_embeds\": 10\n }\n\nuser_embeddings, item_embeddings = get_factors(args)\nprint(user_embeddings[:3])\nprint(item_embeddings[:3])",
"INFO:tensorflow:Using default config.\nINFO:tensorflow:Using config: {'_environment': 'local', '_is_chief': True, '_keep_checkpoint_every_n_hours': 10000, '_num_worker_replicas': 0, '_session_config': None, '_task_type': None, '_eval_distribute': None, '_tf_config': gpu_options {\n per_process_gpu_memory_fraction: 1.0\n}\n, '_master': '', '_log_step_count_steps': 100, '_model_dir': 'gs://qwiklabs-gcp-cbc8684b07fc2dbd-bucket/wals/model_trained', '_cluster_spec': <tensorflow.python.training.server_lib.ClusterSpec object at 0x7f4bd8302f28>, '_device_fn': None, '_keep_checkpoint_max': 5, '_task_id': 0, '_evaluation_master': '', '_save_checkpoints_steps': None, '_protocol': None, '_train_distribute': None, '_save_checkpoints_secs': 600, '_save_summary_steps': 100, '_tf_random_seed': None, '_num_ps_replicas': 0}\n[[ 3.3451824e-06 -1.1986867e-05 4.8447573e-06 -1.5209486e-05\n -1.7004859e-07 1.1976428e-05 9.8887876e-06 7.2386983e-06\n -7.0237149e-07 -7.9796819e-06]\n [-2.5300323e-03 1.4055537e-03 -9.8291773e-04 -4.2533795e-03\n -1.4166030e-03 -1.9530674e-03 8.5932651e-04 -1.5276540e-03\n 2.1342330e-03 1.2041229e-03]\n [ 9.5228699e-21 5.5453966e-21 2.2947056e-21 -5.8859543e-21\n 7.7516509e-21 -2.7640896e-20 2.3587296e-20 -3.9876822e-21\n 1.7312470e-20 2.5409211e-20]]\n[[-1.2125404e-06 -8.6304914e-05 4.4657736e-05 -6.8423047e-05\n 5.8551927e-06 9.7241784e-05 6.6776753e-05 1.6673854e-05\n -1.2708440e-05 -5.1148414e-05]\n [-1.1353870e-01 5.9097271e-02 -4.6105500e-02 -1.5460028e-01\n -1.9166643e-02 -7.3236257e-02 3.5582058e-02 -5.6805085e-02\n 7.5831160e-02 7.5306065e-02]\n [ 7.1989548e-20 4.4574543e-20 6.5149121e-21 -4.6291777e-20\n 8.8196718e-20 -2.3245078e-19 1.9459292e-19 4.0191465e-20\n 1.6273659e-19 2.2836562e-19]]\n"
]
],
[
[
"You can visualize the embedding vectors using dimensional reduction techniques such as PCA.",
"_____no_output_____"
]
],
[
[
"import matplotlib.pyplot as plt\nfrom mpl_toolkits.mplot3d import Axes3D\nfrom sklearn.decomposition import PCA\n\npca = PCA(n_components = 3)\npca.fit(user_embeddings)\nuser_embeddings_pca = pca.transform(user_embeddings)\n\nfig = plt.figure(figsize = (8,8))\nax = fig.add_subplot(111, projection = \"3d\")\nxs, ys, zs = user_embeddings_pca[::150].T\nax.scatter(xs, ys, zs)",
"_____no_output_____"
]
],
[
[
"<pre>\n# Copyright 2018 Google Inc. All Rights Reserved.\n#\n# Licensed under the Apache License, Version 2.0 (the \"License\");\n# you may not use this file except in compliance with the License.\n# You may obtain a copy of the License at\n#\n# http://www.apache.org/licenses/LICENSE-2.0\n#\n# Unless required by applicable law or agreed to in writing, software\n# distributed under the License is distributed on an \"AS IS\" BASIS,\n# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.\n# See the License for the specific language governing permissions and\n# limitations under the License.\n</pre>",
"_____no_output_____"
]
]
] | [
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown"
] | [
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
]
] |
cb00137c2859a191b4abbe35a8f447441e42874b | 3,926 | ipynb | Jupyter Notebook | week02/01_hello_world.ipynb | neolaw84/data_science_using_python_labs | 15bd2445577c5e254831f2acd97ea4b93a224229 | [
"Apache-2.0"
] | 1 | 2022-03-20T12:49:43.000Z | 2022-03-20T12:49:43.000Z | week02/01_hello_world.ipynb | neolaw84/data_science_using_python_labs | 15bd2445577c5e254831f2acd97ea4b93a224229 | [
"Apache-2.0"
] | null | null | null | week02/01_hello_world.ipynb | neolaw84/data_science_using_python_labs | 15bd2445577c5e254831f2acd97ea4b93a224229 | [
"Apache-2.0"
] | null | null | null | 23.650602 | 199 | 0.513245 | [
[
[
"# Welcome back to Data Science using Python\n\nဒီအပါတ်မှာ Python Programming Language ကို အသုံးပြုရပြီး Program တွေ ရေးသားရမှာ ဖြစ်ပါတယ်။",
"_____no_output_____"
]
],
[
[
"# ဒီ cell ကို run လိုက်ပါ။\n!git clone https://github.com/neolaw84/data_science_using_python_labs.git\n%cd week02",
"_____no_output_____"
]
],
[
[
"# Hello World\n\n\"Hello World\" လို့ print ထုတ်တဲ့ statement တကြောင်းထဲပါတဲ့ Python program ကို ရေးကြည့်ရအောင်။\n\n```python\n# beginning of program\nprint(\"Hello World\")\n# end of program\n```\n\nအပေါ်က program ကို အောက်က cell မှာ ကူးရေးပါ။",
"_____no_output_____"
]
],
[
[
"# copy the program here and run\n# you will see the output underneath\n",
"_____no_output_____"
],
[
"%%writefile lab_hello_world.py\n# copy the program here to make the program file called \"lab_hello_world.py\"\n",
"_____no_output_____"
],
[
"# program မှန် မမှန်ကို ဒီ cell ကို run ပြီး စမ်းသပ်ပါ။\n!pytest test_hello_world.py",
"_____no_output_____"
],
[
"# မင်္ဂလာပါ ကမ္ဘာကြီး လို့ output ထုတ်ပေးတဲ့ program ကို ဒီ cell မှာ ရေးကြည့်ပါ။\n# ပြီးရင် run စမ်းကြည့်ပါ။",
"_____no_output_____"
],
[
"%%writefile lab_hello_world_burmese.py\n# အပေါ်က မင်္ဂလာပါ ကမ္ဘာကြီး လို့ output ထုတ်ပေးတဲ့ program ကို ဒီ cell မှာ ပြန်ရေးပါ။\n# this is how to make a program file called \"lab_hello_world_burmese.py\"",
"_____no_output_____"
],
[
"# program မှန် မမှန်ကို ဒီ cell ကို run ပြီး စမ်းသပ်ပါ။\n!pytest test_hello_world_burmese.py",
"_____no_output_____"
],
[
"# \"My name is xxxxx\" လို့ ကိုယ့်နာမည်ကိုယ် output ထုတ်တဲ့ program ရေးကြည့်ပါ။\n",
"_____no_output_____"
],
[
"%%writefile lab_my_name.py\n# now create \"lab_my_name.py\" file containing the above program",
"_____no_output_____"
],
[
"# now test your program file here\n!pytest test_my_name.py",
"_____no_output_____"
]
],
[
[
"အခု အချိန်အထိ print statement တကြောင်းပါတဲ့ program ၃ ပုဒ် တိတိ ရေးသားခဲ့ပြီးပြီ ဖြစ်တယ်။ အဲဒီ program တွေကို ကိုယ်တိုင်လဲ run ပြီး စမ်းသလို ကြိုရေးထားတဲ့ unit test တွေနဲ့လဲ စမ်းသပ်ခဲ့ကြတယ်။\n\nအခုတော့ သီအိုရီ နည်းနည်း ပြောကြဦးစို့။",
"_____no_output_____"
]
]
] | [
"markdown",
"code",
"markdown",
"code",
"markdown"
] | [
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
]
] |
cb001896370063c3812fff36ec0da8f94ff0f151 | 220,687 | ipynb | Jupyter Notebook | head_view_gpt2.ipynb | aavshr/bertviz | f3e2c2459f58976132cb7b4ccf8383ed5e65dce0 | [
"Apache-2.0"
] | null | null | null | head_view_gpt2.ipynb | aavshr/bertviz | f3e2c2459f58976132cb7b4ccf8383ed5e65dce0 | [
"Apache-2.0"
] | null | null | null | head_view_gpt2.ipynb | aavshr/bertviz | f3e2c2459f58976132cb7b4ccf8383ed5e65dce0 | [
"Apache-2.0"
] | null | null | null | 445.832323 | 202,154 | 0.738322 | [
[
[
"from bertviz.pytorch_transformers_attn import GPT2Model, GPT2Tokenizer\nfrom bertviz.head_view_gpt2 import show",
"_____no_output_____"
],
[
"%%javascript\nrequire.config({\n paths: {\n d3: '//cdnjs.cloudflare.com/ajax/libs/d3/3.4.8/d3.min',\n jquery: '//ajax.googleapis.com/ajax/libs/jquery/2.0.0/jquery.min',\n }\n});",
"_____no_output_____"
],
[
"model = GPT2Model.from_pretrained('gpt2')\ntokenizer = GPT2Tokenizer.from_pretrained('gpt2')\ntext = \"The quick brown fox ran after the lazy dogs.\"\nshow(model, tokenizer, text)",
"_____no_output_____"
]
]
] | [
"code"
] | [
[
"code",
"code",
"code"
]
] |
cb0033787afaececef01d60b3da70c4aa943670a | 1,472 | ipynb | Jupyter Notebook | 04-Career-Modules/Interview_Questions/09-Most_Frequent/Unsolved/Most_Frequent.ipynb | anirudhmungre/sneaky-lessons | 8e48015c50865059db96f8cd369bcc15365d66c7 | [
"ADSL"
] | null | null | null | 04-Career-Modules/Interview_Questions/09-Most_Frequent/Unsolved/Most_Frequent.ipynb | anirudhmungre/sneaky-lessons | 8e48015c50865059db96f8cd369bcc15365d66c7 | [
"ADSL"
] | null | null | null | 04-Career-Modules/Interview_Questions/09-Most_Frequent/Unsolved/Most_Frequent.ipynb | anirudhmungre/sneaky-lessons | 8e48015c50865059db96f8cd369bcc15365d66c7 | [
"ADSL"
] | null | null | null | 18.871795 | 75 | 0.516984 | [
[
[
"# Most Frequent Element\n\nWrite a function that finds the most frequent element in an array.\n\n\n`most_frequent([1, 2, 2, 3]) # 2`\n",
"_____no_output_____"
]
],
[
[
"numbers = [1, 2, 2, 3]",
"_____no_output_____"
]
]
] | [
"markdown",
"code"
] | [
[
"markdown"
],
[
"code"
]
] |
cb00337eed4fdc40e181952c76fa1d22fd25086f | 24,167 | ipynb | Jupyter Notebook | PythonDataScienceHandbook-master/notebooks/05.01-What-Is-Machine-Learning.ipynb | BioCogito/sagemakertest | 76733c23287959f5080404b7db3ef53a80cdd83a | [
"MIT"
] | null | null | null | PythonDataScienceHandbook-master/notebooks/05.01-What-Is-Machine-Learning.ipynb | BioCogito/sagemakertest | 76733c23287959f5080404b7db3ef53a80cdd83a | [
"MIT"
] | 2 | 2021-06-08T21:31:36.000Z | 2022-01-13T01:42:22.000Z | PythonDataScienceHandbook-master/notebooks/05.01-What-Is-Machine-Learning.ipynb | BioCogito/sagemakertest | 76733c23287959f5080404b7db3ef53a80cdd83a | [
"MIT"
] | null | null | null | 47.109162 | 396 | 0.687425 | [
[
[
"<!--BOOK_INFORMATION-->\n<img align=\"left\" style=\"padding-right:10px;\" src=\"figures/PDSH-cover-small.png\">\n\n*This notebook contains an excerpt from the [Python Data Science Handbook](http://shop.oreilly.com/product/0636920034919.do) by Jake VanderPlas; the content is available [on GitHub](https://github.com/jakevdp/PythonDataScienceHandbook).*\n\n*The text is released under the [CC-BY-NC-ND license](https://creativecommons.org/licenses/by-nc-nd/3.0/us/legalcode), and code is released under the [MIT license](https://opensource.org/licenses/MIT). If you find this content useful, please consider supporting the work by [buying the book](http://shop.oreilly.com/product/0636920034919.do)!*",
"_____no_output_____"
],
[
"<!--NAVIGATION-->\n< [Machine Learning](05.00-Machine-Learning.ipynb) | [Contents](Index.ipynb) | [Introducing Scikit-Learn](05.02-Introducing-Scikit-Learn.ipynb) >\n\n<a href=\"https://colab.research.google.com/github/jakevdp/PythonDataScienceHandbook/blob/master/notebooks/05.01-What-Is-Machine-Learning.ipynb\"><img align=\"left\" src=\"https://colab.research.google.com/assets/colab-badge.svg\" alt=\"Open in Colab\" title=\"Open and Execute in Google Colaboratory\"></a>\n",
"_____no_output_____"
],
[
"# What Is Machine Learning?",
"_____no_output_____"
],
[
"Before we take a look at the details of various machine learning methods, let's start by looking at what machine learning is, and what it isn't.\nMachine learning is often categorized as a subfield of artificial intelligence, but I find that categorization can often be misleading at first brush.\nThe study of machine learning certainly arose from research in this context, but in the data science application of machine learning methods, it's more helpful to think of machine learning as a means of *building models of data*.\n\nFundamentally, machine learning involves building mathematical models to help understand data.\n\"Learning\" enters the fray when we give these models *tunable parameters* that can be adapted to observed data; in this way the program can be considered to be \"learning\" from the data.\nOnce these models have been fit to previously seen data, they can be used to predict and understand aspects of newly observed data.\nI'll leave to the reader the more philosophical digression regarding the extent to which this type of mathematical, model-based \"learning\" is similar to the \"learning\" exhibited by the human brain.\n\nUnderstanding the problem setting in machine learning is essential to using these tools effectively, and so we will start with some broad categorizations of the types of approaches we'll discuss here.",
"_____no_output_____"
],
[
"## Categories of Machine Learning\n\nAt the most fundamental level, machine learning can be categorized into two main types: supervised learning and unsupervised learning.\n\n*Supervised learning* involves somehow modeling the relationship between measured features of data and some label associated with the data; once this model is determined, it can be used to apply labels to new, unknown data.\nThis is further subdivided into *classification* tasks and *regression* tasks: in classification, the labels are discrete categories, while in regression, the labels are continuous quantities.\nWe will see examples of both types of supervised learning in the following section.\n\n*Unsupervised learning* involves modeling the features of a dataset without reference to any label, and is often described as \"letting the dataset speak for itself.\"\nThese models include tasks such as *clustering* and *dimensionality reduction.*\nClustering algorithms identify distinct groups of data, while dimensionality reduction algorithms search for more succinct representations of the data.\nWe will see examples of both types of unsupervised learning in the following section.\n\nIn addition, there are so-called *semi-supervised learning* methods, which falls somewhere between supervised learning and unsupervised learning.\nSemi-supervised learning methods are often useful when only incomplete labels are available.",
"_____no_output_____"
],
[
"## Qualitative Examples of Machine Learning Applications\n\nTo make these ideas more concrete, let's take a look at a few very simple examples of a machine learning task.\nThese examples are meant to give an intuitive, non-quantitative overview of the types of machine learning tasks we will be looking at in this chapter.\nIn later sections, we will go into more depth regarding the particular models and how they are used.\nFor a preview of these more technical aspects, you can find the Python source that generates the following figures in the [Appendix: Figure Code](06.00-Figure-Code.ipynb).\n",
"_____no_output_____"
],
[
"### Classification: Predicting discrete labels\n\nWe will first take a look at a simple *classification* task, in which you are given a set of labeled points and want to use these to classify some unlabeled points.\n\nImagine that we have the data shown in this figure:",
"_____no_output_____"
],
[
"\n[figure source in Appendix](06.00-Figure-Code.ipynb#Classification-Example-Figure-1)",
"_____no_output_____"
],
[
"Here we have two-dimensional data: that is, we have two *features* for each point, represented by the *(x,y)* positions of the points on the plane.\nIn addition, we have one of two *class labels* for each point, here represented by the colors of the points.\nFrom these features and labels, we would like to create a model that will let us decide whether a new point should be labeled \"blue\" or \"red.\"\n\nThere are a number of possible models for such a classification task, but here we will use an extremely simple one. We will make the assumption that the two groups can be separated by drawing a straight line through the plane between them, such that points on each side of the line fall in the same group.\nHere the *model* is a quantitative version of the statement \"a straight line separates the classes\", while the *model parameters* are the particular numbers describing the location and orientation of that line for our data.\nThe optimal values for these model parameters are learned from the data (this is the \"learning\" in machine learning), which is often called *training the model*.\n\nThe following figure shows a visual representation of what the trained model looks like for this data:",
"_____no_output_____"
],
[
"\n[figure source in Appendix](06.00-Figure-Code.ipynb#Classification-Example-Figure-2)",
"_____no_output_____"
],
[
"Now that this model has been trained, it can be generalized to new, unlabeled data.\nIn other words, we can take a new set of data, draw this model line through it, and assign labels to the new points based on this model.\nThis stage is usually called *prediction*. See the following figure:",
"_____no_output_____"
],
[
"\n[figure source in Appendix](06.00-Figure-Code.ipynb#Classification-Example-Figure-3)",
"_____no_output_____"
],
[
"This is the basic idea of a classification task in machine learning, where \"classification\" indicates that the data has discrete class labels.\nAt first glance this may look fairly trivial: it would be relatively easy to simply look at this data and draw such a discriminatory line to accomplish this classification.\nA benefit of the machine learning approach, however, is that it can generalize to much larger datasets in many more dimensions.\n\nFor example, this is similar to the task of automated spam detection for email; in this case, we might use the following features and labels:\n\n- *feature 1*, *feature 2*, etc. $\\to$ normalized counts of important words or phrases (\"Viagra\", \"Nigerian prince\", etc.)\n- *label* $\\to$ \"spam\" or \"not spam\"\n\nFor the training set, these labels might be determined by individual inspection of a small representative sample of emails; for the remaining emails, the label would be determined using the model.\nFor a suitably trained classification algorithm with enough well-constructed features (typically thousands or millions of words or phrases), this type of approach can be very effective.\nWe will see an example of such text-based classification in [In Depth: Naive Bayes Classification](05.05-Naive-Bayes.ipynb).\n\nSome important classification algorithms that we will discuss in more detail are Gaussian naive Bayes (see [In Depth: Naive Bayes Classification](05.05-Naive-Bayes.ipynb)), support vector machines (see [In-Depth: Support Vector Machines](05.07-Support-Vector-Machines.ipynb)), and random forest classification (see [In-Depth: Decision Trees and Random Forests](05.08-Random-Forests.ipynb)).",
"_____no_output_____"
],
[
"### Regression: Predicting continuous labels\n\nIn contrast with the discrete labels of a classification algorithm, we will next look at a simple *regression* task in which the labels are continuous quantities.\n\nConsider the data shown in the following figure, which consists of a set of points each with a continuous label:",
"_____no_output_____"
],
[
"\n[figure source in Appendix](06.00-Figure-Code.ipynb#Regression-Example-Figure-1)",
"_____no_output_____"
],
[
"As with the classification example, we have two-dimensional data: that is, there are two features describing each data point.\nThe color of each point represents the continuous label for that point.\n\nThere are a number of possible regression models we might use for this type of data, but here we will use a simple linear regression to predict the points.\nThis simple linear regression model assumes that if we treat the label as a third spatial dimension, we can fit a plane to the data.\nThis is a higher-level generalization of the well-known problem of fitting a line to data with two coordinates.\n\nWe can visualize this setup as shown in the following figure:",
"_____no_output_____"
],
[
"\n[figure source in Appendix](06.00-Figure-Code.ipynb#Regression-Example-Figure-2)",
"_____no_output_____"
],
[
"Notice that the *feature 1-feature 2* plane here is the same as in the two-dimensional plot from before; in this case, however, we have represented the labels by both color and three-dimensional axis position.\nFrom this view, it seems reasonable that fitting a plane through this three-dimensional data would allow us to predict the expected label for any set of input parameters.\nReturning to the two-dimensional projection, when we fit such a plane we get the result shown in the following figure:",
"_____no_output_____"
],
[
"\n[figure source in Appendix](06.00-Figure-Code.ipynb#Regression-Example-Figure-3)",
"_____no_output_____"
],
[
"This plane of fit gives us what we need to predict labels for new points.\nVisually, we find the results shown in the following figure:",
"_____no_output_____"
],
[
"\n[figure source in Appendix](06.00-Figure-Code.ipynb#Regression-Example-Figure-4)",
"_____no_output_____"
],
[
"As with the classification example, this may seem rather trivial in a low number of dimensions.\nBut the power of these methods is that they can be straightforwardly applied and evaluated in the case of data with many, many features.\n\nFor example, this is similar to the task of computing the distance to galaxies observed through a telescope—in this case, we might use the following features and labels:\n\n- *feature 1*, *feature 2*, etc. $\\to$ brightness of each galaxy at one of several wave lengths or colors\n- *label* $\\to$ distance or redshift of the galaxy\n\nThe distances for a small number of these galaxies might be determined through an independent set of (typically more expensive) observations.\nDistances to remaining galaxies could then be estimated using a suitable regression model, without the need to employ the more expensive observation across the entire set.\nIn astronomy circles, this is known as the \"photometric redshift\" problem.\n\nSome important regression algorithms that we will discuss are linear regression (see [In Depth: Linear Regression](05.06-Linear-Regression.ipynb)), support vector machines (see [In-Depth: Support Vector Machines](05.07-Support-Vector-Machines.ipynb)), and random forest regression (see [In-Depth: Decision Trees and Random Forests](05.08-Random-Forests.ipynb)).",
"_____no_output_____"
],
[
"### Clustering: Inferring labels on unlabeled data\n\nThe classification and regression illustrations we just looked at are examples of supervised learning algorithms, in which we are trying to build a model that will predict labels for new data.\nUnsupervised learning involves models that describe data without reference to any known labels.\n\nOne common case of unsupervised learning is \"clustering,\" in which data is automatically assigned to some number of discrete groups.\nFor example, we might have some two-dimensional data like that shown in the following figure:",
"_____no_output_____"
],
[
"\n[figure source in Appendix](06.00-Figure-Code.ipynb#Clustering-Example-Figure-2)",
"_____no_output_____"
],
[
"By eye, it is clear that each of these points is part of a distinct group.\nGiven this input, a clustering model will use the intrinsic structure of the data to determine which points are related.\nUsing the very fast and intuitive *k*-means algorithm (see [In Depth: K-Means Clustering](05.11-K-Means.ipynb)), we find the clusters shown in the following figure:",
"_____no_output_____"
],
[
"\n[figure source in Appendix](06.00-Figure-Code.ipynb#Clustering-Example-Figure-2)",
"_____no_output_____"
],
[
"*k*-means fits a model consisting of *k* cluster centers; the optimal centers are assumed to be those that minimize the distance of each point from its assigned center.\nAgain, this might seem like a trivial exercise in two dimensions, but as our data becomes larger and more complex, such clustering algorithms can be employed to extract useful information from the dataset.\n\nWe will discuss the *k*-means algorithm in more depth in [In Depth: K-Means Clustering](05.11-K-Means.ipynb).\nOther important clustering algorithms include Gaussian mixture models (See [In Depth: Gaussian Mixture Models](05.12-Gaussian-Mixtures.ipynb)) and spectral clustering (See [Scikit-Learn's clustering documentation](http://scikit-learn.org/stable/modules/clustering.html)).",
"_____no_output_____"
],
[
"### Dimensionality reduction: Inferring structure of unlabeled data\n\nDimensionality reduction is another example of an unsupervised algorithm, in which labels or other information are inferred from the structure of the dataset itself.\nDimensionality reduction is a bit more abstract than the examples we looked at before, but generally it seeks to pull out some low-dimensional representation of data that in some way preserves relevant qualities of the full dataset.\nDifferent dimensionality reduction routines measure these relevant qualities in different ways, as we will see in [In-Depth: Manifold Learning](05.10-Manifold-Learning.ipynb).\n\nAs an example of this, consider the data shown in the following figure:",
"_____no_output_____"
],
[
"\n[figure source in Appendix](06.00-Figure-Code.ipynb#Dimensionality-Reduction-Example-Figure-1)",
"_____no_output_____"
],
[
"Visually, it is clear that there is some structure in this data: it is drawn from a one-dimensional line that is arranged in a spiral within this two-dimensional space.\nIn a sense, you could say that this data is \"intrinsically\" only one dimensional, though this one-dimensional data is embedded in higher-dimensional space.\nA suitable dimensionality reduction model in this case would be sensitive to this nonlinear embedded structure, and be able to pull out this lower-dimensionality representation.\n\nThe following figure shows a visualization of the results of the Isomap algorithm, a manifold learning algorithm that does exactly this:",
"_____no_output_____"
],
[
"\n[figure source in Appendix](06.00-Figure-Code.ipynb#Dimensionality-Reduction-Example-Figure-2)",
"_____no_output_____"
],
[
"Notice that the colors (which represent the extracted one-dimensional latent variable) change uniformly along the spiral, which indicates that the algorithm did in fact detect the structure we saw by eye.\nAs with the previous examples, the power of dimensionality reduction algorithms becomes clearer in higher-dimensional cases.\nFor example, we might wish to visualize important relationships within a dataset that has 100 or 1,000 features.\nVisualizing 1,000-dimensional data is a challenge, and one way we can make this more manageable is to use a dimensionality reduction technique to reduce the data to two or three dimensions.\n\nSome important dimensionality reduction algorithms that we will discuss are principal component analysis (see [In Depth: Principal Component Analysis](05.09-Principal-Component-Analysis.ipynb)) and various manifold learning algorithms, including Isomap and locally linear embedding (See [In-Depth: Manifold Learning](05.10-Manifold-Learning.ipynb)).",
"_____no_output_____"
],
[
"## Summary\n\nHere we have seen a few simple examples of some of the basic types of machine learning approaches.\nNeedless to say, there are a number of important practical details that we have glossed over, but I hope this section was enough to give you a basic idea of what types of problems machine learning approaches can solve.\n\nIn short, we saw the following:\n\n- *Supervised learning*: Models that can predict labels based on labeled training data\n\n - *Classification*: Models that predict labels as two or more discrete categories\n - *Regression*: Models that predict continuous labels\n \n- *Unsupervised learning*: Models that identify structure in unlabeled data\n\n - *Clustering*: Models that detect and identify distinct groups in the data\n - *Dimensionality reduction*: Models that detect and identify lower-dimensional structure in higher-dimensional data\n \nIn the following sections we will go into much greater depth within these categories, and see some more interesting examples of where these concepts can be useful.\n\nAll of the figures in the preceding discussion are generated based on actual machine learning computations; the code behind them can be found in [Appendix: Figure Code](06.00-Figure-Code.ipynb).",
"_____no_output_____"
],
[
"<!--NAVIGATION-->\n< [Machine Learning](05.00-Machine-Learning.ipynb) | [Contents](Index.ipynb) | [Introducing Scikit-Learn](05.02-Introducing-Scikit-Learn.ipynb) >\n\n<a href=\"https://colab.research.google.com/github/jakevdp/PythonDataScienceHandbook/blob/master/notebooks/05.01-What-Is-Machine-Learning.ipynb\"><img align=\"left\" src=\"https://colab.research.google.com/assets/colab-badge.svg\" alt=\"Open in Colab\" title=\"Open and Execute in Google Colaboratory\"></a>\n",
"_____no_output_____"
]
]
] | [
"markdown"
] | [
[
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown"
]
] |
cb003cbe369b87af5e36ae750ee795615a6de7f3 | 7,585 | ipynb | Jupyter Notebook | data structure/stack and queues/stack/python_stack_practice.ipynb | salviosage/Data-Structures-Algorithms-Nanodegree-playground-o | e1b1221a5809849d410a9eeddeb8b9a992c8f1b6 | [
"MIT"
] | null | null | null | data structure/stack and queues/stack/python_stack_practice.ipynb | salviosage/Data-Structures-Algorithms-Nanodegree-playground-o | e1b1221a5809849d410a9eeddeb8b9a992c8f1b6 | [
"MIT"
] | 1 | 2021-05-10T18:11:07.000Z | 2021-05-10T18:11:07.000Z | data structure/stack and queues/stack/python_stack_practice.ipynb | salviosage/Data-Structures-Algorithms-Nanodegree-playground-o | e1b1221a5809849d410a9eeddeb8b9a992c8f1b6 | [
"MIT"
] | null | null | null | 30.708502 | 986 | 0.518128 | [
[
[
"## Building a Stack in Python",
"_____no_output_____"
],
[
"Before we start let us reiterate they key components of a stack. A stack is a data structure that consists of two main operations: push and pop. A push is when you add an element to the **top of the stack** and a pop is when you remove an element from **the top of the stack**. Python 3.x conviently allows us to demonstate this functionality with a list. When you have a list such as [2,4,5,6] you can decide which end of the list is the bottom and the top of the stack respectivley. Once you decide that, you can use the append, pop or insert function to simulate a stack. We will choose the first element to be the bottom of our stack and therefore be using the append and pop functions to simulate it. Give it a try by implementing the function below!",
"_____no_output_____"
],
[
"#### Try Building a Stack",
"_____no_output_____"
]
],
[
[
"class Stack:\n def __init__(self):\n # TODO: Initialize the Stack\n \n def size(self):\n # TODO: Check the size of the Stack\n \n def push(self, item):\n # TODO: Push item onto Stack\n\n def pop(self):\n # TODO: Pop item off of the Stack",
"_____no_output_____"
]
],
[
[
"#### Test the Stack",
"_____no_output_____"
]
],
[
[
"MyStack = Stack()\n\nMyStack.push(\"Web Page 1\")\nMyStack.push(\"Web Page 2\")\nMyStack.push(\"Web Page 3\")\n\nprint (MyStack.items)\n\nMyStack.pop()\nMyStack.pop()\n\nprint (\"Pass\" if (MyStack.items[0] == 'Web Page 1') else \"Fail\")\n\nMyStack.pop()\n\nprint (\"Pass\" if (MyStack.pop() == None) else \"Fail\")",
"_____no_output_____"
]
],
[
[
"<span class=\"graffiti-highlight graffiti-id_3l78doc-id_l2kcjcz\"><i></i><button>Hide Solution</button></span>",
"_____no_output_____"
]
],
[
[
"# Solution\n\nclass Stack:\n def __init__(self):\n self.items = []\n \n def size(self):\n return len(self.items)\n \n def push(self, item):\n self.items.append(item)\n\n def pop(self):\n if self.size()==0:\n return None\n else:\n return self.items.pop()\n \nMyStack = Stack()\n\nMyStack.push(\"Web Page 1\")\nMyStack.push(\"Web Page 2\")\nMyStack.push(\"Web Page 3\")\n\nprint (MyStack.items)\n\nMyStack.pop()\nMyStack.pop()\n\nprint (\"Pass\" if (MyStack.items[0] == 'Web Page 1') else \"Fail\")\n\nMyStack.pop()\n\nprint (\"Pass\" if (MyStack.pop() == None) else \"Fail\")# Solution\n\nclass Stack:\n def __init__(self):\n self.items = []\n \n def size(self):\n return len(self.items)\n \n def push(self, item):\n self.items.append(item)\n\n def pop(self):\n if self.size()==0:\n return None\n else:\n return self.items.pop()\n \nMyStack = Stack()\n\nMyStack.push(\"Web Page 1\")\nMyStack.push(\"Web Page 2\")\nMyStack.push(\"Web Page 3\")\n\nprint (MyStack.items)\n\nMyStack.pop()\nMyStack.pop()\n\nprint (\"Pass\" if (MyStack.items[0] == 'Web Page 1') else \"Fail\")\n\nMyStack.pop()\n\nprint (\"Pass\" if (MyStack.pop() == None) else \"Fail\")# Solution\n\nclass Stack:\n def __init__(self):\n self.items = []\n \n def size(self):\n return len(self.items)\n \n def push(self, item):\n self.items.append(item)\n\n def pop(self):\n if self.size()==0:\n return None\n else:\n return self.items.pop()\n \nMyStack = Stack()\n\nMyStack.push(\"Web Page 1\")\nMyStack.push(\"Web Page 2\")\nMyStack.push(\"Web Page 3\")\n\nprint (MyStack.items)\n\nMyStack.pop()\nMyStack.pop()\n\nprint (\"Pass\" if (MyStack.items[0] == 'Web Page 1') else \"Fail\")\n\nMyStack.pop()\n\nprint (\"Pass\" if (MyStack.pop() == None) else \"Fail\")\n",
"_____no_output_____"
]
]
] | [
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] | [
[
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
]
] |
cb0047f7a42e0a97d906877ab6d1227fcef94639 | 4,525 | ipynb | Jupyter Notebook | examples/ambassador/custom/ambassador_custom.ipynb | pachyderm/seldon-core | 01fc519a7268053b191c745c5ae6bafa0229d5ea | [
"Apache-2.0"
] | 1 | 2020-12-29T03:28:42.000Z | 2020-12-29T03:28:42.000Z | examples/ambassador/custom/ambassador_custom.ipynb | AminuIsrael/seldon-core | 2092ec5471267c3d69697b659376def20c211027 | [
"Apache-2.0"
] | 231 | 2020-08-10T08:38:42.000Z | 2021-08-02T20:56:49.000Z | examples/ambassador/custom/ambassador_custom.ipynb | AminuIsrael/seldon-core | 2092ec5471267c3d69697b659376def20c211027 | [
"Apache-2.0"
] | null | null | null | 25.138889 | 319 | 0.576354 | [
[
[
"# Custom URL prefix with Seldon and Ambassador\n\nThis notebook shows how you can deploy Seldon Deployments with custom Ambassador configuration.",
"_____no_output_____"
],
[
"## Setup Seldon Core\n\nUset the setup notebook to [Setup Cluster](../../../notebooks/seldon_core_setup.ipynb#Setup-Cluster) with [Ambassador Ingress](../../../notebooks/seldon_core_setup.ipynb#Ambassador) and [Install Seldon Core](../../seldon_core_setup.ipynb#Install-Seldon-Core). Instructions [also online](./seldon_core_setup.html).",
"_____no_output_____"
],
[
"## Launch main model\n\nWe will create a very simple Seldon Deployment with a dummy model image `seldonio/mock_classifier:1.0`. This deployment is named `example`. We will add custom Ambassador config which sets the Ambassador prefix to `/mycompany/ml`\n\nWe must ensure we set the correct service endpoint. Seldon Core creates an endpoint of the form:\n \n`<spec.name>-<predictor.name>.<namespace>:<port>`\n\nWhere\n\n * `<spec-name>` is the name you give to the Seldon Deployment spec: `example` below\n * `<predcitor.name>` is the predictor name in the Seldon Deployment: `single` below\n * `<namespace>` is the namespace your Seldon Deployment is deployed to\n * `<port>` is the port either 8000 for REST or 5000 for gRPC\n \nThis will allow you to set the `service` value in the Ambassador config you create. So for the example below we have:\n\n```\nservice: production-model-example.seldon:8000\n```\n \n ",
"_____no_output_____"
]
],
[
[
"!pygmentize model_custom_ambassador.json",
"_____no_output_____"
],
[
"!kubectl create -f model_custom_ambassador.json",
"_____no_output_____"
],
[
"!kubectl rollout status deploy/$(kubectl get deploy -l seldon-deployment-id=example-custom -o jsonpath='{.items[0].metadata.name}')",
"_____no_output_____"
]
],
[
[
"### Get predictions",
"_____no_output_____"
]
],
[
[
"from seldon_core.seldon_client import SeldonClient\nsc = SeldonClient(deployment_name=\"example-custom\",namespace=\"seldon\")",
"_____no_output_____"
]
],
[
[
"#### REST Request",
"_____no_output_____"
]
],
[
[
"r = sc.predict(gateway=\"ambassador\",transport=\"rest\",gateway_prefix=\"/mycompany/ml\")\nassert(r.success==True)\nprint(r)",
"_____no_output_____"
],
[
"!kubectl delete -f model_custom_ambassador.json",
"_____no_output_____"
]
]
] | [
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] | [
[
"markdown",
"markdown",
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
]
] |
cb006059007c897e062b9f999092265d9e5053f4 | 19,995 | ipynb | Jupyter Notebook | index.ipynb | Taphei/kawai_demo_tapani | 34ac36751a7700231ab38dda51bb870f6330a0a9 | [
"Apache-2.0"
] | null | null | null | index.ipynb | Taphei/kawai_demo_tapani | 34ac36751a7700231ab38dda51bb870f6330a0a9 | [
"Apache-2.0"
] | null | null | null | index.ipynb | Taphei/kawai_demo_tapani | 34ac36751a7700231ab38dda51bb870f6330a0a9 | [
"Apache-2.0"
] | null | null | null | 56.643059 | 813 | 0.661265 | [
[
[
"# hide\n%load_ext nb_black\n# nb_black if using jupyter",
"_____no_output_____"
]
],
[
[
"# Helsinki Machine Learning Project Template\n\n Template for open source ML and predictive analytics projects.",
"_____no_output_____"
],
[
"\n[](https://github.com/psf/black)\n[](https://badge.fury.io/gh/City-of-Helsinki%2Fml_project_template)\n\n\n\n\n",
"_____no_output_____"
],
[
" NOTE: Once you begin your work, rewrite this notebook (index.ipynb) so that it describes your project, and regenerate README by calling `nbdev_build_docs`",
"_____no_output_____"
],
[
"## About\n\nThis is a git repository template for Python-based open source ML and analytics projects.\n\nThe template assumes the concept of Notebook Development.\nThis means, that you do all the data science work inside notebooks.\nThere is no copy-pasting! We use the [nbdev](https://nbdev.fast.ai/) tool to build python modules and doc pages from the notebooks, automatically.\nThis way you always have your code, results and documentation as one. \nNotebooks can be executed with the [papermill](https://papermill.readthedocs.io/) tool for an automatic, \nwell documented model update workflow. Handy, isn't it?\n\nThe template assumes that you divide your machine learning project into 5 parts:\n\n0. Data - loading & preprocessing\n1. Model - Python class code & algorithm development\n2. Loss - model training & evaluation\n3. Workflow - automatic model update (reproduce steps 0.-2.)\n4. API - an interface to interact with a trained model\n\nEach part has their own notebook template, that you can follow to plan and do your development.\n\nIn addition, the template comes with a working Dockerfile and .devcontainer for doing your development easily with any device.\nYou can extend these for your needs and for building a runtime container for your machine learning app.\n\nThe template is completely open source and environment agnostic.\nFollow the installation instructions to create a new, \nindependent repository with clean commit history, \nbut with a copy of all the files and folders presented.\nThe authors of this template will not be contributors to your project,\nalthough we are more hear what you have achieved with it!\nAlso, if you don't like something or know an improvement, your contribution is very welcome!\n\nNote, that updates to the template can not be automatically pulled to child projects.\n\nThe template is developed and maintained by the data and analytics team of the city of Helsinki.\nThe template is published under the Apache-2.0 licence and open source utilization is encouraged!\n",
"_____no_output_____"
],
[
"## Contents\n\nThe core structure of the repository is the following:\n\n ## EDITABLE:\n data/ # Folder for storing data files. Ignored by git by default.\n |- raw_data/ # To store raw data files\n |- preprocessed_data/ # To store cleaned data\n results/ # Save results here. Ignored by git by default.\n |- notebooks/ # Save automatically executed notebooks here\n 00_data.ipynb # Extract, transfer, load data here & define related functions.\n 01_model.ipynb # Create and code test your ML model\n 02_loss.ipynb # Train and evaluate ML model, deploy or save for later use\n 03_workflow.ipynb # Define ML workflow and parameterization\n 04_api.ipynb # Define runtime API for using trained ML model\n project-requirements.in # Add here the Python packages you want to install\n update_install_dev_reqs.sh # run this script to install new python packages\n settings.ini # Project specific settings. Build instructions for lib and docs.\n Dockerfile # Define docker image build instructions\n .devcontainer # Codespaces / VSC dev environment instructions\n\n ## AUTOMATICALLY GENERATED: (Do not edit unless otherwise specified!)\n docs/ # Project documentation (html)\n [your_module]/ # Python module built from the notebooks (follow the installation instructions).\n README.md # The frontpage of your project, generated from index.ipynb\n requirements.txt # dev / default requirements. automatically generated by pip-tools\n min-requirements.txt # lighter requirements without dev tools. automatically generated by pip-tools\n\n ## STATIC NON-EDITABLE: (Edit only if you know what you're doing!)\n base-requirements.in # core tools that every project built based on the template always requires\n requirements.in # development tools + project spesific requirements\n LISENCE # lisence information\n MANIFEST.in # metadata for building python distributable\n setup.py # settings for the python module of your project\n CODE_OF_CONDUCT.md # code of conduct. Please review before contributing.\n ",
"_____no_output_____"
],
[
"## How to install\n\n> Note: if you are doing a project on personal data for the City of Helsinki, contact the data and analytics team of the city before proceeding any further!\n\n### 1. On your GitHub homepage\n\n0. (Create [GitHub account](https://github.com/) if you do not have one already. \n1. Sign into your GitHub homepage\n2. Go to [github.com/City-of-Helsinki/ml_project_template](https://github.com/City-of-Helsinki/ml_project_template) and click the green button that says 'Use this template'.\n3. Give your project a name. Do not use the dash symbol '-', but rather the underscore '_', because the name of the repo will become the name of your Python module.\n4. If you are creating a project for your organization, change the owner of the repo. From the drop down bar, select your organization GitHub account (e.g. City-of-Helsinki). You need to be included as a team member to the GitHub of the organization.\n5. Define your project publicity (you can change this later, but most likely you want to begin with a private repo).\n6. Click 'Create repository from template'\n\nThis will create a new repository for you copying everything from this template, but with clean commit history.\n\n### 2. Setting up your development environment\n\n#### a) Recommended: Codespaces\n\nIf your organization has [Codespaces](https://github.com/features/codespaces) enabled (requires GitHub Enterprise & Azure subscription), you are now ready to begin development. Just launch the repository in a codespace, and a dev container is automatically set up!\n\n#### b) Can't use Codespaces: Local installation with Docker\n\nYou can build a development environment locally with docker.\nThe recommended way is to use VSC in container development mode ([link to instructions](https://code.visualstudio.com/docs/remote/containers)).\n\n#### c) Can't use Docker: Local manual installation\n\nYou can also do your development 'the good old way':\n\n0. Create an SSH key and add it to your github profile ([instructions](https://docs.github.com/en/authentication/connecting-to-github-with-ssh))\n1. Configure your git user name and email adress if you haven't done it already: `git config --global user.name \"Firstname Lastname\" && git config --global user.email \"[email protected]\"`\n2. Clone your new repository: `git clone [email protected]:[repository_owner]/[your_repository]`\n3. Go inside the repository folder: `cd [your_repository]`\n4. Create and activate virtual environment of your choice. Remember to define the Python version to 3.8! (Instructions: [conda](https://docs.conda.io/projects/conda/en/latest/user-guide/tasks/manage-environments.html), [venv](https://packaging.python.org/en/latest/guides/installing-using-pip-and-virtual-environments/))\n5. Install pip-tools: `python -m pip install pip-tools`\n6. Install requirements: `pip-sync requirements.txt`\n7. Create an ipython kernel for running the notebooks: `python -m ipykernel install --user --name python38myenv`\n8. The default development enviroment contains basic Jupyter, and many IDEs have built-in support notebooks. If you wish, you can install JupyterLab by uncommenting it in `requirements.in` and re-running pip-sync. To launch JupyterLab, run `jupyter-lab --allow-root --config .devcontainer/jupyter-server-config.py`\n\n#### d) Can't connect to internet: Offline install with Docker\n\nSometimes you have to work in an environment that can not be connected to the internet, for example for privacy or cybersecurity reasons. In this case, first install the template and all packages that you assume you will require to an environment with internet, and build the docker image as in 2c). Then, save the docker image and transfer it to your offline environment following [these instructions](https://stackoverflow.com/questions/48125169/how-run-docker-images-without-connect-to-internet/48125632#48125632).\n\n### 3. Initializing your project\n\nFew last tweaks before you are good to go:\n\n1. Edit `LICENCE`, `Makefile`, `settings.ini`, `docs/_config.yml` and `docs/_data/topnav.yml` according to your project details. Don't worry - you can continue editing them in the future.\n2. Remove the folder `ml_project_template` with the command `git rm -r ml_project_template`. A new folder with the name of your repository will be created automatically when calling `nbdev_build_lib`.\n3. Recreate the python module: `nbdev_build_lib`. In the future, repeat this step every time you move between notebooks to ensure your python modules are up to date.\n4. Recreate the html doc pages & README: `nbdev_build_docs`. In the future, repeat this step every time you push code to ensure your documentation is up to date.\n5. Make initial commit: `git add . && git commit -m \"initialized repository from City-of-Helsinki/ml_project_template\"`\n6. Push changes `git push -u origin master`\n\nYou are now ready to begin your ML project development. Remember to track your changes with git!\n",
"_____no_output_____"
],
[
"## How to use\n\n1. Install this template as basis of your new project (see above).\n\n2. If you are not working inside a container, remember to activate your virtual environment every time you begin work: `conda activate [environment name]` with anaconda or `source [environment name]/bin/activate` with virtualenv.\n\n3. Develop your ML solution! (Follow the notebooks!)\n\n4. Save your notebooks and call `nbdev_build_lib` to build python modules of your notebooks - needed if you want to share code between notebooks or create a modules.\nThis will export all notebook cells with `# export` tag to corresponding .py files under the module (the folder inside your repository named after your repository).\nDo this every time you make changes to any exportable parts of the code.\n\n5. Save your notebooks and call `nbdev_build_docs` to create doc pages based on your notebooks (see below).\nThis will convert the notebooks into HTML files under `docs/` and update README based on the `index.ipynb`.\nIf you want to host your project pages on GitHub (like [the doc pages of this template](https://city-of-helsinki.github.io/ml_project_template/)), you will have to make your project public and enable github pages in repo > Settings > Pages : set Source to `docs/`.\nAlternatively you can build the pages locally with jekyll.\n",
"_____no_output_____"
],
[
"## Installing & updating project libraries\n\nPython has a rich and wide ecosystem of libraries to help with machine learning tasks among other things.\nPandas, Matplotlib, Scipy, PyTorch to name a few.\nIf base libraries in this template aren't sufficient you can add more with `pip install library`.\nHowever, `pip` command installs libraries into your local Python environment. \nTo achieve consistent reproducibility we need to gather information about requirements into project repository. \nNew libraries are added to **`project-requirements.in`** file. When you change this file remember to run:\n\n```bash\npip-compile --generate-hashes --allow-unsafe -o requirements.txt base-requirements.in requirements.in project-requirements.in\npip-compile --generate-hashes --allow-unsafe -o min-requirements.txt base-requirements.in project-requirements.in\n```\n\nThese update full requirements for development environments and lighter, more focused requirements for server usage.\n\nAfter requirements are updated you should run:\n\n```bash\npip-sync requirements.txt\n```\n\nThis way libraries you and other users will have the same Python environment.\n\n\n NOTE: run `./update_install_dev_reqs.sh` for short - it contains the three above pip commands for updating and installing the requirements!\n\n WARNING: if you don't update package names and versions next time you or anybody else tries to use this project in another environment its code might not work. Worse, it might *seem to* work, but does so incorrectly.\n",
"_____no_output_____"
],
[
"## Ethical aspects\n\nPlease involve ethical consideration in the documentation ML application.\n\nFor example:\n* Can you recognize ethical issues with your ML project?\n* Is there a risk for bias, discrimination, violation of privacy or conflict with the local or global laws?\n* Could your results or algorithms be misused for malicious acts?\n* Can data or model updates include bias in your model?\n* How have you tackled these issues in your implementation?\n* You most certainly make ethical choises in your code. Do you document & highlight them?\n* If you build an actual application, how can contribute if they notice an unresolved ethical issue?",
"_____no_output_____"
],
[
"## How to cite (optional)\n\nIf you are doing a research project, you can add bibtex and other citation templates here.\nYou can also get a doi for your code by adding it to a code archive,\nso your code can be cited directly! Most archives also provide repository badges.\n\nTo cite this work, use:\n\n @misc{sten2022helsinki,\n title = {Helsinki Machine Learning Project Template},\n author = {Nuutti A Sten and Jussi Arpalahti},\n year = {2022},\n howpublished = {City of Helsinki. Available at: \\url{https://github.com/City-of-Helsinki/ml_project_template}}\n }",
"_____no_output_____"
],
[
"## Contributing\n\nSee [CONTRIBUTING.md](https://github.com/City-of-Helsinki/ml_project_template/blob/master/CONTRIBUTING.md) on how to contribute to the development of this template.\n",
"_____no_output_____"
],
[
"## Copyright\n\nCopyright 2022 City-of-Helsinki. Licensed under the Apache License, Version 2.0 (the \"License\");\nyou may not use this project's files except in compliance with the License.\nA copy of the License is provided in the LICENSE file in this repository.\n\nThe Helsinki logo is a registered trademark, and may only be used by the city of Helsinki.\n \n NOTE: If you are using this template for other than city of Helsinki projects, remove the files `favicon.ico` and `company_logo.png` from `docs/assets/images/`.\n\n\n # to remove remove helsinki logo and favicon, run:\n git rm docs/assets/images/favicon.ico docs/assets/images/company_logo.png\n git commit -m \"removed Helsinki logo and favicon\"\n\nThis template was built using [nbdev](https://nbdev.fast.ai/) on top of the fast.ai [nbdev_template](https://github.com/fastai/nbdev_template).",
"_____no_output_____"
]
]
] | [
"code",
"markdown"
] | [
[
"code"
],
[
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown"
]
] |
cb006fa62ea21070dcd5f3a90a853a0190eb51a8 | 34,819 | ipynb | Jupyter Notebook | fine_tuning_roBERTsaikwan.ipynb | Sav-eng/roBERTsaikwan-model | 3f11af874efd402719c54833601ee49ee4205376 | [
"MIT"
] | null | null | null | fine_tuning_roBERTsaikwan.ipynb | Sav-eng/roBERTsaikwan-model | 3f11af874efd402719c54833601ee49ee4205376 | [
"MIT"
] | null | null | null | fine_tuning_roBERTsaikwan.ipynb | Sav-eng/roBERTsaikwan-model | 3f11af874efd402719c54833601ee49ee4205376 | [
"MIT"
] | null | null | null | 35.529592 | 1,612 | 0.501594 | [
[
[
"## Install the packages",
"_____no_output_____"
]
],
[
[
"!pip install -Uqq datasets pythainlp==2.2.4 transformers==4.4.0 tensorflow==2.4.0 tensorflow_text emoji seqeval sentencepiece fuzzywuzzy\n!npx degit --force https://github.com/vistec-AI/thai2transformers#dev",
"_____no_output_____"
],
[
"%load_ext autoreload\n%autoreload 2\n\nimport pythainlp, transformers\npythainlp.__version__, transformers.__version__ #fix pythainlp to stabilize word tokenization for metrics",
"_____no_output_____"
],
[
"import collections\nimport logging\nimport pprint\nimport re\nfrom tqdm.auto import tqdm\n\nimport numpy as np\nimport torch\n\n#datasets\nfrom datasets import (\n load_dataset, \n load_metric, \n concatenate_datasets,\n load_from_disk,\n)\n\n#transformers\nfrom transformers import (\n AutoConfig,\n AutoTokenizer,\n AutoModelForQuestionAnswering,\n TrainingArguments,\n Trainer,\n default_data_collator,\n)\n\n#thai2transformers\nimport thai2transformers\nfrom thai2transformers.metrics import (\n squad_newmm_metric,\n question_answering_metrics,\n)\nfrom thai2transformers.preprocess import (\n prepare_qa_train_features\n)\nfrom thai2transformers.tokenizers import (\n ThaiRobertaTokenizer,\n ThaiWordsNewmmTokenizer,\n ThaiWordsSyllableTokenizer,\n FakeSefrCutTokenizer,\n SEFR_SPLIT_TOKEN\n)\n\nfrom tqdm import tqdm",
"_____no_output_____"
],
[
"model_names = [\n 'wangchanberta-base-att-spm-uncased',\n 'xlm-roberta-base',\n 'bert-base-multilingual-cased',\n 'wangchanberta-base-wiki-newmm',\n 'wangchanberta-base-wiki-ssg',\n 'wangchanberta-base-wiki-sefr',\n 'wangchanberta-base-wiki-spm',\n]\n\ntokenizers = {\n 'wangchanberta-base-att-spm-uncased': AutoTokenizer,\n 'xlm-roberta-base': AutoTokenizer,\n 'bert-base-multilingual-cased': AutoTokenizer,\n 'wangchanberta-base-wiki-newmm': ThaiWordsNewmmTokenizer,\n 'wangchanberta-base-wiki-ssg': ThaiWordsSyllableTokenizer,\n 'wangchanberta-base-wiki-sefr': FakeSefrCutTokenizer,\n 'wangchanberta-base-wiki-spm': ThaiRobertaTokenizer,\n}\npublic_models = ['xlm-roberta-base', 'bert-base-multilingual-cased'] \n#@title Choose Pretrained Model\nmodel_name = \"wangchanberta-base-att-spm-uncased\" #@param [\"wangchanberta-base-att-spm-uncased\", \"xlm-roberta-base\", \"bert-base-multilingual-cased\", \"wangchanberta-base-wiki-newmm\", \"wangchanberta-base-wiki-syllable\", \"wangchanberta-base-wiki-sefr\", \"wangchanberta-base-wiki-spm\"]\n\n#create tokenizer\ntokenizer = tokenizers[model_name].from_pretrained(\n f'airesearch/{model_name}' if model_name not in public_models else f'{model_name}',\n revision='main',\n model_max_length=416,)",
"_____no_output_____"
]
],
[
[
"## Prepare function for calculate metrics",
"_____no_output_____"
]
],
[
[
"!pip install rouge",
"_____no_output_____"
],
[
"from rouge import Rouge \nrouge = Rouge()\ndef cal_rouge_score(hyps, refs, get_average_f1=True):\n '''\n argument: cands, refs [list of string], get_average_f1=True\n return dict of r1, r2, rl score\n if get_average_f1 == True return mean of rouge-1, rouge-2, rouge-L\n '''\n r1 = dict(); r1['precision'] = []; r1['recall'] = []; r1['f1'] = []\n r2 = dict(); r2['precision'] = []; r2['recall'] = []; r2['f1'] = []\n rl = dict(); rl['precision'] = []; rl['recall'] = []; rl['f1'] = []\n for hyp, ref in zip(hyps, refs):\n score = {}\n if(len(hyp)==0 or len(ref)==0):\n score = {\n 'rouge-1': {\n 'p': 0,\n 'r': 0,\n 'f': 0\n },\n 'rouge-2': {\n 'p': 0,\n 'r': 0,\n 'f': 0\n },\n 'rouge-l': {\n 'p': 0,\n 'r': 0,\n 'f': 0\n }\n }\n else: score = rouge.get_scores(hyp, ref)[0]\n r1['precision'].append(score['rouge-1']['p'])\n r1['recall'].append(score['rouge-1']['r'])\n r1['f1'].append(score['rouge-1']['f'])\n \n r2['precision'].append(score['rouge-2']['f'])\n r2['recall'].append(score['rouge-2']['f'])\n r2['f1'].append(score['rouge-2']['f'])\n\n rl['precision'].append(score['rouge-l']['f'])\n rl['recall'].append(score['rouge-l']['f'])\n rl['f1'].append(score['rouge-l']['f'])\n if(get_average_f1==True): return sum(r1['f1'])/len(r1['f1']), sum(r2['f1'])/len(r2['f1']), sum(rl['f1'])/len(rl['f1'])\n else: return r1, r2, rl",
"_____no_output_____"
],
[
"cands = ['test test test test test test bad']\nrefs = ['test test']\n\nr1, r2, rl = cal_rouge_score(cands, refs)\nprint(r1)\nprint(r2)\nprint(rl)",
"_____no_output_____"
]
],
[
[
"## Utility functions for calculate label in our use.",
"_____no_output_____"
]
],
[
[
"def tokenize_with_space(texts, tokenizer):\n output = []\n encoded_texts = tokenizer(texts, max_length=416, truncation=True)\n for text in encoded_texts['input_ids']:\n tokenized_text = \" \".join(tokenizer.convert_ids_to_tokens(text, skip_special_tokens=True))\n if(len(tokenized_text)==0):\n output.append(\"\")\n continue\n if(tokenized_text[0]==\"▁\"): \n tokenized_text = tokenized_text[1:]\n output.append(tokenized_text.strip())\n return output\n\n\ndef selection_start_end(paragraphs_raw, summaries_raw, tokenizer, length_sum_max = 10, metric='rouge-l'):\n \"\"\"\n Select the start position and end postion for each paragraph to make a summary and maximize the Rouge-L score\n Args: \n paragraphs [#number of paragraph, #number of word, #number of character] (must be tokenized with space and space change to '_')\n summaries [#number of summary, #number of word, #number of character] (must be tokenized with space and space change to '_')\n \"\"\"\n \n paragraphs = tokenize_with_space(paragraphs_raw, tokenizer)\n summaries = tokenize_with_space(summaries_raw, tokenizer)\n start_position = []\n end_position = []\n texts_all = []\n for paragraph_raw, summary in zip(paragraphs, summaries):\n paragraph = paragraph_raw.split(\" \")\n len_paragraph = len(paragraph)\n max_score = 0\n s = 0\n e = len_paragraph\n text = \"\"\n for length in range(1, length_sum_max):\n for start_pos in range(len_paragraph-length+1):\n t_summary = \" \".join(paragraph[start_pos:start_pos+length])\n try:\n r1, r2, score = cal_rouge_score([summary], [t_summary])\n if(max_score < score):\n max_score = score\n s = start_pos\n e = start_pos + length\n text = \"\".join(paragraph[s:e])\n except:\n pass\n start_position.append(s)\n end_position.append(e)\n texts_all.append(text)\n return start_position, end_position, texts_all\n",
"_____no_output_____"
],
[
"import collections as coll\n# stopwords = pkgutil.get_data(__package__, 'smart_common_words.txt')\n# stopwords = stopwords.decode('ascii').split('\\n')\n# stopwords = {key.strip(): 1 for key in stopwords}\n\ndef _get_ngrams_count(n, text):\n \"\"\"Calcualtes n-grams.\n Args:\n n: which n-grams to calculate\n text: An array of tokens\n Returns:\n A set of n-grams\n \"\"\"\n ngram_dic = coll.defaultdict(int)\n text_length = len(text)\n max_index_ngram_start = text_length - n\n for i in range(max_index_ngram_start + 1):\n ngram_dic[tuple(text[i:i + n])] += 1\n return ngram_dic\n\ndef _get_ngrams(n, text):\n \"\"\"Calcualtes n-grams.\n Args:\n n: which n-grams to calculate\n text: An array of tokens\n Returns:\n A set of n-grams\n \"\"\"\n ngram_set = set()\n text_length = len(text)\n max_index_ngram_start = text_length - n\n for i in range(max_index_ngram_start + 1):\n ngram_set.add(tuple(text[i:i + n]))\n return ngram_set\n\ndef _get_word_ngrams_list(n, text):\n \"\"\"Calcualtes n-grams.\n Args:\n n: which n-grams to calculate\n text: An array of tokens\n Returns:\n A set of n-grams\n \"\"\"\n text = sum(text, [])\n ngram_set = []\n text_length = len(text)\n max_index_ngram_start = text_length - n\n for i in range(max_index_ngram_start + 1):\n ngram_set.append(tuple(text[i:i + n]))\n return ngram_set\n\ndef _get_word_ngrams(n, sentences, do_count=False):\n \"\"\"Calculates word n-grams for multiple sentences.\n \"\"\"\n assert len(sentences) > 0\n assert n > 0\n\n # words = _split_into_words(sentences)\n\n words = sum(sentences, [])\n # words = [w for w in words if w not in stopwords]\n if do_count:\n return _get_ngrams_count(n, words)\n return _get_ngrams(n, words)\n \ndef cal_rouge(evaluated_ngrams, reference_ngrams):\n reference_count = len(reference_ngrams)\n evaluated_count = len(evaluated_ngrams)\n\n overlapping_ngrams = evaluated_ngrams.intersection(reference_ngrams)\n overlapping_count = len(overlapping_ngrams)\n\n if evaluated_count == 0:\n precision = 0.0\n else:\n precision = overlapping_count / evaluated_count\n\n if reference_count == 0:\n recall = 0.0\n else:\n recall = overlapping_count / reference_count\n\n f1_score = 2.0 * ((precision * recall) / (precision + recall + 1e-8))\n return {\"f\": f1_score, \"p\": precision, \"r\": recall}\n\ndef selection_start_end_r1_r2(doc, abstract, tokenizer, summary_size = 50):\n \"\"\"\n Select the start position and end postion for each paragraph to make a summary and maximize the Rouge-L score\n Args: \n paragraphs [#number of paragraph, #number of word, #number of character] (must be tokenized with space and space change to '_')\n summaries [#number of summary, #number of word, #number of character] (must be tokenized with space and space change to '_')\n \"\"\"\n \n max_rouge = 0.0\n tokenized_doc = tokenize_with_space([doc], tokenizer)[0].split(\" \")\n tokenized_abstract = tokenize_with_space([abstract], tokenizer)[0].split(\" \")\n # abstract = sum(abstract_sent_list, [])\n # abstract = ' '.join(abstract).split()\n # sents = [' '.join(s).split() for s in doc_sent_list]\n evaluated_1grams = _get_word_ngrams_list(1, [tokenized_doc])\n reference_1grams = _get_word_ngrams(1, [tokenized_abstract])\n evaluated_2grams = _get_word_ngrams_list(2, [tokenized_doc])\n reference_2grams = _get_word_ngrams(2, [tokenized_abstract])\n\n\n start = 0\n end = 0\n text = \"\"\n max_rouge = 0\n for s in range(1,summary_size):\n for i in range(len(tokenized_doc)-s+1):\n # if (i in selected):\n # continue\n c = range(i,i+s)\n candidates_1 = set(evaluated_1grams[i:i+s])\n # candidates_1 = set.union(*map(set, candidates_1))\n rouge = cal_rouge(candidates_1, reference_1grams)['f']\n if(s > 1):\n candidates_2 = set(evaluated_1grams[i:i+s-1])\n rouge += cal_rouge(candidates_2, reference_2grams)['f']\n if rouge > max_rouge:\n max_rouge = rouge\n start = i\n end = i+s\n text = \"\".join(tokenized_doc[i:i+s])\n\n return start, end, text",
"_____no_output_____"
]
],
[
[
"## Preprocess data",
"_____no_output_____"
]
],
[
[
"!gdown --id 1-8IU8qyry-yPXwQ7AXz0GHIgn19QKGZP\n!gdown --id 1-J0eqf4ig7cP8bMPRgSFUejshnBFTZoq\n!gdown --id 1-IIJFl4AGNr7rRax4YSQTTm7j12YJ0ya",
"_____no_output_____"
],
[
"import pandas as pd\ndf = pd.read_csv('thaisum.csv')\nval_df = pd.read_csv('validation_set.csv')\ntest_df = pd.read_csv('test_set.csv')\ndf = pd.concat([df, val_df, test_df], axis=0)",
"_____no_output_____"
],
[
"df = df.reset_index(drop=True)\ndf['body'][358868+11000]",
"_____no_output_____"
],
[
"def gold_summary(df, num_train, num_val, num_test):\n return df.iloc[num_train+num_val:num_train+num_val+num_test,:]['summary'].tolist()",
"_____no_output_____"
],
[
"def get_tokenized_df(df):\n df = df.reset_index(drop=True)\n res = pd.DataFrame(columns=['attention_mask', 'input_ids', 'start_positions', 'end_positions'])\n for i in tqdm(range(len(df))):\n sent1 = df['body'][i].lower()\n sent2 = df['summary'][i].lower()\n start, end, _ = selection_start_end_r1_r2(sent1, sent2, tokenizer)\n inp_ids = tokenizer(df['body'][i], max_length=416, truncation=True, padding='max_length')['input_ids']\n att_mask = tokenizer(df['body'][i], max_length=416, truncation=True, padding='max_length')['attention_mask']\n res = res.append({'attention_mask': att_mask, \n 'input_ids': inp_ids, \n 'start_positions': start, \n 'end_positions': end}, ignore_index=True)\n return res\n '''\n return {'input_ids': res['input_ids'].tolist(),\n 'attention_mask': res['attention_mask'].tolist(),\n 'start_positions': res['start_positions'].tolist(),\n 'end_positions': res['end_positions'].tolist()}\n '''\n\ndef get_tokenized_dict(df, num_train, num_val, num_test):\n train_df = df.iloc[:num_train, :]\n val_df = df.iloc[num_train:num_train+num_val, :]\n test_df = df.iloc[num_train+num_val:num_train+num_val+num_test, :]\n return {'train': get_tokenized_df(train_df),\n 'validation': get_tokenized_df(val_df),\n 'test': get_tokenized_df(test_df)}\n\ndef get_tokenized_dict_test_val(df, num_train, num_val, num_test):\n val_df = df.iloc[num_train:num_train+num_val, :]\n test_df = df.iloc[num_train+num_val:num_train+num_val+num_test, :]\n return {'validation': get_tokenized_df(val_df),\n 'test': get_tokenized_df(test_df)}\n\ndef get_tokenized_dict_test(df, num_train, num_val, num_test):\n test_df = df.iloc[num_train+num_val:num_train+num_val+num_test, :]\n return {'test': get_tokenized_df(test_df)}",
"_____no_output_____"
],
[
"tokenize_with_space([df['body'][369868]], tokenizer)",
"_____no_output_____"
]
],
[
[
"Usually tokenizing takes a lot of time, you can choose to tokenize only some part of data by uncommenting.",
"_____no_output_____"
]
],
[
[
"# %%time\ntokenized_datasets = get_tokenized_dict(df, 358868, 11000, 11000)\n# tokenized_datasets = get_tokenized_dict_test_val(df, 358868, 11000, 11000)\n# tokenized_datasets = get_tokenized_dict_test(df, 358868, 11000, 11000)",
"_____no_output_____"
],
[
"gold_summaries = gold_summary(df, 358868, 11000, 11000)",
"_____no_output_____"
]
],
[
[
"You can choose to save the data after preprocessing and load it.",
"_____no_output_____"
]
],
[
[
"# tokenized_datasets['train'].to_json('train.json', orient='records', lines=True)\n# tokenized_datasets['validation'].to_json('/content/drive/MyDrive/validation_true_set.json', orient='records', lines=True)\n# tokenized_datasets['test'].to_json('/content/drive/MyDrive/test_true_lower.json', orient='records', lines=True)\ntokenized_datasets = load_dataset('json', data_files={'train': '/content/drive/MyDrive/train.json', 'validation': '/content/drive/MyDrive/validation_true.json', 'test': '/content/drive/MyDrive/test_true.json'})",
"_____no_output_____"
],
[
"tokenized_datasets",
"_____no_output_____"
],
[
"#8 in datasets['validation'] points to both 8 and 9 in tokenized_datasets['validation'] due to overflowing tokens\ni = 8\nexample = tokenized_datasets['validation'][i]\ncombined_text = tokenizer.decode(example['input_ids'])\nanswer_with_token_idx = tokenizer.decode(example['input_ids'][example['start_positions']:example['end_positions']])\n\n#there are quite a few more \nlen(tokenized_datasets['validation']), answer_with_token_idx, combined_text",
"_____no_output_____"
]
],
[
[
"## Fine-tuning model",
"_____no_output_____"
]
],
[
[
"model = AutoModelForQuestionAnswering.from_pretrained(\n f'airesearch/{model_name}' if model_name not in public_models else f'{model_name}',\n revision='main',)",
"_____no_output_____"
],
[
"batch_size = 16\nlearning_rate = 4e-5\n\nargs = TrainingArguments(\n f\"finetune_thaiSum\",\n evaluation_strategy = \"epoch\",\n learning_rate=learning_rate,\n per_device_train_batch_size=batch_size,\n per_device_eval_batch_size=batch_size*2,\n num_train_epochs=6,\n warmup_ratio=0.15,\n weight_decay=0.01,\n fp16=True,\n save_total_limit=3,\n load_best_model_at_end=True,\n)\n\ndata_collator = default_data_collator",
"_____no_output_____"
],
[
"trainer = Trainer(\n model=model,\n args=args,\n train_dataset=tokenized_datasets[\"train\"],\n eval_dataset=tokenized_datasets[\"validation\"],\n data_collator=data_collator,\n tokenizer=tokenizer,\n)",
"_____no_output_____"
],
[
"trainer.train()",
"_____no_output_____"
],
[
"trainer.save_model(\"/content/drive/MyDrive/finetune_thaiSum4\")",
"_____no_output_____"
]
],
[
[
"## Postprocess and metrics(BERTscore since rouge we already import at the beginning)",
"_____no_output_____"
]
],
[
[
"def post_process_index(data, raw_predictions, tokenizer, n_best_size = 20, max_answer_length=50):\n all_start_logits, all_end_logits = raw_predictions\n predictions = []\n for start_logits, end_logits, example in zip(all_start_logits, all_end_logits, data):\n start_indexes = np.argsort(start_logits)[-1 : -n_best_size - 1 : -1].tolist()\n end_indexes = np.argsort(end_logits)[-1 : -n_best_size - 1 : -1].tolist()\n valid_answers = []\n for start_index in start_indexes:\n for end_index in end_indexes:\n # Don't consider answers with a length that is either < 0 or > max_answer_length.\n if end_index < start_index or end_index - start_index + 1 > max_answer_length:\n continue\n valid_answers.append(\n {\n \"score\": start_logits[start_index] + end_logits[end_index],\n \"text\": tokenizer.decode(example['input_ids'][start_index+1:end_index+1], skip_special_tokens=True)\n }\n )\n if len(valid_answers) > 0:\n best_answer = sorted(valid_answers, key=lambda x: x[\"score\"], reverse=True)[0]\n else:\n # In the very rare edge case we have not a single non-null prediction, we create a fake prediction to avoid failure.\n best_answer = {\"text\": \"\", \"score\": 0.0} \n predictions.append(best_answer[\"text\"])\n return predictions",
"_____no_output_____"
]
],
[
[
"### BERTScore",
"_____no_output_____"
]
],
[
[
"!pip install bert_score==0.3.7",
"_____no_output_____"
],
[
"from bert_score import score\nimport numpy as np\nimport gc",
"_____no_output_____"
],
[
"def cal_bert_score(cands, refs, get_average_f1=True):\n '''\n arguments: cands, refs\n return array of presicion, recall, f1, presicion_average, recall_average, f1_average\n if get_average == True return mean of BERTScore\n '''\n p, r, f1 = score(cands, refs, lang=\"others\", verbose=False)\n p = p.numpy()\n r = r.numpy()\n f1 = f1.numpy()\n if(get_average_f1==True): return f1.mean()\n else: return p, r, f1\n\ndef cal_batch_bert_score(cands, refs, get_average_f1=True, batch_size=8):\n f1_average = []\n for i in tqdm(range(0,len(cands),batch_size)):\n cand_batch = cands[i:i+batch_size]\n ref_batch = refs[i:i+batch_size]\n res = cal_bert_score(cand_batch, ref_batch)\n f1_average.append(res)\n gc.collect()\n print(f1_average)\n return sum(f1_average)/len(f1_average)",
"_____no_output_____"
],
[
"%%time\nrefs = ['เมื่อวันที่ 6 ม.ค.60 ที่ทำเนียบรัฐบาล นายวิษณุ เครืองาม รองนายกรัฐมนตรี กล่าวถึงกรณี ที่ นายสุรชัย เลี้ยงบุญเลิศชัย รองประธานสภานิติบัญญัติแห่งชาติ (สนช.) ออกมาระบุว่า การเลือกตั้งจะถูกเลื่อนออกไปถึงปี 2561 ว่า ขอให้ไปสอบถามกับ สนช. แต่เชื่อว่าคงไม่กล้าพูดอีก เพราะทำให้คนเข้าใจผิด ซึ่งที่ สนช.พูดเนื่องจากผูกกับกฎหมายของกรรมการร่างรัฐธรรมนูญ(กรธ.) ตนจึงไม่ขอวิพากษ์วิจารณ์ แต่รัฐบาลยืนยันว่ายังเดินตามโรดแม็ป ซึ่งโรดแม็ปมองได้สองแบบ คือ มีลำดับขั้นตอนและการกำหนดช่วงเวลา โดยเริ่มต้นจากการประกาศใช้รัฐธรรมนูญ แต่ขณะนี้รัฐธรรมนูญยังไม่ประกาศใช้ จึงยังเริ่มนับหนึ่งไม่ถูก จากนั้นเข้าสู่ขั้นตอนการร่างกฎหมายประกอบร่างรัฐธรรมนูญหรือกฎหมายลูก ภายใน 240 วัน ก่อนจะส่งกลับให้ สนช.พิจารณา ภายใน 2 เดือน\\xa0,นายวิษณุ กล่าวต่อว่า หากมีการแก้ไขก็จะมีการพิจารณาร่วมกับ กรธ.อีก 1 เดือน ก่อนนำขึ้นทูลเกล้าฯ ทรงลงพระปรมาภิไธย ภายใน 90 วัน และจะเข้าสู่การเลือกตั้งภายในระยะเวลา 5 เดือน ซึ่งทั้งหมดนี้คือโรดแม็ปที่ยังเป็นแบบเดิมอยู่ ส่วนเดิมที่กำหนดวันเลือกตั้งไว้ภายในปี 60 นั้น เพราะมาจากสมมติฐานของขั้นตอนเดิมทั้งหมด แต่เมื่อมีเหตุสวรรคตทุกอย่างจึงต้องเลื่อนออกไป ส่วนการพิจารณากฎหมายลูกทั้งหมด 4 ฉบับ ขณะนี้กรธ.พิจารณาแล้วเสร็จ 2 ฉบับ คือ พ.ร.ป.พรรคการเมือง และพ.ร.ป. คณะกรรมการการเลือกตั้ง แต่ พ.ร.ป.การเลือกตั้งควรจะพิจารณาได้เร็วกลับล่าช้า ดังนั้น กรธ.จะต้องออกชี้แจงถึงเหตุผลว่าทำไมพิจารณากฎหมายดังกล่าวล่าช้ากว่ากำหนด ส่งผลให้เกิดข้อสงสัยจนถึงทุกวันนี้ ส่วนกรณีที่ สนช. ระบุว่า มีกฎหมายเข้าสู่การพิจารณาของ สนช.เป็นจำนวนมาก ทำให้ส่งผลกระทบต่อโรดแม็ปนั้น รัฐบาลเคยบอกไว้แล้วว่าในช่วงนี้ของโรดแม็ปกฎหมายจะเยอะกว่าที่ผ่านมา ดังนั้น สนช.จะต้องบริหารจัดการกันเอง เพราะได้มีการเพิ่มสมาชิก สนช.ให้แล้ว.']\ncands = ['เมื่อวันที่ 6 ม.ค.60 ที่ทำเนียบรัฐบาล นายวิษณุ เครืองาม รองนายกรัฐมนตรี กล่าวถึงกรณี ที่ นายสุรชัย เลี้ยงบุญเลิศชัย รองประธานสภานิติบัญญัติแห่งชาติ (สนช.)']\nf1_average = cal_bert_score(cands, refs)",
"_____no_output_____"
],
[
"print(f1_average)",
"_____no_output_____"
]
],
[
[
"### Evaluate",
"_____no_output_____"
]
],
[
[
"def evaluate_rouge(cands, refs, tokenizer):\n cands_tokenized = tokenize_with_space(cands, tokenizer)\n refs_tokenized = tokenize_with_space(refs, tokenizer)\n r1, r2, rl = cal_rouge_score(refs_tokenized, cands_tokenized)\n return r1, r2, rl",
"_____no_output_____"
],
[
"raw_predictions = trainer.predict(tokenized_datasets['test'])",
"_____no_output_____"
],
[
"predictions = post_process_index(tokenized_datasets['test'], raw_predictions[0], tokenizer)",
"_____no_output_____"
],
[
"predictions[:3]",
"_____no_output_____"
],
[
"display(predictions[:3], gold_summaries[:3])",
"_____no_output_____"
],
[
"r1, r2, rl = evaluate_rouge(predictions, gold_summaries, tokenizer)\nprint(r1, r2, rl)",
"_____no_output_____"
],
[
"%%time\nBERTScore = cal_batch_bert_score(predictions, gold_summaries, tokenizer, batch_size=128)",
"_____no_output_____"
],
[
"print(BERTScore)",
"_____no_output_____"
]
]
] | [
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] | [
[
"markdown"
],
[
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
cb007e2eefcaa0caa2d13104d35bfb45b67f9234 | 51,433 | ipynb | Jupyter Notebook | samples/core/get_started/eager.ipynb | TomLisankie/tensorflow-models | a7c338bc2a601fd8edaca20be96e6267502e7708 | [
"Apache-2.0"
] | null | null | null | samples/core/get_started/eager.ipynb | TomLisankie/tensorflow-models | a7c338bc2a601fd8edaca20be96e6267502e7708 | [
"Apache-2.0"
] | null | null | null | samples/core/get_started/eager.ipynb | TomLisankie/tensorflow-models | a7c338bc2a601fd8edaca20be96e6267502e7708 | [
"Apache-2.0"
] | null | null | null | 40.402985 | 1,053 | 0.563432 | [
[
[
"##### Copyright 2018 The TensorFlow Authors.\n\nLicensed under the Apache License, Version 2.0 (the \"License\");",
"_____no_output_____"
]
],
[
[
"#@title Licensed under the Apache License, Version 2.0 (the \"License\");\n# you may not use this file except in compliance with the License.\n# You may obtain a copy of the License at\n#\n# https://www.apache.org/licenses/LICENSE-2.0\n#\n# Unless required by applicable law or agreed to in writing, software\n# distributed under the License is distributed on an \"AS IS\" BASIS,\n# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.\n# See the License for the specific language governing permissions and\n# limitations under the License.",
"_____no_output_____"
]
],
[
[
"# Get Started with Eager Execution\n\n\n<table align=\"left\"><td>\n<a target=\"_blank\" href=\"https://colab.sandbox.google.com/github/tensorflow/models/blob/master/samples/core/get_started/eager.ipynb\">\n <img src=\"https://www.tensorflow.org/images/colab_logo_32px.png\" />Run in Google Colab</a> \n</td><td>\n<a target=\"_blank\" href=\"https://github.com/tensorflow/models/blob/master/samples/core/get_started/eager.ipynb\"><img width=32px src=\"https://www.tensorflow.org/images/GitHub-Mark-32px.png\" />View source on Github</a></td></table>\n\n",
"_____no_output_____"
],
[
"This guide uses machine learning to *categorize* Iris flowers by species. It uses [TensorFlow](https://www.tensorflow.org)'s eager execution to:\n1. Build a model,\n2. Train this model on example data, and\n3. Use the model to make predictions about unknown data.\n\nMachine learning experience isn't required, but you'll need to read some Python code.\n\n## TensorFlow programming\n\nThere are many [TensorFlow APIs](https://www.tensorflow.org/api_docs/python/) available, but start with these high-level TensorFlow concepts:\n\n* Enable an [eager execution](https://www.tensorflow.org/programmers_guide/eager) development environment,\n* Import data with the [Datasets API](https://www.tensorflow.org/programmers_guide/datasets),\n* Build models and layers with TensorFlow's [Keras API](https://keras.io/getting-started/sequential-model-guide/).\n\nThis tutorial is structured like many TensorFlow programs:\n\n1. Import and parse the data sets.\n2. Select the type of model.\n3. Train the model.\n4. Evaluate the model's effectiveness.\n5. Use the trained model to make predictions.\n\nFor more TensorFlow examples, see the [Get Started](https://www.tensorflow.org/get_started/) and [Tutorials](https://www.tensorflow.org/tutorials/) sections. To learn machine learning basics, consider taking the [Machine Learning Crash Course](https://developers.google.com/machine-learning/crash-course/).\n\n## Run the notebook\n\nThis tutorial is available as an interactive [Colab notebook](https://colab.research.google.com) that can execute and modify Python code directly in the browser. The notebook handles setup and dependencies while you \"play\" cells to run the code blocks. This is a fun way to explore the program and test ideas.\n\nIf you are unfamiliar with Python notebook environments, there are a couple of things to keep in mind:\n\n1. Executing code requires connecting to a runtime environment. In the Colab notebook menu, select *Runtime > Connect to runtime...*\n2. Notebook cells are arranged sequentially to gradually build the program. Typically, later code cells depend on prior code cells, though you can always rerun a code block. To execute the entire notebook in order, select *Runtime > Run all*. To rerun a code cell, select the cell and click the *play icon* on the left.",
"_____no_output_____"
],
[
"## Setup program",
"_____no_output_____"
],
[
"### Install the latest version of TensorFlow\n\nThis tutorial uses eager execution, which is available in [TensorFlow 1.8](https://www.tensorflow.org/install/). (You may need to restart the runtime after upgrading.)",
"_____no_output_____"
]
],
[
[
"!pip install --upgrade tensorflow",
"_____no_output_____"
]
],
[
[
"### Configure imports and eager execution\n\nImport the required Python modules—including TensorFlow—and enable eager execution for this program. Eager execution makes TensorFlow evaluate operations immediately, returning concrete values instead of creating a [computational graph](https://www.tensorflow.org/programmers_guide/graphs) that is executed later. If you are used to a REPL or the `python` interactive console, this feels familiar.\n\nOnce eager execution is enabled, it *cannot* be disabled within the same program. See the [eager execution guide](https://www.tensorflow.org/programmers_guide/eager) for more details.",
"_____no_output_____"
]
],
[
[
"from __future__ import absolute_import, division, print_function\n\nimport os\nimport matplotlib.pyplot as plt\n\nimport tensorflow as tf\nimport tensorflow.contrib.eager as tfe\n\ntf.enable_eager_execution()\n\nprint(\"TensorFlow version: {}\".format(tf.VERSION))\nprint(\"Eager execution: {}\".format(tf.executing_eagerly()))",
"_____no_output_____"
]
],
[
[
"## The Iris classification problem\n\nImagine you are a botanist seeking an automated way to categorize each Iris flower you find. Machine learning provides many algorithms to statistically classify flowers. For instance, a sophisticated machine learning program could classify flowers based on photographs. Our ambitions are more modest—we're going to classify Iris flowers based on the length and width measurements of their [sepals](https://en.wikipedia.org/wiki/Sepal) and [petals](https://en.wikipedia.org/wiki/Petal).\n\nThe Iris genus entails about 300 species, but our program will only classify the following three:\n\n* Iris setosa\n* Iris virginica\n* Iris versicolor\n\n<table>\n <tr><td>\n <img src=\"https://www.tensorflow.org/images/iris_three_species.jpg\"\n alt=\"Petal geometry compared for three iris species: Iris setosa, Iris virginica, and Iris versicolor\">\n </td></tr>\n <tr><td align=\"center\">\n <b>Figure 1.</b> <a href=\"https://commons.wikimedia.org/w/index.php?curid=170298\">Iris setosa</a> (by <a href=\"https://commons.wikimedia.org/wiki/User:Radomil\">Radomil</a>, CC BY-SA 3.0), <a href=\"https://commons.wikimedia.org/w/index.php?curid=248095\">Iris versicolor</a>, (by <a href=\"https://commons.wikimedia.org/wiki/User:Dlanglois\">Dlanglois</a>, CC BY-SA 3.0), and <a href=\"https://www.flickr.com/photos/33397993@N05/3352169862\">Iris virginica</a> (by <a href=\"https://www.flickr.com/photos/33397993@N05\">Frank Mayfield</a>, CC BY-SA 2.0).<br/> \n </td></tr>\n</table>\n\nFortunately, someone has already created a [data set of 120 Iris flowers](https://en.wikipedia.org/wiki/Iris_flower_data_set) with the sepal and petal measurements. This is a classic dataset that is popular for beginner machine learning classification problems.",
"_____no_output_____"
],
[
"## Import and parse the training dataset\n\nDownload the dataset file and convert it to a structure that can be used by this Python program.\n\n### Download the dataset\n\nDownload the training dataset file using the [tf.keras.utils.get_file](https://www.tensorflow.org/api_docs/python/tf/keras/utils/get_file) function. This returns the file path of the downloaded file.",
"_____no_output_____"
]
],
[
[
"train_dataset_url = \"http://download.tensorflow.org/data/iris_training.csv\"\n\ntrain_dataset_fp = tf.keras.utils.get_file(fname=os.path.basename(train_dataset_url),\n origin=train_dataset_url)\n\nprint(\"Local copy of the dataset file: {}\".format(train_dataset_fp))",
"_____no_output_____"
]
],
[
[
"### Inspect the data\n\nThis dataset, `iris_training.csv`, is a plain text file that stores tabular data formatted as comma-separated values (CSV). Use the `head -n5` command to take a peak at the first five entries:",
"_____no_output_____"
]
],
[
[
"!head -n5 {train_dataset_fp}",
"_____no_output_____"
]
],
[
[
"From this view of the dataset, notice the following:\n\n1. The first line is a header containing information about the dataset:\n * There are 120 total examples. Each example has four features and one of three possible label names. \n2. Subsequent rows are data records, one *[example](https://developers.google.com/machine-learning/glossary/#example)* per line, where:\n * The first four fields are *[features](https://developers.google.com/machine-learning/glossary/#feature)*: these are characteristics of an example. Here, the fields hold float numbers representing flower measurements.\n * The last column is the *[label](https://developers.google.com/machine-learning/glossary/#label)*: this is the value we want to predict. For this dataset, it's an integer value of 0, 1, or 2 that corresponds to a flower name.\n\nLet's write that out in code:",
"_____no_output_____"
]
],
[
[
"# column order in CSV file\ncolumn_names = ['sepal_length', 'sepal_width', 'petal_length', 'petal_width', 'species']\n\nfeature_names = column_names[:-1]\nlabel_name = column_names[-1]\n\nprint(\"Features: {}\".format(feature_names))\nprint(\"Label: {}\".format(label_name))",
"_____no_output_____"
]
],
[
[
"Each label is associated with string name (for example, \"setosa\"), but machine learning typically relies on numeric values. The label numbers are mapped to a named representation, such as:\n\n* `0`: Iris setosa\n* `1`: Iris versicolor\n* `2`: Iris virginica\n\nFor more information about features and labels, see the [ML Terminology section of the Machine Learning Crash Course](https://developers.google.com/machine-learning/crash-course/framing/ml-terminology).",
"_____no_output_____"
]
],
[
[
"class_names = ['Iris setosa', 'Iris versicolor', 'Iris virginica']",
"_____no_output_____"
]
],
[
[
"### Create a `tf.data.Dataset`\n\nTensorFlow's [Dataset API](https://www.tensorflow.org/programmers_guide/datasets) handles many common cases for loading data into a model. This is a high-level API for reading data and transforming it into a form used for training. See the [Datasets Quick Start guide](https://www.tensorflow.org/get_started/datasets_quickstart) for more information.\n\n\nSince the dataset is a CSV-formatted text file, use the the [make_csv_dataset](https://www.tensorflow.org/api_docs/python/tf/contrib/data/make_csv_dataset) function to parse the data into a suitable format. Since this function generates data for training models, the default behavior is to shuffle the data (`shuffle=True, shuffle_buffer_size=10000`), and repeat the dataset forever (`num_epochs=None`). We also set the [batch_size](https://developers.google.com/machine-learning/glossary/#batch_size) parameter.",
"_____no_output_____"
]
],
[
[
"batch_size = 32\n\ntrain_dataset = tf.contrib.data.make_csv_dataset(\n train_dataset_fp,\n batch_size, \n column_names=column_names,\n label_name=label_name,\n num_epochs=1)",
"_____no_output_____"
]
],
[
[
"The `make_csv_dataset` function returns a `tf.data.Dataset` of `(features, label)` pairs, where `features` is a dictionary: `{'feature_name': value}`\n\nWith eager execution enabled, these `Dataset` objects are iterable. Let's look at a batch of features:",
"_____no_output_____"
]
],
[
[
"features, labels = next(iter(train_dataset))\n\nfeatures",
"_____no_output_____"
]
],
[
[
"Notice that like-features are grouped together, or *batched*. Each example row's fields are appended to the corresponding feature array. Change the `batch_size` to set the number of examples stored in these feature arrays.\n\nYou can start to see some clusters by plotting a few features from the batch:",
"_____no_output_____"
]
],
[
[
"plt.scatter(features['petal_length'],\n features['sepal_length'],\n c=labels,\n cmap='viridis')\n\nplt.xlabel(\"Petal length\")\nplt.ylabel(\"Sepal length\");",
"_____no_output_____"
]
],
[
[
"To simplify the model building step, create a function to repackage the features dictionary into a single array with shape: `(batch_size, num_features)`.\n\nThis function uses the [tf.stack](https://www.tensorflow.org/api_docs/python/tf/stack) method which takes values from a list of tensors and creates a combined tensor at the specified dimension.",
"_____no_output_____"
]
],
[
[
"def pack_features_vector(features, labels):\n \"\"\"Pack the features into a single array.\"\"\"\n features = tf.stack(list(features.values()), axis=1)\n return features, labels",
"_____no_output_____"
]
],
[
[
"Then use the [tf.data.Dataset.map](https://www.tensorflow.org/api_docs/python/tf/data/dataset/map) method to pack the `features` of each `(features,label)` pair into the training dataset:",
"_____no_output_____"
]
],
[
[
"train_dataset = train_dataset.map(pack_features_vector)",
"_____no_output_____"
]
],
[
[
"The features element of the `Dataset` are now arrays with shape `(batch_size, num_features)`. Let's look at the first few examples:",
"_____no_output_____"
]
],
[
[
"features, labels = next(iter(train_dataset))\n\nprint(features[:5])",
"_____no_output_____"
]
],
[
[
"## Select the type of model\n\n### Why model?\n\nA *[model](https://developers.google.com/machine-learning/crash-course/glossary#model)* is the relationship between features and the label. For the Iris classification problem, the model defines the relationship between the sepal and petal measurements and the predicted Iris species. Some simple models can be described with a few lines of algebra, but complex machine learning models have a large number of parameters that are difficult to summarize.\n\nCould you determine the relationship between the four features and the Iris species *without* using machine learning? That is, could you use traditional programming techniques (for example, a lot of conditional statements) to create a model? Perhaps—if you analyzed the dataset long enough to determine the relationships between petal and sepal measurements to a particular species. And this becomes difficult—maybe impossible—on more complicated datasets. A good machine learning approach *determines the model for you*. If you feed enough representative examples into the right machine learning model type, the program will figure out the relationships for you.\n\n### Select the model\n\nWe need to select the kind of model to train. There are many types of models and picking a good one takes experience. This tutorial uses a neural network to solve the Iris classification problem. *[Neural networks](https://developers.google.com/machine-learning/glossary/#neural_network)* can find complex relationships between features and the label. It is a highly-structured graph, organized into one or more *[hidden layers](https://developers.google.com/machine-learning/glossary/#hidden_layer)*. Each hidden layer consists of one or more *[neurons](https://developers.google.com/machine-learning/glossary/#neuron)*. There are several categories of neural networks and this program uses a dense, or *[fully-connected neural network](https://developers.google.com/machine-learning/glossary/#fully_connected_layer)*: the neurons in one layer receive input connections from *every* neuron in the previous layer. For example, Figure 2 illustrates a dense neural network consisting of an input layer, two hidden layers, and an output layer:\n\n<table>\n <tr><td>\n <img src=\"https://www.tensorflow.org/images/custom_estimators/full_network.png\"\n alt=\"A diagram of the network architecture: Inputs, 2 hidden layers, and outputs\">\n </td></tr>\n <tr><td align=\"center\">\n <b>Figure 2.</b> A neural network with features, hidden layers, and predictions.<br/> \n </td></tr>\n</table>\n\nWhen the model from Figure 2 is trained and fed an unlabeled example, it yields three predictions: the likelihood that this flower is the given Iris species. This prediction is called *[inference](https://developers.google.com/machine-learning/crash-course/glossary#inference)*. For this example, the sum of the output predictions is 1.0. In Figure 2, this prediction breaks down as: `0.03` for *Iris setosa*, `0.95` for *Iris versicolor*, and `0.02` for *Iris virginica*. This means that the model predicts—with 95% probability—that an unlabeled example flower is an *Iris versicolor*.",
"_____no_output_____"
],
[
"### Create a model using Keras\n\nThe TensorFlow [tf.keras](https://www.tensorflow.org/api_docs/python/tf/keras) API is the preferred way to create models and layers. This makes it easy to build models and experiment while Keras handles the complexity of connecting everything together.\n\nThe [tf.keras.Sequential](https://www.tensorflow.org/api_docs/python/tf/keras/Sequential) model is a linear stack of layers. Its constructor takes a list of layer instances, in this case, two [Dense](https://www.tensorflow.org/api_docs/python/tf/keras/layers/Dense) layers with 10 nodes each, and an output layer with 3 nodes representing our label predictions. The first layer's `input_shape` parameter corresponds to the number of features from the dataset, and is required.",
"_____no_output_____"
]
],
[
[
"model = tf.keras.Sequential([\n tf.keras.layers.Dense(10, activation=\"relu\", input_shape=(4,)), # input shape required\n tf.keras.layers.Dense(10, activation=\"relu\"),\n tf.keras.layers.Dense(3)\n])",
"_____no_output_____"
]
],
[
[
"The *[activation function](https://developers.google.com/machine-learning/crash-course/glossary#activation_function)* determines the output shape of each node in the layer. These non-linearities are important—without them the model would be equivalent to a single layer. There are many [available activations](https://www.tensorflow.org/api_docs/python/tf/keras/activations), but [ReLU](https://developers.google.com/machine-learning/crash-course/glossary#ReLU) is common for hidden layers.\n\nThe ideal number of hidden layers and neurons depends on the problem and the dataset. Like many aspects of machine learning, picking the best shape of the neural network requires a mixture of knowledge and experimentation. As a rule of thumb, increasing the number of hidden layers and neurons typically creates a more powerful model, which requires more data to train effectively.",
"_____no_output_____"
],
[
"### Using the model\n\nLet's have a quick look at what this model does to a batch of features:",
"_____no_output_____"
]
],
[
[
"predictions = model(features)\npredictions[:5]",
"_____no_output_____"
]
],
[
[
"Here, each example returns a [logit](https://developers.google.com/machine-learning/crash-course/glossary#logit) for each class. \n\nTo convert these logits to a probability for each class, use the [softmax](https://developers.google.com/machine-learning/crash-course/glossary#softmax) function:",
"_____no_output_____"
]
],
[
[
"tf.nn.softmax(predictions[:5])",
"_____no_output_____"
]
],
[
[
"Taking the `tf.argmax` across classes gives us the predicted class index. But, the model hasn't been trained yet, so these aren't good predictions.",
"_____no_output_____"
]
],
[
[
"print(\"Prediction: {}\".format(tf.argmax(predictions, axis=1)))\nprint(\" Labels: {}\".format(labels))",
"_____no_output_____"
]
],
[
[
"## Train the model\n\n*[Training](https://developers.google.com/machine-learning/crash-course/glossary#training)* is the stage of machine learning when the model is gradually optimized, or the model *learns* the dataset. The goal is to learn enough about the structure of the training dataset to make predictions about unseen data. If you learn *too much* about the training dataset, then the predictions only work for the data it has seen and will not be generalizable. This problem is called *[overfitting](https://developers.google.com/machine-learning/crash-course/glossary#overfitting)*—it's like memorizing the answers instead of understanding how to solve a problem.\n\nThe Iris classification problem is an example of *[supervised machine learning](https://developers.google.com/machine-learning/glossary/#supervised_machine_learning)*: the model is trained from examples that contain labels. In *[unsupervised machine learning](https://developers.google.com/machine-learning/glossary/#unsupervised_machine_learning)*, the examples don't contain labels. Instead, the model typically finds patterns among the features.",
"_____no_output_____"
],
[
"### Define the loss and gradient function\n\nBoth training and evaluation stages need to calculate the model's *[loss](https://developers.google.com/machine-learning/crash-course/glossary#loss)*. This measures how off a model's predictions are from the desired label, in other words, how bad the model is performing. We want to minimize, or optimize, this value.\n\nOur model will calculate its loss using the [tf.keras.losses.categorical_crossentropy](https://www.tensorflow.org/api_docs/python/tf/losses/sparse_softmax_cross_entropy) function which takes the model's class probability predictions and the desired label, and returns the average loss across the examples.",
"_____no_output_____"
]
],
[
[
"def loss(model, x, y):\n y_ = model(x)\n return tf.losses.sparse_softmax_cross_entropy(labels=y, logits=y_)\n\n\nl = loss(model, features, labels)\nprint(\"Loss test: {}\".format(l))",
"_____no_output_____"
]
],
[
[
"Use the [tf.GradientTape](https://www.tensorflow.org/api_docs/python/tf/GradientTape) context to calculate the *[gradients](https://developers.google.com/machine-learning/crash-course/glossary#gradient)* used to optimize our model. For more examples of this, see the [eager execution guide](https://www.tensorflow.org/programmers_guide/eager).",
"_____no_output_____"
]
],
[
[
"def grad(model, inputs, targets):\n with tf.GradientTape() as tape:\n loss_value = loss(model, inputs, targets)\n return loss_value, tape.gradient(loss_value, model.trainable_variables)",
"_____no_output_____"
]
],
[
[
"### Create an optimizer\n\nAn *[optimizer](https://developers.google.com/machine-learning/crash-course/glossary#optimizer)* applies the computed gradients to the model's variables to minimize the `loss` function. You can think of the loss function as a curved surface (see Figure 3) and we want to find its lowest point by walking around. The gradients point in the direction of steepest ascent—so we'll travel the opposite way and move down the hill. By iteratively calculating the loss and gradient for each batch, we'll adjust the model during training. Gradually, the model will find the best combination of weights and bias to minimize loss. And the lower the loss, the better the model's predictions.\n\n<table>\n <tr><td>\n <img src=\"https://cs231n.github.io/assets/nn3/opt1.gif\" width=\"70%\"\n alt=\"Optimization algorthims visualized over time in 3D space.\">\n </td></tr>\n <tr><td align=\"center\">\n <b>Figure 3.</b> Optimization algorithms visualized over time in 3D space. (Source: <a href=\"http://cs231n.github.io/neural-networks-3/\">Stanford class CS231n</a>, MIT License)<br/> \n </td></tr>\n</table>\n\nTensorFlow has many [optimization algorithms](https://www.tensorflow.org/api_guides/python/train) available for training. This model uses the [tf.train.GradientDescentOptimizer](https://www.tensorflow.org/api_docs/python/tf/train/GradientDescentOptimizer) that implements the *[stochastic gradient descent](https://developers.google.com/machine-learning/crash-course/glossary#gradient_descent)* (SGD) algorithm. The `learning_rate` sets the step size to take for each iteration down the hill. This is a *hyperparameter* that you'll commonly adjust to achieve better results.",
"_____no_output_____"
],
[
"Let's setup the optimizer and the `global_step` counter:",
"_____no_output_____"
]
],
[
[
"optimizer = tf.train.GradientDescentOptimizer(learning_rate=0.01)\n\nglobal_step = tf.train.get_or_create_global_step()",
"_____no_output_____"
]
],
[
[
"We'll use this to calculate a single optimization step:",
"_____no_output_____"
]
],
[
[
"loss_value, grads = grad(model, features, labels)\n\nprint(\"Step: {}, Initial Loss: {}\".format(global_step.numpy(),\n loss_value.numpy()))\n\noptimizer.apply_gradients(zip(grads, model.variables), global_step)\n\nprint(\"Step: {}, Loss: {}\".format(global_step.numpy(),\n loss(model, features, labels).numpy()))",
"_____no_output_____"
]
],
[
[
"### Training loop\n\nWith all the pieces in place, the model is ready for training! A training loop feeds the dataset examples into the model to help it make better predictions. The following code block sets up these training steps:\n\n1. Iterate each *epoch*. An epoch is one pass through the dataset.\n2. Within an epoch, iterate over each example in the training `Dataset` grabbing its *features* (`x`) and *label* (`y`).\n3. Using the example's features, make a prediction and compare it with the label. Measure the inaccuracy of the prediction and use that to calculate the model's loss and gradients.\n4. Use an `optimizer` to update the model's variables.\n5. Keep track of some stats for visualization.\n6. Repeat for each epoch.\n\nThe `num_epochs` variable is the amount of times to loop over the dataset collection. Counter-intuitively, training a model longer does not guarantee a better model. `num_epochs` is a *[hyperparameter](https://developers.google.com/machine-learning/glossary/#hyperparameter)* that you can tune. Choosing the right number usually requires both experience and experimentation.",
"_____no_output_____"
]
],
[
[
"## Note: Rerunning this cell uses the same model variables\n\n# keep results for plotting\ntrain_loss_results = []\ntrain_accuracy_results = []\n\nnum_epochs = 201\n\nfor epoch in range(num_epochs):\n epoch_loss_avg = tfe.metrics.Mean()\n epoch_accuracy = tfe.metrics.Accuracy()\n\n # Training loop - using batches of 32\n for x, y in train_dataset:\n # Optimize the model\n loss_value, grads = grad(model, x, y)\n optimizer.apply_gradients(zip(grads, model.variables),\n global_step)\n\n # Track progress\n epoch_loss_avg(loss_value) # add current batch loss\n # compare predicted label to actual label\n epoch_accuracy(tf.argmax(model(x), axis=1, output_type=tf.int32), y)\n\n # end epoch\n train_loss_results.append(epoch_loss_avg.result())\n train_accuracy_results.append(epoch_accuracy.result())\n \n if epoch % 50 == 0:\n print(\"Epoch {:03d}: Loss: {:.3f}, Accuracy: {:.3%}\".format(epoch,\n epoch_loss_avg.result(),\n epoch_accuracy.result()))",
"_____no_output_____"
]
],
[
[
"### Visualize the loss function over time",
"_____no_output_____"
],
[
"While it's helpful to print out the model's training progress, it's often *more* helpful to see this progress. [TensorBoard](https://www.tensorflow.org/programmers_guide/summaries_and_tensorboard) is a nice visualization tool that is packaged with TensorFlow, but we can create basic charts using the `matplotlib` module.\n\nInterpreting these charts takes some experience, but you really want to see the *loss* go down and the *accuracy* go up.",
"_____no_output_____"
]
],
[
[
"fig, axes = plt.subplots(2, sharex=True, figsize=(12, 8))\nfig.suptitle('Training Metrics')\n\naxes[0].set_ylabel(\"Loss\", fontsize=14)\naxes[0].plot(train_loss_results)\n\naxes[1].set_ylabel(\"Accuracy\", fontsize=14)\naxes[1].set_xlabel(\"Epoch\", fontsize=14)\naxes[1].plot(train_accuracy_results);",
"_____no_output_____"
]
],
[
[
"## Evaluate the model's effectiveness\n\nNow that the model is trained, we can get some statistics on its performance.\n\n*Evaluating* means determining how effectively the model makes predictions. To determine the model's effectiveness at Iris classification, pass some sepal and petal measurements to the model and ask the model to predict what Iris species they represent. Then compare the model's prediction against the actual label. For example, a model that picked the correct species on half the input examples has an *[accuracy](https://developers.google.com/machine-learning/glossary/#accuracy)* of `0.5`. Figure 4 shows a slightly more effective model, getting 4 out of 5 predictions correct at 80% accuracy:\n\n<table cellpadding=\"8\" border=\"0\">\n <colgroup>\n <col span=\"4\" >\n <col span=\"1\" bgcolor=\"lightblue\">\n <col span=\"1\" bgcolor=\"lightgreen\">\n </colgroup>\n <tr bgcolor=\"lightgray\">\n <th colspan=\"4\">Example features</th>\n <th colspan=\"1\">Label</th>\n <th colspan=\"1\" >Model prediction</th>\n </tr>\n <tr>\n <td>5.9</td><td>3.0</td><td>4.3</td><td>1.5</td><td align=\"center\">1</td><td align=\"center\">1</td>\n </tr>\n <tr>\n <td>6.9</td><td>3.1</td><td>5.4</td><td>2.1</td><td align=\"center\">2</td><td align=\"center\">2</td>\n </tr>\n <tr>\n <td>5.1</td><td>3.3</td><td>1.7</td><td>0.5</td><td align=\"center\">0</td><td align=\"center\">0</td>\n </tr>\n <tr>\n <td>6.0</td> <td>3.4</td> <td>4.5</td> <td>1.6</td> <td align=\"center\">1</td><td align=\"center\" bgcolor=\"red\">2</td>\n </tr>\n <tr>\n <td>5.5</td><td>2.5</td><td>4.0</td><td>1.3</td><td align=\"center\">1</td><td align=\"center\">1</td>\n </tr>\n <tr><td align=\"center\" colspan=\"6\">\n <b>Figure 4.</b> An Iris classifier that is 80% accurate.<br/> \n </td></tr>\n</table>",
"_____no_output_____"
],
[
"### Setup the test dataset\n\nEvaluating the model is similar to training the model. The biggest difference is the examples come from a separate *[test set](https://developers.google.com/machine-learning/crash-course/glossary#test_set)* rather than the training set. To fairly assess a model's effectiveness, the examples used to evaluate a model must be different from the examples used to train the model.\n\nThe setup for the test `Dataset` is similar to the setup for training `Dataset`. Download the CSV text file and parse that values, then give it a little shuffle:",
"_____no_output_____"
]
],
[
[
"test_url = \"http://download.tensorflow.org/data/iris_test.csv\"\n\ntest_fp = tf.keras.utils.get_file(fname=os.path.basename(test_url),\n origin=test_url)",
"_____no_output_____"
],
[
"test_dataset = tf.contrib.data.make_csv_dataset(\n train_dataset_fp,\n batch_size, \n column_names=column_names,\n label_name='species',\n num_epochs=1,\n shuffle=False)\n\ntest_dataset = test_dataset.map(pack_features_vector)",
"_____no_output_____"
]
],
[
[
"### Evaluate the model on the test dataset\n\nUnlike the training stage, the model only evaluates a single [epoch](https://developers.google.com/machine-learning/glossary/#epoch) of the test data. In the following code cell, we iterate over each example in the test set and compare the model's prediction against the actual label. This is used to measure the model's accuracy across the entire test set.",
"_____no_output_____"
]
],
[
[
"test_accuracy = tfe.metrics.Accuracy()\n\nfor (x, y) in test_dataset:\n logits = model(x)\n prediction = tf.argmax(logits, axis=1, output_type=tf.int32)\n test_accuracy(prediction, y)\n\nprint(\"Test set accuracy: {:.3%}\".format(test_accuracy.result()))",
"_____no_output_____"
]
],
[
[
"We can see on the last batch, for example, the model is usually correct:",
"_____no_output_____"
]
],
[
[
"tf.stack([y,prediction],axis=1)",
"_____no_output_____"
]
],
[
[
"## Use the trained model to make predictions\n\nWe've trained a model and \"proven\" that it's good—but not perfect—at classifying Iris species. Now let's use the trained model to make some predictions on [unlabeled examples](https://developers.google.com/machine-learning/glossary/#unlabeled_example); that is, on examples that contain features but not a label.\n\nIn real-life, the unlabeled examples could come from lots of different sources including apps, CSV files, and data feeds. For now, we're going to manually provide three unlabeled examples to predict their labels. Recall, the label numbers are mapped to a named representation as:\n\n* `0`: Iris setosa\n* `1`: Iris versicolor\n* `2`: Iris virginica",
"_____no_output_____"
]
],
[
[
"predict_dataset = tf.convert_to_tensor([\n [5.1, 3.3, 1.7, 0.5,],\n [5.9, 3.0, 4.2, 1.5,],\n [6.9, 3.1, 5.4, 2.1]\n])\n\npredictions = model(predict_dataset)\n\nfor i, logits in enumerate(predictions):\n class_idx = tf.argmax(logits).numpy()\n p = tf.nn.softmax(logits)[class_idx]\n name = class_names[class_idx]\n print(\"Example {} prediction: {} ({:4.1f}%)\".format(i, name, 100*p))",
"_____no_output_____"
]
],
[
[
"These predictions look good!\n\nTo dig deeper into machine learning models, take a look at the TensorFlow [Programmer's Guide](https://www.tensorflow.org/programmers_guide/) and check out the [community](https://www.tensorflow.org/community/).",
"_____no_output_____"
]
]
] | [
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown"
] | [
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
]
] |
cb008c55f82c5c2b853f46b878b0f23bf64fd148 | 6,436 | ipynb | Jupyter Notebook | Chained Comparison Operators.ipynb | Coslate/Python_Exercise | 9c400aa99d65136398e6b2944bd3e7886953b090 | [
"MIT"
] | null | null | null | Chained Comparison Operators.ipynb | Coslate/Python_Exercise | 9c400aa99d65136398e6b2944bd3e7886953b090 | [
"MIT"
] | null | null | null | Chained Comparison Operators.ipynb | Coslate/Python_Exercise | 9c400aa99d65136398e6b2944bd3e7886953b090 | [
"MIT"
] | null | null | null | 15.287411 | 53 | 0.424953 | [
[
[
"1 < 2 < 3",
"_____no_output_____"
],
[
"1 < 2 and 2 < 3",
"_____no_output_____"
],
[
"1 < 2 and 2 < 3",
"_____no_output_____"
],
[
"1 < 2 or 3 < 1",
"_____no_output_____"
],
[
"1 < 2 < 3",
"_____no_output_____"
],
[
"1 < 2 > 0.1",
"_____no_output_____"
],
[
"1 > 0 < -1",
"_____no_output_____"
],
[
"1 < 3 > 2",
"_____no_output_____"
],
[
"1 == 2",
"_____no_output_____"
],
[
"2 < 3 ",
"_____no_output_____"
],
[
"1 == 2 or 2 < 3",
"_____no_output_____"
],
[
"1 == 2 or 2 > 3",
"_____no_output_____"
],
[
"1 == 1 or 100 == 1",
"_____no_output_____"
],
[
"(1 == 1) or (100 == 1)",
"_____no_output_____"
],
[
"not(1==1)",
"_____no_output_____"
],
[
"not(100==1)",
"_____no_output_____"
],
[
"with open(\"./gg.txt\", \"w\") as file_out :\n file_out.write(\"shit\\n\")",
"_____no_output_____"
],
[
"pwd",
"_____no_output_____"
]
]
] | [
"code"
] | [
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
cb00916bc030d50506cf7489b6eb7b7d867d972f | 26,220 | ipynb | Jupyter Notebook | batch-processing/how_to_timelapse/how_to_timelapse.ipynb | j-desloires/eo-learn-examples | fc534dc8c6cea79f25ce905ea5a3956146372106 | [
"MIT"
] | 6 | 2022-02-23T09:28:10.000Z | 2022-03-24T15:30:24.000Z | batch-processing/how_to_timelapse/how_to_timelapse.ipynb | j-desloires/eo-learn-examples | fc534dc8c6cea79f25ce905ea5a3956146372106 | [
"MIT"
] | 2 | 2022-02-03T10:52:00.000Z | 2022-03-19T22:25:09.000Z | batch-processing/how_to_timelapse/how_to_timelapse.ipynb | j-desloires/eo-learn-examples | fc534dc8c6cea79f25ce905ea5a3956146372106 | [
"MIT"
] | 1 | 2022-02-03T09:55:29.000Z | 2022-02-03T09:55:29.000Z | 34.68254 | 690 | 0.563577 | [
[
[
"\n# How to make the perfect time-lapse of the Earth\n\nThis tutorial shows a detail coverage of making time-lapse animations from satellite imagery like a pro.",
"_____no_output_____"
],
[
"<h1>Table of Contents<span class=\"tocSkip\"></span></h1>\n<div class=\"toc\"><ul class=\"toc-item\"><li><span><a href=\"#0.-Prerequisites\" data-toc-modified-id=\"0.-Prerequisites-1\">0. Prerequisites</a></span></li><li><span><a href=\"#1.-Removing-clouds\" data-toc-modified-id=\"1.-Removing-clouds-2\">1. Removing clouds</a></span></li><li><span><a href=\"#2.-Applying-co-registration\" data-toc-modified-id=\"2.-Applying-co-registration-3\">2. Applying co-registration</a></span></li><li><span><a href=\"#3.-Large-Area-Example\" data-toc-modified-id=\"3.-Large-Area-Example-4\">3. Large Area Example</a></span></li><li><span><a href=\"#4.-Split-Image\" data-toc-modified-id=\"4.-Split-Image-5\">4. Split Image</a></span></li></ul></div>\n\nNote: This notebook requires an installation of additional packages `ffmpeg-python` and `ipyleaflet`.",
"_____no_output_____"
]
],
[
[
"%load_ext autoreload\n%autoreload 2\n\nimport datetime as dt\nimport json\nimport os\nimport subprocess\nfrom concurrent.futures import ProcessPoolExecutor\nfrom datetime import date, datetime, time, timedelta\nfrom functools import partial\nfrom glob import glob\n\nimport ffmpeg\nimport geopandas as gpd\nimport imageio\nimport matplotlib.pyplot as plt\nimport numpy as np\nimport pandas as pd\nimport shapely\nfrom ipyleaflet import GeoJSON, Map, basemaps\nfrom shapely.geometry import Polygon\nfrom tqdm.auto import tqdm\n\nfrom eolearn.core import (EOExecutor, EOPatch, EOTask, FeatureType,\n LinearWorkflow, LoadTask, OverwritePermission,\n SaveTask, ZipFeatureTask)\nfrom eolearn.coregistration import ECCRegistration\nfrom eolearn.features import LinearInterpolation, SimpleFilterTask\nfrom eolearn.io import ExportToTiff, ImportFromTiff, SentinelHubInputTask\nfrom eolearn.mask import CloudMaskTask\n\nfrom sentinelhub import (CRS, BatchSplitter, BBox, BBoxSplitter,\n DataCollection, Geometry, MimeType, SentinelHubBatch,\n SentinelHubRequest, SHConfig, bbox_to_dimensions)\n",
"_____no_output_____"
]
],
[
[
"## 0. Prerequisites\n\nIn order to set everything up and make the credentials work, please check [this notebook](https://github.com/sentinel-hub/eo-learn/blob/master/examples/io/SentinelHubIO.ipynb).",
"_____no_output_____"
]
],
[
[
"class AnimateTask(EOTask):\n def __init__(self, image_dir, out_dir, out_name, feature=(FeatureType.DATA, 'RGB'), scale_factor=2.5, duration=3, dpi=150, pad_inches=None, shape=None):\n self.image_dir = image_dir\n self.out_name = out_name\n self.out_dir = out_dir\n self.feature = feature\n self.scale_factor = scale_factor\n self.duration = duration\n self.dpi = dpi\n self.pad_inches = pad_inches\n self.shape = shape\n \n def execute(self, eopatch):\n images = np.clip(eopatch[self.feature]*self.scale_factor, 0, 1)\n fps = len(images)/self.duration\n subprocess.run(f'rm -rf {self.image_dir} && mkdir {self.image_dir}', shell=True)\n \n for idx, image in enumerate(images):\n if self.shape:\n fig = plt.figure(figsize=(self.shape[0], self.shape[1]))\n plt.imshow(image)\n plt.axis(False)\n plt.savefig(f'{self.image_dir}/image_{idx:03d}.png', bbox_inches='tight', dpi=self.dpi, pad_inches = self.pad_inches)\n plt.close()\n \n # video related\n stream = ffmpeg.input(f'{self.image_dir}/image_*.png', pattern_type='glob', framerate=fps)\n stream = stream.filter('pad', w='ceil(iw/2)*2', h='ceil(ih/2)*2', color='white')\n split = stream.split()\n video = split[0]\n \n # gif related\n palette = split[1].filter('palettegen', reserve_transparent=True, stats_mode='diff')\n gif = ffmpeg.filter([split[2], palette], 'paletteuse', dither='bayer', bayer_scale=5, diff_mode='rectangle')\n \n # save output\n os.makedirs(self.out_dir, exist_ok=True)\n video.output(f'{self.out_dir}/{self.out_name}.mp4', crf=15, pix_fmt='yuv420p', vcodec='libx264', an=None).run(overwrite_output=True)\n gif.output(f'{self.out_dir}/{self.out_name}.gif').run(overwrite_output=True)\n return eopatch",
"_____no_output_____"
]
],
[
[
"## 1. Removing clouds",
"_____no_output_____"
]
],
[
[
"# https://twitter.com/Valtzen/status/1270269337061019648\nbbox = BBox(bbox=[-73.558102,45.447728,-73.488750,45.491908], crs=CRS.WGS84)\nresolution = 10\ntime_interval = ('2018-01-01', '2020-01-01')\nprint(f'Image size: {bbox_to_dimensions(bbox, resolution)}')\n\ngeom, crs = bbox.geometry, bbox.crs\nwgs84_geometry = Geometry(geom, crs).transform(CRS.WGS84)\ngeometry_center = wgs84_geometry.geometry.centroid\n\nmap1 = Map(\n basemap=basemaps.Esri.WorldImagery,\n center=(geometry_center.y, geometry_center.x),\n zoom=13\n)\n\narea_geojson = GeoJSON(data=wgs84_geometry.geojson)\nmap1.add_layer(area_geojson)\n\nmap1",
"_____no_output_____"
],
[
"download_task = SentinelHubInputTask(\n bands = ['B04', 'B03', 'B02'],\n bands_feature = (FeatureType.DATA, 'RGB'),\n resolution=resolution,\n maxcc=0.9,\n time_difference=timedelta(minutes=120),\n data_collection=DataCollection.SENTINEL2_L2A,\n max_threads=10,\n mosaicking_order='leastCC',\n additional_data=[\n (FeatureType.MASK, 'CLM'),\n (FeatureType.MASK, 'dataMask')\n ]\n)\n\ndef valid_coverage_thresholder_f(valid_mask, more_than=0.95):\n coverage = np.count_nonzero(valid_mask)/np.prod(valid_mask.shape)\n return coverage > more_than\n\nvalid_mask_task = ZipFeatureTask({FeatureType.MASK: ['CLM', 'dataMask']}, (FeatureType.MASK, 'VALID_DATA'),\n lambda clm, dm: np.all([clm == 0, dm], axis=0))\n\nfilter_task = SimpleFilterTask((FeatureType.MASK, 'VALID_DATA'), valid_coverage_thresholder_f)\n\nname = 'clm_service'\nanim_task = AnimateTask(image_dir = './images', out_dir = './animations', out_name=name, duration=5, dpi=200)\n\nparams = {'MaxIters': 500}\ncoreg_task = ECCRegistration((FeatureType.DATA, 'RGB'), channel=2, params=params)\n\nname = 'clm_service_coreg'\nanim_task_after = AnimateTask(image_dir='./images', out_dir='./animations', out_name=name, duration=5, dpi=200)",
"_____no_output_____"
],
[
"workflow = LinearWorkflow(\n download_task,\n valid_mask_task,\n filter_task,\n anim_task,\n coreg_task,\n anim_task_after\n)\n\nresult = workflow.execute({\n download_task: {'bbox': bbox, 'time_interval': time_interval}\n})",
"_____no_output_____"
]
],
[
[
"## 2. Applying co-registration",
"_____no_output_____"
]
],
[
[
"bbox = BBox(bbox=[34.716, 30.950, 34.743, 30.975], crs=CRS.WGS84)\nresolution = 10\ntime_interval = ('2020-01-01', '2021-01-01')\nprint(f'BBox size: {bbox_to_dimensions(bbox, resolution)}')\n\ngeom, crs = bbox.geometry, bbox.crs\nwgs84_geometry = Geometry(geom, crs).transform(CRS.WGS84)\ngeometry_center = wgs84_geometry.geometry.centroid\n\nmap1 = Map(\n basemap=basemaps.Esri.WorldImagery,\n center=(geometry_center.y, geometry_center.x),\n zoom=14\n)\n\narea_geojson = GeoJSON(data=wgs84_geometry.geojson)\nmap1.add_layer(area_geojson)\n\nmap1",
"_____no_output_____"
],
[
"download_task_l2a = SentinelHubInputTask(\n bands = ['B04', 'B03', 'B02'],\n bands_feature = (FeatureType.DATA, 'RGB'),\n resolution=resolution,\n maxcc=0.9,\n time_difference=timedelta(minutes=120),\n data_collection=DataCollection.SENTINEL2_L2A,\n max_threads=10,\n additional_data=[\n (FeatureType.MASK, 'dataMask', 'dataMask_l2a')\n ]\n)\n\ndownload_task_l1c = SentinelHubInputTask(\n bands_feature = (FeatureType.DATA, 'BANDS'),\n resolution=resolution,\n maxcc=0.9,\n time_difference=timedelta(minutes=120),\n data_collection=DataCollection.SENTINEL2_L1C,\n max_threads=10,\n additional_data=[\n (FeatureType.MASK, 'dataMask', 'dataMask_l1c')\n ]\n)\n\ndata_mask_merge = ZipFeatureTask({FeatureType.MASK: ['dataMask_l1c', 'dataMask_l2a']}, (FeatureType.MASK, 'dataMask'),\n lambda dm1, dm2: np.all([dm1, dm2], axis=0))\n\ncloud_masking_task = CloudMaskTask(\n data_feature=(FeatureType.DATA, 'BANDS'),\n is_data_feature='dataMask',\n all_bands=True,\n processing_resolution=120,\n mono_features=None,\n mask_feature='CLM',\n average_over=16,\n dilation_size=12,\n mono_threshold=0.2\n)\n\nvalid_mask_task = ZipFeatureTask({FeatureType.MASK: ['CLM', 'dataMask']}, (FeatureType.MASK, 'VALID_DATA'),\n lambda clm, dm: np.all([clm == 0, dm], axis=0))\n\nfilter_task = SimpleFilterTask((FeatureType.MASK, 'VALID_DATA'), valid_coverage_thresholder_f)\n\nname = 'wo_coreg_anim'\nanim_task_before = AnimateTask(image_dir='./images', out_dir='./animations', out_name=name, duration=5, dpi=200)\n\n\nparams = {'MaxIters': 500}\ncoreg_task = ECCRegistration((FeatureType.DATA, 'RGB'), channel=2, params=params)\n\nname = 'coreg_anim'\nanim_task_after = AnimateTask(image_dir='./images', out_dir='./animations', out_name=name, duration=5, dpi=200)",
"_____no_output_____"
],
[
"workflow = LinearWorkflow(\n download_task_l2a,\n download_task_l1c,\n data_mask_merge,\n cloud_masking_task,\n valid_mask_task,\n filter_task,\n anim_task_before,\n coreg_task,\n anim_task_after\n)\n\nresult = workflow.execute({\n download_task_l2a: {'bbox': bbox, 'time_interval': time_interval}\n})",
"_____no_output_____"
]
],
[
[
"## 3. Large Area Example",
"_____no_output_____"
]
],
[
[
"bbox = BBox(bbox=[21.4,-20.0,23.9,-18.0], crs=CRS.WGS84)\ntime_interval = ('2017-09-01', '2019-04-01')\n# time_interval = ('2017-09-01', '2017-10-01')\nresolution = 640\nprint(f'BBox size: {bbox_to_dimensions(bbox, resolution)}')\n\ngeom, crs = bbox.geometry, bbox.crs\nwgs84_geometry = Geometry(geom, crs).transform(CRS.WGS84)\ngeometry_center = wgs84_geometry.geometry.centroid\n\nmap1 = Map(\n basemap=basemaps.Esri.WorldImagery,\n center=(geometry_center.y, geometry_center.x),\n zoom=8\n)\n\narea_geojson = GeoJSON(data=wgs84_geometry.geojson)\nmap1.add_layer(area_geojson)\n\nmap1",
"_____no_output_____"
],
[
"download_task_l2a = SentinelHubInputTask(\n bands = ['B04', 'B03', 'B02'],\n bands_feature = (FeatureType.DATA, 'RGB'),\n resolution=resolution,\n maxcc=0.9,\n time_difference=timedelta(minutes=120),\n data_collection=DataCollection.SENTINEL2_L2A,\n max_threads=10,\n additional_data=[\n (FeatureType.MASK, 'dataMask', 'dataMask_l2a')\n ],\n aux_request_args={'dataFilter': {'previewMode': 'PREVIEW'}}\n)\n\ndownload_task_l1c = SentinelHubInputTask(\n bands_feature = (FeatureType.DATA, 'BANDS'),\n resolution=resolution,\n maxcc=0.9,\n time_difference=timedelta(minutes=120),\n data_collection=DataCollection.SENTINEL2_L1C,\n max_threads=10,\n additional_data=[\n (FeatureType.MASK, 'dataMask', 'dataMask_l1c')\n ],\n aux_request_args={'dataFilter': {'previewMode': 'PREVIEW'}}\n)\n\ndata_mask_merge = ZipFeatureTask({FeatureType.MASK: ['dataMask_l1c', 'dataMask_l2a']}, (FeatureType.MASK, 'dataMask'),\n lambda dm1, dm2: np.all([dm1, dm2], axis=0))\n\ncloud_masking_task = CloudMaskTask(\n data_feature='BANDS',\n is_data_feature='dataMask',\n all_bands=True,\n processing_resolution=resolution,\n mono_features=('CLP', 'CLM'),\n mask_feature=None,\n mono_threshold=0.3,\n average_over=1,\n dilation_size=4\n)\n\nvalid_mask_task = ZipFeatureTask({FeatureType.MASK: ['CLM', 'dataMask']}, (FeatureType.MASK, 'VALID_DATA'),\n lambda clm, dm: np.all([clm == 0, dm], axis=0))\n\nresampled_range = ('2018-01-01', '2019-01-01', 10)\ninterp_task = LinearInterpolation(\n feature=(FeatureType.DATA, 'RGB'),\n mask_feature=(FeatureType.MASK, 'VALID_DATA'),\n resample_range=resampled_range,\n bounds_error=False\n)\n\nname = 'botswana_single_raw'\nanim_task_raw = AnimateTask(image_dir='./images', out_dir='./animations', out_name=name, duration=5, dpi=200)\n\nname = 'botswana_single'\nanim_task = AnimateTask(image_dir='./images', out_dir='./animations', out_name=name, duration=3, dpi=200)",
"_____no_output_____"
],
[
"workflow = LinearWorkflow(\n download_task_l2a,\n# anim_task_raw\n download_task_l1c,\n data_mask_merge,\n cloud_masking_task,\n valid_mask_task,\n interp_task,\n anim_task\n)\n\nresult = workflow.execute({\n download_task_l2a:{'bbox': bbox, 'time_interval': time_interval},\n})",
"_____no_output_____"
]
],
[
[
"## 4. Split Image",
"_____no_output_____"
]
],
[
[
"bbox = BBox(bbox=[21.3,-20.0,24.0,-18.0], crs=CRS.WGS84)\ntime_interval = ('2018-09-01', '2020-04-01')\nresolution = 120\n\nbbox_splitter = BBoxSplitter([bbox.geometry], bbox.crs, (6,5))\nbbox_list = np.array(bbox_splitter.get_bbox_list())\ninfo_list = np.array(bbox_splitter.get_info_list())\nprint(f'{len(bbox_list)} patches of size: {bbox_to_dimensions(bbox_list[0], resolution)}')\n\ngdf = gpd.GeoDataFrame(None, crs=int(bbox.crs.epsg), geometry=[bbox.geometry for bbox in bbox_list])\n\ngeom, crs = gdf.unary_union, CRS.WGS84\nwgs84_geometry = Geometry(geom, crs).transform(CRS.WGS84)\ngeometry_center = wgs84_geometry.geometry.centroid\n\nmap1 = Map(\n basemap=basemaps.Esri.WorldImagery,\n center=(geometry_center.y, geometry_center.x),\n zoom=8\n)\n\nfor geo in gdf.geometry:\n area_geojson = GeoJSON(data=Geometry(geo, crs).geojson)\n map1.add_layer(area_geojson)\n\nmap1",
"_____no_output_____"
],
[
"download_task = SentinelHubInputTask(\n bands = ['B04', 'B03', 'B02'],\n bands_feature = (FeatureType.DATA, 'RGB'),\n resolution=resolution,\n maxcc=0.9,\n time_difference=timedelta(minutes=120),\n data_collection=DataCollection.SENTINEL2_L2A,\n max_threads=10,\n additional_data=[\n (FeatureType.MASK, 'CLM'),\n (FeatureType.DATA, 'CLP'),\n (FeatureType.MASK, 'dataMask')\n ]\n)\n\nvalid_mask_task = ZipFeatureTask([(FeatureType.MASK, 'dataMask'), (FeatureType.MASK, 'CLM'), (FeatureType.DATA, 'CLP')], (FeatureType.MASK, 'VALID_DATA'),\n lambda dm, clm, clp: np.all([dm, clm == 0, clp/255 < 0.3], axis=0))\n\nresampled_range = ('2019-01-01', '2020-01-01', 10)\ninterp_task = LinearInterpolation(\n feature=(FeatureType.DATA, 'RGB'),\n mask_feature=(FeatureType.MASK, 'VALID_DATA'),\n resample_range=resampled_range,\n bounds_error=False\n)\n\nexport_r = ExportToTiff(feature=(FeatureType.DATA, 'RGB'), folder='./tiffs/', band_indices=[0])\nexport_g = ExportToTiff(feature=(FeatureType.DATA, 'RGB'), folder='./tiffs/', band_indices=[1])\nexport_b = ExportToTiff(feature=(FeatureType.DATA, 'RGB'), folder='./tiffs/', band_indices=[2])\n\nconvert_to_uint16 = ZipFeatureTask([(FeatureType.DATA, 'RGB')], (FeatureType.DATA, 'RGB'),\n lambda x: (x*1e4).astype(np.uint16))",
"_____no_output_____"
],
[
"os.system('rm -rf ./tiffs && mkdir ./tiffs')\n\nworkflow = LinearWorkflow(\n download_task,\n valid_mask_task,\n interp_task,\n convert_to_uint16,\n export_r,\n export_g,\n export_b\n)\n\n# Execute the workflow\nexecution_args = []\nfor idx, bbox in enumerate(bbox_list):\n execution_args.append({\n download_task: {'bbox': bbox, 'time_interval': time_interval},\n export_r: {'filename': f'r_patch_{idx}.tiff'},\n export_g: {'filename': f'g_patch_{idx}.tiff'},\n export_b: {'filename': f'b_patch_{idx}.tiff'}\n })\n \nexecutor = EOExecutor(workflow, execution_args, save_logs=True)\nexecutor.run(workers=10, multiprocess=False)\nexecutor.make_report()",
"_____no_output_____"
],
[
"# spatial merge\nsubprocess.run(f'gdal_merge.py -n 0 -a_nodata 0 -o tiffs/r.tiff -co compress=LZW tiffs/r_patch_*.tiff && rm -rf tiffs/r_patch_*.tiff', shell=True);\nsubprocess.run(f'gdal_merge.py -n 0 -a_nodata 0 -o tiffs/g.tiff -co compress=LZW tiffs/g_patch_*.tiff && rm -rf tiffs/g_patch_*.tiff', shell=True);\nsubprocess.run(f'gdal_merge.py -n 0 -a_nodata 0 -o tiffs/b.tiff -co compress=LZW tiffs/b_patch_*.tiff && rm -rf tiffs/b_patch_*.tiff', shell=True);",
"_____no_output_____"
],
[
"dates = pd.date_range('2019-01-01', '2020-01-01', freq='10D').to_pydatetime()\nimport_r = ImportFromTiff((FeatureType.DATA, 'R'), f'tiffs/r.tiff', timestamp_size=len(dates))\nimport_g = ImportFromTiff((FeatureType.DATA, 'G'), f'tiffs/g.tiff', timestamp_size=len(dates))\nimport_b = ImportFromTiff((FeatureType.DATA, 'B'), f'tiffs/b.tiff', timestamp_size=len(dates))\n\nmerge_bands_task = ZipFeatureTask({FeatureType.DATA: ['R', 'G', 'B']}, (FeatureType.DATA, 'RGB'),\n lambda r, g, b: np.moveaxis(np.array([r[...,0], g[...,0], b[...,0]]), 0, -1))\n \ndef temporal_ma_f(f):\n k = np.array([0.05, 0.6, 1, 0.6, 0.05])\n k = k/np.sum(k)\n w = len(k)//2\n return np.array([np.sum([f[(i-w+j)%len(f)]*k[j] for j in range(len(k))], axis=0) for i in range(len(f))])\n\ntemporal_smoothing = ZipFeatureTask([(FeatureType.DATA, 'RGB')], (FeatureType.DATA, 'RGB'), temporal_ma_f)\n\nname = 'botswana_multi_ma'\nanim_task = AnimateTask(image_dir='./images', out_dir='./animations', out_name=name, duration=3,\n dpi=400, scale_factor=3.0/1e4)",
"_____no_output_____"
],
[
"workflow = LinearWorkflow(\n import_r,\n import_g,\n import_b,\n merge_bands_task,\n temporal_smoothing,\n anim_task\n)\n\nresult = workflow.execute()",
"_____no_output_____"
]
],
[
[
"## 5. Batch request",
"_____no_output_____"
],
[
"Use the evalscript from the [custom scripts repository](https://github.com/sentinel-hub/custom-scripts/tree/master/sentinel-2/interpolated_time_series) and see how to use it in the batch example in our [sentinelhub-py](https://github.com/sentinel-hub/sentinelhub-py/blob/master/examples/batch_processing.ipynb) library.",
"_____no_output_____"
]
]
] | [
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown"
] | [
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown",
"markdown"
]
] |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.